Hardware Architectures for Real-Time Medical Imaging
Abstract
1. Introduction
2. Evolution of Hardware Architectures and Their Applications in Real-Time Medical Imaging
2.1. Central Processing Units (CPUs)
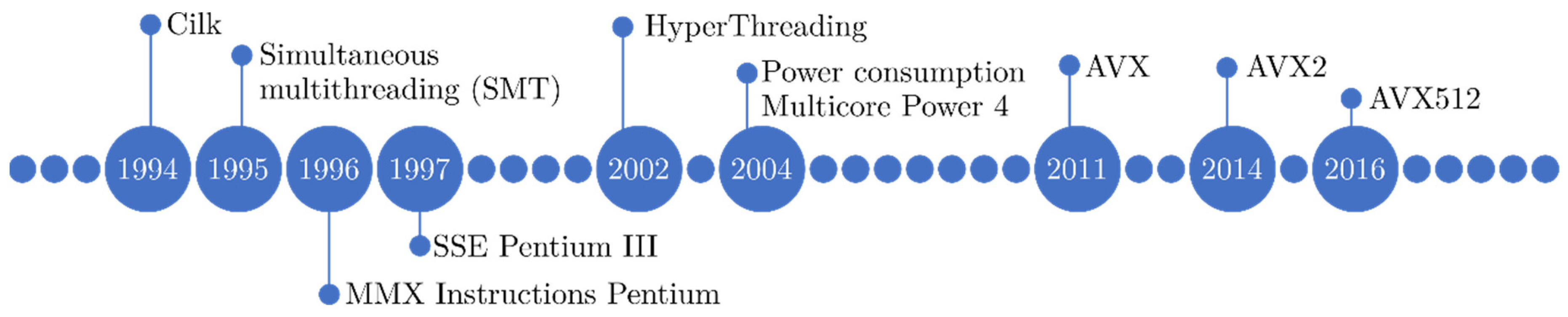
2.1.1. Evolution of CPU Architectures
2.1.2. CPU Architectures in Medical Imaging
2.2. Graphics Processing Units (GPUs)
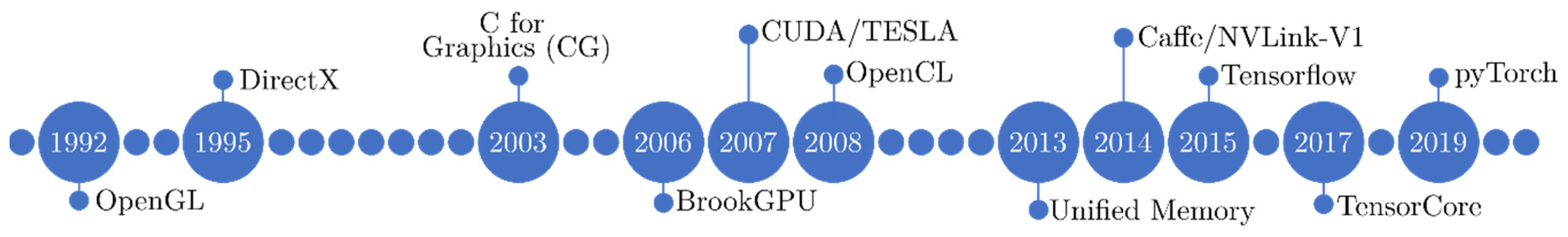
2.2.1. Evolution of GPU Architectures
2.2.2. GPU Architectures in Medical Imaging
2.3. Digital Signal Processors (DSPs)
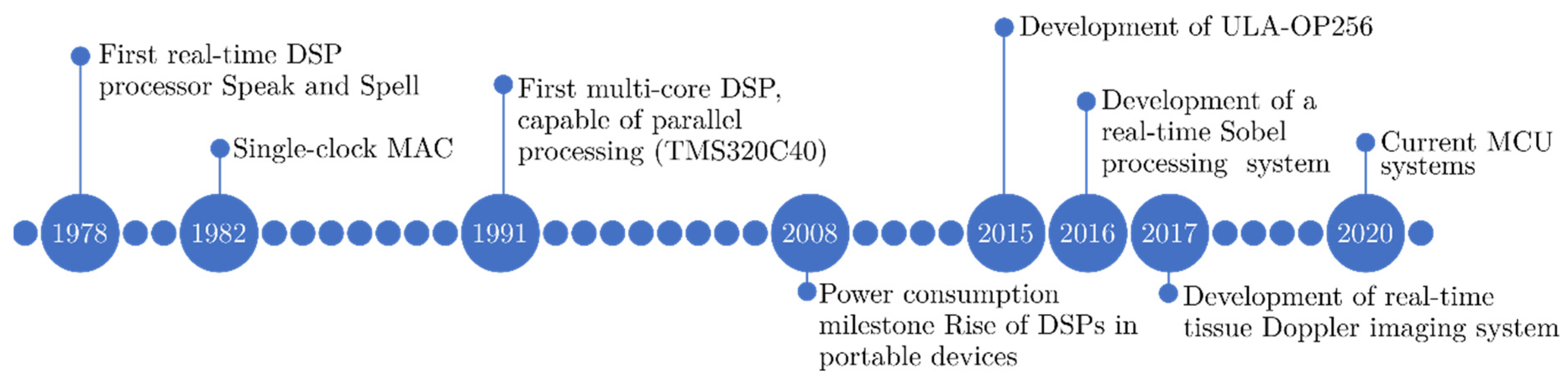
2.3.1. Evolution of DSPs Architectures
2.3.2. DSPs in Medical Imaging
2.4. Field Programmable Gate Arrays (FPGAs)
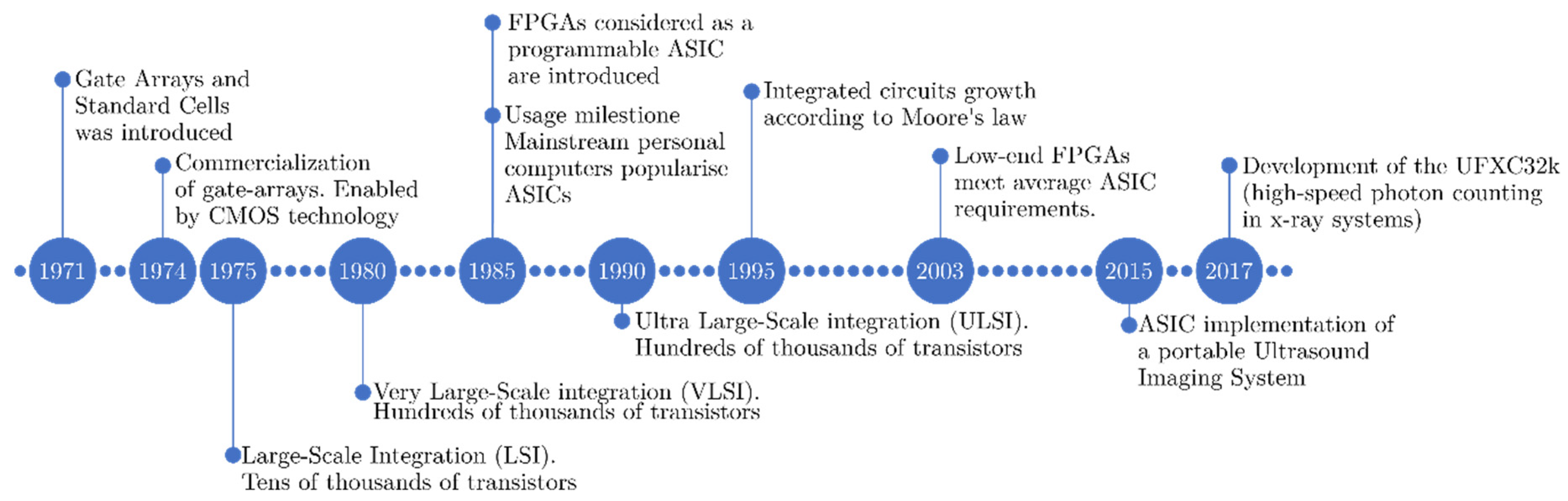
2.4.1. Evolution of FPGA Architectures
2.4.2. FPGA in Medical Imaging
2.5. Application Specific Integrated Circuits (ASICs)
2.5.1. Evolution of ASICs
2.5.2. ASICs in Medical Imaging
3. Discussion
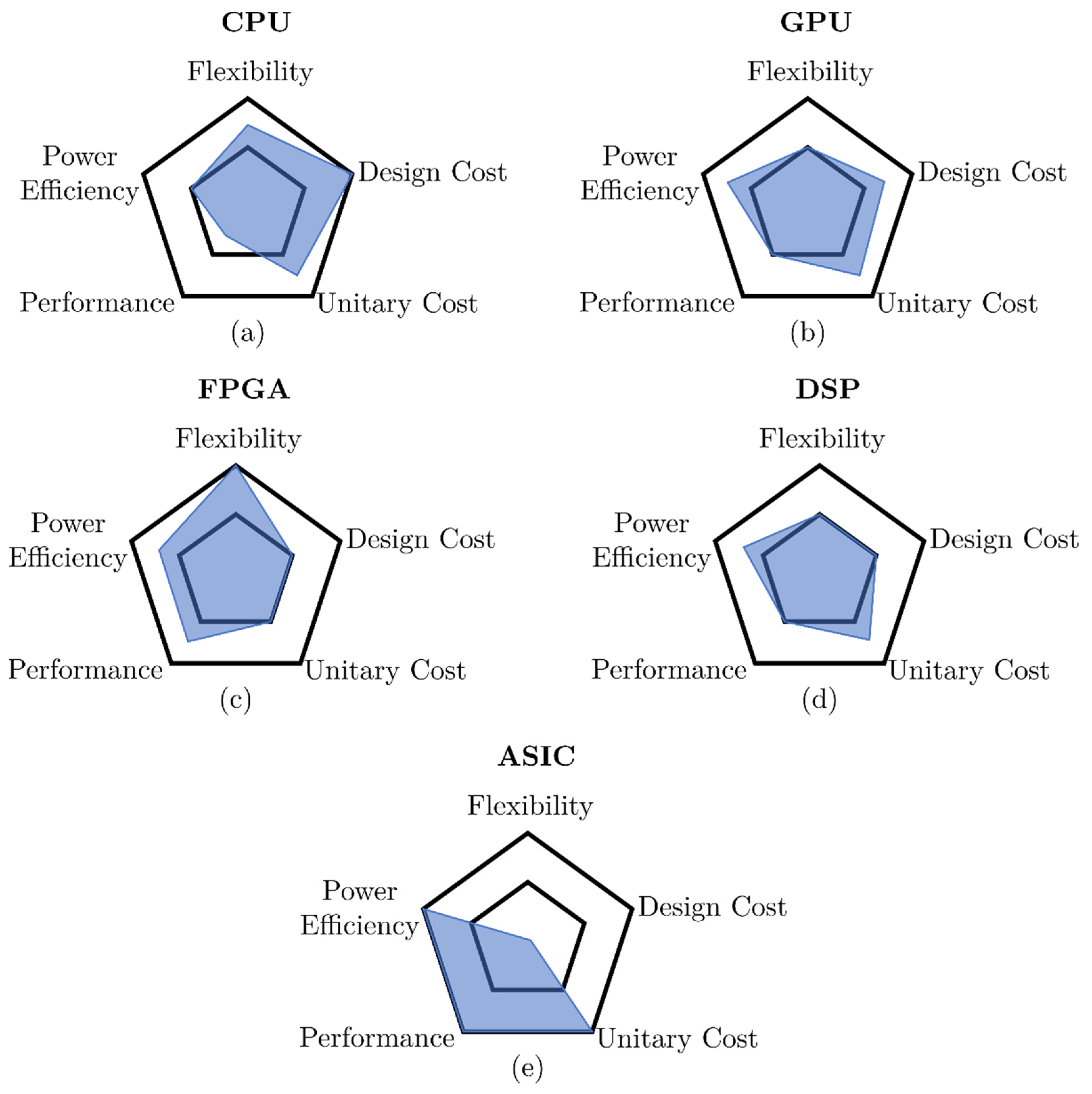
3.1. Hardware Architectures Comparison
- Flexibility: Ability to adapt the platform to different scenarios and use cases while keeping performance high enough to be a competitive option (higher is better).
- Design cost: Economic cost of resources needed to produce and program the device. This includes coding easiness (higher rating means lower requirements).
- Unitary cost: Cost per manufactured unit. This cost does not include coding or implementation expenses and is influenced by common manufacturing runs (higher rating means lower cost).
- Performance: Ability to complete a given (appropriate) task (higher is better).
- Power efficiency: Power required to operate (higher rating means lower power consumption).
3.2. Advantages and Disadvantages of Hardware Architectures for Real-Time Medical Imaging
- Lower level: It deals with pixel operation functions, differences between images and filtering using kernels. These operations are usually repetitive and exhaustive processes that tend to increase the operations number despite the simplicity of the operations.
- Intermediate level: The segmentation, the movement estimation and the extraction or coincidence of characteristics (i.e., image registration). It requires a large use of operations and memory.
- Top level: It deals with the interpretation. This function requires prior knowledge of the environment and even the application of Artificial Intelligence techniques.
- They are an adequate platform for prototyping due to the existence of many ubiquitous optimized libraries for tackling image processing and mathematical problems.
- There exist standards (OpenMP) to facilitate multithreading programming.
- They offer good performance in problems that require serial processing.
- They can be part of computation clusters or supercomputers.
- A combination of multicores and vectorization is needed to obtain the best speed up. This implies a deep knowledge of hardware and software and heavily timed optimization processes.
- Difficult to control the exclusivity of the memory, as there are more processes running on the same operating system.
- They offer a good performance/cost ratio.
- Coding productivity has been improved in recent years.
- GPU architectures have been adapted to cover different algorithms that were not suitable at the beginning, such as branching or bank conflicts in shared memory.
- Many high optimized libraries have been developed for many tasks such as machine learning, signal, and image processing, etc., increasing their suitability for medical imaging problems.
- More efficiency in medical image processing tasks, generally, due to GPU ecosystems being designed and optimized for working with large amounts of data.
- Manageability of cloud ecosystems with powerful GPUs, which eliminate the necessity of having local equipment.
- Data transfer between general and dedicated memory limits some real-time applications, generating a memory transfer/exchange bottleneck.
- Difficulty to migrate code into other GPUs vendors, unless using OpenCL and HIP.
- Requirement of deep knowledge of CPU and GPU programming languages for specialized medical imaging solutions involving GPUs.
- Not all image processing tasks perform better on a GPU than on a CPU.
- The development costs of a portable system are relatively low, helping to lower the prices of each part of the imaging equipment.
