Droplet Digital PCR Analysis of Liquid Biopsy Samples Unveils the Diagnostic Role of hsa-miR-133a-3p and hsa-miR-375-3p in Oral Cancer
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Samples Included in the Study
2.2. RNA Extraction and miRNAs Retrotranscription
2.3. Analysis of miRNAs Expression Through Droplet Digital PCR
2.4. Bioinformatics Analyses
2.5. Statistical Analyses
3. Results
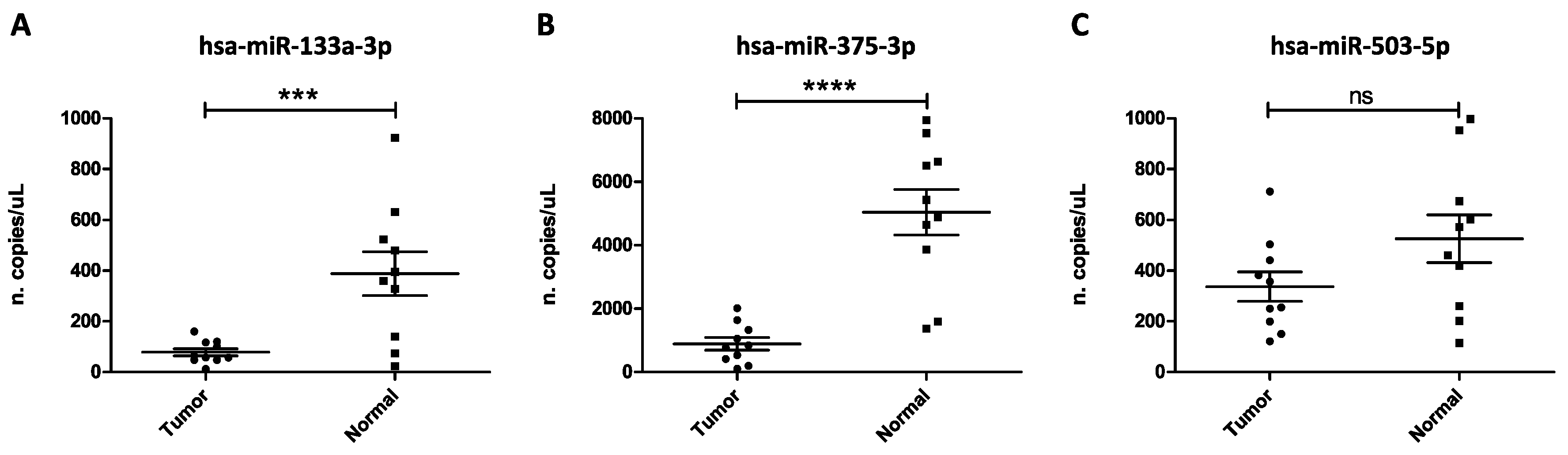
3.1. Analysis of miRNA Expression Levels in Liquid Biopsy Samples of Oral Cancer Patients and Healthy Controls
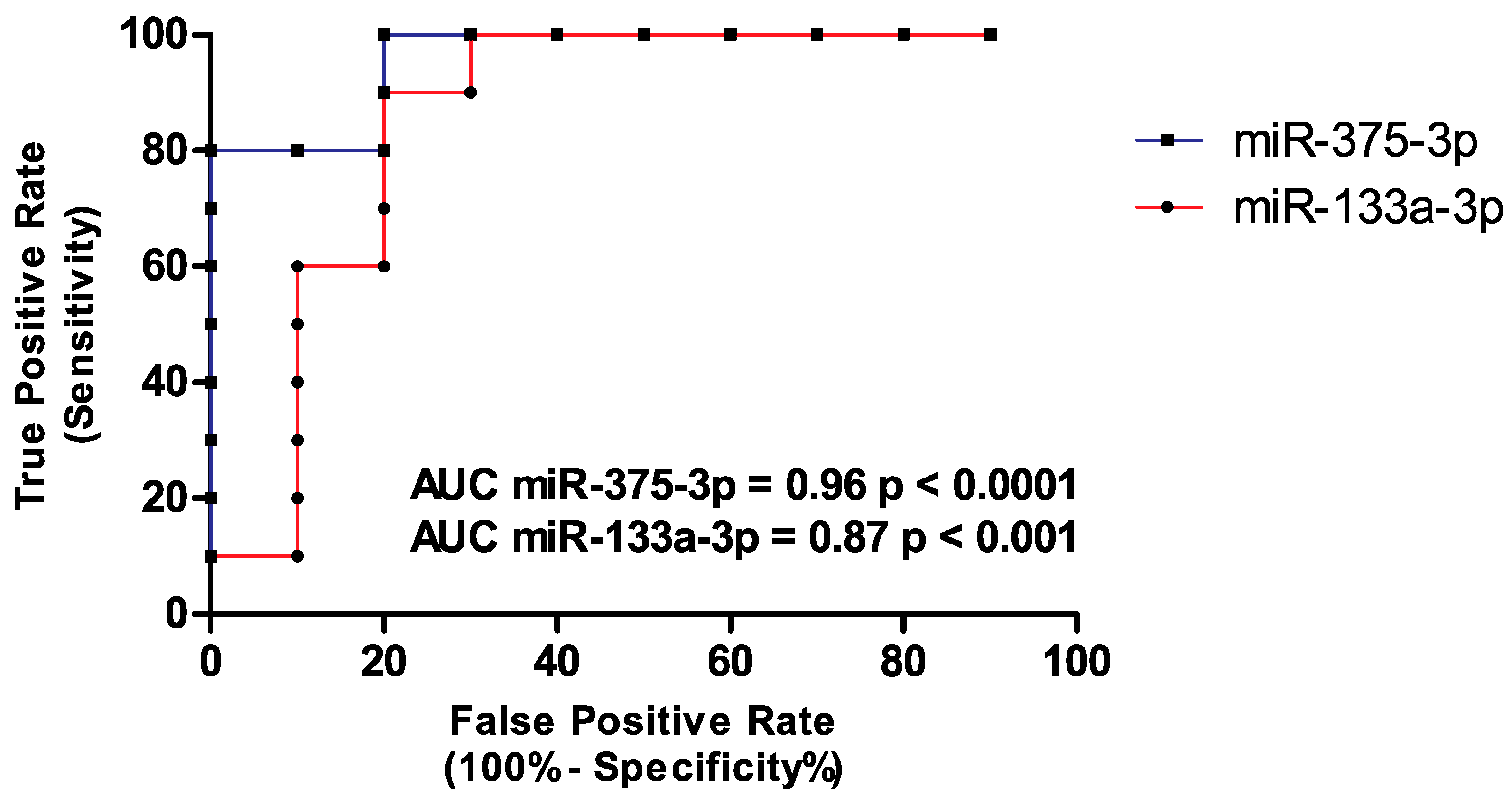
3.2. Diagnostic Value of hsa-miR-133a-3p and hsa-miR-375-3p
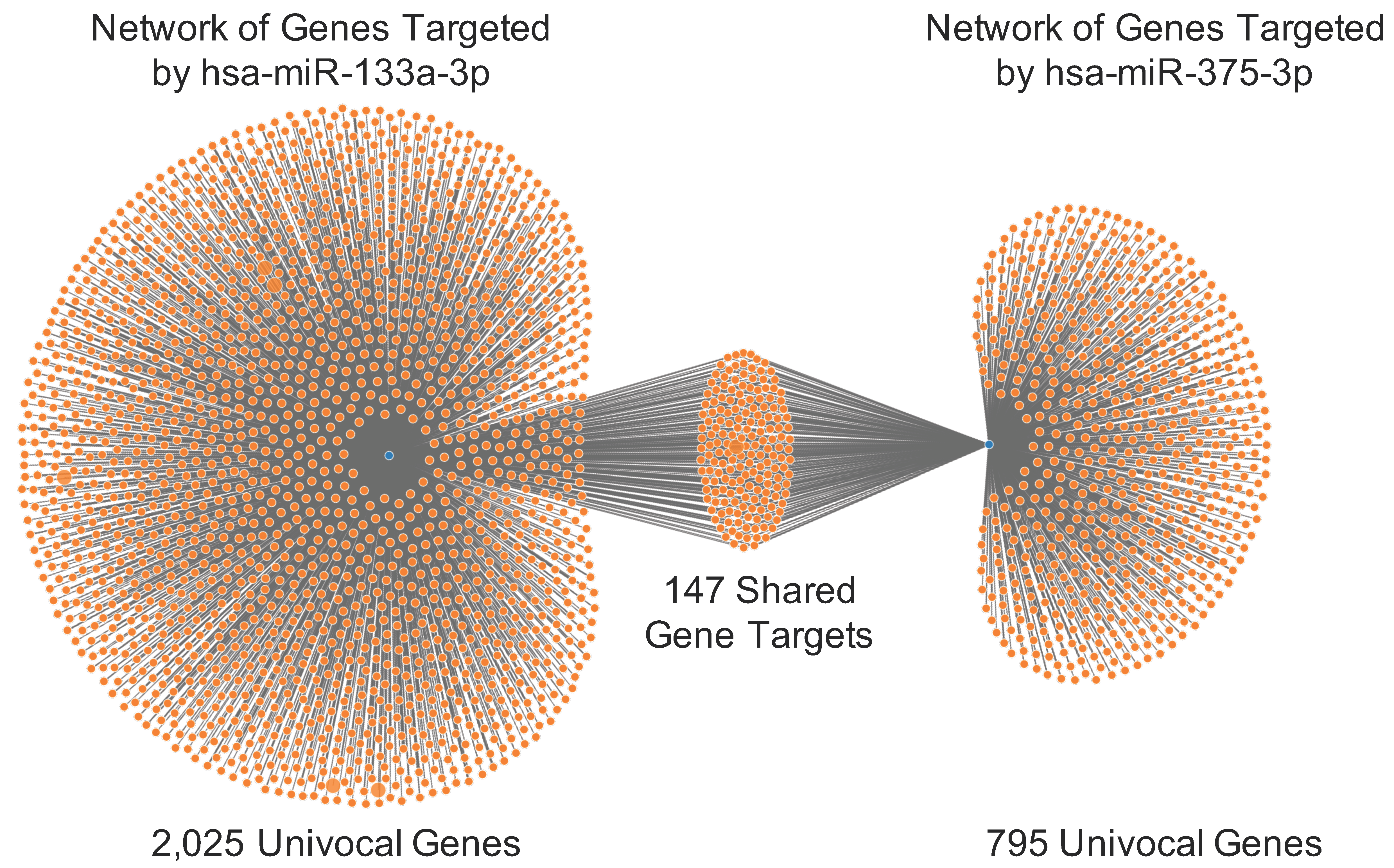
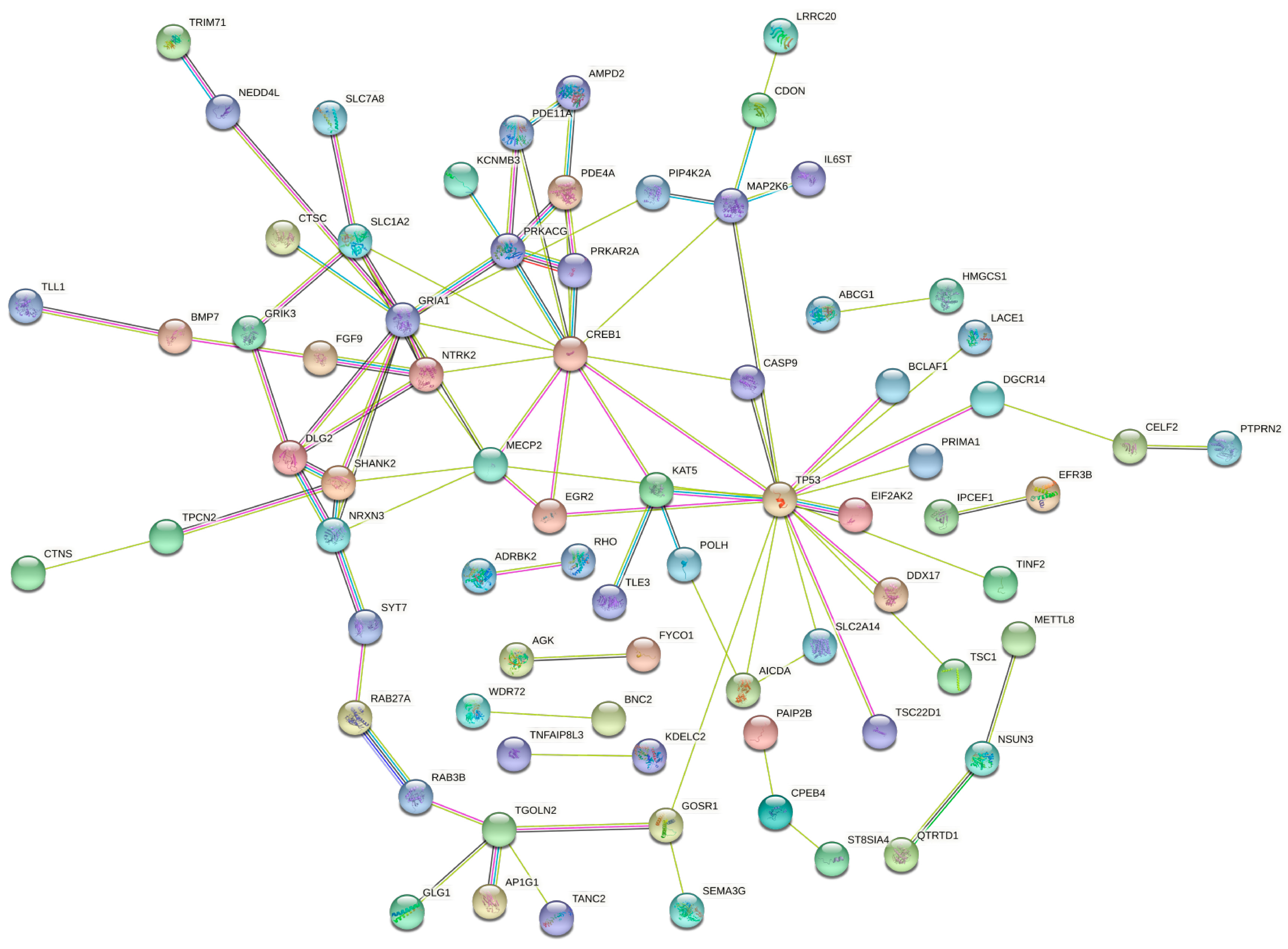
3.3. Functional Role of hsa-miR-133a-3p and hsa-miR-375-3p in Oral Cancer
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ferlay, J.; Colombet, M.; Soerjomataram, I.; Mathers, C.; Parkin, D.M.; Piñeros, M.; Znaor, A.; Bray, F. Estimating the global cancer incidence and mortality in 2018: GLOBOCAN sources and methods. Int. J. Cancer 2018, 144, 1941–1953. [Google Scholar] [CrossRef] [PubMed]
- Sarode, S.; Sarode, S.C.; Maniyar, N.; Sharma, N.K.; Yerwadekar, S.; Patil, S. Recent trends in predictive biomarkers for determining malignant potential of oral potentially malignant disorders. Oncol. Rev. 2019, 13, 424. [Google Scholar] [CrossRef] [PubMed]
- Grafton-Clarke, C.; Chen, K.W.; Wilcock, J. Diagnosis and referral delays in primary care for oral squamous cell cancer: A systematic review. Br. J. Gen. Pr. 2018, 69, e112–e126. [Google Scholar] [CrossRef] [PubMed]
- Akbulut, N.; Oztas, B.; Kursun, S.; Evirgen, S. Delayed diagnosis of oral squamous cell carcinoma: A case series. J. Med. Case Rep. 2011, 5, 291. [Google Scholar] [CrossRef]
- La Rosa, G.R.M.; Gattuso, G.; Pedullà, E.; Rapisarda, E.; Nicolosi, D.; Salmeri, M. Association of oral dysbiosis with oral cancer development. Oncol. Lett. 2020, 19, 3045–3058. [Google Scholar] [CrossRef] [PubMed]
- Kumar, M.; Nanavati, R.; Modi, T.G.; Dobariya, C. Oral cancer: Etiology and risk factors: A review. J. Cancer Res. Ther. 2016, 12, 458–463. [Google Scholar] [CrossRef] [PubMed]
