Antibiotic Resistance Characteristics of Pseudomonas aeruginosa Isolated from Keratitis in Australia and India
Abstract
:1. Introduction
2. Results
2.1. Antibiotic Susceptibility
2.2. General Features of the Genomes
2.3. Acquired Resistance Genes
2.4. Possession of exoU and Mutations in the DNA Mismatch Repair System
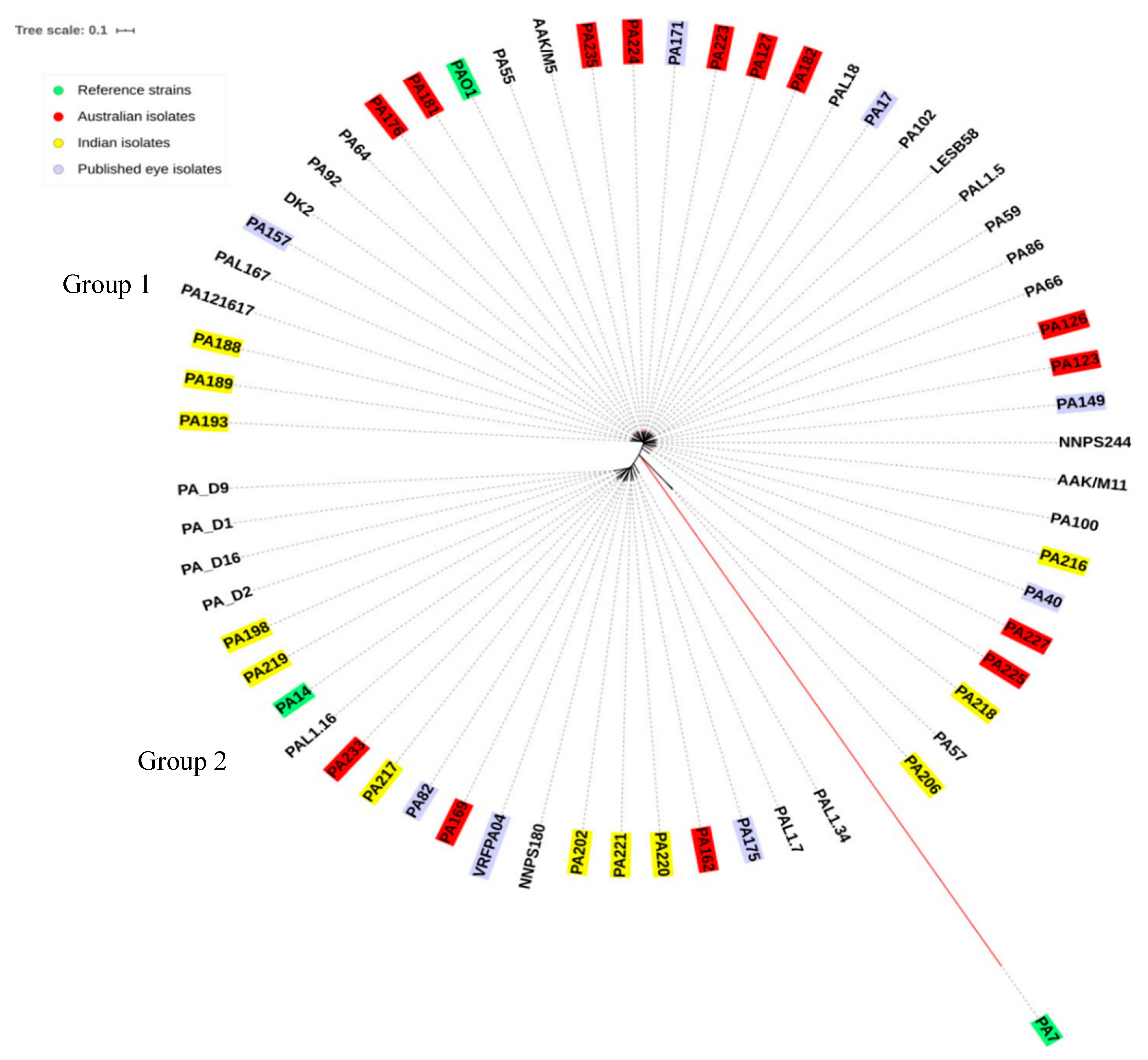
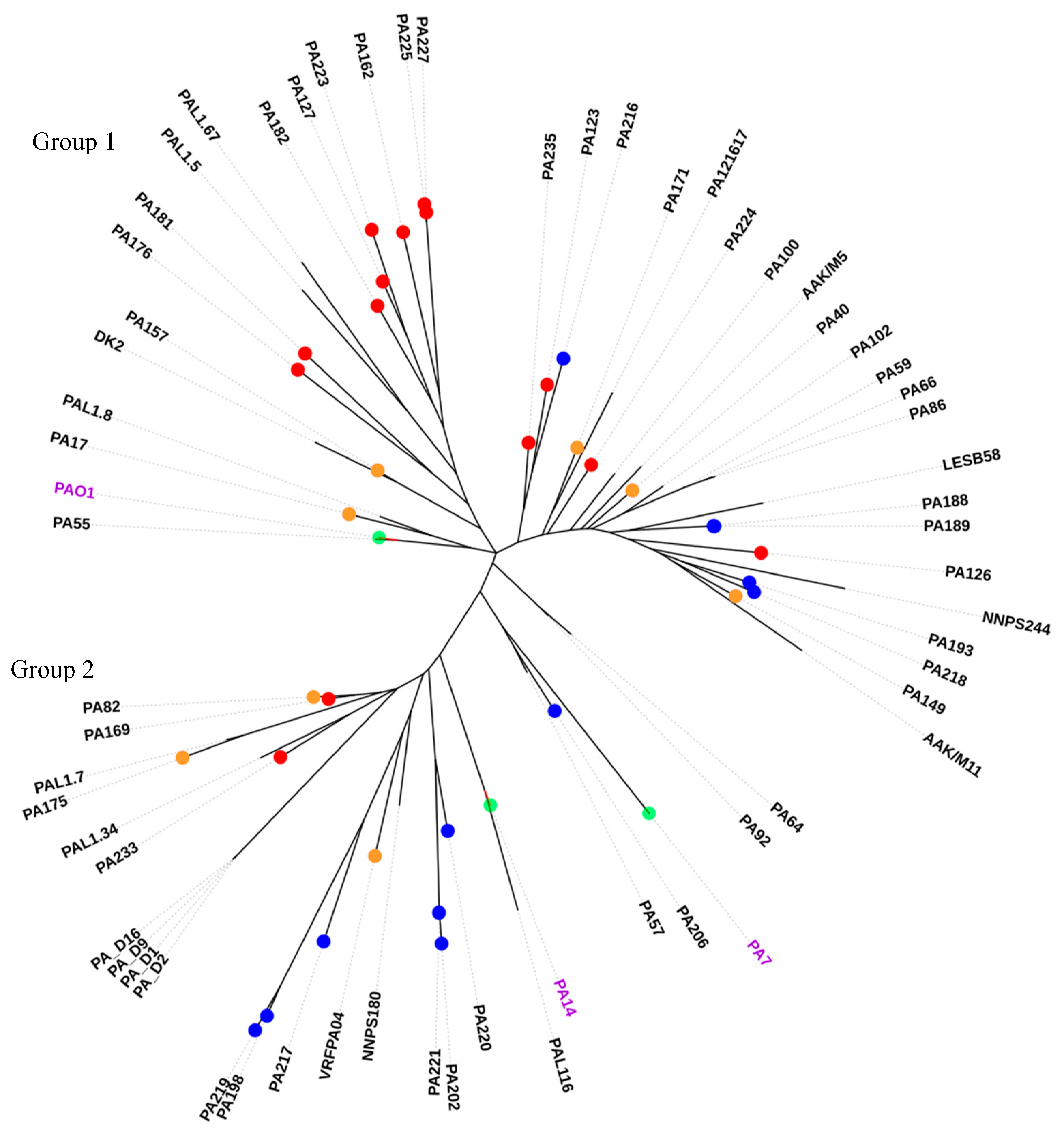
2.5. Sequence Type Analysis and Phylogenetics
3. Discussion
4. Materials and Methods
4.1. P. aeruginosa Strains and Susceptibility Testing
4.2. Genomic Sequencing
4.3. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Richards, M.J.; Edwards, J.R.; Culver, D.H.; Gaynes, R.P. Nosocomial infections in medical intensive care units in the United States. National Nosocomial Infections Surveillance System. Crit. Care Med. 1999, 27, 887–892. [Google Scholar] [CrossRef] [PubMed]
- Abjani, F.; Khan, N.A.; Jung, S.Y.; Siddiqui, R. Status of the effectiveness of contact lens disinfectants in Malaysia against keratitis-causing pathogens. Exp. Parasitol. 2017, 183, 187–193. [Google Scholar] [CrossRef] [PubMed]
- Stapleton, F.; Carnt, N. Contact lens-related microbial keratitis: How have epidemiology and genetics helped us with pathogenesis and prophylaxis. Eye (Lond.) 2012, 26, 185–193. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Trouillet, J.L.; Vuagnat, A.; Combes, A.; Kassis, N.; Chastre, J.; Gibert, C. Pseudomonas aeruginosa ventilator-associated pneumonia: Comparison of episodes due to piperacillin-resistant versus piperacillin-susceptible organisms. Clin. Infect. Dis. 2002, 34, 1047–1054. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Green, M.; Apel, A.; Stapleton, F. Risk factors and causative organisms in microbial keratitis. Cornea 2008, 27, 22–27. [Google Scholar] [CrossRef]
- Hooi, S.H.; Hooi, S.T. Culture-proven bacterial keratitis in a Malaysian general hospital. Med. J. Malays. 2005, 60, 614–623. [Google Scholar]
- Parmar, P.; Salman, A.; Kalavathy, C.M.; Kaliamurthy, J.; Thomas, P.A.; Jesudasan, C.A. Microbial keratitis at extremes of age. Cornea 2006, 25, 153–158. [Google Scholar] [CrossRef]
- Sharma, N.; Sinha, R.; Singhvi, A.; Tandon, R. Pseudomonas keratitis after laser in situ keratomileusis. J. Cataract. Refract. Surg. 2006, 32, 519–521. [Google Scholar] [CrossRef]
- Chatterjee, S.; Agrawal, D. Multi-drug resistant Pseudomonas aeruginosa keratitis and its effective treatment with topical colistimethate. Indian J. Ophthalmol. 2016, 64, 153–157. [Google Scholar] [CrossRef]
- Livermore, D.M. beta-Lactamases in laboratory and clinical resistance. Clin. Microbiol. Rev. 1995, 8, 557–584. [Google Scholar] [CrossRef]
- Li, X.-Z.; Ma, D.; Livermore, D.M.; Nikaido, H. Role of efflux pump (s) in intrinsic resistance of Pseudomonas aeruginosa: Active efflux as a contributing factor to beta-lactam resistance. Antimicrob. Agents Chemother. 1994, 38, 1742–1752. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Poole, K. Pseudomonas aeruginosa: Resistance to the max. Front. Microbiol. 2011, 2, 65. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hocquet, D.; Vogne, C.; El Garch, F.; Vejux, A.; Gotoh, N.; Lee, A.; Lomovskaya, O.; Plésiat, P. MexXY-OprM efflux pump is necessary for adaptive resistance of Pseudomonas aeruginosa to aminoglycosides. Antimicrob. Agents Chemother. 2003, 47, 1371–1375. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Livermore, D.M. Multiple mechanisms of antimicrobial resistance in Pseudomonas aeruginosa: Our worst nightmare? Clin. Infect. Dis. 2002, 34, 634–640. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lomholt, J.A.; Kilian, M. Ciprofloxacin susceptibility of Pseudomonas aeruginosa isolates from keratitis. Br. J. Ophthalmol. 2003, 87, 1238–1240. [Google Scholar] [CrossRef] [Green Version]