- Generally, medical image processing tasks involve sequential algorithms, and the chip architecture of DSPs is optimized to do it. Multi-core DSPs have added the ability to implement coarse-grained parallelism for algorithms with a low to medium level of complexity.
- Power consumption in DSPs is relatively low. There are DSP families specially designed for mobile and portable applications, being a great choice for portable imaging equipment, such as portable ultrasound imaging equipment.
- Development time for simple single-core DSP computer vision and image processing algorithms is generally relatively short. There are free libraries and operative systems available for imaging processing.
- DSPs used to incorporate standard communications ports (e.g., USB, SATA, RS-232 and ethernet). It facilitates storage and transmission tasks between medical devices.
- Some DSPs are compatible with audio and video codecs, making them suitable for video processing applications on portable devices.
- Multi-core DSPs are designed for low to medium complexity HPC applications mostly. Thus, they are not suitable for high-speed or high-data throughput applications, and their use is not recommended for imaging devices that generate great amounts of data.
- DSPs are better suited for sequential processing but not a suitable option for increasing processing speed in massively parallel imaging algorithms.
- It is generally not efficient to use DSP in conjunction with CPU in PCs. Both have a similar sequential processing nature, while CPU programming is easier and more efficient than using DSP.
- It is possible to perform massive arithmetic operations per clock cycle, and these operations can be performed in parallel. As stated before, for the case of GPUs, this is of great interest, as a huge amount of medical image processing and analysis algorithms are inherently parallelizable.
- Different image algorithms implementations or designs are feasible using the same hardware, which provides flexibility of application.
- The designer can create prototypes quickly, simulate, measure time constraints, and modify them, making FPGAs a very interesting option for real-time medical image processing applications using High-Level Synthesis (HLS) tools, such as Vitis HLS.
- The programmable logic of current FPGAs is huge, allowing more complex algorithms to be implemented in hardware. Thus, FPGAs are excellent devices to implement functions and operations, such as convolutions, or even neural networks for machine learning and DL.
- In the mixed hardware/software architectures, it is possible to implement a hardware solution versus a software solution to achieve some algorithm acceleration.
- The integration of the recent FPGA devices includes RAM memory, ARM processors and even DSP engines on the same die. This makes it possible to develop more complex applications by developing heterogeneous architectures where a large part of the elements contained in the FPGA are used.
- Image processing algorithms are often highly complex and computationally expensive, requiring many logical resources. Thus, it is necessary to consider the cost of FPGAs since the more programmable logic integrated into the FPGA, the more expensive the device.
- Although it is possible to create prototypes quickly, it has long design cycles to optimize the resources consumption and achieve the time constraints, as well as making sure that the hardware designs reach a balance between area occupied and parallelization. In addition, the compilation and synthesis processes can take some time, depending on the design, and are related to the designer experience
- The chip size is reduced, resulting in ASICs with high levels of customization. Many circuits can be integrated on the same chip, making it ideal for high-speed applications.
- Libraries in the case of standard cells reduce development time and design complexity.
- Minimal signal routing and timing issues since there is minimal routing to interconnect the different circuits.
- Low power consumption. The ASICs are designed to use all parts of the circuit, which implies high energy efficiency.
- ASICs in general have a high cost and development time.
- The design of ASICs requires greater design skill and complexity, which increases the price per unit.
- ASICs have limited programming flexibility because they are custom chips created for a specific application.
- ASICs require more time to market.
