Microbial Involvement in Carbon Transformation via CH4 and CO2 in Saline Sedimentary Pool
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
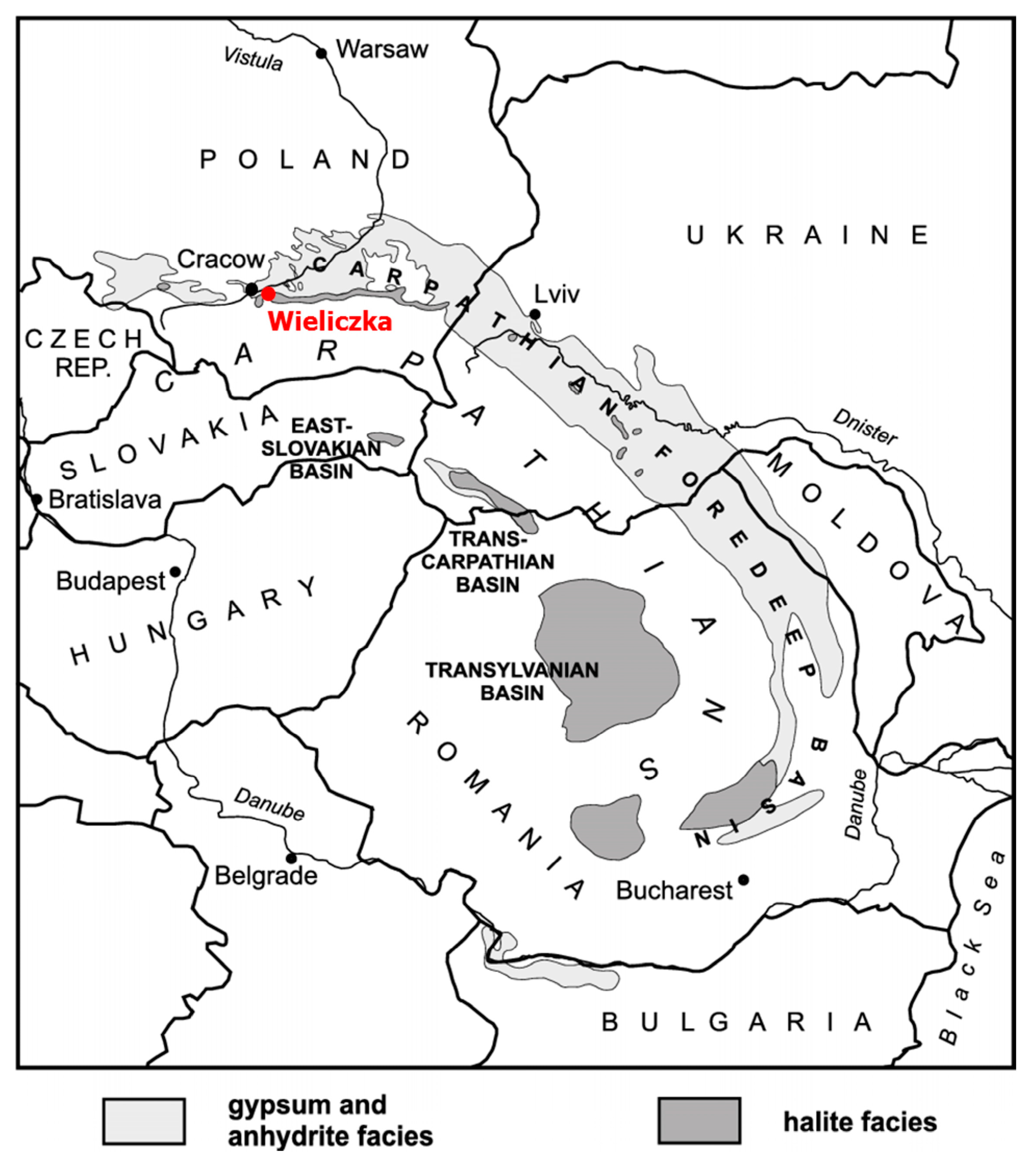
2.1. Study Site
2.2. Sample Collection and Processing
2.3. Isotopic Analysis
2.4. Present-Day Methane and Carbon Dioxide Turnover
2.5. Microbial Diversity and Molecular Techniques
2.6. Data Analysis
3. Results
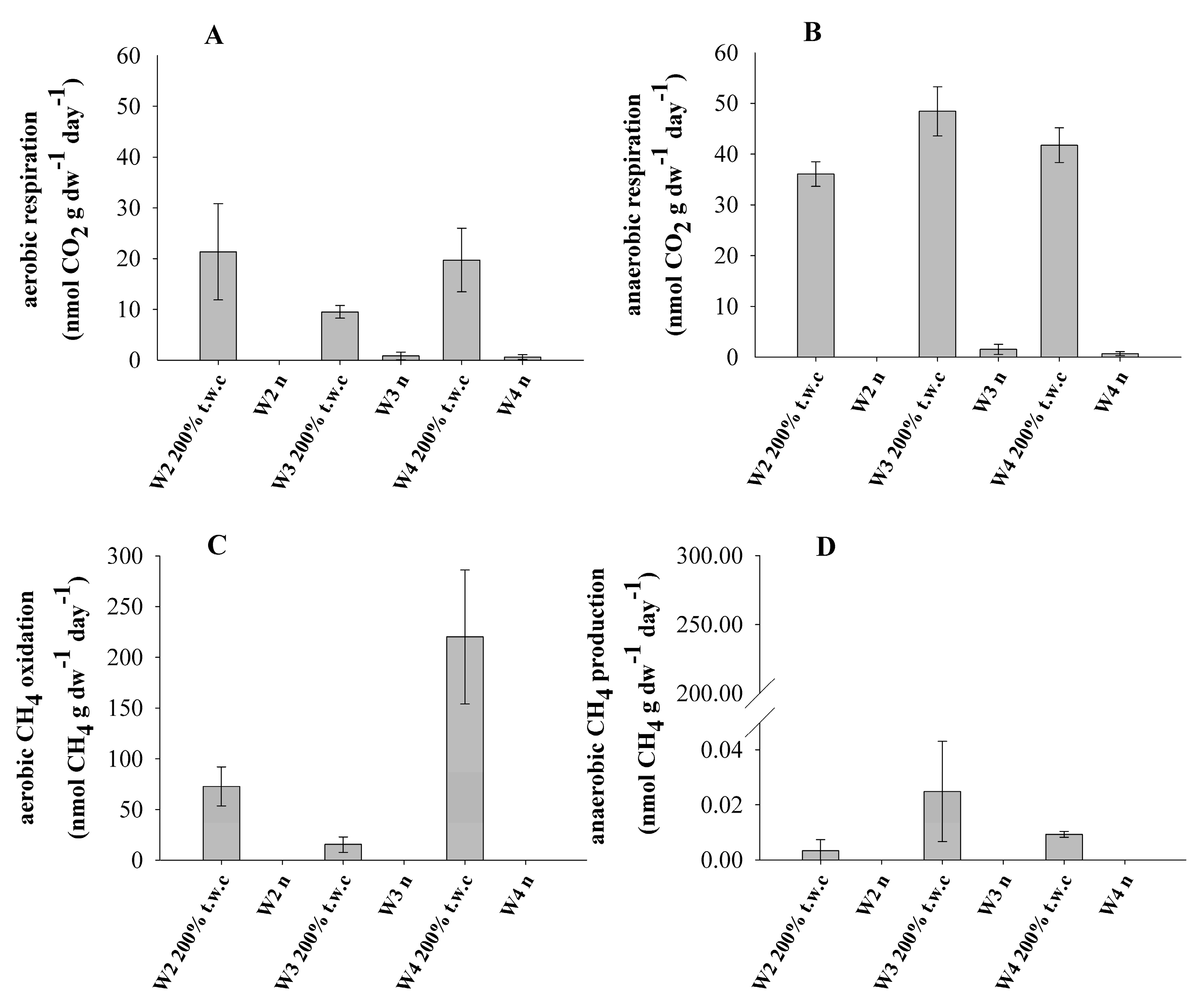
3.1. Methane Turnover and Respiration Rates
3.2. Microbial Identification
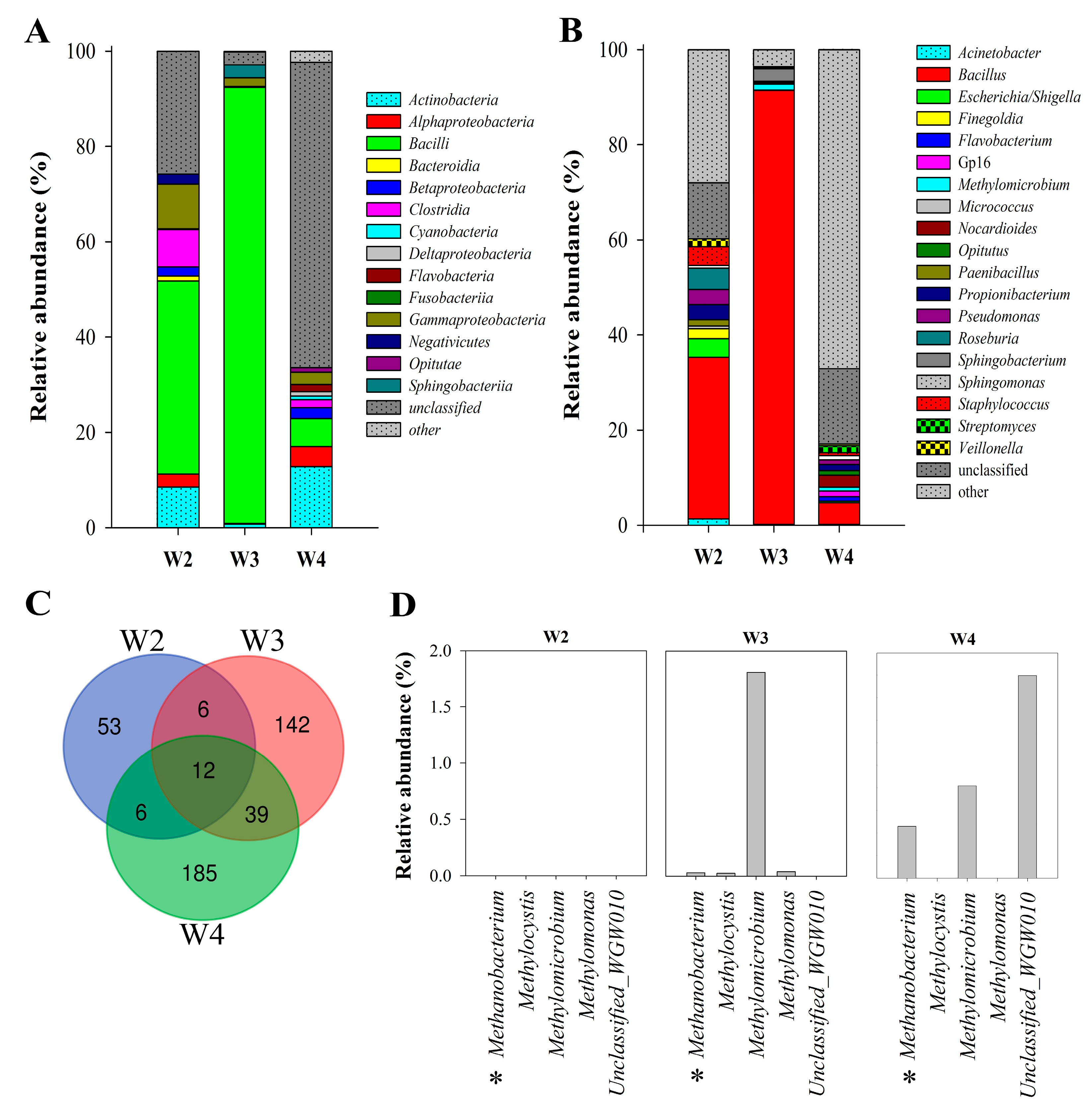
3.2.1. Biodiversity
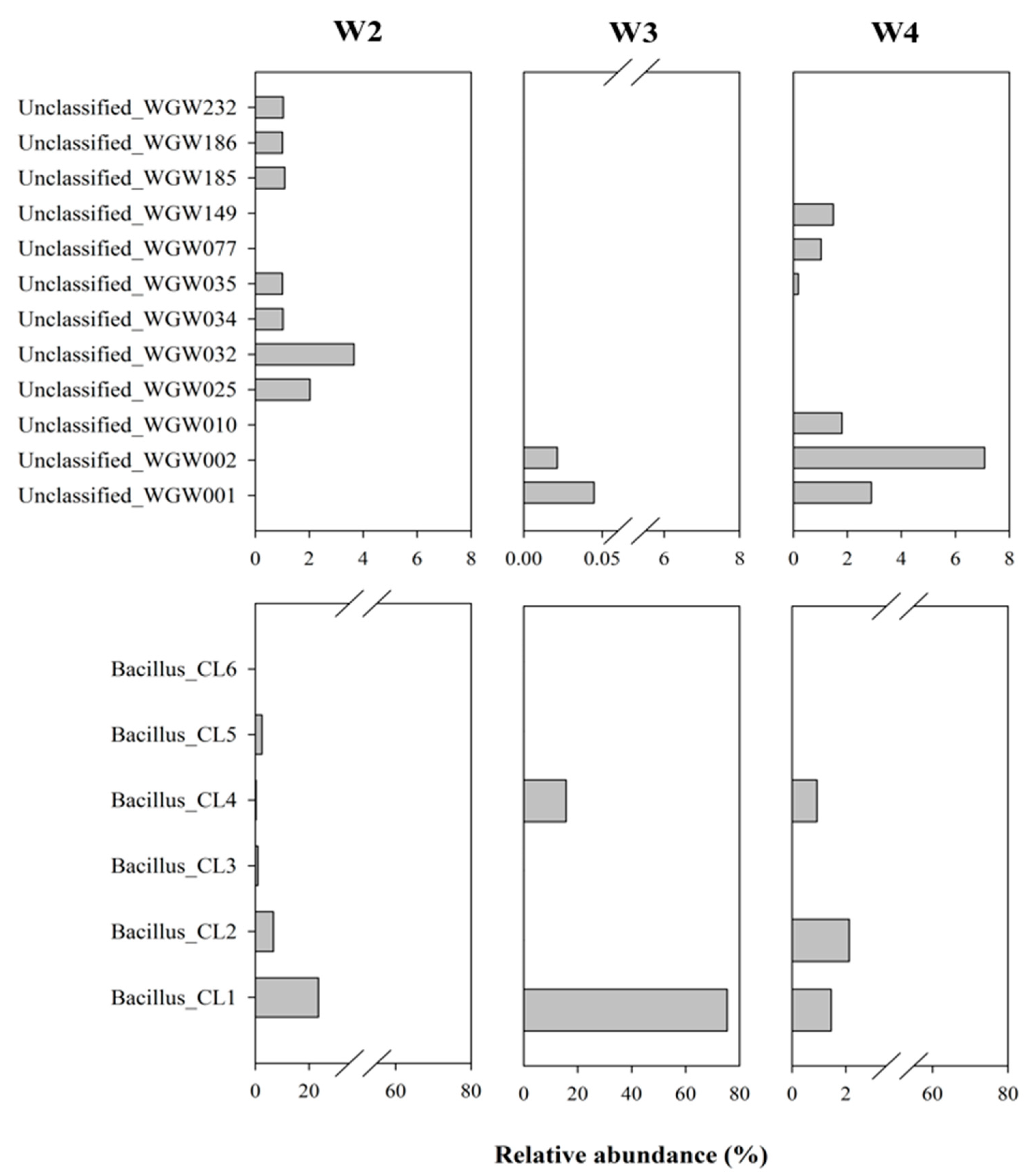
3.2.2. Microbial Community Structure
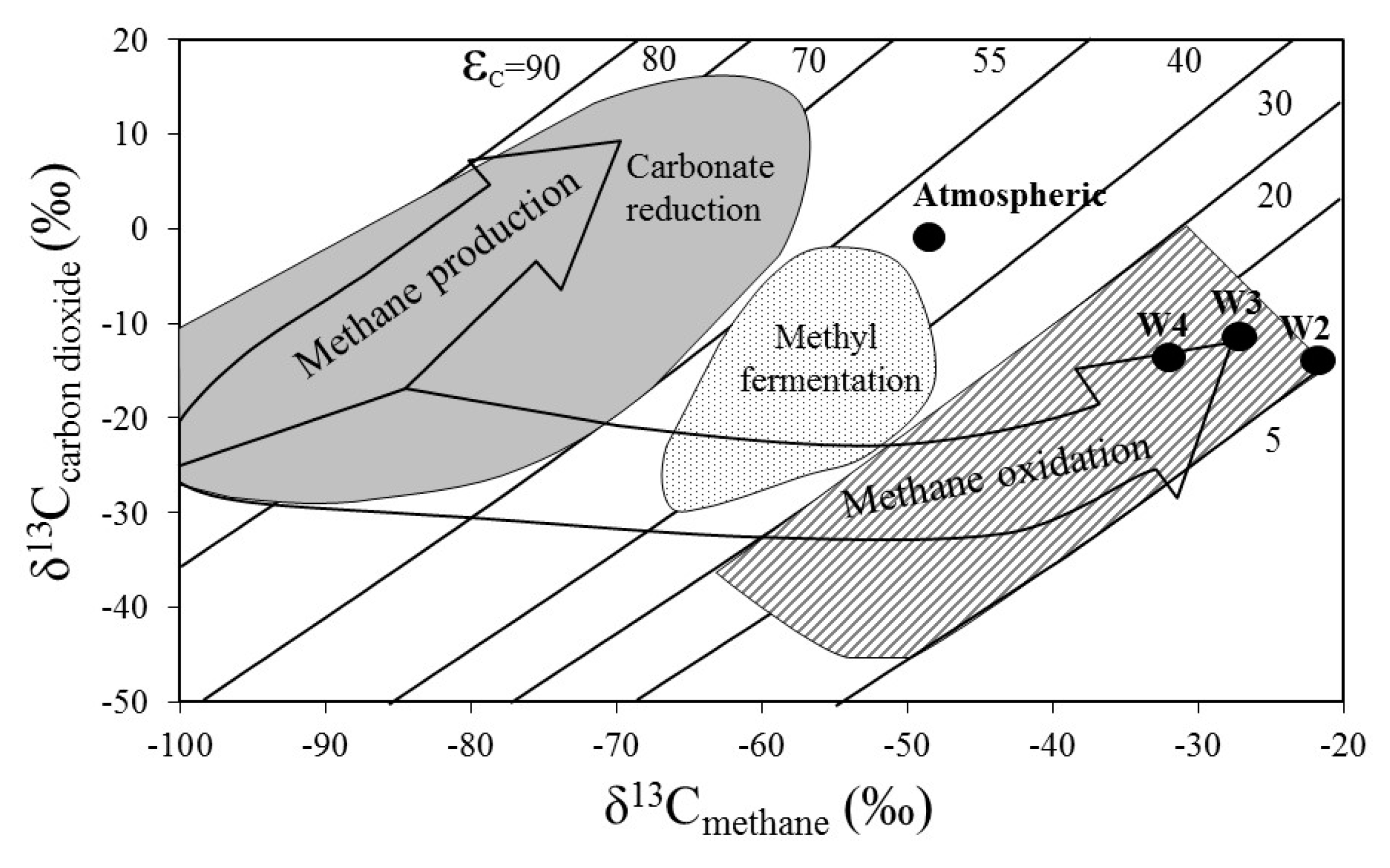
3.3. Stable Methane and Carbon Dioxide Isotopes
4. Discussion
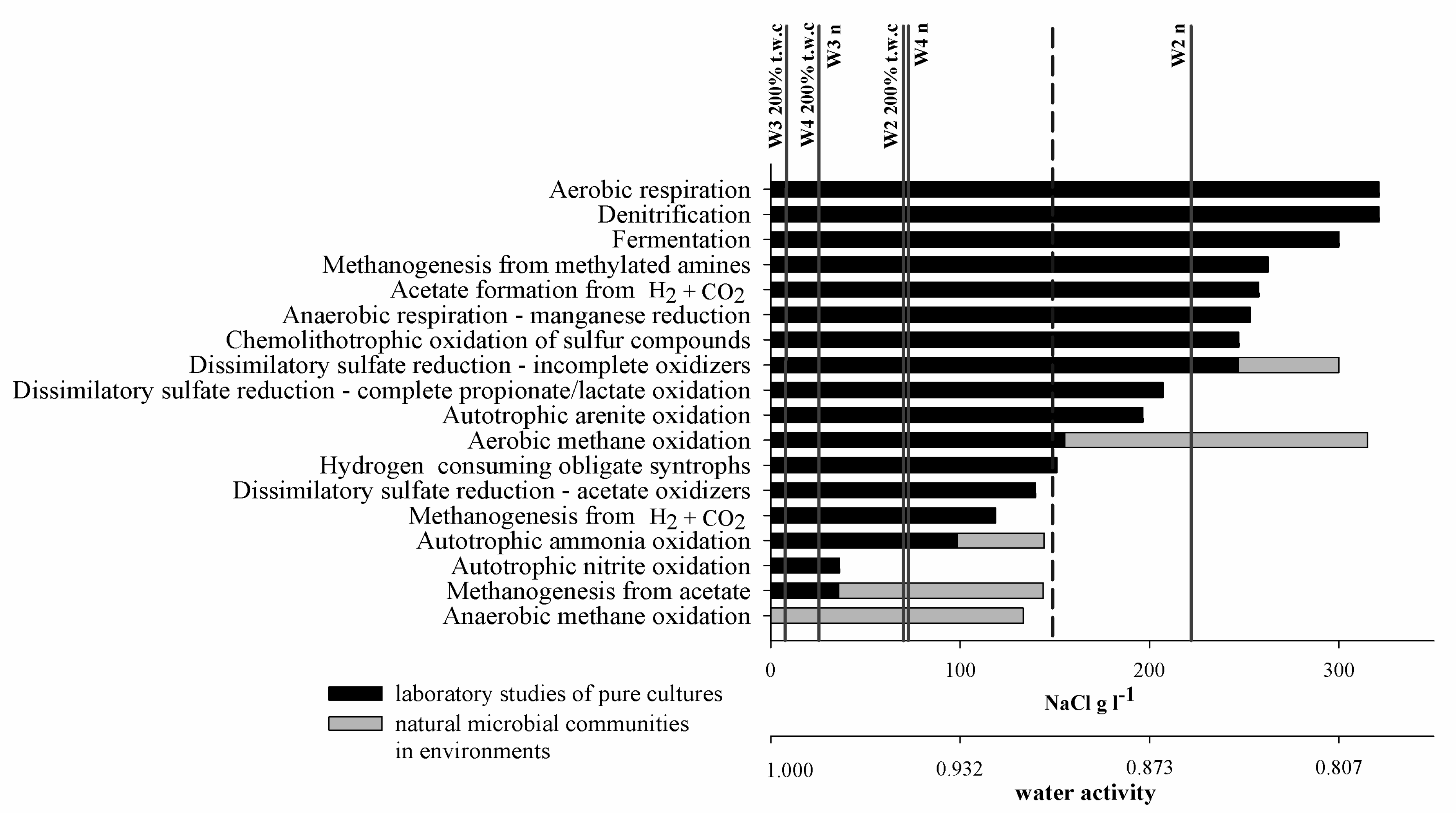
4.1. The Role of Salinity in Methane Metabolism
4.2. Microbial Respiration in Saline Sedimentary Pool
4.3. Microbial Diversity
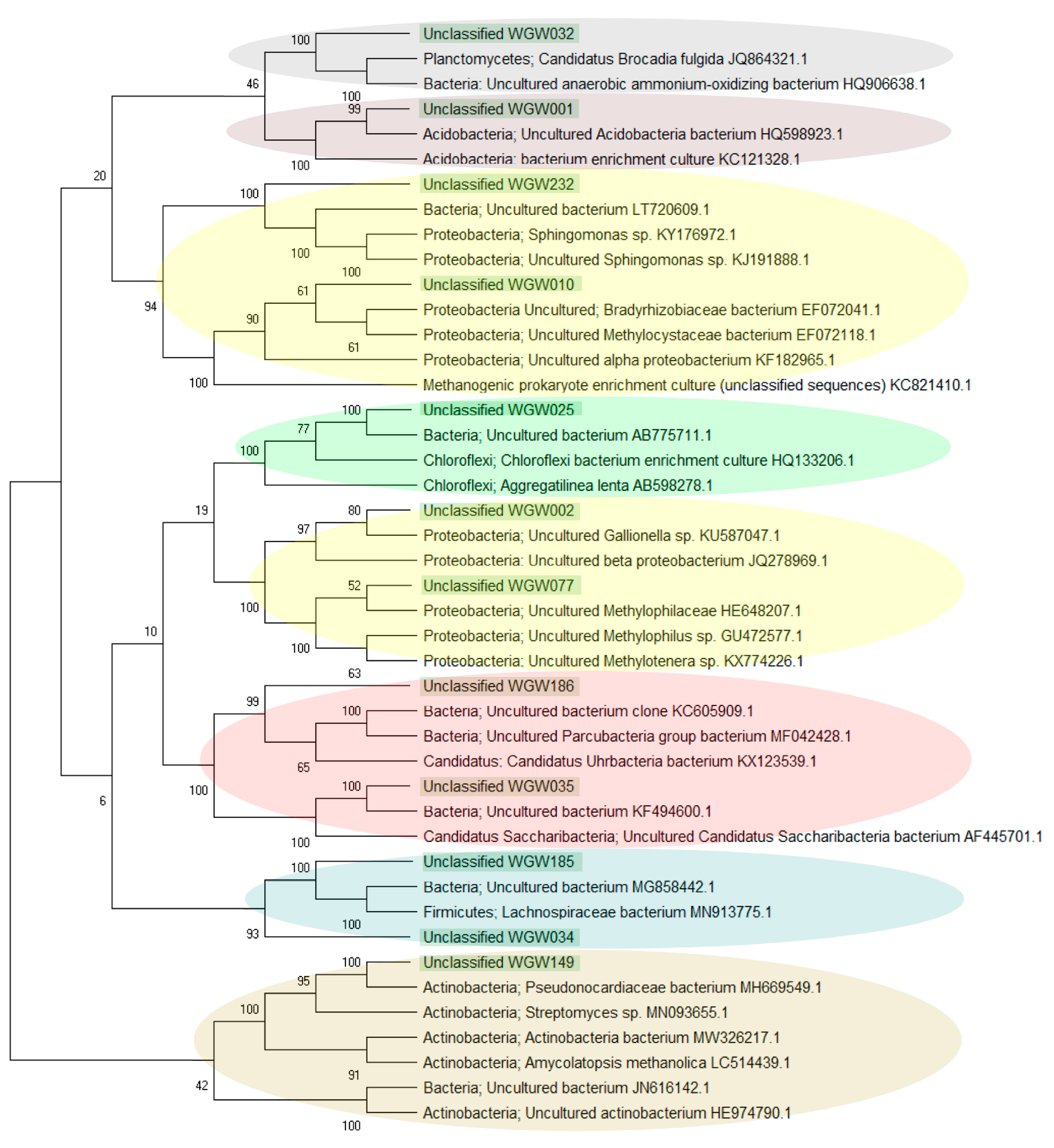
4.3.1. Prevalence of Unclassified Bacteria in Saline Sedimentary Pool
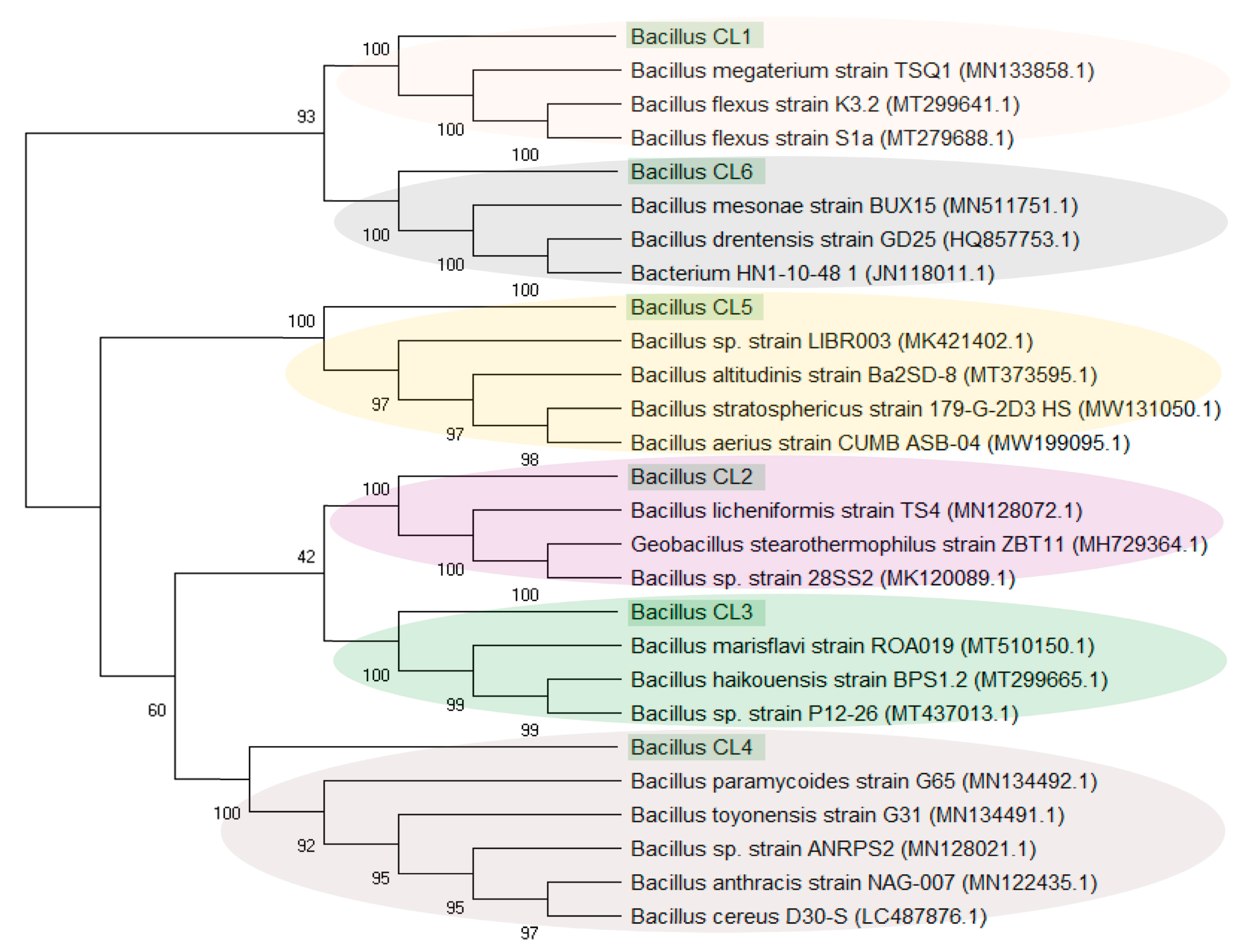
4.3.2. Ubiquity of Bacillus-like Bacteria
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Olivier, J.; Peters, J. Trends in Global CO2 and Total Greenhouse Gas Emissions; 2019 Report; PBL Netherlands Environmental Assessment Agency: The Hague, The Netherlands, 2020. [Google Scholar]
- Dunn, R.J.H.; Stanitski, D.M.; Gobron, N.; Willett, K.M. Global climate. Bull. Am. Meteorol. Soc. 2020, 101, S9–S128. [Google Scholar] [CrossRef]
