Microsyntenic Clusters Reveal Conservation of lncRNAs in Chordates Despite Absence of Sequence Conservation
Abstract
1. Introduction
2. Materials and Methods
2.1. Amphioxus and Human Coding and lincRNA Datasets
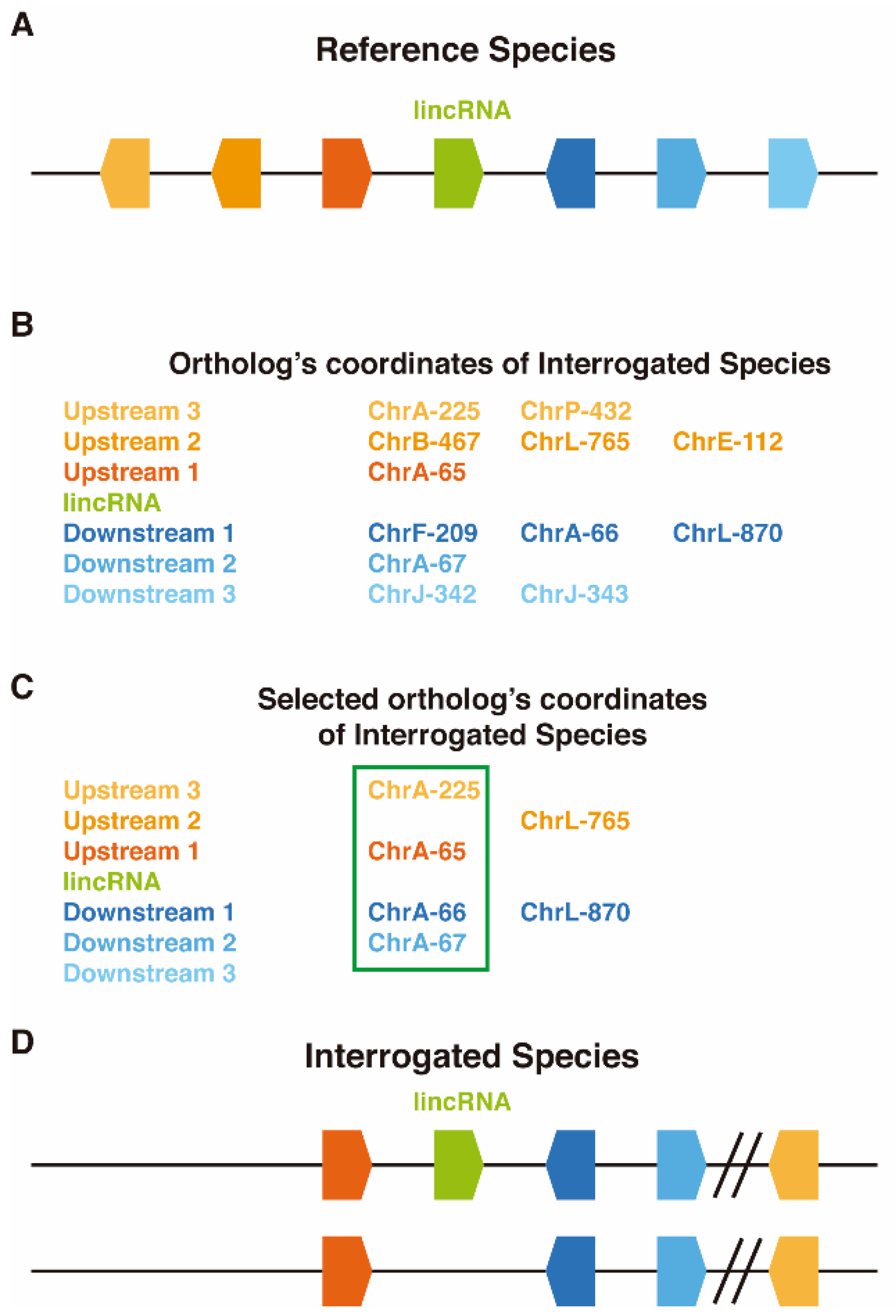
2.2. LincOFinder (lincRNA Orthology Finder)
2.3. Xenopus Embryos and MO Injections
2.4. Xenopus RNA Extraction, cDNA Synthesis, PCR and Quantitative PCR
2.5. Xenopus Whole-Mount In Situ Hybridization
2.6. Amphioxus Embryo Collection and Whole-Mount In Situ Hybridization
3. Results and Discussion
3.1. Conserved Microsynteny Clusters
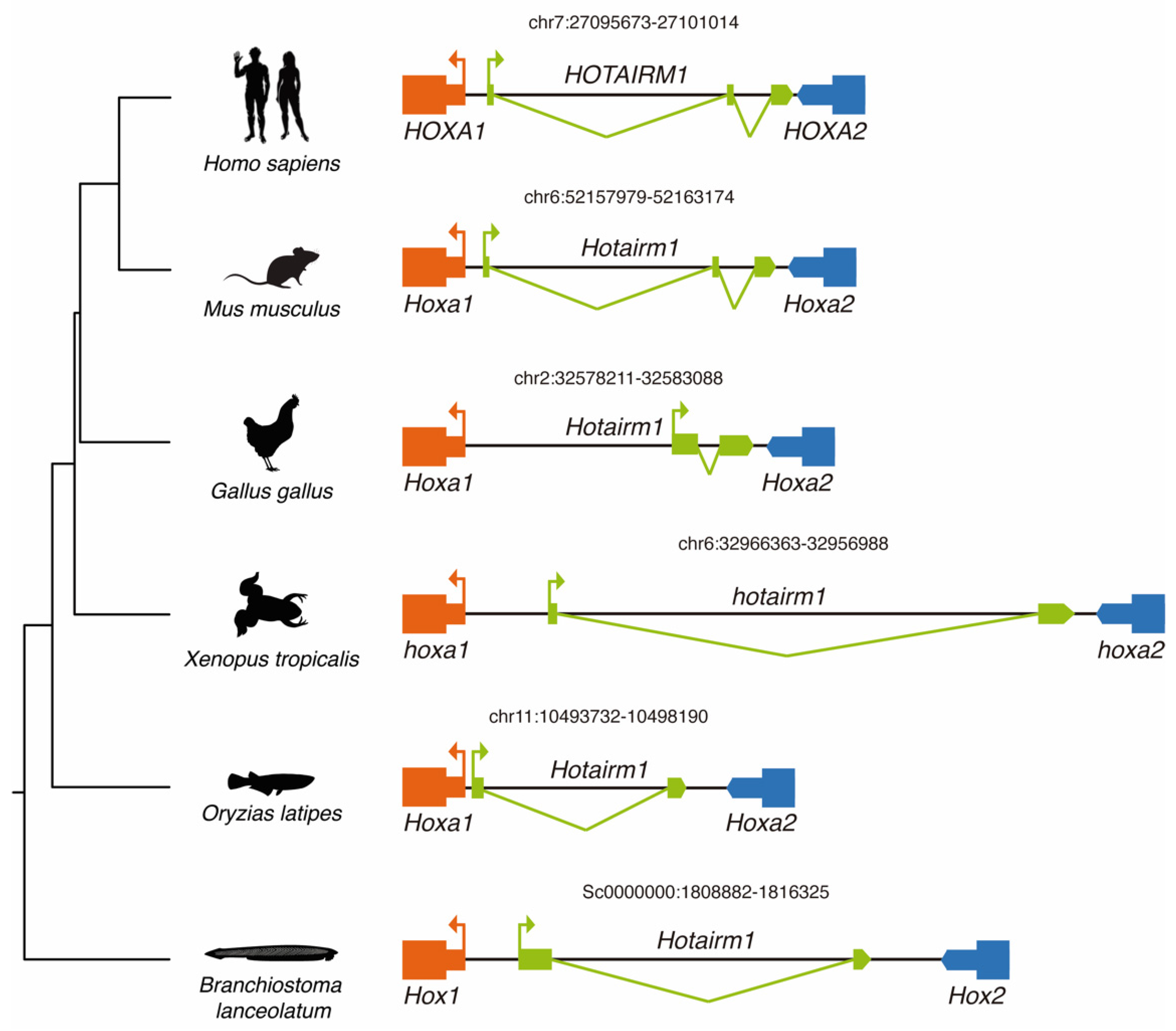
3.2. Conservation of HOTAIRM1 across Chordates
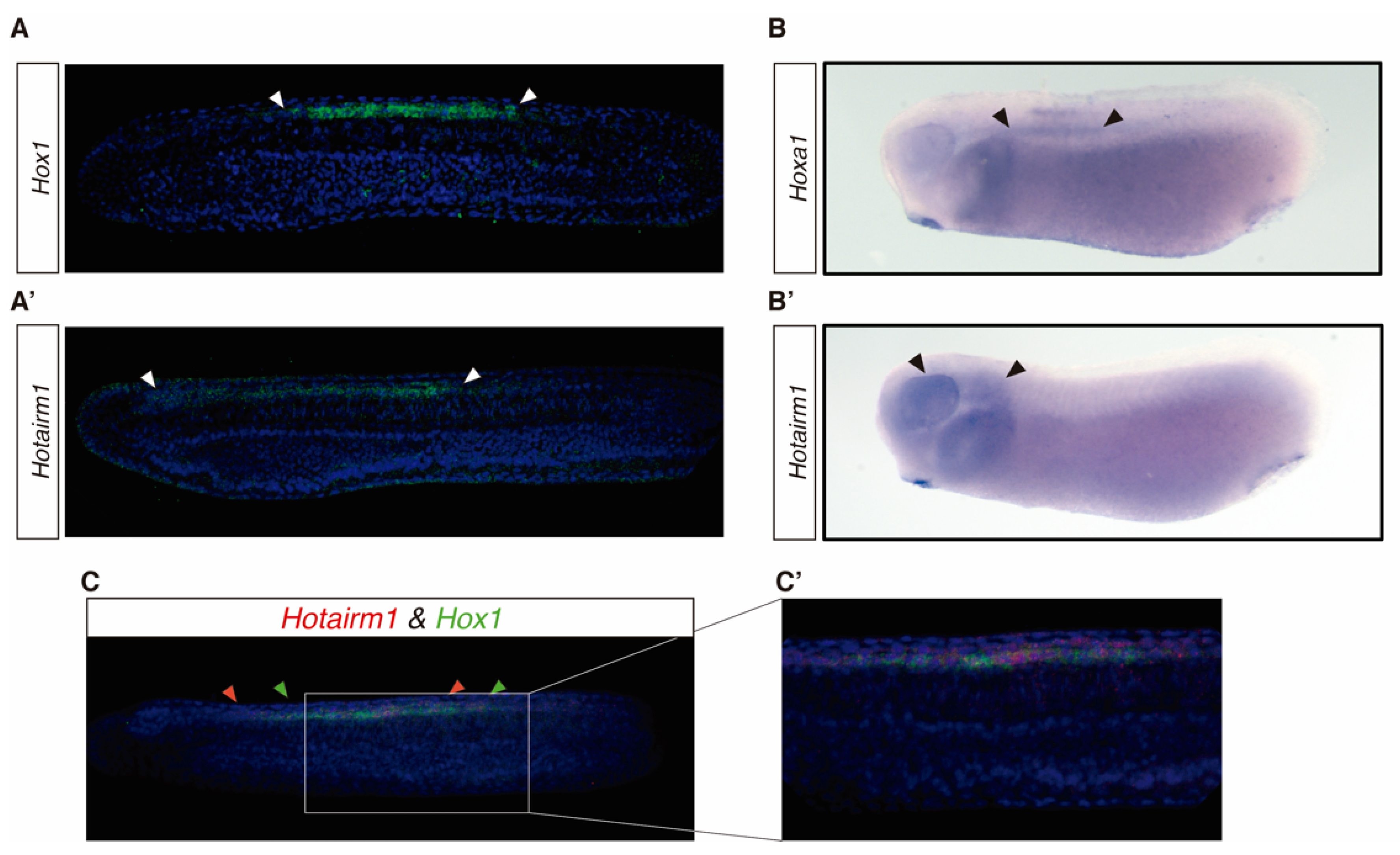
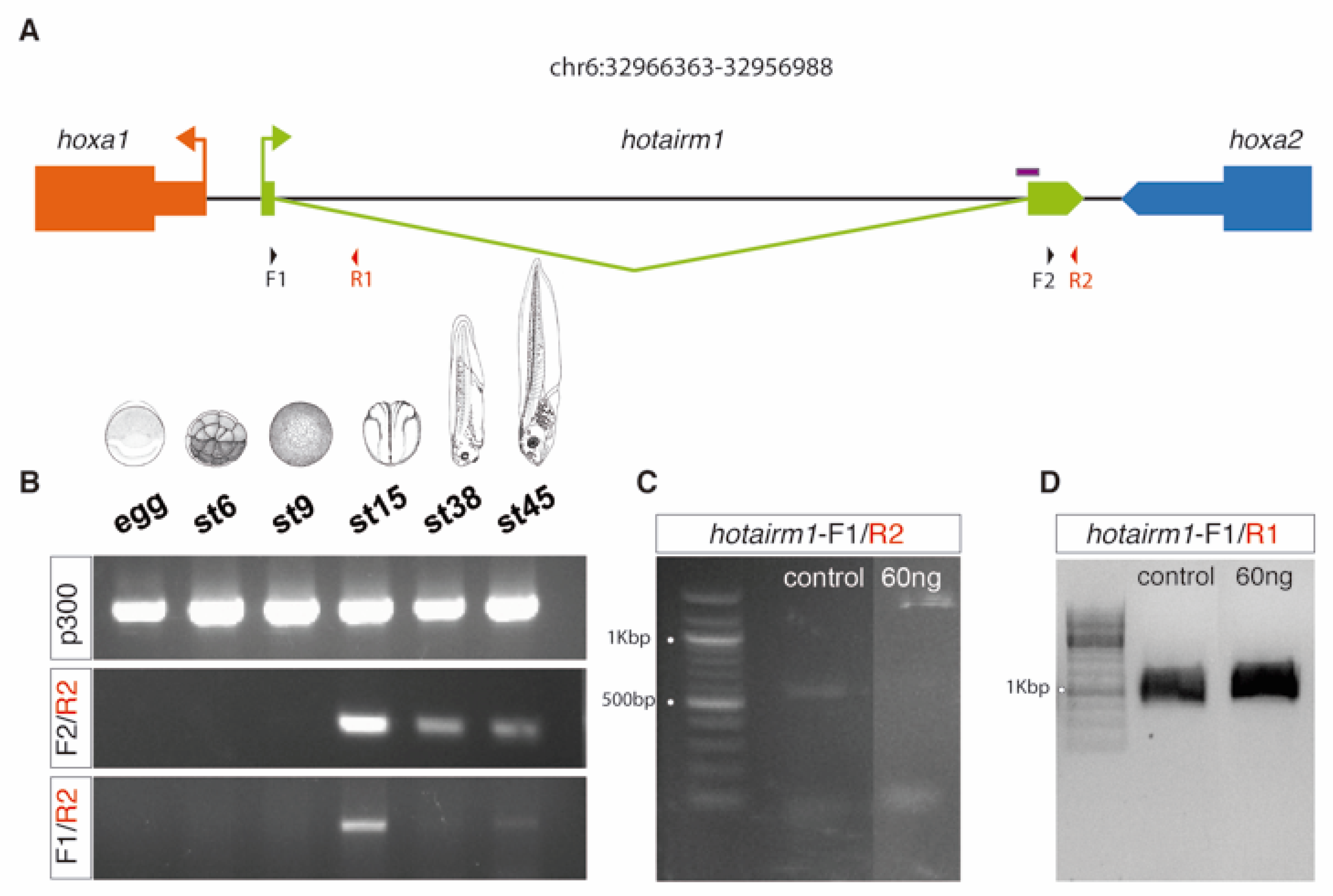
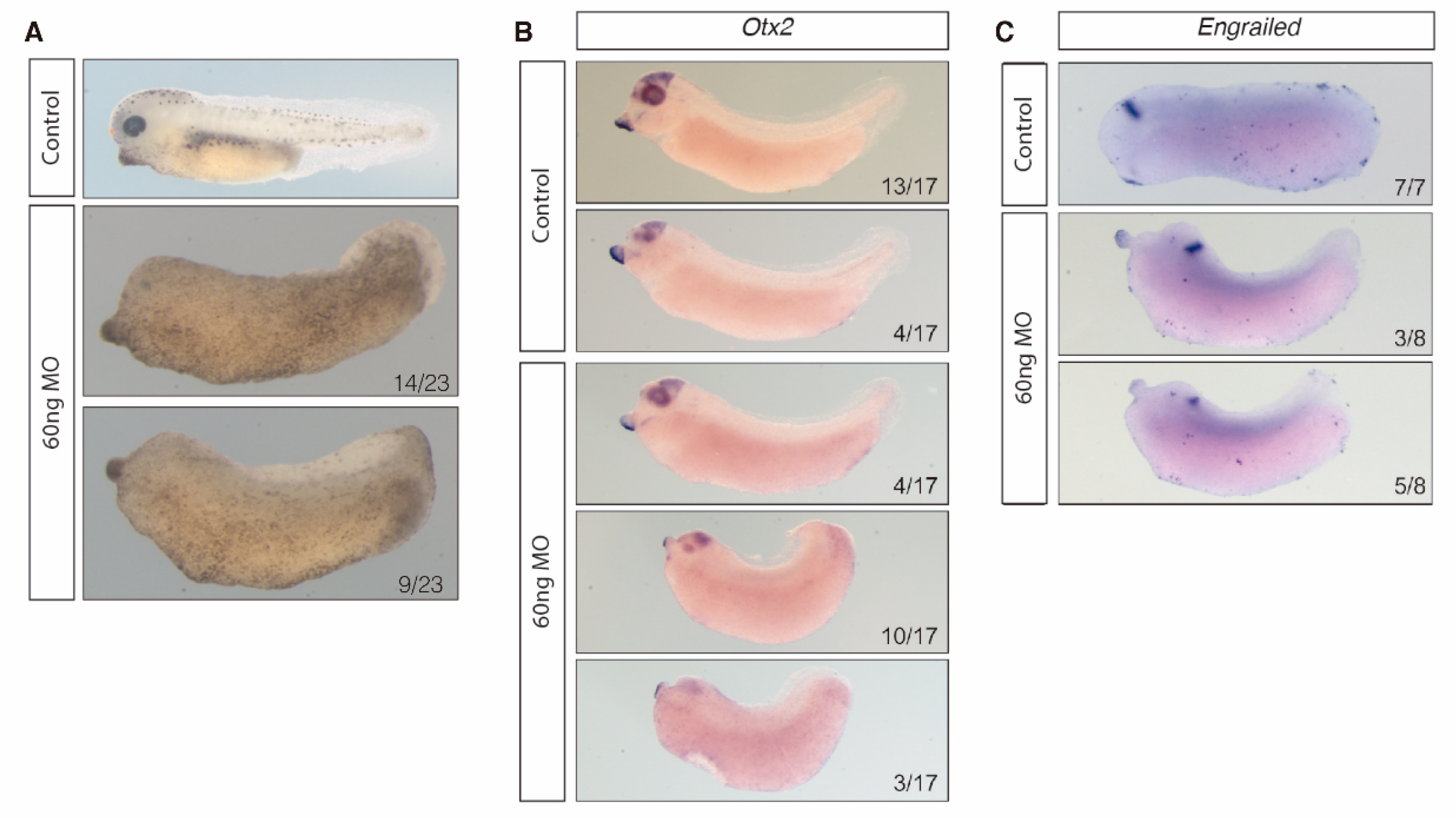
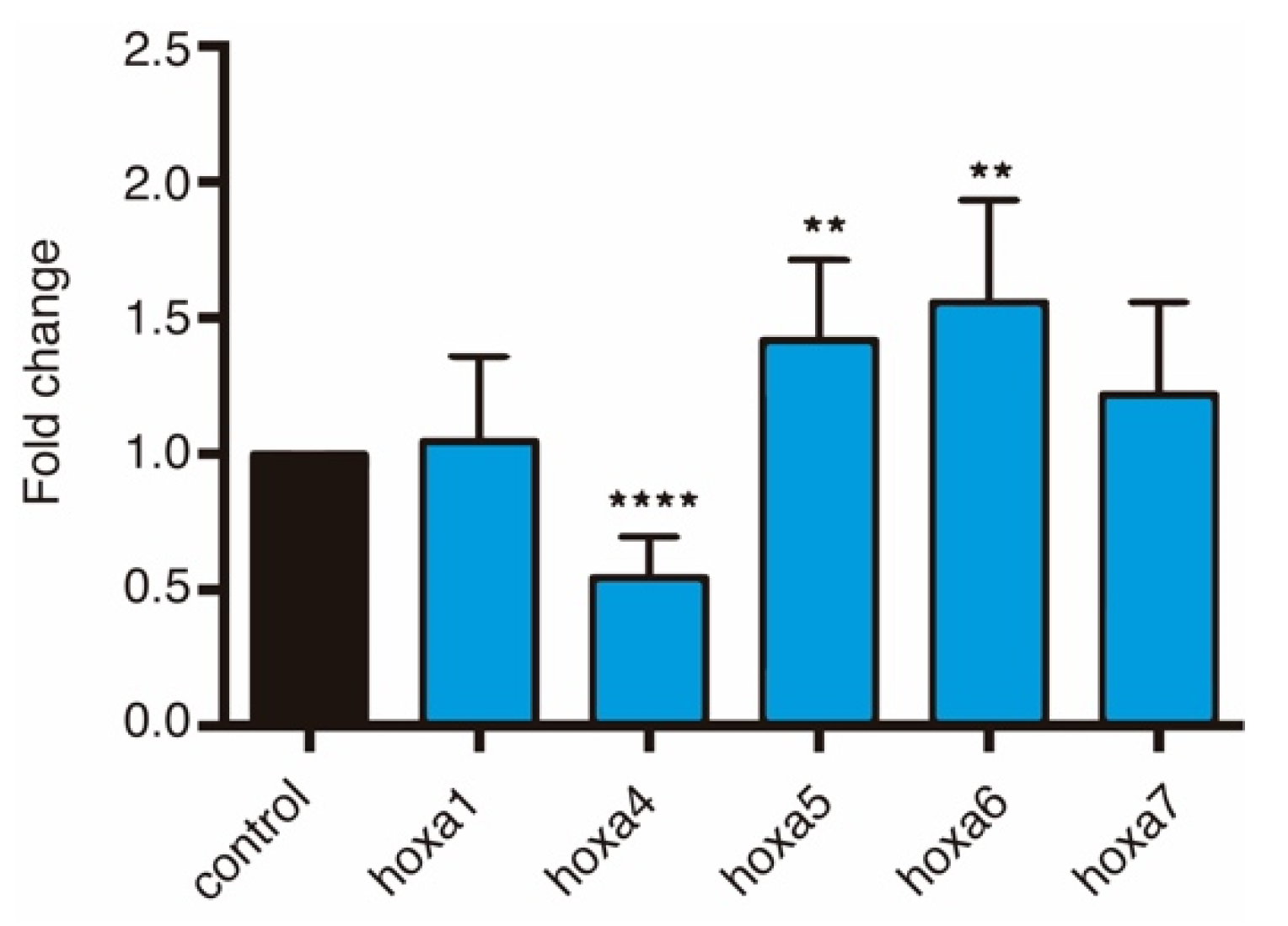
3.3. HOTAIRM1 Expression Patterns in Amphioxus and Xenopus
3.4. HOTAIRM1 Function and Expression Conservation
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Holland, P.W.H.; Garcia-Fernàndez, J.; Williams, N.A.; Sidow, A. Gene duplications and the origins of vertebrate development. Development 1994, 1994, 125–133. [Google Scholar]
- Schmitz, J.F.; Zimmer, F.; Bornberg-Bauer, E. Mechanisms of transcription factor evolution in Metazoa. Nucleic Acids Res. 2016, 44, 6287–6297. [Google Scholar] [CrossRef] [PubMed]
- Morris, K.V.; Mattick, J.S. The rise of regulatory RNA. Nat. Rev. Genet. 2014, 15, 423–437. [Google Scholar] [CrossRef] [PubMed]
- Zampetaki, A.; Albrecht, A.; Steinhofel, K. Long Non-coding RNA Structure and Function: Is There a Link? Front. Physiol. 2018, 9, 1201. [Google Scholar] [CrossRef] [PubMed]