3.3. Concluding Remarks and Outlook to the Future
Author Contributions
Funding
Conflicts of Interest
References
- Iglehart, J.K. The New Era of Medical Imaging. N. Engl. J. Med. 2006, 355, 1502. [Google Scholar] [CrossRef]
- Munn, Z.; Jordan, Z. The patient experience of high technology medical imaging: A systematic review of the qualitative evidence. Radiography 2011, 17, 323–331. [Google Scholar] [CrossRef]
- Suetens, P. Fundamentals of Medical Imaging, 3rd ed.; Cambridge University Press: Cambridge, UK, 2017. [Google Scholar]
- Beyer, T.; Pichler, B. A decade of combined imaging: From a PET attached to a CT to a PET inside an MR. Eur. J. Nucl. Med. Mol. Imaging 2009, 36, 1–2. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Cal-Gonzalez, J.; Rausch, I.; Sundar, L.K.S.; Lassen, M.L.; Muzik, O.; Moser, E.; Papp, L.; Beyer, T. Hybrid imaging: Instrumentation and data processing. Front. Phys. 2018, 6. [Google Scholar] [CrossRef]
- Saini, S.; Seltzer, S.E.; Bramson, R.T.; Levine, L.A.; Kelly, P.; Jordan, P.F.; Chîango, B.F.; Thrall, J.H. Technical cost of radiologic examinations: Analysis across imaging modalities. Radiology 2000, 216, 269–272. [Google Scholar] [CrossRef]
- Sistrom, C.L.; McKay, N.L. Costs, charges, and revenues for hospital diagnostic imaging procedures: Differences by modality and hospital characteristics. J. Am. Coll. Radiol. 2005, 2, 511–519. [Google Scholar] [CrossRef]
- HajiRassouliha, A.; Taberner, A.J.; Nash, M.P.; Nielsen, P.M.F. Suitability of recent hardware accelerators (DSPs, FPGAs, and GPUs) for computer vision and image processing algorithms. Signal Process. Image Commun. 2018, 68, 101–119. [Google Scholar] [CrossRef]
- Talib, M.A.; Majzoub, S.; Nasir, Q.; Jamal, D. A Systematic Literature Review on Hardware Implementation of Artificial Intelligence Algorithms. J. Supercomput. 2021, 77. [Google Scholar] [CrossRef]
- Mittal, S. Vibhu A survey of accelerator architectures for 3D convolution neural networks. J. Syst. Archit. 2021, 115, 102041. [Google Scholar] [CrossRef]
- Alzubaidi, L.; Zhang, J.; Humaidi, A.J.; Al-Dujaili, A.; Duan, Y.; Al-Shamma, O.; Santamaría, J.; Fadhel, M.A.; Al-Amidie, M.; Farhan, L. Review of Deep Learning: Concepts, CNN Architectures, Challenges, Applications, Future Directions. J. Big Data 2021, 8. [Google Scholar] [CrossRef]
- Eklund, A.; Dufort, P.; Forsberg, D.; LaConte, S.M. Medical image processing on the GPU--Past, present and future. Med. Image Anal. 2013, 17, 1073–1094. [Google Scholar] [CrossRef]
- Shi, L.; Liu, W.; Zhang, H.; Xie, Y.; Wang, D. A survey of GPU-based medical image computing techniques. Quant. Imaging Med. Surg. 2012, 2, 188. [Google Scholar]
- Wang, H.; Peng, H.; Chang, Y.; Liang, D. A survey of GPU-based acceleration techniques in MRI reconstructions. Quant. Imaging Med. Surg. 2018, 8, 196–208. [Google Scholar] [CrossRef]
- Smistad, E.; Falch, T.L.; Bozorgi, M.; Elster, A.C.; Lindseth, F. Medical image segmentation on GPUs—A comprehensive review. Med. Image Anal. 2015, 20, 1–18. [Google Scholar] [CrossRef]
- Fluck, O.; Vetter, C.; Wein, W.; Kamen, A.; Preim, B.; Westermann, R. A survey of medical image registration on graphics hardware. Comput. Methods Programs Biomed. 2011, 104, e45–e57. [Google Scholar] [CrossRef]
- Shams, R.; Sadeghi, P.; Kennedy, R.; Hartley, R. A survey of medical image registration on multicore and the GPU. IEEE Signal Process. Mag. 2010, 27, 50–60. [Google Scholar] [CrossRef]
- Passaretti, D.; Joseph, J.M.; Pionteck, T. Survey on FPGAs in medical radiology applications: Challenges, architectures and programming models. In Proceedings of the 2019 International Conference on Field-Programmable Technology (ICFPT), Tianjin, China, 9–13 December 2019; pp. 279–282. [Google Scholar] [CrossRef]
- Moses, J.; Selvathi, D. A survey on FPGA implementation of medical image registration. Asian J. Inf. Technol. 2014, 13, 46–52. [Google Scholar]
- Ballabriga, R.; Alozy, J.; Bandi, F.N.; Campbell, M.; Egidos, N.; Fernandez-Tenllado, J.M.; Heijne, E.H.M.; Kremastiotis, I.; Llopart, X.; Madsen, B.J.; et al. Photon Counting Detectors for X-Ray Imaging with Emphasis on CT. IEEE Trans. Radiat. Plasma Med. Sci. 2021, 5, 422–440. [Google Scholar] [CrossRef]
- Gepner, P.; Kowalik, M.F. Multi-core processors: New way to achieve high system performance. In Proceedings of the International Symposium on Parallel Computing in Electrical Engineering (PARELEC’06), Bialystok, Poland, 13–17 September 2006; pp. 9–13. [Google Scholar]
- Mittal, M.; Peleg, A.; Weiser, U. MMX Technology Architecture Overview. Intel Technol. J. 1997, 1. Available online: https://www.intel.com/content/dam/www/public/us/en/documents/research/1997-vol01-iss-3-intel-technology-journal.pdf (accessed on 15 October 2021).
- Oberman, S.; Favor, G.; Weber, F. AMD 3DNow! technology: Architecture and implementations. IEEE Micro 1999, 19, 37–48. [Google Scholar] [CrossRef]
- Lomont, C. Introduction to intel advanced vector extensions. Intel White Pap. 2011, 1–21. [Google Scholar]
- Jeffers, J.; Reinders, J.; Sodani, A. Intel Xeon Phi Processor High Performance Programming: Knights Landing Edition; Morgan Kaufmann: San Francisco, CA, USA, 2016. [Google Scholar]
- Hennessy, J.L.; Patterson, D.A. Cumputer Architecture, Sixth Edition: A Quantitative Approach; Morgan Kaufmann Publishers Inc.: San Francisco, CA, USA, 2017; ISBN 0128119055. [Google Scholar]
- Asanovic, K.; Bodik, R.; Catanzaro, B.C.; Gebis, J.J.; Husbands, P.; Keutzer, K.; Patterson, D.A.; Plishker, W.L.; Shalf, J.; Williams, S.W.; et al. The Landscape of Parallel Computing Research: A View from Berkeley; Technical Report UCB/EECS-2006-183; EECS Department, University of California: Berkeley, CA, USA, 2006. [Google Scholar]
- Barlas, G. Multicore and GPU Programming. Elsevier Morgan-Kaufmann Publishers Inc.: Burlington, MA, USA, 2014. [Google Scholar]
- Amdahl, G.M. Validity of the single processor approach to achieving large scale computing capabilities. In Proceedings of the Spring Joint Computer Conference, Atlantic City, NJ, USA, 18–20 April 1967; pp. 483–485. [Google Scholar] [CrossRef]
- Hill, M.D.; Marty, M.R. Amdahl’s Law in the Multicore Era. Comput. Long. Beach. Calif. 2008, 41, 33–38. [Google Scholar] [CrossRef]
- Flynn, M.J. Very High-speed Computing Systems. Proc. IEEE 1966, 54, 1901–1909. [Google Scholar] [CrossRef]
- Roy, A.; Xu, J.; Chowdhury, M.H. Multi-core processors: A new way forward and challenges. In Proceedings of the 2008 International Conference on Microelectronics, Sharjah, United Arab Emirates, 14–17 December 2008; pp. 454–457. [Google Scholar] [CrossRef]
- IEEE. IEEE P1003.1c/D10; Draft Standard for Information Technology—Portable Operating Systems Interface (POSIX); The Institute of Electrical & Electronics Engineers: New York, NY, USA, 1994. [Google Scholar]
- OpenMP Application Programming Interface, OpenMP Specification. Available online: https://www.openmp.org/wp-content/uploads/OpenMP-API-Specification-5.0.pdf (accessed on 15 October 2021).
- Reinders, J. Intel Threading Building BLOCKS-outfitting C++ for Multi-Core Processor Parallelism; O’Reilly Media, Inc.: Sebastopol, CA, USA; ISBN 978-0-596-51480-8.