- Dlugokencky, E. Global Monitoring Laboratory—Carbon Cycle Greenhouse Gases. Available online: https://www.esrl.noaa.gov/gmd/ccgg/trends_ch4/ (accessed on 21 March 2021).
- Etminan, M.; Myhre, G.; Highwood, E.J.; Shine, K.P. Radiative forcing of carbon dioxide, methane, and nitrous oxide: A significant revision of the methane radiative forcing. Geophys. Res. Lett. 2016, 43, 12614–12623. [Google Scholar] [CrossRef]
- Hoegh-Guldberg, O.; Jacob, D.; Bindi, M.; Brown, S.; Camilloni, I.; Diedhiou, A.; Djalante, R.; Ebi, K.; Engelbrecht, F.; Guiot, J.; et al. Chapter 3: Impacts of 1.5oC Global Warming on Natural and Human Systems; IPCC Secretariat: Geneva, Switzerland, 2018; pp. 175–311. [Google Scholar]
- Walkiewicz, A.; Brzezińska, M.; Wnuk, E.; Jabłoński, B. Soil properties and not high CO2 affect CH4 production and uptake in periodically waterlogged arable soils. J. Soils Sediments 2020, 20, 1231–1240. [Google Scholar] [CrossRef]
- Stępniewska, Z.; Goraj, W.; Sochaczewska, A.; Kuźniar, A.; Pytlak, A.; Malec, M. Changes in atmospheric CH4, O3, NO2, SO2 concentration dynamics in Lublin in years 2007–2009. Acta Agrophysica 2014, 21, 361–373. [Google Scholar]
- Anantharaman, K.; Brown, C.T.; Hug, L.A.; Sharon, I.; Castelle, C.J.; Probst, A.J.; Thomas, B.C.; Singh, A.; Wilkins, M.J.; Karaoz, U.; et al. Thousands of microbial genomes shed light on interconnected biogeochemical processes in an aquifer system. Nat. Commun. 2016, 7, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Fernandez, M.; Simone, D.; Wu, X.; Soler, L.; Nilsson, E.; Holmfeldt, K.; Lantz, H.; Bertilsson, S.; Dopson, M. Metatranscriptomes reveal that all three domains of life are active but are dominated by bacteria in the fennoscandian crystalline granitic continental deep biosphere. MBio 2018, 9, e01792-18. [Google Scholar] [CrossRef] [PubMed]
