Unlocking the Potential of Quantum Machine Learning to Advance Drug Discovery
Abstract
1. Introduction
- Which QML models are used in drug discovery?
- Are QML versus ML algorithms more efficient in terms of time and accuracy in drug discovery?
2. Preliminaries—Background
2.1. Drug Discovery
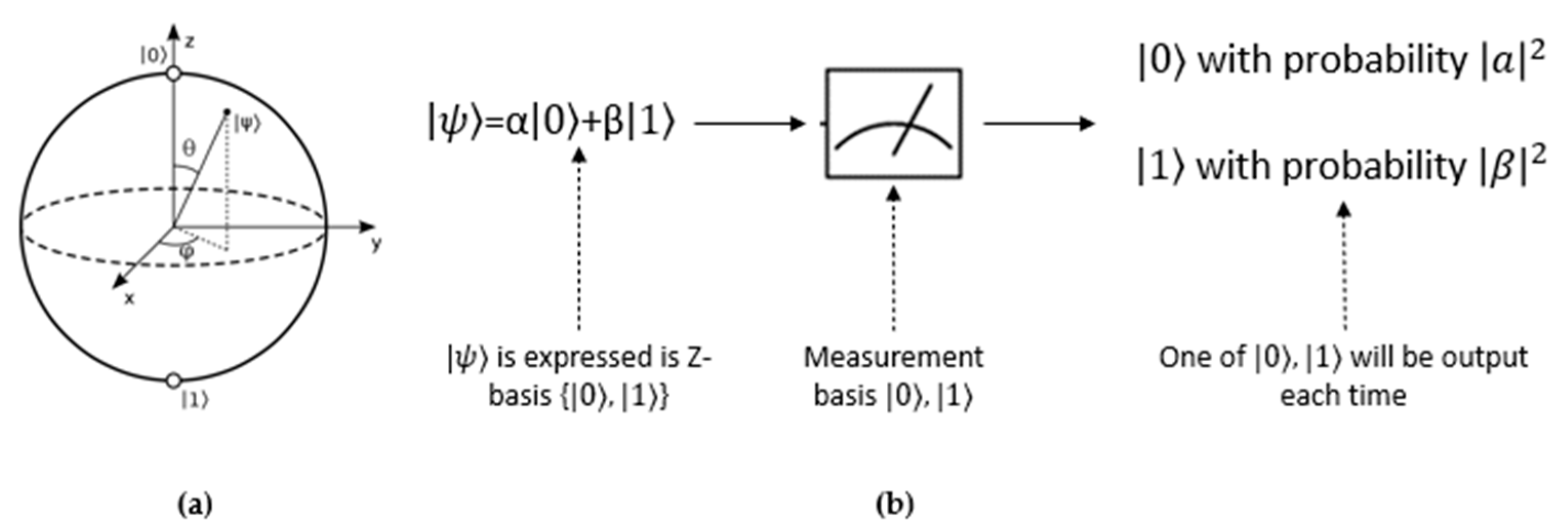
2.2. Quantum Computing
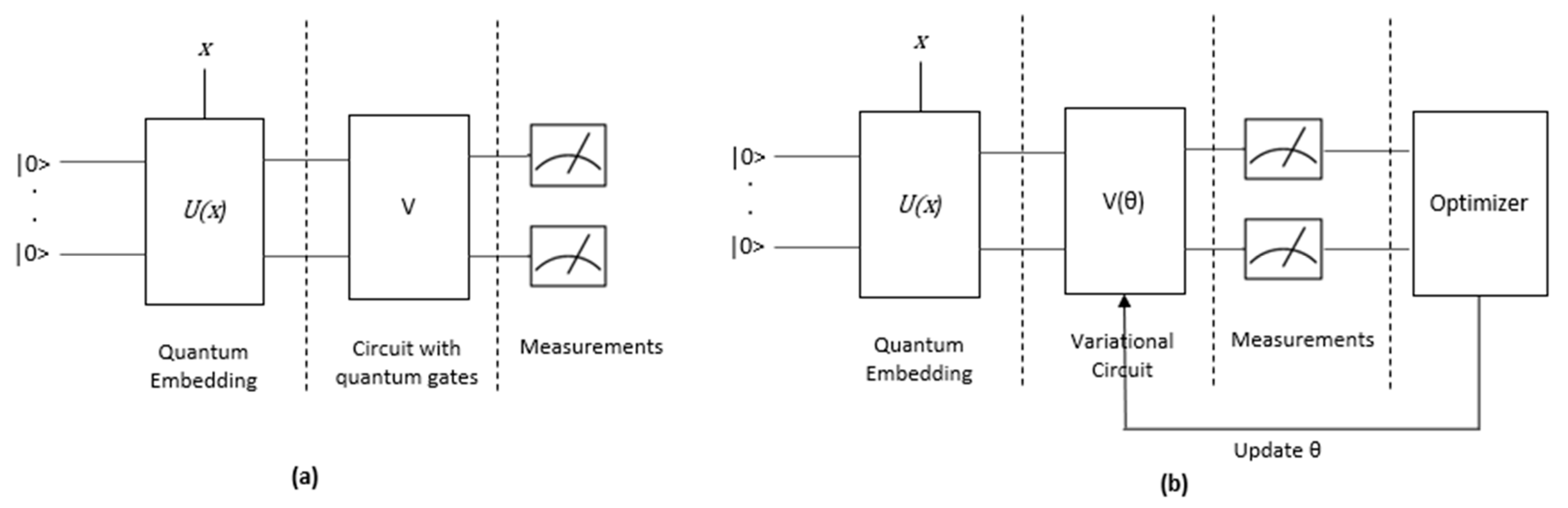
2.3. Quantum Machine Learning (QML)
2.4. Quantum Machine Learning (QML) Algorithms
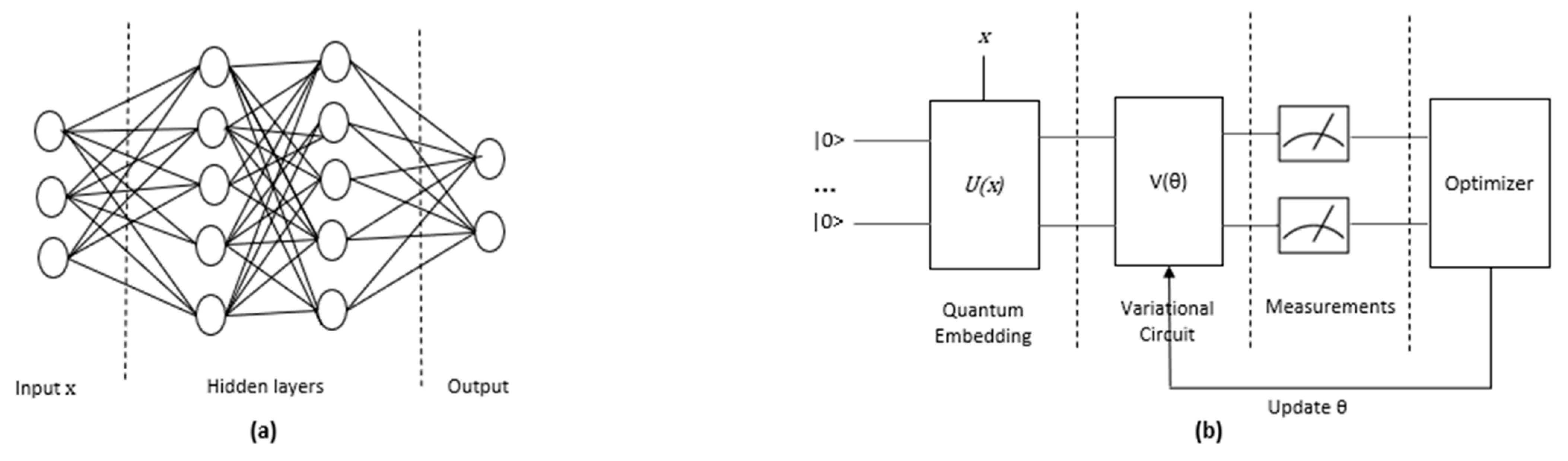
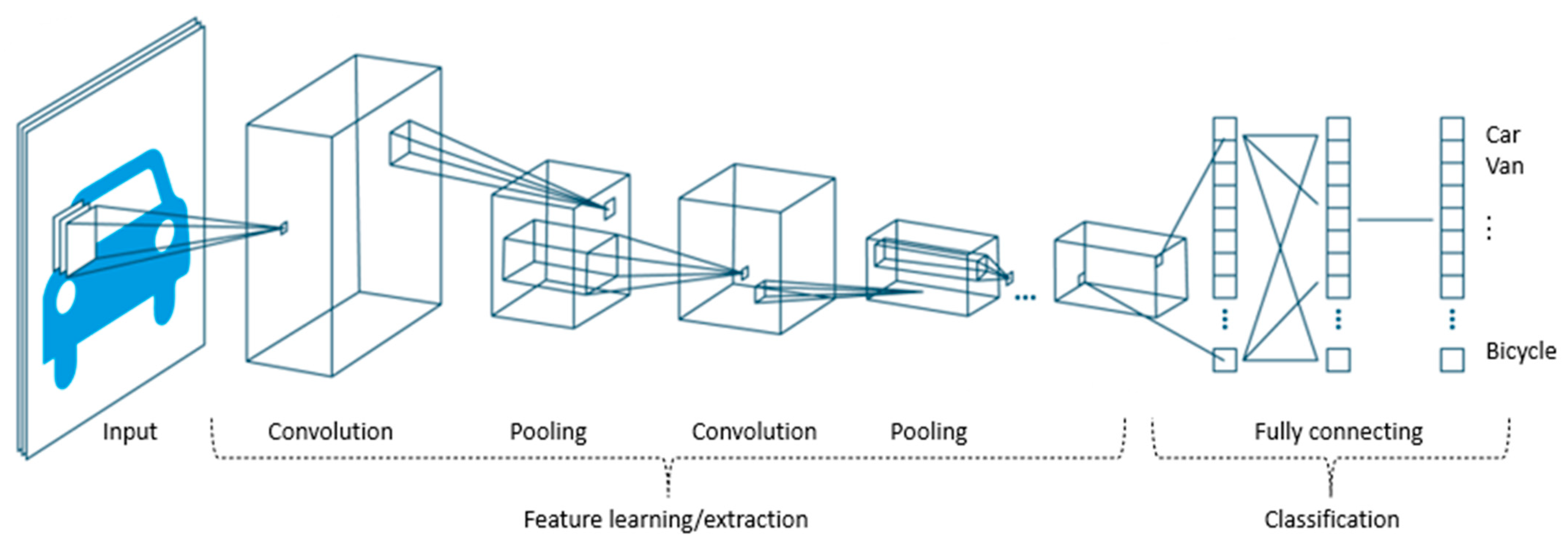
2.4.1. QNN
2.4.2. QGAN
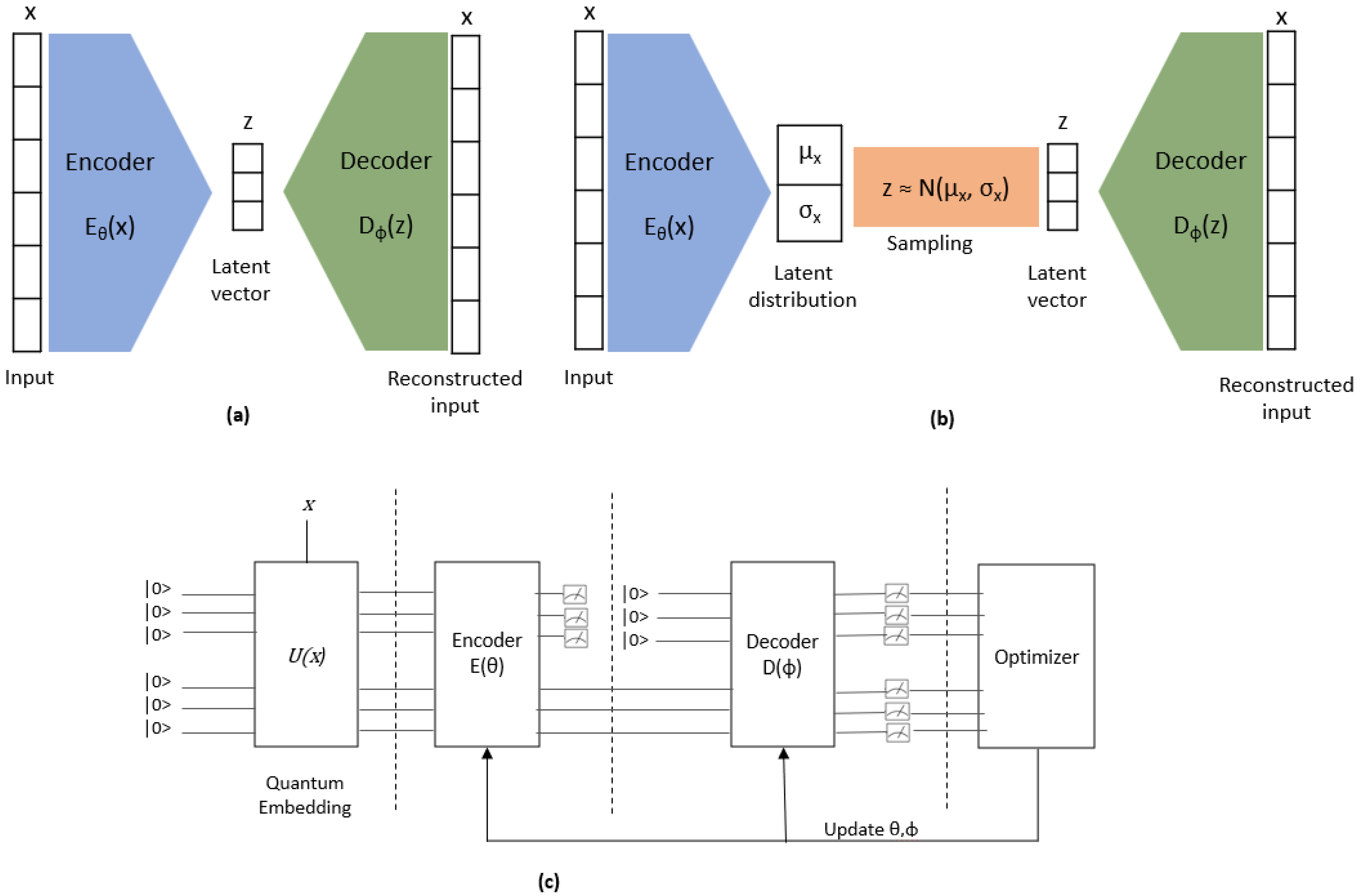
2.4.3. QVAE/AE
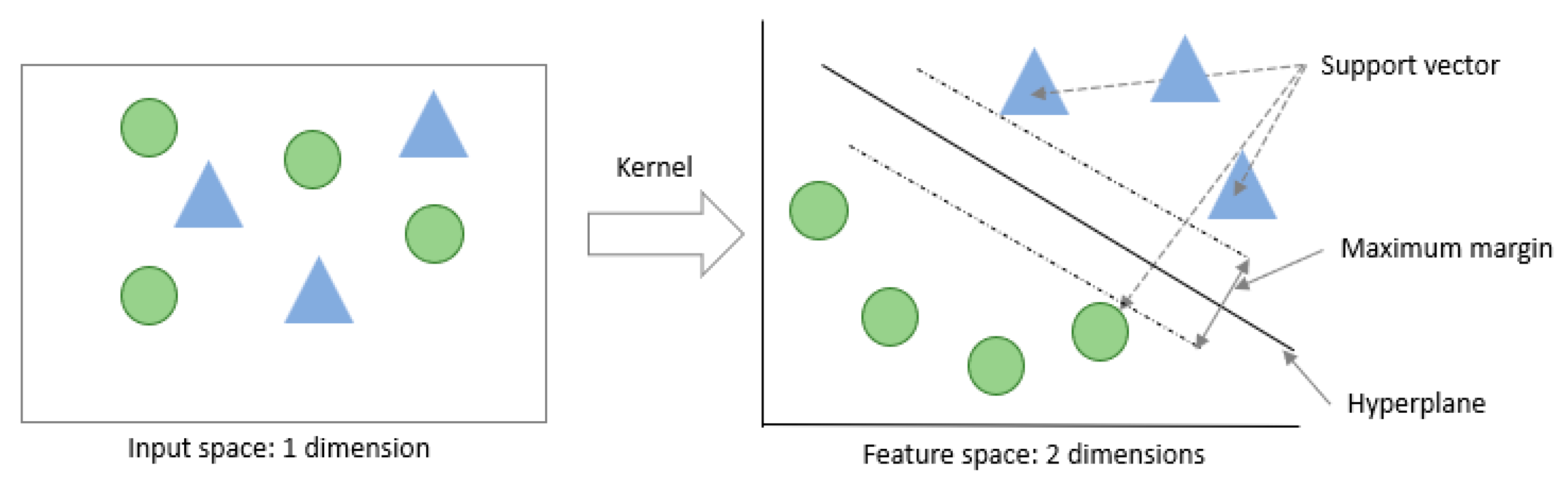
2.4.4. QSVM
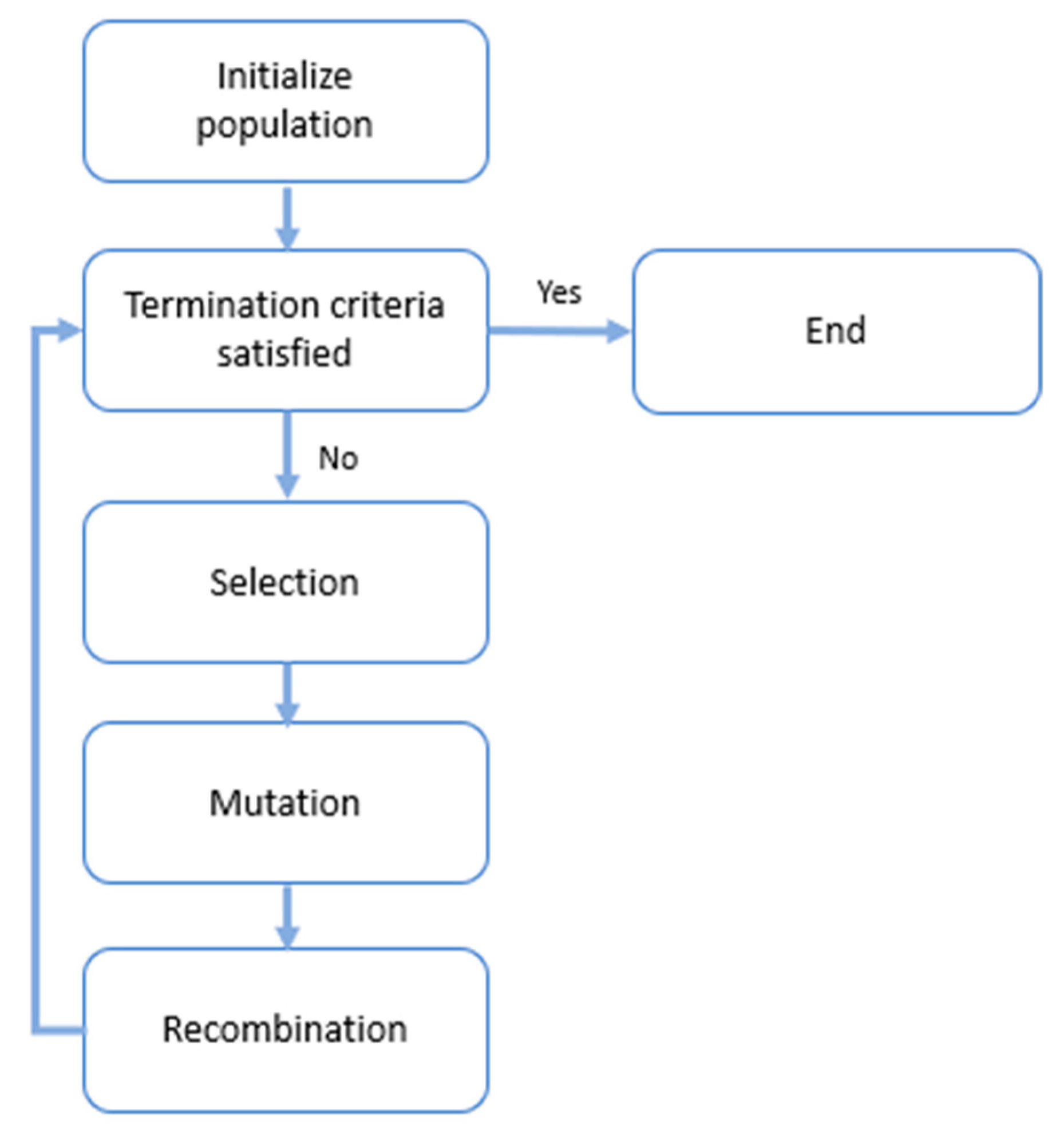
2.4.5. Quantum Genetic Algorithms
2.4.6. Quantum Linear and No-Linear Regression
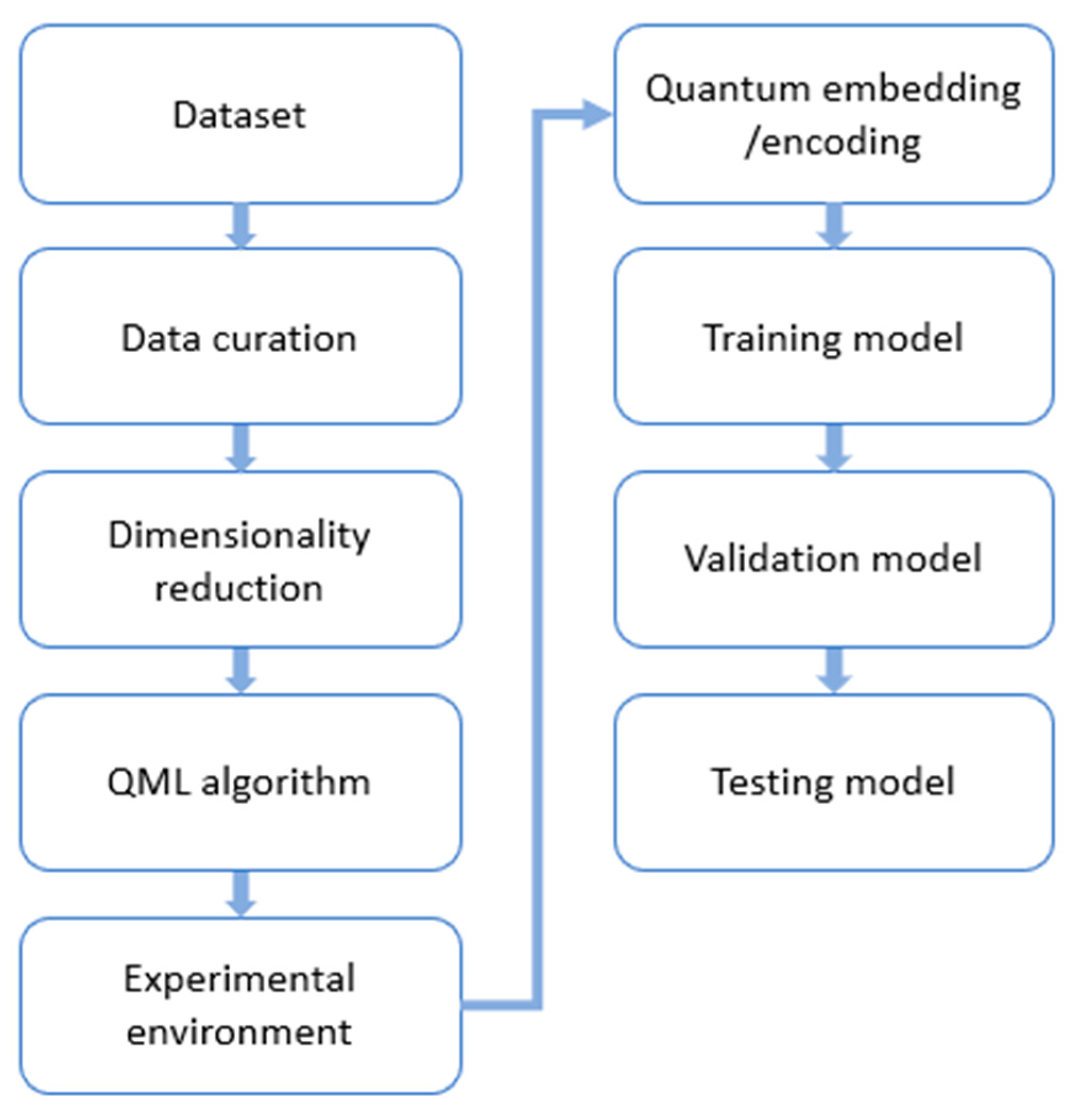
3. Methodology
- “quantum machine learning” AND (“drug discovery” OR “drug design” OR “drug development”)
4. QML Applications in Drug Discovery
4.1. QNN
4.2. QGAN
4.3. QVAE/QAE
4.4. QSVM
4.5. Quantum Genetic Algorithms
4.6. Quantum Linear and No-Linear Regression
5. Discussion
6. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Carracedo-Reboredo, P.; Liñares-Blanco, J.; Rodríguez-Fernández, N.; Cedrón, F.; Novoa, F.J.; Carballal, A.; Maojo, V.; Pazos, A.; Fernandez-Lozano, C. A review on machine learning approaches and trends in drug discovery. Comput. Struct. Biotechnol. J. 2021, 19, 4538–4558. [Google Scholar] [CrossRef]
- Sliwoski, G.; Kothiwale, S.; Meiler, J.; Lowe, E.W. Computational methods in drug discovery. Pharmacol. Res. 2014, 66, 334–395. [Google Scholar] [CrossRef]
- Zinner, M.; Dahlhausen, F.; Boehme, P.; Ehlers, J.; Bieske, L.; Fehring, L. Quantum computing’s potential for drug discovery: Early stage industry dynamics. Drug Discov. Today 2021, 26, 1680–1688. [Google Scholar] [CrossRef]
- Zinner, M.; Dahlhausen, F.; Boehme, P.; Ehlers, J.; Bieske, L.; Fehring, L. Toward the institutionalization of quantum computing in pharmaceutical research. Drug Discov. Today 2022, 27, 378–383. [Google Scholar] [CrossRef]
- Feynman, R.P. Quantum mechanical computers. Found. Phys. 1986, 16, 507–532. [Google Scholar] [CrossRef]
- Lloyd, S. Universal quantum simulators. Science 1996, 273, 1073–1078. [Google Scholar] [CrossRef]