- Marr, D.; Binns, F. Hyper-Threading Technology Architecture and Microarchitecture. Intel Technol. J. 2002, 1–12. [Google Scholar]
- Membarth, R.; Hannig, F.; Teich, J.; Körner, M.; Eckert, W. Frameworks for Multi-core Architectures: A Comprehensive Evaluation Using 2D/3D Image Registration. In Architecture of Computing Systems—ARCS 2011; Berekovic, M., Fornaciari, W., Brinkschulte, U., Silvano, C., Eds.; Springer: Berlin/Heidelberg, Germany, 2011; pp. 62–73. [Google Scholar]
- Ekström, S.; Pilia, M.; Kullberg, J.; Ahlström, H.; Strand, R.; Malmberg, F. Faster dense deformable image registration by utilizing both CPU and GPU. J. Med. Imaging 2021, 8, 1–15. [Google Scholar] [CrossRef]
- Kegel, P.; Schellmann, M.; Gorlatch, S. Comparing programming models for medical imaging on multi-core systems. Concurr. Comput. Pract. Exp. 2011, 23, 1051–1065. [Google Scholar] [CrossRef]
- Kalamkar, D.D.; Trzaskoz, J.D.; Sridharan, S.; Smelyanskiy, M.; Kim, D.; Manduca, A.; Shu, Y.; Bernstein, M.A.; Kaul, B.; Dubey, P. High performance non-uniform FFT on modern X86-based multi-core systems. In Proceedings of the 2012 IEEE 26th International Parallel and Distributed Processing Symposium 2012, Shanghai, China, 21–25 May 2012; pp. 449–460. [Google Scholar] [CrossRef]
- Saxena, S.; Sharma, N.; Sharma, S. An intelligent system for segmenting an abdominal image in multi core architecture. In Proceedings of the 2013 10th International Conference and Export on Emerging Technologies for a Smarter World (CEWIT), Melville, NY, USA, 21–22 October 2013. [Google Scholar] [CrossRef]
- Alsmirat, M.A.; Jararweh, Y.; Al-Ayyoub, M.; Shehab, M.A.; Gupta, B.B. Accelerating compute intensive medical imaging segmentation algorithms using hybrid CPU-GPU implementations. Multimed. Tools Appl. 2017, 76, 3537–3555. [Google Scholar] [CrossRef]
- Valsalan, P.; Sriramakrishnan, P.; Sridhar, S.; Charlyn, G.; Latha, P.; Priya, A.; Ramkumar, S.; Robert, S.A.; Rajendran, T. Knowledge based fuzzy c-means method for rapid brain tissues segmentation of magnetic resonance imaging scans with CUDA enabled GPU machine. J. Ambient Intell. Humaniz. Comput. 2020, 1, 3. [Google Scholar] [CrossRef]
- Vaze, S.; Xie, W.; Namburete, A.I.L. Low-Memory CNNs Enabling Real-Time Ultrasound Segmentation Towards Mobile Deployment. IEEE J. Biomed. Health Inform. 2020, 24, 1059–1069. [Google Scholar] [CrossRef] [PubMed]
- Kirk, D.B.; Wen-Mei, W.H. Programming Massively Parallel Processors: A Hands-on Approach; Morgan kaufmann: San Francisco, CA, USA, 2016. [Google Scholar]
- Wilt, N. The Cuda Handbook: A Comprehensive Guide to Gpu Programming; CreateSpace Independent Publishing Platform: Scotts Valley, CA, USA, 2017; ISBN 9781548845162. [Google Scholar]
- Woo, M.; Neider, J.; Davis, T.; Shreiner, D. Opengl Programming Guide: The Official Guide To Learning Opengl; Version 1.2; Addison-Wesley: Boston, MA, USA, 1999; ISBN 978-0-201-60458-0. [Google Scholar]
- Microsoft. DirectX Developer Site. Available online: https://docs.microsoft.com/en-us/windows/win32/directx (accessed on 15 October 2021).
- Owens, J.D.; Luebke, D.; Govindaraju, N.; Harris, M.; Krüger, J.; Lefohn, A.E.; Purcell, T.J. A survey of general-purpose computation on graphics hardware. Comput. Graph. Forum 2007, 26, 80–113. [Google Scholar] [CrossRef]
- Cook, S. CUDA Programming: A Developer’s Guide to Parallel Computing with GPUs. Elsevier Morgan-Kaufmann Publishers Inc.: Burlington, MA, USA, 2013; 012415334. [Google Scholar]
- Otterness, N.; Anderson, J.H. AMD GPUs as an alternative to NVIDIA for supporting real-time workloads. In Leibniz International Proceedings in Informatics (LIPIcs), Proceedings of the 32nd Euromicro Conference on Real-Time Systems (ECRTS 2020), Virtual, 7– 10 July 2020; Völp, M., Ed.; Schloss Dagstuhl–Leibniz-Zentrum für Informatik: Dagstuhl, Germany, 2020; Volume 165, pp. 10:1–10:23. [Google Scholar] [CrossRef]
- Nvidia Corporation, Nvidia NVIDIA A100 Tensor Core GPU Architecture Unprecedented Acceleration at Every Scale (White Paper). Available online: https://images.nvidia.cn/aem-dam/en-zz/Solutions/data-center/nvidia-ampere-architecture-whitepaper.pdf (accessed on 15 October 2021).
- Nvidia Corporation. NVIDIA® NVLink TM High-Speed Interconnect: Application Performance (White Paper). Available online: http://info.nvidianews.com/rs/nvidia/images/NVIDIA%20NVLink%20High-Speed%20Interconnect%20Application%20Performance%20Brief.pdf (accessed on 15 October 2021).
- Jia, Z.; Maggioni, M.; Staiger, B.; Scarpazza, D.P. Dissecting the NVIDIA Volta GPU Architecture via Microbenchmarking. arXiv 2018, arXiv:cs.DC/1804.06826. [Google Scholar]
- Bell, N.; Garland, M. Efficient Sparse Matrix-Vector Multiplication On Cuda. Available online: https://www.nvidia.com/docs/IO/66889/nvr-2008-004.pdf (accessed on 15 October 2021).
- Volkov, V. Understanding Latency Hiding On Gpus. Ph.D. Thesis, University of California Berkeley, Berkeley, CA, USA, 2016. [Google Scholar]
- Bell, N.; Hoberock, J. Thrust: A Productivity-Oriented Library for CUDA. In GPU Computing Gems JADE Edition; Elsevier: Amsterdam, The Netherlands, 2012; pp. 359–371. [Google Scholar]
- Merrill III, D.G. Allocation-Oriented Algorithm Design with Application to Gpu Computing; University of Virginia: Charlottesville, VA, USA, 2011. [Google Scholar]
- Armyr, D.; Doherty, D. MathWorks. Prototyping Algorithms and Testing CUDA Kernels in MATLAB. Available online: https://es.mathworks.com/company/newsletters/articles/prototyping-algorithms-and-testing-cuda-kernels-in-matlab.html (accessed on 15 October 2021).
- Munshi, A.; Gaster, B.; Mattson, T.G.; Ginsburg, D. OpenCL Programming Guide; Pearson Education: London, UK, 2011. [Google Scholar]
- Welcome to AMD ROCm™ Platform. AMD Corporation. OCm Documentation. Available online: https://rocmdocs.amd.com/en/latest/ (accessed on 15 October 2021).
- The OpenACC© Application Programming Interface. Version 3.0. Nov. 2019. Available online: https://www.openacc.org/sites/default/files/inline-images/Specification/OpenACC.3.0.pdf (accessed on 15 October 2021).