- Dam, H.T.; Vollmers, J.; Sobol, M.S.; Cabezas, A.; Kaster, A.K. Targeted cell sorting combined with single cell genomics captures low abundant microbial dark matter with higher sensitivity than metagenomics. Front. Microbiol. 2020, 11, 1377. [Google Scholar] [CrossRef]
- Zhou, Z.; Liu, Y.; Pan, J.; Cron, B.R.; Toner, B.M.; Anantharaman, K.; Breier, J.A.; Dick, G.J.; Li, M. Gammaproteobacteria mediating utilization of methyl-, sulfur- and petroleum organic compounds in deep ocean hydrothermal plumes. ISME J. 2020, 14, 3136–3148. [Google Scholar] [CrossRef]
- Inagaki, F.; Hinrichs, K.U.; Kubo, Y.; Bowles, M.W.; Heuer, V.B.; Hong, W.L.; Hoshino, T.; Ijiri, A.; Imachi, H.; Ito, M.; et al. Exploring deep microbial life in coal-bearing sediment down to ∼2.5 km below the ocean floor. Science 2015, 349, 420–424. [Google Scholar] [CrossRef]
- Pytlak, A.; Stepniewska, Z.; Kuźniar, A.; Szafranek-Nakonieczna, A.; Wolińska, A.; Banach, A. Potential for aerobic methane oxidation in carboniferous coal measures. Geomicrobiol. J. 2014, 31, 737–747. [Google Scholar] [CrossRef]
- Thomas, C.; Grossi, V.; Antheaume, I.; Ariztegui, D. Recycling of archaeal biomass as a new strategy for extreme life in Dead Sea deep sediments. Geology 2019, 47, 479–482. [Google Scholar] [CrossRef]
