Abstract
The global demand for herbal cosmetics is vastly increasing due to their health benefits and relative safety. Glycyrrhiza spp. extracts are used in cosmetic preparations due to their skin-whitening, antisensitizing, and anti-inflammatory properties. The aim of this work is to computationally evaluate the bioactive constituents of licorice (Glycyrrhiza glabra L.) that possess antioxidant, anti-inflammatory, and dermatocosmetic activities, and elucidate the dynamics of their molecular targets. The used methods are skin permeability prediction, target prediction, molecular docking, and molecular dynamic simulation (MDS). The results show that, at a skin permeation cut-off value of −6.0 cm/s, nine phytoconstituents of licorice (furfuraldehyde, glucoliquiritin apioside, glycyrrhizin, isoliquiritin, licopyranocoumarin, licuraside, liquiritigenin, liquiritin, and liquiritin apioside) were workable. Molecular target prediction results indicate probability for tyrosinase, 11-beta-hydroxysteroid dehydrogenase 1 (HSD11B1), monoamine oxidase B, steroid 5-alpha-reductase 1, and cyclo-oxygenase-1. On the basis of molecular docking, glucoliquiritin apioside and glycyrrhizin had the best antioxidant, anti-inflammation, and dermatocosmetic activities. MDS results show that the complexes had good stability, and MMGBSA results indicate that the complexes had satisfactory binding energy. Overall, this study demonstrates that licorice extracts are potential antioxidants that could enhance histological dermal and epidermal properties, and reduce the level of inflammatory and wrinkling markers.
1. Introduction
Cosmetics are any bioactive-containing preparation that is intended for use on the external surface area of human or animal bodies with the aim of cleansing, perfuming, protecting, or treating certain diseases [1]. Cosmetic products from natural sources such as plants usually do not pose health risks, but due to exposure to some hazardous agents in the environment such as allergens, toxins, carcinogens, and endocrine disruptors, there is a need for the authentication of plant materials for cosmetic applications [2]. The global demand for herbal cosmetics is vastly increasing due to their health benefits [3]. There is ongoing research in the cosmetic industry to discover new tropically sourced products and ingredients as their raw materials that often have functional properties due to the differential climatic and topographical settings [4].
Phytocosmetics is a segment of cosmetology that utilizes plant species for cosmetic purposes such as beautification and medication for skin diseases, which include abscesses, boils, eczema burns, pimples, and ringworms [5,6]. Particularly, the use of specific beauty and medication recipes remains common in countries such as Nigeria [7], Cameroon [8], South Africa [5], Sri Lanka [3], France [9], Morocco, Brazil and Portugal [10], and Egypt, Pakistan, and India [11].
Glycyrrhiza spp. are herbaceous perennial plants that grow in subtropical and temperate zones. The Glycyrrhiza genus belonging to the Leguminosae family (also known as Fabaceae) consists of more than 30 globally widely distributed species. In the Glycyrrhiza genus, G. glabra L., G. uralensis Fisch., and G. inflata Bat. are the most investigated species with diverse nutritional and pharmacological benefits [12]. Glycyrrhiza spp. extracts are used in cosmetic preparations due to their skin-whitening, antisensitizing, and anti-inflammatory properties [13,14].
Licorice (Glycyrrhiza glabra L., Fabaceae) contains a wide array of natural bioactive products. According to Husain et al. [15], 50% of the dry weight of licorice roots is due to water-soluble compounds, sugars (5–15% glucose, mannitol, and sucrose), starch (25–30%), glycyrrhizin (10–16%), amines (1–2% asparagine, betaine, and choline), and sterols (stigmasterol and β-sitosterol). An array of licorice phytochemicals were investigated for their whitening and antioxidant effects in the treatment of pigmented skin disease. Bioactive compounds in licorice root extracts (such as liquiritigenin, isoliquiritigenin, liquiritin, isoliquiritin, liquiritin apioside, licuraside glucoliquiritin apioside, and glabridin) could protect the skin against oxidative stress injuries, and efficiently reduce the symptoms of atopic dermatitis [16,17,18,19,20].
There are some dermatological dysfunctions that have no actual treatment, including melasma, lentigines, postinflammatory hyperpigmentation, age spots, solar lentigo, Prurigo pigmentosa, Café-au-lait, Linea niagra, and freckles [21]. Computational or in silico techniques are fundamental to nonanimal chemical safety assessment, as they could be applied to internal exposure, hazard identification [22], and unraveled bioactive compounds that could be useful in treating untreated medical conditions. In silico tools and resources have recently gained relevance in the toxicokinetic study of cosmetic ingredients to provide insights and serve as the foundation of the next generation of risk assessment [23]. The aim of this study is to computationally evaluate the constituents of licorice that possessed antioxidant, anti-inflammatory, and dermatocosmetic activities, and elucidate the dynamics of their molecular targets.
2. Materials and Methods
2.1. Ligand Preparation
Major phytochemicals in Glycyrrhiza glabra (licorice) were identified from the literature [20,24,25], and their structures were obtained from the NCBI PubChem Compound database (https://pubchem.ncbi.nlm.nih.gov/) (accessed on 21 January 2023) in SMILES format.
2.2. In Silico Pharmacokinetics
The SMILESs of each of the ligands were used for in silico absorption, distribution, metabolism, and excretion (ADME) screening on a SwissADME server (http://www.swissadme.ch) (accessed on 7 February 2023) [26] that was performed with the default parameters. Predicted skin permeation log kp from the in silico pharmacokinetics was based on a model by Potts and Guy [27] according to the following equation:
where MW is the molecular weight of the compound, and log kow (or log Po/w) is the octanol-water partition coefficient, a physicochemical constant used to describe the lipophilicity of the penetrant [28]. Compounds with high skin permeation were noted for further analysis. Hierarchical clustering analysis was also performed on a ChemMine web server (http://chemmine.ucr.edu/) (accessed on 18 April 2023) as previously described by Fatoki et al. [29] using the SMILESs of the ligands.
log kp (cm/s) = 0.71 ∗ log kow − 0.0061 ∗ MW − 6.3
2.3. In Silico Target Prediction
The selected ligands that possessed a high skin-permeability coefficient based on the predicted pharmacokinetics were used for target prediction on a SwissTargetPrediction server (http://www.swisstargetprediction.ch/) (accessed on 13 February 2023), where Homo sapiens was designated as the target organism [30].
2.4. Molecular Docking Studies
Molecular docking studies were conducted as described by Fatoki et al. [29]. Briefly, the three-dimensional structures of 10 standard molecular target proteins for antioxidant (superoxide dismutase and glutathione peroxidase), anti-inflammation (11B-hydroxysteroid dehydrogenase 1, lipoxygenase, cyclo-oxygenase and inducible nitric oxide synthase), and dermatocosmetic (tyrosinase, collagenase, hyaluronidase and elastase) activities were obtained from www.rcsb.org/pdb on the basis of literature reports [9,10,31]. The structure of ligands that possessed high skin permeation were subjected to 3D structure optimization using ACDLab/Chemsketch software and saved in mol format. PyMol software was used for ligand file conversion from mol into pdb and for the preparation of protein chain A with the removal of water and existing ligands. Both ligands and proteins were prepared for docking using AutoDock Tools (ADT) v1.5.6 [32] with the default settings, and the output file was saved in pdbqt format. Molecular docking program AutoDock Vina v1.2.3 [33,34] was used for the docking experiment. After docking, close interactions of the binding of the target with the ligands were analyzed and visualized using ezLigPlot on an ezCADD web server (https://dxulab.org/software) (accessed on 18 February 2023) [35].
2.5. Protein–Protein Interaction Analysis
To established the relationship between the predicted targets of licorice phytochemicals with high skin permeability, the gene IDs of 10 standard molecular target proteins for anti-inflammation, antioxidant, and dermatocosmetic activities were analyzed on the basis of literature reports [9,10,31] in combination with the predicted targets in humans for the protein–protein interaction (PPI) profile on a STRING web server (https://string-db.org/) (accessed on 13 February 2023) [36].
2.6. Protein–Ligand Molecular Dynamics Simulation
Molecular dynamics simulations were performed for 100 nanoseconds using Desmond, Schrödinger LLC [37,38,39]. The initial stages of the protein and ligand complexes for molecular dynamics simulation were obtained from the docking studies. Protein–ligand complexes were preprocessed using Maestro’s protein preparation wizard, which also included complex optimization and minimization. All systems were prepared with the System Builder tool. A solvent model with an orthorhombic box was selected as the Transferable Intermolecular Interaction Potential 3 Points (TIP3P). The Optimized Potential for Liquid Simulations (OPLS)-2005 force field was used in the simulation [40]. The models were made neutral by adding 0.15 M NaCl counterions to mimic physiological conditions. The NPT ensemble (isothermal–isobaric: moles (N), pressure (P), and temperature (T) were conserved) at 300 K temperature and 1 atm pressure was select for a complete simulation. The models were relaxed before the simulation. The trajectories were saved after every 100 ps during the simulation, and the post-simulation analysis of the trajectories was conducted to determine the root-mean-square deviation (RMSD), root-mean-square fluctuation (RMSF), radius of gyration (Rg), solvent accessibility surface area (SASA), and protein–ligand interaction profile [37,38]. Prime molecular mechanics/generalized Born surface area (MMGBSA) was calculated as follows:
where protein* means a protein from the optimized complex; ligand* means “a ligand from the optimized complex; NS means no strain, which is the binding/interaction energy without accounting for conformational receptor and ligand changes needed to form the complex [41,42].
