Deep Functional Profiling Facilitates the Evaluation of the Antibacterial Potential of the Antibiotic Amicoumacin
Abstract
1. Introduction
2. Results
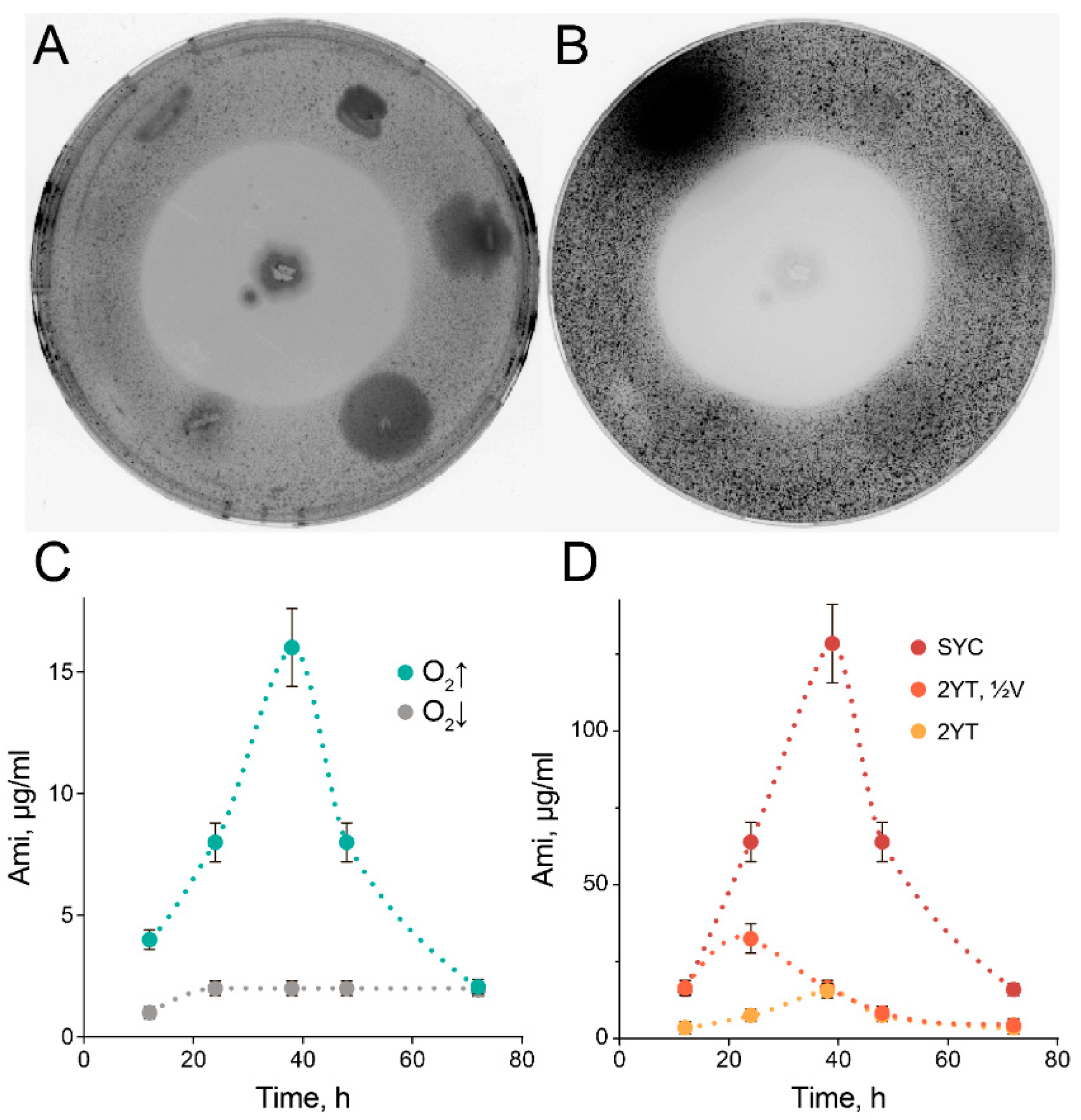
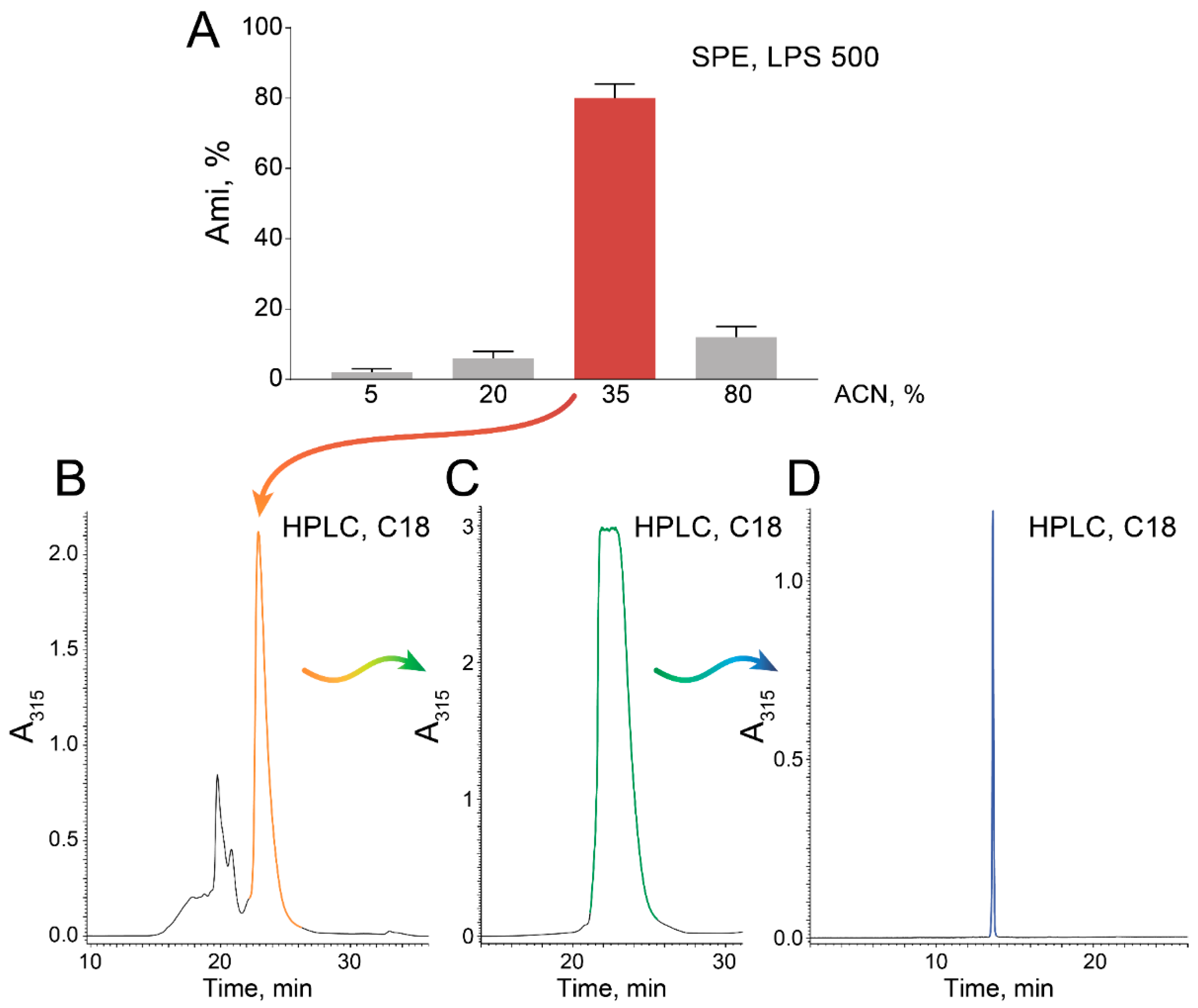
2.1. High-Scale Ami Production and Purification
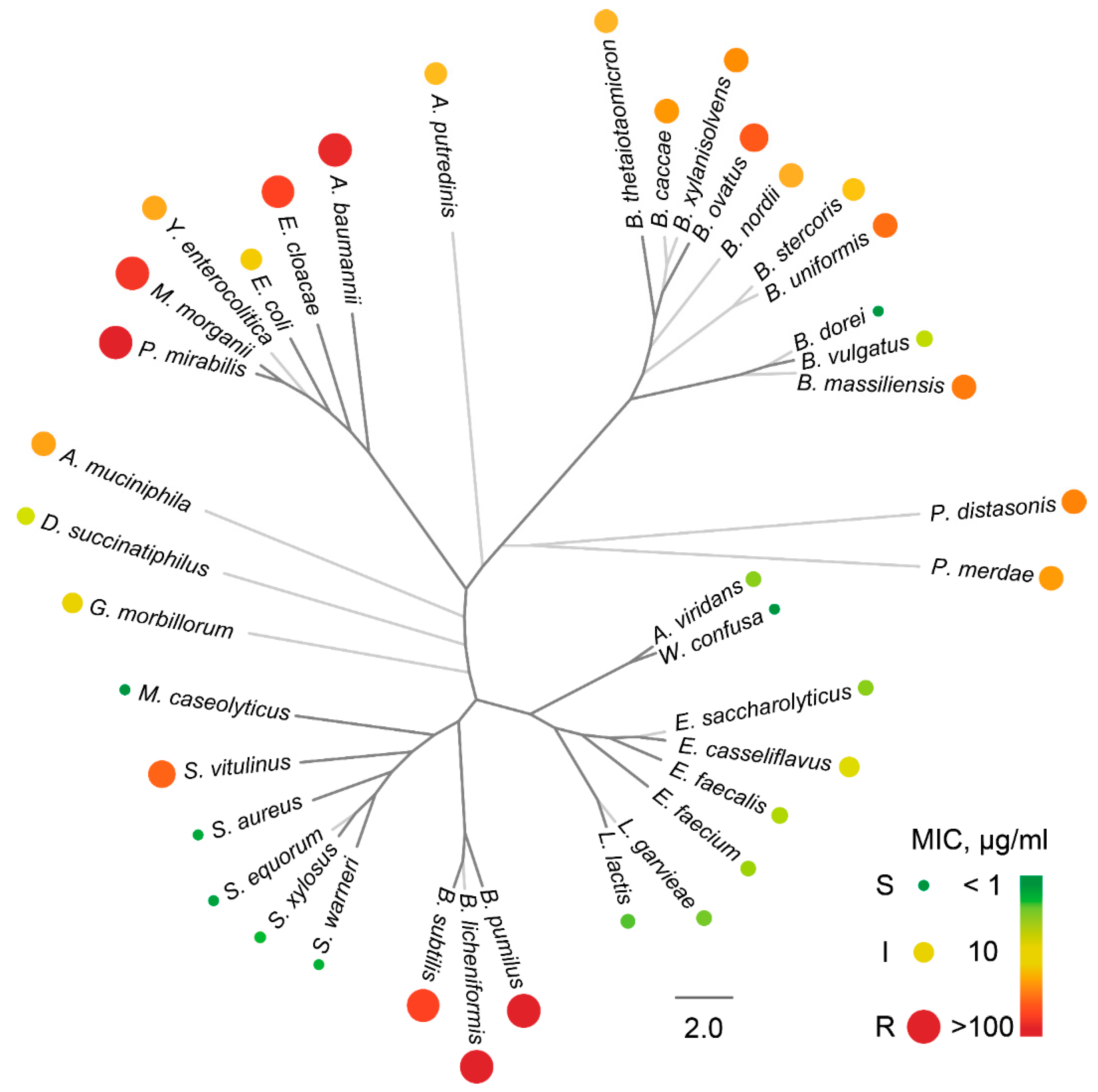