- Trembath-Reichert, E.; Morono, Y.; Ijiri, A.; Hoshino, T.; Dawson, K.S.; Inagaki, F.; Orphan, V.J. Methyl-compound use and slow growth characterize microbial life in 2-km-deep subseafloor coal and shale beds. Proc. Natl. Acad. Sci. USA 2017, 114, E9206–E9215. [Google Scholar] [CrossRef]
- Donadei, S.; Schneider, G.S. Compressed air energy storage in underground formations. In Storing Energy: With Special Reference to Renewable Energy Sources; Elsevier Inc.: Hannover, Germany, 2016; pp. 113–133. ISBN 9780128034408. [Google Scholar]
- Kietäväinen, R.; Purkamo, L. The origin, source, and cycling of methane in deep crystalline rock biosphere. Front. Microbiol. 2015, 6, 725. [Google Scholar] [CrossRef] [PubMed]
- Drake, H.; Heim, C.; Roberts, N.M.W.; Zack, T.; Tillberg, M.; Broman, C.; Ivarsson, M.; Whitehouse, M.J.; Åström, M.E. Isotopic evidence for microbial production and consumption of methane in the upper continental crust throughout the Phanerozoic eon. Earth Planet. Sci. Lett. 2017, 470, 108–118. [Google Scholar] [CrossRef]
- Stępniewska, Z.; Goraj, W.; Wolińska, A.; Szafranek-Nakonieczna, A.; Banach, A.; Górski, A. Methanotrophic activity of rocks surrounding Badenian salts in the “Wieliczka” Salt Mine. Carpathian J. Earth Environ. Sci. 2018, 13, 107–119. [Google Scholar] [CrossRef]
- Mills, C.T.; Amano, Y.; Slater, G.F.; Dias, R.F.; Iwatsuki, T.; Mandernack, K.W. Microbial carbon cycling in oligotrophic regional aquifers near the Tono Uranium Mine, Japan as inferred from δ13C and Δ14C values of in situ phospholipid fatty acids and carbon sources. Geochim. Cosmochim. Acta 2010, 74, 3785–3805. [Google Scholar] [CrossRef]