MMGBSA ΔGbind = ΔGcomplex − ΔGprotein − ΔGligand
MMGBSA ΔGbind (NS) = ΔGcomplex − ΔGprotein* − ΔGligand* or
MMGBSA ΔGbind (NS) = MMGBSA ΔGbind − ΔGprotein_strain − ΔGligand_strain
3. Results
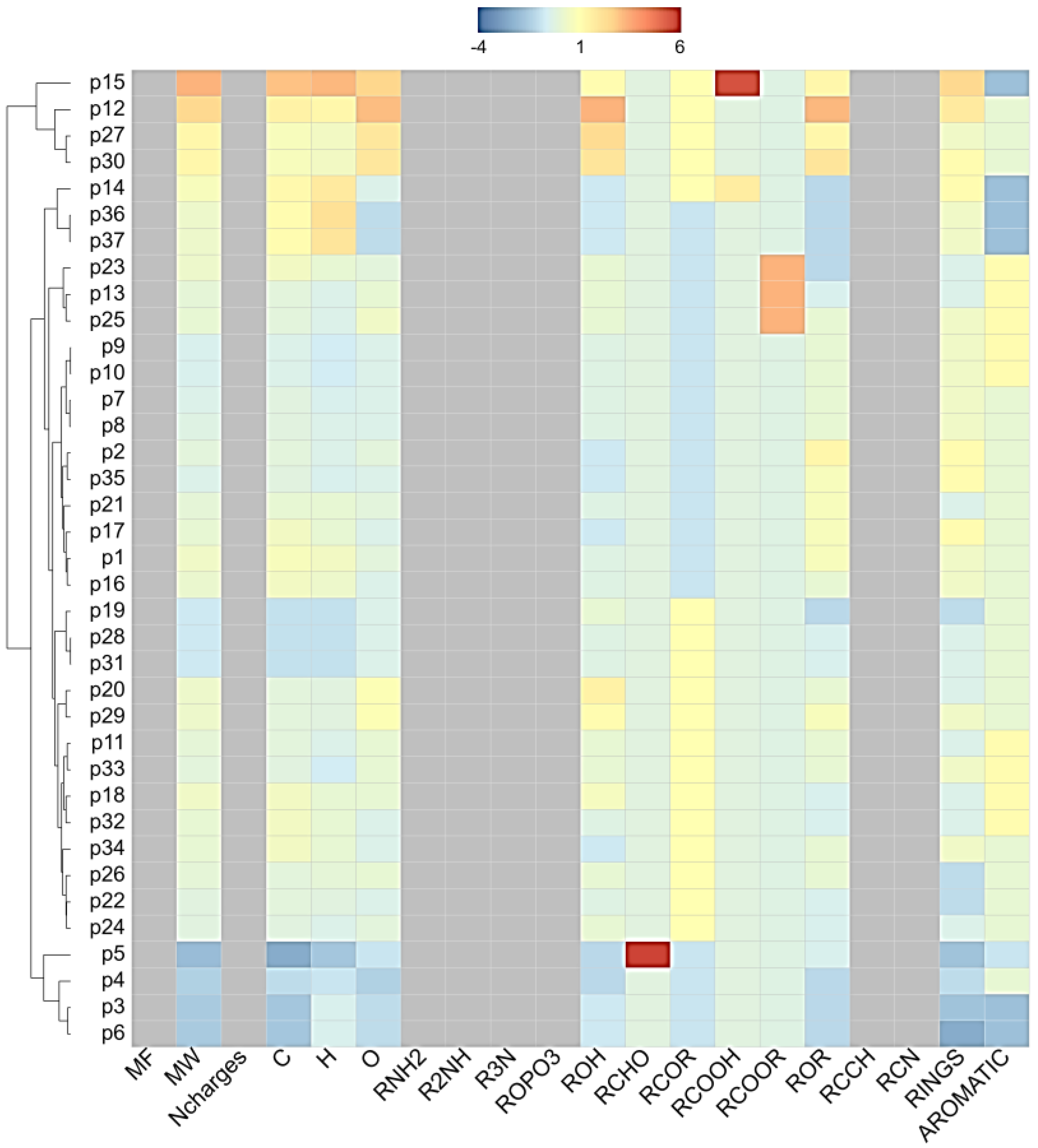
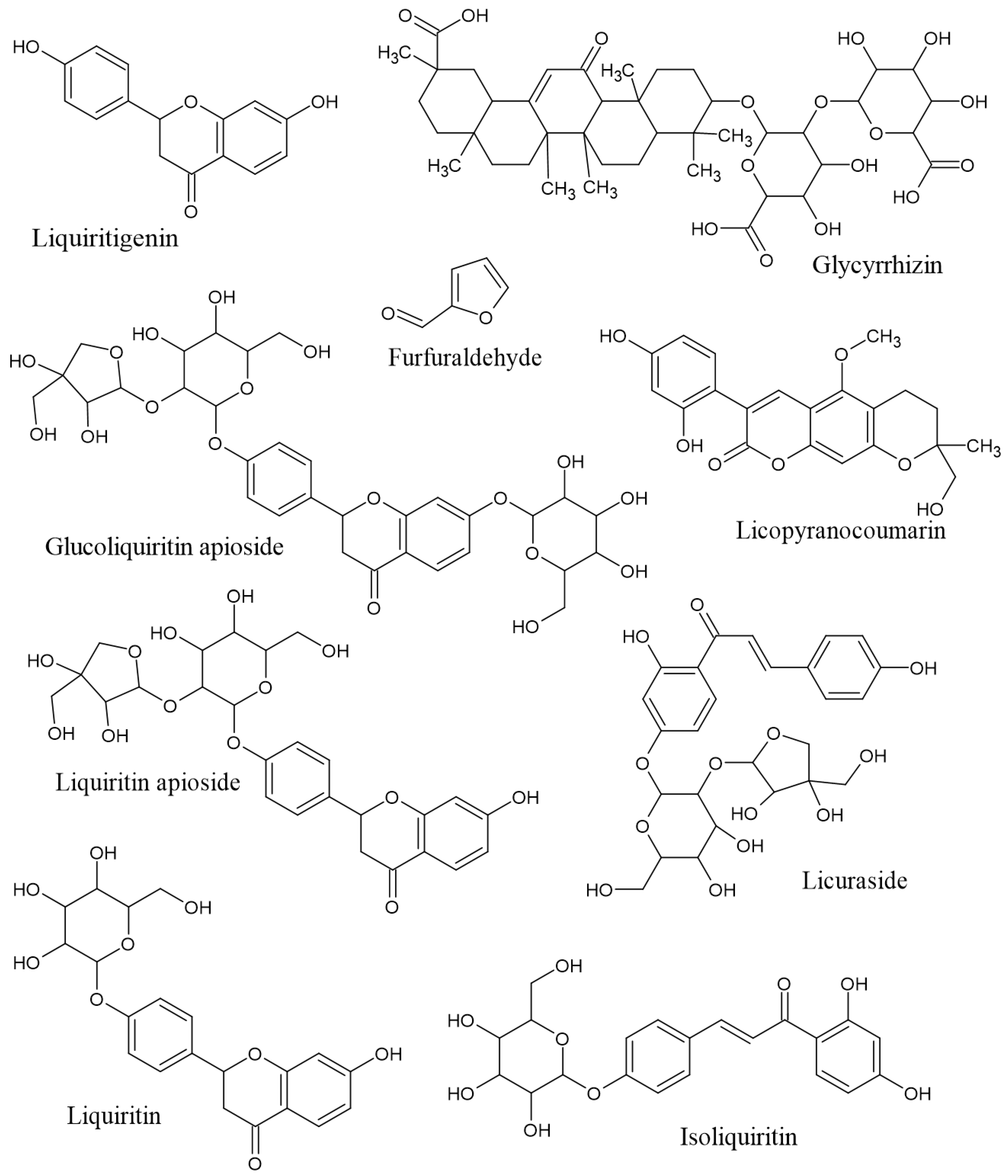
Using skin permeation (log kP) at a cut-off value of –6.0 cm/s, 9 phytoconstituents of licorice (furfuraldehyde, glucoliquiritin apioside, glycyrrhizin, isoliquiritin, licopyranocoumarin, licuraside, liquiritigenin (4′,7-dihydroxyflavanone), liquiritin, and liquiritin apioside) were capable of dermatocosmetic activity (Table 1). The overall association based on the cluster of the physicochemical properties is presented in Figure 1. In relation to the ADME results, the clustering results reveal that glucoliquiritin apioside (p12), glycyrrhizin (p15, triterpenoid saponins), licuraside (p27, flavonoid glycoside) and liquiritin apioside (p30, flavanone apioside) serve as lead markers for cosmetic purposes, followed by isoliquiritin (p20, flavanoid glucoside) and liquiritin (p29, flavanone glucoside), liquiritigenin (p28, flavanone), lycopyranocoumarin (p25, isoflavonoid coumarin), and furfuraldehyde (p5, aldehyde). Most of these nine compounds showed unique patterns of low gastrointestinal absorption and were not permeable through the blood–brain barrier. The chemical structures of these 9 phytoconstituents of licorice are shown in Figure 2.

Table 1.
Predicted pharmacokinetic properties of licorice phytochemicals.
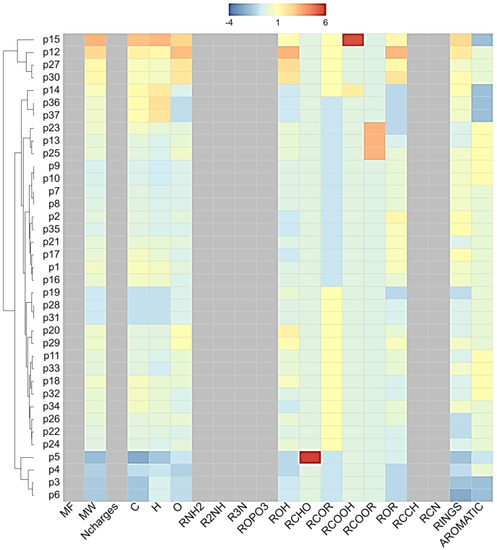
Figure 1.
Hierarchical clustering results. p1-p37 is equivalent to SN in Table 1. The used parameter options are as follows: heat map: distance matrix, Linkage method: average. Physicochemical-property heat map: ChemmineR Properties. Color and display property values: Z scores.
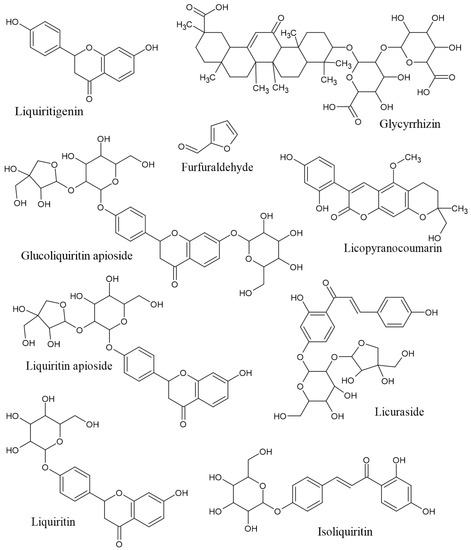
Figure 2.
Chemical structures of the nine compounds in licorice with potential dermatocosmetic activities.
The identified molecular targets with probability greater than or equal to 40% in this study were cytochrome P450 19A1, 11-beta-hydroxysteroid dehydrogenases 1 and 2, monoamine oxidase B, estradiol 17-beta-dehydrogenase 1, estrogen receptors alpha and beta, carbonic anhydrase IV/VII/XII, and carbonyl reductase (Table 2). The most represented molecular targets in the 9 compounds were cytochrome P450 19A1, epoxide hydratase, tyrosinase, monoamine oxidase B, Steroid 5-alpha-reductase 1 and cyclo-oxygenase-1.

Table 2.
