Comprehensive Analysis of WUSCEL-Related Homeobox Gene Family in Ramie (Boehmeria nivea) Indicates Its Potential Role in Adventitious Root Development
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Identification of WOX Genes in Ramie
2.2. Multiple Sequence Alignment and Phylogenetic Analysis
2.3. Chromosomal Location, Synteny Analysis, and Gene Duplication
2.4. Gene Structure and Motif Analysis
2.5. Expression Patterns Analysis
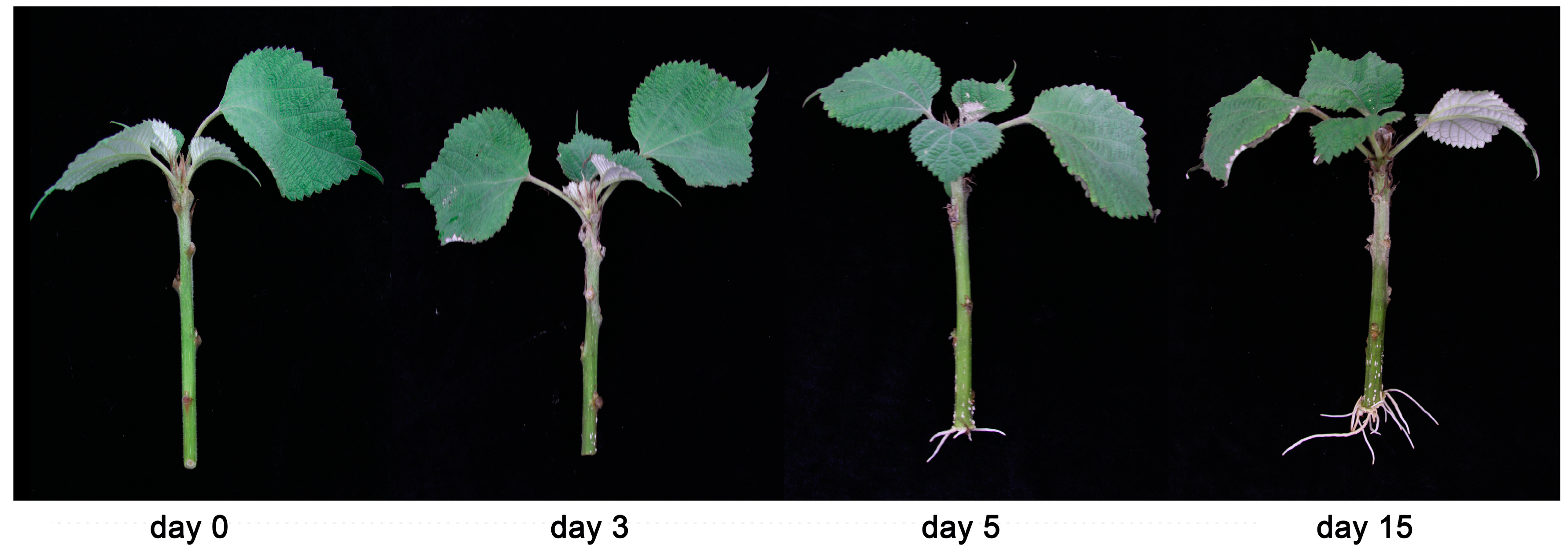
2.6. Adventitious Root Induction and Phenotypic Data
2.7. Cis-Acting Element Analysis of the BnWOX14 in Ramie Elite Genetic Resources
3. Results
3.1. Identification of Ramie WOX Genes
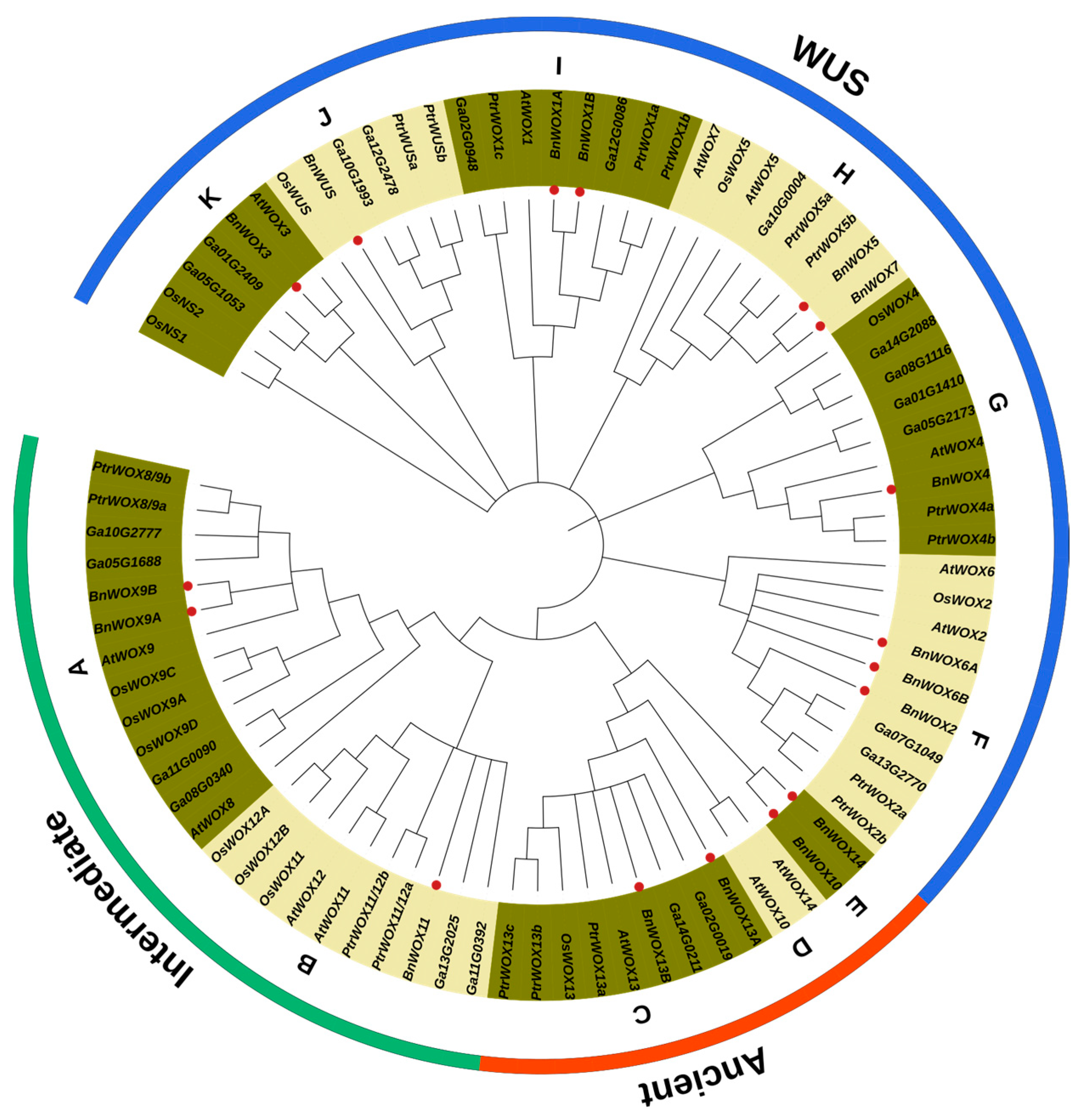
3.2. Multiple Sequence Alignment and Phylogenetic Analysis
3.3. Chromosomal Location
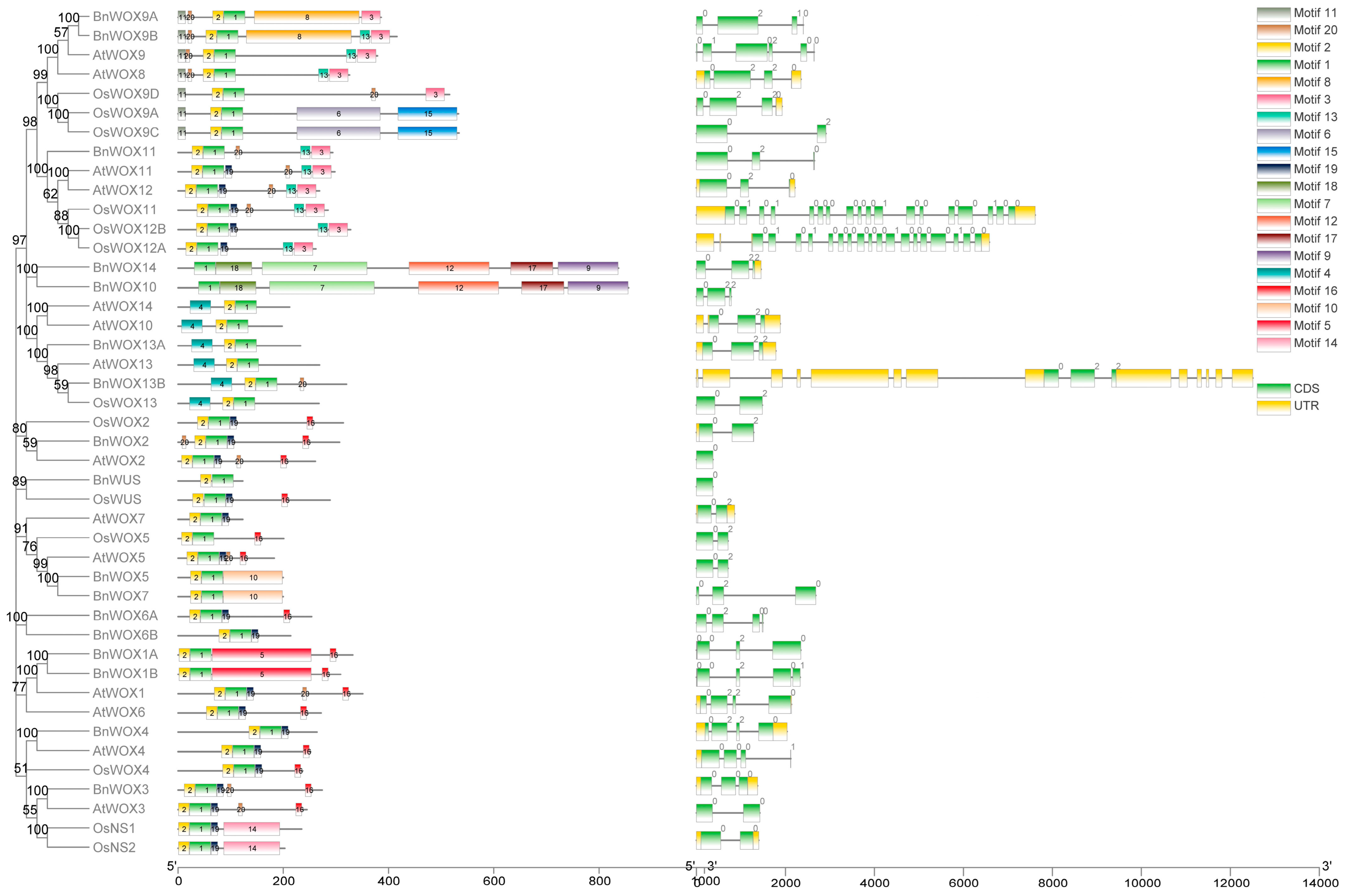
3.4. Gene Structure and Motif Analysis
3.5. Gene Duplications and Synteny Analysis
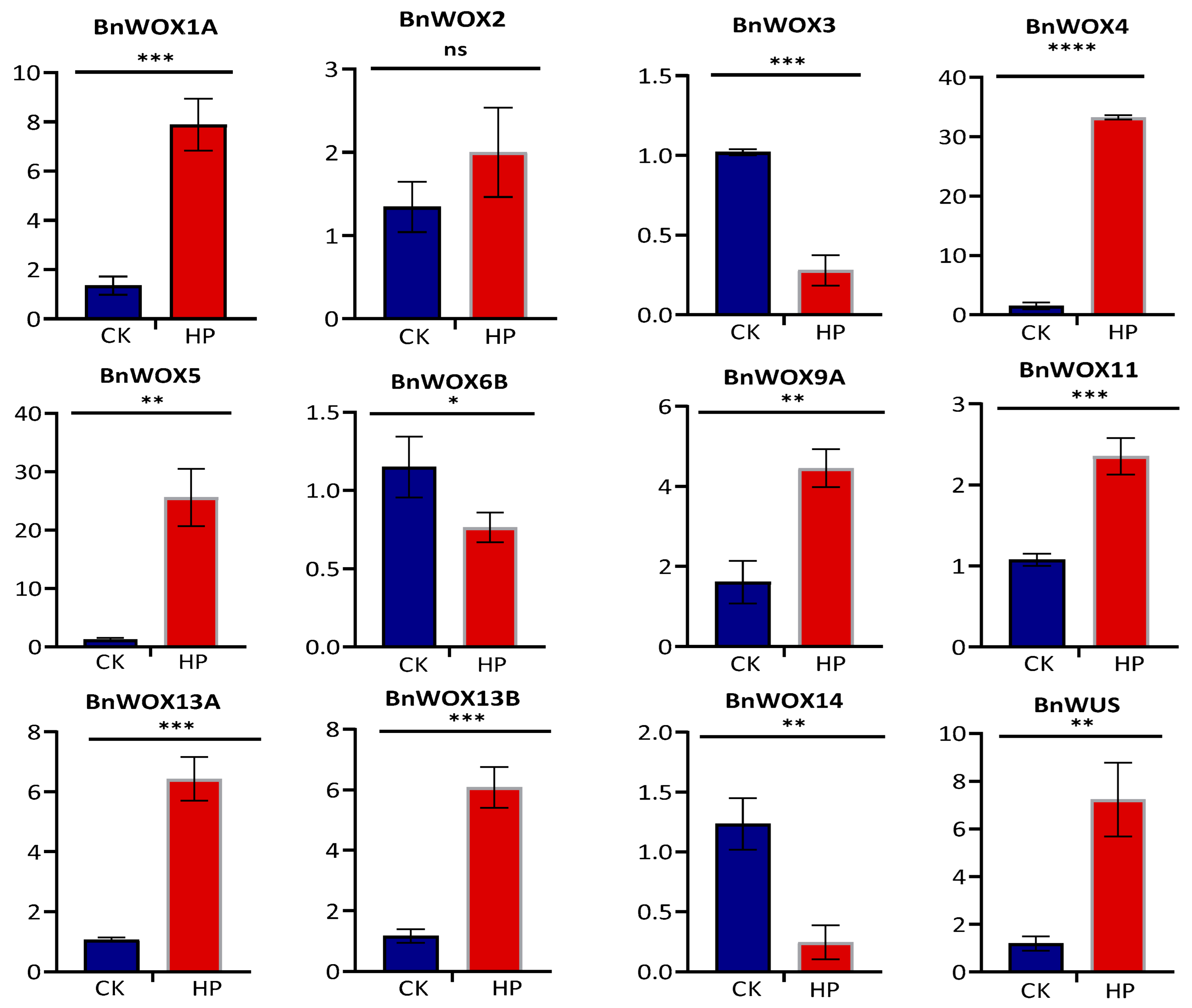
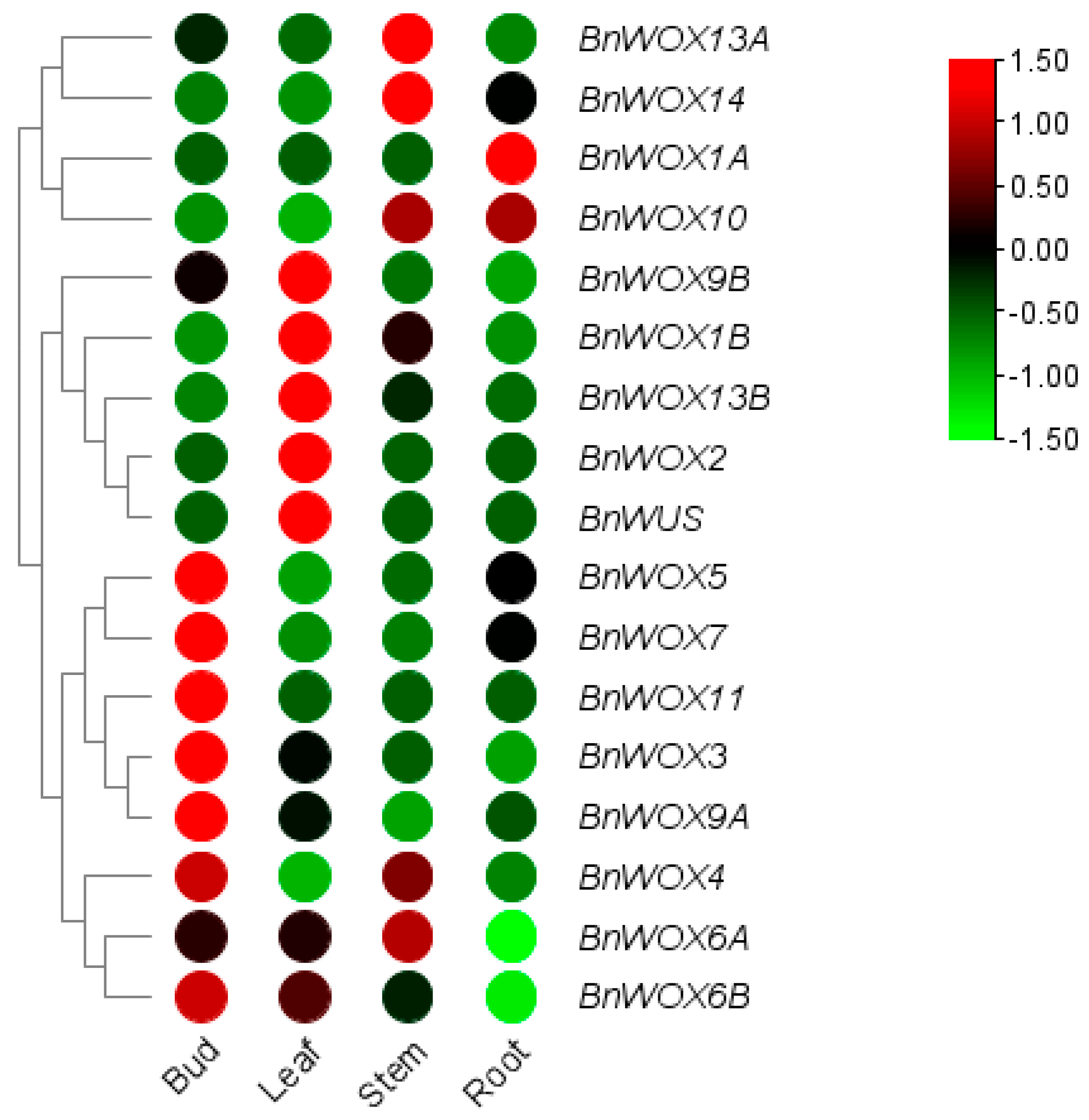
3.6. Expression Pattern Analysis of the BnWOX Genes in Ramie Stem Cuttings of Hydroponic Elite Cultivar
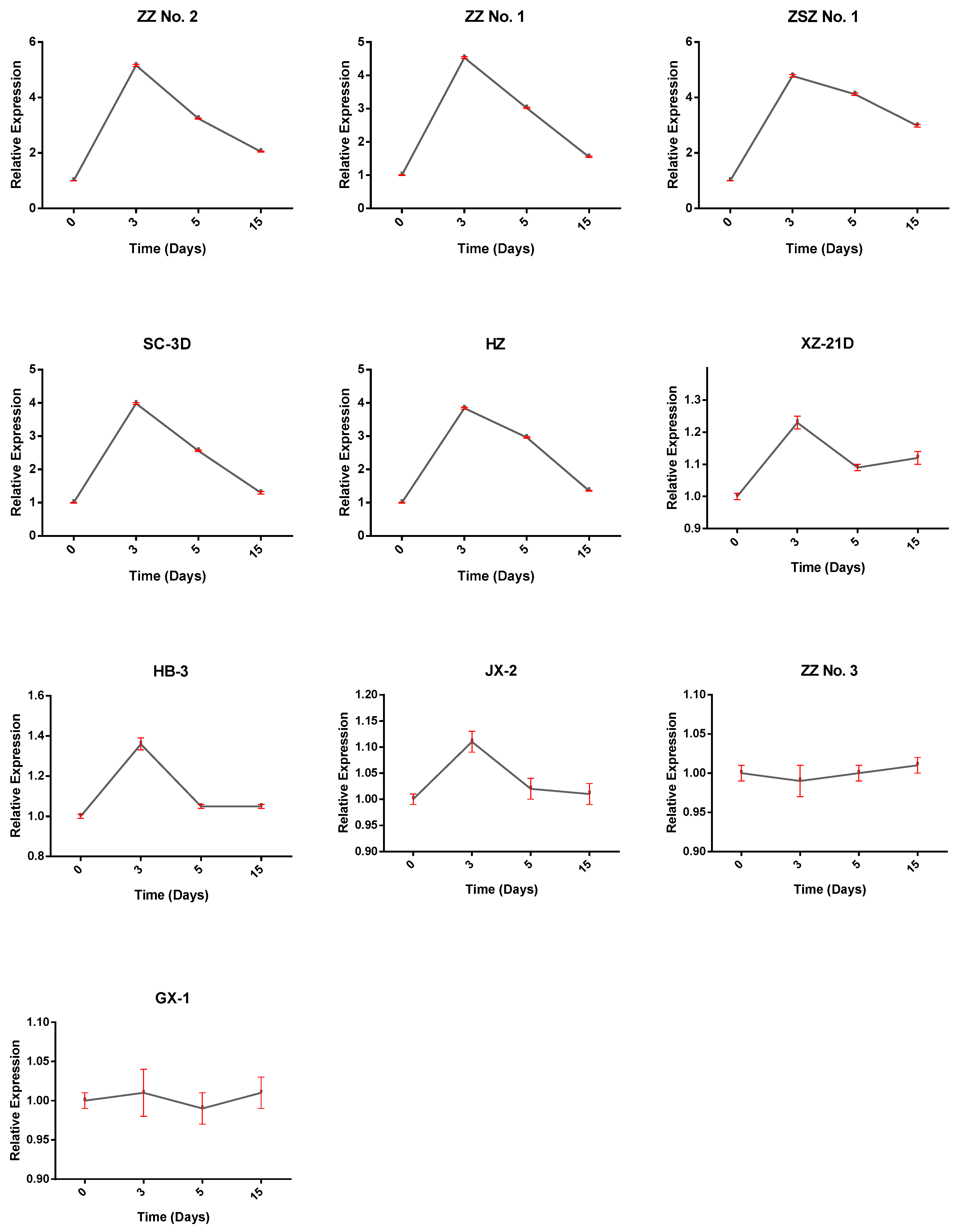
3.7. Expression of BnWOX14 and Adventitious Root Development in Different Ramie Genetic Resource
3.8. Cis-Acting Element Analysis of the BnWOX14 in the Elite Genetic Resources
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Chen, F.; He, J.; Wang, X.; Lv, T.; Liu, C.; Liao, L.; Li, Z.; Zhou, J.; He, B.; Qiu, H.; et al. Effect of Dietary Ramie Powder at Various Levels on the Growth Performance, Meat Quality, Serum Biochemical Indices and Antioxidative Capacity of Yanling White Geese. Animals 2022, 12, 2045. [Google Scholar] [CrossRef] [PubMed]
- Gao, G.; Xiong, H.; Chen, J.; Chen, K.; Chen, P.; Yu, C.; Zhu, A. Hydroponic Method for Ramie and Removal of Nitrogen and Phosphorus from Livestock Wastewater. Int. J. Phytoremediation 2018, 20, 545–551. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.; Guo, B.; Yu, C.; Chen, P.; Chen, J.; Gao, G.; Wang, X.; Zhu, A. Comparative Transcriptome Analysis Provides New Insights into the Molecular Regulatory Mechanism of Adventitious Root Formation in Ramie (Boehmeria nivea L.). Plants 2021, 10, 160. [Google Scholar] [CrossRef]
- Abubakar, A.S.; Yahaya, S.U.; Shaibu, A.S.; Yahaya, S.U.; Ibrahim, H.; Ibrahim, A.K.; Lawan, Z.M.; Isa, A.M. In Vitro Propagation of Sweet Potato (Ipomoea batatas (L.) Lam.) Cultivars. Agric. Sci. Dig. ASD 2018, 38, 17–21. [Google Scholar] [CrossRef]
- Bellini, C.; Pacurar, D.I.; Perrone, I. Adventitious Roots and Lateral Roots: Similarities and Differences. Annu. Rev. Plant Biol. 2014, 65, 639–666. [Google Scholar] [CrossRef] [PubMed]
- Sorin, C.; Negroni, L.; Balliau, T.; Corti, H.; Jacquemot, M.-P.; Davanture, M.; Sandberg, G.; Zivy, M.; Bellini, C. Proteomic Analysis of Different Mutant Genotypes of Arabidopsis Led to the Identification of 11 Proteins Correlating with Adventitious Root Development. Plant Physiol. 2006, 140, 349–364. [Google Scholar] [CrossRef]