- Willcox, M.D. Review of resistance of ocular isolates of Pseudomonas aeruginosa and staphylococci from keratitis to ciprofloxacin, gentamicin and cephalosporins. Clin. Exp. Optom. 2011, 94, 161–168. [Google Scholar] [CrossRef]
- Poonsuk, K.; Tribuddharat, C.; Chuanchuen, R. Aminoglycoside resistance mechanisms in Pseudomonas aeruginosa isolates from non-cystic fibrosis patients in Thailand. Can. J. Microbiol. 2013, 59, 51–56. [Google Scholar] [CrossRef]
- Diver, J.M.; Schollaardt, T.; Rabin, H.R.; Thorson, C.; Bryan, L.E. Persistence mechanisms in Pseudomonas aeruginosa from cystic fibrosis patients undergoing ciprofloxacin therapy. Antimicrob. Agents Chemother. 1991, 35, 1538–1546. [Google Scholar] [CrossRef] [Green Version]
- Guzvinec, M.; Izdebski, R.; Butic, I.; Jelic, M.; Abram, M.; Koscak, I.; Baraniak, A.; Hryniewicz, W.; Gniadkowski, M.; Andrasevic, A.T. Sequence types 235, 111, and 132 predominate among multidrug-resistant Pseudomonas aeruginosa clinical isolates in Croatia. Antimicrob. Agents Chemother. 2014, 58, 6277–6283. [Google Scholar] [CrossRef] [Green Version]
- Phillips, R.M.; Six, D.A.; Dennis, E.A.; Ghosh, P. In vivo phospholipase activity of the Pseudomonas aeruginosa cytotoxin ExoU and protection of mammalian cells with phospholipase A2 inhibitors. J. Biol. Chem. 2003, 278, 41326–41332. [Google Scholar] [CrossRef] [Green Version]
- Finck-Barbançon, V.; Goranson, J.; Zhu, L.; Sawa, T.; Wiener-Kronish, J.P.; Fleiszig, S.M.J.; Wu, C.; Mende-Mueller, L.; Frank, D.W. ExoU expression by Pseudomonas aeruginosa correlates with acute cytotoxicity and epithelial injury. Mol. Microbiol. 1997, 25, 547–557. [Google Scholar] [CrossRef] [PubMed]
- Subedi, D.; Vijay, A.K.; Willcox, M. Overview of mechanisms of antibiotic resistance in Pseudomonas aeruginosa: An ocular perspective. Clin. Exp. Optom. 2018, 101, 162–171. [Google Scholar] [CrossRef] [Green Version]
- Maciá, M.D.; Blanquer, D.; Togores, B.; Sauleda, J.; Pérez, J.L.; Oliver, A. Hypermutation is a key factor in development of multiple-antimicrobial resistance in Pseudomonas aeruginosa strains causing chronic lung infections. Antimicrob. Agents Chemother. 2005, 49, 3382–3386. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Modrich, P. Mechanisms and biological effects of mismatch repair. Annu. Rev. Genet. 1991, 25, 229–253. [Google Scholar] [CrossRef]
- Oliver, A.; Baquero, F.; Blazquez, J. The mismatch repair system (mutS, mutL and uvrD genes) in Pseudomonas aeruginosa: Molecular characterization of naturally occurring mutants. Mol. Microbiol. 2002, 43, 1641–1650. [Google Scholar] [CrossRef]
- Alsan, M.; Schoemaker, L.; Eggleston, K.; Kammili, N.; Kolli, P.; Bhattacharya, J. Out-of-pocket health expenditures and antimicrobial resistance in low-income and middle-income countries: An economic analysis. Lancet Infect. Dis. 2015, 15, 1203–1210. [Google Scholar] [CrossRef] [Green Version]
