1. Introduction
Machine learning algorithms learn from observations whose class labels are already defined, and they make predictions about new instances based on the models constructed during the training process [
1]. There are three main approaches to machine learning: supervised, unsupervised and semi-supervised [
2].
Supervised learning infers classification functions from training labeled data. Major supervised learning approaches are multi-layer perceptron neural networks, decision tree learning, support vector machines and symbolic machine learning algorithms [
3]. Unsupervised learning infers functions from the hidden structure of unlabeled data. The major unsupervised learning approaches are clustering (k-means, mixture models and hierarchical clustering), anomaly detection and neural networking (Hebbian learning and generative adversarial networks) [
3]. Semi-supervised learning infers classification functions from a large amount of unlabeled data, together with a small amount of labeled data [
4]. It thus falls between supervised learning and unsupervised learning. The main semi-supervised learning methods are generative models, low-density separation, graph-based methods and heuristic approaches [
3].
Among supervised learning models, the support vector machine (SVM) proposed by Boser, Guyon and Vapnik in 1992 [
5] is one of the best-known classification methods. SVMs are capable of solving complicated tasks, like local minima, overfitting and high dimension, while delivering an outstanding performance [
6]. Important elements for the efficient use of SVMs are data preprocessing, the selection of the correct kernel, and optimized SVM and kernel setting [
5]. The main limitation with SVMs is the difficulty of resolving quadratic programming when the number of data points is large [
6].
SVM classification algorithms fall into two classes: linear and nonlinear [
7]. A linear classifier is the dot product of a data vector and a weight vector, with the addition of a bias value. Assuming
x is a vector,
w is a weight vector and
b is the bias value, Equation (1) is the discrimination function of a two-class linear classifier [
5].
If
b = 0, the points
x such that
are all vectors to
w that go through the origin. In two dimensions, it is a line; and in three dimensions, a plane and, generally, a hyperplane. The bias
b is a translation of the hyperplane from its origin [
8].
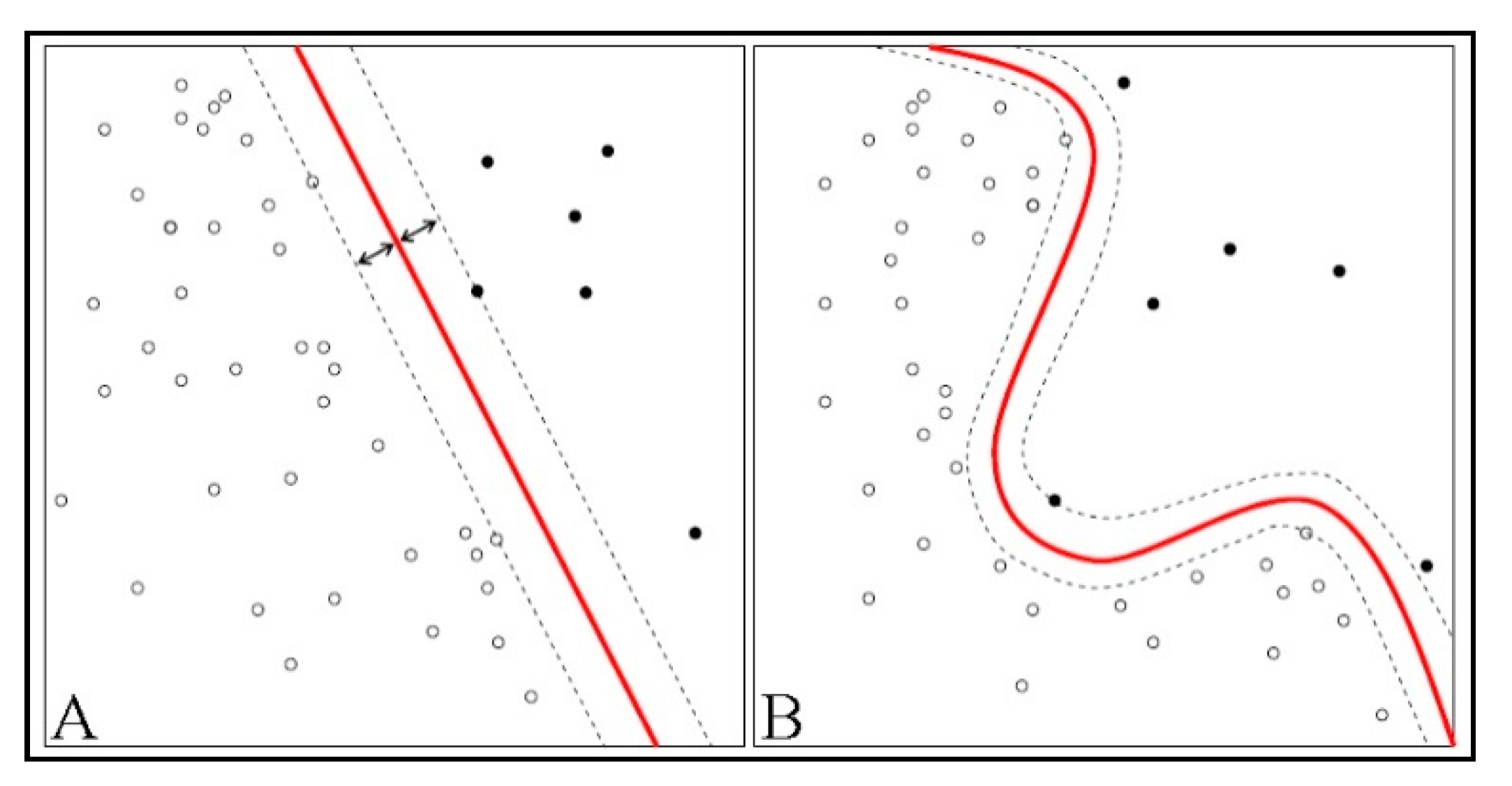
If the data are not separable by a straight line (a hyperplane)—as shown in
Figure 1B, above—a non-straight line separates the instances. This type of data is in nonlinear format, and the classifier for it is called nonlinear [
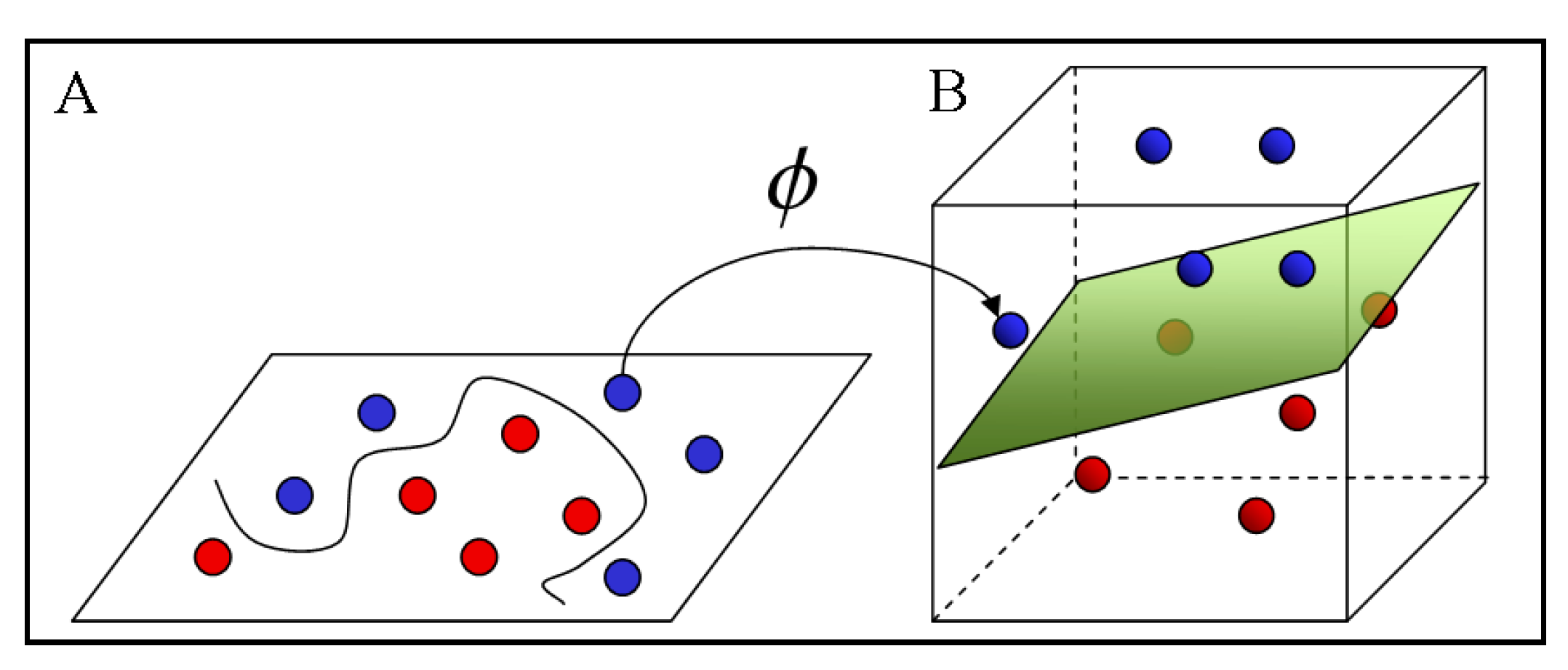
5]. The advantage of linear classifiers is that the training algorithm is simpler and it scales with the number of training instances. Linear classifiers, as illustrated in
Figure 2, can be converted into nonlinear ones by mapping the input data from space,
x, to the feature space, F, by means of a nonlinear function, ϕ. Equation (2) is the nonlinear discrimination function in feature space, F [
9].