- Yang, S.; Ma, S.; Qiu, R.; Guang, Q.; Lv, Y.; Dong, Y.; Wu, J.; Song, L. Comparative Analysis of RNA-Seq Data Reveals Adventitious Root Development Is Mediated by ChIFNα in Lotus japonicus. Biotechnol. Biotechnol. Equip. 2021, 35, 179–195. [Google Scholar] [CrossRef]
- Lakehal, A.; Bellini, C. Control of Adventitious Root Formation: Insights into Synergistic and Antagonistic Hormonal Interactions. Physiol Plant. 2019, 165, 90–100. [Google Scholar] [CrossRef] [PubMed]
- Steffens, B.; Rasmussen, A. The Physiology of Adventitious Roots. Plant Physiol. 2016, 170, 603–617. [Google Scholar] [CrossRef]
- Hong, C.; Yun, F.; JianMei, Z.; ZhenGuo, X.; Chao, L.; QiRong, G. Research advance of plant root biology. World For. Res. 2013, 26, 25–29. [Google Scholar]
- Gutierrez, L.; Bussell, J.D.; Păcurar, D.I.; Schwambach, J.; Păcurar, M.; Bellini, C. Phenotypic Plasticity of Adventitious Rooting in Arabidopsis Is Controlled by Complex Regulation of AUXIN RESPONSE FACTOR Transcripts and MicroRNA Abundance. Plant Cell 2009, 21, 3119–3132. [Google Scholar] [CrossRef]
- Krauss, K.W.; Allen, J.A.; Cahoon, D.R. Differential Rates of Vertical Accretion and Elevation Change among Aerial Root Types in Micronesian Mangrove Forests. Estuar. Coast. Shelf Sci. 2003, 56, 251–259. [Google Scholar] [CrossRef]
- Liu, W.; Xu, L. Recruitment of IC-WOX Genes in Root Evolution. Trends Plant Sci. 2018, 23, 490–496. [Google Scholar] [CrossRef] [PubMed]
- Zuo, J.; Niu, Q.-W.; Frugis, G.; Chua, N.-H. The WUSCHEL Gene Promotes Vegetative-to-Embryonic Transition in Arabidopsis. Plant J. 2002, 30, 349–359. [Google Scholar] [CrossRef] [PubMed]
- Deveaux, Y.; Toffano-Nioche, C.; Claisse, G.; Thareau, V.; Morin, H.; Laufs, P.; Moreau, H.; Kreis, M.; Lecharny, A. Genes of the Most Conserved WOX Clade in Plants Affect Root and Flower Development in Arabidopsis. BMC Evol. Biol. 2008, 8, 291. [Google Scholar] [CrossRef] [PubMed]
- Kieffer, M.; Stern, Y.; Cook, H.; Clerici, E.; Maulbetsch, C.; Laux, T.; Davies, B. Analysis of the Transcription Factor WUSCHEL and Its Functional Homologue in Antirrhinum Reveals a Potential Mechanism for Their Roles in Meristem Maintenance. Plant Cell 2006, 18, 560–573. [Google Scholar] [CrossRef] [PubMed]
- Shi, L.; Wang, K.; Du, L.; Song, Y.; Li, H.; Ye, X. Genome-Wide Identification and Expression Profiling Analysis of WOX Family Protein-Encoded Genes in Triticeae Species. Int. J. Mol. Sci. 2021, 22, 9325. [Google Scholar] [CrossRef]
- Wu, X.; Dabi, T.; Weigel, D. Requirement of Homeobox Gene STIMPY/WOX9 for Arabidopsis Meristem Growth and Maintenance. Curr. Biol. 2005, 15, 436–440. [Google Scholar] [CrossRef]