- Brudnik, K.; Czop, M.; Motyka, J.; D’Obyrn, K.; Rogoż, M.; Witczak, S. The complex hydrogeology of the unique Wieliczka salt mine. Przegląd Geol. 2010, 58, 787–796. [Google Scholar]
- Hohenegger, J.; Ćorićstjepan, S.; Wagreich, M. Timing of the middle miocene badenian stage of the central paratethys. Geol. Carpathica 2014, 65, 55–66. [Google Scholar] [CrossRef]
- De Leeuw, A.; Bukowski, K.; Krijgsman, W.; Kuiper, K.F. Age of the Badenian salinity crisis; Impact of Miocene climate variability on the circum-mediterranean region. Geology 2010, 38, 715–718. [Google Scholar] [CrossRef]
- Peryt, T.M. The beginning, development and termination of the Middle Miocene Badenian salinity crisis in Central Paratethys. Sediment. Geol. 2006, 188, 379–396. [Google Scholar] [CrossRef]
- Garlicki, A. Salt mines at Bochnia and Wieliczka. Przegląd Geol. 2008, 56, 663–669. [Google Scholar]
- Wachowiak, J.; Pawlikowski, M.; Wilkosz, P. Lithostratigraphy of Zechstein evaporites of the central and north-western parts of the Mogilno Salt Diapir, based on boreholes Z-9 and Z-17. Geol. Geophys. Environ. 2012, 38, 115–151. [Google Scholar] [CrossRef][Green Version]
- Babel, M.; Bogucki, A. The Badenian evaporite basin of the northern Carpathian Foredeep as a model of a meromictic selenite basin. Geol. Soc. Lond. Spec. Publ. 2007, 285, 219–246. [Google Scholar] [CrossRef]
- Pokorný, R.; Olejníková, P.; Balog, M.; Zifčák, P.; Hölker, U.; Janssen, M.; Bend, J.; Höfer, M.; Holienčin, R.; Hudecová, D.; et al. Characterization of microorganisms isolated from lignite excavated from the Záhorie coal mine (southwestern Slovakia). Res. Microbiol. 2005, 156, 932–943. [Google Scholar] [CrossRef]