When mapping analogies are applied to input vectors in the d-dimensional space, for feature space, F, the dimensionality is quadratic in d. This entails a quadratic increase in the memory storage required for each feature and in the time needed to compute the classifier’s discrimination function. For low data dimensions, this quadratic increase in complexity may be acceptable, but at higher data dimensions, it becomes problematic and, if higher degree monomials are used, completely unworkable [
7]. To overcome this issue, kernel methods are devised to avoid explicit mapping of the data into high-dimensional feature space [
5].
In 1963, Vapnik initially proposed a linear classifier, using a maximum-margin hyperplane algorithm [
3]. In 1964, Aizerman et al., for the first time, proposed the kernel method. Subsequently, in 1992, Boser et al. put forward the method of creating nonlinear classifiers by applying the kernel method to maximum-margin hyperplanes.
The strength of the kernel method is that it can convert any algorithm that is expressible in terms of dot products between two vectors into nonlinear format [
10]. A kernel function indicates an inner product within the feature space, and it is generally denoted as in Equation (3), below.
An applicable kernel function is symmetric and continuous, and it ideally has a positive definite Gram matrix: The kernel matrix holds only non-negative values. Algorithms which can operate with kernel functions include support vector machines (SVM), kernel perceptrons, Gaussian processes, canonical correlation analysis, principal component analysis (PCA), ridge regression, linear adaptive filters and spectral clustering [
10]. The main SVM kernel types are linear, polynomial, radial basis function and sigmoid kernels [
12,
13].
Due to the popularity of SVM, a substantial number of specially designed solvers has been developed for SVM optimization [
14]. Current software products for SVM training algorithms fall into two categories: (1) software for delivering SVM solver services, like LIBSVM [
15] and SVMlight [
16]; and (2) machine learning libraries that support various classification methods for essential tasks, like feature selection and preprocessing. Examples of some current popular SVM classifier products are Elefant [
17], The Spider [
18], Orange [
19], Weka [
20], Plearn [
21], Shogun [
22], Lush [
23], PyML [
24] and RapidMiner [
25].
2. Literature Review
Kernels are designed to project input data into feature space, aiming to achieve a hyperplane that efficiently separates data of different classes [
26]. Since this paper proposes an SVM kernel, we first review, below, relevant previous research on the SVM kernels currently in use, focusing on their description, the equations they employ, their applications and, finally, their advantages and disadvantages.
Linear kernels are the simplest kernel function, classifying data points in two classes by use of a straight separator line [
26]. Since the classification is carried out in a linear format, no mapping of data points to feature space is required. The function is the inner product (
x,
y), with the addition of an optional constant,
c, called bias [
26] (Equation (4)).
The main advantages of linear kernels are the quickness of their training and classification processes (since they do not use kernel operations); their low cost; the lower risk of overfitting than with nonlinear kernels; and the lower number of optimization parameters required than with nonlinear kernels [
3,
10]. Linear kernels can outperform nonlinear ones when the number of features is large relative to the number of training samples, and also when there is a small number of features but the training set is large. Against that, their main disadvantage is that, if the features are not linearly separable—by a hyperplane—nonlinear kernels, such as the Gaussian one, will usually produce better classifications [
3,
10].