- Cao, Y.; Romero, J.; Aspuru-Guzik, A. Potential of quantum computing for drug discovery. IBM J. Res. Dev. 2018, 62, 6:1–6:20. [Google Scholar] [CrossRef]
- Sajeev, V.; Vyshnavi, A.H.; Namboori, P.K. Thyroid Cancer Prediction Using Gene Expression Profile, Pharmacogenomic Variants and Quantum Image Processing in Deep Learning Platform-A Theranostic Approach. In Proceedings of the 2020 International Conference for Emerging Technology, Belgaum, India, 5–7 June 2020; pp. 1–5. [Google Scholar]
- Li, T.Y.; Mekala, V.R.; Ng, K.L.; Su, C.F. Classification of Tumor Metastasis Data by Using Quantum kernel-based Algorithms. In Proceedings of the IEEE 22nd International Conference on Bioinformatics and Bioengineering, Taichung, Taiwan, 7–9 November 2022; pp. 351–354. [Google Scholar]
- Houssein, E.H.; Abohashima, Z.; Elhoseny, M.; Mohamed, W.M. Hybrid quantum-classical convolutional neural network model for COVID-19 prediction using chest X-ray images. J. Comput. Des. Eng. 2022, 9, 343–363. [Google Scholar] [CrossRef]
- Shahwar, T.; Zafar, J.; Almogren, A.; Zafar, H.; Rehman, A.U.; Shafiq, M.; Hamam, H. Automated detection of Alzheimer’s via hybrid classical quantum neural networks. Electronics 2022, 11, 721. [Google Scholar] [CrossRef]
- Alam, M.; Ghosh, S. Qnet: A scalable and noise-resilient quantum neural network architecture for noisy intermediate-scale quantum computers. Front. Phys. 2022, 9, 702. [Google Scholar] [CrossRef]
- Amin, J.; Sharif, M.; Fernandes, S.L.; Wang, S.H.; Saba, T.; Khan, A.R. Breast microscopic cancer segmentation and classification using unique 4-qubit-quantum model. Microsc. Res. Technol. 2022, 85, 1926–1936. [Google Scholar] [CrossRef] [PubMed]
- Ullah, U.; Maheshwari, D.; Gloyna, H.H.; Garcia-Zapirain, B. Severity Classification of COVID-19 Patients Data using Quantum Machine Learning Approaches. In Proceedings of the 2022 International Conference on Electrical, Computer, Communications and Mechatronics Engineering, Male, Malvides, 16–18 November 2022; pp. 1–6. [Google Scholar]
- Sengupta, K.; Srivastava, P.R. Quantum algorithm for quicker clinical prognostic analysis: An application and experimental study using CT scan images of COVID-19 patients. BMC Med. Inform. Decis. Mak. 2021, 21, 227. [Google Scholar] [CrossRef] [PubMed]