- Wan, Y.; Kertesz, M.; Spitale, R.C.; Segal, E.; Chang, H.Y. Understanding the transcriptome through RNA structure. Nat. Rev. Genet. 2011, 12, 641–655. [Google Scholar] [CrossRef] [PubMed]
- Ponting, C.P.; Oliver, P.L.; Reik, W. Evolution and functions of long noncoding RNAs. Cell 2009, 136, 629–641. [Google Scholar] [CrossRef]
- Fico, A.; Fiorenzano, A.; Pascale, E.; Patriarca, E.J.; Minchiotti, G. Long non-coding RNA in stem cell pluripotency and lineage commitment: functions and evolutionary conservation. Cell. Mol. Life Sci. 2019, 76, 1459–1471. [Google Scholar] [CrossRef]
- Diederichs, S. The four dimensions of noncoding RNA conservation. Trends Genet. 2014, 30, 121–123. [Google Scholar] [CrossRef]
- Jathar, S.; Kumar, V.; Srivastava, J.; Tripathi, V. Technological developments in lncRNA biology. In Advances in Experimental Medicine and Biology; Springer: Singapore, 2017; Vol. 1008, pp. 283–323. [Google Scholar]
- Rivas, E.; Clements, J.; Eddy, S.R. A statistical test for conserved RNA structure shows lack of evidence for structure in lncRNAs. Nat. Methods 2017, 14, 45–48. [Google Scholar] [CrossRef]
- Neme, R.; Tautz, D. Fast turnover of genome transcription across evolutionary time exposes entire non-coding DNA to de novo gene emergence. Elife 2016, 5. [Google Scholar] [CrossRef]
- Garcia-Fernandez, J.; Benito-Gutierrez, E. It’s a long way from amphioxus: descendants of the earliest chordate. Bioessays 2009, 31, 665–675. [Google Scholar] [CrossRef]
- Paps, J.; Holland, P.W.H.; Shimeld, S.M. A genome-wide view of transcription factor gene diversity in chordate evolution: less gene loss in amphioxus? Brief. Funct. Genomics 2012, 11, 177–186. [Google Scholar] [CrossRef][Green Version]