Polynomial kernels are a commonly used kernel function with SVMs that enable the learning of nonlinear models. In the feature space, these characterize similarities between training input vectors and polynomials of the original variables [
3,
10]. In addition to features of input samples, polynomial kernels normally use a combination of features, which, for the purposes of regression analysis, are called interaction features [
3,
10].
Equation (5), below, denotes the polynomial kernel function when the degree of polynomials is N. The variables
x and
y are the input space vectors, and
c ≥ 0 is an influence tradeoff parameter between lower order and higher order polynomial terms. If
c = 0, then the kernel is homogeneous. Numerical instability is the problem of polynomial kernels. If
< 1, by increasing the value of N, the kernel function declines toward zero, while if
, it tends toward infinity.
A radial basis function (RBF), or Gaussian kernel, is a function which classifies data on a radial basis [
3,
10]. The separator function with two input space feature vectors of
x and
y is defined as in Equation (6), below.
is the squared Euclidean distance between the two input space feature vectors, while
is an optional value parameter. Equation (7) is a simplified version of the RBF kernel function that substitutes
with
. The function value decreases by distance, ranging between zero (within the limit) and 1 (if
x =
y).
How RBF kernels behave depends to a large extent on the selection of the gamma parameter. An overly large gamma can lead to overfitting, while a small gamma can constrain the model and render it unable to capture the shape or complexity of the data [
3,
10]. In addition, this type of kernel is robust against adversarial noise and in predictions. However, it has more limitations than neural networks [
27,
28].
Sigmoid kernels, also known as hyperbolic tangent and multilayer perceptron (MLP) kernels, are an SVM classifier inspired by neural networks. A bipolar sigmoid function is also used as an activation function for the artificial neurons [
3,
10]. In SVMs, sigmoid kernels are the equivalent of a two-layer perceptron neural network. A Sigmoid function kernel is shown in Equation (8), below.
The value of parameter
is generally
, with N representing the data dimension and
c the intercept constant. In certain ranges of
and
c, a sigmoid kernel behaves like an RBF. The operation
x.y is dot product between the two vectors. The application of Sigmoid kernels is similar to that of RBFs and depends on the chosen level of cross-validation [
27,
28]. It is an appropriate kernel to use particularly for nonlinear classification in two dimensions or when the number of dimensions is high.
The main advantages of sigmoid kernels are (1) the differentiability at all points of the domain; (2) the fast training process; and (3) presenting a choice of different nonlinearity levels by choosing the sophistication and amount in the membership function [
29]. The main drawback of sigmoid is their limited applicability and the fact that they only outperform RBFs in a limited number of cases [
29].
The four SVM kernels listed above are not the only ones. There is also a range of less-frequently used SVM kernels, including fisher, graph, string, tree, path, Fourier, B-spline, cosine, multiquadric, wave, log, cauchy, Tstudent, thin-plate and wavelet-SVM kernels (Harr, Daubechies, Coiflet and Symlet) [
30].
4. Fuzzy Weighted SVM Kernel
A number of statistical and probability functions and the related abbreviations are used in this research.
Table 1, below, lists and briefly describes these for clarity and consistency.
4.1. Principal Probability and Statistical Functions
In the theory of probability, normal or Gaussian distribution is defined as a continuous probability distribution that is based on the premise that data distribution will converge to the normal, especially in natural science, if a sufficiently large number of observations is made. As this normal distribution takes the form of a bell when plotted on a graph, it is also often informally called the bell curve.
Standard deviation (generally denoted as ) is a statistical function used to quantify the amount of variation or dispersion in a given dataset. Moreover, it is generally used to define the level of confidence in the accuracy of a given set of data. A high value for standard deviation means that the data are spread over a wide range, whereas a low value indicates that most of the data are close to the mean.