- Beynon, R.A.; Lang, S.; Schimansky, S.; Penfold, C.M.; Waylen, A.; Thomas, S.J.; Pawlita, M.; Waterboer, T.; Martin, R.M.; May, M.; et al. Tobacco smoking and alcohol drinking at diagnosis of head and neck cancer and all-cause mortality: Results from head and neck 5000, a prospective observational cohort of people with head and neck cancer. Int. J. Cancer 2018, 143, 1114–1127. [Google Scholar] [CrossRef]
- Wang, H.; Wang, X.; Xu, J.; Wang, L.; Liu, C. The role of cigarette smoking and alcohol consumption in the differentiation of oral squamous cell carcinoma for the males in China. J. Cancer Res. Ther. 2015, 11, 141. [Google Scholar] [CrossRef]
- She, Y.; Nong, X.; Zhang, M.; Wang, M. Epstein-Barr virus infection and oral squamous cell carcinoma risk: A meta-analysis. PLoS ONE 2017, 12, e0186860. [Google Scholar] [CrossRef] [PubMed]
- Bakri, M.M.; Hussaini, H.M.; Holmes, A.R.; Cannon, R.D.; Rich, A.M. Revisiting the association between candidal infection and carcinoma, particularly oral squamous cell carcinoma. J. Oral Microbiol. 2010, 2. [Google Scholar] [CrossRef]
- Kreimer, A.R.; Clifford, G.; Boyle, P.; Franceschi, S. Human Papillomavirus Types in Head and Neck Squamous Cell Carcinomas Worldwide: A Systematic Review. Cancer Epidemiol. Biomark. Prev. 2005, 14, 467–475. [Google Scholar] [CrossRef]
- Karpiński, T.M. Role of Oral Microbiota in Cancer Development. Microorganisms 2019, 7, 20. [Google Scholar] [CrossRef] [PubMed]
- Bhatt, A.P.; Redinbo, M.R.; Bultman, S.J. The role of the microbiome in cancer development and therapy. CA A Cancer J. Clin. 2017, 67, 326–344. [Google Scholar] [CrossRef]
- Polo, A.; Crispo, A.; Cerino, P.; Falzone, L.; Candido, S.; Giudice, A.; De Petro, G.; Ciliberto, G.; Montella, M.; Budillon, A.; et al. Environment and bladder cancer: Molecular analysis by interaction networks. Oncotarget 2017, 8, 65240–65252. [Google Scholar] [CrossRef]
- Herceg, Z.; Hainaut, P. Genetic and epigenetic alterations as biomarkers for cancer detection, diagnosis and prognosis. Mol. Oncol. 2007, 1, 26–41. [Google Scholar] [CrossRef]
- D’Souza, W.; Saranath, D. OMICS, Oral Cancer Molecular Landscapes, and Clinical Practice. OMICS: A J. Integr. Biol. 2017, 21, 689–703. [Google Scholar] [CrossRef]
- Bavle, R.M.; Venugopal, R.; Konda, P.; Muniswamappa, S.; Makarla, S. Molecular Classification of Oral Squamous Cell Carcinoma. J. Clin. Diagn. Res. 2016, 10, ZE18–ZE21. [Google Scholar] [CrossRef] [PubMed]