- Sánchez, M.; Sivaraman, S. News Media Reporting of Antimicrobial Resistance in Latin America and India. In Antimicrobial Resistance in Developing Countries; Sosa, A.d.J., Byarugaba, D.K., Amábile-Cuevas, C.F., Hsueh, P.-R., Kariuki, S., Okeke, I.N., Eds.; Springer: New York, NY, USA, 2010; pp. 525–537. [Google Scholar]
- Van Boeckel, T.P.; Gandra, S.; Ashok, A.; Caudron, Q.; Grenfell, B.T.; Levin, S.A.; Laxminarayan, R. Global antibiotic consumption 2000 to 2010: An analysis of national pharmaceutical sales data. Lancet Infect. Dis. 2014, 14, 742–750. [Google Scholar] [CrossRef]
- Khan, M.; Stapleton, F.; Willcox, M.D.P. Susceptibility of contact lens-related Pseudomonas aeruginosa keratitis isolates to multipurpose disinfecting solutions, disinfectants, and antibiotics. Transl. Vis. Sci. Technol. 2020, 9, 2. [Google Scholar] [CrossRef] [Green Version]
- Choy, M.H.; Stapleton, F.; Willcox, M.D.P.; Zhu, H. Comparison of virulence factors in Pseudomonas aeruginosa strains isolated from contact lens- and non-contact lens-related keratitis. J. Med. Microbiol. 2008, 57, 1539–1546. [Google Scholar] [CrossRef] [PubMed]
- Subedi, D.; Vijay, A.K.; Kohli, G.S.; Rice, S.A.; Willcox, M. Comparative genomics of clinical strains of Pseudomonas aeruginosa strains isolated from different geographic sites. Sci. Rep. 2018, 8, 15668. [Google Scholar] [CrossRef] [Green Version]
- Porter, G.; Grills, N. Medication misuse in India: A major public health issue in India. J. Public Health (Oxf.) 2016, 38, e150–e157. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wilhelmus, K.R.; Abshire, R.L.; Schlech, B.A. Influence of fluoroquinolone susceptibility on the therapeutic response of fluoroquinolone-treated bacterial keratitis. Arch. Ophthalmol. 2003, 121, 1229–1233. [Google Scholar] [CrossRef] [PubMed]
- Keay, L.; Edwards, K.; Naduvilath, T.; Taylor, H.R.; Snibson, G.R.; Forde, K.; Stapleton, F. Microbial keratitis predisposing factors and morbidity. Ophthalmology 2006, 113, 109–116. [Google Scholar] [CrossRef] [PubMed]
- Keay, L.; Edwards, K.; Dart, J.; Stapleton, F. Grading contact lens-related microbial keratitis: Relevance to disease burden. Optom. Vis. Sci. 2008, 85, 531–537. [Google Scholar] [CrossRef]
- Murugan, N.; Malathi, J.; Umashankar, V.; Madhavan, H.N. Unraveling genomic and phenotypic nature of multidrug-resistant (MDR) Pseudomonas aeruginosa VRFPA04 isolated from keratitis patient. Microbiol. Res. 2016, 193, 140–149. [Google Scholar]
- Belotti, P.T.; Thabet, L.; Laffargue, A.; Andre, C.; Coulange-Mayonnove, L.; Arpin, C.; Messadi, A.; M’zali, F.; Quentin, C.; Dubois, V. Description of an original integron encompassing blaVIM-2, qnrVC1 and genes encoding bacterial group II intron proteins in Pseudomonas aeruginosa. J. Antimicrob. Chemother. 2015, 70, 2237–2240. [Google Scholar] [CrossRef] [Green Version]
- Kocsis, B.; Toth, A.; Gulyas, D.; Ligeti, B.; Katona, K.; Rokusz, L.; Szabo, D. Acquired qnrVC1 and blaNDM-1 resistance markers in an international high-risk Pseudomonas aeruginosa ST773 clone. J. Med. Microbiol. 2019, 68, 336–338. [Google Scholar] [CrossRef]
- Tauch, A.; Schluter, A.; Bischoff, N.; Goesmann, A.; Meyer, F.; Puhler, A. The 79,370-bp conjugative plasmid pB4 consists of an IncP-1beta backbone loaded with a chromate resistance transposon, the strA-strB streptomycin resistance gene pair, the oxacillinase gene bla(NPS-1), and a tripartite antibiotic efflux system of the resistance-nodulation-division family. Mol. Genet. Genom. 2003, 268, 570–584. [Google Scholar]