- Putnam, N.H.; Butts, T.; Ferrier, D.E.K.; Furlong, R.F.; Hellsten, U.; Kawashima, T.; Robinson-Rechavi, M.; Shoguchi, E.; Terry, A.; Yu, J.-K.; et al. The amphioxus genome and the evolution of the chordate karyotype. Nature 2008, 453, 1064–1071. [Google Scholar] [CrossRef]
- Marlétaz, F.; Firbas, P.N.; Maeso, I.; Tena, J.J.; Bogdanovic, O.; Perry, M.; Wyatt, C.D.R.; de la Calle-Mustienes, E.; Bertrand, S.; Burguera, D.; et al. Amphioxus functional genomics and the origins of vertebrate gene regulation. Nature 2018, 564, 64–70. [Google Scholar] [CrossRef]
- Bertrand, S.; Escriva, H.; Williams, N.A.; Holland, N.D.; Holland, L.Z. Evolutionary crossroads in developmental biology: amphioxus. Development 2011, 138, 4819–4830. [Google Scholar] [CrossRef]
- Pegueroles, C.; Iraola-Guzmán, S.; Chorostecki, U.; Ksiezopolska, E.; Saus, E.; Gabaldón, T. Transcriptomic analyses reveal groups of co-expressed, syntenic lncRNAs in four species of the genus Caenorhabditis. RNA Biol. 2019, 16, 320–329. [Google Scholar] [CrossRef]
- Bush, S.J.; Muriuki, C.; McCulloch, M.E.B.; Farquhar, I.L.; Clark, E.L.; Hume, D.A. Cross-species inference of long non-coding RNAs greatly expands the ruminant transcriptome. Genet. Sel. Evol. 2018, 50, 20. [Google Scholar] [CrossRef]
- Zerbino, D.R.; Achuthan, P.; Akanni, W.; Amode, M.R.; Barrell, D.; Bhai, J.; Billis, K.; Cummins, C.; Gall, A.; Girón, C.G.; et al. Ensembl 2018. Nucleic Acids Res. 2018, 46, D754–D761. [Google Scholar] [CrossRef]
- Emms, D.M.; Kelly, S. OrthoFinder: solving fundamental biases in whole genome comparisons dramatically improves orthogroup inference accuracy. Genome Biol. 2015, 16, 157. [Google Scholar] [CrossRef]
