Impact of Gut Dysbiosis on Neurohormonal Pathways in Chronic Kidney Disease
Abstract
1. Introduction
2. Gut Microbiota and Symbiotic Benefits
3. Mechanisms of Gut Dysbiosis in CKD
3.1. Alterations in the GI Tract Biochemical Environment
3.2. Diet
3.3. Medications
4. Disruption of the Intestinal Epithelial Barrier
4.1. Urea Toxicity
4.2. Hemodialysis-Associated Disruption of the Intestinal Barrier
4.3. Gut Wall Inflammation and Oxidative Stress
5. Dysbiosis as a Major Source of Uremic Toxins in CKD
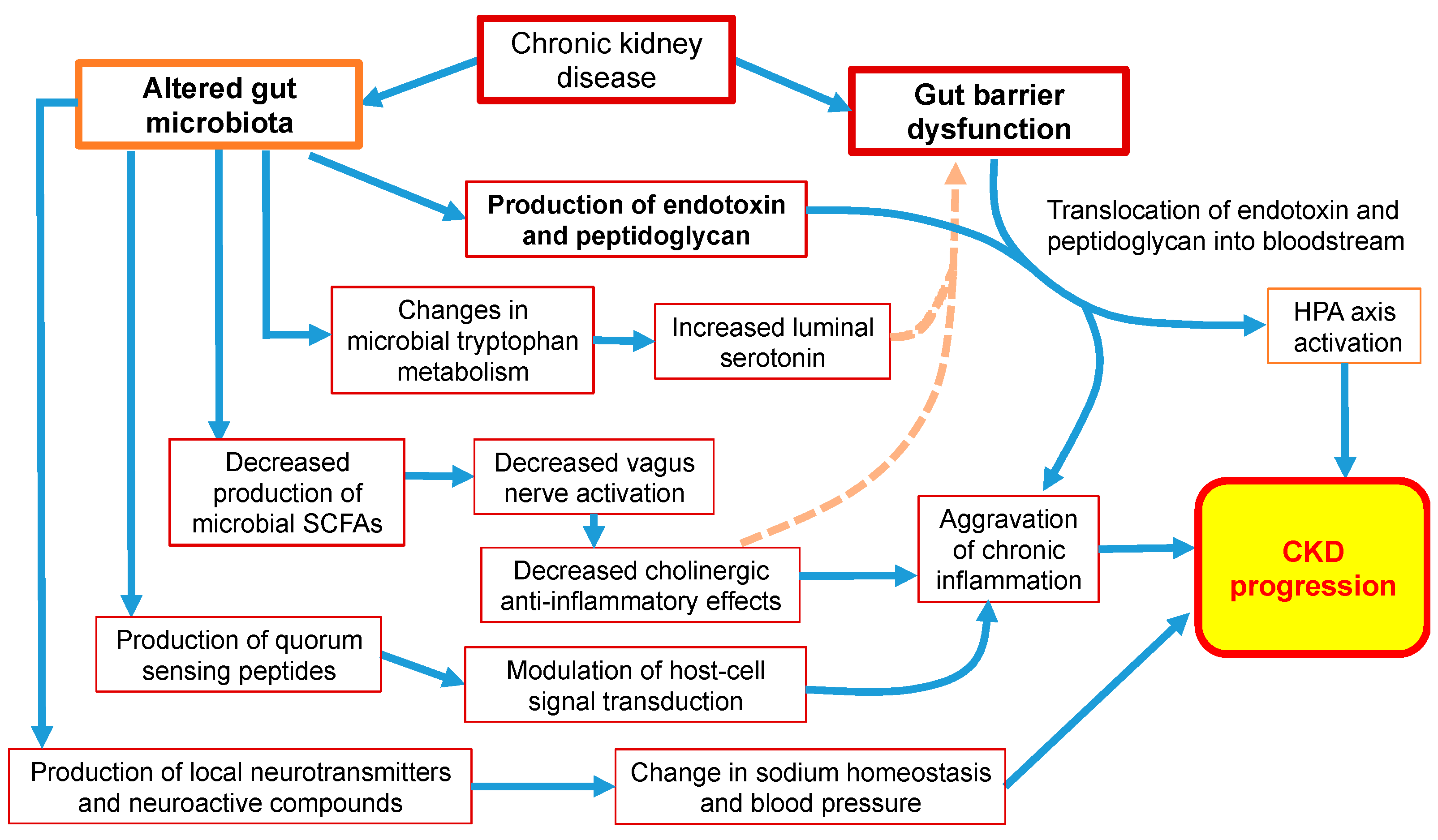
6. The Effect of Dysbiosis on Neuroendocrine Pathways in CKD Patients
6.1. Hypothalamic-Pituitary-Adrenal (HPA) Axis
6.2. Induction of Release of Gut Hormones
6.3. Production of Neurotransmitters and Neuroactive Compounds
6.4. Tryptophan Metabolism
6.5. Bacterial Hormone-Like Compounds
6.6. Cholinergic Anti-Inflammatory Pathway
7. Strategies to Attenuate Gut Dysbiosis in CKD
7.1. Balanced Diet
7.2. Prebiotics
7.3. Intestinal Alpha-Glycosidase Inhibition
7.4. Probiotics
7.5. Synbiotics
8. Summary
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Rafieian-Kopaei, M.; Beigrezaei, S.; Nasri, H.; Kafeshani, M. Soy protein and chronic kidney disease: An updated review. Int. J. Prev. Med. 2017, 8, 105. [Google Scholar] [PubMed]
- Tang, E.; Bansal, A.; Novak, M.; Mucsi, I. Patient-reported outcomes in patients with chronic kidney disease and kidney transplant-Part 1. Front. Med. 2017, 4, 254. [Google Scholar] [CrossRef] [PubMed]
- Hill, N.R.; Fatoba, S.T.; Oke, J.L.; Hirst, J.A.; O’Callaghan, C.A.; Lasserson, D.S.; Richard Hobbs, F.D. Global prevalence of chronic kidney disease—A systematic review and meta-analysis. PLoS ONE 2016, 11, e0158765. [Google Scholar] [CrossRef] [PubMed]
- Stevens, P.E.; Levin, A. Kidney disease: Improving Global Outcomes Chronic Kidney Disease Guideline Development Work Group, M. Evaluation and management of chronic kidney disease: Synopsis of the kidney disease: Improving global outcomes 2012 clinical practice guideline. Ann. Intern. Med. 2013, 158, 825–830. [Google Scholar] [CrossRef] [PubMed]
- Xia, J.; Wang, L.; Ma, Z.; Zhong, L.; Wang, Y.; Gao, Y.; He, L.; Su, X. Cigarette smoking and chronic kidney disease in the general population: A systematic review and meta-analysis of prospective cohort studies. Nephrol. Dial. Transplant. 2017, 32, 475–487. [Google Scholar] [CrossRef] [PubMed]
- Romagnani, P.; Remuzzi, G.; Glassock, R.; Levin, A.; Jager, K.J.; Tonelli, M.; Massy, Z.; Wanner, C.; Anders, H.J. Chronic kidney disease. Nat. Rev. Dis. Primers 2017, 3, 17088. [Google Scholar] [CrossRef] [PubMed]
- Glassock, R.J.; Denic, A.; Rule, A.D. The conundrums of chronic kidney disease and aging. J. Nephrol. 2017, 30, 477–483. [Google Scholar] [CrossRef]
- Westland, R.; Schreuder, M.F.; van Goudoever, J.B.; Sanna-Cherchi, S.; van Wijk, J.A. Clinical implications of the solitary functioning kidney. Clin. J. Am. Soc. Nephrol. 2014, 9, 978–986. [Google Scholar] [CrossRef]
- Vos, T.; Barber, R.M.; Bell, B.; Bertozzi-Villa, A.; Biryukov, S.; Bolliger, I.; Charlson, F.; Davis, A.; Degenhardt, L.; Dicker, D.; et al. Global, regional, and national incidence, prevalence, and years lived with disability for 301 acute and chronic diseases and injuries in 188 countries, 1990–2013: A systematic analysis for the Global Burden of Disease Study 2013. Lancet 2015, 386, 743–800. [Google Scholar] [CrossRef]
- Thomas, E.A.; Pawar, B.; Thomas, A. A prospective study of cutaneous abnormalities in patients with chronic kidney disease. Indian J. Nephrol. 2012, 22, 116–120. [Google Scholar] [CrossRef]
- Kaplan, J.M.; Sharma, N.; Dikdan, S. Hypoxia-Inducible Factor and Its Role in the Management of Anemia in Chronic Kidney Disease. Int. J. Mol. Sci. 2018, 19, 389. [Google Scholar] [CrossRef] [PubMed]
- Vaziri, N.D.; Yuan, J.; Rahimi, A.; Ni, Z.; Said, H. Subramanian VS. Disintegration of colonic epithelial tight junction in uremia: A likely cause of CKD-associated inflammation. Nephrol. Dial. Transplant. 2012, 27, 2686–2693. [Google Scholar] [CrossRef] [PubMed]
- Vaziri, N.D. CKD impairs barrier function and alters microbial flora of the intestine: A major link to inflammation and uremic toxicity. Curr. Opin. Nephrol. Hypertens. 2012, 21, 587–592. [Google Scholar] [CrossRef] [PubMed]
- Nigam, G.; Camacho, M.; Chang, E.T.; Riaz, M. Exploring sleep disorders in patients with chronic kidney disease. Nat. Sci. Sleep 2018, 10, 35–43. [Google Scholar] [CrossRef]
- Malluche, H.H.; Porter, D.S.; Pienkowski, D. Evaluating bone quality in patients with chronic kidney disease. Nat. Rev. Nephrol. 2013, 9, 671–680. [Google Scholar] [CrossRef] [PubMed]
- Ketteler, M.; Wanner, C. Chronic kidney disease—Update 2018. Dtsch. Med. Wochenschr. 2018, 143, 169–173. [Google Scholar] [PubMed]
- Gilligan, S.; Raphael, K.L. Hyperkalemia and hypokalemia in CKD: Prevalence, risk factors, and clinical outcomes. Adv. Chronic Kidney Dis. 2017, 24, 315–318. [Google Scholar] [CrossRef] [PubMed]
- Langston, C. Managing fluid and electrolyte disorders in kidney disease. Vet. Clin. North. Am. Small Anim. Pract. 2017, 47, 471–490. [Google Scholar] [CrossRef] [PubMed]
- Vaziri, N.D. Gut microbial translocation in the pathogenesis of systemic inflammation in patients with end-stage renal disease. Dig. Dis. Sci. 2014, 59, 2020–2022. [Google Scholar] [CrossRef] [PubMed]
- Slee, A.D. Exploring metabolic dysfunction in chronic kidney disease. Nutr. Metab. (Lond) 2012, 9, 36. [Google Scholar] [CrossRef]
- Siezen, R.J.; Kleerebezem, M. The human gut microbiome: Are we our enterotypes? Microb. Biotechnol. 2011, 4, 550–553. [Google Scholar] [CrossRef] [PubMed]
- Malnick, S.; Melzer, E. Human microbiome: From the bathroom to the bedside. World J. Gastrointest. Pathophysiol. 2015, 6, 79–85. [Google Scholar] [CrossRef] [PubMed]
- Vaziri, N.D.; Wong, J.; Pahl, M.; Piceno, YM.; Yuan, J.; Desantis, T.Z.; Ni, Z.; Nguyen, T.H.; Andersen, G.L. Chronic kidney disease alters intestinal microbial flora. Kidney Int. 2013, 83, 308–315. [Google Scholar] [CrossRef] [PubMed]
- Gerritsen, J.; Smidt, H.; Rijkers, G.T.; de Vos, W.M. Intestinal microbiota in human health and disease: The impact of probiotics. Genes Nutr. 2011, 6, 209–240. [Google Scholar] [CrossRef] [PubMed]
- Selber-Hnatiw, S.; Rukundo, B.; Ahmadi, M.; Akoubi, H.; Al-Bizri, H.; Aliu, A.F.; Ambeaghen, T.U.; Avetisyan, L.; Bahar, I.; Baird, A.; et al. Human gut microbiota: Toward an ecology of disease. Front. Microbiol. 2017, 8, 1265. [Google Scholar] [CrossRef] [PubMed]
- Shin, N.R.; Whon, T.W.; Bae, J.W. Proteobacteria: Microbial signature of dysbiosis in gut microbiota. Trends Biotechnol. 2015, 33, 496–503. [Google Scholar] [CrossRef]
- Qin, J.; Li, R.; Raes, J.; Arumugam, M.; Burgdorf, K.S.; Manichanh, C.; Nielsen, T.; Pons, N.; Levenez, F.; Yamada, T.; et al. A human gut microbial gene catalogue established by metagenomic sequencing. Nature. 2010, 464, 59–65. [Google Scholar] [CrossRef]