- Kumar, Y.; Koul, A.; Sisodia, P.S.; Shafi, J.; Kavita, V.; Gheisari, M.; Davoodi, M.B. Heart failure detection using quantum-enhanced machine learning and traditional machine learning techniques for internet of artificially intelligent medical things. Wirel. Commun. Mob. Comput. 2021, 2021, 1616725. [Google Scholar] [CrossRef]
- Robert, A.; Barkoutsos, P.K.; Woerner, S.; Tavernelli, I. Resource-efficient quantum algorithm for protein folding. npj Quantum Inf. 2021, 7, 38. [Google Scholar] [CrossRef]
- Outeiral, C.; Strahm, M.; Shi, J.; Morris, G.M.; Benjamin, S.C.; Deane, C.M. The prospects of quantum computing in computational molecular biology. Wiley Interdiscip. Rev. Comput. Mol. Sci. 2021, 11, e1481. [Google Scholar] [CrossRef]
- Von Lilienfeld, O.A.; Müller, K.R.; Tkatchenko, A. Exploring chemical compound space with quantum-based machine learning. Nat. Rev. Chem. 2020, 4, 347–358. [Google Scholar] [CrossRef]
- Singh, J.; Bhangu, K.S. Contemporary Quantum Computing Use Cases: Taxonomy, Review and Challenges. Arch. Comput. Methods Eng. 2023, 30, 615–638. [Google Scholar] [CrossRef]
- Cordier, B.A.; Sawaya, N.P.; Guerreschi, G.G.; McWeeney, S.K. Biology and medicine in the landscape of quantum advantages. J. R. Soc. Interface 2022, 19, 20220541. [Google Scholar] [CrossRef]
- Marchetti, L.; Nifosì, R.; Martelli, P.L.; Da Pozzo, E.; Cappello, V.; Banterle, F.; Trincavelli, M.L.; Martini, C.; D’Elia, M. Quantum computing algorithms: Getting closer to critical problems in computational biology. Brief. Bioinform. 2022, 23, bbac437. [Google Scholar] [CrossRef]
- Sajjan, M.; Li, J.; Selvarajan, R.; Sureshbabu, S.H.; Kale, S.S.; Gupta, R.; Singh, V.; Kais, S. Quantum machine learning for chemistry and physics. Chem. Soc. Rev. 2022, 51, 6475–6573. [Google Scholar] [CrossRef]
- McArdle, S.; Endo, S.; Aspuru-Guzik, A.; Benjamin, S.C.; Yuan, X. Quantum computational chemistry. Rev. Mod. Phys. 2020, 92, 015003. [Google Scholar] [CrossRef]
- Avramouli, M.; Savvas, I.; Vasilaki, A.; Garani, G.; Xenakis, A. Quantum Machine Learning in Drug Discovery: Current State and Challenges. In Proceedings of the 26th Pan-Hellenic Conference on Informatics, Athens, Greece, 25–27 November 2022; pp. 394–401. [Google Scholar]