- Bogdanovic, O.; Alexis, M.S.; Tena, J.J.; Maeso, I.; Fernandez-Minan, A.; Fraser, H.B.; Gomez-Skarmeta, J.L.; Roy, S.W.; Irimia, M.; de la Calle-Mustienes, E. Extensive conservation of ancient microsynteny across metazoans due to cis-regulatory constraints. Genome Res. 2012, 22, 2356–2367. [Google Scholar]
- Sokal, R.R. A statistical method for evaluating systematic relationships. Univ. Kansas, Sci. Bull. 1958, 38, 1409–1438. [Google Scholar]
- Kent, W.J.; Sugnet, C.W.; Furey, T.S.; Roskin, K.M.; Pringle, T.H.; Zahler, A.M.; Haussler, D. The human genome browser at UCSC. Genome Res. 2002, 12, 996–1006. [Google Scholar] [CrossRef]
- Monsoro-Burq, A.H. A Rapid Protocol for Whole-Mount In Situ Hybridization on Xenopus Embryos. Cold Spring Harb. Protoc. 2007, 2007. [Google Scholar] [CrossRef]
- Fuentes, M.; Benito, E.; Bertrand, S.; Paris, M.; Mignardot, A.; Godoy, L.; Jimenez-Delgado, S.; Oliveri, D.; Candiani, S.; Hirsinger, E.; et al. Insights into spawning behavior and development of the european amphioxus (Branchiostoma lanceolatum). J. Exp. Zool. Part B Mol. Dev. Evol. 2007, 308B, 484–493. [Google Scholar] [CrossRef]
- Choi, H.M.T.; Schwarzkopf, M.; Fornace, M.E.; Acharya, A.; Artavanis, G.; Stegmaier, J.; Cunha, A.; Pierce, N.A. Third-generation in situ hybridization chain reaction: multiplexed, quantitative, sensitive, versatile, robust. Development 2018, 145, dev165753. [Google Scholar] [CrossRef]
- Ulitsky, I. Evolution to the rescue: using comparative genomics to understand long non-coding RNAs. Nat. Rev. Genet. 2016, 17, 601–614. [Google Scholar] [CrossRef]
- Yu, H.; Lindsay, J.; Feng, Z.-P.; Frankenberg, S.; Hu, Y.; Carone, D.; Shaw, G.; Pask, A.J.; O’Neill, R.; Papenfuss, A.T.; et al. Evolution of coding and non-coding genes in HOX clusters of a marsupial. BMC Genomics 2012, 13, 251. [Google Scholar] [CrossRef]