- Vaziri, N.D.; Zhao, Y.Y.; Pahl, M.V. Altered intestinal microbial flora and impaired epithelial barrier structure and function in CKD: The nature, mechanisms, consequences and potential treatment. Nephrol. Dial. Transplant. 2016, 31, 737–746. [Google Scholar] [CrossRef]
- Turroni, F.; Ribbera, A.; Foroni, E.; van Sinderen, D.; Ventura, M. Human gut microbiota and bifidobacteria: From composition to functionality. Antonie Van Leeuwenhoek 2008, 94, 35–50. [Google Scholar] [CrossRef]
- Zoetendal, E.G.; Vaughan, E.E.; de Vos, W.M. A microbial world within us. Mol. Microbiol. 2006, 59, 1639–1650. [Google Scholar] [CrossRef]
- Backhed, F.; Ley, R.E.; Sonnenburg, J.L.; Peterson, D.A.; Gordon, J.I. Host-bacterial mutualism in the human intestine. Science 2005, 307, 1915–1920. [Google Scholar] [CrossRef] [PubMed]
- Capurso, G.; Lahner, E. The interaction between smoking, alcohol and the gut microbiome. Best Pract. Res. Clin. Gastroenterol. 2017, 31, 579–588. [Google Scholar] [CrossRef] [PubMed]
- Wypych, T.P.; Marsland, B.J. Diet hypotheses in light of the microbiota revolution: New perspectives. Nutrients 2017, 9, 537. [Google Scholar] [CrossRef] [PubMed]
- Ursell, L.K.; Clemente, J.C.; Rideout, J.R.; Gevers, D.; Caporaso, J.G.; Knight, R. The interpersonal and intrapersonal diversity of human-associated microbiota in key body sites. J. Allergy Clin. Immunol. 2012, 129, 1204–1208. [Google Scholar] [CrossRef] [PubMed]
- Wong, J.; Piceno, Y.M.; Desantis, T.Z.; Pahl, M.; Andersen, G.L.; Vaziri, N.D. Expansion of urease- and uricase-containing, indole- and p-cresol-forming and contraction of short-chain fatty acid-producing intestinal microbiota in ESRD. Am. J. Nephrol. 2014, 39, 230–237. [Google Scholar] [CrossRef] [PubMed]
- Rossi, M.; Amaretti, A.; Raimondi, S. Folate production by probiotic bacteria. Nutrients 2011, 3, 118–134. [Google Scholar] [CrossRef] [PubMed]
- Carbonero, F.; Benefiel, A.C.; Alizadeh-Ghamsari, A.H.; Gaskins, H.R. Microbial pathways in colonic sulfur metabolism and links with health and disease. Front. Physiol. 2012, 3, 448. [Google Scholar] [CrossRef] [PubMed]
- Pahl, M.V.; Vaziri, N.D. The Chronic kidney disease—Colonic axis. Semin. Dial. 2015, 28, 459–463. [Google Scholar] [CrossRef] [PubMed]
- den Besten, G.; van Eunen, K.; Groen, A.K.; Venema, K.; Reijngoud, D.J.; Bakker, B.M. The role of short-chain fatty acids in the interplay between diet, gut microbiota, and host energy metabolism. J. Lipid Res. 2013, 54, 2325–2340. [Google Scholar] [CrossRef]
- Miyamoto, J.; Kasubuchi, M.; Nakajima, A.; Irie, J.; Itoh, H.; Kimura, I. The role of short-chain fatty acid on blood pressure regulation. Curr. Opin. Nephrol. Hypertens. 2016, 25, 379–383. [Google Scholar] [CrossRef] [PubMed]
- Cani, P.D.; Knauf, C. How gut microbes talk to organs: The role of endocrine and nervous routes. Mol. Metab. 2016, 5, 743–752. [Google Scholar] [CrossRef] [PubMed]
- Pluznick, J. A novel SCFA receptor, the microbiota, and blood pressure regulation. Gut Microbes 2014, 5, 202–207. [Google Scholar] [CrossRef] [PubMed]
- Mell, B.; Jala, V.R.; Mathew, A.V.; Byun, J.; Waghulde, H.; Zhang, Y.; Haribabu, B.; Vijay-Kumar, M.; Pennathur, S.; Joe, B. Evidence for a link between gut microbiota and hypertension in the Dahl rat. Physiol. Genomics 2015, 47, 187–197. [Google Scholar] [CrossRef] [PubMed]
- Vaziri, N.D.; Liu, S.M.; Lau, W.L.; Khazaeli, M.; Nazertehrani, S.; Farzaneh, SH.; Kieffer, D.A.; Adams, S.H.; Martin, R.J. High amylose resistant starch diet ameliorates oxidative stress, inflammation, and progression of chronic kidney disease. PLoS ONE 2014, 9, e114881. [Google Scholar] [CrossRef] [PubMed]
- Lau, W.L.; Vaziri, N.D. The Leaky Gut and Altered Microbiome in Chronic Kidney Disease. J. Ren. Nutr. 2017, 27, 458–461. [Google Scholar] [CrossRef]
- Thomas, S.; Izard, J.; Walsh, E.; Batich, K.; Chongsathidkiet, P.; Clarke, G.; Sela, D.A.; Muller, A.J.; Mullin, J.M.; Albert, K.; et al. The host microbiome regulates and maintains human health: A primer and perspective for non-microbiologists. Cancer Res. 2017, 77, 1783–1812. [Google Scholar] [CrossRef] [PubMed]
- Kim, C.H.; Park, J.; Kim, M. Gut microbiota-derived short-chain Fatty acids, T cells, and inflammation. Immun. Netw. 2014, 14, 277–288. [Google Scholar] [CrossRef] [PubMed]
- Wei, B.; Wingender, G.; Fujiwara, D.; Chen, D.Y.; McPherson, M.; Brewer, S.; Borneman, J.; Kronenberg, M.; Braun, J. Commensal microbiota and CD8+ T cells shape the formation of invariant NKT cells. J. Immunol. 2010, 184, 1218–1226. [Google Scholar] [CrossRef]
- Kang, Y.; Cai, Y. Gut microbiota and obesity: Implications for fecal microbiota transplantation therapy. Hormones 2017, 16, 223–234. [Google Scholar] [CrossRef] [PubMed]
- Weiss, G.A.; Hennet, T. Mechanisms and consequences of intestinal dysbiosis. Cell Mol. Life Sci. 2017, 74, 2959–2977. [Google Scholar] [CrossRef]
- Saltzman, E.T.; Palacios, T.; Thomsen, M.; Vitetta, L. Intestinal microbiome shifts, dysbiosis, inflammation, and non-alcoholic fatty liver disease. Front. Microbiol. 2018, 9, 61. [Google Scholar] [CrossRef] [PubMed]
- Minato, T.; Maeda, T.; Fujisawa, Y.; Tsuji, H.; Nomoto, K.; Ohno, K.; Hirayama, M. Progression of Parkinson’s disease is associated with gut dysbiosis: Two-year follow-up study. PLoS ONE 2017, 12, e0187307. [Google Scholar] [CrossRef] [PubMed]
- Moustafa, A.; Li, W.; Anderson, E.L.; Wong, E.H.M.; Dulai, P.S.; Sandborn, W.J.; Biggs, W.; Yooseph, S.; Jones, M.B.; Venter, J.C.; et al. Genetic risk, dysbiosis, and treatment stratification using host genome and gut microbiome in inflammatory bowel disease. Clin. Transl. Gastroenterol. 2018, 9, e132. [Google Scholar] [CrossRef] [PubMed]
- Carding, S.; Verbeke, K.; Vipond, D.T.; Corfe, B.M.; Owen, L.J. Dysbiosis of the gut microbiota in disease. Microb. Ecol. Health Dis. 2015, 26, 26191. [Google Scholar] [CrossRef] [PubMed]
- Raskov, H.; Burcharth, J.; Pommergaard, H.C. Linking gut microbiota to colorectal cancer. J. Cancer 2017, 8, 3378–3395. [Google Scholar] [CrossRef] [PubMed]
- Lopez, A.; Hansmannel, F.; Kokten, T.; Bronowicki, J.P.; Melhem, H.; Sokol, H.; Peyrin-Biroulet, L. Microbiota in digestive cancers: Our new partner? Carcinogenesis 2017, 38, 1157–1166. [Google Scholar] [CrossRef]
- Koliada, A.; Syzenko, G.; Moseiko, V.; Budovska, L.; Puchkov, K.; Perederiy, V.; Gavalko, Y.; Dorofeyev, A.; Romanenko, M.; Tkach, S.; et al. Association between body mass index and Firmicutes/Bacteroidetes ratio in an adult Ukrainian population. BMC Microbiol. 2017, 17, 120. [Google Scholar] [CrossRef]
- Adnan, S.; Nelson, J.W.; Ajami, N.J.; Venna, V.R.; Petrosino, J.F.; Bryan, R.M., Jr.; Durgan, D.J. Alterations in the gut microbiota can elicit hypertension in rats. Physiol. Genomics 2017, 49, 96–104. [Google Scholar] [CrossRef]
- Strati, F.; Cavalieri, D.; Albanese, D.; De Felice, C.; Donati, C.; Hayek, J.; Jousson, O.; Leoncini, S.; Renzi, D.; Calabrò, A.; et al. New evidences on the altered gut microbiota in autism spectrum disorders. Microbiome 2017, 5, 24. [Google Scholar] [CrossRef]
- Chung, C.S.; Chang, P.F.; Liao, C.H.; Lee, T.H.; Chen, Y.; Lee, Y.C.; Wu, M.S.; Wang, H.P.; Ni, Y.H. Differences of microbiota in small bowel and faeces between irritable bowel syndrome patients and healthy subjects. Scand. J. Gastroenterol. 2016, 51, 410–419. [Google Scholar] [CrossRef]