- Singh, D.B. (Ed.) Computer-Aided Drug Design; Springer: Singapore, 2020. [Google Scholar]
- Murray, C.W.; Verdonk, M.L.; Rees, D.C. Experiences in fragment-based drug discovery. Trends Pharmacol. Sci. 2012, 33, 224–232. [Google Scholar] [CrossRef] [PubMed]
- Preskill, J. Quantum computing in the NISQ era and beyond. Quantum 2018, 2, 79. [Google Scholar] [CrossRef]
- Cerezo, M.; Arrasmith, A.; Babbush, R.; Benjamin, S.C.; Endo, S.; Fujii, K.; McClean, J.R.; Mitarai, K.; Yuan, X.; Cincio, L.; et al. Variational quantum algorithms. Nat. Rev. Phys. 2021, 3, 625–644. [Google Scholar] [CrossRef]
- Mensa, S.; Sahin, E.; Tacchino, F.; Barkoutsos, P.K.; Tavernelli, I. Quantum machine learning framework for virtual screening in drug discovery: A prospective quantum advantage. Mach. Learn. Sci. Technol. 2023, 4, 015023. [Google Scholar] [CrossRef]
- Beaudoin, C.; Kundu, S.; Topaloglu, R.O.; Ghosh, S. Quantum Machine Learning for Material Synthesis and Hardware Security. In Proceedings of the 41st IEEE/ACM International Conference on Computer-Aided Design, San Diego, CA, USA, 30 October–3 November 2022; pp. 1–7. [Google Scholar]
- Batra, K.; Zorn, K.M.; Foil, D.H.; Minerali, E.; Gawriljuk, V.O.; Lane, T.R.; Ekins, S. Quantum machine learning algorithms for drug discovery applications. J. Chem. Inf. Model 2021, 61, 2641–2647. [Google Scholar] [CrossRef]
- Lim, M.A.; Yang, S.; Mai, H.; Cheng, A.C. Exploring deep learning of quantum chemical properties for absorption, distribution, metabolism, and excretion predictions. J. Chem. Inf. Model. 2022, 62, 6336–6341. [Google Scholar] [CrossRef]
- Isert, C.; Atz, K.; Jiménez-Luna, J.; Schneider, G. QMugs, quantum mechanical properties of drug-like molecules. Sci. Data 2022, 9, 273. [Google Scholar] [CrossRef]
- Reddy, P.; Bhattacherjee, A.B. A hybrid quantum regression model for the prediction of molecular atomization energies. Mach. Learn. Sci. Technol. 2021, 2, 025019. [Google Scholar] [CrossRef]
- Li, J.; Ghosh, S. Scalable variational quantum circuits for autoencoder-based drug discovery. In Proceedings of the 2022 Design, Automation and Test in Europe Conference and Exhibition (DATE), Antwerp, Belgium, 14–23 March 2022; pp. 340–345. [Google Scholar]