- Gardner, P.P.; Fasold, M.; Burge, S.W.; Ninova, M.; Hertel, J.; Kehr, S.; Steeves, T.E.; Griffiths-Jones, S.; Stadler, P.F. Conservation and Losses of Non-Coding RNAs in Avian Genomes. PLoS One 2015, 10, e0121797. [Google Scholar] [CrossRef]
- Wang, X.Q.D.; Dostie, J. Reciprocal regulation of chromatin state and architecture by HOTAIRM1 contributes to temporal collinear HOXA gene activation. Nucleic Acids Res. 2017, 45, 1091–1104. [Google Scholar] [CrossRef]
- Zhang, X.; Lian, Z.; Padden, C.; Gerstein, M.B.; Rozowsky, J.; Snyder, M.; Gingeras, T.R.; Kapranov, P.; Weissman, S.M.; Newburger, P.E. A myelopoiesis-associated regulatory intergenic noncoding RNA transcript within the human HOXA cluster. Blood 2009, 113, 2526–2534. [Google Scholar] [CrossRef]
- Sekigami, Y.; Kobayashi, T.; Omi, A.; Nishitsuji, K.; Ikuta, T.; Fujiyama, A.; Satoh, N.; Saiga, H. Hox gene cluster of the ascidian, Halocynthia roretzi, reveals multiple ancient steps of cluster disintegration during ascidian evolution. Zool. Lett. 2017, 3, 17. [Google Scholar] [CrossRef]
- Pascual-Anaya, J.; Sato, I.; Sugahara, F.; Higuchi, S.; Paps, J.; Ren, Y.; Takagi, W.; Ruiz-Villalba, A.; Ota, K.G.; Wang, W.; et al. Hagfish and lamprey Hox genes reveal conservation of temporal colinearity in vertebrates. Nat. Ecol. Evol. 2018, 2, 859–866. [Google Scholar] [CrossRef]
- Dehal, P.; Boore, J.L. Two Rounds of Whole Genome Duplication in the Ancestral Vertebrate. PLoS Biol. 2005, 3, e314. [Google Scholar] [CrossRef]
- Esfandi, F.; Taheri, M.; Omrani, M.D.; Shadmehr, M.B.; Arsang-Jang, S.; Shams, R.; Ghafouri-Fard, S. Expression of long non-coding RNAs (lncRNAs) has been dysregulated in non-small cell lung cancer tissues. BMC Cancer 2019, 19, 222. [Google Scholar] [CrossRef]
