Starch/Poly(glycerol-adipate) Nanocomposites: A Novel Oral Drug Delivery Device
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Methods
2.2.1. PGA and Cy5-PGA Synthesis
2.2.2. PGA NP Preparation and Characterization by Dynamic Light Scattering (DLS)
2.2.3. Casting of Films
2.2.4. In vitro Static Digestion of Starch/Blue PGA Nanocomposites
2.2.5. Dynamic Digestion
2.2.6. NP Release Profile Evaluation
2.2.7. Cell Culture Conditions
2.2.8. Cytotoxicity of Study Formulations
2.2.9. Minimum Inhibitory Concentration (MIC)
2.2.10. Molecular Dynamics simulations (MD)
3. Results and Discussion
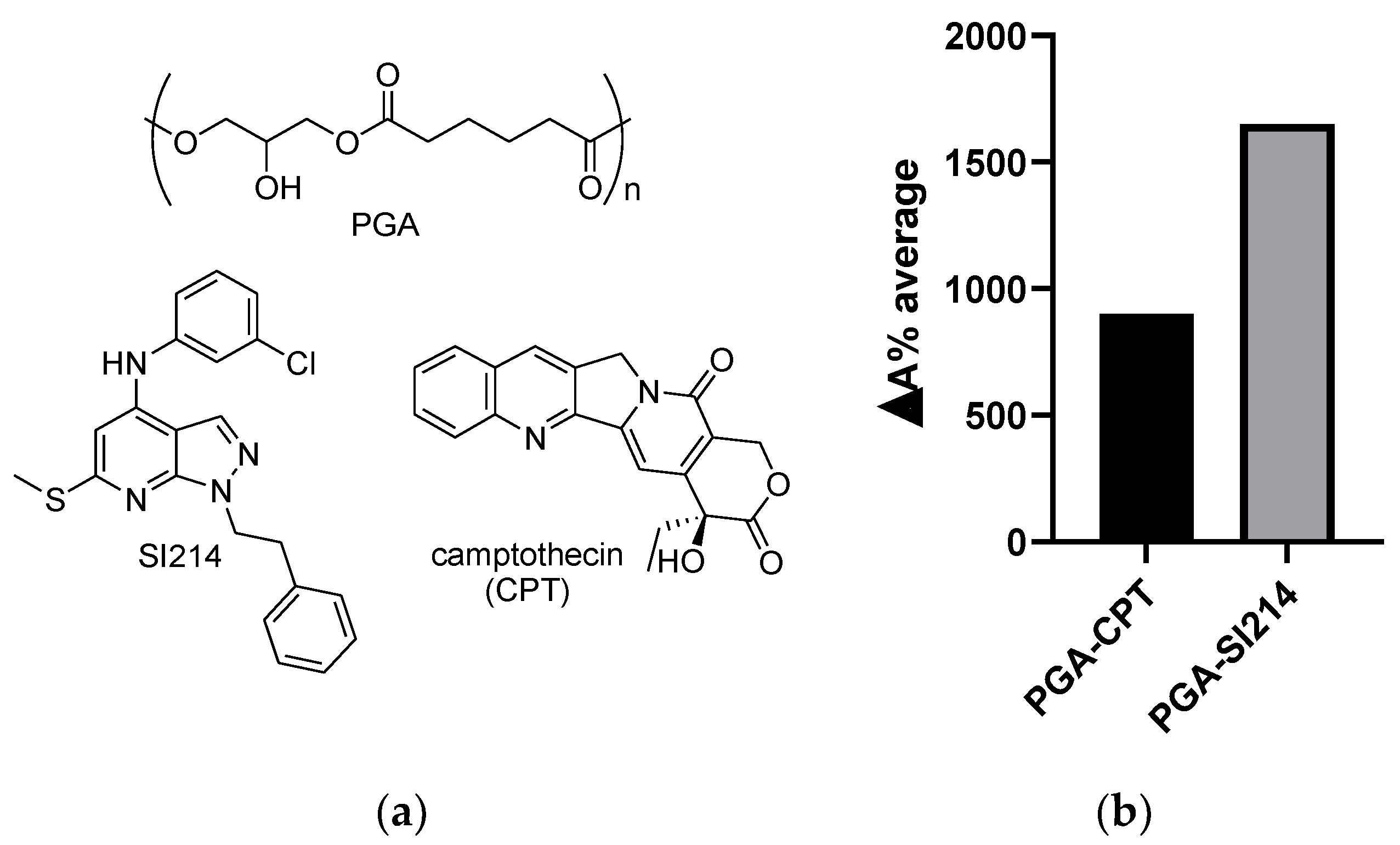
3.1. NPs’ Formation and Drug Water Solubility Enhancement
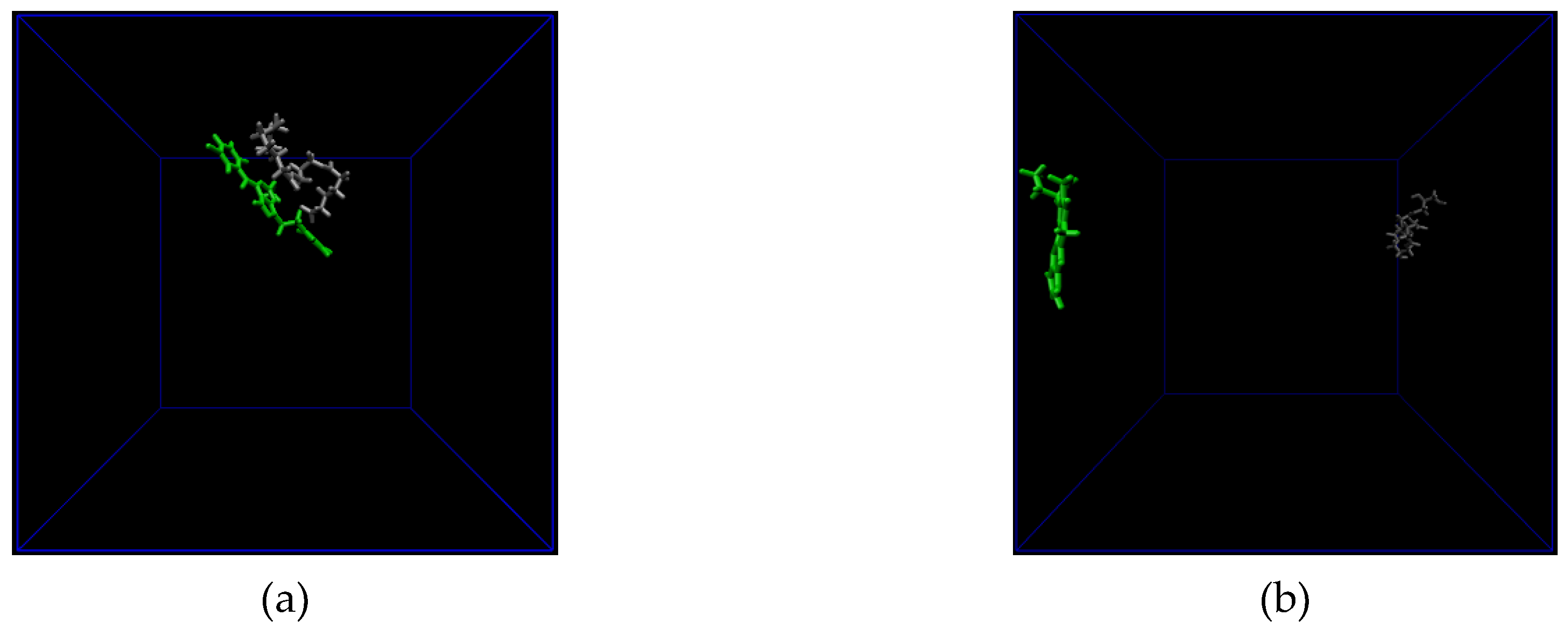
3.2. All Atom Molecular Dynamics (MD) Simulation
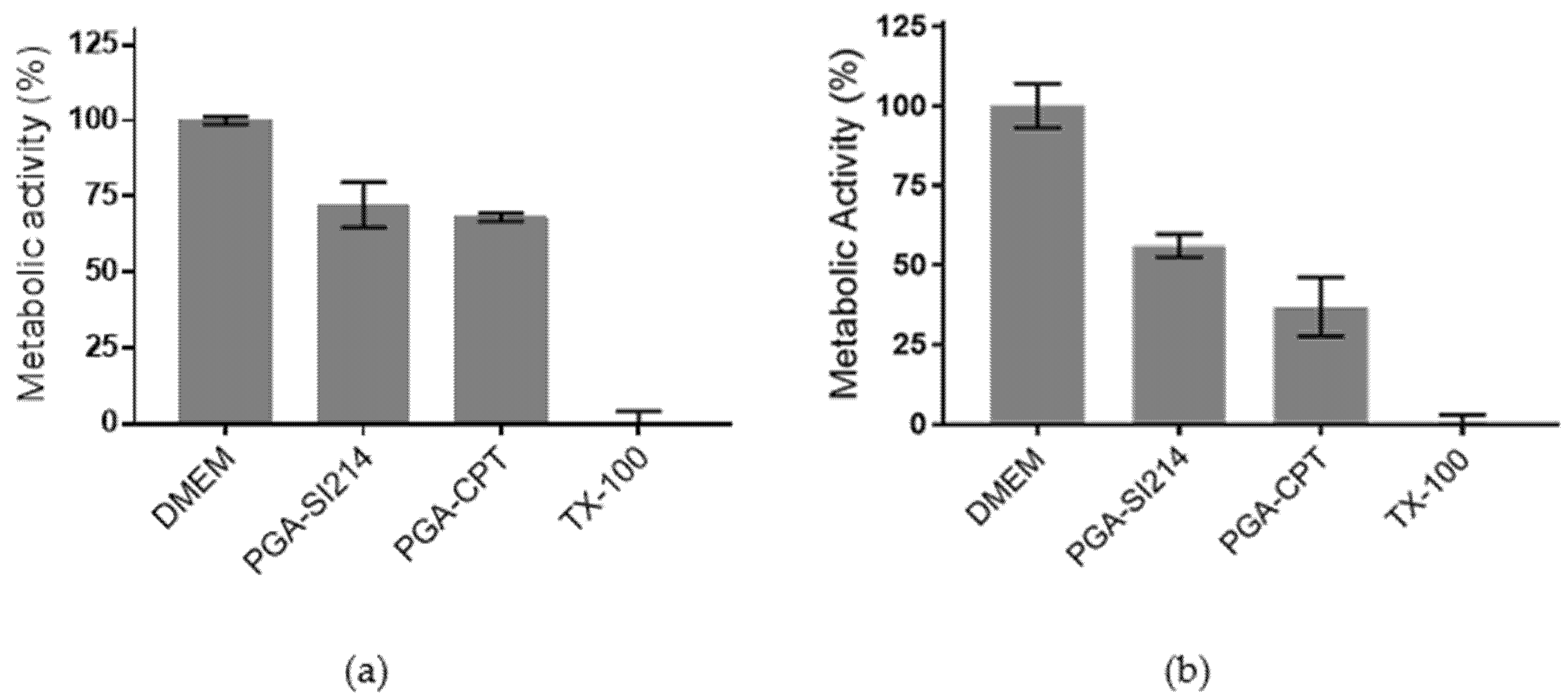