- Vaziri, N.D.; Suematsu, Y.; Shimomura, A. Uremic toxins and gut microbiome. Nihon Jinzo Gakkai Shi 2017, 535–544. [Google Scholar]
- Jiang, S.; Xie, S.; Lv, D.; Wang, P.; He, H.; Zhang, T.; Zhou, Y.; Lin, Q.; Zhou, H.; Jiang, J.; et al. Alteration of the gut microbiota in Chinese population with chronic kidney disease. Sci. Rep. 2017, 7, 2870. [Google Scholar] [CrossRef] [PubMed]
- Cigarran Guldris, S.; Gonzalez Parra, E.; Cases Amenos, A. Gut microbiota in chronic kidney disease. Nefrologia 2017, 37, 9–19. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Li, J.; Yu, J.; Wang, Y.; Lu, J.; Shang, E.X.; Zhu, Z.; Guo, J.; Duan, J. Disorder of gut amino acids metabolism during CKD progression is related with gut microbiota dysbiosis and metagenome change. J. Pharm. Biomed. Anal. 2018, 149, 425–435. [Google Scholar] [CrossRef]
- Rossi, M.; Johnson, D.W.; Campbell, K.L. The kidney-gut axis: Implications for nutrition care. J. Ren. Nutr. 2015, 25, 399–403. [Google Scholar] [CrossRef]
- Ramezani, A.; Raj, D.S. The gut microbiome, kidney disease, and targeted interventions. J. Am. Soc. Nephrol. 2014, 25, 657–670. [Google Scholar] [CrossRef]
- Hida, M.; Aiba, Y.; Sawamura, S.; Suzuki, N.; Satoh, T.; Koga, Y. Inhibition of the accumulation of uremic toxins in the blood and their precursors in the feces after oral administration of Lebenin, a lactic acid bacteria preparation, to uremic patients undergoing hemodialysis. Nephron 1996, 74, 349–355. [Google Scholar] [CrossRef]
- De Angelis, M.; Montemurno, E.; Piccolo, M.; Vannini, L.; Lauriero, G.; Maranzano, V.; Gozzi, G.; Serrazanetti, D.; Dalfino, G.; Gobbetti, M.; et al. Microbiota and metabolome associated with immunoglobulin A nephropathy (IgAN). PLoS ONE 2014, 9, e99006. [Google Scholar] [CrossRef]
- Wang, F.; Jiang, H.; Shi, K.; Ren, Y.; Zhang, P.; Cheng, S. Gut bacterial translocation is associated with microinflammation in end-stage renal disease patients. Nephrology (Carlton) 2012, 17, 733–738. [Google Scholar] [CrossRef]
- Wang, F.; Zhang, P.; Jiang, H.; Cheng, S. Gut bacterial translocation contributes to microinflammation in experimental uremia. Dig. Dis. Sci. 2012, 57, 2856–2862. [Google Scholar] [CrossRef]
- Jiang, S.; Xie, S.; Lv, D.; Zhang, Y.; Deng, J.; Zeng, L.; Chen, Y. A reduction in the butyrate producing species Roseburia spp. and Faecalibacterium prausnitzii is associated with chronic kidney disease progression. Antonie van Leeuwenhoek 2016, 109, 1389–1396. [Google Scholar] [CrossRef] [PubMed]
- Vaziri, N.D.; Yuan, J.; Norris, K. Role of urea in intestinal barrier dysfunction and disruption of epithelial tight junction in chronic kidney disease. Am. J. Nephrol. 2013, 37, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Lau, W.L.; Vaziri, N.D. Urea, a true uremic toxin: The empire strikes back. Clin. Sci. 2017, 131, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Fathallah-Shaykh, S.A.; Cramer, M.T. Uric acid and the kidney. Pediatr. Nephrol. 2014, 29, 999–1008. [Google Scholar] [CrossRef] [PubMed]
- Ermer, T.; Eckardt, K.U.; Aronson, P.S.; Knauf, F. Oxalate, inflammasome, and progression of kidney disease. Curr. Opin. Nephrol. Hypertens. 2016, 25, 363–371. [Google Scholar] [CrossRef] [PubMed]
- Poesen, R.; Meijers, B.; Evenepoel, P. The colon: An overlooked site for therapeutics in dialysis patients. Semin. Dial. 2013, 26, 323–332. [Google Scholar] [CrossRef]
- Kortman, G.A.M.; Reijnders, D.; Swinkels, D.W. Oral iron supplementation: Potential implications for the gut microbiome and metabolome in patients with CKD. Hemodial. Int. 2017, 21, S28–S36. [Google Scholar] [CrossRef]
- Lau, W.L.; Vaziri, N.D.; Nunes, A.C.F.; Comeau, A.M.; Langille, M.G.I.; England, W.; Khazaeli, M.; Suematsu, Y.; Phan, J.; Whiteson, K. The phosphate binder ferric citrate alters the gut microbiome in rats with chronic kidney disease. J. Pharmacol. Exp. Ther. 2018, 367, 452–460. [Google Scholar] [CrossRef]
- Vangay, P.; Ward, T.; Gerber, J.S.; Knights, D. Antibiotics, pediatric dysbiosis, and disease. Cell Host Microbe 2015, 17, 553–564. [Google Scholar] [CrossRef]
- Gonçalves, S.; Pecoits-Filho, R.; Perreto, S.; Barberato, S.H.; Stinghen, A.E.; Lima, E.G.; Fuerbringer, R.; Sauthier, S.M.; Riella, M.C. Associations between renal function, volume status and endotoxaemia in chronic kidney disease patients. Nephrol. Dial. Transplant. 2006, 21, 2788–2794. [Google Scholar] [CrossRef]
- Szeto, C.C.; Kwan, B.C.; Chow, K.M.; Lai, K.B.; Chung, K.Y.; Leung, C.B.; Li, P.K. Endotoxemia is related to systemic inflammation and atherosclerosis in peritoneal dialysis patients. Clin. J. Am. Soc. Nephrol. 2008, 3, 431–436. [Google Scholar] [CrossRef] [PubMed]
- Raj, D.S.; Carrero, J.J.; Shah, V.O.; Qureshi, A.R.; Barany, P.; Heimburger, O.; Lindholm, B.; Ferguson, J.; Moseley, P.L.; Stenvinkel, P. Soluble CD14 levels, interleukin 6, and mortality among prevalent hemodialysis patients. Am. J. Kidney Dis. 2009, 54, 1072–1080. [Google Scholar] [CrossRef] [PubMed]
- Magnusson, M.; Magnusson, K.E.; Sundqvist, T.; Denneberg, T. Increased intestinal permeability to differently sized polyethylene glycols in uremic rats: Effects of low- and high-protein diets. Nephron 1990, 56, 306–311. [Google Scholar] [CrossRef] [PubMed]
- Magnusson, M.; Magnusson, K.E.; Sundqvist, T.; Denneberg, T. Impaired intestinal barrier function measured by differently sized polyethylene glycols in patients with chronic renal failure. Gut 1991, 32, 754–759. [Google Scholar] [CrossRef] [PubMed]
- de Almeida Duarte, J.B.; de Aguilar-Nascimento, J.E.; Nascimento, M.; Nochi, R.J., Jr. Bacterial translocation in experimental uremia. Urol. Res. 2004, 32, 266–270. [Google Scholar] [CrossRef] [PubMed]
- Vaziri, N.D.; Dure-Smith, B.; Miller, R.; Mirahmadi, M.K. Pathology of gastrointestinal tract in chronic hemodialysis patients: An autopsy study of 78 cases. Am. J. Gastroenterol. 1985, 80, 608–611. [Google Scholar] [PubMed]
- Vaziri, N.D.; Yuan, J.; Nazertehrani, S.; Ni, Z.; Liu, S. Chronic kidney disease causes disruption of gastric and small intestinal epithelial tight junction. Am. J. Nephrol. 2013, 38, 99–103. [Google Scholar] [CrossRef]
- Ritz, E. Intestinal-renal syndrome: Mirage or reality? Blood Purif. 2011, 31, 70–76. [Google Scholar] [CrossRef]
- Vaziri, N.D.; Goshtasbi, N.; Yuan, J.; Jellbauer, S.; Moradi, H.; Raffatellu, M.; Kalantar-Zadeh, K. Uremic plasma impairs barrier function and depletes the tight junction protein constituents of intestinal epithelium. Am. J. Nephrol. 2012, 36, 438–443. [Google Scholar] [CrossRef]
- Shi, K.; Wang, F.; Jiang, H.; Liu, H.; Wei, M.; Wang, Z.; Xie, L. Gut bacterial translocation may aggravate microinflammation in hemodialysis patients. Dig. Dis. Sci. 2014, 59, 2109–2117. [Google Scholar] [CrossRef]
- Bossola, M.; Sanguinetti, M.; Scribano, D.; Zuppi, C.; Giungi, S.; Luciani, G.; Torelli, R.; Posteraro, B.; Fadda, G.; Tazza, L. Circulating bacterial-derived DNA fragments and markers of inflammation in chronic hemodialysis patients. Clin. J. Am. Soc. Nephrol. 2009, 4, 379–385. [Google Scholar] [PubMed]