- Li, Q.; Dong, C.; Cui, J.; Wang, Y.; Hong, X. Over-expressed lncRNA HOTAIRM1 promotes tumor growth and invasion through up-regulating HOXA1 and sequestering G9a/EZH2/Dnmts away from the HOXA1 gene in glioblastoma multiforme. J. Exp. Clin. Cancer Res. 2018, 37, 265. [Google Scholar] [CrossRef]
- Song, L.; Zhang, S.; Duan, C.; Ma, S.; Hussain, S.; Wei, L.; Chu, M. Genome-wide identification of lncRNAs as novel prognosis biomarkers of glioma. J. Cell. Biochem. 2019. [Google Scholar] [CrossRef]
- Lin, M.; Pedrosa, E.; Shah, A.; Hrabovsky, A.; Maqbool, S.; Zheng, D.; Lachman, H.M. RNA-Seq of Human Neurons Derived from iPS Cells Reveals Candidate Long Non-Coding RNAs Involved in Neurogenesis and Neuropsychiatric Disorders. PLoS One 2011, 6, e23356. [Google Scholar] [CrossRef]
- Albuixech-Crespo, B.; López-Blanch, L.; Burguera, D.; Maeso, I.; Sánchez-Arrones, L.; Moreno-Bravo, J.A.; Somorjai, I.; Pascual-Anaya, J.; Puelles, E.; Bovolenta, P.; et al. Molecular regionalization of the developing amphioxus neural tube challenges major partitions of the vertebrate brain. PLoS Biol. 2017, 15, e2001573. [Google Scholar] [CrossRef]
- Albuixech-Crespo, B.; Herrera-Úbeda, C.; Marfany, G.; Irimia, M.; Garcia-Fernàndez, J. Origin and evolution of the chordate central nervous system: insights from amphioxus genoarchitecture. Int. J. Dev. Biol. 2017, 61, 655–664. [Google Scholar] [CrossRef]
- Schubert, M.; Holland, N.D.; Laudet, V.; Holland, L.Z. A retinoic acid-Hox hierarchy controls both anterior/posterior patterning and neuronal specification in the developing central nervous system of the cephalochordate amphioxus. Dev. Biol. 2006, 296, 190–202. [Google Scholar] [CrossRef]