Molecular targets of dermally active constituents predicted from SwissTargetPrediction.
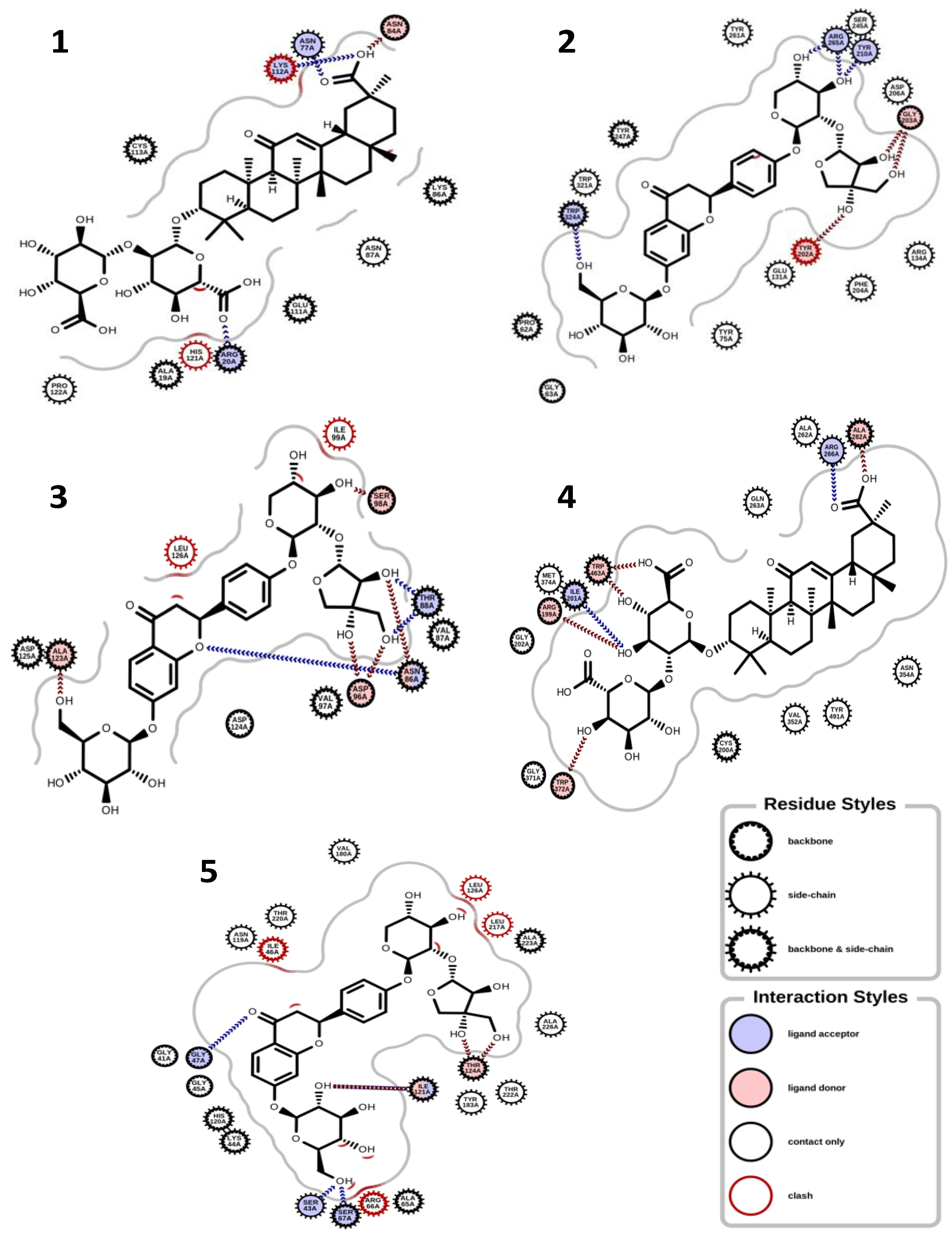
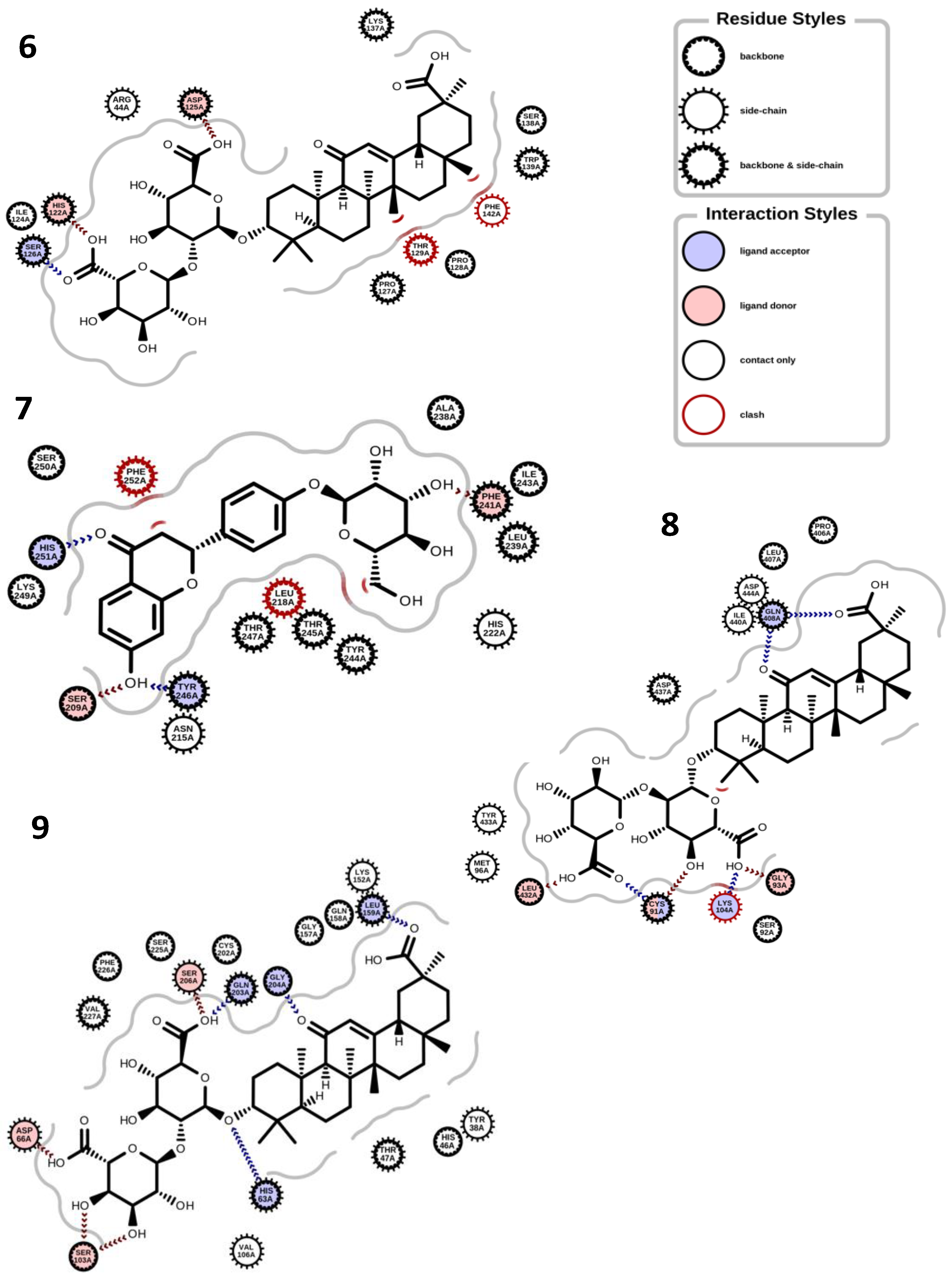
In terms of the molecular docking of the 9 compounds, glucoliquiritin apioside and glycyrrhizin had the best antioxidant, anti-inflammation, and dermatocosmetic activities (Table 3). The binding affinity of glucoliquiritin apioside with 11β-hydroxysteroid dehydrogenase 1 was 10.900 kcal.mol−1, followed by that of isoliquiritin (−9.553 kcal.mol−1), while glucoliquiritin apioside had binding affinity of −9.964 kcal.mol−1, which was less than that of standard tannic acid (−15.980 kcal.mol−1), and higher than those of kojic acid and quercetin. Glycyrrhizin had the highest binding affinity against elastase (−10.100 kcal.mol−1), cyclo-oxygenase (−9.427 kcal.mol−1), tyrosinase (−8.768 kcal.mol−1), glutathione peroxidase (−8.409 kcal.mol−1) and lipoxygenase (−8.979 kcal.mol−1), followed by glucoliquiritin apioside. Glycyrrhizin had the highest binding affinity for inducible nitric oxide synthase (−10.510 kcal.mol−1), followed by liquiritigenin (−9.840 kcal.mol−1). Moreover, liquiritin and liquiritin apioside showed nearly the same high binding affinity against collagenase (about −9.9 kcal.mol−1), but less than that of standard tannic acid (−13.110 kcal.mol−1), and higher than that of kojic acid and quercetin. Overall, furfuraldehyde showed the lowest binding affinities, while glucoliquiritin apioside, glycyrrhizin, liquiritin, and liquiritin apioside had the best multitarget binding affinities. The interactions of some of the docking poses are shown in Figure 3 and Figure 4.

Table 3.
Molecular docking properties and binding affinity scores.
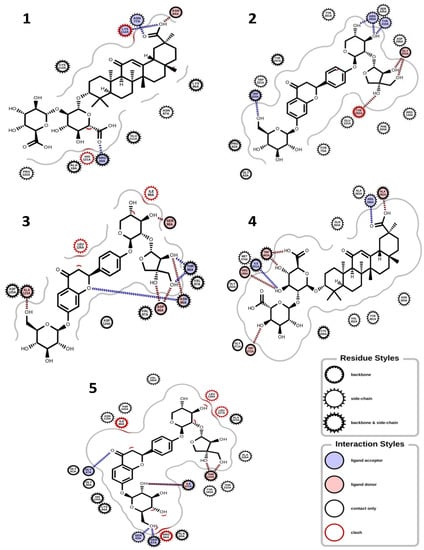
Figure 3.
Binding interactions. (1) Glycyrrhizin and glutathione peroxidase (PDB ID: 2F8A); (2) glucoliquiritin apioside and hyaluronidase (PDB ID: 2PE4); (3) glucoliquiritin apioside and superoxide dismutase (PDB ID: 3HFF); (4) glycyrrhizin and Inducible nitric oxide synthase (PDB ID: 4CX7); (5) glucoliquiritin apioside and 11B-hydroxysteroid dehydrogenase 1 (PDB ID: 4YYZ).