- Sundin, G.W.; Bender, C.L. Dissemination of the strA-strB streptomycin-resistance genes among commensal and pathogenic bacteria from humans, animals, and plants. Mol. Ecol. 1996, 5, 133–143. [Google Scholar] [CrossRef]
- Zhu, H.; Conibear, T.C.; Bandara, R.; Aliwarga, Y.; Stapleton, F.; Willcox, M.D. Type III secretion system-associated toxins, proteases, serotypes, and antibiotic resistance of Pseudomonas aeruginosa isolates associated with keratitis. Curr. Eye Res. 2006, 31, 297–306. [Google Scholar] [CrossRef]
- Lakkis, C.; Fleiszig, S.M. Resistance of Pseudomonas aeruginosa isolates to hydrogel contact lens disinfection correlates with cytotoxic activity. J. Clin. Microbiol. 2001, 39, 1477–1486. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Borkar, D.S.; Acharya, N.R.; Leong, C.; Lalitha, P.; Srinivasan, M.; Oldenburg, C.E.; Cevallos, V.; Lietman, T.M.; Evans, D.J.; Fleiszig, S.M. Cytotoxic clinical isolates of Pseudomonas aeruginosa identified during the Steroids for Corneal Ulcers Trial show elevated resistance to fluoroquinolones. BMC Ophthalmol. 2014, 14, 54. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tiraby, J.G.; Fox, M.S. Marker discrimination in transformation and mutation of Pneumococcus. Proc. Natl. Acad. Sci. USA 1973, 70, 3541–3545. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Acharya, S.; Foster, P.L.; Brooks, P.; Fishel, R. The coordinated functions of the E. coli MutS and MutL proteins in mismatch repair. Mol. Cell 2003, 12, 233–246. [Google Scholar] [CrossRef]
- Rayssiguier, C.; Thaler, D.S.; Radman, M. The barrier to recombination between Escherichia coli and Salmonella typhimurium is disrupted in mismatch-repair mutants. Nature 1989, 342, 396–401. [Google Scholar] [CrossRef]
- Chopra, I.; O’Neill, A.J.; Miller, K. The role of mutators in the emergence of antibiotic-resistant bacteria. Drug Resist. Updates 2003, 6, 137–145. [Google Scholar] [CrossRef]
- Sato, H.; Frank, D.W.; Hillard, C.J.; Feix, J.B.; Pankhaniya, R.R.; Moriyama, K.; Finck-Barbançon, V.; Buchaklian, A.; Lei, M.; Long, R.M.; et al. The mechanism of action of the Pseudomonas aeruginosa-encoded type III cytotoxin, ExoU. EMBO J. 2003, 22, 2959–2969. [Google Scholar] [CrossRef] [Green Version]
- Kulasekara, B.R.; Kulasekara, H.D.; Wolfgang, M.C.; Stevens, L.; Frank, D.W.; Lory, S. Acquisition and evolution of the exoU locus in Pseudomonas aeruginosa. J. Bacteriol. 2006, 188, 4037–4050. [Google Scholar] [CrossRef] [Green Version]
- Freschi, L.; Jeukens, J.; Kukavica-Ibrulj, I.; Boyle, B.; Dupont, M.J.; Laroche, J.; Larose, S.; Maaroufi, H.; Fothergill, J.L.; Moore, M.; et al. Clinical utilization of genomics data produced by the international Pseudomonas aeruginosa consortium. Front. Microbiol. 2015, 6, 1036. [Google Scholar] [CrossRef] [Green Version]
- Winstanley, C.; Langille, M.G.; Fothergill, J.L.; Kukavica-Ibrulj, I.; Paradis-Bleau, C.; Sanschagrin, F.; Thomson, N.R.; Winsor, G.L.; Quail, M.A.; Lennard, N.; et al. Newly introduced genomic prophage islands are critical determinants of in vivo competitiveness in the Liverpool Epidemic Strain of Pseudomonas aeruginosa. Genome Res. 2009, 19, 12–23. [Google Scholar] [CrossRef] [Green Version]