- Jia, Y.; Shelhamer, E.; Donahue, J.; Karayev, S.; Long, J.; Girshick, R.; Guadarrama, S.; Darrell, T. Caffe: Convolutional Architecture for Fast Feature Embedding. In Proceedings of the 22nd ACM International Conference on Multimedia, Orlando, FL, USA, 3–7 November 2014; pp. 675–678. [Google Scholar] [CrossRef]
- Abadi, M.; Agarwal, A.; Barham, P.; Brevdo, E.; Chen, Z.; Citro, C.; Corrado, G.S.; Davis, A.; Dean, J.; Devin, M.; et al. TensorFlow: Large-Scale Machine Learning on Heterogenous Distributed Systems. arXiv 2016, arXiv:cs.DC/1603.04467]. [Google Scholar]
- Paszke, A.; Gross, S.; Massa, F.; Lerer, A.; Bradbury, J.; Chanan, G.; Killeen, T.; Lin, Z.; Gimelshein, N.; Antiga, L.; et al. PyTorch: An imperative style, high-performance deep learning library. Adv. Neural Inf. Process. Syst. 2019, 32, 8026–8037. [Google Scholar]
- Johnson, D.H.; Narayan, S.; Flask, C.A.; Wilson, D.L. Improved fat-water reconstruction algorithm with graphics hardware acceleration. J. Magn. Reson. Imaging 2010, 31, 457–465. [Google Scholar] [CrossRef]
- Herraiz, J.L.; España, S.; Cabido, R.; Montemayor, A.S.; Desco, M.; Vaquero, J.J.; Udias, J.M. GPU-based fast iterative reconstruction of fully 3-D PET Sinograms. IEEE Trans. Nucl. Sci. 2011, 58, 2257–2263. [Google Scholar] [CrossRef]
- Alcaín, E.; Torrado-Carvajal, A.; Montemayor, A.S.; Malpica, N. Real-time patch-based medical image modality propagation by GPU computing. J. Real-Time Image Process. 2017, 13, 193–204. [Google Scholar] [CrossRef]
- Torrado-Carvajal, A.; Herraiz, J.L.; Alcain, E.; Montemayor, A.S.; Garcia-Canamaque, L.; Hernandez-Tamames, J.A.; Rozenholc, Y.; Malpica, N. Fast patch-based pseudo-CT synthesis from T1-weighted MR images for PET/MR attenuation correction in brain studies. J. Nucl. Med. 2016, 57, 136–143. [Google Scholar] [CrossRef]
- Punithakumar, K.; Boulanger, P.; Noga, M. A GPU-Accelerated Deformable Image Registration Algorithm With Applications to Right Ventricular Segmentation. IEEE Access 2017, 5, 20374–20382. [Google Scholar] [CrossRef]
- Florimbi, G.; Fabelo, H.; Torti, E.; Ortega, S.; Marrero-Martin, M.; Callico, G.M.; Danese, G.; Leporati, F. Towards Real-Time Computing of Intraoperative Hyperspectral Imaging for Brain Cancer Detection Using Multi-GPU Platforms. IEEE Access 2020, 8, 8485–8501. [Google Scholar] [CrossRef]
- Torti, E.; Leon, R.; Salvia, M.L.; Florimbi, G.; Martinez-Vega, B.; Fabelo, H.; Ortega, S.; Callicó, G.M.; Leporati, F. Parallel classification pipelines for skin cancer detection exploiting hyperspectral imaging on hybrid systems. Electronics 2020, 9, 1503. [Google Scholar] [CrossRef]
- Zachariadis, O.; Teatini, A.; Satpute, N.; Gómez-Luna, J.; Mutlu, O.; Elle, O.J.; Olivares, J. Accelerating B-spline interpolation on GPUs: Application to medical image registration. Comput. Methods Programs Biomed. 2020, 193, 105431. [Google Scholar] [CrossRef]
- Milshteyn, E.; Guryev, G.; Torrado-Carvajal, A.; Adalsteinsson, E.; White, J.K.; Wald, L.L.; Guerin, B. Individualized SAR calculations using computer vision-based MR segmentation and a fast electromagnetic solver. Magn. Reson. Med. 2021, 85, 429–443. [Google Scholar] [CrossRef]
- Brunn, M.; Himthani, N.; Biros, G.; Mehl, M.; Mang, A. Fast GPU 3D diffeomorphic image registration. J. Parallel Distrib. Comput. 2021, 149, 149–162. [Google Scholar] [CrossRef]
- Martinez-Girones, P.M.; Vera-Olmos, J.; Gil-Correa, M.; Ramos, A.; Garcia-Cañamaque, L.; Izquierdo-Garcia, D.; Malpica, N.; Torrado-Carvajal, A. Franken-ct: Head and neck mr-based pseudo-ct synthesis using diverse anatomical overlapping mr-ct scans. Appl. Sci. 2021, 11, 3508. [Google Scholar] [CrossRef]
- Frantz, G.; Brantingham, L. Signal Core: A Short History of the Digital Signal Processor. IEEE Solid-State Circuits Mag. 2012, 4, 16–20. [Google Scholar] [CrossRef]
- Karam, L.; Alkamal, I.; Gatherer, A.; Frantz, G.; Anderson, D.; Evans, B. Trends in multicore DSP platforms. IEEE Signal. Process. Mag. 2009, 26, 38–49. [Google Scholar] [CrossRef]
- Eyre, J.; Bier, J. The evolution of DSP processors. IEEE Signal Processing Magazine 2000, 17, 43–51. [Google Scholar] [CrossRef]
- Frantz, G. Digital signal processor trends. IEEE Micro 2000, 20, 52–59. [Google Scholar] [CrossRef]
- Illgner, K. DSPs for image and video processing. Signal Process. 2000, 80, 2323–2336. [Google Scholar] [CrossRef]
- Narendra, C.P.; Kumar, K.M.R. Low power MAC architecture for DSP applications. In Proceedings of the International Conference on Circuits, Communication, Control and Computing, Bangalore, India, 21–22 November 2014; pp. 404–407. [Google Scholar] [CrossRef]
- Simar, R. The TMS320C40: A DSP for parallel processing. In Proceedings of the ICASSP 91: 1991 International Conference on Acoustics, Speech, and Signal Processing, Toronto, ON, Canada, 14–17 April 1991; pp. 1089–1092. [Google Scholar]
- Fridman, J.; Greenfield, Z. The TigerSHARC DSP architecture. IEEE Micro 2000, 20, 66–76. [Google Scholar] [CrossRef]
- Takala, J. General-Purpose DSP Processors. In Handbook of Signal Processing Systems; Bhattacharyya, S., Deprettere, E., Leupers, R., Takala, J., Eds.; Springer: Boston, MA, USA, 2010. [Google Scholar] [CrossRef]
- Patterson, J.; Dixon, J. Optimizing Power Consumption in DSP Designs (White Paper). Available online: https://www.ti.com/lit/wp/spry089/spry089.pdf?ts=1639187652517&ref_url=https%253A%252F%252Fwww.google.com.hk%252F (accessed on 15 October 2021).
- Rao, A.; Nagori, S. Optimizing Video Encoders with TI DSPs (White Paper). Available online: https://www.ti.com/lit/wp/spry106/spry106.pdf?ts=1639187749440&ref_url=https%253A%252F%252Fwww.google.com.hk%252F (accessed on 15 October 2021).
- Bergsagel, J.; Leconte, L. Super high resolution displays empowered by the OMAP4470 mobile processor (White paper). Available online: https://www.ti.com/lit/wp/swpy028/swpy028.pdf (accessed on 15 October 2021).
- Chaoui, J.; Cyr, K.; De Gregorio, S.; Giacalone, J.P.; Webb, J.; Masse, Y. Open multimedia application platform: Enabling multimedia applications in third generation wireless terminals through a combined Risc/Dsp architecture. In Proceedings of the IEEE International Conference on Acoustics, Speech, and Signal Processing, Salt Lake City, UT, USA, 7–11 May 2001; Volume 2, pp. 1009–1012. [Google Scholar] [CrossRef]
- Bahri, N.; Belhadj, N.; Ben Ayed, M.A.; Masmoudi, N.; Grandpierre, T.; Akil, M. Real-time H264/AVC high definition video encoder on a multicore DSP TMS320C6678. In Proceedings of the International Conference on Computer Vision and Image Analysis Applications, Sousse, Tunisia, 18–20 January 2015; pp. 1–6. [Google Scholar]
- Igual, F.D.; Ali, M.; Friedmann, A.; Stotzer, E.; Wentz, T.; van de Geijn, R.A. Unleashing the high-performance and low-power of multi-core DSPs for general-purpose HPC. In Proceedings of the SC’12 International Conference on High Performance Computing, Networking, Storage and Analysis, Salt Lake City, UT, USA, 25 February 2012; pp. 1–11. [Google Scholar]
- Rao, M.V.G.; Kumar, P.R.; Prasad, A.M. Implementation of real time image processing system with FPGA and DSP. In Proceedings of the 2016 International Conference on Microelectronics, Computing and Communications (MicroCom), Durgapur, India, 23–25 January 2016; pp. 4–7. [Google Scholar] [CrossRef]
- NXP Semiconductors. Crossover Embedded Processors-Bridging the Gap between Performance and Usability (White Paper). Available online: https://www.nxp.com/docs/en/white-paper/I.MXRT1050WP.pdf (accessed on 15 October 2021).
- Zhong, M.; Yang, Y.; Zhou, Y.; Octavian, P.M.; Chandrasekar, M.; Venkat, B.G.; Manikandan, C.; Balaji, V.S.; Saravanan, S.; Elamaran, V. Advanced Digital Signal Processing Techniques on the Classification of the Heart Sound Signals. J. Med. Imaging Heal. Inform. 2020, 10, 2010–2015. [Google Scholar] [CrossRef]
- Berg, R.; König, L.; Rühaak, J.; Lausen, R.; Fischer, B. Highly efficient image registration for embedded systems using a distributed multicore DSP architecture. J. Real-Time Image Process. 2018, 14, 341–361. [Google Scholar] [CrossRef]
- Chandrashekar, L.; Sreedevi, A. A Multi-Objective Enhancement Technique for Poor Contrast Magnetic Resonance Images of Brain Glioblastomas. Procedia Comput. Sci. 2020, 171, 1770–1779. [Google Scholar] [CrossRef]
- Ueng, S.K.; Yen, C.L.; Chen, G.Z. Ultrasound image enhancement using structure-based filtering. Comput. Math. Methods Med. 2014, 1–14. [Google Scholar] [CrossRef]
- Ali, M.; Magee, D.; Dasgupta, U. Signal Processing Overview of Ultrasound Systems for Medical Imaging. Available online: https://www.k-space.org/ymk/sprab12.pdf (accessed on 15 October 2021).