- Li, J.; Alam, M.; Congzhou, M.S.; Wang, J.; Dokholyan, N.V.; Ghosh, S. Drug discovery approaches using quantum machine learning. In Proceedings of the 58th ACM/IEEE Design Automation Conference, San Francisco, CA, USA, 5–9 December 2021; pp. 1356–1359. [Google Scholar]
- Suzuki, T.; Katouda, M. Predicting toxicity by quantum machine learning. J. Phys. Commun. 2020, 4, 125012. [Google Scholar] [CrossRef]
- Li, J.; Topaloglu, R.O.; Ghosh, S. Quantum generative models for small molecule drug discovery. IEEE Trans. Autom. Sci. Eng. 2021, 2, 3103308. [Google Scholar] [CrossRef]
- Darwish, S.M.; Shendi, T.A.; Younes, A. Chemometrics approach for the prediction of chemical compounds’ toxicity degree based on quantum inspired optimization with applications in drug discovery. Chemometr. Intell. Lab. Syst. 2019, 193, 103826. [Google Scholar] [CrossRef]
- Jain, S.; Ziauddin, J.; Leonchyk, P.; Yenkanchi, S.; Geraci, J. Quantum and classical machine learning for the classification of non-small-cell lung cancer patients. SN Appl. Sci. 2020, 2, 1088. [Google Scholar] [CrossRef]
- Khan, T.M.; Robles-Kelly, A. Machine learning: Quantum vs. classical. IEEE Access 2020, 8, 219275–219294. [Google Scholar] [CrossRef]
- Tacchino, F.; Macchiavello, C.; Gerace, D.; Bajoni, D. An artificial neuron implemented on an actual quantum processor. npj Quantum Inf. 2019, 5, 26. [Google Scholar] [CrossRef]
- Zhao, J.; Zhang, Y.H.; Shao, C.P.; Wu, Y.C.; Guo, G.C.; Guo, G.P. Building quantum neural networks based on a swap test. Phys. Rev. A 2019, 100, 012334. [Google Scholar] [CrossRef]
- Zhao, C.; Gao, X.S. Qdnn: Deep neural networks with quantum layers. Quantum Mach. Intell. 2021, 3, 15. [Google Scholar] [CrossRef]
- Cong, I.; Choi, S.; Lukin, M.D. Quantum convolutional neural networks. Nat. Phys. 2019, 15, 1273–1278. [Google Scholar] [CrossRef]
- Henderson, M.; Shakya, S.; Pradhan, S.; Cook, T. Quanvolutional neural networks: Powering image recognition with quantum circuits. Quantum Mach. Intell. 2020, 2, 2. [Google Scholar] [CrossRef]
- Chen, S.Y.; Yoo, S.; Fang, Y.L. Quantum long short-term memory. In Proceedings of the 2022 IEEE International Conference on Acoustics, Speech and Signal Processing, Singapore, 22–27 May 2022; pp. 8622–8626. [Google Scholar]