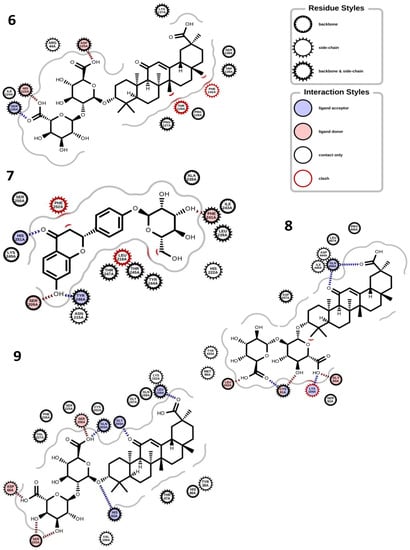
Figure 4.
Binding interactions. (6) Glycyrrhizin and cyclo-oxygenase (PDB ID: 5KIR); (7) liquiritin and collagenase (PDB ID: 5UWL); (8) glycyrrhizin and tyrosinase (PDB ID: AF-P14679-F1); (9) glycyrrhizin and elastase (PDB ID: AF-Q9UNI1-F1).
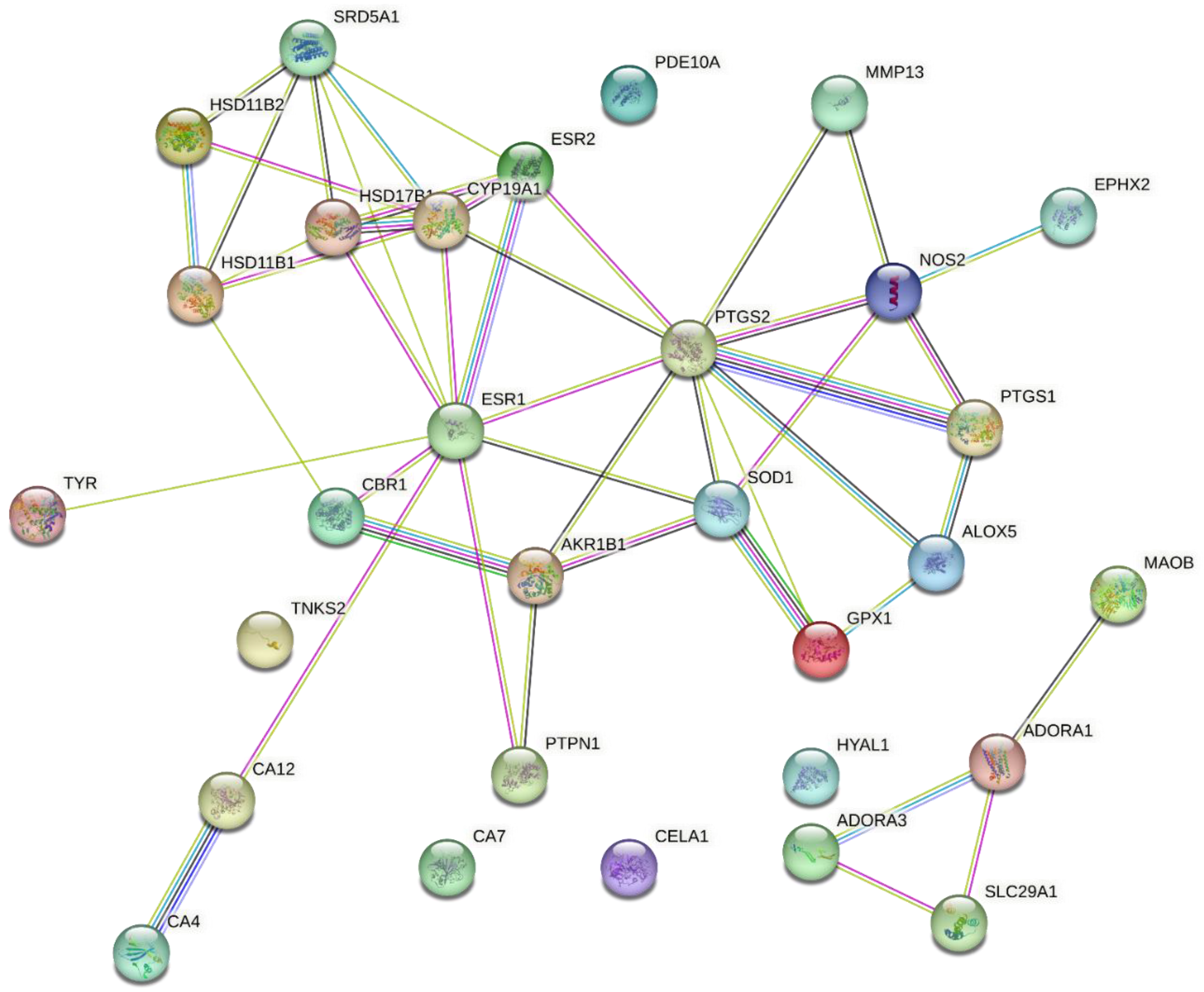
We retrieved the protein–protein interaction network for the predicted molecular targets of the potential dermally active compounds and standard molecular targets for antioxidant, anti-inflammatory, and dermatocosmetic activities from the literature. As shown in Figure 5, a relationship exists among estrogen receptor alpha (ESR1), cytochrome P450 19A1 (CYP19A1), cyclo-oxygenase 2 (PTGS2), inducible nitric oxide synthase (NOS2), superoxide dismutase 1 (SOD1), and collagenase (MMP13).
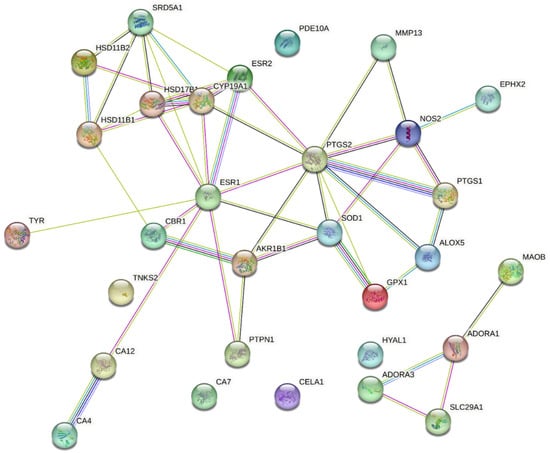
Figure 5.
Protein–protein interaction network for predicted and standard molecular targets.
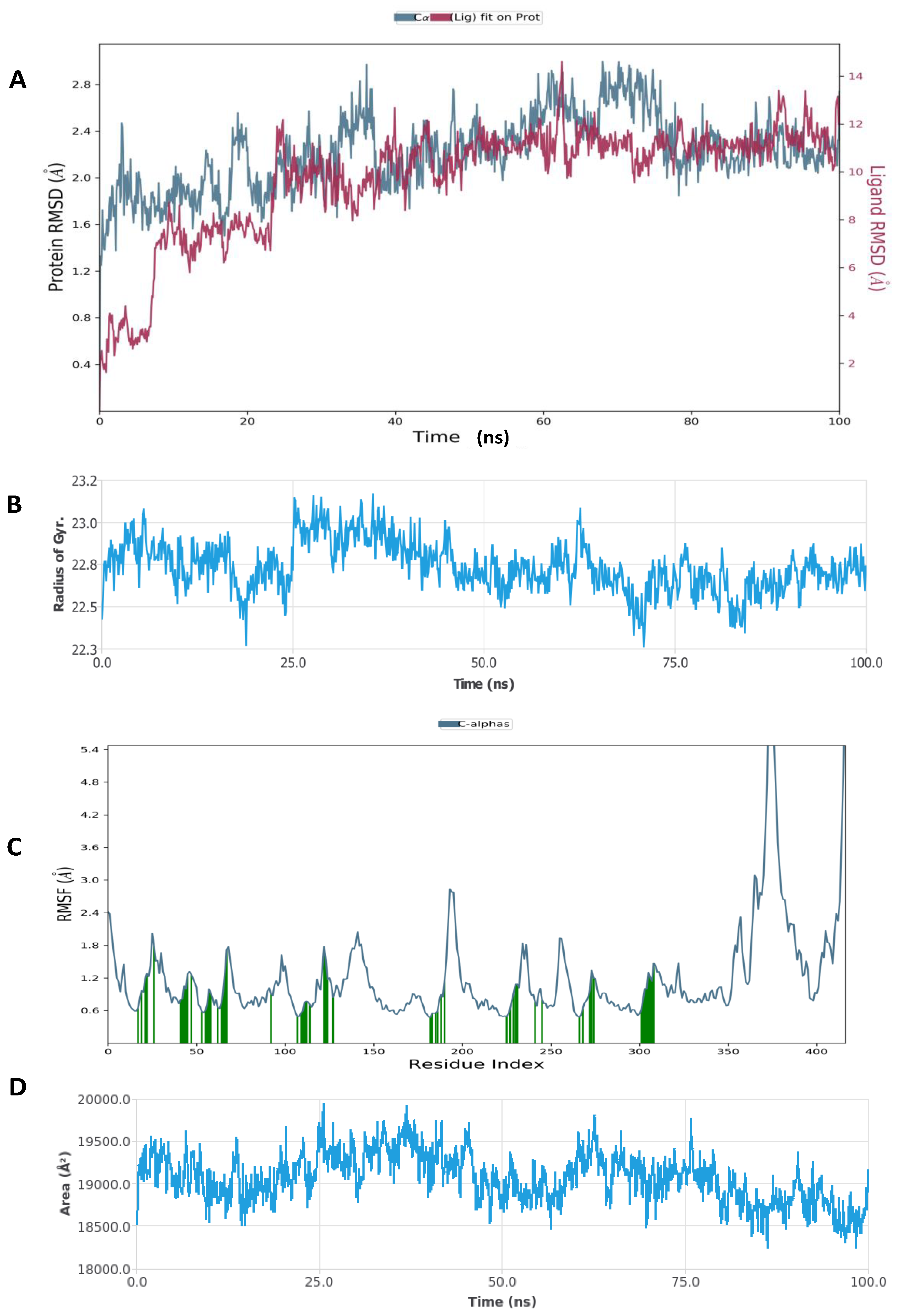
The Desmond simulation package in Schrodinger was used for the MD simulation of two selected results of the docked complex (Figure 6, Figure 7, Figure 8 and Figure 9). The hyaluronidase protein complex with glucoliquiritin apioside had an RMSD of about 1.6 Å, and the protein was quite stable during 20–100 ns of simulation time, while the RMSD ligand was stable at 25–100 ns (Figure 6). Overall, the ligand was stable during the simulation. Hyaluronidase had Rg < 0.9 Å, the RMSF was mostly significant at the 190–200 and C-terminal amino acid residues, and the total SASA was about 1800 Å2. Figure 7 shows the high interaction of hyaluronidase with glucoliquiritin apioside that occurred on ASN39, GLY63, ILE73, SER76, SER77, GLN78, ASP129, GLU131, TYR202, ASP292 and TRP324 amino acid residues, and the profiles of glucoliquiritin apioside during the simulation.