- Nusrat, A.; Turner, J.R.; Madara, J.L. Molecular physiology and pathophysiology of tight junctions. IV. Regulation of tight junctions by extracellular stimuli: Nutrients, cytokines, and immune cells. Am. J. Physiol. Gastrointest. Liver Physiol. 2000, 279, G851–G857. [Google Scholar] [CrossRef] [PubMed]
- Al-Sadi, R.; Boivin, M.; Ma, T. Mechanism of cytokine modulation of epithelial tight junction barrier. Front. Biosci. (Landmark Ed.) 2009, 14, 2765–2778. [Google Scholar] [CrossRef] [PubMed]
- Bruewer, M.; Samarin, S.; Nusrat, A. Inflammatory bowel disease and the apical junctional complex. Ann. N. Y. Acad. Sci. 2006, 1072, 242–252. [Google Scholar] [CrossRef]
- Shen, L.; Turner, J.R. Role of epithelial cells in initiation and propagation of intestinal inflammation. Eliminating the static: Tight junction dynamics exposed. Am. J. Physiol. Gastrointest. Liver Physiol. 2006, 290, G577–G582. [Google Scholar] [CrossRef] [PubMed]
- Lorentz, C.A.; Liang, Z.; Meng, M.; Chen, C.W.; Yoseph, B.P.; Breed, E.R.; Mittal, R.; Klingensmith, N.J.; Farris, A.B.; Burd, E.M.; et al. Myosin light chain kinase knockout improves gut barrier function and confers a survival advantage in polymicrobial sepsis. Mol. Med. 2017, 23, 155–165. [Google Scholar] [CrossRef] [PubMed]
- Cunningham, K.E.; Turner, J.R. Myosin light chain kinase: Pulling the strings of epithelial tight junction function. Ann. N. Y. Acad. Sci. 2012, 1258, 34–42. [Google Scholar] [CrossRef] [PubMed]
- Lau, W.L.; Savoj, J.; Nakata, M.B.; Vaziri, N.D. Altered microbiome in chronic kidney disease: Systemic effects of gut-derived uremic toxins. Clin. Sci. (Lond) 2018, 132, 509–522. [Google Scholar] [CrossRef]
- Jourde-Chiche, N.; Dou, L.; Cerini, C.; Dignat-George, F.; Vanholder, R.; Brunet, P. Protein-bound toxins--update 2009. Semin. Dial. 2009, 22, 334–339. [Google Scholar] [CrossRef]
- Martinez, A.W.; Recht, N.S.; Hostetter, T.H.; Meyer, T.W. Removal of P-cresol sulfate by hemodialysis. J. Am. Soc. Nephrol. 2005, 16, 3430–3436. [Google Scholar]
- Ufnal, M.; Zadlo, A.; Ostaszewski, R. TMAO: A small molecule of great expectations. Nutrition 2015, 31, 1317–1323. [Google Scholar] [PubMed]
- Zimmerman, L.; Jörnvall, H.; Bergström, J. Phenylacetylglutamine and hippuric acid in uremic and healthy subjects. Nephron 1990, 55, 265–271. [Google Scholar] [PubMed]
- Niwa, T.; Takeda, N.; Tatematsu, A.; Maeda, K. Accumulation of indoxyl sulfate, an inhibitor of drug-binding, in uremic serum as demonstrated by internal-surface reversed-phase liquid chromatography. Clin. Chem. 1988, 34, 2264–2267. [Google Scholar] [PubMed]
- Vanholder, R.; De Smet, R.; Glorieux, G.; Argilés, A.; Baurmeister, U.; Brunet, P.; Clark, W.; Cohen, G.; De Deyn, P.P.; Deppisch, R.; et al. Review on uremic toxins: Classification, concentration, and interindividual variability. Kidney Int. 2003, 63, 1934–1943. [Google Scholar] [CrossRef] [PubMed]
- Liabeuf, S.; Barreto, D.V.; Barreto, F.C.; Meert, N.; Glorieux, G.; Schepers, E.; Temmar, M.; Choukroun, G.; Vanholder, R.; Massy, Z.A. European Uraemic Toxin Work Group (EUTox). Free p-cresylsulphate is a predictor of mortality in patients at different stages of chronic kidney disease. Nephrol. Dial. Transplant. 2010, 25, 1183–1191. [Google Scholar] [PubMed]
- Barreto, F.C.; Barreto, D.V.; Liabeuf, S.; Meert, N.; Glorieux, G.; Temmar, M.; Choukroun, G.; Vanholder, R.; Massy, Z.A. European Uremic Toxin Work Group (EUTox). Serum indoxyl sulfate is associated with vascular disease and mortality in chronic kidney disease patients. Clin. J. Am. Soc. Nephrol. 2009, 4, 1551–1558. [Google Scholar] [CrossRef]
- Wang, Z.; Klipfell, E.; Bennett, B.J.; Koeth, R.; Levison, B.S.; Dugar, B.; Feldstein, A.E.; Britt, E.B.; Fu, X.; Chung, Y.M.; et al. Gut flora metabolism of phosphatidylcholine promotes cardiovascular disease. Nature 2011, 472, 57–63. [Google Scholar] [CrossRef]
- Prokopienko, A.J.; Nolin, T.D. Microbiota-derived uremic retention solutes: Perpetrators of altered nonrenal drug clearance in kidney disease. Expert Rev. Clin. Pharmacol. 2018, 11, 71–82. [Google Scholar] [CrossRef]
- Stubbs, J.R.; House, J.A.; Ocque, A.J.; Zhang, S.; Johnson, C.; Kimber, C.; Schmidt, K.; Gupta, A.; Wetmore, J.B.; Nolin, T.D.; et al. Serum trimethylamine-N-oxide is elevated in CKD and correlates with coronary atherosclerosis burden. J. Am. Soc. Nephrol. 2016, 27, 305–313. [Google Scholar] [CrossRef]
- Lekawanvijit, S.; Adrahtas, A.; Kelly, D.J.; Kompa, A.R.; Wang, B.H.; Krum, H. Does indoxyl sulfate, a uraemic toxin, have direct effects on cardiac fibroblasts and myocytes? Eur. Heart J. 2010, 31, 1771–1779. [Google Scholar] [CrossRef]
- Yisireyili, M.; Shimizu, H.; Saito, S.; Enomoto, A.; Nishijima, F.; Niwa, T. Indoxyl sulfate promotes cardiac fibrosis with enhanced oxidative stress in hypertensive rats. Life Sci. 2013, 92, 1180–1185. [Google Scholar] [CrossRef] [PubMed]
- Wing, M.R.; Patel, S.S.; Ramezani, A.; Raj, D.S. Gut microbiome in chronic kidney disease. Exp. Physiol. 2016, 101, 471–477. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.H.; Mitch, W.E. Mechanisms of muscle wasting in chronic kidney disease. Nat. Rev. Nephrol. 2014, 10, 504–516. [Google Scholar] [CrossRef] [PubMed]
- Enoki, Y.; Watanabe, H.; Arake, R.; Sugimoto, R.; Imafuku, T.; Tominaga, Y.; Ishima, Y.; Kotani, S.; Nakajima, M.; Tanaka, M.; et al. Indoxyl sulfate potentiates skeletal muscle atrophy by inducing the oxidative stress-mediated expression of myostatin and atrogin-1. Sci. Rep. 2016, 6, 32084. [Google Scholar] [CrossRef] [PubMed]
- Sato, E.; Mori, T.; Mishima, E.; Suzuki, A.; Sugawara, S.; Kurasawa, N.; Saigusa, D.; Miura, D.; Morikawa-Ichinose, T.; Saito, R.; et al. Metabolic alterations by indoxyl sulfate in skeletal muscle induce uremic sarcopenia in chronic kidney disease. Sci. Rep. 2016, 6, 36618. [Google Scholar] [CrossRef] [PubMed]
- Sudo, N.; Chida, Y.; Aiba, Y.; Sonoda, J.; Oyama, N.; Yu, X.N.; Kubo, C.; Koga, Y. Postnatal microbial colonization programs the hypothalamic-pituitary-adrenal system for stress response in mice. J. Physiol. 2004, 558, 263–275. [Google Scholar] [CrossRef]
- Mudd, A.T.; Berding, K.; Wang, M.; Donovan, S.M.; Dilger, R.N. Serum cortisol mediates the relationship between fecal Ruminococcus and brain N-acetylaspartate in the young pig. Gut Microb. 2017, 8, 589–600. [Google Scholar] [CrossRef]
- O’Mahony, S.M.; Clarke, G.; Borre, Y.E.; Dinan, T.G.; Cryan, J.F. Serotonin, tryptophan metabolism and the brain-gut-microbiome axis. Behav. Brain Res. 2015, 277, 32–48. [Google Scholar] [CrossRef]
- Tan, J.; McKenzie, C.; Potamitis, M.; Thorburn, A.N.; Mackay, C.R.; Macia, L. The role of short-chain fatty acids in health and disease. Adv. Immunol. 2014, 121, 91–119. [Google Scholar]
- Arentsen, T.; Raith, H.; Qian, Y.; Forssberg, H.; Diaz Heijtz, R. Host microbiota modulates development of social preference in mice. Microb. Ecol. Health Dis. 2015, 26, 29719. [Google Scholar] [CrossRef]