3.3. Bioactivity of the Loaded NPs against Cancer Cells and Bacteria
3.4. Film Formation and NPs Entrapment
3.5. Blue-NPs’ Release from Film in Different Buffers
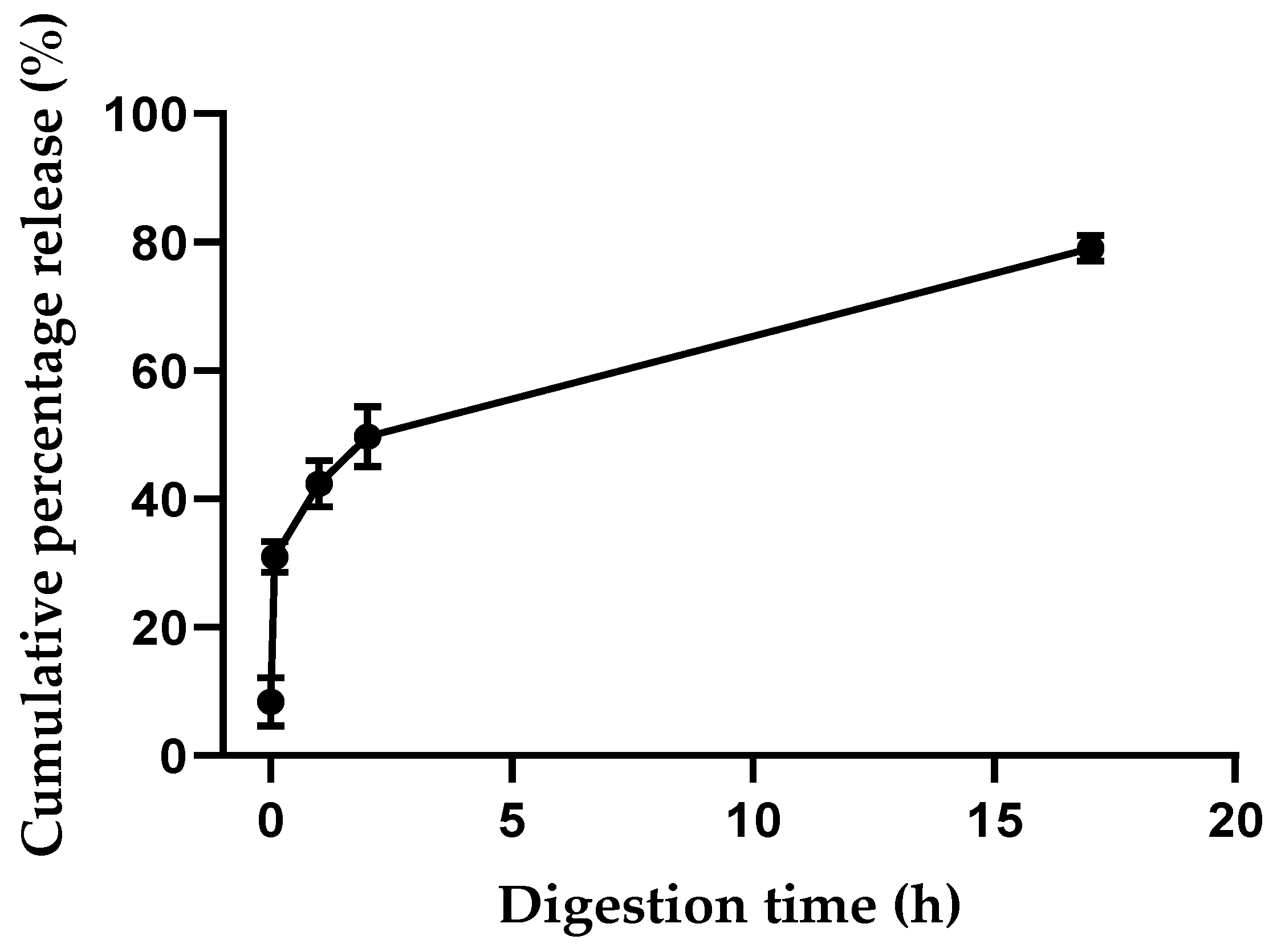
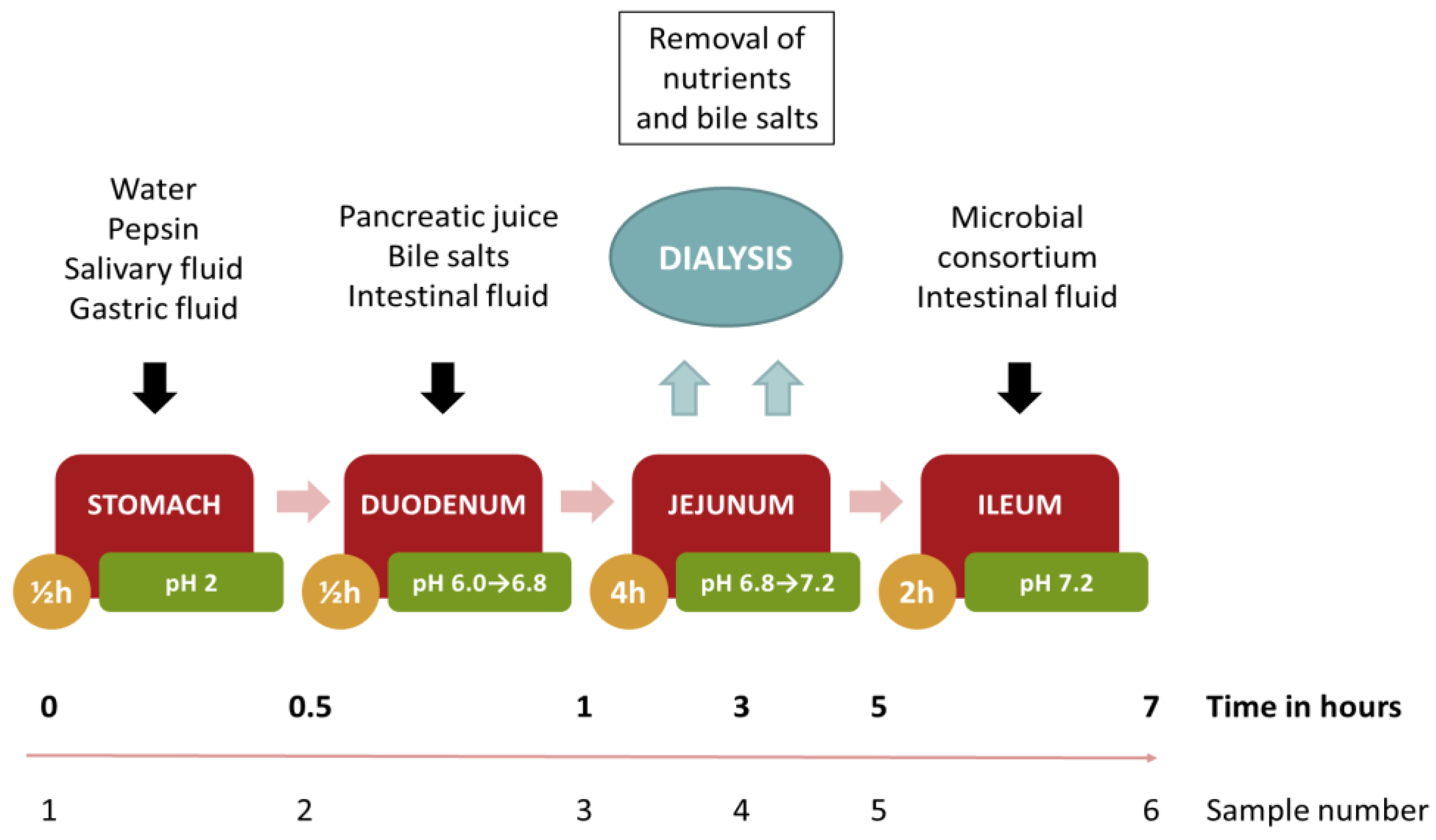
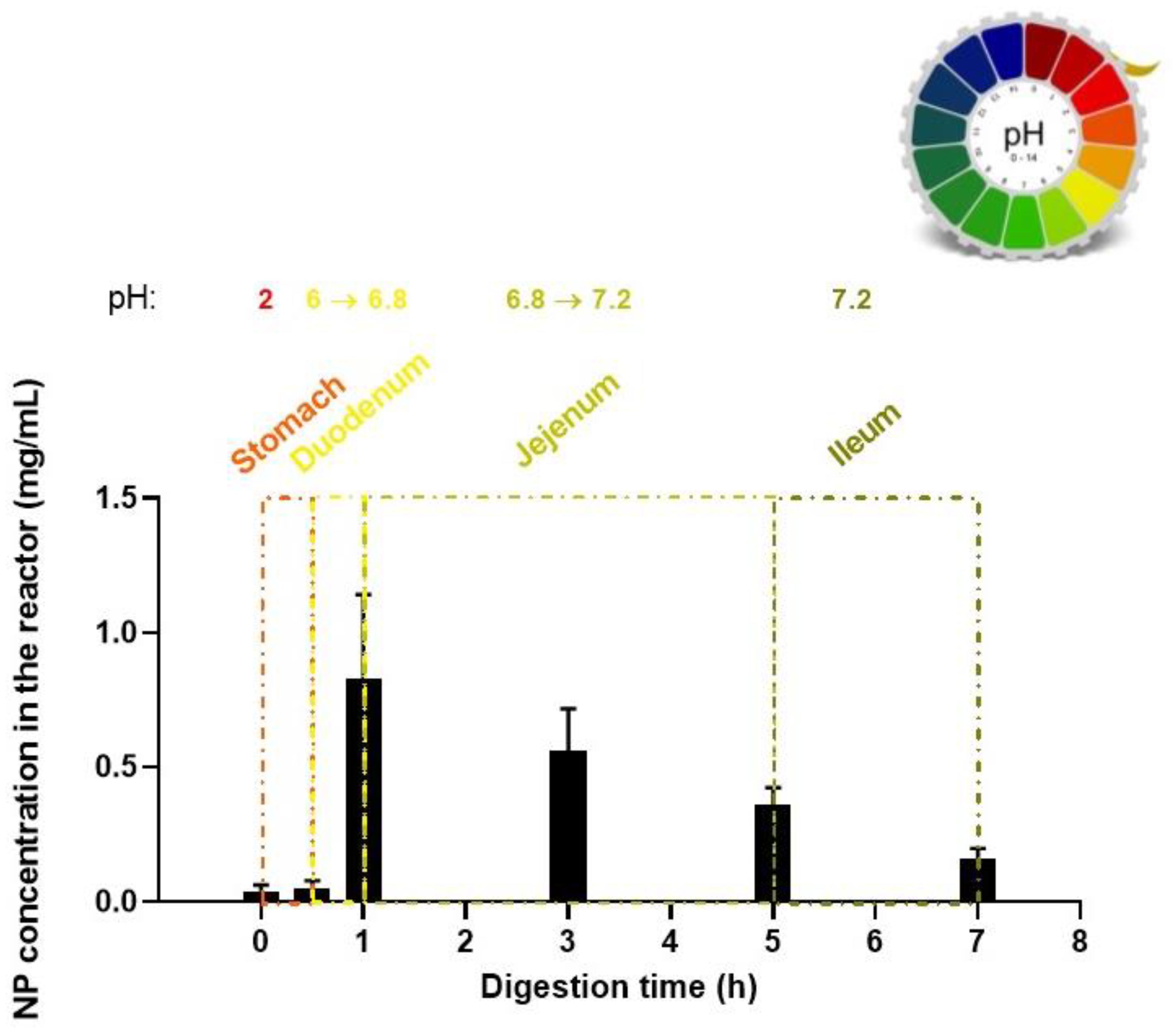
3.6. Digestion Model of the Edible Film
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Liu, L.S.; Kost, J.; Yan, F.; Spiro, R.C. Hydrogels from biopolymer hybrid for biomedical, food, and functional food applications. Polymers (Basel) 2012. [Google Scholar] [CrossRef]
- Swainson, S.M.E.; Styliari, I.D.; Taresco, V.; Garnett, M.C. Poly (glycerol adipate) (PGA), an enzymatically synthesized functionalizable polyester and versatile drug delivery carrier: A literature update. Polymers (Basel) 2019, 11, 1561. [Google Scholar] [CrossRef] [PubMed]