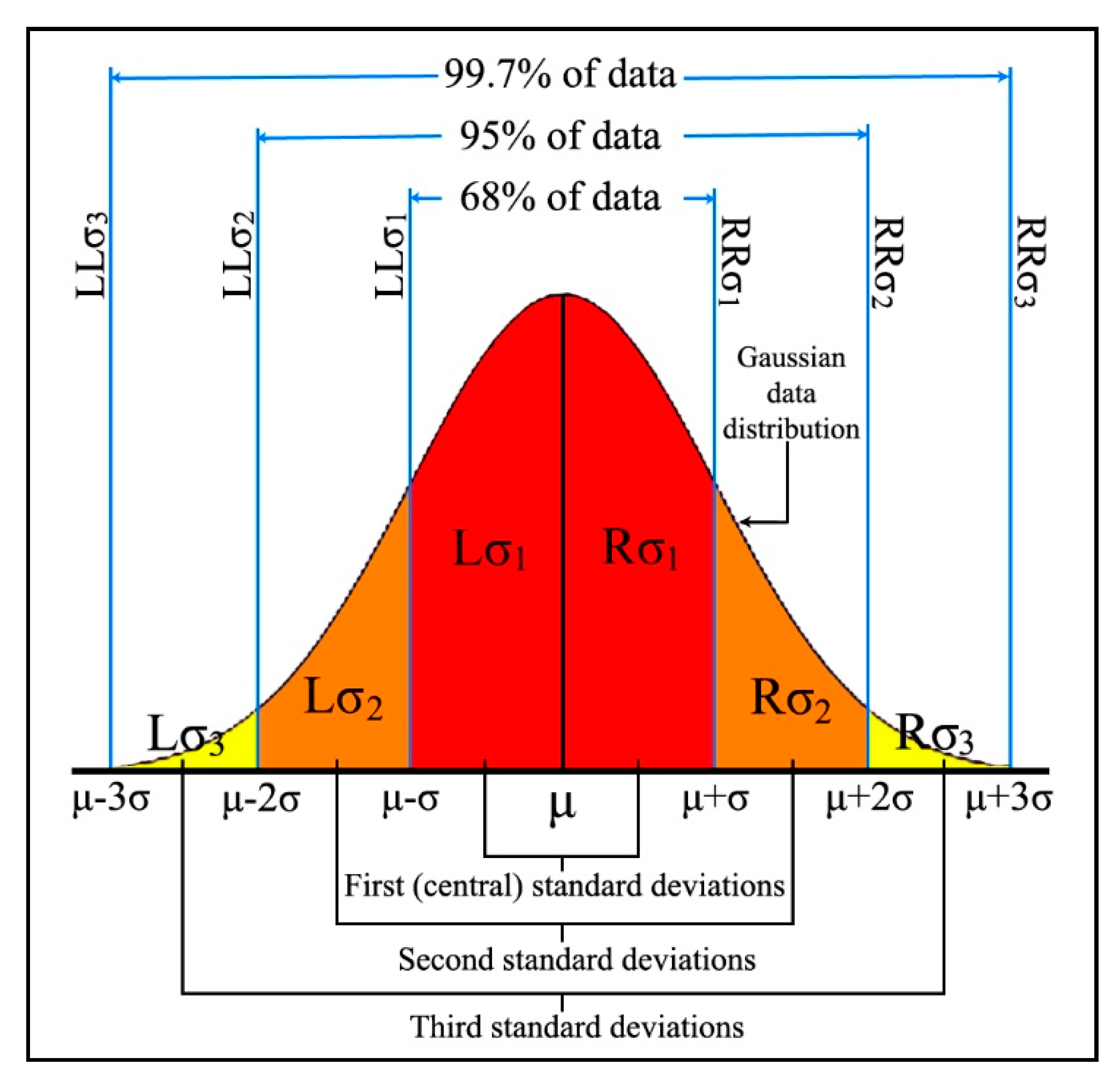
In statistics, the three-sigma, 68-95-99.7, or empirical rule, describes the density of data within standard deviation bands at both sides of the mean point. Specifically, in Gaussian distribution 68.27% (Equation (9)), 95.45% (Equation (10)) and 99.73% (Equation (11)) of the data values should be located within a distance of one, two and three standard deviation bands from both sides of the mean point, respectively.
4.2. Data Classification
In the proposed kernel of this research, the role of standard deviation is to quantify the Gaussian distributed data, while the three-sigma rule defines the importance of the data placed in each band. According to the three-sigma rule, the data located in the first standard deviation bands are the most reliable. However, reliability of data declines from the second standard deviation bands toward the third bands, and any data lying beyond the third bands are considered as noise and, accordingly, ignored.
Figure 3, below, presents an illustration of quantified Gaussian data distribution. The first, second and third standard deviation bands are highlighted in different colors, with the three right and left bands labeled as
,
and
and
,
and
respectively.