- Pailoor, R.; Pradhan, D. Digital signal processor (DSP) for portable ultrasound. Tex. Instrum. Appl. Rep. SPRAB18A 2008, 1–11. [Google Scholar]
- Boni, E.; Bassi, L.; Dallai, A.; Giannini, G.; Guidi, F.; Meacci, V.; Matera, R.; Ramalli, A.; Ricci, S.; Scaringella, M.; et al. ULA-OP 256: A portable high-performance research scanner. In Proceedings of the 2015 IEEE International Ultrasonics Symposium (IUS), Taipei, Taiwan, 21–24 October 2015; pp. 1–4. [Google Scholar]
- Ricci, S.; Ramalli, A.; Bassi, L.; Boni, E.; Tortoli, P. Real-time blood velocity vector measurement over a 2-D region. IEEE Trans. Ultrason. Ferroelectr. Freq. Control. 2017, 65, 201–209. [Google Scholar] [CrossRef] [PubMed]
- Ramalli, A.; Guidi, F.; Dallai, A.; Boni, E.; Tong, L.; D’hooge, J.; Tortoli, P. High frame rate, wide-angle tissue doppler imaging in real-time. In Proceedings of the 2017 IEEE International Ultrasonics Symposium (IUS), Washington, DC, USA, 6–9 September 2017; pp. 1–4. [Google Scholar]
- Tortoli, P.; Lenge, M.; Righi, D.; Ciuti, G.; Liebgott, H.; Ricci, S. Comparison of carotid artery blood velocity measurements by vector and standard Doppler approaches. Ultrasound Med. Biol. 2015, 41, 1354–1362. [Google Scholar] [CrossRef]
- Arulkumar, M.; Chandrasekaran, M. An Improved VLSI Design of the ALU Based FIR Filter for Biomedical Image Filtering Application. Curr. Med. Imaging 2021, 17, 276–287. [Google Scholar] [CrossRef]
- Beddad, B.; Hachemi, K. Application of brain tumor detection on DSP environment using TMS320C6713 DSK. In Proceedings of the 2017 40th International Conference on Telecommunications and Signal Processing (TSP), Barcelona, Spain, 5–7 July 2017; pp. 599–603. [Google Scholar]
- A Brief History of FPGA|Make. Available online: https://makezine.com/2019/10/11/a-brief-history-of-fpga/ (accessed on 15 October 2021).
- Farooq, U.; Marrakchi, Z.; Mehrez, H. Tree-Based Heterogeneous FPGA Architectures: Application Specific Exploration and Optimization; Springer: New York, NY, USA, 2012; ISBN 9781461435945. [Google Scholar]
- Anusudha, K.; Naguboina, G.C. Design and implementation of PAL and PLA using reversible logic on FPGA SPARTAN 3E. In Proceedings of the 2017 40th International Conference on Telecommunications and Signal Processing (TSP), Barcelona, Spain, 5–7 July 2017; pp. 1–6. [Google Scholar] [CrossRef]
- Grout, I. Digital Systems Design with FPGAs and CPLDs; Newnes: Oxford, UK, 2008; ISBN 9780080558509. [Google Scholar]
- Kuon, I.; Rose, J. Measuring the Gap between FPGAs and ASICs. IEEE Trans. Comput. Des. Integr. Circuits Syst. 2007, 26, 203–215. [Google Scholar] [CrossRef]
- Trimberger, S.M.S. Three Ages of FPGAs: A Retrospective on the First Thirty Years of FPGA Technology: This Paper Reflects on How Moore’s Law Has Driven the Design of FPGAs Through Three Epochs: The Age of Invention, the Age of Expansion, and the Age of Accumulation. IEEE Solid-State Circuits Mag. 2018, 10, 16–29. [Google Scholar] [CrossRef]
- Wolf, W. FPGA-Based System Design; Prentice Hall PTR Pearson Education: Upper Saddle River, NJ, USA, 2004; ISBN 978-0137033485. [Google Scholar]
- Ahmad, S.; Boppana, V.; Ganusov, I.; Kathail, V.; Rajagopalan, V.; Wittig, R. A 16-nm multiprocessing system-on-chip field-programmable gate array platform. IEEE Micro 2016, 36, 48–62. [Google Scholar] [CrossRef]
- Ahmad, S.; Subramanian, S.; Boppana, V.; Lakka, S.; Ho, F.-H.; Knopp, T.; Noguera, J.; Singh, G.; Wittig, R. Xilinx first 7 nm device: Versal AI core (VC1902). In Proceedings of the 2019 IEEE Hot Chips 31 Symposium (HCS), Cupertino, CA, USA, 18–20 August 2019; pp. 1–28. [Google Scholar]
- Voogel, M.; Frans, Y.; Ouellette, M.; Coppens, J.; Ahmad, S.; Dastidar, J.; Mohsen, E.; Dada, F.; Thompson, M.; Wittig, R.; et al. Xilinx VersalTM Premium. In Proceedings of the 2020 IEEE Hot Chips 32 Symposium (HCS), Palo Alto, CA, USA, 16–18 August 2020; pp. 1–46. [Google Scholar]
- Xilinx, Inc. Versal, the First Adaptive Compute Acceleration Platform (ACAP) (White Paper). Available online: https://www.xilinx.com/support/documentation/white_papers/wp505-versal-acap.pdf (accessed on 15 October 2021).
- Gaide, B.; Gaitonde, D.; Ravishankar, C.; Bauer, T. Xilinx Adaptive Compute Acceleration Platform: Versal™ Architecture. Proceedings of the 2019 ACM/SIGDA International Symposium on Field-Programmable Gate Arrays; Association for Computing Machinery: New York, NY, USA, 2019; FPGA ’19; pp. 84–93. [Google Scholar] [CrossRef]
- Dillinger, P.; Vogelbruch, J.F.; Leinen, J.; Suslov, S.; Patzak, R.; Winkler, H.; Schwan, K. FPGA Based Real-Time Image Segmentation for Medical Systems and Data Processing. IEEE Trans. Nuclear Sci. 2006, 53. [Google Scholar] [CrossRef]
- Jiang, R.M.; Crookes, D. FPGA implementation of 3D discrete wavelet transform for real-time medical imaging. In Proceedings of the 2007 18th European Conference on Circuit Theory and Design, Seville, Spain, 27–30 August 2007; pp. 519–522. [Google Scholar]
- Korcyl, G.; Bialas, P.; Curceanu, C.; Czerwinski, E.; Dulski, K.; Flak, B.; Gajos, A.; Glowacz, B.; Gorgol, M.; Hiesmayr, B.C.; et al. Evaluation of Single-Chip, Real-Time Tomographic Data Processing on FPGA SoC Devices. IEEE Trans. Med. Imaging 2018, 37, 2526–2535. [Google Scholar] [CrossRef] [PubMed]
- Gebhardt, P.; Weissler, B.; Zinke, M.; Kiessling, F.; Marsden, P.K.; Schulz, V. FPGA-based singles and coincidences processing pipeline for integrated digital PET/MR detectors. In Proceedings of the 2012 IEEE Nuclear Science Symposium and Medical Imaging Conference Record (NSS/MIC), Anaheim, CA, USA, 27 October–3 November 2012; pp. 2479–2482. [Google Scholar] [CrossRef]
- Marjanovic, J.; Reber, J.; Brunner, D.O.; Engel, M.; Kasper, L.; Dietrich, B.E.; Vionnet, L.; Pruessmann, K.P. A Reconfigurable Platform for Magnetic Resonance Data Acquisition and Processing. IEEE Trans. Med. Imaging 2020, 39, 1138–1148. [Google Scholar] [CrossRef] [PubMed]
- Kim, G.D.; Yoon, C.; Kye, S.B.; Lee, Y.; Kang, J.; Yoo, Y.; Song, T.K. A single FPGA-based portable ultrasound imaging system for point-of-care applications. IEEE Trans. Ultrason. Ferroelectr. Freq. Control 2012, 59, 1386–1394. [Google Scholar] [CrossRef]
- Zhang, Z.; Xin, Y.; Liu, B.; Li, W.X.Y.; Lee, K.H.; Ng, C.F.; Stoyanov, D.; Cheung, R.C.C.; Kwok, K.W. FPGA-based high-performance collision detection: An enabling technique for image-guided robotic surgery. Front. Robot. AI 2016, 3, 1–14. [Google Scholar] [CrossRef]
- Mittal, S.; Gupta, S.; Dasgupta, S. FPGA: An efficient and promising platform for real-time image processing applications. In Proceedings of the National Conference On Research and Development In Hardware Systems (CSI-RDHS), Kolkata, India, 20–21 June 2008. [Google Scholar]
- Mondal, P.; Biswal, P.K.; Banerjee, S. FPGA based accelerated 3D affine transform for real-time image processing applications. Comput. Electr. Eng. 2016, 49, 69–83. [Google Scholar] [CrossRef]
- Conficconi, D.; D’Arnese, E.; Del Sozzo, E.; Sciuto, D.; Santambrogio, M.D. A framework for customizable Fpga-based image registration accelerators. In Proceedings of the FPGA’21: The 2021 ACM/SIGDA International Symposium on Field-Programmable Gate Arrays, Virtual Event, USA, 28 February–2 March 2021; pp. 251–261. [Google Scholar] [CrossRef]
- Mondal, P.; Banerjee, S. FPGA-accelerated adaptive projection-based image registration. J. Real-Time Image Process. 2021, 18, 113–125. [Google Scholar] [CrossRef]
- Gebhardt, P.; Wehner, J.; Weissler, B.; Botnar, R.; Marsden, P.K.; Schulz, V. FPGA-based RF interference reduction techniques for simultaneous PET-MRI. Phys. Med. Biol. 2016, 61, 3500–3526. [Google Scholar] [CrossRef]