- Sagnelli; Cavanagh; Xu; Swainson; Blennow; Duncan; Taresco; Howdle Starch/poly (glycerol-adipate) nanocomposite film as novel biocompatible materials. Coatings 2019, 9, 482. [CrossRef]
- Taresco, V.; Creasey, R.G.; Kennon, J.; Mantovani, G.; Alexander, C.; Burley, J.C.; Garnett, M.C. Variation in structure and properties of poly(glycerol adipate) via control of chain branching during enzymatic synthesis. Polymer (Guildf) 2016, 89, 41–49. [Google Scholar] [CrossRef]
- Gaucher, G.; Satturwar, P.; Jones, M.C.; Furtos, A.; Leroux, J.C. Polymeric micelles for oral drug delivery. Eur. J. Pharm. Biopharm. 2010, 76, 147–158. [Google Scholar] [CrossRef]
- Feitosa, R.C.; Geraldes, D.C.; Beraldo-De-Araújo, V.L.; Costa, J.S.R.; Oliveira-Nascimento, L. Pharmacokinetic aspects of nanoparticle-in-matrix drug delivery systems for oral/buccal delivery. Front. Pharmacol. 2019, 10, 1057. [Google Scholar] [CrossRef]
- Vasanthan, T.; Hoover, R. Barley starch: Production, properties, modification and uses. Starch 2009, 601–628. [Google Scholar]
- Pérez, S.; Bertoft, E. The molecular structures of starch components and their contribution to the architecture of starch granules: A comprehensive review. Starch/Staerke 2010, 62, 389–420. [Google Scholar] [CrossRef]
- Sagnelli, D.; Hebelstrup, K.H.; Leroy, E.; Rolland-Sabaté, A.; Guilois, S.; Kirkensgaard, J.J.K.; Mortensen, K.; Lourdin, D.; Blennow, A. Plant-crafted starches for bioplastics production. Carbohydr. Polym. 2016, 152, 398–408. [Google Scholar] [CrossRef] [PubMed]
- Blennow, A.; Engelsen, S.B. Helix-breaking news: Fighting crystalline starch energy deposits in the cell. Trends Pant Sci 2010, 15, 236–240. [Google Scholar] [CrossRef]
- Roskoski, R. Properties of FDA-approved small molecule protein kinase inhibitors. Pharmacol. Res. 2019, 144, 19–50. [Google Scholar] [CrossRef] [PubMed]
- FDA-Approved Protein Kinase Inhibitors/US Food and Drug Administration Approved Small Molecule Protein Kinase Inhibitors. Available online: http://webcache.googleusercontent.com/search?q=cache:http://www.brimr.org/PKI/PKIs.htm (accessed on 22 October 2019).
- Search of: Kinase Inhibitors | Recruiting, Not yet Recruiting, Active, not Recruiting, Enrolling by Invitation Studies - List Results - ClinicalTrials.gov. Available online: https://clinicaltrials.gov/ct2/results?term=kinase+inhibitors&Search=Apply&recrs=b&recrs=a&recrs=f&recrs=d&age_v=&gndr=&type=&rslt= (accessed on 21 October 2019).