- Falzone, L.; Grimaldi, M.; Celentano, E.; Augustin, L.S.A.; Libra, M. Identification of Modulated MicroRNAs Associated with Breast Cancer, Diet, and Physical Activity. Cancers 2020, 12, 2555. [Google Scholar] [CrossRef]
- Irimie, A.I.; Ciocan, C.; Gulei, D.; Mehterov, N.; Atanasov, A.G.; Dudea, D.; Berindan-Neagoe, I. Current Insights into Oral Cancer Epigenetics. Int. J. Mol. Sci. 2018, 19, 670. [Google Scholar] [CrossRef] [PubMed]
- Hema, K.N.; Smitha, T.; Sheethal, H.S.; Mirnalini, S.A. Epigenetics in oral squamous cell carcinoma. J. Oral Maxillofac. Pathol. 2017, 21, 252–259. [Google Scholar] [CrossRef]
- Falzone, L.; Salemi, R.; Travali, S.; Scalisi, A.; McCubrey, J.A.; Candido, S.; Libra, M. MMP-9 overexpression is associated with intragenic hypermethylation of MMP9 gene in melanoma. Aging 2016, 8, 933–944. [Google Scholar] [CrossRef]
- Bertoli, G.; Cava, C.; Castiglioni, I. MicroRNAs: New Biomarkers for Diagnosis, Prognosis, Therapy Prediction and Therapeutic Tools for Breast Cancer. Theranostics 2015, 5, 1122–1143. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Falzone, L.; Lupo, G.; La Rosa, G.R.M.; Crimi, S.; Anfuso, C.D.; Salemi, R.; Rapisarda, E.; Libra, M.; Candido, S. Identification of Novel MicroRNAs and Their Diagnostic and Prognostic Significance in Oral Cancer. Cancers 2019, 11, 610. [Google Scholar] [CrossRef]
- Filetti, V.; Falzone, L.; Rapisarda, V.; Caltabiano, R.; Graziano, A.C.E.; Ledda, C.; Loreto, C. Modulation of microRNA expression levels after naturally occurring asbestiform fibers exposure as a diagnostic biomarker of mesothelial neoplastic transformation. Ecotoxicol. Environ. Saf. 2020, 198, 110640. [Google Scholar] [CrossRef] [PubMed]
- Battaglia, R.; Palini, S.; Vento, M.E.; La Ferlita, A.; Leonardi, A.A.; Caroppo, L.E.; Borzì, P.; Falzone, L.; Barbagallo, D.; Ragusa, M.; et al. Identification of extracellular vesicles and characterization of miRNA expression profiles in human blastocoel fluid. Sci. Rep. 2019, 9, 84. [Google Scholar] [CrossRef]
- Sticht, C.; De La Torre, C.; Parveen, A.; Gretz, N. miRWalk: An online resource for prediction of microRNA binding sites. PLoS ONE 2018, 13, e0206239. [Google Scholar] [CrossRef]