2.2. The Activity Spectrum of Ami
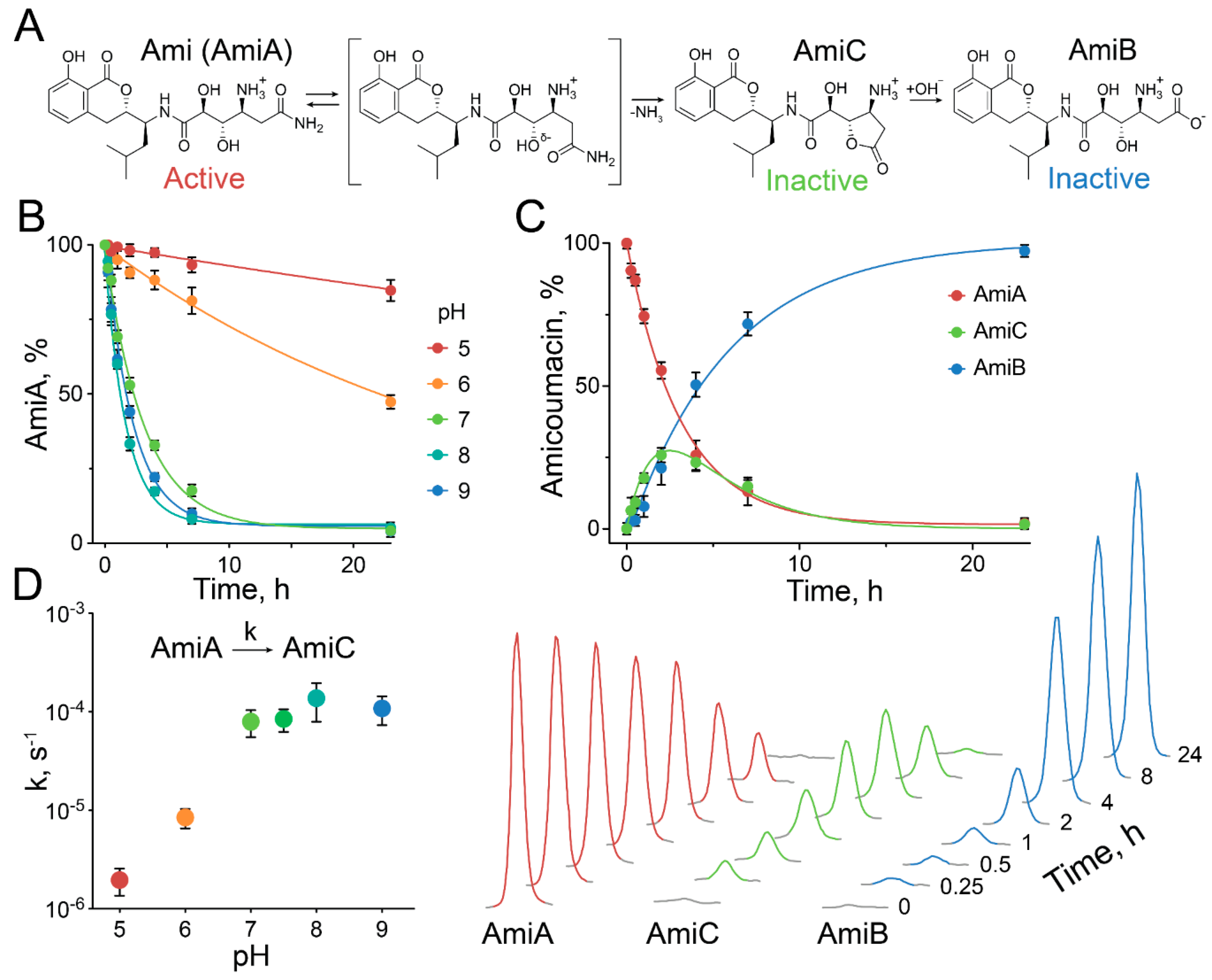
2.3. Exceptional Instability of Ami
3. Discussion
4. Materials and Methods
4.1. Microbiota Collection and Storage
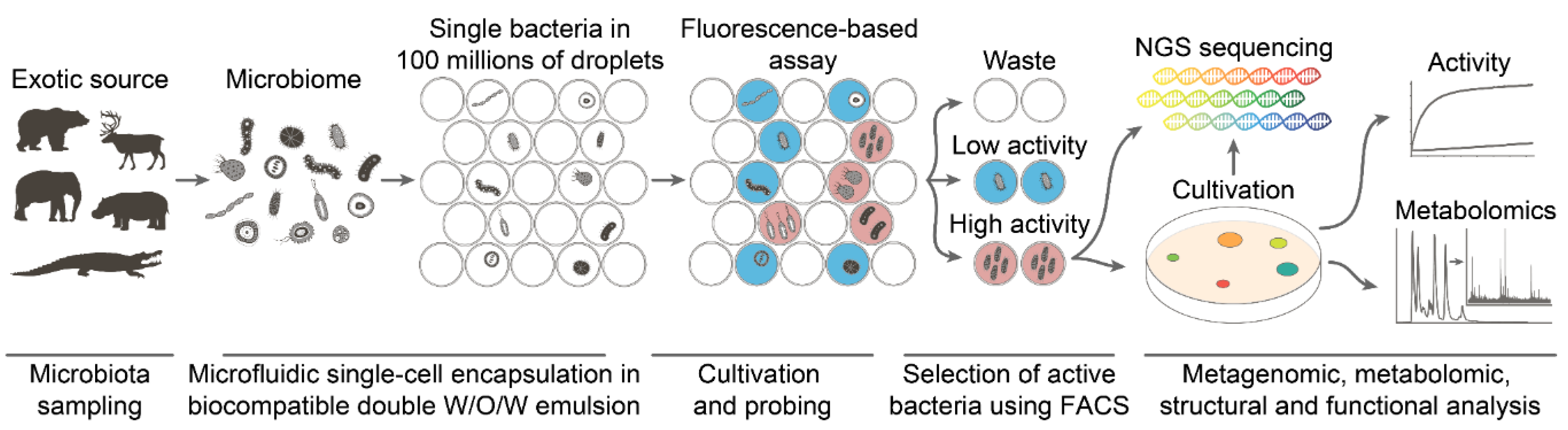
4.2. Deep Functional Profiling
4.3. NGS Sequencing and Bioinformatics
4.4. Identification of Bacteria using Mass Spectrometry
4.5. Antimicrobial Activity
4.6. Ami Production and Purification
4.7. Ami Hydrolysis
Author Contributions
Funding
Conflicts of Interest
References
- Fleischmann, C.; Scherag, A.; Adhikari, N.K.J.; Hartog, C.S.; Tsaganos, T.; Schlattmann, P.; Angus, D.C.; Reinhart, K. Assessment of Global Incidence and Mortality of Hospital-treated Sepsis. Current Estimates and Limitations. Am. J. Respir. Crit. Care Med. 2016, 193, 259–272. [Google Scholar] [CrossRef] [PubMed]
- Global Action Plan on Antimicrobial Resistance; WHO Press: Geneva, Switzerland, 2015.