- Bhullar, K.S.; Lagarón, N.O.; McGowan, E.M.; Parmar, I.; Jha, A.; Hubbard, B.P.; Rupasinghe, H.P.V. Kinase-targeted cancer therapies: Progress, challenges and future directions. Mol. Cancer 2018, 17, 48. [Google Scholar] [CrossRef] [PubMed]
- Herbrink, M.; Nuijen, B.; Schellens, J.H.M.; Beijnen, J.H. Variability in bioavailability of small molecular tyrosine kinase inhibitors. Cancer Treat. Rev. 2015, 41, 412–422. [Google Scholar] [CrossRef] [PubMed]
- Pharmacokinetic Considerations and Challenges in Oral Anticancer Drug Therapy | Perspective Article | Pharmaceutical Journal. Available online: https://www.pharmaceutical-journal.com/research/perspective-article/pharmacokinetic-considerations-and-challenges-in-oral-anticancer-drug-therapy/20206478.article?firstPass=false (accessed on 22 October 2019).
- Matteoni, S.; Abbruzzese, C.; Matarrese, P.; De Luca, G.; Mileo, A.M.; Miccadei, S.; Schenone, S.; Musumeci, F.; Haas, T.L.; Sette, G.; et al. The kinase inhibitor SI113 induces autophagy and synergizes with quinacrine in hindering the growth of human glioblastoma multiforme cells. J. Exp. Clin. Cancer Res. 2019, 38, 202. [Google Scholar] [CrossRef] [PubMed]
- Fallacara, A.L.; Passannanti, R.; Mori, M.; Iovenitti, G.; Musumeci, F.; Greco, C.; Crespan, E.; Kissova, M.; Maga, G.; Tarantelli, C.; et al. Identification of a new family of pyrazolo[3,4-d]pyrimidine derivatives as multitarget Fyn-Blk-Lyn inhibitors active on B- and T-lymphoma cell lines. Eur. J. Med. Chem. 2019, 181, 111545. [Google Scholar] [CrossRef]
- Tintori, C.; Fallacara, A.L.; Radi, M.; Zamperini, C.; Dreassi, E.; Crespan, E.; Maga, G.; Schenone, S.; Musumeci, F.; Brullo, C.; et al. Combining X-ray crystallography and molecular modeling toward the optimization of pyrazolo[3,4-d]pyrimidines as potent c-Src inhibitors active in vivo against Neuroblastoma. J. Med. Chem. 2015, 58, 347–361. [Google Scholar] [CrossRef]
- Radi, M.; Tintori, C.; Musumeci, F.; Brullo, C.; Zamperini, C.; Dreassi, E.; Fallacara, A.L.; Vignaroli, G.; Crespan, E.; Zanoli, S.; et al. Design, synthesis, and biological evaluation of pyrazolo[3,4-d]pyrimidines active in vivo on the Bcr-Abl T315I mutant. J. Med. Chem. 2013, 56, 5382–5394. [Google Scholar] [CrossRef]
- Radi, M.; Brullo, C.; Crespan, E.; Tintori, C.; Musumeci, F.; Biava, M.; Schenone, S.; Dreassi, E.; Zamperini, C.; Maga, G.; et al. Identification of potent c-Src inhibitors strongly affecting the proliferation of human neuroblastoma cells. Bioorg. Med. Chem. Lett. 2011, 21, 5928–5933. [Google Scholar] [CrossRef]
- Sanna, M.; Sicilia, G.; Alazzo, A.; Singh, N.; Musumeci, F.; Schenone, S.; Spriggs, K.A.; Burley, J.C.; Garnett, M.C.; Taresco, V.; et al. Water solubility enhancement of pyrazolo[3,4-d]pyrimidine derivatives via miniaturized polymer-drug microarrays. ACS Med. Chem. Lett. 2018, 9, 193–197. [Google Scholar] [CrossRef]
- El-Sayed, A.M.; Ibrahim, S.M.; Soltan, M.K.; Abo-Kul, M.E. Synthesis and antimicrobial activity of newly synthesized 4-substituted-pyrazolo[3,4-d]pyrimidine derivatives. Med. Chem. Res. 2017, 26, 1107–1116. [Google Scholar] [CrossRef]
- Bakavoli, M.; Bagherzadeh, G.; Vaseghifar, M.; Shiri, A.; Pordel, M.; Mashreghi, M.; Pordeli, P.; Araghi, M. Molecular iodine promoted synthesis of new pyrazolo[3,4-d]pyrimidine derivatives as potential antibacterial agents. Eur. J. Med. Chem. 2010, 45, 647–650. [Google Scholar] [CrossRef] [PubMed]
- Ali, A.; Taylor, G.E.; Ellsworth, K.; Harris, G.; Painter, R.; Silver, L.L.; Young, K. Novel pyrazolo[3,4-d]pyrimidine-based inhibitors of Staphlococcus aureus DNA polymerase III: Design, synthesis, and biological evaluation. J. Med. Chem. 2003, 46, 1824–1830. [Google Scholar] [CrossRef] [PubMed]
- Pommier, Y.; Leo, E.; Zhang, H.; Marchand, C. DNA topoisomerases and their poisoning by anticancer and antibacterial drugs. Chem. Biol. 2010, 17, 421–433. [Google Scholar] [CrossRef] [PubMed]
- Dong, Q.; Luo, J.; Qiu, W.; Cai, L.; Anjum, S.; Li, B.; Hou, M.; Xie, G.; Sun, G. Inhibitory effect of camptothecin against rice bacterial brown stripe pathogen acidovorax avenae subsp. avenae RS-2. Molecules 2016, 21, 978. [Google Scholar] [CrossRef] [PubMed]