- Amin, M.H.; Andriyash, E.; Rolfe, J.; Kulchytskyy, B.; Melko, R. Quantum boltzmann machine. Phys. Rev. X 2018, 8, 021050. [Google Scholar] [CrossRef]
- Ngo, T.A.; Nguyen, T.; Thang, T.C. A Survey of Recent Advances in Quantum Generative Adversarial Networks. Electronics 2023, 12, 856. [Google Scholar] [CrossRef]
- Romero, J.; Olson, J.P.; Aspuru-Guzik, A. Quantum autoencoders for efficient compression of quantum data. Quantum Sci. Technol. 2017, 2, 045001. [Google Scholar] [CrossRef]
- Khoshaman, A.; Vinci, W.; Denis, B.; Andriyash, E.; Sadeghi, H.; Amin, M.H. Quantum variational autoencoder. Quantum Sci. Technol. 2018, 4, 014001. [Google Scholar] [CrossRef]
- Rebentrost, P.; Mohseni, M.; Lloyd, S. Quantum support vector machine for big data classification. Phys. Rev. Lett. 2014, 113, 130503. [Google Scholar] [CrossRef]
- Lahoz-Beltra, R. Quantum genetic algorithms for computer scientists. Computers 2016, 5, 24. [Google Scholar] [CrossRef]
- Moher, D.; Shamseer, L.; Clarke, M.; Ghersi, D.; Liberati, A.; Petticrew, M.; Shekelle, P.; Stewart, L.A. Preferred reporting items for systematic review and meta-analysis protocols (PRISMA-P) 2015 statement. Syst. Rev. 2015, 4, 1. [Google Scholar] [CrossRef] [PubMed]
- Lau, B.; Emani, P.S.; Chapman, J.; Yao, L.; Lam, T.; Merrill, P.; Warrell, J.; Gerstein, M.B.; Lam, H.Y. Insights from incorporating quantum computing into drug design workflows. Bioinformatics 2023, 39, btac789. [Google Scholar] [CrossRef]
- Wang, X.; Wang, X.; Zhang, S. Adverse Drug Reaction Detection from Social Media Based on Quantum Bi-LSTM with Attention. IEEE Access 2022, 11, 16194–16202. [Google Scholar] [CrossRef]
- Smith, E.A.; Horan, W.P.; Demolle, D.; Schueler, P.; Fu, D.J.; Anderson, A.E.; Geraci, J.; Butlen-Ducuing, F.; Link, J.; Khin, N.A.; et al. Using Artificial Intelligence-based Methods to Address the Placebo Response in Clinical Trials. Innov. Clin. Neurosci. 2022, 19, 60–70. [Google Scholar]
- Ganesh, R. Computational identification of inhibitors of MSUT-2 using quantum machine learning and molecular docking for the treatment of Alzheimer’s disease. Alzheimers Dement. 2021, 17, 1. [Google Scholar] [CrossRef]




| NN | LSTM | QBi-LSTM | LNS, CNNWEF, RNN, and ATT-RNN | |