- Thomas, P.D.; Muruganujan, A.; Ebert, D.; Huang, X.; Thomas, P.D. PANTHER version 14: More genomes, a new PANTHER GO-slim and improvements in enrichment analysis tools. Nucleic Acids Res. 2018, 47, D419–D426. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2018, 47, D607–D613. [Google Scholar] [CrossRef]
- Ahmed, M.; Nguyên, H.; Lai, T.H.; Kim, D.R. miRCancerdb: A database for correlation analysis between microRNA and gene expression in cancer. BMC Res. Notes 2018, 11, 1–7. [Google Scholar] [CrossRef]
- Tang, Z.; Li, C.; Kang, B.; Gao, G.; Li, C.; Zhang, Z. GEPIA: A web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017, 45, W98–W102. [Google Scholar] [CrossRef]
- Goldman, M.; Craft, B.; Hastie, M.; Repečka, K.; McDade, F.; Kamath, A.; Banerjee, A.; Luo, Y.; Rogers, D.; Brooks, A.N.; et al. Visualizing and interpreting cancer genomics data via the Xena platform. Nat. Biotechnol. 2020, 38, 675–678. [Google Scholar] [CrossRef]
- Saxena, S.; Sankhla, B.; Sundaragiri, K.S.; Bhargava, A. A Review of Salivary Biomarker: A Tool for Early Oral Cancer Diagnosis. Adv. Biomed. Res. 2017, 6, 90. [Google Scholar] [CrossRef]
- Yang, K.; Yuan, C.; Tang, H.; Chen, D. Diagnostic values of serum tumor markers Cyfra21-1, SCCAg, ferritin, CEA, CA19-9, and AFP in oral/oropharyngeal squamous cell carcinoma. OncoTargets Ther. 2016, 9, 3381–3386. [Google Scholar] [CrossRef]
- John, M.A.R.S.; Li, Y.; Zhou, X.; Denny, P.; Ho, C.-M.; Montemagno, C.; Shi, W.; Qi, F.; Wu, B.; Sinha, U.; et al. Interleukin 6 and Interleukin 8 as Potential Biomarkers for Oral Cavity and Oropharyngeal Squamous Cell Carcinoma. Arch. Otolaryngol. Head Neck Surg. 2004, 130, 929–935. [Google Scholar] [CrossRef]
- Amiri-Dashatan, N.; Koushki, M.; Jalilian, A.; Ahmadi, N.A.; Tavirani, M.R. Integrated Bioinformatics Analysis of mRNAs and miRNAs Identified Potential Biomarkers of Oral Squamous Cell Carcinoma. Asian Pac. J. Cancer Prev. 2020, 21, 1841–1848. [Google Scholar] [CrossRef]
- Tuaeva, N.O.; Falzone, L.; Porozov, Y.B.; Nosyrev, A.E.; Trukhan, V.M.; Kovatsi, L.; Spandidos, D.A.; Drakoulis, N.; Kalogeraki, A.; Mamoulakis, C.; et al. Translational Application of Circulating DNA in Oncology: Review of the Last Decades Achievements. Cells 2019, 8, 1251. [Google Scholar] [CrossRef]