- Burke, T.G.; Mi, Z. The structural basis of camptothecin interactions with human serum albumin: Impact on drug stability. J. Med. Chem. 1994, 37, 40–46. [Google Scholar] [CrossRef] [PubMed]
- Khan, A.R.; Magnusson, J.P.; Watson, S.; Grabowska, A.M.; Wilkinson, R.W.; Alexander, C.; Pritchard, D. Camptothecin prodrug block copolymer micelles with high drug loading and target specificity. Polym. Chem. 2014, 5, 5320–5329. [Google Scholar] [CrossRef]
- Fessi, H.; Puisieux, F.; Devissaguet, J.P.; Ammoury, N.; Benita, S. Nanocapsule formation by interfacial polymer deposition following solvent displacement. Int. J. Pharm. 1989, 55, R1–R4. [Google Scholar] [CrossRef]
- Sagnelli, D.; Chessa, S.; Mandalari, G.; Di Martino, M.; Sorndech, W.; Mamone, G.; Vincze, E.; Buillon, G.; Nielsen, D.S.; Wiese, M.; et al. Low glycaemic index foods from wild barley and amylose-only barley lines. J. Funct. Foods 2018, 40, 408–416. [Google Scholar] [CrossRef]
- Cieplak, T.; Wiese, M.; Nielsen, S.; Van De Wiele, T.; Van Den Berg, F.; Nielsen, D.S. The Smallest Intestine (TSI) - A low volume in vitro model of the small intestine with increased throughput. FEMS Microbiol. Lett. 2018, 365, fny231. [Google Scholar] [CrossRef]
- Malde, A.K.; Zuo, L.; Breeze, M.; Stroet, M.; Poger, D.; Nair, P.C.; Oostenbrink, C.; Mark, A.E. An Automated force field Topology Builder (ATB) and repository: Version 1.0. J. Chem. Theory Comput. 2011, 7, 4026–4037. [Google Scholar] [CrossRef]
- Koziara, K.B.; Stroet, M.; Malde, A.K.; Mark, A.E. Testing and validation of the Automated Topology Builder (ATB) version 2.0: Prediction of hydration free enthalpies. J. Comput. Aided. Mol. Des. 2014, 28, 221–233. [Google Scholar] [CrossRef] [PubMed]
- Schmid, N.; Eichenberger, A.P.; Choutko, A.; Riniker, S.; Winger, M.; Mark, A.E.; Van Gunsteren, W.F. Definition and testing of the GROMOS force-field versions 54A7 and 54B7. Eur. Biophys. J. 2011, 40, 843–856. [Google Scholar] [CrossRef] [PubMed]
- Geerke, D.P.; Van Gunsteren, W.F. Force field evaluation for biomolecular simulation: Fenthalpies of solvation of polar and apolar compounds in various solvents. ChemPhysChem 2006, 7, 671–678. [Google Scholar] [CrossRef] [PubMed]
- Berendsen, H.J.C.; Postma, J.P.M.; van Gunsteren, W.F.; Hermans, J. Interaction models for water in relation to protein hydration. In Intermolecular Forces: Proceedings of the Fourteenth Jerusalem Symposium on Quantum Chemistry and Biochemistry Held in Jerusalem, Israel, April 13-16, 1981; Pullman, B., Ed.; Springer: Dordrecht, The Netherlands, 1981; pp. 331–342. [Google Scholar]
- Darden, T.; York, D.; Pedersen, L. Particle mesh Ewald: An N·log(N) method for Ewald sums in large systems. J. Chem. Phys. 1993, 98, 10089–10092. [Google Scholar] [CrossRef]
- Essmann, U.; Perera, L.; Berkowitz, M.L.; Darden, T.; Lee, H.; Pedersen, L.G. A smooth particle mesh Ewald method. J. Chem. Phys. 1995, 103, 8577–8593. [Google Scholar] [CrossRef]
- Gordhan, D.; Swainson, S.M.E.; Pearce, A.K.; Styliari, I.D.; Lovato, T.; Burley, J.C.; Garnett, M.C.; Taresco, V. Poly(glycerol adipate): From a functionalized nanocarrier to a polymeric-prodrug matrix to create amorphous solid dispersions. J. Pharm. Sci. 2019. [Google Scholar] [CrossRef]
- Abushrida, A.; Elhuni, I.; Taresco, V.; Marciani, L.; Stolnik, S.; Garnett, M.C. A simple and efficient method for polymer coating of iron oxide nanoparticles. J. Drug Deliv. Sci. Technol. 2020, 55, 101460. [Google Scholar] [CrossRef]
- Swainson, S.M.E.; Taresco, V.; Pearce, A.K.; Clapp, L.H.; Ager, B.; McAllister, M.; Bosquillon, C.; Garnett, M.C. Exploring the enzymatic degradation of poly(glycerol adipate). Eur. J. Pharm. Biopharm. 2019, 142, 377–386. [Google Scholar] [CrossRef]
- Madeira Do, J.; Foralosso, R.; Yilmaz, G.; Mastrotto, F.; King, P.J.S.; Xerri, R.M.; He, Y.; Van Der Walle, C.F.; Fernandez-Trillo, F.; Laughton, C.A.; et al. Poly(triazolyl methacrylate) glycopolymers as potential targeted unimolecular nanocarriers. Nanoscale 2019, 11, 21155. [Google Scholar] [CrossRef]