- Neuman, H.; Debelius, J.W.; Knight, R.; Koren, O. Microbial endocrinology: The interplay between the microbiota and the endocrine system. FEMS Microbiol. Rev. 2015, 39, 509–521. [Google Scholar] [CrossRef] [PubMed]
- Kim, N.; Yun, M.; Oh, Y.J.; Choi, H.J. Mind-altering with the gut: Modulation of the gut-brain axis with probiotics. J. Microbiol. 2018, 56, 172–182. [Google Scholar] [CrossRef] [PubMed]
- Afsar, B.; Vaziri, N.D.; Aslan, G.; Tarim, K.; Kanbay, M. Gut hormones and gut microbiota: Implications for kidney function and hypertension. J. Am. Soc. Hypertens 2016, 10, 954–961. [Google Scholar] [CrossRef] [PubMed]
- Bravo, J.A.; Forsythe, P.; Chew, M.V.; Escaravage, E.; Savignac, H.M.; Dinan, T.G.; Bienenstock, J.; Cryan, J.F. Ingestion of Lactobacillus strain regulates emotional behavior and central GABA receptor expression in a mouse via the vagus nerve. Proc. Natl. Acad. Sci. USA 2011, 108, 16050–16055. [Google Scholar] [CrossRef] [PubMed]
- Farzi, A.; Frohlich, E.E.; Holzer, P. Gut microbiota and the neuroendocrine system. Neurotherapeutics 2018, 15, 5–22. [Google Scholar] [CrossRef]
- Asao, T.; Oki, K.; Yoneda, M.; Tanaka, J.; Kohno, N. Hypothalamic-pituitary-adrenal axis activity is associated with the prevalence of chronic kidney disease in diabetic patients. Endocr. J. 2016, 63, 119–126. [Google Scholar] [CrossRef]
- de Punder, K.; Pruimboom, L. Stress induces endotoxemia and low-grade inflammation by increasing barrier permeability. Front. Immunol. 2015, 6, 223. [Google Scholar] [CrossRef]
- Kelly, J.R.; Kennedy, P.J.; Cryan, J.F.; Dinan, T.G.; Clarke, G.; Hyland, N.P. Breaking down the barriers: The gut microbiome, intestinal permeability and stress-related psychiatric disorders. Front. Cell Neurosci. 2015, 9, 392. [Google Scholar] [CrossRef]
- Larraufie, P.; Martin-Gallausiaux, C.; Lapaque, N.; Dore, J.; Gribble, F.M.; Reimann, F.; Blottiere, H.M. SCFAs strongly stimulate PYY production in human enteroendocrine cells. Sci. Rep. 2018, 8, 74. [Google Scholar] [CrossRef]
- Shih, P.A.; Wang, L.; Chiron, S.; Wen, G.; Nievergelt, C.; Mahata, M.; Khandrika, S.; Rao, F.; Fung, M.M.; Mahata, S.K.; et al. Peptide YY (PYY) gene polymorphisms in the 3’-untranslated and proximal promoter regions regulate cellular gene expression and PYY secretion and metabolic syndrome traits in vivo. J. Clin. Endocrinol. Metab. 2009, 94, 4557–4566. [Google Scholar] [CrossRef]
- VanDeVoorde, R.G.; Mitsnefes, M.M. Hypertension and CKD. Adv. Chronic Kidney Dis. 2011, 18, 355–361. [Google Scholar] [CrossRef]
- Stenvinkel, P.; Zoccali, C.; Ikizler, T.A. Obesity in CKD--what should nephrologists know? J. Am. Soc. Nephrol. 2013, 24, 1727–1736. [Google Scholar] [CrossRef] [PubMed]
- Desbonnet, L.; Garrett, L.; Clarke, G.; Kiely, B.; Cryan, J.F.; Dinan, T.G. Effects of the probiotic Bifidobacterium infantis in the maternal separation model of depression. Neuroscience 2010, 170, 1179–1188. [Google Scholar] [CrossRef]
- Yano, J.M.; Yu, K.; Donaldson, G.P.; Shastri, G.G.; Ann, P.; Ma, L.; Nagler, C.R.; Ismagilov, R.F.; Mazmanian, S.K.; Hsiao, E.Y. Indigenous bacteria from the gut microbiota regulate host serotonin biosynthesis. Cell 2015, 161, 264–276. [Google Scholar] [CrossRef] [PubMed]
- Gregg, L.P.; Hedayati, S.S. Management of traditional cardiovascular risk factors in CKD: What are the data? Am. J. Kidney Dis. 2018, 72, 728–744. [Google Scholar] [CrossRef] [PubMed]
- Jimenez, E.; Ladero, V.; Chico, I.; Maldonado-Barragan, A.; Lopez, M.; Martin, V.; Fernández, L.; Fernández, M.; Álvarez, M.A.; Torres, C.; et al. Antibiotic resistance, virulence determinants and production of biogenic amines among enterococci from ovine, feline, canine, porcine and human milk. BMC Microbiol. 2013, 13, 288. [Google Scholar] [CrossRef]
- Lyte, M. Probiotics function mechanistically as delivery vehicles for neuroactive compounds: Microbial endocrinology in the design and use of probiotics. Bioessays 2011, 33, 574–581. [Google Scholar] [CrossRef]
- Shishov, V.A.; Kirovskaia, T.A.; Kudrin, V.S.; Oleskin, A.V. Amine neuromediators, their precursors, and oxidation products in the culture of Escherichia coli K-12. Prikl Biokhim Mikrobiol. 2009, 45, 550–554. [Google Scholar] [CrossRef]
- Camilleri, M. Serotonin in the gastrointestinal tract. Curr. Opin. Endocrinol. Diabetes Obes. 2009, 16, 53–59. [Google Scholar] [CrossRef]
- Lee, J.H.; Lee, J. Indole as an intercellular signal in microbial communities. FEMS Microbiol. Rev. 2010, 34, 426–444. [Google Scholar] [CrossRef]
- Li, G.; Young, K.D. Indole production by the tryptophanase TnaA in Escherichia coli is determined by the amount of exogenous tryptophan. Microbiology 2013, 159, 402–410. [Google Scholar] [CrossRef] [PubMed]
- Raboni, S.; Bettati, S.; Mozzarelli, A. Tryptophan synthase: A mine for enzymologists. Cell Mol. Life Sci. 2009, 66, 2391–2403. [Google Scholar] [CrossRef] [PubMed]
- Yanofsky, C. RNA-based regulation of genes of tryptophan synthesis and degradation, in bacteria. RNA 2007, 13, 1141–1154. [Google Scholar] [CrossRef] [PubMed]
- Daneman, R.; Rescigno, M. The gut immune barrier and the blood-brain barrier: Are they so different? Immunity 2009, 31, 722–735. [Google Scholar] [CrossRef] [PubMed]
- Park, H.; Shin, H.; Lee, K.; Holzapfel, W. Autoinducer-2 properties of kimchi are associated with lactic acid bacteria involved in its fermentation. Int. J. Food Microbiol. 2016, 225, 38–42. [Google Scholar] [CrossRef] [PubMed]
- Karavolos, M.H.; Winzer, K.; Williams, P.; Khan, C.M. Pathogen espionage: Multiple bacterial adrenergic sensors eavesdrop on host communication systems. Mol. Microbiol. 2013, 87, 455–465. [Google Scholar] [CrossRef] [PubMed]
- Bellezza, I.; Peirce, M.J.; Minelli, A. Cyclic dipeptides: From bugs to brain. Trends Mol. Med. 2014, 20, 551–558. [Google Scholar] [CrossRef] [PubMed]
- Modaresi, A.; Nafar, M.; Sahraei, Z. Oxidative stress in chronic kidney disease. Iran. J. Kidney Dis. 2015, 9, 165–179. [Google Scholar] [PubMed]
- Marietta, E.; Horwath, I.; Taneja, V. Microbiome, immunomodulation, and the neuronal system. Neurotherapeutics 2018, 15, 23–30. [Google Scholar] [CrossRef] [PubMed]
- Bonaz, B.; Bazin, T.; Pellissier, S. The vagus nerve at the interface of the microbiota-gut-brain axis. Front. Neurosci. 2018, 12, 49. [Google Scholar] [CrossRef]
- Raybould, H.E. Gut chemosensing: Interactions between gut endocrine cells and visceral afferents. Auton Neurosci. 2010, 153, 41–46. [Google Scholar] [CrossRef] [PubMed]
- Tracey, K.J. Physiology and immunology of the cholinergic antiinflammatory pathway. J. Clin. Investig. 2007, 117, 289–296. [Google Scholar] [CrossRef] [PubMed]
- Edwards, J.K. Acute kidney injury: Vagus nerve stimulation may prevent AKI. Nat. Rev. Nephrol. 2016, 12, 376. [Google Scholar] [CrossRef] [PubMed]
- Task Force of the European Society of Cardiology and the North American Society of Pacing and Electrophysiology. Heart rate variability. Standards of measurement, physiological interpretation, and clinical use. Eur. Heart J. 1996, 17, 354–381. [Google Scholar]