|---|---|---|---|---|
| Weighted mQNN, cQNN, and qisQNN | [56] [≈] Precision [GenoDock, Platinum]/ [Rigetti, ibmq_bogota, PennyLane’s, Simulation]/ [AUC, F1, Sensitivity, Precision] | |||
| QLSTM | [31] [−] Training accuracy [31] [+] Testing accuracy [USPTO-50k]/ [simulation PannyLane]/ [accuracy, Loss] | |||
| QBi-LSTMA | [57] [+] Performance [TwiMed, TwitterADR]/ [simulation]/ [Precision, Recall rate, F1-score] | [57] [+] Performance [TwiMed, TwitterADR]/ [simulation]/ [Precision, Recall rate, F1-score] | [57] [+] Accuracy [57] [+] Training time [TwiMed, TwitterADR]/ [simulation]/ [Precision, Recall rate, F1-score] |
| RBFNN | H-QFT-Based Hybrid QNN | Single-Layer CNN + MLP | Linear Regression/Logistic Regression/SVM Random Forest/XGBoost/Neural Network | |
|---|---|---|---|---|
| Q-RBFNN | [35] [≈] Accuracy [QM7]/ [simulation with noise]/ [MAE, RMSE, R2, Pearson correlation] | |||
| H-QNN | [35] [−] Accuracy [QM7]/ [simulation with noise]/ [L1-Loss] | |||
| single-layer QuanNN + MLP | [37] [+] Accuracy [TOUGH-C1]/ [simulation]/ [Cross Entropy] | |||
| QRBM | [41] [≈] Accuracy [Kuner,Golumbic]/ [D-Wave]/ [accuracy] |
| MolGAN | QGAN-HG | |
|---|---|---|
| QGAN-HG | [37,39] [+] Training epochs [37,39] [−] Training time [QM9, moderately reduced parameters]/ [Simulation,ibm_quito]/ [Fréchet Distance/training epochs/time] | |
| QGAN-HG | [37,39] [−] Training epochs [37,39] [−] Training time [QM9, highly reduced parameters]/ [Simulation, ibm_quito]/ [Fréchet Distance/training epochs/time] | |
| P-QGAN-HG | [39] [+] Learning accuracy [39] [+] Training time [37,39] [+] Training epochs [QM9, highly reduced parameters]/ [Simulation,ibm_quito]/ [Fréchet Distance/training epochs/time] |
| VAE/AE | VAE | |
|---|---|---|
| BQ-VAE/AE | [36] [−] Accuracy [36] [≈] time training (epochs) [QM9] [non-normalized molecule]/ [simulation]/ [ accuracy, time training (epochs)] | |
| BQ-VAE/AE | [36] [−] Accuracy [36] [+] time training (epochs) [QM9] [normalized molecule]/ [simulation]/ [accuracy, time training (epochs)] | |
| SQ-VAE/AE | [36] [+] reconstruction accuracy [36] [+] sampling accuracy [PDBbind]/ [simulation]/ [accuracy, time training (epochs)] | |
| Hybrid QVAE | [37] [−] Time learning [TOURCH1]/ [simulation]/ [L2 Loss, time] |
