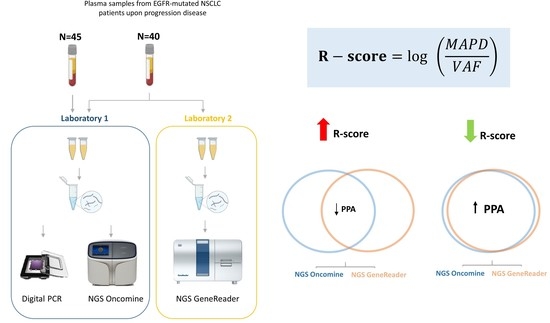
R-Score: A New Parameter to Assess the Quality of Variants’ Calls Assessed by NGS Using Liquid Biopsies
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Patients and Samples
2.2. Laboratory Procedures
2.3. Parameters
2.4. Statistical Analysis
3. Results
3.1. EGFR Mutation Detection by dPCR and NGS-Oncomine
3.2. VAF and MAPD Involvement in the Agreement between dPCR and NGS-Oncomine Calls
3.3. R-Score Is a Useful Parameter to Select Reliable Variant Calls
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Provencio, M.; Torrente, M.; Calvo, V.; Gutiérrez, L.; Pérez-Callejo, D.; Pérez-Barrios, C.; Barquín, M.; Royuela, A.; Rodriguez-Alfonso, B.; Sotelo, M.; et al. Dynamic circulating tumor DNA quantificaton for the individualization of non-small-cell lung cancer patients treatment. Oncotarget 2017, 8, 60291–60298. [Google Scholar] [CrossRef] [Green Version]
- Ettinger, D.S.; Wood, D.E.; Aisner, D.L.; Akerley, W.; Bauman, J.; Chirieac, L.R.; D’Amico, T.A.; Decamp, M.M.; Dilling, T.J.; Dobelbower, M.; et al. Non-small cell lung cancer, version 5.2017: Clinical practice guidelines in oncology. JNCCN J. Natl. Compr. Cancer Netw. 2017, 15, 504–535. [Google Scholar] [CrossRef]
- Pérez-Callejo, D.; Romero, A.; Provencio, M.; Torrente, M. Liquid biopsy based biomarkers in non-small cell lung cancer for diagnosis and treatment monitoring. Transl. Lung Cancer Res. 2016, 5, 455–465. [Google Scholar] [CrossRef] [Green Version]
- Romero, A.; Serna-Blasco, R.; Calvo, V.; Provencio, M. Use of Liquid Biopsy in the Care of Patients with Non-Small Cell Lung Cancer. Curr. Treat. Options Oncol. 2021, 22, 86. [Google Scholar] [CrossRef]
- Provencio, M.; Serna-Blasco, R.; Franco, F.; Calvo, V.; Royuela, A.; Auglytė, M.; Sánchez-Hernández, A.; de Julián Campayo, M.; García-Girón, C.; Dómine, M.; et al. Analysis of circulating tumour DNA to identify patients with epidermal growth factor receptor–positive non-small cell lung cancer who might benefit from sequential tyrosine kinase inhibitor treatment. Eur. J. Cancer 2021, 149, 61–72. [Google Scholar] [CrossRef] [PubMed]
- McLean, A.; Barnes, D.; Troy, L. Diagnosing Lung Cancer: The Complexities of Obtaining a Tissue Diagnosis in the Era of Minimally Invasive and Personalised Medicine. J. Clin. Med. 2018, 7, 163. [Google Scholar] [CrossRef] [Green Version]
- Schrank, Z.; Chhabra, G.; Lin, L.; Iderzorig, T.; Osude, C.; Khan, N.; Kuckovic, A.; Singh, S.; Miller, R.J.; Puri, N. Current molecular-targeted therapies in NSCLC and their mechanism of resistance. Cancers 2018, 10, 224. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hallin, J.; Engstrom, L.D.; Hargi, L.; Calinisan, A.; Aranda, R.; Briere, D.M.; Sudhakar, N.; Bowcut, V.; Baer, B.R.; Ballard, J.A.; et al. The KRASG12C inhibitor MRTX849 provides insight toward therapeutic susceptibility of KRAS-mutant cancers in mouse models and patients. Cancer Discov. 2020, 10, 54–71. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hong, D.S.; Fakih, M.G.; Strickler, J.H.; Desai, J.; Durm, G.A.; Shapiro, G.I.; Falchook, G.S.; Price, T.J.; Sacher, A.; Denlinger, C.S.; et al. KRAS G12C Inhibition with Sotorasib in Advanced Solid Tumors. N. Engl. J. Med. 2020, 383, 1207–1217. [Google Scholar] [CrossRef] [PubMed]
- Stewart, E.L.; Tan, S.Z.; Liu, G.; Tsao, M.S. Known and putative mechanisms of resistance to EGFR targeted therapies in NSCLC patients with EGFR mutations-a review. Transl. Lung Cancer Res. 2015, 4, 67–81. [Google Scholar] [PubMed]
- Dal Maso, A.; Lorenzi, M.; Roca, E.; Pilotto, S.; Macerelli, M.; Polo, V.; Cecere, F.L.; Del Conte, A.; Nardo, G.; Buoro, V.; et al. Clinical Features and Progression Pattern of Acquired T790M-positive Compared With T790M-negative EGFR Mutant Non–small-cell Lung Cancer: Catching Tumor and Clinical Heterogeneity Over Time Through Liquid Biopsy. Clin. Lung Cancer 2020, 21, 1–14.e3. [Google Scholar] [CrossRef] [Green Version]