- Bonaz, B.; Sinniger, V.; Pellissier, S. Vagal tone: Effects on sensitivity, motility, and inflammation. Neurogastroenterol. Motil. 2016, 28, 455–462. [Google Scholar] [CrossRef]
- Montemurno, E.; Cosola, C.; Dalfino, G.; Daidone, G.; De Angelis, M.; Gobbetti, M. What would you like to eat, Mr CKD microbiota? A mediterranean diet, please! Kidney Blood Press Res. 2014, 39, 114–123. [Google Scholar] [CrossRef] [PubMed]
- Mekki, K.; Bouzidi-bekada, N.; Kaddous, A.; Bouchenak, M. Mediterranean diet improves dyslipidemia and biomarkers in chronic renal failure patients. Food Funct. 2010, 1, 110–115. [Google Scholar] [CrossRef]
- Pignanelli, M.; Just, C.; Bogiatzi, C.; Dinculescu, V.; Gloor, G.B.; Allen-Vercoe, E.; Reid, G.; Urquhart, B.L.; Ruetz, K.N.; Velenosi, T.J.; et al. Mediterranean diet score: Associations with metabolic products of the intestinal microbiome, carotid plaque burden, and renal function. Nutrients 2018, 10, 779. [Google Scholar] [CrossRef]
- Lee, H.S.; Lee, K.B.; Hyun, Y.Y.; Chang, Y.; Ryu, S.; Choi, Y. DASH dietary pattern and chronic kidney disease in elderly Korean adults. Eur. J. Clin. Nutr. 2017, 71, 755–761. [Google Scholar] [CrossRef]
- Esgalhado, M.; Kemp, J.A.; Damasceno, N.R.; Fouque, D.; Mafra, D. Short-chain fatty acids: A link between prebiotics and microbiota in chronic kidney disease. Future Microbiol. 2017, 12, 1413–1425. [Google Scholar] [CrossRef]
- Salehi-Abargouei, A.; Ghiasvand, R.; Hariri, M. Prebiotics, prosynbiotics and synbiotics: Can they reduce plasma oxidative stress parameters? A systematic review. Probiotics Antimicrob Proteins 2017, 9, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Valcheva, R.; Dieleman, L.A. Prebiotics: Definition and protective mechanisms. Best Pract. Res. Clin. Gastroenterol. 2016, 30, 27–37. [Google Scholar] [CrossRef] [PubMed]
- Slavin, J. Fiber and prebiotics: Mechanisms and health benefits. Nutrients 2013, 5, 1417–1435. [Google Scholar] [CrossRef] [PubMed]
- Kieffer, D.A.; Piccolo, B.D.; Vaziri, N.D.; Liu, S.; Lau, W.L.; Khazaeli, M.; Nazertehrani, S.; Moore, M.E.; Marco, M.L.; Martin, R.J.; Adams, S.H. Resistant starch alters gut microbiome and metabolomic profiles concurrent with amelioration of chronic kidney disease in rats. Am. J. Physiol Renal Physiol. 2016, 310, F857–F871. [Google Scholar] [CrossRef] [PubMed]
- DiNicolantonio, J.J.; Bhutani, J.; O’Keefe, J.H. Acarbose: Safe and effective for lowering postprandial hyperglycaemia and improving cardiovascular outcomes. Open Heart 2015, 2, e000327. [Google Scholar] [CrossRef] [PubMed]
- Weaver, G.A.; Tangel, C.T.; Krause, J.A.; Parfitt, M.M.; Jenkins, P.L.; Rader, J.M.; Lewis, B.A.; Miller, T.L.; Wolin, M.J. Acarbose enhances human colonic butyrate production. J. Nutr. 1997, 127, 717–723. [Google Scholar] [CrossRef] [PubMed]
- Xu, G.D.; Cai, L.; Ni, Y.S.; Tian, S.Y.; Lu, Y.Q.; Wang, L.N.; Chen, L.L.; Ma, W.Y.; Deng, S.P. Comparisons of effects on intestinal short-chain fatty acid concentration after exposure of two glycosidase inhibitors in mice. Biol. Pharm. Bull. 2018, 41, 1024–1033. [Google Scholar] [CrossRef]
- Harrison, D.E.; Strong, R.; Allison, D.B.; Ames, B.N.; Astle, C.M.; Atamna, H.; Fernandez, E.; Flurkey, K.; Javors, M.A.; Nadon, N.L.; et al. Acarbose, 17-α-estradiol, and nordihydroguaiaretic acid extend mouse lifespan preferentially in males. Aging Cell 2014, 13, 273–282. [Google Scholar] [CrossRef]
- Strong, R.; Miller, R.A.; Antebi, A.; Astle, C.M.; Bogue, M.; Denzel, M.S.; Fernandez, E.; Flurkey, K.; Hamilton, K.L.; Lamming, D.W.; et al. Longer lifespan in male mice treated with a weakly estrogenic agonist, an antioxidant, an α-glucosidase inhibitor or a Nrf2-inducer. Aging Cell 2016, 15, 872–884. [Google Scholar] [CrossRef]
- Zhang, X.; Fang, Z.; Zhang, C.; Xia, H.; Jie, Z.; Han, X.; Chen, Y.; Ji, L. Effects of acarbose on the gut microbiota of prediabetic patients: A randomized, double-blind, controlled crossover trial. Diabetes Ther. 2017, 8, 293–307. [Google Scholar] [CrossRef]
- Ranganathan, N.; Ranganathan, P.; Friedman, E.A.; Joseph, A.; Delano, B.; Goldfarb, D.S.; Tam, P.; Rao, A.V.; Anteyi, E.; Musso, C.G. Pilot study of probiotic dietary supplementation for promoting healthy kidney function in patients with chronic kidney disease. Adv. Ther. 2010, 27, 634–647. [Google Scholar] [CrossRef] [PubMed]
- Borges, N.A.; Carmo, F.L.; Stockler-Pinto, M.B.; de Brito, J.S.; Dolenga, C.J.; Ferreira, D.C.; Nakao, L.S.; Rosado, A.; Fouque, D.; Mafra, D. Probiotic supplementation in chronic kidney disease: A double-blind, randomized, placebo-controlled trial. J. Ren. Nutr. 2018, 28, 28–36. [Google Scholar] [CrossRef] [PubMed]
- Natarajan, R.; Pechenyak, B.; Vyas, U.; Ranganathan, P.; Weinberg, A.; Liang, P.; Mallappallil, MC.; Norin, A.J.; Friedman, E.A.; Saggi, S.J. Randomized controlled trial of strain-specific probiotic formulation (Renadyl) in dialysis patients. Biomed. Res. Int. 2014, 2014, 568571. [Google Scholar] [CrossRef] [PubMed]
- Neef, A.; Sanz, Y. Future for probiotic science in functional food and dietary supplement development. Curr. Opin. Clin. Nutr. Metab. Care 2013, 16, 679–687. [Google Scholar] [CrossRef] [PubMed]
- El Hage, R.; Hernandez-Sanabria, E.; Van de Wiele, T. Emerging trends in “smart probiotics”: Functional consideration for the development of novel health and industrial applications. Front. Microbiol. 2017, 8, 1889. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Lin, S.; Vanhoutte, P.M.; Woo, C.W.; Xu, A. Akkermansia muciniphila protects against atherosclerosis by preventing metabolic endotoxemia-induced inflammation in apoe-/-mice. Circulation 2016, 133, 2434–2446. [Google Scholar] [CrossRef]
- Troy, E.B.; Kasper, D.L. Beneficial effects of Bacteroides fragilis polysaccharides on the immune system. Front. Biosci. 2010, 15, 25–34. [Google Scholar] [CrossRef]
- Engels, C.; Ruscheweyh, H.J.; Beerenwinkel, N.; Lacroix, C.; Schwab, C. The common gut microbe eubacterium hallii also contributes to intestinal propionate formation. Front. Microbiol. 2016, 7, 713. [Google Scholar] [CrossRef]
- Cani, P.D.; Van Hul, M. Novel opportunities for next-generation probiotics targeting metabolic syndrome. Curr. Opin. Biotechnol. 2015, 32, 21–27. [Google Scholar] [CrossRef]
- Atarashi, K.; Tanoue, T.; Shima, T.; Imaoka, A.; Kuwahara, T.; Momose, Y.; Cheng, G.; Yamasaki, S.; Saito, T.; Ohba, Y.; et al. Induction of colonic regulatory T cells by indigenous Clostridium species. Science 2011, 331, 337–341. [Google Scholar] [CrossRef]
- Rossi, M.; Johnson, D.W.; Morrison, M.; Pascoe, E.M.; Coombes, J.S.; Forbes, J.M.; Szeto, C.C.; McWhinney, B.C.; Ungerer, J.P.; Campbell, K.L. Synbiotics easing renal failure by improving gut microbiology (synergy): A randomized trial. Clin. J. Am. Soc. Nephrol. 2016, 11, 223–231. [Google Scholar] [CrossRef] [PubMed]
- Nakabayashi, I.; Nakamura, M.; Kawakami, K.; Ohta, T.; Kato, I.; Uchida, K.; Yoshida, M. Effects of synbiotic treatment on serum level of p-cresol in haemodialysis patients: A preliminary study. Nephrol. Dial. Transplant. 2011, 26, 1094–1098. [Google Scholar] [CrossRef] [PubMed]