- Klockgether, J.; Cramer, N.; Wiehlmann, L.; Davenport, C.F.; Tummler, B. Pseudomonas aeruginosa genomic structure and diversity. Front. Microbiol. 2011, 2, 150. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Roy, P.H.; Tetu, S.G.; Larouche, A.; Elbourne, L.; Tremblay, S.; Ren, Q.; Dodson, R.; Harkins, D.; Shay, R.; Watkins, K.; et al. Complete genome sequence of the multiresistant taxonomic outlier Pseudomonas aeruginosa PA7. PLoS ONE 2010, 5, e8842. [Google Scholar] [CrossRef] [PubMed]
- Freschi, L.; Vincent, A.T.; Jeukens, J.; Emond-Rheault, J.G.; Kukavica-Ibrulj, I.; Dupont, M.J.; Charette, S.J.; Boyle, B.; Levesque, R.C. The Pseudomonas aeruginosa pan-genome provides new insights on its population structure, horizontal gene transfer, and pathogenicity. Genome Biol. Evol. 2019, 11, 109–120. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ozer, E.A.; Allen, J.P.; Hauser, A.R. Characterization of the core and accessory genomes of Pseudomonas aeruginosa using bioinformatic tools Spine and AGEnt. BMC Genom. 2014, 15, 737. [Google Scholar] [CrossRef] [Green Version]
- Valot, B.; Guyeux, C.; Rolland, J.Y.; Mazouzi, K.; Bertrand, X.; Hocquet, D. What it takes to be a Pseudomonas aeruginosa? The core genome of the opportunistic pathogen updated. PLoS ONE 2015, 10, e0126468. [Google Scholar] [CrossRef] [PubMed]
- Wolfgang, M.C.; Kulasekara, B.R.; Liang, X.; Boyd, D.; Wu, K.; Yang, Q.; Miyada, C.G.; Lory, S. Conservation of genome content and virulence determinants among clinical and environmental isolates of Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 2003, 100, 8484–8489. [Google Scholar] [CrossRef] [Green Version]
- Smits, T.H.M. The importance of genome sequence quality to microbial comparative genomics. BMC Genom. 2019, 20, 662. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [Green Version]
- EUCAST. Breakpoint Tables for Interpretation of MICs and Zone Diameters; Version 6.0; The European Committee on Antimicrobial Susceptibility Testing: Växjö, Sweden, 2019. [Google Scholar]
- Patel, J.B.; Cockerill, F.R.; Alder, J.; Bradford, P.A.; Eliopoulos, G.M.; Hardy, D.J.; Hindler, J.A.; Jenkins, S.G.; Lewis, J.S.; Miller, L.A.; et al. Performance Standards for Antimicrobial Susceptibility Testing; Twenty-Fourth Informational Supplement; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2014. [Google Scholar]
- Nurk, S.; Bankevich, A.; Antipov, D.; Gurevich, A.; Korobeynikov, A.; Lapidus, A.; Prjibelsky, A.; Pyshkin, A.; Sirotkin, A.; Sirotkin, Y.; et al. Assembling Genomes and Mini-metagenomes from Highly Chimeric Reads. In Research in Computational Molecular Biology RECOMB 2013; Deng, M., Jiang, R., Sun, F., Zhang, X., Eds.; Lecture Notes in Computer Science; Springer: Berlin/Heidelberg, Germany, 2013. [Google Scholar]
- Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics (Oxf. Engl.) 2014, 30, 2068–2069. [Google Scholar] [CrossRef]
- Page, A.J.; Cummins, C.A.; Hunt, M.; Wong, V.K.; Reuter, S.; Holden, M.T.; Fookes, M.; Falush, D.; Keane, J.A.; Parkhill, J. Roary: Rapid large-scale prokaryote pan genome analysis. Bioinformatics 2015, 31, 3691–3693. [Google Scholar] [CrossRef]