- Zeljic, K.; Jovanovic, I.; Jovanovic, J.; Magic, Z.; Stankovic, A.; Supic, G. MicroRNA meta-signature of oral cancer: Evidence from a meta-analysis. Upsala J. Med Sci. 2018, 123, 43–49. [Google Scholar] [CrossRef] [PubMed]
- Zhou, C.; Ye, M.; Ni, S.; Li, Q.; Ye, D.; Li, J.; Shen, Z.; Deng, H. DNA methylation biomarkers for head and neck squamous cell carcinoma. Epigenetics 2018, 13, 398–409. [Google Scholar] [CrossRef]
- Zhang, H.; Li, T.; Zheng, L.; Huang, X. Biomarker MicroRNAs for Diagnosis of Oral Squamous Cell Carcinoma Identified Based on Gene Expression Data and MicroRNA-mRNA Network Analysis. Comput. Math. Methods. Med. 2017, 2017, 9803018. [Google Scholar] [CrossRef] [PubMed]
- Huang, G.-Z.; Wu, Q.-Q.; Zheng, Z.-N.; Shao, T.-R.; Lv, X.-Z. Identification of Candidate Biomarkers and Analysis of Prognostic Values in Oral Squamous Cell Carcinoma. Front. Oncol. 2019, 9, 1054. [Google Scholar] [CrossRef]
- Falzone, L.; Gattuso, G.; Lombardo, C.; Lupo, G.; Grillo, C.M.; Spandidos, D.A.; Libra, M.; Salmeri, M. Droplet digital PCR for the detection and monitoring of Legionella pneumophila. Int. J. Mol. Med. 2020, 46, 1777–1782. [Google Scholar] [CrossRef]
- Falzone, L.; Musso, N.; Gattuso, G.; Bongiorno, D.; Palermo, C.I.; Scalia, G.; Libra, M.; Stefani, S. Sensitivity assessment of droplet digital PCR for SARS-CoV-2 detection. Int. J. Mol. Med. 2020, 46, 957–964. [Google Scholar] [CrossRef] [PubMed]
- Salemi, R.; Falzone, L.; Madonna, G.; Polesel, J.; Cinà, D.; Mallardo, D.; Ascierto, P.A.; Libra, M.; Candido, S. MMP-9 as a Candidate Marker of Response to BRAF Inhibitors in Melanoma Patients With BRAFV600E Mutation Detected in Circulating-Free DNA. Front. Pharmacol. 2018, 9, 856. [Google Scholar] [CrossRef]
- Costa, E.F.D.; Lopes-Aguiar, L.; Nogueira, G.S.; Lima, T.R.P.; Rinck-Junior, J.A.; Lourenço, G.J.; Lima, C.S.P. CASP9 c.-1339A > G and CASP3 c.-1191A > G polymorphisms alter susceptibility and clinical aspects of head and neck squamous cell carcinoma. Head Neck 2019, 41, 2665–2670. [Google Scholar] [CrossRef]
- Freier, K.; Sticht, C.; Hofele, C.; Flechtenmacher, C.; Stange, D.E.; Puccio, L.; Toedt, G.; Radlwimmer, B.; Lichter, P.; Joos, S. Recurrent coamplification of cytoskeleton-associated genesEMS1 andSHANK2 withCCND1 in oral squamous cell carcinoma. Genes Chromosom. Cancer 2005, 45, 118–125. [Google Scholar] [CrossRef] [PubMed]