- Zhang, Y.; Liu, Y.; Wang, X.; Wang, R.; Chen, X.; Wang, S.; Wei, H.; Wei, Z. PtrWOX13A Promotes Wood Formation and Bioactive Gibberellins Biosynthesis in Populus Trichocarpa. Front. Plant Sci. 2022, 13, 835035. [Google Scholar] [CrossRef]
- Kong, X.; Lu, S.; Tian, H.; Ding, Z. WOX5 Is Shining in the Root Stem Cell Niche. Trends Plant Sci. 2015, 20, 601–603. [Google Scholar] [CrossRef]
- Wang, L.-Q.; Li, Z.; Wen, S.-S.; Wang, J.-N.; Zhao, S.-T.; Lu, M.-Z. WUSCHEL-Related Homeobox Gene PagWOX11/12a Responds to Drought Stress by Enhancing Root Elongation and Biomass Growth in Poplar. J. Exp. Bot. 2020, 71, 1503–1513. [Google Scholar] [CrossRef] [PubMed]
- Rathour, M.; Sharma, A.; Kaur, A.; Upadhyay, S.K. Genome-Wide Characterization and Expression and Co-Expression Analysis Suggested Diverse Functions of WOX Genes in Bread Wheat. Heliyon 2020, 6, e05762. [Google Scholar] [CrossRef]
- Wang, Z.; Cai, Q.; Xia, H.; Han, B.; Li, M.; Wang, Y.; Zhu, M.; Jiao, C.; Wang, D.; Zhu, J.; et al. Genome-Wide Identification and Comparative Analysis of WOX Genes in Four Euphorbiaceae Species and Their Expression Patterns in Jatropha curcas. Front. Genet. 2022, 13, 878554. [Google Scholar] [CrossRef] [PubMed]
- Yang, T.; Gao, T.; Wang, C.; Wang, X.; Chen, C.; Tian, M.; Yang, W. In Silico Genome Wide Identification and Expression Analysis of the WUSCHEL-Related Homeobox Gene Family in Medicago sativa. Genom. Inf. 2022, 20, e19. [Google Scholar] [CrossRef]
- Zhang, X.; Zong, J.; Liu, J.; Yin, J.; Zhang, D. Genome-Wide Analysis of WOX Gene Family in Rice, Sorghum, Maize, Arabidopsis and Poplar. J. Integr. Plant Biol. 2010, 52, 1016–1026. [Google Scholar] [CrossRef]
- Gallois, J.-L.; Nora, F.R.; Mizukami, Y.; Sablowski, R. WUSCHEL Induces Shoot Stem Cell Activity and Developmental Plasticity in the Root Meristem. Genes Dev. 2004, 18, 375–380. [Google Scholar] [CrossRef] [PubMed]
- Kamiya, N.; Nagasaki, H.; Morikami, A.; Sato, Y.; Matsuoka, M. Isolation and Characterization of a Rice WUSCHEL-Type Homeobox Gene That Is Specifically Expressed in the Central Cells of a Quiescent Center in the Root Apical Meristem. Plant J. 2003, 35, 429–441. [Google Scholar] [CrossRef]
- Park, S.O.; Zheng, Z.; Oppenheimer, D.G.; Hauser, B.A. The PRETTY FEW SEEDS2 Gene Encodes an Arabidopsis Homeodomain Protein That Regulates Ovule Development. Development 2005, 132, 841–849. [Google Scholar] [CrossRef]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant 2020, 13, 1194–1202. [Google Scholar] [CrossRef]
- Marchler-Bauer, A.; Lu, S.; Anderson, J.B.; Chitsaz, F.; Derbyshire, M.K.; DeWeese-Scott, C.; Fong, J.H.; Geer, L.Y.; Geer, R.C.; Gonzales, N.R.; et al. CDD: A Conserved Domain Database for the Functional Annotation of Proteins. Nucleic Acids Res. 2011, 39, D225–D229. [Google Scholar] [CrossRef] [PubMed]