- Szafranek-Nakonieczna, A.; Zheng, Y.; Słowakiewicz, M.; Pytlak, A.; Polakowski, C.; Kubaczyński, A.; Bieganowski, A.; Banach, A.; Wolińska, A.; Stępniewska, Z. Methanogenic potential of lignites in Poland. Int. J. Coal Geol. 2018, 196, 201–210. [Google Scholar] [CrossRef]
- Wolińska, A.; Pytlak, A.; Stepniewska, Z.; Kuźniar, A.; Piasecki, C. Identification of methanotrophic bacteria community in the Jastrzebie-Moszczenica coal mine by fluorescence in situ hybridization and PCR techniques. Pol. J. Environ. Stud. 2013, 22, 275–282. [Google Scholar]
- Szafranek-Nakonieczna, A.; Pytlak, A.; Grządziel, J.; Kubaczyński, A.; Banach, A.; Górski, A.; Goraj, W.; Kuzńiar, A.; Gałązka, A.; Stępniewska, Z. Changes in the substrate source reveal novel interactions in the sediment-derived methanogenic microbial community. Int. J. Mol. Sci. 2019, 20, 4415. [Google Scholar] [CrossRef]
- Klindworth, A.; Pruesse, E.; Schweer, T.; Peplies, J.; Quast, C.; Horn, M.; Glöckner, F.O. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 2013, 41, e1. [Google Scholar] [CrossRef]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing 2016; R Foundation for Statistical Computing: Vienna, Austria, 2013. [Google Scholar]
- Wright, E.S. RDP v16 Modified Training Set for 16S rRNA Classification; 2019. Available online: http://www2.decipher.codes/Classification/TrainingSets/RDP_v16-mod_March2018.RData (accessed on 9 March 2021).