- Zhou, G.; Liu, Z.; Myers, J.N. TP53Mutations in Head and Neck Squamous Cell Carcinoma and Their Impact on Disease Progression and Treatment Response. J. Cell. Biochem. 2016, 117, 2682–2692. [Google Scholar] [CrossRef] [PubMed]
- He, B.; Lin, X.; Tian, F.; Yu, W.; Qiao, B. MiR-133a-3p Inhibits Oral Squamous Cell Carcinoma (OSCC) Proliferation and Invasion by Suppressing COL1A1. J. Cell. Biochem. 2018, 119, 338–346. [Google Scholar] [CrossRef]
- Brown, A.L.; Al-Samadi, A.; Sperandio, M.; Soares, A.B.; Teixeira, L.N.; Martinez, E.F.; Demasi, A.P.D.; Araújo, V.C.; Leivo, I.; Salo, T.; et al. MiR-455-3p, miR-150 and miR-375 are aberrantly expressed in salivary gland adenoid cystic carcinoma and polymorphous adenocarcinoma. J. Oral Pathol. Med. 2019, 48, 840–845. [Google Scholar] [CrossRef]
- Shi, W.; Yang, J.; Li, S.; Shan, X.; Liu, X.; Hua, H.; Zhao, C.; Feng, Z.; Cai, Z.; Zhang, L.; et al. Potential involvement of miR-375 in the premalignant progression of oral squamous cell carcinoma mediated via transcription factor KLF5. Oncotarget 2015, 6, 40172–40185. [Google Scholar] [CrossRef]
- Shiiba, M.; Uzawa, K.; Tanzawa, H. MicroRNAs in Head and Neck Squamous Cell Carcinoma (HNSCC) and Oral Squamous Cell Carcinoma (OSCC). Cancers 2010, 2, 653–669. [Google Scholar] [CrossRef]
- Gissi, D.; Tarsitano, A.; Gabusi, A.; Rossi, R.; Attardo, G.; Lenzi, J.; Marchetti, C.; Montebugnoli, L.; Foschini, M.P.; Morandi, L. 13-gene DNA Methylation Analysis from Oral Brushing: A Promising Non Invasive Tool in the Follow-up of Oral Cancer Patients. J. Clin. Med. 2019, 8, 2107. [Google Scholar] [CrossRef] [PubMed]
- Salta, S.; Nunes, S.P.; Fontes-Sousa, M.; Lopes, P.; Freitas, M.; Caldas, M.; Antunes, L.; Castro, F.; Antunes, P.; De Sousa, S.P.; et al. A DNA Methylation-Based Test for Breast Cancer Detection in Circulating Cell-Free DNA. J. Clin. Med. 2018, 7, 420. [Google Scholar] [CrossRef]
- Hamam, R.; Hamam, D.; Alsaleh, K.A.; Kassem, M.; Zaher, W.; Alfayez, M.; Aldahmash, A.; Alajez, N.M. Circulating microRNAs in breast cancer: Novel diagnostic and prognostic biomarkers. Cell Death Dis. 2017, 8, e3045. [Google Scholar] [CrossRef]