- Zhou, H.; Machupalli, R.; Mandal, M. Efficient FPGA implementation of automatic nuclei detection in histopathology images. J. Imaging 2019, 5, 21. [Google Scholar] [CrossRef]
- Othman, M.F.B.; Abdullah, N.; Rusli, N.A.B.A. An overview of MRI brain classification using FPGA implementation. In Proceedings of the 2010 IEEE Symposium on Industrial Electronics and Applications (ISIEA), Penang, Malaysia, 3–5 October 2010; pp. 623–628. [Google Scholar] [CrossRef]
- Sanaullah, A.; Yang, C.; Alexeev, Y.; Yoshii, K.; Herbordt, M.C. Real-time data analysis for medical diagnosis using FPGA-accelerated neural networks. BMC Bioinform. 2018, 19, 19–31. [Google Scholar] [CrossRef]
- Li, L.; Wyrwicz, A.M. Design of an MR image processing module on an FPGA chip. J. Magn. Reson. 2015, 255, 51–58. [Google Scholar] [CrossRef]
- Siddiqui, M.F.; Reza, A.W.; Shafique, A.; Omer, H.; Kanesan, J. FPGA implementation of real-time SENSE reconstruction using pre-scan and Emaps sensitivities. Magn. Reson. Imaging 2017, 44, 82–91. [Google Scholar] [CrossRef]
- Inam, O.; Basit, A.; Qureshi, M.; Omer, H. FPGA-based hardware accelerator for SENSE (a parallel MR image reconstruction method). Comput. Biol. Med. 2020, 117, 103598. [Google Scholar] [CrossRef] [PubMed]
- Liang, X.; Binghe, S.; Yueping, M.; Ruyan, Z. A digital magnetic resonance imaging spectrometer using digital signal processor and field programmable gate array. Rev. Sci. Instrum. 2013, 84, 054702. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.; Cong, J. Acceleration of EM-Based 3D CT Reconstruction Using FPGA. IEEE Trans. Biomed. Circuits Syst. 2016, 10, 754–767. [Google Scholar] [CrossRef]
- Windisch, D.; Knodel, O.; Juckeland, G.; Hampel, U.; Bieberle, A. FPGA-based Real-Time Data Acquisition for Ultrafast X-Ray Computed Tomography. IEEE Trans. Nucl. Sci. 2021. [Google Scholar] [CrossRef]
- Goel, G.; Gondhalekar, A.; Qi, J.; Zhang, Z.; Cao, G.; Feng, W. ComputeCOVID19+: Accelerating COVID-19 diagnosis and monitoring via high-performance deep Learning on CT images. In Proceedings of the 50th International Conference on Parallel Processing; Association for Computing Machinery, Lemont, IL, USA, 9–12 August 2021. [Google Scholar]
- Gong, K.; Berg, E.; Cherry, S.R.; Qi, J. Machine Learning in PET: From Photon Detection to Quantitative Image Reconstruction. Proc. IEEE 2020, 108, 51–68. [Google Scholar] [CrossRef]
- Won, J.Y.; Lee, J.S. Highly Integrated FPGA-Only Signal Digitization Method Using Single-Ended Memory Interface Input Receivers for Time-of-Flight PET Detectors. IEEE Trans. Biomed. Circuits Syst. 2018, 12, 1401–1409. [Google Scholar] [CrossRef]
- Won, J.Y.; Park, H.; Lee, S.; Son, J.-W.; Chung, Y.; Ko, G.B.; Kim, K.Y.; Song, J.; Seo, S.; Ryu, Y.; et al. Development and Initial Results of a Brain PET Insert for Simultaneous 7-Tesla PET/MRI Using an FPGA-Only Signal Digitization Method. IEEE Trans. Med. Imaging 2021, 40, 1579–1590. [Google Scholar] [CrossRef]
- Assef, A.A.; Ferreira, B.M.; Maia, J.M.; Costa, E.T. Modeling and FPGA-based implementation of an efficient and simple envelope detector using a Hilbert Transform FIR filter for ultrasound imaging applications. Res. Biomed. Eng. 2018, 34, 87–92. [Google Scholar] [CrossRef][Green Version]
- Wu, X.; Sanders, J.L.; Zhang, X.; Yamaner, F.Y.; Oralkan, Ö. An FPGA-Based Backend System for Intravascular Photoacoustic and Ultrasound Imaging. IEEE Trans. Ultrason. Ferroelectr. Freq. Control 2019, 66, 45–56. [Google Scholar] [CrossRef] [PubMed]
- Nasser, Y.; Lorandel, J.; Prévotet, J.-C.; Hélard, M. RTL to Transistor Level Power Modeling and Estimation Techniques for FPGA and ASIC: A Survey. IEEE Trans. Comput. Des. Integr. Circuits Syst. 2020, 40, 479–493. [Google Scholar] [CrossRef]
- Weste, N.H.E.; Harris, D. CMOS VLSI Design: A Circuits and Systems Perspective; Pearson Education India: Upper Saddle River, NJ, USA, 2015. [Google Scholar]
- Rabaey, J.M.; Chandrakasan, A.P.; Nikolić, B. Digital Integrated Circuits: A Design Perspective; Pearson education: Upper Saddle River, NJ, USA, 2003; Volume 7. [Google Scholar]
- Zahiri, B. Structured ASICs: Opportunities and challenges. In Proceedings of the 21st International Conference on Computer Design, San Jose, CA, USA, 13–15 October 2003; pp. 404–409. [Google Scholar] [CrossRef]
- Nekoogar, F. From ASICs to SOCs: A Practical Approach; Prentice Hall Professional: Hoboken, NJ, USA, 2003. [Google Scholar]
- Kmon, P.; Szczygiel, R.; Maj, P.; Grybos, P.; Kleczek, R. High-speed readout solution for single-photon counting ASICs. J. Instrum. 2016, 11, C02057. [Google Scholar] [CrossRef]
- Sundberg, C.; Persson, M.U.; Sjölin, M.; Wikner, J.J.; Danielsson, M. Silicon photon-counting detector for full-field CT using an ASIC with adjustable shaping time. J. Med. Imaging 2020, 7, 53503. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.; Persson, M.; Chen, H.; Karlsson, S.; Danielsson, M.; Svensson, C.; Bornefalk, H. Evaluation of a second-generation ultra-fast energy-resolved ASIC for photon-counting spectral CT. IEEE Trans. Nucl. Sci. 2013, 60, 437–445. [Google Scholar] [CrossRef]
- Zannoni, E.M.; Wilson, M.D.; Bolz, K.; Goede, M.; Lauba, F.; Schöne, D.; Zhang, J.; Veale, M.C.; Verhoeven, M.; Meng, L.J. Development of a multi-detector readout circuitry for ultrahigh energy resolution single-photon imaging applications. Nucl. Instrum. Methods Phys. Res. Sect. A Accel. Spectrometers Detect. Assoc. Equip. 2020, 981, 164531. [Google Scholar] [CrossRef]
- Jones, L.; Seller, P.; Wilson, M.; Hardie, A. HEXITEC ASIC-a pixellated readout chip for CZT detectors. Nucl. Instrum. Methods Phys. Res. Sect. A Accel. Spectrometers Detect. Assoc. Equip. 2009, 604, 34–37. [Google Scholar] [CrossRef]
- Pushkar; Junnarkar, S. Development of high resolution modular four side buttable small field of view detectors for three dimensional gamma imaging. J. Nucl. Med. 2021, 62 (Suppl. 1), 3038. [Google Scholar]
- Kang, J.; Yoon, C.; Lee, J.; Kye, S.B.; Lee, Y.; Chang, J.H.; Kim, G.D.; Yoo, Y.; Song, T.K. A System-on-Chip Solution for Point-of-Care Ultrasound Imaging Systems: Architecture and ASIC Implementation. IEEE Trans. Biomed. Circuits Syst. 2016, 10, 412–423. [Google Scholar] [CrossRef]
- Kim, T.; Fool, F.; Dos Santos, D.S.; Chang, Z.-Y.; Noothout, E.; Vos, H.J.; Bosch, J.G.; Verweij, M.D.; de Jong, N.; Pertijs, M.A.P. Design of an ultrasound transceiver asic with a switching-artifact reduction technique for 3D carotid artery imaging. Sensors 2021, 21, 150. [Google Scholar] [CrossRef]
- Rothberg, J.M.; Ralston, T.S.; Rothberg, A.G.; Martin, J.; Zahorian, J.S.; Alie, S.A.; Sanchez, N.J.; Chen, K.; Chen, C.; Thiele, K.; et al. Ultrasound-on-chip platform for medical imaging, analysis, and collective intelligence. Proc. Natl. Acad. Sci. USA 2021, 118, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Sheng, D.; Liu, C.-H.; Chen, S.-Y.; Song, B.-Y.; Chiu, Y.-C.; Cai, M.-H. DLL-based transmit beamforming IC for high-frequency ultrasound medical imaging system. In Proceedings of the 2021 IEEE International Conference on Consumer Electronics-Taiwan (ICCE-TW), Penghu, Taiwan, China; 2021; pp. 1–2. [Google Scholar]
- Cela, J.M.; Freixas, L.; Lagares, J.I.; Marin, J.; Martinez, G.; Navarrete, J.; Oller, J.C.; Perez, J.M.; Rato-Mendes, P.; Sarasola, I.; et al. A Compact Detector Module Design Based on FlexToT ASICs for Time-of-Flight PET-MR. IEEE Trans. Radiat. Plasma Med. Sci. 2018, 2, 549–553. [Google Scholar] [CrossRef]