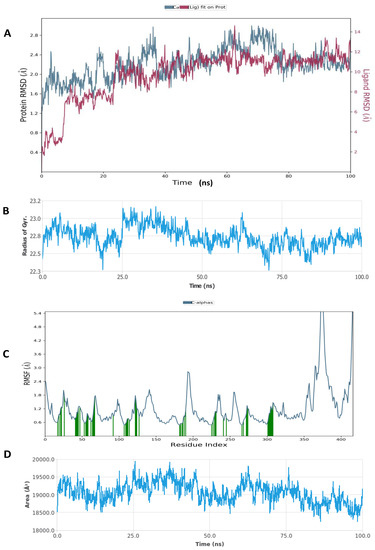
Figure 6.
Molecular dynamic simulation (MDS) results. (A) RMSD of hyaluronidase (2PE4) with glucoliquiritin apioside. (B) Rg of hyaluronidase. (C) RMSF of hyaluronidase. (D) SASA of hyaluronidase.
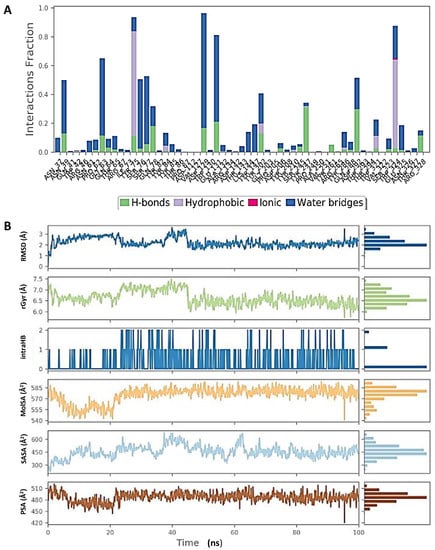
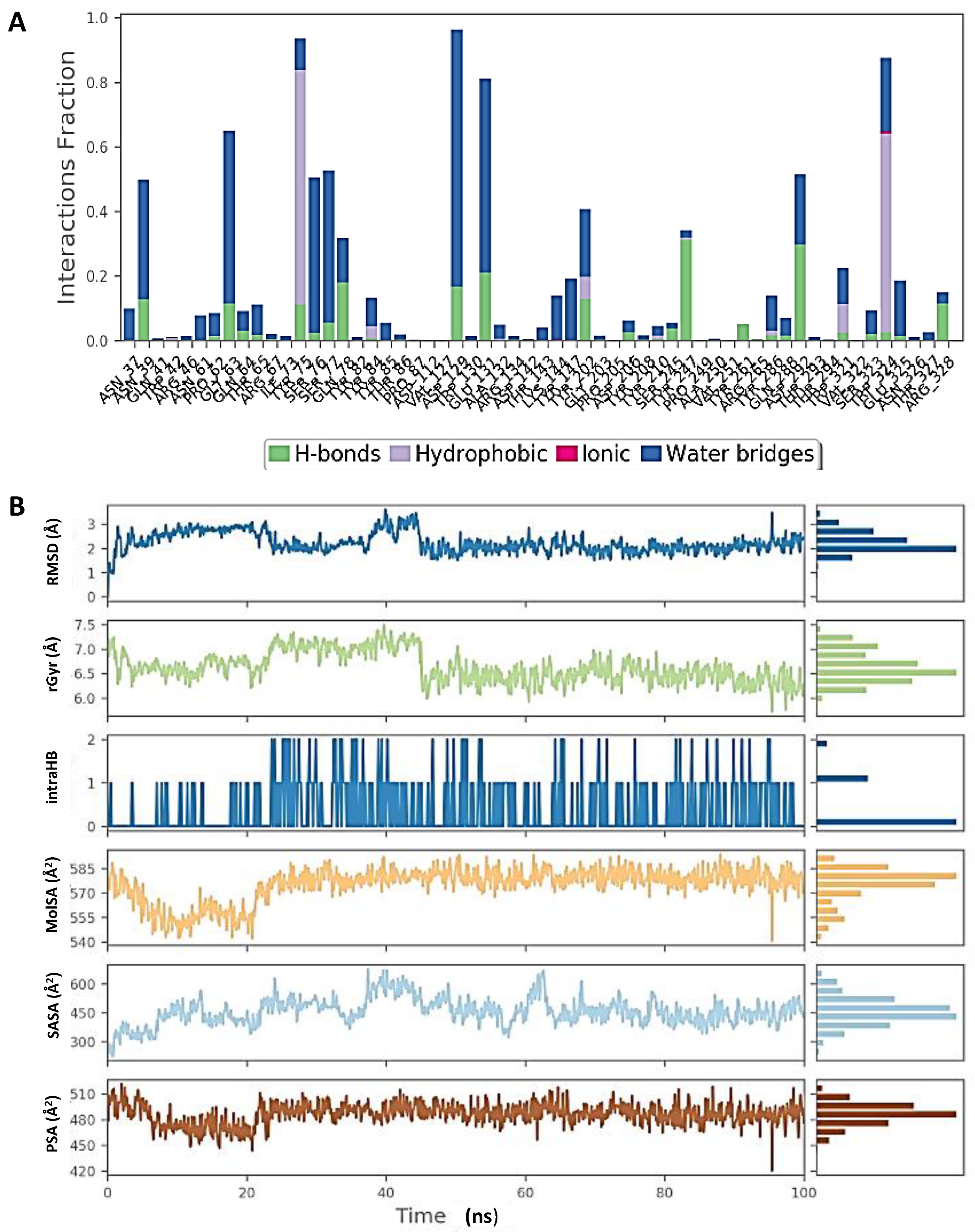
Figure 7.
Molecular dynamic simulation (MDS) results. (A) Interaction profile of the contact of hyaluronidase (2PE4) with glucoliquiritin apioside. (B) Ligand (glucoliquiritin apioside) profile (RMSD, Rg, intramolecular hydrogen bonds (intraHB), molecular surface area (MolSA), SASA, and polar surface area (PSA) during simulation.
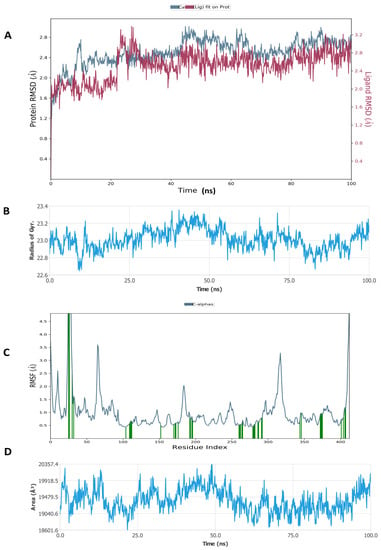
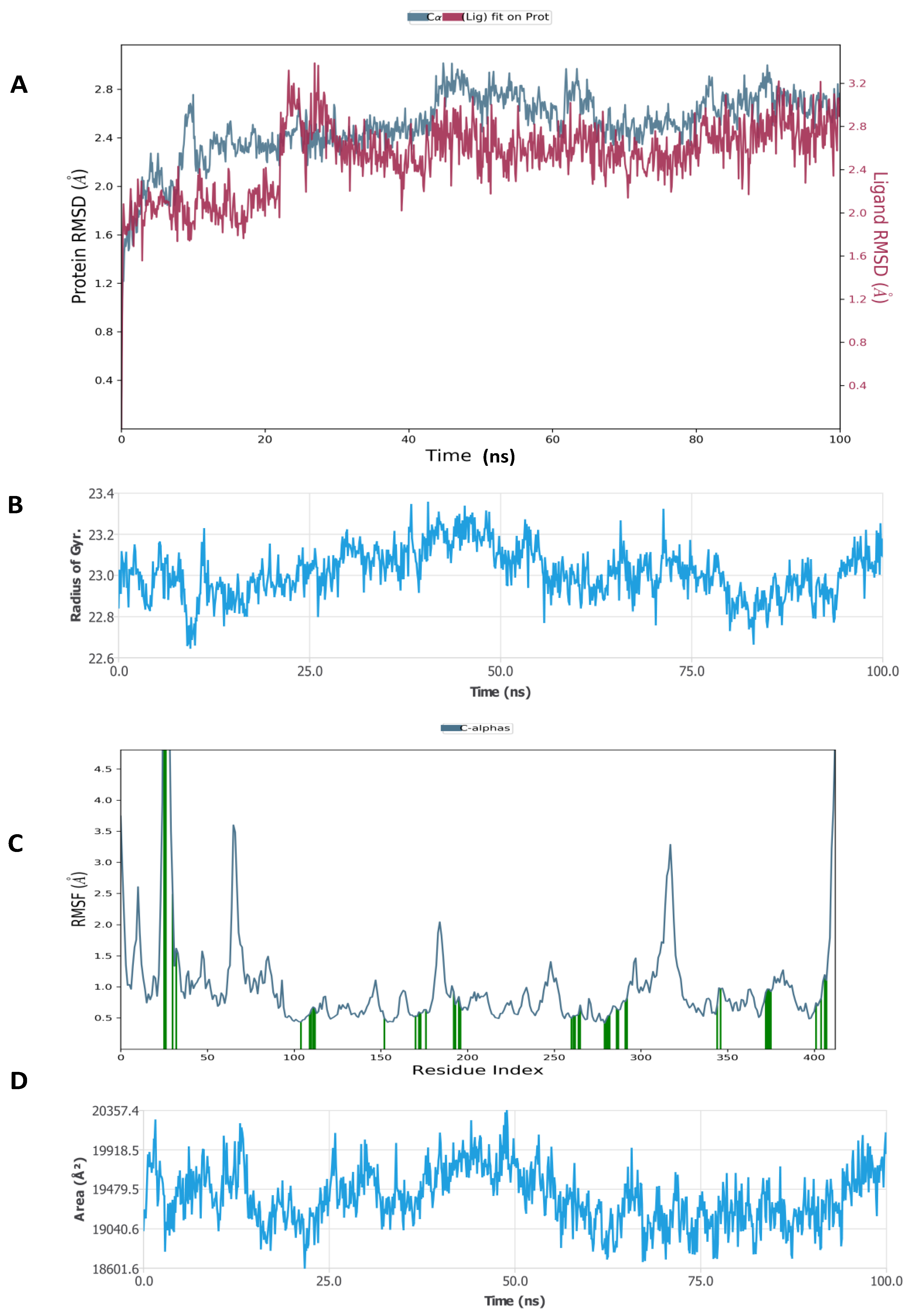
Figure 8.