- Hafsi, S.; Candido, S.; Maestro, R.; Falzone, L.; Soua, Z.; Bonavida, B.; Spandidos, D.A.; Libra, M. Correlation between the overexpression of Yin Yang 1 and the expression levels of miRNAs in Burkitt’s lymphoma: A computational study. Oncol. Lett. 2016, 11, 1021–1025. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Li, Z.; Xuan, Z.; Xu, P.; Wang, W.; Chen, Z.; Wang, S.; Sun, G.; Xu, J.; Xu, Z. Novel role of miR-133a-3p in repressing gastric cancer growth and metastasis via blocking autophagy-mediated glutaminolysis. J. Exp. Clin. Cancer Res. 2018, 37, 1–22. [Google Scholar] [CrossRef] [PubMed]
- Tang, Y.; Pan, J.; Huang, S.; Peng, X.; Zou, X.; Luo, Y.; Ren, D.; Zhang, X.; Li, R.; He, P.; et al. Downregulation of miR-133a-3p promotes prostate cancer bone metastasis via activating PI3K/AKT signaling. J. Exp. Clin. Cancer Res. 2018, 37, 1–16. [Google Scholar] [CrossRef]
- Liang, H.; Yang, X.; Wen, D.; Gao, L.; Zhang, X.; Ye, Z.; Luo, J.; Li, Z.; He, Y.; Pang, Y.; et al. Utility of miR-133a-3p as a diagnostic indicator for hepatocellular carcinoma: An investigation combined with GEO, TCGA, meta-analysis and bioinformatics. Mol. Med. Rep. 2017, 17, 1469–1484. [Google Scholar] [CrossRef]
- Xu, X.; Chen, X.; Xu, M.; Liu, X.; Pan, B.; Qin, J.; Xu, T.; Zeng, K.; Pan, Y.; He, B.; et al. miR-375-3p suppresses tumorigenesis and partially reverses chemoresistance by targeting YAP1 and SP1 in colorectal cancer cells. Aging 2019, 11, 7357–7385. [Google Scholar] [CrossRef]
- He, S.; Shi, J.; Mao, J.; Luo, X.; Liu, W.; Liu, R.; Yang, F. The expression of miR-375 in prostate cancer: A study based on GEO, TCGA data and bioinformatics analysis. Pathol. Res. Pr. 2019, 215, 152375. [Google Scholar] [CrossRef]

| Patients’ Characteristics | Cancer Patients (N. 10) | Normal Controls (N. 10) | ||
|---|---|---|---|---|
| N. | % | N. | % | |
| Sex | ||||
| Male | 6 | 60 | 9 | 90 |
| Female | 4 | 40 | 1 | 10 |
| Age | ||||
| <45 | 0 | 0 | 0 | 0 |
| 45–59 | 3 | 30 | 6 | 60 |
| >60 | 7 | 70 | 4 | 40 |
| Smoke | ||||
| Yes | 4 | 40 | 3 | 30 |
| No | 4 | 40 | 5 | 50 |
| Ex-smoker | 2 | 20 | 2 | 20 |
| Alchol | ||||
| Yes | 3 | 30 | 1 | 10 |
| No | 7 | 70 | 9 | 90 |
| Family History | ||||
| Yes | 6 | 70 | 3 | 30 |
| No | 4 | 30 | 7 | 70 |