- Mackenzie, R.; Booth, J.; Alexander, C.; Garnett, M.C.; Laughton, C.A. Multiscale modeling of drug-polymer nanoparticle assembly identifies parameters influencing drug encapsulation efficiency. J. Chem. Theory Comput. 2015, 11, 2705–2713. [Google Scholar] [CrossRef]
- Turpin, E.R.; Taresco, V.; Al-Hachami, W.A.; Booth, J.; Treacher, K.; Tomasi, S.; Alexander, C.; Burley, J.; Laughton, C.A.; Garnett, M.C. In silico screening for solid dispersions: The trouble with solubility parameters and χFH. Mol. Pharm. 2018, 15, 4654–4667. [Google Scholar] [CrossRef] [PubMed]
- Brattain, M.G.; Fine, W.D.; Khaled, F.M.; Thompson, J.; Brattain, D.E. Heterogeneity of malignant cells from a human colonic carcinoma. Cancer Res. 1981, 41, 1751–1756. [Google Scholar] [PubMed]
- Leibovitz, A.; Stinson, J.C.; McCombs, W.B.; McCoy, C.E.; Mazur, K.C.; Mabry, N.D. Classification of human colorectal adenocarcinoma cell lines. Cancer Res. 1976, 36, 4562–4569. [Google Scholar] [PubMed]
- Ahmed, D.; Eide, P.W.; Eilertsen, I.A.; Danielsen, S.A.; Eknæs, M.; Hektoen, M.; Lind, G.E.; Lothe, R.A. Epigenetic and genetic features of 24 colon cancer cell lines. Oncogenesis 2013, 2, e71. [Google Scholar] [CrossRef]
- Anderson, G.R.; Winter, P.S.; Lin, K.H.; Nussbaum, D.P.; Cakir, M.; Stein, E.M.; Soderquist, R.S.; Crawford, L.; Leeds, J.C.; Newcomb, R.; et al. A Landscape of therapeutic cooperativity in KRAS mutant cancers reveals principles for controlling tumor evolution. Cell Rep. 2017, 20, 999–1015. [Google Scholar] [CrossRef]
- van Zandwijk, N.; Mathy, A.; Boerrigter, L.; Ruijter, H.; Tielen, I.; de Jong, D.; Baas, P.; Burgers, S.; Nederlof, P. EGFR and KRAS mutations as criteria for treatment with tyrosine kinase inhibitors: Retro- and prospective observations in non-small-cell lung cancer. Ann. Oncol. 2007, 18, 99–103. [Google Scholar] [CrossRef]
- Lee, M.; Young Kim, S.; Kim, J.; Kim, H.-S.; Kim, S.-M.; Kim, E.J. Mitogen-activated protein kinase phosphatase-1 inhibition and sustained extracellular signal-regulated kinase 1/2 activation in camptothecin-induced human colon cancer cell death. Cancer Biol. Ther. 2013, 14, 1007–1015. [Google Scholar] [CrossRef][Green Version]
- Washington, N.; Washington, C.; Wilson, C.W. Physiological Pharmaceutics, Barriers to Drug Absorption, 2nd ed.; Taylor and Francis Series in Pharmaceutics; CRC Press: Boca Raton, FL, USA, 2000. [Google Scholar]


| Size (nm) | PdI | Z-potential (mV) | |
|---|---|---|---|
| PGA | 175.0 ± 1.9 | 0.033 | −12.0 ± 0.5 |
| PGA-SI214 | 172.3 ± 1.8 | 0.030 | −15.2 ± 0.9 |
| PGA-CPT | 184.1 ± 2.0 | 0.027 | −17.6 ± 0.5 |
| SA01 | SA02 | EC07 | EC19 | |
|---|---|---|---|---|
| SI214 1 | n.d. | n.d. | n.d. | n.d. |
| PGA-SI214 | 500 µg/mL | 500 µg/mL | 250 µg/mL | 250 µg/mL |
| CPT 1 | n.d. | n.d. | n.d. | n.d. |
| PGA-CPT | 500 µg/mL | 500 µg/mL | 250 µg/mL | 250 µg/mL |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vestri, A.; Pearce, A.K.; Cavanagh, R.; Styliari, I.D.; Sanders, C.; Couturaud, B.; Schenone, S.; Taresco, V.; Jakobsen, R.R.; Howdle, S.M.; et al. Starch/Poly(glycerol-adipate) Nanocomposites: A Novel Oral Drug Delivery Device. Coatings 2020, 10, 125. https://doi.org/10.3390/coatings10020125
Vestri A, Pearce AK, Cavanagh R, Styliari ID, Sanders C, Couturaud B, Schenone S, Taresco V, Jakobsen RR, Howdle SM, et al. Starch/Poly(glycerol-adipate) Nanocomposites: A Novel Oral Drug Delivery Device. Coatings. 2020; 10(2):125. https://doi.org/10.3390/coatings10020125
Chicago/Turabian StyleVestri, Ambra, Amanda K. Pearce, Robert Cavanagh, Ioanna D. Styliari, Carlos Sanders, Benoit Couturaud, Silvia Schenone, Vincenzo Taresco, Rasmus R. Jakobsen, Steven M. Howdle, and et al. 2020. "Starch/Poly(glycerol-adipate) Nanocomposites: A Novel Oral Drug Delivery Device" Coatings 10, no. 2: 125. https://doi.org/10.3390/coatings10020125
APA StyleVestri, A., Pearce, A. K., Cavanagh, R., Styliari, I. D., Sanders, C., Couturaud, B., Schenone, S., Taresco, V., Jakobsen, R. R., Howdle, S. M., Musumeci, F., & Sagnelli, D. (2020). Starch/Poly(glycerol-adipate) Nanocomposites: A Novel Oral Drug Delivery Device. Coatings, 10(2), 125. https://doi.org/10.3390/coatings10020125