- Punta, M.; Coggill, P.C.; Eberhardt, R.Y.; Mistry, J.; Tate, J.; Boursnell, C.; Pang, N.; Forslund, K.; Ceric, G.; Clements, J.; et al. The Pfam Protein Families Database. Nucleic Acids Res. 2012, 40, D290–D301. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. 20 Years of the SMART Protein Domain Annotation Resource. Nucleic Acids Res. 2018, 46, D493–D496. [Google Scholar] [CrossRef]
- Cao, Y.; Han, Y.; Meng, D.; Li, G.; Li, D.; Abdullah, M.; Jin, Q.; Lin, Y.; Cai, Y. Genome-Wide Analysis Suggests the Relaxed Purifying Selection Affect the Evolution of WOX Genes in Pyrus bretschneideri, Prunus persica, Prunus mume, and Fragaria vesca. Front. Genet. 2017, 8, 78. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Che, Q.; Cheng, C.; Yuan, Y.; Wang, Y. Genome-Wide Identification of WOX Gene Family in Apple and a Functional Analysis of MdWOX4b during Adventitious Root Formation. J. Integr. Agric. 2022, 21, 1332–1345. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Waterhouse, A.M.; Procter, J.B.; Martin, D.M.A.; Clamp, M.; Barton, G.J. Jalview Version 2—A Multiple Sequence Alignment Editor and Analysis Workbench. Bioinformatics 2009, 25, 1189–1191. [Google Scholar] [CrossRef]
- Crooks, G.E.; Hon, G.; Chandonia, J.-M.; Brenner, S.E. WebLogo: A Sequence Logo Generator: Figure 1. Genome Res. 2004, 14, 1188–1190. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Bork, P. Interactive Tree of Life (ITOL) v5: An Online Tool for Phylogenetic Tree Display and Annotation. Nucleic Acids Res. 2021, 49, W293–W296. [Google Scholar] [CrossRef]
- Krzywinski, M.I.; Schein, J.E.; Birol, I.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: An Information Aesthetic for Comparative Genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef]
- Wang, Y.; Tang, H.; DeBarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.-H.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A Toolkit for Detection and Evolutionary Analysis of Gene Synteny and Collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef]
- Wang, D.; Zhang, Y.; Zhang, Z.; Zhu, J.; Yu, J. KaKs_Calculator 2.0: A Toolkit Incorporating Gamma-Series Methods and Sliding Window Strategies. Genom. Proteom. Bioinform. 2010, 8, 77–80. [Google Scholar] [CrossRef]
- Bailey, T.L.; Johnson, J.; Grant, C.E.; Noble, W.S. The MEME Suite. Nucleic Acids Res. 2015, 43, W39–W49. [Google Scholar] [CrossRef]
- Qiu, X.; Zhao, H.; Abubakar, A.S.; Shao, D.; Chen, J.; Chen, P.; Yu, C.; Wang, X.; Chen, K.; Zhu, A. Genome-Wide Analysis of AP2/ERF Gene Superfamily in Ramie (Boehmeria nivea L.) Revealed Their Synergistic Roles in Regulating Abiotic Stress Resistance and Ramet Development. Int. J. Mol. Sci. 2022, 23, 15117. [Google Scholar] [CrossRef] [PubMed]
- Feng, X.; Abubakar, A.S.; Chen, K.; Yu, C.; Zhu, A.; Chen, J.; Gao, G.; Wang, X.; Mou, P.; Chen, P. Genome-Wide Analysis of R2R3-MYB Transcription Factors in Boehmeria nivea (L.) Gaudich Revealed Potential Cadmium Tolerance and Anthocyanin Biosynthesis Genes. Front. Genet. 2023, 14, 1080909. [Google Scholar] [CrossRef]