- Zieger, E.; Candiani, S.; Garbarino, G.; Croce, J.C.; Schubert, M. Roles of Retinoic Acid Signaling in Shaping the Neuronal Architecture of the Developing Amphioxus Nervous System. Mol. Neurobiol. 2018, 55, 5210–5229. [Google Scholar] [CrossRef]
- McNulty, C.L.; Peres, J.N.; Bardine, N.; van den Akker, W.M.R.; Durston, A.J. Knockdown of the complete Hox paralogous group 1 leads to dramatic hindbrain and neural crest defects. Development 2005, 132, 2861–2871. [Google Scholar] [CrossRef]
| Orthologous lincRNAs 1 | State of the Cluster in Human 2 | Human Orthologous lincRNA 3 |
|---|---|---|
| BL20528|Sc0000000|28|+ | * Conserved microsynteny | ENST00000623777.1_1 |
| BL38782|Sc0000000|30|+ | * Conserved microsynteny | HOTAIRM1 |
| BL90848|Sc0000001|150|− | Correct order but strands inverted. In addition, there are two lncRNAs surrounding the cluster | AL354977.2 |
| BL79733|Sc0000007|52|+ | One gene with the strand inverted | BANCR |
| One gene with the strand inverted | TCONS_12_00008513 | |
| BL84418|Sc0000009|86|+ | * Conserved microsynteny | TCONS_00027655 |
| BL91140|Sc0000010|144|− | Couple of lincRNAs in amphioxus, synteny conserved in the coding genes, but not in the lincRNA | TCONS_00006308 |
| BL91143|Sc0000010|145|- | * Conserved microsynteny | RP11-181G12.4 |
| BL53024|Sc0000015|55|+ | One gene with the strand inverted | TCONS_00027115 |
| BL82992|Sc0000016|15|− | One gene with the strand inverted | TCONS_00000550 |
| BL78145|Sc0000039|54|+ | * Conserved microsynteny | TCONS_00024711 |
| BL55463|Sc0000050|45|+ | * Conserved microsynteny | TCONS_00011710 |
| BL68900|Sc0000072|2|+ | Synteny conserved in the coding genes | AI219887 |
| BL54861|Sc0000089|4|− | Problematic region with several lincRNAs and massive distances | AC109136.1 |