| SVM | H-QSVM | Data Re-Uploading Classifier on CC | |
|---|---|---|---|
| QSVM | [32] [−] Accuracy [SARS-CoV-2 (Vero cell), small dataset]/ [ibmq_rochester]/ [Accuracy] | [32] [+] Accuracy [SARS-CoV-2 (Vero cell), small dataset]/ [ibmq_rochester]/ [Accuracy] | |
| QSVM | [30] [+] ROC in cases [LIT-PCBA, COVID-19]/ [simulation]/ [ROC] [30] [+] ROC in cases [ADRB2, COVID-19]/ [IBM Quantum Montreal, IBM Quantum Guadalupe]/ [ROC] | ||
| Data Re-uploading Classifier on QC | [32] [−] Accuracy [32] [+] Time [M. tuberculosis Inhibition -small dataset]/ [ ibmq_rochester]/ [accuracy, run time] [32] [−] Accuracy [32] [+] Time [cathepsin B(Pubchem)/ Krabbe disease (Pubchem)/ plague (Pubchem)/ M. tuberculosis (Pubchem)/ hERG]/ [simulation]/ [accuracy, run time] |
| CGP and RBF-NN | |
|---|---|
| QIGP | [40] [+] Accuracy [40] [+] Τime [221 phenols]/ [simulation]/ [R, cosine similarities, MSE] |
| Classical Linear Regression | MLR/RBF-NN | |
|---|---|---|
| Q Linear Regression | [35] [≈]Accuracy [QM7]/ [simulation with noise]/ [MAE, RMSE, R2, Pearson correlation] | |
| PQC No Linear Regression | [38] [+] performance [221 phenols/ [simulation]/ [Rtrain2, Rval2, MSEtrain, MSEval RMStrain, RMSval] |
| Time | Accuracy/Precision | |
|---|---|---|
| QML superiority | 7 | 12 |
| QML-ML equivalence | 2 | 4 |
| ML superiority | 4 | 6 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Avramouli, M.; Savvas, I.K.; Vasilaki, A.; Garani, G. Unlocking the Potential of Quantum Machine Learning to Advance Drug Discovery. Electronics 2023, 12, 2402. https://doi.org/10.3390/electronics12112402
Avramouli M, Savvas IK, Vasilaki A, Garani G. Unlocking the Potential of Quantum Machine Learning to Advance Drug Discovery. Electronics. 2023; 12(11):2402. https://doi.org/10.3390/electronics12112402
Chicago/Turabian StyleAvramouli, Maria, Ilias K. Savvas, Anna Vasilaki, and Georgia Garani. 2023. "Unlocking the Potential of Quantum Machine Learning to Advance Drug Discovery" Electronics 12, no. 11: 2402. https://doi.org/10.3390/electronics12112402
APA StyleAvramouli, M., Savvas, I. K., Vasilaki, A., & Garani, G. (2023). Unlocking the Potential of Quantum Machine Learning to Advance Drug Discovery. Electronics, 12(11), 2402. https://doi.org/10.3390/electronics12112402