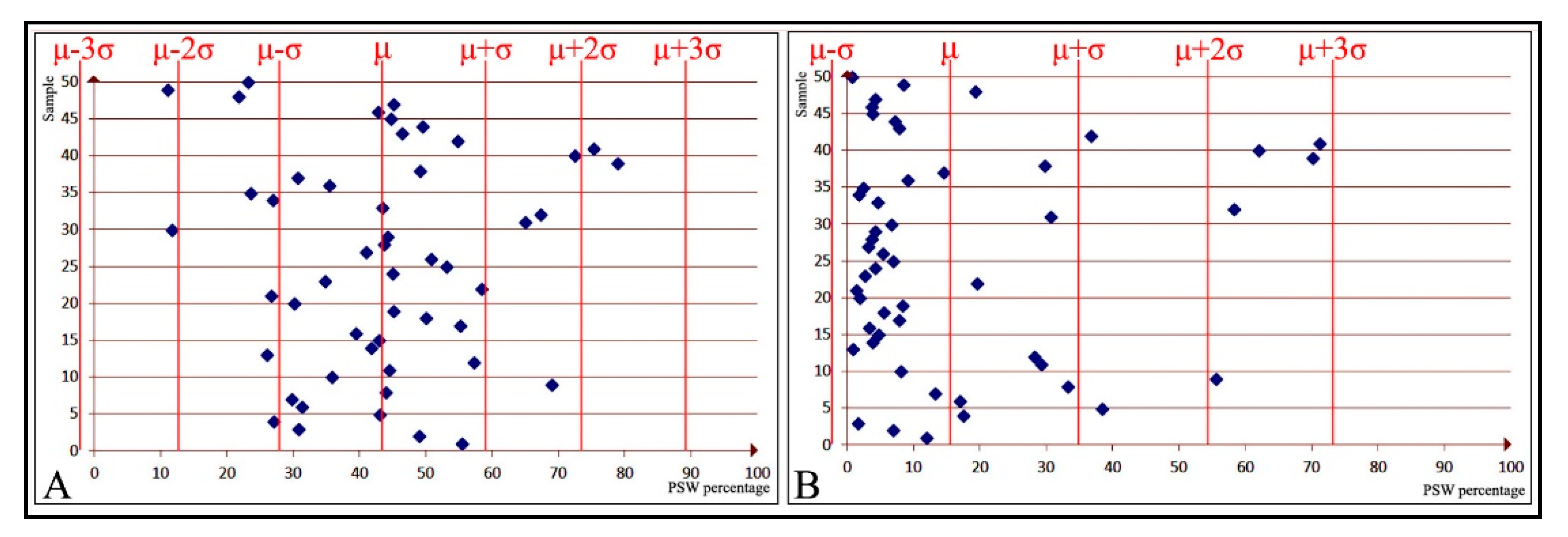
When data are normally distributed according to the three-sigma rule, the central (first) standard deviation bands hold the highest density, while this declines toward the outer bands. If, however, the data are abnormally distributed, they may not comply with the three-sigma rule: the data density in some of the bands might be higher or lower than the expected normal values, and some bands might not even exist.
Figure 4A, below, illustrates a normal data distribution, with a higher density within the
and
bands and a lower density in the more distant bands.
Figure 4B, on the other hand, displays an instance of abnormally distributed data: a high density within
, but the rest of the data dispersed only across the right bands. In
Figure 4B, not only do the
and
bands not exist, but the
border has had to be relocated to the point of 0. However, it should be noted that sometimes border relocation, as is the case with
in
Figure 4A, might be necessary, even if the data are overall normally distributed.
The criteria for calculating the left and right borders of the left and right standard deviation bands, respectively, are shown in Equations (12) and (13), below. The mean and max denote the need to relocate borders, while the word “Null” indicates that, due to abnormal data distribution, the band does not exist.
4.3. Membership Function
On the discoursed universe of X, the fuzzy membership function of set A is defined as , where the mapping values of X elements varies between 0 and 1. This membership degree quantifies the extent of membership of element X within fuzzy set A. In fuzzy classification, when the membership criteria resemble a polygon, the function is normally denoted as P(x) rather than .