- Romero, A.; Serna-Blasco, R.; Alfaro, C.; Sánchez-Herrero, E.; Barquín, M.; Turpin, M.C.; Chico, S.; Sanz-Moreno, S.; Rodrigez-Festa, A.; Laza-Briviesca, R.; et al. ctDNA analysis reveals different molecular patterns upon disease progression in patients treated with osimertinib. Transl. Lung Cancer Res. 2020, 9, 532–540. [Google Scholar] [CrossRef]
- Provencio, M.; Torrente, M.; Calvo, V.; Pérez-Callejo, D.; Gutiérrez, L.; Franco, F.; Pérez-Barrios, C.; Barquín, M.; Royuela, A.; García-García, F.; et al. Prognostic value of quantitative ctDNA levels in non small cell lung cancer patients. Oncotarget 2018, 9, 488–494. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Salk, J.J.; Schmitt, M.W.; Loeb, L.A. Enhancing the accuracy of next-generation sequencing for detecting rare and subclonal mutations. Nat. Rev. Genet. 2018, 19, 269–285. [Google Scholar] [CrossRef]
- Kivioja, T.; Vähärautio, A.; Karlsson, K.; Bonke, M.; Enge, M.; Linnarsson, S.; Taipale, J. Counting absolute numbers of molecules using unique molecular identifiers. Nat. Methods 2012, 9, 72–74. [Google Scholar] [CrossRef] [PubMed]
- Romero, A.; Jantus-Lewintre, E.; García-Peláez, B.; Royuela, A.; Insa, A.; Cruz, P.; Collazo, A.; Pérez Altozano, J.; Vidal, O.J.; Diz, P.; et al. Comprehensive cross-platform comparison of methods for non-invasive EGFR mutation testing: Results of the RING observational trial. Mol. Oncol. 2021, 15, 43–56. [Google Scholar] [CrossRef] [PubMed]
- Ion Reporter™ Software 5.12 USER GUIDE. Available online: https://assets.thermofisher.com/TFS-Assets/LSG/manuals/MAN0018032_IonReporterSoftware_5_12_UG.pdf (accessed on 5 July 2021).
- Deveson, I.W.; Gong, B.; Lai, K.; LoCoco, J.S.; Richmond, T.A.; Schageman, J.; Zhang, Z.; Novoradovskaya, N.; Willey, J.C.; Jones, W.; et al. Evaluating the analytical validity of circulating tumor DNA sequencing assays for precision oncology. Nat. Biotechnol. 2021, 1–14. [Google Scholar] [CrossRef]
- Le Tourneau, C.; Delord, J.P.; Gonçalves, A.; Gavoille, C.; Dubot, C.; Isambert, N.; Campone, M.; Trédan, O.; Massiani, M.A.; Mauborgne, C.; et al. Molecularly targeted therapy based on tumour molecular profiling versus conventional therapy for advanced cancer (SHIVA): A multicentre, open-label, proof-of-concept, randomised, controlled phase 2 trial. Lancet Oncol. 2015, 16, 1324–1334. [Google Scholar] [CrossRef]
- Fisher, K.E.; Zhang, L.; Wang, J.; Smith, G.H.; Newman, S.; Schneider, T.M.; Pillai, R.N.; Kudchadkar, R.R.; Owonikoko, T.K.; Ramalingam, S.S.; et al. Clinical Validation and Implementation of a Targeted Next-Generation Sequencing Assay to Detect Somatic Variants in Non-Small Cell Lung, Melanoma, and Gastrointestinal Malignancies. J. Mol. Diagn. 2016, 18, 299–315. [Google Scholar] [CrossRef]
- Cottrell, C.E.; Al-Kateb, H.; Bredemeyer, A.J.; Duncavage, E.J.; Spencer, D.H.; Abel, H.J.; Lockwood, C.M.; Hagemann, I.S.; O’Guin, S.M.; Burcea, L.C.; et al. Validation of a next-generation sequencing assay for clinical molecular oncology. J. Mol. Diagn. 2014, 16, 89–105. [Google Scholar] [CrossRef]
- Provencio, M.; Pérez-Barrios, C.; Barquin, M.; Calvo, V.; Franco, F.; Sánchez, E.; Sánchez, R.; Marsden, D.; Cristóbal Sánchez, J.; Martin Acosta, P.; et al. Next-generation sequencing for tumor mutation quantification using liquid biopsies. Clin. Chem. Lab. Med. 2019, 58, 306–313. [Google Scholar] [CrossRef]
- Sánchez-Herrero, E.; Serna-Blasco, R.; Ivanchuk, V.; García-Campelo, R.; Dómine Gómez, M.; Sánchez, J.M.; Massutí, B.; Reguart, N.; Camps, C.; Sanz-Moreno, S.; et al. NGS-based liquid biopsy profiling identifies mechanisms of resistance to ALK inhibitors: A step toward personalized NSCLC treatment. Mol. Oncol. 2021, 15, 2363–2376. [Google Scholar] [CrossRef]