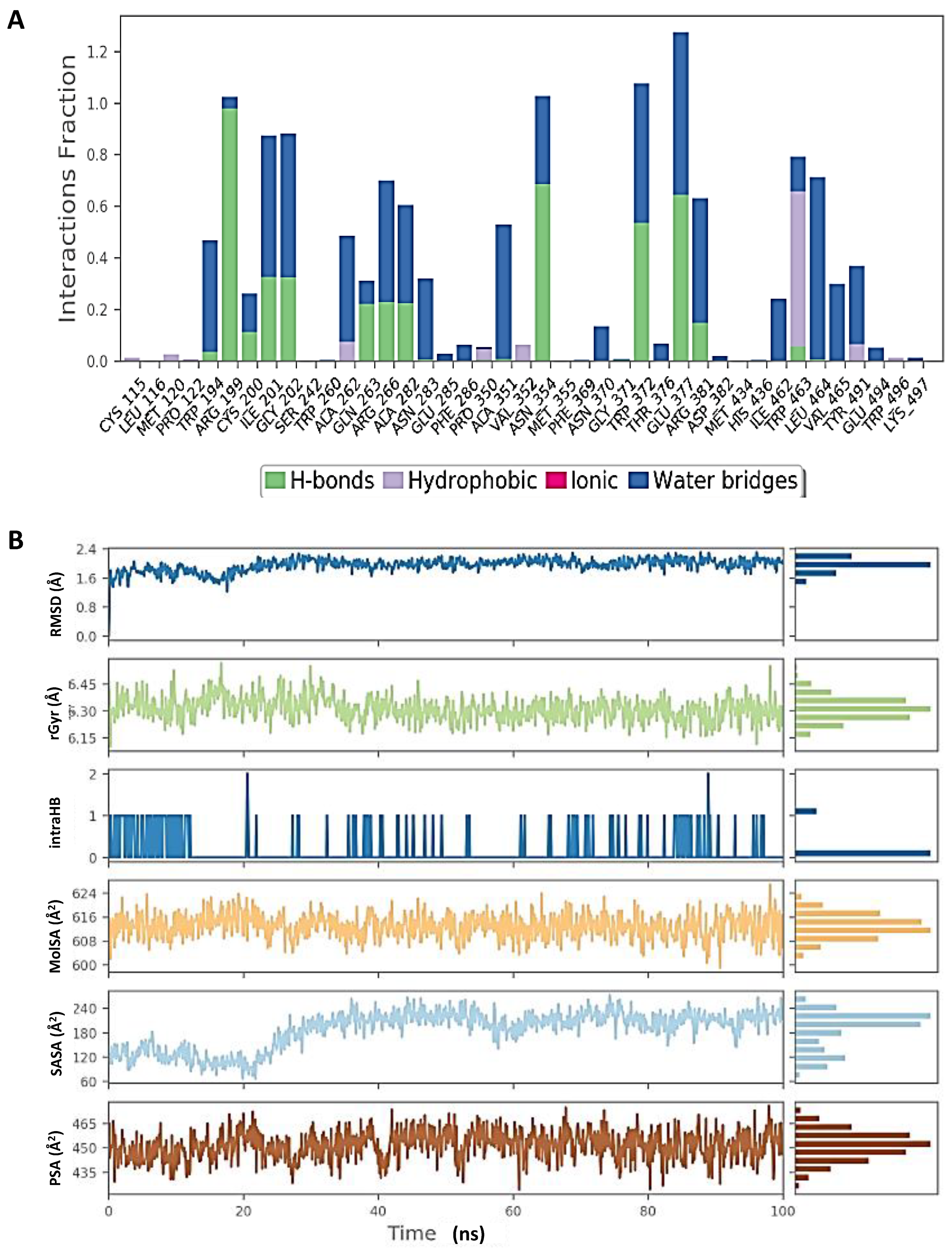
Molecular dynamic simulation (MDS) results. (A) RMSD of inducible nitric oxide synthase (iNOS) (4CX7) with glycyrrhizin. (B) Rg of iNOS. (C) RMSF of iNOS. (D) SASA of iNOS.
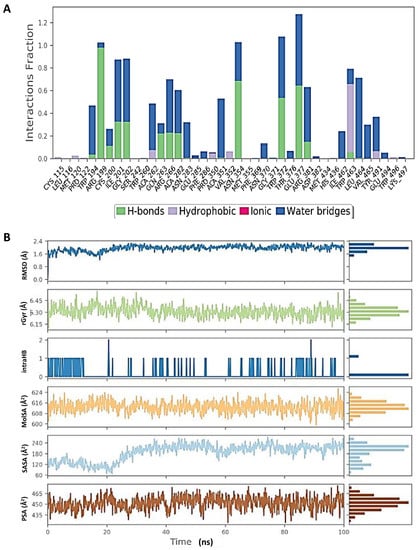
Figure 9.
Molecular dynamic simulation (MDS) results. (A) Interaction profile of the contact of inducible nitric oxide synthase (iNOS) (4CX7) with glycyrrhizin. (B) Ligand (glycyrrhizin) profile (RMSD, Rg, intramolecular hydrogen bonds (intraHB), molecular surface area (MolSA), SASA, and polar surface area (PSA)) during simulation.
Figure 8 shows that the RMSD of the inducible nitric oxide synthase (iNOS) complex with glycyrrhizin was about 1.6 Å, and the protein and ligand were both stable during 25–100 ns of the simulation time. In addition, Rg < 0.8 Å, RMSF was mostly significant at the N-terminal, 60–70, 170–190, and 310–330 amino acid residues, and total SASA was about 2000 Å2 for myeloperoxidase. Figure 9 shows the high interaction of iNOS with glycyrrhizin that occurred on the PRO122, TRP194, ARG199, ILE201, GLY202, ALA262, ARG266, ALA282, ALA351, ASN354, TRP372, ARG381, TRP463 and LEU464 amino acid residues, and the profiles of glycyrrhizin during the simulation. The binding free energies of all complexes were calculated using MMGBSA at 0 and 100 ns. The results indicate a change in the binding energy of the complex of glucoliquiritin apioside and hyaluronidase from −73.732 to −43.085 kcal.mol−1, while the binding energy of the glycyrrhizin and inducible nitric oxide synthase (iNOS) complex decreased from −91.602 to −74.874 kcal.mol−1 (Table 4 and Table 5).

Table 4.
Prime MMGBSA binding energy of the interaction of hyaluronidase and glucoliquiritin apioside before and after molecular dynamics simulation.

Table 5.
Prime MMGBSA binding energy of interaction of inducible nitric oxide synthase (iNOS) and glycyrrhizin before and after molecular dynamics simulation.
4. Discussion
Therapeutic indication was reported for Glycyrrhiza glabra regarding its antiaging, anti-inflammatory, and antioxidant properties on the basis of its inhibitory extract activity on some enzymes such as elastase and tyrosinase, which led to increased collagen and elastin synthesis [31]. Antiaging activity is due to the free-radical scavenging action and the inhibition of lipoperoxidation by the herbal extracts [43]. Licorice phytochemicals are good anti-inflammatory agents that are useful for treating skin irritations, and in cosmetics for acne and sunburns [44].
Most of the selected licorice constituents for dermatocosmetic effects exerted good ADME properties such as low gastrointestinal absorption, not being BBB permeable and substrates of P-glycoprotein, and having log Kp values that are close to those of kojic acid (−7.62 cm/s) and quercetin (−7.05 cm/s), which were among the standards used in this study; these features ensure high dermal retention and low systemic bioavailability, and thus low side effects. Skin permeability (Kp) describes the rate of chemical permeation through the outermost layer in the stratum corneum of the epidermis [28]. A high log koct value indicates high lipophilicity and is proportional to a qualitative indicator of penetration [28]. Therefore, substances with high lipophilicity persist in the lipophilic part of the skin, thus being useful for cosmetic purposes. The low GI absorption of cosmetic ingredients is generally desirable because it minimizes the potential for systemic exposure and related health risks.
According to Tuli et al. [45], targets (enzymes, biomarkers, and transcription factors) implicated by major licorice phytochemicals include cyclo-oxygenase-2 (COX-2), HMGP 1, inducible nitric oxide synthase (iNOS), interleukin-6 (IL-6), IL-10, tumor necrosis factor-α (TNF-α), TGF-β, prostaglandin E2 (PGE2), myeloperoxidase (MPO), and nuclear factor-κB (NF-κB). Bioactive licorice compounds also modulate several signaling pathways, including AMPK, PI3K/Akt, MAPK, AGE-RAGE, NLRP3, and NF-kB [46].
Moreover, the current findings indicate that glycyrrhizin could modulate HSD11B1, HSD11B2, and HSD17B, while liquiritigenin could modulate COX-1 and possessed high binding affinity for COX-2. Licorice extracts could reduce the activities of HSD enzymes, causing greater amounts of cortisol to be produced in humans and ultimately interacting with mineralocorticoid receptors [24]. In vitro and in vivo experiments showed that licorice extracts possess therapeutic properties against colon cancer by inhibiting HSD11B2 and enhancing the glucocorticoid-mediated suppression of the cyclo-oxygenase 2 (COX-2) signaling pathway [47,48]. Glucoliquiritin apioside and glycyrrhizin have high binding affinities for hyaluronidase and elastase, whereas liquiritin and liquiritin apioside showed high binding affinities for collagenase. Hyaluronidases are a family of enzymes that catalyze hyaluronic acid, and are widely distributed in the body and particularly at the periphery of collagen and elastin fibers, which is an indication of their major role in skin aging [49]. Elastase is a serine protease that preferentially digests elastin, the highly elastic protein that works together with collagen to give the skin its shape and firmness [50]. Collagenases are enzymes that cleave collagen molecules within their helical region and are more generally involved in the degradation of extracellular matrix components, thus leading to skin wrinkles [51].
Tyrosinase is a copper-containing oxidase that plays a key role in melanogenesis, controlling the production of melanin, and it is mainly involved in the hydroxylation of L-tyrosine into L-DOPA (L-3,4-dihydroxyphenylalanine, monophenolase activity) and its further oxidation into dopaquinone (diphenolase activity). This enzyme’s tyrosinase inhibitors are greatly concerning in the development of skin whitening agents [52,53]. Recently, a study indicated the synergistic effects of licorice combined with zinc in the treatment of pigmented skin disease, and that the combined preparation decreased tyrosinase, tyrosinase-related protein-1, microphthalmia-associated transcription factor, melanin formation, and cutaneous tissue injury [54]. A computational study also showed that glabridin and its semisynthetic derivatives are potential tyrosinase inhibitors that possess higher binding affinities than that of kojic acid [55].