In
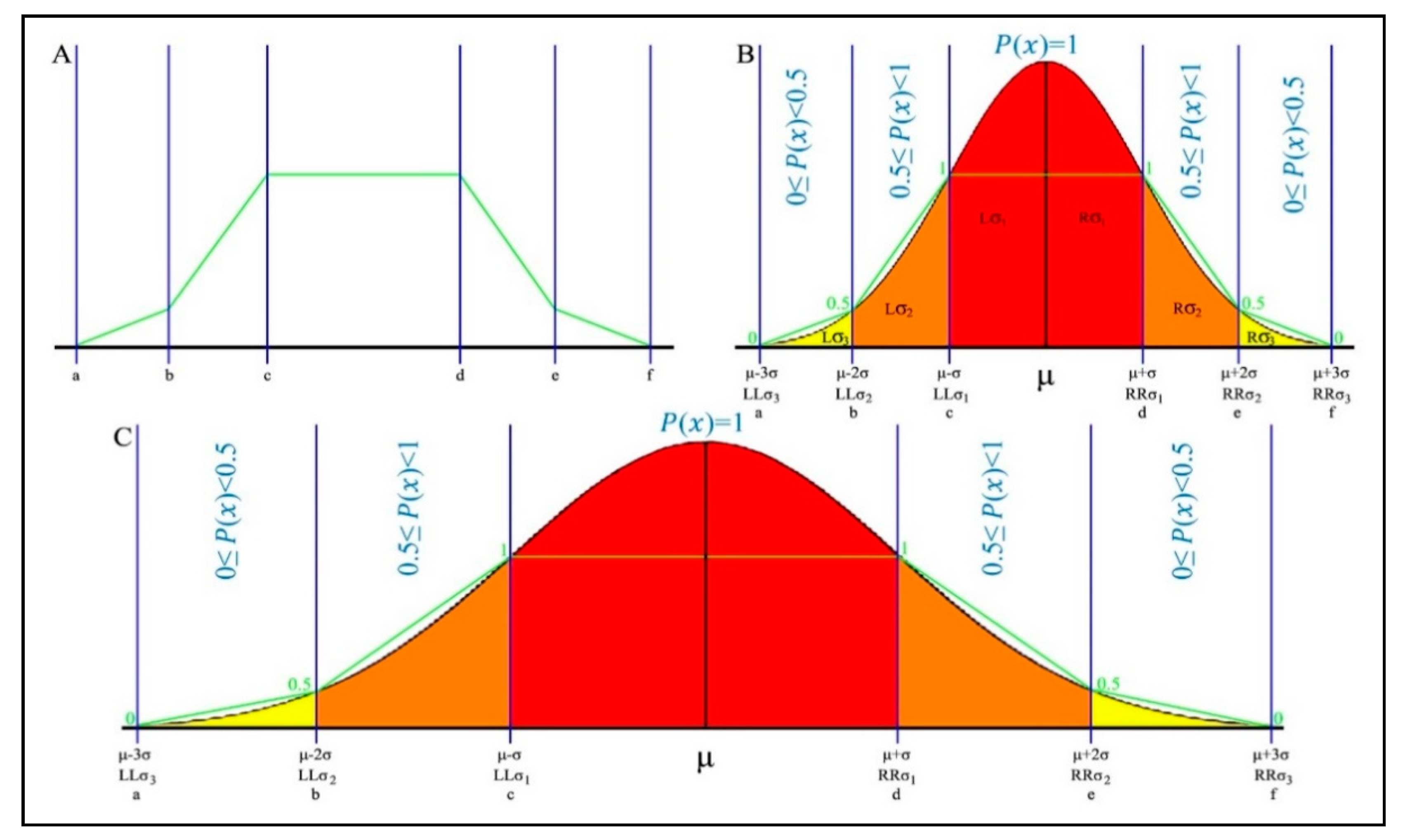
Figure 5A, above, the polygon’s limits are respectively labeled as a, b, c, d, e and f, where a < b < c < d < e < f. In
Figure 5B,C, the polygonal shape of section A is mapped onto the Gaussian distribution, which is segmented according to standard deviation and the three-sigma rule. This mapping is an essential part of the designed classifier. The central standard deviation bands, as can be seen in the above figure, hold the most reliable data and are the densest bands. The inflation points of c and d are mapped onto the boundaries of the central bands. The whole area between these points receives the full membership degree of 1, as it holds the most reliable data. The lower limit a and upper limit f are respectively mapped onto
and
, as the leftmost and rightmost borders of reliable data. The areas between
and
(a and c), as well as between
and
(d and f), are divided into two equal parts. The leftmost and rightmost bands (a to b and e to f) receive a membership degree equal or bigger than 0 and less than 0.5, as these hold the least reliable data, and the bands at sides of the central area (b to c and d to e) receive a membership degree equal or bigger than 0.5 and less than 1, as a medium-to-high reliable portion of data, depending on their precise location along the axis.