- Treangen, T.J.; Ondov, B.D.; Koren, S.; Phillippy, A.M. The Harvest suite for rapid core-genome alignment and visualization of thousands of intraspecific microbial genomes. Genome Biol. 2014, 15, 524. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zankari, E.; Hasman, H.; Cosentino, S.; Vestergaard, M.; Rasmussen, S.; Lund, O.; Aarestrup, F.M.; Larsen, M.V. Identification of acquired antimicrobial resistance genes. J. Antimicrob. Chemother. 2012, 67, 2640–2644. [Google Scholar] [CrossRef] [PubMed]
- Seeman, T. Snippy: Fast Bacterial Variant Calling from NGS Reads; GitHub Australia: Melbourne, Australia, 2015. [Google Scholar]
| Strain Number | Fluoroquinolones * | Aminoglycosides | β-Lactams | Poly-Peptide | ||||
|---|---|---|---|---|---|---|---|---|
| 2nd Generation | 3rd Generation | Penicillin 4th Generation | Carba-Penem | Cephalosporin 3rd Generation | ||||
|
Cipro µg/mL ≤1, 2, ≥4 # |
Levo µg/mL ≤2, 4, ≥8 |
Genta µg/mL ≤4, 8, ≥16 |
Tobra µg/mL ≤4, 8, ≥16 |
Pipera µg/mL ≤16 |
Imi µg/mL ≤2, 4, ≥8 |
Ceftaz µg/mL ≤8, 16, ≥32 |
PMB µg/mL ≤2, 4, ≥8 | |
| MIC/ MBC | MIC/ MBC | MIC/ MBC | MIC/ MBC | MIC/ MBC | MIC/ MBC | MIC/ MBC | MIC/ MBC | |
| 123 | 1/1 | 1/1 | 0.25/0.5 | 4/4 | 8/16 | 4(I)/8 | 2/2 | 1280(R)/1280 |
| 126 | 0.5/1 | 0.5/1 | 0.5/1 | 0.25/0.5 | 8/16 | 8(R)/16 | 128(R)/256 | 1/1 |
| 127 | 1/2 | 0.25/1 | 2/4 | 32(R)/128 | 4/16 | 4(I)/8 | 128(R)/256 | 0.5/1 |
| 162 | 0.5/1 | 0.5/1 | 0.25/0.5 | 0.25/1 | 8/8 | 4(I)/4 | 2/4 | 0.25/0.5 |
| 169 | 2(I)/4 | 0.25/0.5 | 0.25/0.5 | 0.25/0.5 | 4/8 | 2/4 | 1/2 | 0.25/0.25 |
| 176 | 0.5/1 | 0.25/0.5 | 0.25/0.5 | 0.25/0.5 | 4/8 | 2/8 | 2/4 | 0.25/0.5 |
| 181 | 1/4 | 0.25/0.5 | 0.25/0.5 | 0.25/0.5 | 32(R)/64 | 4(I)/8 | 16(I)/32 | 0.5/1 |
| 182 | 1/2 | 0.25/0.5 | 0.25/0.5 | 0.25/0.5 | 4/8 | 8(R)/16 | 1/2 | 0.25/0.5 |
| 223 | 64(R)/128 | 1/2 | 0.5/1 | 0.5/1 | 160(R)/320 | 1/2 | 16(I)/32 | 2/4 |
| 224 | 16(R)/32 | 1/2 | 0.25/0.5 | 0.25/0.5 | 8/16 | 64(R)/128 | 16(I)/32 | 1/2 |
| 225 | 64(R)/128 | 16(R)/32 | 0.5/2 | 1/2 | 16/32 | 64(R)/128 | 8/16 | 0.25/0.5 |
| 227 | 64(R)/128 | 64(R)/128 | 0.5/1 | 0.25/1 | 16/32 | 16(R)/32 | 16(I)/32 | 0.25/0.5 |
| 233 | 8(R)/16 | 1/2 | 1/2 | 105/1 | 16/32 | 4(I)/8 | 160(I)/320 | 0.5/1 |
| 235 | 16(R)/32 | 0.5/1 | 2/4 | 0.5/1 | 64(R)/128 | 4(I)/8 | 64(I)/128 | 0.25 |
| 188 | 2(I)/4 | 1/2 | 0.5/1 | 32(R)/64 | 16/65 | 0.5/1 | 4/8 | 2/4 |
| 189 | 0.25/1 | 1/2 | 0.25/0.5 | 16(R)/32 | 4/8 | 2/1 | 8/16 | 2/4 |
| 193 | 1/1 | 0.25/1 | 0.25/0.25 | 0.25/0.5 | 4/8 | 2/4 | 2/2 | 0.5/1 |
| 198 | 1280(R)/2560 | 320(R)/1280 | 2560(R)/5120 | 16(R)/16 | 8/8 | 1/2 | 8/8 | 4(I)/4 |
| 202 | 640(R)/1280 | 320(R)/640 | 8(I)/32 | 320(R)/640 | 16/64 | 8(R)/32 | 8/32 | 0.25/0.25 |
| 206 | 1/1 | 0.5/0.5 | 1/1 | 0.25/0.5 | 8/8 | 2/4 | 2/4 | 0.25/0.5 |
| 216 | 64(R)/128 | 4 (I)/8 | 1/2 | 0.5/2 | 160(R)/320 | 16(R)/32 | 64(R)/128 | 64(R)/128 |
| 217 | 64(R)/128 | 32(R)/64 | 1/2 | 1/2 | 64(R)/128 | 8(R)/16 | 32(R)/64 | 0.25/1 |
| 218 | 8(R)/16 | 1/2 | 0.5/1 | 0.5/1 | 160(R)/320 | 8(R)/16 | 64(R)/128 | 1/4 |
| 219 | ≥5120(R)/≥5120 | 640(R)/1280 | ≥5120(R)/≥5120 | 1280(R)/2560 | 2560(R)/5120 | 40(R)/80 | 16(I)/32 | 0.25/1 |
| 220 | 2(I)/4 | 0.25/0.5 | 0.5/1 | 0.5/1 | 0.25/0.5 | 8(R)/16 | 160(R)/320 | 8(R)/16 |
| 221 | 2560(R)/5120 | 2560(R)/5120 | 2560(R)/5120 | 2560(R)/5120 | 64(R)/128 | 16(R)/32 | 32(R)/64 | 0.25/1 |
| Genes | Australian Isolates | Indian Isolates | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 123 | 126 | 127 | 162 | 169 | 176 | 181 | 182 | 223 | 224 | 225 | 227 | 233 | 235 | 188 | 189 | 193 | 198 | 202 | 206 | 216 | 217 | 218 | 219 | 220 | 221 | |