Both glucoliquiritin apioside and glycyrrhizin had high binding affinities for lipoxygenase, whereas glycyrrhizin and liquiritigenin were predicted with high binding affinity for iNOS. Lipoxygenase, an iron-containing enzyme catalyzing the deoxygenation of polyunsaturated fatty acids into the corresponding hydroperoxides, plays a key role in inflammation [56]. A study reported that licorice extract cream with 10% concentration was more effective in lightening the skin than the concentrations of 20% and 40% were [57].
A molecular dynamics (MD) simulation was performed to determine the variation in the protein–ligand system at the atomic level, and articulate on the stability of the protein–ligand complex in a dynamic environment [58]. An RMSD of about 1.6 Å was obtained for both complexes investigated in this study, which indicates that the proteins had undergone relatively small conformational changes and were, thus, stable during the simulation. In addition, Rg < 0.9 Å, which demonstrates the compactness of the protein and the protein–ligand complex. Total SASA was in the range of 1800–2000 Å2, which indicates the surface area of proteins covered by polar and nonpolar interactions, and declines with an increment in macromolecular compactness. RMSF is useful in characterizing local changes along a protein chain. A study revealed that the tyrosinase–kojic acid complex had an RMSD of about 3.5 Å, an average Rg of nearly 0.5 Å, and SASA of approximately 2500 Å [55]. The binding potential of the ligand was quantitatively estimated using free binding energy calculation analysis with MM-GBSA [38,40,42]. The binding free energy clearly showed that the complexes were stable before and after the simulation with lesser binding energy, which could easily aid the metabolism from dermal compartments; glycyrrhizin and glucoliquiritin apioside were moderately bound to inducible nitric oxide synthase (iNOS) and hyaluronidase, respectively.
5. Conclusions
This study demonstrated that licorice (Glycyrrhiza glabra) comprises some active phytochemicals (such as glucoliquiritin apioside, glycyrrhizin, isoliquiritin, liquiritin, and liquiritin apioside) that possess high skin-permeability properties. These selected phytochemicals in licorice are potential antioxidants that enhanced dermal and epidermal histological properties, and reduced the level of inflammatory and wrinkling markers. Overall, glucoliquiritin apioside and glycyrrhizin had the best antioxidant, anti-inflammation, and dermatocosmetic activities. Although computational methods are invaluable tools in the development and safety assessment of cosmetic products, they have some limitations, which include the limited availability of data, the inadequate understanding of complex biological systems (e.g., effects on gene expression), limited predictive power (e.g., algorithms and models for prediction), variability between individuals (e.g., skin type, age, and ethnicity), and an incomplete understanding of safety endpoints (e.g., long-term effects of exposure). Therefore, in vitro and in vivo studies, and computational modeling, particularly physiological pharmacokinetics/toxicokinetics (PBPK/PBTK) in the dermal route, are required to validate these molecular pharmacological activities of licorice constituents in terms of their relevance as cosmetics.
Author Contributions
Conceptualization, T.H.F. and B.O.A.; investigation, T.H.F.; validation, T.H.F. and B.O.A.; writing—original draft preparation, T.H.F.; writing—review and editing, B.O.A. and A.O.A.; supervision, B.O.A.; project administration, B.O.A. and A.O.A.; funding acquisition, B.O.A. and A.O.A. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
All data associated with the current study are included in this article.
Acknowledgments
The institutional support from the Federal University Oye-Ekiti (Nigeria), the North-West University (South Africa), and the University of KwaZulu-Natal (South Africa) is sincerely appreciated.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Aftel, M. Essence and Alchemy: A Book of Perfume; Macmillan: London, UK, 2002. [Google Scholar]
- Drouet, S.; Garros, L.; Hano, C.; Tungmunnithum, D.; Renouard, S.; Hagège, D.; Maunit, B.; Lainé, E. A critical view of different botanical, molecular, and chemical techniques used in authentication of plant materials for cosmetic applications. Cosmetics 2018, 5, 30. [Google Scholar] [CrossRef]
- Gamage, D.G.N.D.; Dharmadasa, R.M.; Abeysinghe, D.C.; Wijesekara, R.G.S.; Prathapasinghe, G.A.; Someya, T. Global perspective of plant-based cosmetic industry and possible contribution of Sri Lanka to the development of herbal cosmetics. Evid.-Based Complement. Altern. Med. 2022, 2022, 9940548. [Google Scholar] [CrossRef]
- Lubbe, A.; Verpoorte, R. Cultivation of medicinal and aromatic plants for specialty industrial materials. Ind. Crops Prod. 2011, 34, 785–801. [Google Scholar] [CrossRef]
- Ndhlovu, P.T.; Mooki, O.; Mbeng, W.O.; Aremu, A.O. Plant species used for cosmetic and cosmeceutical purposes by the Vhavenda women in Vhembe District Municipality, Limpopo, South Africa. South Afr. J. Bot. 2019, 122, 422–431. [Google Scholar] [CrossRef]
- Pieroni, A.; Quave, C.L.; Villanelli, M.L.; Mangino, P.; Sabbatini, G.; Santini, L.; Boccetti, T.; Profili, M.; Ciccioli, T.; Rampa, L.G. Ethnopharmacognostic survey on the natural ingredients used in folk cosmetics, cosmeceuticals and remedies for healing skin diseases in the inland Marches, Central-Eastern Italy. J. Ethnopharmacol. 2004, 91, 331–344. [Google Scholar] [CrossRef]
- Fred-Jaiyesimi, A.; Ajibesin, K.K.; Tolulope, O.; Gbemisola, O. Ethnobotanical studies of folklore phytocosmetics of South West Nigeria. Pharm. Biol. 2015, 53, 313–318. [Google Scholar] [CrossRef]
- Fedoung, E.F.; Zra, T.; Biyegue, C.F.N.; Bissoue, A.N.; Baraye, S.; Tsabang, N. Herbal cosmetics knowledge of Arab-Choa and Kotoko ethnic groups in the semi-arid areas of far north Cameroon: Ethnobotanical assessment and phytochemical review. Cosmetics 2018, 5, 31. [Google Scholar] [CrossRef]
- Plainfossé, H.; Burger, P.; Azoulay, S.; Landreau, A.; Verger-Dubois, G.; Fernandez, X. Development of a natural anti-age ingredient based on Quercus pubescens Willd. leaves extract—A case study. Cosmetics 2018, 5, 15. [Google Scholar] [CrossRef]
- Mechqoq, H.; Hourfane, S.; Yaagoubi, M.E.; Hamdaoui, A.E.; Msanda, F.; Almeida, J.R.G.d.S.; Rocha, J.M.; Aouad, N.E. Phytochemical screening, and in vitro evaluation of the antioxidant and dermocosmetic activities of four Moroccan plants: Halimium antiatlanticum, Adenocarpus artemisiifolius, Pistacia lentiscus and Leonotis nepetifolia. Cosmetics 2022, 9, 94. [Google Scholar] [CrossRef]
- Goyal, A.; Sharma, A.; Kaur, J.; Kumari, S.; Garg, M.; Sindhu, R.K.; Rahman, M.H.; Akhtar, M.F.; Tagde, P.; Najda, A.; et al. Bioactive-Based Cosmeceuticals: An update on emerging trends. Molecules 2022, 27, 828. [Google Scholar] [CrossRef]
- Cerulli, A.; Masullo, M.; Montoro, P.; Piacente, S. Licorice (Glycyrrhiza glabra, G. uralensis, and G. inflata) and their constituents as active cosmeceutical ingredients. Cosmetics 2022, 9, 7. [Google Scholar] [CrossRef]
- Vanitha, M.; Soundhari, C. Isolation and characterisation of mushroom tyrosinase and screening of herbal extracts for anti-tyrosinase activity. Int. J. ChemTech Res. 2017, 10, 1156–1167. [Google Scholar]
- Wang, C.; Chen, L.; Xu, C.; Shi, J.; Chen, S.; Tan, M.; Chen, J.; Zou, L.; Chen, C.; Liu, Z.; et al. A comprehensive review for phytochemical, pharmacological, and biosynthesis studies on Glycyrrhiza spp. Am. J. Chin. Med. 2020, 48, 17–45. [Google Scholar] [CrossRef]
- Husain, I.; Bala, K.; Khan, I.A.; Khan, S.I. A review on phytochemicals, pharmacological activities, drug interactions, and associated toxicities of licorice (Glycyrrhiza sp.). Food Front. 2021, 2, 449–485. [Google Scholar] [CrossRef]
- Simmler, C.; Pauli, G.F.; Chen, S.N. Phytochemistry and biological properties of glabridin. Fitoterapia 2013, 90, 160–184. [Google Scholar] [CrossRef] [PubMed]
- Mostafa, D.M.; Ammar, N.M.; Abd El-Alim, S.H.; El-anssary, A.A. Transdermal microemulsions of Glycyrrhiza glabra L.: Characterization, stability and evaluation of antioxidant potential. Drug Deliv. 2014, 21, 130–139. [Google Scholar] [CrossRef] [PubMed]
- Castangia, I.; Caddeo, C.; Manca, M.L.; Casu, L.; Latorre, A.C.; Díez-Sales, O.; Ruiz-Saurí, A.; Bacchetta, G.; Fadda, A.M.; Manconi, M. Delivery of liquorice extract by liposomes and hyalurosomes to protect the skin against oxidative stress injuries. Carbohydr. Polym. 2015, 134, 657–663. [Google Scholar] [CrossRef]
- Rizzato, G.; Scalabrin, E.; Radaelli, M.; Capodaglio, G.; Piccolo, O. A new exploration of licorice metabolome. Food Chem. 2017, 221, 959–968. [Google Scholar] [CrossRef]
- Pastorino, G.; Cornara, L.; Soares, S.; Rodrigues, F.; Oliveira, M.B.P.P. Liquorice (Glycyrrhiza glabra): A phytochemical and pharmacological review. Phytother. Res. 2018, 32, 2323–2339. [Google Scholar] [CrossRef]