- Andrade-Oliveira, V.; Amano, M.T.; Correa-Costa, M.; Castoldi, A.; Felizardo, R.J.; de Almeida, D.C.; Bassi, E.J.; Moraes-Vieira, P.M.; Hiyane, M.I.; Rodas, A.C.; Peron, J.P.; et al. Gut bacteria products prevent aki induced by ischemia-reperfusion. J. Am. Soc. Nephrol. 2015, 26, 1877–1888. [Google Scholar] [CrossRef] [PubMed]
- Bliss, D.Z.; Stein, T.P.; Schleifer, C.R.; Settle, R.G. Supplementation with gum arabic fiber increases fecal nitrogen excretion and lowers serum urea nitrogen concentration in chronic renal failure patients consuming a low-protein diet. Am. J. Clin. Nutr. 1996, 63, 392–398. [Google Scholar] [CrossRef] [PubMed]
- Younes, H.; Egret, N.; Hadj-Abdelkader, M.; Remesy, C.; Demigne, C.; Gueret, C.; Deteix, P.; Alphonse, J.C. Fermentable carbohydrate supplementation alters nitrogen excretion in chronic renal failure. J. Ren. Nutr. 2006, 16, 67–74. [Google Scholar] [CrossRef] [PubMed]
- Sirich, T.L.; Plummer, N.S.; Gardner, C.D.; Hostetter, T.H.; Meyer, T.W. Effect of increasing dietary fiber on plasma levels of colon-derived solutes in hemodialysis patients. Clin. J. Am. Soc. Nephrol. 2014, 9, 1603–1610. [Google Scholar] [CrossRef]
- Xie, L.M.; Ge, Y.Y.; Huang, X.; Zhang, Y.Q.; Li, J.X. Effects of fermentable dietary fiber supplementation on oxidative and inflammatory status in hemodialysis patients. Int. J. Clin. Exp. Med. 2015, 8, 1363–1369. [Google Scholar]
- Poesen, R.; Evenepoel, P.; de Loor, H.; Delcour, J.A.; Courtin, C.M.; Kuypers, D.; Augustijns, P.; Verbeke, K.; Meijers, B. The influence of prebiotic arabinoxylan oligosaccharides on microbiota derived uremic retention solutes in patients with chronic kidney disease: A randomized controlled trial. PLoS ONE 2016, 11, e0153893. [Google Scholar] [CrossRef]
- Ranganathan, N.; Patel, B.; Ranganathan, P.; Marczely, J.; Dheer, R.; Chordia, T.; Dunn, S.R.; Friedman, E.A. Probiotic amelioration of azotemia in 5/6th nephrectomized Sprague-Dawley rats. Sci. World J. 2005, 5, 652–660. [Google Scholar] [CrossRef]
- Ranganathan, N.; Patel, B.G.; Ranganathan, P.; Marczely, J.; Dheer, R.; Pechenyak, B.; Dunn, S.R.; Verstraete, W.; Decroos, K.; Mehta, R.; et al. In vitro and in vivo assessment of intraintestinal bacteriotherapy in chronic kidney disease. ASAIO J. 2006, 52, 70–79. [Google Scholar] [CrossRef]
- Prakash, S.; Chang, T.M. Microencapsulated genetically engineered live E. coli DH5 cells administered orally to maintain normal plasma urea level in uremic rats. Nat. Med. 1996, 2, 883–887. [Google Scholar] [CrossRef] [PubMed]
- Lippi, I.; Perondi, F.; Ceccherini, G.; Marchetti, V.; Guidi, G. Effects of probiotic VSL#3 on glomerular filtration rate in dogs affected by chronic kidney disease: A pilot study. Can. Vet. J. 2017, 58, 1301–1305. [Google Scholar] [PubMed]
- Ranganathan, N.; Friedman, E.A.; Tam, P.; Rao, V.; Ranganathan, P.; Dheer, R. Probiotic dietary supplementation in patients with stage 3 and 4 chronic kidney disease: A 6-month pilot scale trial in Canada. Curr. Med. Res. Opin. 2009, 25, 1919–1930. [Google Scholar] [CrossRef] [PubMed]
- Taki, K.; Takayama, F.; Niwa, T. Beneficial effects of Bifidobacteria in a gastroresistant seamless capsule on hyperhomocysteinemia in hemodialysis patients. J. Ren. Nutr. 2005, 15, 77–80. [Google Scholar] [CrossRef] [PubMed]
- Takayama, F.; Taki, K.; Niwa, T. Bifidobacterium in gastro-resistant seamless capsule reduces serum levels of indoxyl sulfate in patients on hemodialysis. Am. J. Kidney Dis. 2003, 41, S142–S145. [Google Scholar] [CrossRef] [PubMed]
- Ando, Y.; Miyata, Y.; Tanba, K.; Saito, O.; Muto, S.; Kurosu, M.; Homma, S.; Kusano, E.; Asano, Y. Effect of oral intake of an enteric capsule preparation containing Bifidobacterium longum on the progression of chronic renal failure. Nihon Jinzo Gakkai shi. 2003, 45, 759–764. [Google Scholar] [PubMed]
- Simenhoff, M.L.; Dunn, S.R.; Zollner, G.P.; Fitzpatrick, M.E.; Emery, S.M.; Sandine, W.E.; Ayres, J.W. Biomodulation of the toxic and nutritional effects of small bowel bacterial overgrowth in end-stage kidney disease using freeze-dried Lactobacillus acidophilus. Miner. Electrol. Metab. 1996, 22, 92–96. [Google Scholar]
- Wang, I.K.; Wu, Y.Y.; Yang, Y.F.; Ting, I.W.; Lin, C.C.; Yen, T.H.; Chen, J.H.; Wang, C.H.; Huang, C.C.; Lin, H.C. The effect of probiotics on serum levels of cytokine and endotoxin in peritoneal dialysis patients: A randomised, double-blind, placebo-controlled trial. Benef. Microb. 2015, 6, 423–430. [Google Scholar] [CrossRef] [PubMed]
- Miranda Alatriste, P.V.; Urbina Arronte, R.; Gomez Espinosa, C.O.; Espinosa Cuevas Mde, L. Effect of probiotics on human blood urea levels in patients with chronic renal failure. Nutr. Hosp. 2014, 29, 582–590. [Google Scholar] [PubMed]
- Tao, S.; Tao, S.; Cheng, Y.; Liu, J.; Ma, L.; Fu, P. Effects of probiotic supplements on the progression of chronic kidney disease: A meta-analysis. Nephrology 2018. [Google Scholar] [CrossRef] [PubMed]
- Jia, L.; Jia, Q.; Yang, J.; Jia, R.; Zhang, H. Efficacy of probiotics supplementation on chronic kidney disease: A systematic review and meta-analysis. Kidney Blood Press Res. 2018, 43, 1623–1635. [Google Scholar] [CrossRef] [PubMed]
- Dehghani, H.; Heidari, F.; Mozaffari-Khosravi, H.; Nouri-Majelan, N.; Dehghani, A. Synbiotic supplementations for azotemia in patients with chronic kidney disease: A randomized controlled trial. Iran. J. Kidney Dis. 2016, 10, 351–357. [Google Scholar] [PubMed]
- Pavan, M. Influence of prebiotic and probiotic supplementation on the progression of chronic kidney disease. Minerva Urol. Nefrol. 2016, 68, 222–226. [Google Scholar] [PubMed]
- Guida, B.; Germano, R.; Trio, R.; Russo, D.; Memoli, B.; Grumetto, L.; Barbato, F.; Cataldi, M. Effect of short-term synbiotic treatment on plasma p-cresol levels in patients with chronic renal failure: A randomized clinical trial. Nutr. Metab. Cardiovasc. Dis. 2014, 24, 1043–1049. [Google Scholar] [CrossRef] [PubMed]
- McFarlane, C.; Ramos, C.I.; Johnson, D.W.; Campbell, K.L. Prebiotic, probiotic, and synbiotic supplementation in chronic kidney disease: A systematic review and meta-analysis. J. Ren. Nutr. 2018, 30191–30192, 1051–2276. [Google Scholar] [CrossRef] [PubMed]

| Species | Dietary Intervention | Study Type | Outcomes | References |
|---|---|---|---|---|
| Prebiotics | ||||
| Mice | Short-chain fatty acids (acetate, propionate, and butyrate, pH 7.4 diluted in PBS) | Pilot study | Delayed progression of chronic kidney disease. Improved mitochondrial biogenesis. Reduced local and systemic inflammation, cellular oxidative stress, cell infiltration/activation and apoptosis. | [183] |
| Rat | Amylose maize resistant starch | Original research study | Attenuation of oxidative stress and inflammation. Delayed progression of chronic kidney disease. | [44] |
| Rat | High amylose maize-resistant starch type 2 (HAMRS2) | Original research study | Reduction in serum and urine indoxyl sulfate levels. Reduction in urine p-cresol level. Improvements in kidney function indexes and amelioration of chronic kidney disease outcomes. | [164] |