- Li, B.; Dewey, C.N. RSEM: Accurate Transcript Quantification from RNA-Seq Data with or without a Reference Genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef]
- Sharma, H.; Sharma, A.; Rajput, R.; Sidhu, S.; Dhillon, H.; Verma, P.C.; Pandey, A.; Upadhyay, S.K. Molecular Characterization, Evolutionary Analysis, and Expression Profiling of BOR Genes in Important Cereals. Plants 2022, 11, 911. [Google Scholar] [CrossRef]
- Kaur, A.; Sharma, A.; Madhu; Dixit, S.; Singh, K.; Upadhyay, S.K. OSCA Genes in Bread Wheat: Molecular Characterization, Expression Profiling, and Interaction Analyses Indicated Their Diverse Roles during Development and Stress Response. Int. J. Mol. Sci. 2022, 23, 14867. [Google Scholar] [CrossRef] [PubMed]
- Ramkumar, T.R.; Kanchan, M.; Upadhyay, S.K.; Sembi, J.K. Identification and Characterization of WUSCHEL-Related Homeobox (WOX) Gene Family in Economically Important Orchid Species Phalaenopsis Equestris and Dendrobium Catenatum. Plant Gene 2018, 14, 37–45. [Google Scholar] [CrossRef]
- Gehring, W.J.; Müller, M.; Affolter, M.; Percival-Smith, A.; Billeter, M.; Qian, Y.Q.; Otting, G.; Wüthrich, K. The Structure of the Homeodomain and Its Functional Implications. Trends Genet. 1990, 6, 323–329. [Google Scholar] [CrossRef]
- Yang, Z.; Gong, Q.; Qin, W.; Yang, Z.; Cheng, Y.; Lu, L.; Ge, X.; Zhang, C.; Wu, Z.; Li, F. Genome-Wide Analysis of WOX Genes in Upland Cotton and Their Expression Pattern under Different Stresses. BMC Plant Biol. 2017, 17, 113. [Google Scholar] [CrossRef] [PubMed]
- Feng, X.; Abubakar, A.S.; Yu, C.; Zhu, A.; Chen, J.; Chen, K.; Gao, G.; Wang, X.; Mou, P.; Shao, D.; et al. Analysis of WRKY Resistance Gene Family in Boehmeria nivea (L.) Gaudich: Crosstalk Mechanisms of Secondary Cell Wall Thickening and Cadmium Stress. Front. Plant Sci. 2022, 13, 812988. [Google Scholar] [CrossRef]
- Abubakar, A.S.; Feng, X.; Gao, G.; Yu, C.; Chen, J.; Chen, K.; Wang, X.; Mou, P.; Shao, D.; Chen, P.; et al. Genome Wide Characterization of R2R3 MYB Transcription Factor from Apocynum Venetum Revealed Potential Stress Tolerance and Flavonoid Biosynthesis Genes. Genomics 2022, 114, 110275. [Google Scholar] [CrossRef] [PubMed]
- Lynch, M.; Conery, J.S. The Evolutionary Fate and Consequences of Duplicate Genes. Science 2000, 290, 1151–1155. [Google Scholar] [CrossRef] [PubMed]
- Chandler, J.; Nardmann, J.; Werr, W. Plant Development Revolves around Axes. Trends Plant Sci. 2008, 13, 78–84. [Google Scholar] [CrossRef] [PubMed]
- Abubakar, A.S.; Hamza, A.M.; Tadda, S.A.; Lado, A. Variability in growth characters and anti-nutritional properties of sickle pod (Senna obtusifolia L.). FUDMA J. Sci. 2018, 2, 12–16. [Google Scholar]
- Gilmartin, P.M.; Chua, N.H. Spacing between GT-1 Binding Sites within a Light-Responsive Element Is Critical for Transcriptional Activity. Plant Cell 1990, 2, 447–455. [Google Scholar] [CrossRef] [PubMed]