- Baurin, N.; Arnoult, E.; Scior, T.; Do, Q.T.; Bernard, P. Preliminary screening of some tropical plants for anti-tyrosinase activity. J. Ethnopharmacol. 2002, 82, 155–158. [Google Scholar] [CrossRef]
- Madden, J.C.; Enoch, S.J.; Paini, A.; Cronin, M.T.D. A review of in silico tools as alternatives to animal testing: Principles, resources and applications. Altern. Lab. Anim. 2020, 48, 146–172. [Google Scholar] [CrossRef] [PubMed]
- Cronin, M.T.D.; Enoch, S.J.; Madden, J.C.; Rathman, J.F.; Richarz, A.N.; Yang, C. A review of in silico toxicology approaches to support the safety assessment of cosmetics-related materials. Comput. Toxicol. 2022, 21, 100213. [Google Scholar] [CrossRef]
- Hasan, K.; Ara, I.; Mondal, M.S.A.; Kabir, Y. Phytochemistry, pharmacological activity, and potential health benefits of Glycyrrhiza glabra. Heliyon 2021, 7, e07240. [Google Scholar] [CrossRef] [PubMed]
- Wahab, S.; Annadurai, S.; Abullais, S.S.; Das, G.; Ahmad, W.; Ahmad, M.F.; Kandasamy, G.; Vasudevan, R.; Ali, M.S.; Amir, M. Glycyrrhiza glabra (Licorice): A comprehensive review on its phytochemistry, biological activities, clinical evidence and toxicology. Plants 2021, 10, 2751. [Google Scholar] [CrossRef] [PubMed]
- Daina, A.; Michielin, O.; Zoete, V. SwissADME: A free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci. Rep. 2017, 7, 42717. [Google Scholar] [CrossRef]
- Potts, R.O.; Guy, R.H. Predicting skin permeability. Pharm. Res. 1992, 9, 663–669. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.-P.; Chen, C.-C.; Huang, C.-W.; Chang, Y.-C. Evaluating molecular properties involved in transport of small molecules in Stratum corneum: A quantitative structure-activity relationship for skin permeability. Molecules 2018, 23, 911. [Google Scholar] [CrossRef]
- Fatoki, T.; Chukwuejim, S.; Ibraheem, O.; Oke, C.; Ejimadu, B.; Olaoye, I.; Oyegbenro, O.; Salami, T.; Basorun, R.; Oluwadare, O.; et al. Harmine and 7,8-dihydroxyflavone synergistically suitable for amyotrophic lateral sclerosis management: An insilico study. Res. Results Pharmacol. 2022, 8, 49–61. [Google Scholar] [CrossRef]
- Daina, A.; Michielin, O.; Zoete, V. SwissTargetPrediction: Updated data and new features for efficient prediction of protein targets of small molecules. Nucleic Acids Res. 2019, 47, W357–W364. [Google Scholar] [CrossRef]
- Ciganovic, P.K.; Tomczyk, J.M.; Koncic, M.Z. Glycerolic licorice extracts as active cosmeceutical ingredients: Extraction optimization, chemical characterization, and biological activity. Antioxidants 2019, 8, 445. [Google Scholar] [CrossRef]
- Morris, G.M.; Huey, R.; Lindstrom, W.; Sanner, M.F.; Belew, R.K.; Goodsell, D.S.; Olson, A.J. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J. Comput. Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef] [PubMed]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef] [PubMed]
- Eberhardt, J.; Santos-Martins, D.; Tillack, A.F.; Forli, S. AutoDock Vina 1.2.0: New docking methods, expanded force field, and python bindings. J. Chem. Inf. Model. 2021, 61, 3891–3898. [Google Scholar] [CrossRef] [PubMed]
- Tao, A.; Huang, Y.; Shinohara, Y.; Caylor, M.L.; Pashikanti, S.; Xu, D. ezCADD: A rapid 2D/3D visualization-enabled web modeling environment for democratizing computer-aided drug design. J. Chem. Inf. Model. 2019, 59, 18–24. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Gable, A.L.; Nastou, K.C.; Lyon, D.; Kirsch, R.; Pyysalo, S.; Doncheva, N.T.; Legeay, M.; Fang, T.; Bork, P.; et al. The STRING database in 2021: Customizable protein–protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 2021, 49, D605–D612. [Google Scholar] [CrossRef]
- Bowers, K.J.; Chow, D.E.; Xu, H.; Dror, R.O.; Eastwood, M.P.; Gregersen, B.A.; Klepeis, J.L.; Kolossvary, I.; Moraes, M.A.; Sacerdoti, F.D.; et al. Molecular dynamics—Scalable algorithms for molecular dynamics simulations on commodity clusters. In Proceedings of the 2006 ACM/IEEE Conference on Supercomputing—SC’06, Tampa, FL, USA, 11–17 November 2006. [Google Scholar]
- Schrödinger Release 2023-1. Desmond molecular dynamics system, D.E. Shaw research, New York, NY, 2021. Maestro-Desmond interoperability tools, Schrodinger LLC, New York, NY, 2021. Available online: https://www.schrodinger.com/products/desmond (accessed on 25 February 2023).
- Jawarkar, R.D.; Sharma, P.; Jain, N.; Gandhi, A.; Mukerjee, N.; Al-Mutairi, A.A.; Zaki, M.E.A.; Al-Hussain, S.A.; Samad, A.; Masand, V.H.; et al. QSAR, molecular docking, MD simulation and MMGBSA calculations approaches to recognize concealed pharmacophoric features requisite for the optimization of ALK Tyrosine Kinase Inhibitors as Anticancer Leads. Molecules 2022, 27, 4951. [Google Scholar] [CrossRef]
- Shivakumar, D.; Williams, J.; Wu, Y.; Damm, W.; Shelley, J.; Sherman, W. Prediction of absolute solvation free energies using molecular dynamics free energy perturbation and the OPLS force field. J. Chem. Theory Comput. 2010, 6, 1509–1519. [Google Scholar] [CrossRef]
- Schrödinger Resources: What Do All the Prime MM-GBSA Energy Properties Mean? Available online: www.schrodinger.com/kb/1875 (accessed on 6 March 2023).
- Zhang, X.; Perez-Sanchez, H.; Lightstone, F.C. A Comprehensive Docking and MM/GBSA Rescoring Study of Ligand Recognition upon Binding Antithrombin. Curr. Top. Med. Chem. 2017, 17, 1631–1639. [Google Scholar] [CrossRef]
- Marks, A. Herbal extracts in cosmetics. Agro-Food-Ind. Hi-Tech. 1997, 8, 28–31. [Google Scholar]
- Aburjai, T.; Natsheh, F.M. Plants Used in Cosmetics. Phytother. Res. 2003, 17, 987–1000. [Google Scholar] [CrossRef]
- Tuli, H.S.; Garg, V.K.; Mehta, J.K.; Kaur, G.; Mohapatra, R.K.; Dhama, K.; Sak, K.; Kumar, A.; Varol, M.; Aggarwal, D.; et al. Licorice (Glycyrrhiza glabra L.)-derived phytochemicals target multiple signaling pathways to confer oncopreventive and oncotherapeutic effects. OncoTarg. Ther. 2022, 15, 1419–1448. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Jiang, Y.; Zhang, Z.; Hou, J.; Tian, S.; Liu, Y. The anti-diabetic activity of licorice, a widely used Chinese herb. J. Ethnopharmacol. 2020, 263, 113216. [Google Scholar] [CrossRef] [PubMed]
- Stewart, P.M.; Prescott, S.M. Can licorice lick colon cancer? J. Clin. Investig. 2009, 119, 760–763. [Google Scholar] [CrossRef]
- Zhang, M.Z.; Xu, J.; Yao, B.; Yin, H.; Cai, Q.; Shrubsole, M.J.; Chen, X.; Kon, V.; Zheng, W.; Pozzi, A.; et al. Inhibition of 11-β-hydroxysteroid dehydrogenase type II selectively blocks the tumor COX-2 pathway and suppresses colon carcinogenesis in mice and humans. J. Clin. Investig. 2009, 119, 876–885. [Google Scholar] [CrossRef]
- Papakonstantinou, E.; Roth, M.; Karakiulakis, G. Hyaluronic acid: A key molecule in skin aging. Derm.-Endocrinol. 2012, 4, 253–258. [Google Scholar] [CrossRef]
- Debelle, L.; Tamburro, A.M. Elastin: Molecular description and function. Int. J. Biochem. Cell Biol. 1999, 31, 261–272. [Google Scholar] [CrossRef]
- Roy, A.; Sahu, R.K.; Matlam, M.; Deshmukh, V.K.; Dwivedi, J.; Jha, A.K. In vitro techniques to assess the proficiency of skin care cosmetic formulations. Pharmacogn. Rev. 2013, 7, 97–106. [Google Scholar] [PubMed]
- Burger, P.; Landreau, A.; Azoulay, S.; Michel, T.; Fernandez, X. Skin whitening cosmetics: Feedback and challenges in the development of natural skin lighteners. Cosmetics 2016, 3, 36. [Google Scholar] [CrossRef]
- Fatoki, T.H.; Ibraheem, O.; Adeseko, C.J.; Afolabi, B.L.; Momodu, D.U.; Sanni, D.M.; Enibukun, J.M.; Ogunyemi, I.O.; Adeoye, A.O.; Ugboko, H.U.; et al. Melanogenesis, its regulatory process, and insights on biomedical, biotechnological, and pharmacological potentials of melanin as antiviral biochemical. Bioint Res. Appl. Chem. 2021, 11, 11969–11984. [Google Scholar] [CrossRef]
- Wang, J.-Y.; Xie, X.-Y.; Deng, Y.; Yang, H.-Q.; Du, X.-S.; Liu, P.; Du, Y. Licorice zinc suppresses melanogenesis via inhibiting the activation of P38/MAPK and JNK signaling pathway in C57BL/6J mice skin. Acta Cirúrgica Bras. 2022, 37, e371002. [Google Scholar] [CrossRef]
- Kumari, A.; Kumar, R.; Sulabh, G.; Singh, P.; Kumar, J.; Singh, V.K.; Ojha, K.K. In silico ADMET, molecular docking and molecular simulation-based study of glabridin’s natural and semisynthetic derivatives as potential tyrosinase inhibitors. Adv. Tradit. Med. 2022, 23, 1–19. [Google Scholar] [CrossRef]
- Lu, W.; Zhao, X.; Xu, Z.; Dong, N.; Zou, S.; Shen, X.; Huang, J. Development of a new colorimetric assay for lipoxygenase activity. Anal. Biochem. 2013, 441, 162–168. [Google Scholar] [CrossRef]
- Rakhmini, A.; Ilyas, F.S.; Muchtar, S.V.; Patellongi, I.J.; Djawad, K.; Alam, G. Comparison of 10%, 20% and 40% licorice extract cream as skin lightening agent. Int. J. Med. Rev. Case Rep. 2018, 2, 131–135. [Google Scholar]
- Saini, G.; Dalal, V.; Gupta, D.N.; Sharma, N.; Kumar, P.; Sharma, A.K. A molecular docking and dynamic approach to screen inhibitors against ZnuA1 of Candidatus Liberibacter asiaticus. Mol. Simul. 2021, 47, 510–525. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).