| Aminoglycoside resistance genes | ||||||||||||||||||||||||||
| aph(3’)-IIb | ||||||||||||||||||||||||||
| aph(6)-Id | l; | |||||||||||||||||||||||||
| rmtD2 | ||||||||||||||||||||||||||
| rmtB | ||||||||||||||||||||||||||
| aph(3’)-VI | ||||||||||||||||||||||||||
| aph(3’)-lIb | ||||||||||||||||||||||||||
| aph(3’’)-Ib | ||||||||||||||||||||||||||
| aac(6’)-Ib3 | ||||||||||||||||||||||||||
| aac(3)-IId | ||||||||||||||||||||||||||
| aadA1 | ||||||||||||||||||||||||||
| aac(6’)-Ib-cr | ||||||||||||||||||||||||||
| Fluoroquinolone resistance genes | ||||||||||||||||||||||||||
| crpP | ||||||||||||||||||||||||||
| qnrVC1 | ||||||||||||||||||||||||||
| Beta-lactamase resistance genes | ||||||||||||||||||||||||||
| blaPAO | ||||||||||||||||||||||||||
| blaLCR-1 | ||||||||||||||||||||||||||
| blaOXA-485 | ||||||||||||||||||||||||||
| blaOXA-486 | ||||||||||||||||||||||||||
| blaOXA-488 | ||||||||||||||||||||||||||
| blaOXA-396 | ||||||||||||||||||||||||||
| blaOXA-395 | ||||||||||||||||||||||||||
| blaOXA-50 | ||||||||||||||||||||||||||
| blaOXA-10 | ||||||||||||||||||||||||||
| blaTEM-1B | ||||||||||||||||||||||||||
| blaVIM-2 | ||||||||||||||||||||||||||
| blaPME-1 | ||||||||||||||||||||||||||
| blaPAU-1 | ||||||||||||||||||||||||||
| Sulphonamide, tetracycline, macrolide, fosfomycin, and chloramphenicol resistance genes | ||||||||||||||||||||||||||
| sul1 | ||||||||||||||||||||||||||
| tet(G) | ||||||||||||||||||||||||||
| mph(E) | ||||||||||||||||||||||||||
| mph(A) | ||||||||||||||||||||||||||
| msr(E)* | ||||||||||||||||||||||||||
| fosA | ||||||||||||||||||||||||||
| catB | ||||||||||||||||||||||||||
| P. aeruginosa Isolates | Total Variants | Variant Complex | Variants Insertions | Variants Deletions | Variants MNP | Variant SNP |
|---|---|---|---|---|---|---|
| 123 | 28,279 | 1593 | 187 | 163 | 398 | 25,938 |
| 126 | 26,258 | 1416 | 164 | 159 | 355 | 24,164 |
| 127 | 25,760 | 1362 | 163 | 176 | 391 | 23,668 |
| 162 | 50,999 | 3481 | 281 | 257 | 951 | 46,029 |
| 169 | 50,283 | 3359 | 269 | 245 | 922 | 45,488 |
| 176 | 26,065 | 1372 | 168 | 161 | 342 | 24,022 |
| 181 | 22,536 | 1063 | 162 | 133 | 283 | 20,895 |
| 182 | 25,684 | 1359 | 172 | 180 | 368 | 23,605 |
| 223 | 25,672 | 1358 | 167 | 176 | 402 | 23,568 |
| 224 | 26,376 | 1435 | 163 | 165 | 353 | 24,260 |
| 225 | 28,070 | 1566 | 167 | 156 | 385 | 25,796 |
| 227 | 28,000 | 1560 | 162 | 154 | 370 | 25,754 |
| 233 | 52,392 | 3590 | 285 | 263 | 956 | 47,298 |
| 235 | 24,919 | 1349 | 162 | 171 | 354 | 22,883 |
| 188 | 25,833 | 1435 | 164 | 154 | 351 | 23,729 |
| 189 | 25,910 | 1458 | 165 | 155 | 365 | 23,767 |
| 193 | 26,567 | 1445 | 180 | 147 | 389 | 24,406 |
| 198 | 50,631 | 3503 | 280 | 236 | 945 | 45,667 |
| 202 | 49,981 | 3461 | 257 | 236 | 902 | 45,125 |
| 206 | 76,180 | 6449 | 336 | 371 | 1653 | 67,271 |
| 216 | 28,166 | 1548 | 183 | 164 | 433 | 25,838 |
| 217 | 51,119 | 3575 | 290 | 226 | 944 | 46,084 |
| 218 | 29,161 | 1676 | 182 | 181 | 430 | 26,692 |
| 219 | 50,507 | 3484 | 273 | 237 | 925 | 45,588 |
| 220 | 50,180 | 3452 | 267 | 234 | 894 | 45,332 |
| 221 | 50,030 | 3477 | 260 | 237 | 906 | 45,150 |
| P. aeruginosa Isolates | Type III Secretion System Genes | mutL | mutS | uvrD |
|---|---|---|---|---|
| 123 | exoU | 1 SNP | 0 | 1 complex |
| 126 | exoU/exoS | 0 | 0 | 0 |