- Druege, U. Plant Hormone Homeostasis, Signaling, and Function during Adventitious Root Formation in Cuttings. Front. Plant Sci. 2016, 7, 381. [Google Scholar] [CrossRef]
- Xu, M.; Xie, W.; Huang, M. Two WUSCHEL-Related HOMEOBOX Genes, PeWOX11a and PeWOX11b, Are Involved in Adventitious Root Formation of Poplar. Physiol Plant. 2015, 155, 446–456. [Google Scholar] [CrossRef]
- Abarca, D.; Pizarro, A.; Hernández, I.; Sánchez, C.; Solana, S.P.; Carneros, E.; Díaz-Sala, C. The GRAS Gene Family in Pine: Transcript Expression Patterns Associated with the Maturation-Related Decline of Competence to Form Adventitious Roots. BMC Plant Biol. 2014, 14, 354. [Google Scholar] [CrossRef]
- Zhao, Y.; Hu, Y.; Dai, M.; Huang, L.; Zhou, D.-X. The WUSCHEL-Related Homeobox Gene WOX11 Is Required to Activate Shoot-Borne Crown Root Development in Rice. Plant Cell 2009, 21, 736–748. [Google Scholar] [CrossRef]
- Kyo, M.; Maida, K.; Nishioka, Y.; Matsui, K. Coexpression of WUSCHEL Related Homeobox (WOX) 2 with WOX8 or WOX9 Promotes Regeneration from Leaf Segments and Free Cells in Nicotiana Tabacum L. Plant Biotechnol. 2018, 35, 23–30. [Google Scholar] [CrossRef] [PubMed]
- Cho, S.; Yoo, S.; Zhang, H.; Pandeya, D.; Koh, H.; Hwang, J.; Kim, G.; Paek, N. The Rice Narrow Leaf2 and Narrow Leaf3 Loci Encode WUSCHEL-related Homeobox 3 A (O s WOX 3 A) and Function in Leaf, Spikelet, Tiller and Lateral Root Development. New Phytol. 2013, 198, 1071–1084. [Google Scholar] [CrossRef]
- Ishiwata, A.; Ozawa, M.; Nagasaki, H.; Kato, M.; Noda, Y.; Yamaguchi, T.; Nosaka, M.; Shimizu-Sato, S.; Nagasaki, A.; Maekawa, M.; et al. Two WUSCHEL-Related Homeobox Genes, Narrow Leaf2 and Narrow Leaf3, Control Leaf Width in Rice. Plant Cell Physiol. 2013, 54, 779–792. [Google Scholar] [CrossRef]
- Hu, X.; Xu, L. Transcription Factors WOX11/12 Directly Activate WOX5/7 to Promote Root Primordia Initiation and Organogenesis. Plant Physiol. 2016, 172, 2363–2373. [Google Scholar] [CrossRef]
- van der Graaff, E.; Laux, T.; Rensing, S.A. The WUS Homeobox-Containing (WOX) Protein Family. Genome Biol. 2009, 10, 248. [Google Scholar] [CrossRef] [PubMed]
- Malik, M.R.; Wang, F.; Dirpaul, J.M.; Zhou, N.; Polowick, P.L.; Ferrie, A.M.R.; Krochko, J.E. Transcript Profiling and Identification of Molecular Markers for Early Microspore Embryogenesis in Brassica napus. Plant Physiol. 2007, 144, 134–154. [Google Scholar] [CrossRef] [PubMed]
- Minh-Thu, P.-T.; Kim, J.S.; Chae, S.; Jun, K.M.; Lee, G.-S.; Kim, D.-E.; Cheong, J.-J.; Song, S.I.; Nahm, B.H.; Kim, Y.-K. A WUSCHEL Homeobox Transcription Factor, OsWOX13, Enhances Drought Tolerance and Triggers Early Flowering in Rice. Mol. Cells 2018, 41, 781–798. [Google Scholar] [CrossRef]