| Human | Gum arabic (highly fermentable fiber) | Clinical trial | Significant decrease in serum urea nitrogen. Significant increase in fecal bacterial mass and fecal nitrogen content. | [184] |
| Human | Fermentable carbohydrate | Clinical trial | Significant increase in stool nitrogen excretion. Significant decrease in the urinary nitrogen excretion. Unchanged total nitrogen excreted by the two routes. Significant decrease in plasma urea levels. | [185] |
| Human | Resistant starch | Clinical trial | Significant reduction in plasma indoxyl sulfate. Insignificant reduction in plasma p-cresyl sulfate. | [186] |
| Human | Soluble dietary fiber | Clinical trial | Significant decrease in total cholesterol (TC), low-density lipoprotein (LDL), and TC: LDL ratio. Significant decrease in malondialdehyde, tumor necrosis factor (TNF)-α, interleukin (IL)-6, IL-8, and C-reactive protein levels. No changes in triglycerides, high-density lipoprotein, Cu–Zn superoxide dismutase, and glutathione peroxidase levels. | [187] |
| Human | Arabinoxylan oligosaccharides | Clinical trial | No significant effect on serum p-cresyl sulfate, p-cresyl glucuronide, indoxyl sulfate and phenylacetylglutamine. Small, albeit significant decrease in serum trimethylamine N-oxide. No change in the urinary excretion of p-cresyl sulfate, p-cresyl glucuronide, indoxyl sulfate phenylacetylglutamine, and trimethylamine N-oxide. No significant change in homeostatic model assessment. No influence on microbiota-derived uremic retention solutes and insulin resistance. | [188] |
| Probiotics | ||||
| Rat | Various combinations of Bacillus pasteurii, Sporolac, Kibow cocktail, CHR Hansen Cocktail, and Econorm | Pilot study | Improved survival. Reduction in blood urea nitrogen levels. Delayed progression of chronic kidney disease. | [189] |
| Rat | Soil-borne alkalophilic urease-positive bacterium Sporosarcina pasteurii | Pilot study | Reduced blood urea nitrogen levels. Improved survival. | [190] |
| Rat | Escherichia coli DH5 given with urease | Original research study | Reduction of the high plasma urea level to normal | [191] |
| Dog | VSL#3 supplementation | Original research study | Significant increase in estimated glomerular filtration rate. | [192] |
| Human | L. acidophilus, S. thermophilus and B. longum | Clinical trial | Significant reduction in blood urea nitrogen levels. Improved quality-of-life scores. | [171] |
| Human | L. acidophilus, S. thermophilus, B. longum | Clinical trial | Significant reduction of blood urea nitrogen. Moderate reduction in uric acid levels. Insignificant changes in serum creatinine. Improved quality of life scores. | [193] |
| Human | B. longum | Clinical trial | Significant decrease in predialysis serum levels of homocysteine, indoxyl sulfate, and triglycerides. | [194] |
| Human | B. longum | Clinical trial | Reduction in serum indoxyl sulfate. | [195] |
| Human | B. longum | Clinical trial | Delayed progression of chronic kidney disease. | [196] |
| Human | Lebenin (antibiotic-resistant lactic acid bacteria) | Clinical trial | Reduction in levels of uremic toxins (especially the plasma level of indican). | [67] |
| Human | L. acidophilus | Clinical trial | Reduction of serum dimethylamine and nitrosodimethylamine. Improved nutritional status. | [197] |
| Human | Streptococcus thermophilus, Lactobacillus acidophilus and Bifidobacteria longum | Clinical trial | Significant increase in serum urea nitrogen. Reduction in fecal pH. No effect on inflammatory markers and gut microbiome profile. | [172] |
| Human | Bifobacterium bifidum, Bifidobacterium catenulatum, Bifidobacterium longum and Lactobacillus plantarum | Clinical trial | Significant reduction in serum TNF-α, IL-5, IL-6, and endotoxin. Significant increase in serum IL-10 levels. | [198] |
| Human | S. thermophilus, L. acidophilus, and B. longum | Clinical trial | Non-significant improvement in quality-of-life scores. Non-significant reduction of serum indoxyl glucuronide and C-reactive protein. | [173] |
| Human | Lactobacillus casei shirota | Clinical trial | >10% decrease in serum urea concentrations. | [199] |
| Human | Probiotics | Meta-analysis | Significant reduction in urea level in non-dialysis patients but no change in dialysis patients. No effects on uric acid, C-reactive protein, creatinine, and estimated glomerular filtration rate. | [200] |
| Human | Probiotics | Meta-analysis | Decrease in p-cresyl sulfate. Increase in IL-6. No effects on serum creatinine, blood urine nitrogen, C-reactive protein and hemoglobin levels. | [201] |
| Synbiotics | ||||
| Human | Prebiotics; galactooligosaccharides Probiotics: Lactobacillus casei strain Shirota and Bifidobacterium breve strain Yakult | Clinical trial | Significant decrease in serum p-cresol level. Normalization of bowel habits. | [182] |
| Human | Prebiotics: inulin high performance, fructo-oligosaccharides, and galactooligosaccharides Probiotics: Lactobacillus, Bifidobacteria, and Streptococcus species | Clinical trial | Significant decrease in serum p-cresyl sulfate. Favorable modification of the stool microbiome. | [181] |
| Human | Prebiotics: Fructooligosaccharides Probiotics: Lactobacilus casei, Lactobacilus acidophilus, Lactobacilus bulgarigus, Lactobacilus rhamnosus, Bifidobacterium breve, Bifidobacterium longum, and Streptococcus thermophilus | Clinical trial | Significant reduction in blood urea nitrogen levels. | [202] |
| Human | Prebiotics: Fructooligosaccharides Probiotics: Streptococcus thermophiles, Lactobacillus acidophilus, Bifidobacterium longum | Clinical trial | Significant lowering of the rate of decline in estimated glomerular filtration rate. | [203] |
| Human | Commercial symbiotic formulation: Probinul neutro | Clinical trial | Significant reduction in total plasma p-cresol level. | [204] |
| Human | Prebiotic and Probiotics | Meta-analysis | Synbiotic interventions significantly increased Bifidobacterium in gut microbiota, but had little or no effect on serum urea nitrogen, indoxyl sulfate, and p-cresyl sulfate. Prebiotic supplementation may slightly reduce serum urea concentration. | [205] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jazani, N.H.; Savoj, J.; Lustgarten, M.; Lau, W.L.; Vaziri, N.D. Impact of Gut Dysbiosis on Neurohormonal Pathways in Chronic Kidney Disease. Diseases 2019, 7, 21. https://doi.org/10.3390/diseases7010021
Jazani NH, Savoj J, Lustgarten M, Lau WL, Vaziri ND. Impact of Gut Dysbiosis on Neurohormonal Pathways in Chronic Kidney Disease. Diseases. 2019; 7(1):21. https://doi.org/10.3390/diseases7010021
Chicago/Turabian StyleJazani, Nima H., Javad Savoj, Michael Lustgarten, Wei Ling Lau, and Nosratola D. Vaziri. 2019. "Impact of Gut Dysbiosis on Neurohormonal Pathways in Chronic Kidney Disease" Diseases 7, no. 1: 21. https://doi.org/10.3390/diseases7010021
APA StyleJazani, N. H., Savoj, J., Lustgarten, M., Lau, W. L., & Vaziri, N. D. (2019). Impact of Gut Dysbiosis on Neurohormonal Pathways in Chronic Kidney Disease. Diseases, 7(1), 21. https://doi.org/10.3390/diseases7010021