| 127 | exoU | 0 | 1 MNP | 1 MNP, 1 complex |
| 162 | exoU | 1 SNP | 0 | 2 SNP, 1 MNP, 2 complexes |
| 169 | exoU | 1 SNP | 1 complex | 2 SNP, 2 MNP, 1 complex |
| 176 | exoS | 1 SNP | 0 | 1 SNP |
| 181 | exoS | 0 | 0 | 0 |
| 182 | exoS | 0 | 0 | 1 MNP 1complex |
| 223 | exoS | 0 | 1 SNP | 1 MNP, 1 complex |
| 224 | exoS | 1 SNP | 0 | 1 MNP, 1 complex |
| 225 | exoS | 0 | 0 | 2 SNP, 2 MNP, 1 complex |
| 227 | exoS | 0 | 0 | 2 SNP, 2 MNP, 1 complex |
| 233 | exoU | 0 | 0 | 1 MNP, 1 complex |
| 235 | exoS | 0 | 0 | 0 |
| 188 | exoS | 0 | 0 | 1 MNP, 1 complex |
| 189 | exoS | 1 SNP | 0 | 1 MNP, 1 complex |
| 193 | exoS | 0 | 0 | 0 |
| 198 | exoU | 2 SNP | 0 | 1 SNP, 3 complexes |
| 202 | exoU | 1 SNP | 1 complex | 1 SNP, 2 MNP, 2 complexes |
| 206 | exoS | 1 MNP | 1 complex | 0 |
| 216 | exoS | 0 | 0 | 0 |
| 217 | exoU | 2 SNP | 1 complex | 1 SNP, 2 MNP, 1 complex |
| 218 | exoS | 0 | 0 | 0 |
| 219 | exoU | 2 SNP | 0 | 1 SNP, 1 MNP, 2 complexes |
| 220 | exoU | 1 SNP | 1 complex | 1 SNP, 2 MNP, 2 complexes |
| 221 | exoU | 1 SNP | 1 complex | 1 SNP, 2 MNP, 2 complexes |
| P. aeruginosa Isolates | Sequence Types | Core Genes | Shell Genes | Pan/Total Genes |
|---|---|---|---|---|
| 123 | ST218 | 5496 | 508 | 6004 |
| 126 | ST2726 | 5483 | 712 | 6195 |
| 127 | ST845 | 5483 | 938 | 6421 |
| 162 | ST298 | 5439 | 905 | 6344 |
| 169 | ST1027 | 5456 | 694 | 6150 |
| 176 | ST709 | 5547 | 1112 | 6659 |
| 181 | ST244 | 5588 | 1047 | 6662 |
| 182 | ST27 | 5486 | 1096 | 6582 |
| 223 | ST17 | 5471 | 1232 | 6703 |
| 224 | ST168 | 5483 | 607 | 6090 |
| 225 ¤ | ST233 | 5515 | 1338 | 6853 |
| 227 ¤ | ST233 | 5493 | 1304 | 6797 |
| 233 | NEWST | 5440 | 624 | 6064 |
| 235 | ST262 | 5470 | 540 | 6010 |
| 188 * | ST491 | 5490 | 535 | 6025 |
| 189 * | ST491 | 5492 | 531 | 6023 |
| 193 | ST760 | 5490 | 594 | 6084 |
| 198 † | ST308 | 5454 | 1428 | 6882 |
| 202 # | ST316 | 5425 | 1505 | 6930 |
| 206 | NEWST | 5331 | 1084 | 6415 |
| 216 | ST1527 | 5480 | 1488 | 6968 |
| 217 | ST1047 | 5448 | 1173 | 6621 |
| 218 | ST3083 | 5513 | 488 | 6001 |
| 219 † | ST308 | 5451 | 1796 | 7247 |
| 220 # | ST316 | 5430 | 948 | 6378 |
| 221 # | ST316 | 5425 | 1511 | 6936 |
| PA7 | ST1196 | 3599 | 4586 | 8185 |
| PA14 | ST253 | 5436 | 790 | 6226 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Khan, M.; Stapleton, F.; Summers, S.; Rice, S.A.; Willcox, M.D.P. Antibiotic Resistance Characteristics of Pseudomonas aeruginosa Isolated from Keratitis in Australia and India. Antibiotics 2020, 9, 600. https://doi.org/10.3390/antibiotics9090600
Khan M, Stapleton F, Summers S, Rice SA, Willcox MDP. Antibiotic Resistance Characteristics of Pseudomonas aeruginosa Isolated from Keratitis in Australia and India. Antibiotics. 2020; 9(9):600. https://doi.org/10.3390/antibiotics9090600
Chicago/Turabian StyleKhan, Mahjabeen, Fiona Stapleton, Stephen Summers, Scott A. Rice, and Mark D. P. Willcox. 2020. "Antibiotic Resistance Characteristics of Pseudomonas aeruginosa Isolated from Keratitis in Australia and India" Antibiotics 9, no. 9: 600. https://doi.org/10.3390/antibiotics9090600
APA StyleKhan, M., Stapleton, F., Summers, S., Rice, S. A., & Willcox, M. D. P. (2020). Antibiotic Resistance Characteristics of Pseudomonas aeruginosa Isolated from Keratitis in Australia and India. Antibiotics, 9(9), 600. https://doi.org/10.3390/antibiotics9090600