- Ikeda, M.; Mitsuda, N.; Ohme-Takagi, M. Arabidopsis WUSCHEL Is a Bifunctional Transcription Factor That Acts as a Repressor in Stem Cell Regulation and as an Activator in Floral Patterning. Plant Cell 2009, 21, 3493–3505. [Google Scholar] [CrossRef] [PubMed]
- Prince, V.E.; Pickett, F.B. Splitting Pairs: The Diverging Fates of Duplicated Genes. Nat. Rev. Genet. 2002, 3, 827–837. [Google Scholar] [CrossRef]
- Denis, E.; Kbiri, N.; Mary, V.; Claisse, G.; Conde e Silva, N.; Kreis, M.; Deveaux, Y. WOX14 Promotes Bioactive Gibberellin Synthesis and Vascular Cell Differentiation in Arabidopsis. Plant J. 2017, 90, 560–572. [Google Scholar] [CrossRef]
- Garg, T.; Singh, Z.; Chennakesavulu, K.; Mushahary, K.K.K.; Dwivedi, A.K.; Varapparambathu, V.; Singh, H.; Singh, R.S.; Sircar, D.; Chandran, D.; et al. Species-Specific Function of Conserved Regulators in Orchestrating Rice Root Architecture. Development 2022, 149, dev.200381. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez, L.; Mongelard, G.; Floková, K.; Păcurar, D.I.; Novák, O.; Staswick, P.; Kowalczyk, M.; Păcurar, M.; Demailly, H.; Geiss, G.; et al. Auxin Controls Arabidopsis Adventitious Root Initiation by Regulating Jasmonic Acid Homeostasis. Plant Cell 2012, 24, 2515–2527. [Google Scholar] [CrossRef]
- Wu, C.-C.; Li, F.-W.; Kramer, E.M. Large-Scale Phylogenomic Analysis Suggests Three Ancient Superclades of the WUSCHEL-RELATED HOMEOBOX Transcription Factor Family in Plants. PLoS ONE 2019, 14, e0223521. [Google Scholar] [CrossRef] [PubMed]
- Lischweski, S. Jasmonates Act Positively in Adventitious Root Formation in Petunia Cuttings. BMC Plant Biol. 2015, 15, 229. [Google Scholar] [CrossRef] [PubMed]
- Rasmussen, A.; Hosseini, S.A.; Hajirezaei, M.-R.; Druege, U.; Geelen, D. Adventitious Rooting Declines with the Vegetative to Reproductive Switch and Involves a Changed Auxin Homeostasis. J. Exp. Bot. 2015, 66, 1437–1452. [Google Scholar] [CrossRef]

| Gene Name | Gene ID | Chromosome Position | No. A. Acids | M.wt. (Da) | pI | GRAVY | Subcellular Location |
|---|---|---|---|---|---|---|---|
| BnWOX1A | Maker00062999 | Chr10:2590680-2593026 | 332 | 37,374 | 8.16 | −0.956 | Nuclear |
| BnWOX1B | Maker00074103 | Chr:2485054-2487384 | 309 | 34,964 | 8.85 | −1.032 | Nuclear |
| BnWOX2 | Maker00030742 | Chr3: 7583477-7584960 | 307 | 34,320 | 6.70 | −0.825 | Nuclear |
| BnWOX3 | Maker00031089 | Chr3: 8055078-8056503 | 274 | 31,317 | 8.42 | −0.942 | Nuclear |
| BnWOX4 | Maker00002967 | Chr13: 2041939-2044060 | 264 | 29,331 | 9.54 | −0.826 | Nuclear |
| BnWOX5 | Maker00093460 | Chr1:16210128-16210838 | 200 | 22,959 | 8.56 | −0.714 | Nuclear |
| BnWOX6A | Maker00053871 | Chr7: 3319544-3322227 | 254 | 28,885 | 8.58 | −0.525 | Nuclear |
| BnWOX6B | Maker00021009 | Chr7: 1475052-1476549 | 214 | 24,453 | 9.80 | −0.658 | Nuclear |
| BnWOX7 | Maker00093478 | Chr1: 16182683-16183394 | 200 | 22,984 | 8.78 | −0.746 | Nuclear |
| BnWOX9A | Maker00080289 | Chr2: 4246373-4248777 | 386 | 42,823 | 7.15 | −0.746 | Nuclear |