| Tumor Stage | ||||
| T1 | 1 | 10 | NA | |
| T2 | 2 | 20 | ||
| T3 | 2 | 20 | ||
| T4 | 5 | 50 | ||
| Lymph Nodes | ||||
| Positive | 2 | 20 | NA | |
| Negative | 8 | 80 | ||
| Recurrence | ||||
| Yes | 3 | 30 | NA | |
| No | 7 | 70 | ||
| Gene | GEPIA Analysis | miRCancerdb Correlation | ||
|---|---|---|---|---|
| Log2FC | Adj. p-Value | hsa-miR-133a-3p | hsa-miR-375-3p | |
| DLG2 | −1.259 | 5.55 × 10−51 | 0.19 | 0.34 |
| CPEB4 | −1.165 | 5.14 × 10−9 | 0.42 | 0.17 |
| AGFG2 | −1.126 | 5.69 × 10−16 | / | / |
| PAIP2B | −1.065 | 5.1 × 10−27 | 0.29 | 0.5 |
| ABCG1 | 1.008 | 8.69 × 10−11 | / | / |
| HOXC11 | 1.144 | 1.79 × 10−15 | −0.14 | −0.11 |
| SYT7 | 1.162 | 4.75 × 10−3 | / | / |
| CMTM3 | 1.381 | 3.08 × 10−13 | / | / |
| SLC7A8 | 1.626 | 8.78 × 10−10 | / | / |
| EIF2AK2 | 1.644 | 1.34 × 10−20 | −0.18 | −0.19 |
| WDR72 | 1.693 | 1.08 × 10−4 | / | / |
| CTSC | 1.72 | 1.88 × 10−23 | −0.14 | −0.16 |
| NPNT | 1.869 | 5.04 × 10−17 | / | / |
| PDPN | 2.566 | 1.09 × 10−19 | / | / |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Crimi, S.; Falzone, L.; Gattuso, G.; Grillo, C.M.; Candido, S.; Bianchi, A.; Libra, M. Droplet Digital PCR Analysis of Liquid Biopsy Samples Unveils the Diagnostic Role of hsa-miR-133a-3p and hsa-miR-375-3p in Oral Cancer. Biology 2020, 9, 379. https://doi.org/10.3390/biology9110379
Crimi S, Falzone L, Gattuso G, Grillo CM, Candido S, Bianchi A, Libra M. Droplet Digital PCR Analysis of Liquid Biopsy Samples Unveils the Diagnostic Role of hsa-miR-133a-3p and hsa-miR-375-3p in Oral Cancer. Biology. 2020; 9(11):379. https://doi.org/10.3390/biology9110379
Chicago/Turabian StyleCrimi, Salvatore, Luca Falzone, Giuseppe Gattuso, Caterina Maria Grillo, Saverio Candido, Alberto Bianchi, and Massimo Libra. 2020. "Droplet Digital PCR Analysis of Liquid Biopsy Samples Unveils the Diagnostic Role of hsa-miR-133a-3p and hsa-miR-375-3p in Oral Cancer" Biology 9, no. 11: 379. https://doi.org/10.3390/biology9110379
APA StyleCrimi, S., Falzone, L., Gattuso, G., Grillo, C. M., Candido, S., Bianchi, A., & Libra, M. (2020). Droplet Digital PCR Analysis of Liquid Biopsy Samples Unveils the Diagnostic Role of hsa-miR-133a-3p and hsa-miR-375-3p in Oral Cancer. Biology, 9(11), 379. https://doi.org/10.3390/biology9110379