- Murali, A.; Bhargava, A.; Wright, E.S. IDTAXA: A novel approach for accurate taxonomic classification of microbiome sequences. Microbiome 2018, 6, 1–14. [Google Scholar] [CrossRef] [PubMed]
- McMurdie, P.J.; Holmes, S. Phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 2013, 8, e61217. [Google Scholar] [CrossRef] [PubMed]
- Rognes, T.; Flouri, T.; Nichols, B.; Quince, C.; Mahé, F. VSEARCH: A versatile open source tool for metagenomics. PeerJ 2016, 4, e2584. [Google Scholar] [CrossRef] [PubMed]
- Větrovský, T.; Baldrian, P.; Morais, D. SEED 2: A user-friendly platform for amplicon high-throughput sequencing data analyses. Bioinformatics 2018, 34, 2292–2294. [Google Scholar] [CrossRef] [PubMed]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [CrossRef]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef] [PubMed]
- Nei, M.; Kumar, S. Molecular Evolution and Phylogenetics; Oxford University Press: New York, NY, USA, 2020. [Google Scholar]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547. [Google Scholar] [CrossRef]
- Whiticar, M.J. Carbon and hydrogen isotope systematics of bacterial formation and oxidation of methane. Chem. Geol. 1999, 161, 291–314. [Google Scholar] [CrossRef]
- Whiticar, M.J. Stable isotope geochemistry of coals, humic kerogens and related natural gases. Int. J. Coal Geol. 1996, 32, 191–215. [Google Scholar] [CrossRef]
- Manzoni, S.; Schimel, J.P.; Porporato, A. Responses of soil microbial communities to water stress: Results from a meta-analysis. Ecology 2012, 93, 930–938. [Google Scholar] [CrossRef]
- Moyano, F.E.; Manzoni, S.; Chenu, C. Responses of soil heterotrophic respiration to moisture availability: An exploration of processes and models. Soil Biol. Biochem. 2013, 59, 72–85. [Google Scholar] [CrossRef]
- Hallsworth, J.E.; Yakimov, M.M.; Golyshin, P.N.; Gillion, J.L.M.; D’Auria, G.; De Lima Alves, F.; La Cono, V.; Genovese, M.; McKew, B.A.; Hayes, S.L.; et al. Limits of life in MgCl2-containing environments: Chaotropicity defines the window. Environ. Microbiol. 2007, 9, 801–813. [Google Scholar] [CrossRef] [PubMed]
- Stevenson, A.; Burkhardt, J.; Cockell, C.S.; Cray, J.A.; Dijksterhuis, J.; Fox-Powell, M.; Kee, T.P.; Kminek, G.; Mcgenity, T.J.; Timmis, K.N.; et al. Multiplication of microbes below 0.690 water activity: Implications for terrestrial and extraterrestrial life. Environ. Microbiol. 2015, 17, 257–277. [Google Scholar] [CrossRef]
- Vergara-Fernández, A.; Morales, P.; Scott, F.; Guerrero, S.; Yañez, L.; Mau, S.; Aroca, G. Methane biodegradation and enhanced methane solubilization by the filamentous fungi Fusarium solani. Chemosphere 2019, 226, 24–35. [Google Scholar] [CrossRef]
- Oren, A. Diversity of halophiles. In Extremophiles Handbook; Springer: Tokyo, Japan, 2011; pp. 309–325. [Google Scholar]
- Oren, A. Life at high salt concentrations, intracellular KCl concentrations, and acidic proteomes. Front. Microbiol. 2013, 4, 315. [Google Scholar] [CrossRef]
- Oren, A. The bioenergetic basis for the decrease in metabolic diversity at increasing salt concentrations: Implications for the functioning of salt lake ecosystems. In Saline Lakes. Developments in Hydrobiology; Melack, J.M., Jellison, R., Herbst, D.B., Eds.; Springer: Dordrecht, the Netherlands, 2001; Volume 162, pp. 61–72. [Google Scholar]
- Zhuang, G.C.; Elling, F.J.; Nigro, L.M.; Samarkin, V.; Joye, S.B.; Teske, A.; Hinrichs, K.U. Multiple evidence for methylotrophic methanogenesis as the dominant methanogenic pathway in hypersaline sediments from the Orca Basin, Gulf of Mexico. Geochim. Cosmochim. Acta 2016, 187, 1–20. [Google Scholar] [CrossRef]
- Lazar, C.S.; Parkes, R.J.; Cragg, B.A.; L’Haridon, S.; Toffin, L. Methanogenic diversity and activity in hypersaline sediments of the centre of the Napoli mud volcano, Eastern Mediterranean Sea. Environ. Microbiol. 2011, 13, 2078–2091. [Google Scholar] [CrossRef]
- Magnabosco, C.; Lin, L.-H.; Dong, H.; Bomberg, M.; Ghiorse, W.; Stan-Lotter, H.; Pedersen, K.; Kieft, T.L.; van Heerden, E.; Onstott, T.C. The biomass and biodiversity of the continental subsurface. Nat. Geosci. 2018, 11, 707–717. [Google Scholar] [CrossRef]
- Osburn, M.R.; LaRowe, D.E.; Momper, L.M.; Amend, J.P. Chemolithotrophy in the continental deep subsurface: Sanford underground research facility (SURF), USA. Front. Microbiol. 2014, 5, 610. [Google Scholar] [CrossRef]
- Chivian, D.; Brodie, E.L.; Alm, E.J.; Culley, D.E.; Dehal, P.S.; DeSantis, T.Z.; Gihring, T.M.; Lapidus, A.; Lin, L.H.; Lowry, S.R.; et al. Environmental genomics reveals a single-species ecosystem deep within earth. Science 2008, 322, 275–278. [Google Scholar] [CrossRef]
- Kalwasińska, A.; Krawiec, A.; Deja-Sikora, E.; Golebiewski, M.; Kosobucki, P.; Brzezinska, M.S.; Walczak, M. Microbial diversity in deep-subsurface hot brines of Northwest Poland: From community structure to isolate characteristics. Appl. Environ. Microbiol. 2020, 86, e00252-20. [Google Scholar] [CrossRef] [PubMed]
- Lerm, S.; Westphal, A.; Miethling-Graff, R.; Alawi, M.; Seibt, A.; Wolfgramm, M.; Würdemann, H. Thermal effects on microbial composition and microbiologically induced corrosion and mineral precipitation affecting operation of a geothermal plant in a deep saline aquifer. Extremophiles 2013, 17, 311–327. [Google Scholar] [CrossRef]
- Sogin, M.L.; Morrison, H.G.; Huber, J.A.; Welch, D.M.; Huse, S.M.; Neal, P.R.; Arrieta, J.M.; Herndl, G.J. Microbial diversity in the deep sea and the underexplored “rare biosphere”. Proc. Natl. Acad. Sci. USA 2006, 103, 12115–12120. [Google Scholar] [CrossRef] [PubMed]
- Bartram, A.K.; Lynch, M.D.J.; Stearns, J.C.; Moreno-Hagelsieb, G.; Neufeld, J.D. Generation of multimillion-sequence 16S rRNA gene libraries from complex microbial communities by assembling paired-end Illumina reads. Appl. Environ. Microbiol. 2011, 77, 3846–3852. [Google Scholar] [CrossRef] [PubMed]
- Kotlar, H.K.; Lewin, A.; Johansen, J.; Throne-Holst, M.; Haverkamp, T.; Markussen, S.; Winnberg, A.; Ringrose, P.; Aakvik, T.; Ryeng, E.; et al. High coverage sequencing of DNA from microorganisms living in an oil reservoir 2.5 kilometres subsurface. Environ. Microbiol. Rep. 2011, 3, 674–681. [Google Scholar] [CrossRef] [PubMed]