- Merker, J.D.; Oxnard, G.R.; Compton, C.; Diehn, M.; Hurley, P.; Lazar, A.J.; Lindeman, N.; Lockwood, C.M.; Rai, A.J.; Schilsky, R.L.; et al. Circulating tumor DNA analysis in patients with cancer: American society of clinical oncology and college of American pathologists joint review. J. Clin. Oncol. 2018, 142, 1242–1253. [Google Scholar]
- Gandara, D.R.; Paul, S.M.; Kowanetz, M.; Schleifman, E.; Zou, W.; Li, Y.; Rittmeyer, A.; Fehrenbacher, L.; Otto, G.; Malboeuf, C.; et al. Blood-based tumor mutational burden as a predictor of clinical benefit in non-small-cell lung cancer patients treated with atezolizumab. Nat. Med. 2018, 24, 1441–1448. [Google Scholar] [CrossRef]
- Rizvi, N.A.; Hellmann, M.D.; Snyder, A.; Kvistborg, P.; Makarov, V.; Havel, J.J.; Lee, W.; Yuan, J.; Wong, P.; Ho, T.S.; et al. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 2015, 348, 124–128. [Google Scholar] [CrossRef] [Green Version]
- Kowanetz, M.; Zou, W.; Shames, D.S.; Cummings, C.; Rizvi, N.; Spira, A.I.; Frampton, G.M.; Leveque, V.; Flynn, S.; Mocci, S.; et al. Tumor mutation load assessed by FoundationOne (FM1) is associated with improved efficacy of atezolizumab (atezo) in patients with advanced NSCLC. Ann. Oncol. 2016, 27, vi23. [Google Scholar] [CrossRef]
- Chalmers, Z.R.; Connelly, C.F.; Fabrizio, D.; Gay, L.; Ali, S.M.; Ennis, R.; Schrock, A.; Campbell, B.; Shlien, A.; Chmielecki, J.; et al. Analysis of 100,000 human cancer genomes reveals the landscape of tumor mutational burden. Genome Med. 2017, 9, 34. [Google Scholar] [CrossRef] [PubMed]
- Merino, D.M.; McShane, L.M.; Fabrizio, D.; Funari, V.; Chen, S.J.; White, J.R.; Wenz, P.; Baden, J.; Barrett, J.C.; Chaudhary, R.; et al. Establishing guidelines to harmonize tumor mutational burden (TMB): In silico assessment of variation in TMB quantification across diagnostic platforms: Phase I of the Friends of Cancer Research TMB Harmonization Project. J. Immunother. Cancer 2020, 8, e000147. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- O’Leary, B.; Hrebien, S.; Beaney, M.; Fribbens, C.; Garcia-Murillas, I.; Jiang, J.; Li, Y.; Bartlett, C.H.; André, F.; Loibl, S.; et al. Comparison of beaming and droplet digital PCR for circulating tumor DNA analysis. Clin. Chem. 2019, 65, 1405–1413. [Google Scholar] [CrossRef]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Serna-Blasco, R.; Sánchez-Herrero, E.; Berrocal Renedo, M.; Calabuig-Fariñas, S.; Molina-Vila, M.Á.; Provencio, M.; Romero, A. R-Score: A New Parameter to Assess the Quality of Variants’ Calls Assessed by NGS Using Liquid Biopsies. Biology 2021, 10, 954. https://doi.org/10.3390/biology10100954
Serna-Blasco R, Sánchez-Herrero E, Berrocal Renedo M, Calabuig-Fariñas S, Molina-Vila MÁ, Provencio M, Romero A. R-Score: A New Parameter to Assess the Quality of Variants’ Calls Assessed by NGS Using Liquid Biopsies. Biology. 2021; 10(10):954. https://doi.org/10.3390/biology10100954
Chicago/Turabian StyleSerna-Blasco, Roberto, Estela Sánchez-Herrero, María Berrocal Renedo, Silvia Calabuig-Fariñas, Miguel Ángel Molina-Vila, Mariano Provencio, and Atocha Romero. 2021. "R-Score: A New Parameter to Assess the Quality of Variants’ Calls Assessed by NGS Using Liquid Biopsies" Biology 10, no. 10: 954. https://doi.org/10.3390/biology10100954
APA StyleSerna-Blasco, R., Sánchez-Herrero, E., Berrocal Renedo, M., Calabuig-Fariñas, S., Molina-Vila, M. Á., Provencio, M., & Romero, A. (2021). R-Score: A New Parameter to Assess the Quality of Variants’ Calls Assessed by NGS Using Liquid Biopsies. Biology, 10(10), 954. https://doi.org/10.3390/biology10100954