- Sánchez, D.; Gómez, S.; Mauricio, J.; Freixas, L.; Sanuy, A.; Guixé, G.; López, A.; Manera, R.; Marin, J.; Pérez, J.M.; et al. HRFlexToT: A High Dynamic Range ASIC for Time-of-Flight Positron Emission Tomography. IEEE Trans. Radiat. Plasma Med. Sci. 2021. [Google Scholar] [CrossRef]
- Nemallapudi, M.V.; Rahman, A.; Chen, A.E.-F.; Lee, S.-C.; Lin, C.-H.; Chu, M.-L.; Chou, C.-Y. Positron emitter depth distribution in PMMA irradiated with 130 MeV protons measured using TOF-PET detectors. IEEE Trans. Radiat. Plasma Med. Sci. 2021, 7311, 1. [Google Scholar] [CrossRef]
- Shen, W.; Harion, T.; Schultz-Coulon, H.C. STiC an ASIC CHIP for silicon-photomultiplier fast timing discrimination. In Proceedings of the IEEE Nuclear Science Symposuim & Medical Imaging Conference, Knoxville, TN, USA, 30 October–6 November 2010; pp. 406–408. [Google Scholar] [CrossRef]
- Gandhare, S.; Karthikeyan, B. Survey on FPGA architecture and recent applications. In Proceedings of the 2019 International Conference on Vision Towards Emerging Trends in Communication and Networking (ViTECoN), Vellore, India, 30–31 March 2019; pp. 1–4. [Google Scholar]
- Romoth, J.; Porrmann, M.; Rückert, U. Survey of FPGA applications in the period 2000–2015 (Tech. Rep.). Center of Excellence Cognitive Interaction Technology, Bielefeld University: Bielefeld, Germany, 2017; pp. 1–43. [Google Scholar]
- Wong, H.S.P.; Akarvardar, K.; Antoniadis, D.; Bokor, J.; Hu, C.; King-Liu, T.J.; Mitra, S.; Plummer, J.D.; Salahuddin, S. A Density Metric for Semiconductor Technology [Point of View]. Proc. IEEE 2020, 108, 478–482. [Google Scholar] [CrossRef]
- Al-Ali, F.; Gamage, T.D.; Nanayakkara, H.W.T.S.; Mehdipour, F.; Ray, S.K. Novel casestudy and benchmarking of AlexNet for edge AI: From CPU and GPU to FPGA. In Proceedings of the 2020 IEEE Canadian Conference on Electrical and Computer Engineering (CCECE), London, ON, Canada, 30 August–2 September 2020; pp. 1–4. [Google Scholar]
- Xiong, C.; Xu, N. Performance comparison of BLAS on CPU, GPU and FPGA. In Proceedings of the 2020 IEEE 9th Joint International Information Technology and Artificial Intelligence Conference (ITAIC), Chongqing, China, 11–13 December 2020; Volume 9, pp. 193–197. [Google Scholar]
- Asano, S.; Maruyama, T.; Yamaguchi, Y. Performance comparison of FPGA, GPU and CPU in image processing. In Proceedings of the 2009 International Conference on Field Programmable Logic and Applications, Prague, Czech Republic, 31 August–2 September 2009; pp. 126–131. [Google Scholar] [CrossRef]

| CPU | GPU | DSP | FPGA | ASIC | |
|---|---|---|---|---|---|
| Central Processing Unit | Graphics Processing Unit | Digital Signal Processors | Field Programmable Gate Arrays | Application-Specific Integrated Circuit | |
| Overview | Traditional sequential processor, optimized for executing general computational instructions. | Designed mainly for graphical outputs; now used in a wide range of computationally heavy applications (e.g., training AI models). | Dedicated to processing digital signals such as audio. Designed to perform mathematical functions such as addition and subtraction at high speed with minimal energy consumption. | IP blocks and logic elements that can be modified or configured in the field by the designer. | A customized embedded circuit designed for a specific application. |
| Processing | Single and Multicore MCUs MPUs. | Thousands of identical processor cores. | A parallel architecture is suitable as it is scalable and allows performance based on the number of processors in the system. Current multicore DSPs can be combined with other structures (e.g., CPU cores (ARM) and DSP cores). | SoCs include hard or soft IP cores, and they are configured for application. | Configured for specific application, and they could include third-party IP cores. |
| Programming | OSes, multitude of high-level languages via APIs; OpenMP, Cilk++, TBB, RapidMind and OpenCL; assembly language. | OpenCL, HIP (AMD) and NVIDIA CUDA API allow general-purpose programming (e.g., C, C++, Python). | C, C++ | Traditionally HDL (Verilog VHDL); The newer systems include C/C++ via OpenCL and Vitis. | Application specific. For example, Google Coral TPUs to work with Tensorflow tensors or CPU manufacturers such as Intel include tools with ASICs. |
| Peripherals | Analog and digital peripherals in MCUs include digital bus interfaces. | Very limited, e.g., only cache memory and interconnection among GPUs (multi-GPUs). | Program memory, data memory, BUS. | Transceiver blocks and configurable I/O banks. | Tailored to application. ASICs can include standard connectors such as USB, Thunderbolt, Ethernet, etc. |
| Strengths | High versatility, multitasking and ease of programming. | High-performance processing in specific applications, such as video processing, image and signal analysis and neural network training, deep learning. | They have an optimized architecture to perform the computational operations in digital signal processing with low cost and high performance per watt. | Application-specific configuration, and this configuration could be changed for new fields. High performance per watt. Allows parallel operation. | Tailor-made for the application with an optimal combination of performance and energy consumption. |
| Weaknesses | The operating system adds a large overhead. It is optimized for sequential processing with limited parallelism. | Not suitable for all programs or algorithms. It is energy-intensive and becomes hot. Problems must be adapted to take advantage of parallelism. | They barely offer any flexibility. | Long development time and difficult programming. Low performance in sequential operations. Not optimal for floating-point operations. | Long development time with a high price tag. Cannot be changed once designed. |
| Most common medical usage | Image reconstruction, registration, segmentation. | 3D PET image reconstruction, EM simulations, Pseudo-CT synthesis with MRI. | Ultrasound filtering, image registration, image enhancement. | Optimized 3D MRI image segmentation, TC/PET/MRI image reconstruction, image registration | Time-of-flight (TOF) PET, X-ray hybrid pixel detector, CZT and CdTe detectors for SPECT, specific US solutions. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Alcaín, E.; Fernández, P.R.; Nieto, R.; Montemayor, A.S.; Vilas, J.; Galiana-Bordera, A.; Martinez-Girones, P.M.; Prieto-de-la-Lastra, C.; Rodriguez-Vila, B.; Bonet, M.; et al. Hardware Architectures for Real-Time Medical Imaging. Electronics 2021, 10, 3118. https://doi.org/10.3390/electronics10243118
Alcaín E, Fernández PR, Nieto R, Montemayor AS, Vilas J, Galiana-Bordera A, Martinez-Girones PM, Prieto-de-la-Lastra C, Rodriguez-Vila B, Bonet M, et al. Hardware Architectures for Real-Time Medical Imaging. Electronics. 2021; 10(24):3118. https://doi.org/10.3390/electronics10243118
Chicago/Turabian StyleAlcaín, Eduardo, Pedro R. Fernández, Rubén Nieto, Antonio S. Montemayor, Jaime Vilas, Adrian Galiana-Bordera, Pedro Miguel Martinez-Girones, Carmen Prieto-de-la-Lastra, Borja Rodriguez-Vila, Marina Bonet, and et al. 2021. "Hardware Architectures for Real-Time Medical Imaging" Electronics 10, no. 24: 3118. https://doi.org/10.3390/electronics10243118
APA StyleAlcaín, E., Fernández, P. R., Nieto, R., Montemayor, A. S., Vilas, J., Galiana-Bordera, A., Martinez-Girones, P. M., Prieto-de-la-Lastra, C., Rodriguez-Vila, B., Bonet, M., Rodriguez-Sanchez, C., Yahyaoui, I., Malpica, N., Borromeo, S., Machado, F., & Torrado-Carvajal, A. (2021). Hardware Architectures for Real-Time Medical Imaging. Electronics, 10(24), 3118. https://doi.org/10.3390/electronics10243118