- Lecroq, B.; Lejzerowicz, F.; Bachar, D.; Christen, R.; Esling, P.; Baerlocher, L.; Østerås, M.; Farinelli, L.; Pawlowski, J. Ultra-deep sequencing of foraminiferal microbarcodes unveils hidden richness of early monothalamous lineages in deep-sea sediments. Proc. Natl. Acad. Sci. USA 2011, 108, 13177–13182. [Google Scholar] [CrossRef] [PubMed]
- Furtak, K.; Grządziel, J.; Gałązka, A.; Niedźwiecki, J. Prevalence of unclassified bacteria in the soil bacterial community from floodplain meadows (fluvisols) under simulated flood conditions revealed by a metataxonomic approachss. Catena 2020, 188, 104448. [Google Scholar] [CrossRef]
- Dyda, M.; Pyzik, A.; Wilkojc, E.; Kwiatkowska-Kopka, B.; Sklodowska, A. Sklodowska bacterial and fungal diversity inside the medieval building constructed with sandstone plates and lime mortar as an example of the microbial colonization of a nutrient-limited extreme environment (Wawel royal castle, Krakow, Poland). Microorganisms 2019, 7, 416. [Google Scholar] [CrossRef]
- Lynch, M.D.J.; Neufeld, J.D. SSUnique: Detecting sequence novelty in microbiome surveys. mSystems 2016, 1, e00133-16. [Google Scholar] [CrossRef] [PubMed]
- Williams, M.M.; Domingo, J.W.S.; Meckes, M.C.; Kelty, C.A.; Rochon, H.S. Phylogenetic diversity of drinking water bacteria in a distribution system simulator. J. Appl. Microbiol. 2004, 96, 954–964. [Google Scholar] [CrossRef]
- Fajardo-Cavazos, P.; Nicholson, W. Bacillus endospores isolated from granite: Close molecular relationships to globally distributed Bacillus spp. from endolithic and extreme environments. Appl. Environ. Microbiol. 2006, 72, 2856–2863. [Google Scholar] [CrossRef] [PubMed]
- Bressuire-Isoard, C.; Broussolle, V.; Carlin, F. Sporulation environment influences spore properties in Bacillus: Evidence and insights on underlying molecular and physiological mechanisms. FEMS Microbiol. Rev. 2018, 42, 614–626. [Google Scholar] [CrossRef] [PubMed]
- Cortesão, M.; Fuchs, F.M.; Commichau, F.M.; Eichenberger, P.; Schuerger, A.C.; Nicholson, W.L.; Setlow, P.; Moeller, R. Bacillus subtilis Spore Resistance to Simulated Mars Surface Conditions. Front. Microbiol. 2019, 10, 333. [Google Scholar] [CrossRef]
- Moissl-Eichinger, C.; Cockell, C.; Rettberg, P. Venturing into new realms? Microorganisms in spacea. FEMS Microbiol. Rev. 2016, 40, 722–737. [Google Scholar] [CrossRef] [PubMed]
- Donio, M.B.S.; Ronica, S.F.A.; Viji, V.T.; Velmurugan, S.; Jenifer, J.A.; Michaelbabu, M.; Citarasu, T. Isolation and characterization of halophilic Bacillus sp. BS3 able to produce pharmacologically important biosurfactants. Asian Pac. J. Trop. Med. 2013, 6, 876–883. [Google Scholar] [CrossRef]
- Lenhart, J.S.; Schroeder, J.W.; Walsh, B.W.; Simmons, L.A. DNA repair and genome maintenance in Bacillus subtilis. Microbiol. Mol. Biol. Rev. 2012, 76, 530–564. [Google Scholar] [CrossRef]
- Stincone, P.; Brandelli, A. Marine bacteria as source of antimicrobial compounds. Crit. Rev. Biotechnol. 2020, 40, 306–319. [Google Scholar] [CrossRef]
- Lim, J.-M.; Jeon, C.O.; Lee, S.-M.; Lee, J.-C.; Xu, L.-H.; Jiang, C.-L.; Kim, C.-J. Bacillus salarius sp. nov., a halophilic, spore-forming bacterium isolated from a salt lake in China. Int. J. Syst. Evol. Microbiol. 2006, 56, 373–377. [Google Scholar] [CrossRef]
| Sample | Lithology | Location in Formation | Depth (m) |
|---|---|---|---|
| W2 | Clayey salt (zuber) with veins of fibrous salt and lenses of gypsum and anhydrite | layers in spiza salts | 110 |
| W3 | Siltstone and sandstone | layers in spiza salts | 110 |
| W4 | Siltstone with veins of fibrous salt and lenses of anhydrite | layers in green salt | 130 |
| Treatment | Headspace Composition | Solution Composition * |
|---|---|---|
| Methane production | N2 | sterilized ddH2O |
| Anaerobic methane oxidation | 10% CH4 (v/v) in N2 | NaSO4 (5mM) in sterilized ddH2O |
| Anaerobic respiration | N2 | sterilized ddH2O |
| Aerobic methane oxidation | 10% CH4 (v/v) in air | sterilized ddH2O |
| Aerobic respiration | ambient air | sterilized ddH2O |
| Parameter | W2 | W3 | W4 | ||||
|---|---|---|---|---|---|---|---|
| Mean | ±SD | Mean | ±SD | Mean | ±SD | ||
| EC | mS cm−1 | 114.27 | 3.58 | 12.74 | 0.03 | 37.30 | 1.30 |
| TOC | % | 3.35 | 0.04 | 1.06 | 0.49 | 0.76 | 0.05 |
| N-NO3 | mg kg−1 | 0.00 | 0.00 | 0.00 | 0.00 | 0.40 | 0.02 |
| N-NH4 | mg kg−1 | 40.70 | 0.12 | 37.75 | 0.15 | 69.62 | 0.70 |
| P-PO4 | mg kg−1 | 0.42 | 0.10 | 0.19 | 0.01 | 0.18 | 0.02 |
| S-SO4 | mg g−1 | 25.84 | 2.90 | 18.77 | 1.09 | 21.41 | 2.22 |
| Fe | g kg−1 | 10.21 | 1.02 | 106.05 | 36.89 | 103.27 | 52.36 |
| Mn | g kg−1 | 0.43 | 0.04 | 2.49 | 0.10 | 0.83 | 0.11 |
| Surface area | m² g−1 | 14.75 | 0.02 | 19.96 | 0.03 | 20.31 | 0.06 |
| Pore volume | cm³ g−1 | 0.03 | 0.01 | 0.05 | 0.01 | 0.05 | 0.01 |
| Moisture | % | 4.32 | 0.02 | 10.01 | 0.11 | 7.39 | 0.13 |
| Total water capacity | % | 31.52 | 1.22 | 51.94 | 3.34 | 42.03 | 1.36 |
| NaCl | g L−1 | 221.02 | 3.5 | 24.64 | 0.00 | 72.15 | 1.3 |
| Sample | Observed | Shannon | Simpson | Simpson D’ (Diversity) | InvSimpson | Fisher |
|---|---|---|---|---|---|---|
| W2 | 77 | 3.234 | 0.872 | 0.128 | 7.819 | 8.246 |
| W3 | 199 | 0.621 | 0.166 | 0.834 | 1.199 | 23.639 |
| W4 | 242 | 4.987 | 0.987 | 0.013 | 75.955 | 29.951 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Goraj, W.; Szafranek-Nakonieczna, A.; Grządziel, J.; Polakowski, C.; Słowakiewicz, M.; Zheng, Y.; Gałązka, A.; Stępniewska, Z.; Pytlak, A. Microbial Involvement in Carbon Transformation via CH4 and CO2 in Saline Sedimentary Pool. Biology 2021, 10, 792. https://doi.org/10.3390/biology10080792
Goraj W, Szafranek-Nakonieczna A, Grządziel J, Polakowski C, Słowakiewicz M, Zheng Y, Gałązka A, Stępniewska Z, Pytlak A. Microbial Involvement in Carbon Transformation via CH4 and CO2 in Saline Sedimentary Pool. Biology. 2021; 10(8):792. https://doi.org/10.3390/biology10080792
Chicago/Turabian StyleGoraj, Weronika, Anna Szafranek-Nakonieczna, Jarosław Grządziel, Cezary Polakowski, Mirosław Słowakiewicz, Yanhong Zheng, Anna Gałązka, Zofia Stępniewska, and Anna Pytlak. 2021. "Microbial Involvement in Carbon Transformation via CH4 and CO2 in Saline Sedimentary Pool" Biology 10, no. 8: 792. https://doi.org/10.3390/biology10080792
APA StyleGoraj, W., Szafranek-Nakonieczna, A., Grządziel, J., Polakowski, C., Słowakiewicz, M., Zheng, Y., Gałązka, A., Stępniewska, Z., & Pytlak, A. (2021). Microbial Involvement in Carbon Transformation via CH4 and CO2 in Saline Sedimentary Pool. Biology, 10(8), 792. https://doi.org/10.3390/biology10080792