| BnWOX9B | Maker00047861 | Chr2: 7182597-7185242 | 416 | 45,763 | 7.81 | −0.595 | Nuclear |
| BnWOX10 | Maker00082428 | Chr13: 9054047-906063 | 856 | 93,650 | 5.80 | −0.115 | Nuclear/Plasma membrane |
| BnWOX11 | Maker00086436 | Chr14: 2642861-2645770 | 294 | 32,603 | 5.67 | −0.452 | Nuclear |
| BnWOX13A | Maker00063698 | Contig314_1: 496112-497997 | 233 | 26,515 | 4.91 | −0.819 | Nuclear |
| BnWOX13B | Maker00008406 | Chr12: 12559425-12571929 | 320 | 36,059 | 5.56 | −0.953 | Nuclear |
| BnWOX14 | Maker00091405 | Chr2: 18352039-18359653 | 837 | 91,817 | 6.03 | −0.119 | Nuclear/Plasma membrane |
| BnWUS | Maker00008321 | Chr12: 11915820-11916191 | 123 | 14,384 | 10.00 | −1.395 | Nuclear |
| Germination of Root Primordium | Development of Adventitious Roots | ||
|---|---|---|---|
| Average Number of Primordium after 3 Days | Average Number of Primordium after 5 Days | Average Length (cm) after 15 Days | |
| ZZ No. 2 | 3 ± 0.83 b | 12 ± 1.48 a | 8.56 ± 1.09 a |
| ZZ No. 1 | 1 ± 0.45 c | 10 ± 1.22 b | 6.22 ± 1.13 b |
| ZSZ No. 1 | 5 ± 0.98 a | 7 ± 1.22 c | 4.74 ± 0.55 c |
| SC-3D | 3 ± 0.00 b | 7 ± 1.96 c | 4.25 ± 0.54 c |
| HZ | 1 ± 0.00 c | 4 ± 0.45 d | 3.98 ± 0.44 c |
| XZ-21D | 0 | 2 ± 0.00 f | 0.33 ± 0.01 d |
| HB-3 | 0 | 2 ± 0.44 f | 0.33 ± 0.01 d |
| JX-2 | 0 | 3 ± 0.44 e | 0.32 ± 0.01 d |
| ZZ No. 3 | 0 | 3 ± 0.00 e | 0.30 ± 0.01 d |
| GX-1 | 0 | 3 ± 1.09 e | 0.30 ± 0.01 d |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abubakar, A.S.; Wu, Y.; Chen, F.; Zhu, A.; Chen, P.; Chen, K.; Qiu, X.; Huang, X.; Zhao, H.; Chen, J.; et al. Comprehensive Analysis of WUSCEL-Related Homeobox Gene Family in Ramie (Boehmeria nivea) Indicates Its Potential Role in Adventitious Root Development. Biology 2023, 12, 1475. https://doi.org/10.3390/biology12121475
Abubakar AS, Wu Y, Chen F, Zhu A, Chen P, Chen K, Qiu X, Huang X, Zhao H, Chen J, et al. Comprehensive Analysis of WUSCEL-Related Homeobox Gene Family in Ramie (Boehmeria nivea) Indicates Its Potential Role in Adventitious Root Development. Biology. 2023; 12(12):1475. https://doi.org/10.3390/biology12121475
Chicago/Turabian StyleAbubakar, Aminu Shehu, Yongmei Wu, Fengming Chen, Aiguo Zhu, Ping Chen, Kunmei Chen, Xiaojun Qiu, Xiaoyu Huang, Haohan Zhao, Jikang Chen, and et al. 2023. "Comprehensive Analysis of WUSCEL-Related Homeobox Gene Family in Ramie (Boehmeria nivea) Indicates Its Potential Role in Adventitious Root Development" Biology 12, no. 12: 1475. https://doi.org/10.3390/biology12121475
APA StyleAbubakar, A. S., Wu, Y., Chen, F., Zhu, A., Chen, P., Chen, K., Qiu, X., Huang, X., Zhao, H., Chen, J., & Gao, G. (2023). Comprehensive Analysis of WUSCEL-Related Homeobox Gene Family in Ramie (Boehmeria nivea) Indicates Its Potential Role in Adventitious Root Development. Biology, 12(12), 1475. https://doi.org/10.3390/biology12121475