| Problematic region with several lincRNAs and massive distances | AC124852.1 | |
| BL41904|Sc0000219|3|− | * Conserved microsynteny | TCONS_00007813 |
| BL72725|Sc0000229|14|+ | * Conserved microsynteny | BC043517 |
| * Conserved microsynteny | LINC00114 | |
| BL59605|Sc0000234|6|− | * Conserved microsynteny | TCONS_00011870 |
| BL38170|Sc0000240|5|+ | * Conserved microsynteny | LOC100132215 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Herrera-Úbeda, C.; Marín-Barba, M.; Navas-Pérez, E.; Gravemeyer, J.; Albuixech-Crespo, B.; Wheeler, G.N.; Garcia-Fernàndez, J. Microsyntenic Clusters Reveal Conservation of lncRNAs in Chordates Despite Absence of Sequence Conservation. Biology 2019, 8, 61. https://doi.org/10.3390/biology8030061
Herrera-Úbeda C, Marín-Barba M, Navas-Pérez E, Gravemeyer J, Albuixech-Crespo B, Wheeler GN, Garcia-Fernàndez J. Microsyntenic Clusters Reveal Conservation of lncRNAs in Chordates Despite Absence of Sequence Conservation. Biology. 2019; 8(3):61. https://doi.org/10.3390/biology8030061
Chicago/Turabian StyleHerrera-Úbeda, Carlos, Marta Marín-Barba, Enrique Navas-Pérez, Jan Gravemeyer, Beatriz Albuixech-Crespo, Grant N. Wheeler, and Jordi Garcia-Fernàndez. 2019. "Microsyntenic Clusters Reveal Conservation of lncRNAs in Chordates Despite Absence of Sequence Conservation" Biology 8, no. 3: 61. https://doi.org/10.3390/biology8030061
APA StyleHerrera-Úbeda, C., Marín-Barba, M., Navas-Pérez, E., Gravemeyer, J., Albuixech-Crespo, B., Wheeler, G. N., & Garcia-Fernàndez, J. (2019). Microsyntenic Clusters Reveal Conservation of lncRNAs in Chordates Despite Absence of Sequence Conservation. Biology, 8(3), 61. https://doi.org/10.3390/biology8030061