Figure 5B,C represents the short-tail and long-tail Gaussian data distributions, respectively. In both distributions, because of the three-sigma rule, the two left and the two right polygon sides follow slope of the distribution, resulting in the achievement of a more accurate fuzzy membership degree. This is designed to maximize classification accuracy. The criteria for calculating the polygonal fuzzy membership function are shown in Equation (14), below.
Based on the mapping inflation points of
Figure 5B,C, the equivalent values of the limits a, b, c, d, e and f are substituted in Equation (14) and then simplified to construct Equation (15).
However, data do not always appear in a single feature. In such circumstances, the membership degree is the average of the membership degrees of all relevant features. To support multidimensional features, Equation (15) is modified into Equation (16), below, which allows for the calculation of the membership degree of the given sample,
x, in the
ith feature of
k features.
The probability rule of sum is used when we calculate the probability of a union of events, by adding the distinct probabilities together. In other words, it gives the total probability of the final result when there are mutually exclusive events. The summation also could be interpreted as a weighted average—a value that is helpful for problem-solving. Using this rule, Equation (17), below, is the final equation which, along with Equation (16), calculates the normalized membership degree of the sample,
x, in
k features.
4.4. Noise Filtering
The designed classifier eliminates and mitigates natural data noise in two ways. First of all, it separates reliable data from noisy data, based on Gaussian data distribution and the three-sigma rule. In other words, it ignores any data beyond the reliable boundary. Second, it determines the influence of any given data in decisions, by computing their membership degree . In simple terms, data located in the central bands are more influential than data located in the side bands.
4.5. Reference Profiles
The PFW kernel records the statistical properties of all classes of the training data and stores these based on the structure presented in
Table 2, below. The number of profiles corresponds to the number of data classes, and these are then used to classify future given instances. The extracted statistics from each feature of each data group are divided into six standard deviation bands. L
and
(
≤
≤
) hold the most trustworthy data for the
ith feature, and hence the membership degree
(
) for these bands equals 1. The membership degree for
(
≤
<
) and
(
<
≤
) is 0.5 ≤
(
) < 1; and for the third bands,
(
≤
<
) and
(
<
≤
), the value is 0 ≤
(
) < 0.5. In cases where a band does not exist, because of abnormal data distribution, the associated value range in the profile is replaced with “Null”.
At classification time, PFW kernel calculates the membership degree of the given instance in comparison with each one of the profiles, according to Equation (17). Its class label is then detected based on the profile that achieves the highest membership degree.
6. Conclusions
Supervised learning is a class of machine learning in which a function is inferred from labeled training data. The inferred function then determines the class label of unseen instances. One type of supervised learning, SVM (support vector machine), predicts the categories of new examples, based on a given set of training examples. SVM maps the data into a space in which the categories are dividable by a clear gap. However, such mapping is not always sufficient to produce clearly separable categories. In such cases, a further step is required to transform the data from their original space to feature space, increasing the computational cost of training and classification.
Our PFW (polygonal fuzzy weighted) kernel is designed to map data accurately, without transforming them to feature space, even when groups are not linearly separable. The kernel accurately predicts the class of new instances in the original space, even if the groups have overlapping areas. It is structured based on Gaussian distribution, standard deviation, the three-sigma rule and a polygonal fuzzy membership function. PFW generates one profile for each data class, based on the statistics extracted during the training process. For the purposes of classification, the new instances are then compared with the reference profiles, based on a polygonal fuzzy membership function: The highest degree of membership achieved defines the prediction result.
To test the accuracy of our PFW kernel, we used it together with RBF and conventional linear kernels as classification engines for PSW steganalysis of the same set of 1333 images, whose classification feature data had overlapping data groups. The results showed that the PFW clearly outperformed the linear kernel in predicting the class labels of the images—yielding a minimum of 26% and as much as 40% higher classification accuracy. PFW kernel in two out of three image classes outperformed RBF kernel by 3% and 9%, while RBF delivered better results, by 1% in one class label.