- Lewis, K. Platforms for antibiotic discovery. Nat. Rev. Drug Discov. 2013, 12, 371–387. [Google Scholar] [CrossRef] [PubMed]
- Hatzenpichler, R.; Krukenberg, V.; Spietz, R.L.; Jay, Z.J. Next-generation physiology approaches to study microbiome function at single cell level. Nat. Rev. Genet. 2020, 18, 241–256. [Google Scholar] [CrossRef] [PubMed]
- Terekhov, S.S.; Smirnov, I.V.; Malakhova, M.V.; Samoilov, A.E.; Manolov, A.I.; Nazarov, A.S.; Danilov, D.V.; Dubiley, S.A.; Osterman, I.A.; Rubtsova, M.P.; et al. Ultrahigh-throughput functional profiling of microbiota communities. Proc. Natl. Acad. Sci. USA 2018, 115, 9551–9556. [Google Scholar] [CrossRef]
- Terekhov, S.S.; Smirnov, I.V.; Stepanova, A.V.; Bobik, T.V.; Mokrushina, Y.A.; Ponomarenko, N.A.; Belogurov, A.A.; Rubtsova, M.P.; Kartseva, O.V.; Gomzikova, M.O.; et al. Microfluidic droplet platform for ultrahigh-throughput single-cell screening of biodiversity. Proc. Natl. Acad. Sci. USA 2017, 114, 2550–2555. [Google Scholar] [CrossRef]
- Itoh, J.; Omoto, S.; Shomura, T.; Nishizawa, N.; Miyado, S.; Yuda, Y.; Shibata, U.; Inouye, S. Amicoumacin-A, a new antibiotic with strong antiinflammatory and antiulcer activity. J. Antibiot. 1981, 34, 611–613. [Google Scholar] [CrossRef]
- Itoh, J.; Shomura, T.; Omoto, S.; Miyado, S.; Yuda, Y.; Shibata, U.; Inouye, S. Isolation, Physicochemical Properties and Biological Activities of Amicoumacins Produced by Bacillus pumilus. Agricul. Biolog. Chem. 1982, 46, 1255–1259. [Google Scholar] [CrossRef]
- Itoh, J.; Omoto, S.; Nishizawa, N.; Kodama, Y.; Inouye, S. Chemical Structures of Amicoumacins Produced by Bacillus pumilus. Agricul. Biolog. Chem. 1982, 46, 2659–2665. [Google Scholar] [CrossRef]
- Lama, A.; Pané-Farré, J.; Chon, T.; Wiersma, A.M.; Sit, C.S.; Vederas, J.C.; Hecker, M.; Nakano, M.M. Response of Methicillin-Resistant Staphylococcus aureus to Amicoumacin A. PLoS ONE 2012, 7, e34037. [Google Scholar] [CrossRef]
- Polikanov, Y.S.; Osterman, I.A.; Szal, T.; Tashlitsky, V.N.; Serebryakova, M.V.; Kusochek, P.; Bulkley, D.; Malanicheva, I.A.; Efimenko, T.; Efremenkova, O.V.; et al. Amicoumacin a inhibits translation by stabilizing mRNA interaction with the ribosome. Mol. Cell 2014, 56, 531–540. [Google Scholar] [CrossRef]
- Prokhorova, I.V.; Akulich, K.A.; Makeeva, D.S.; Osterman, I.A.; Skvortsov, D.A.; Sergiev, P.V.; Dontsova, O.A.; Yusupova, G.; Yusupov, M.M.; Dmitriev, S.E. Amicoumacin A induces cancer cell death by targeting the eukaryotic ribosome. Sci. Rep. 2016, 6, 27720. [Google Scholar] [CrossRef] [PubMed]
- Pinchuk, I.V.; Bressollier, P.; Verneuil, B.; Fenet, B.; Sorokulova, I.B.; Mégraud, F.; Urdaci, M.C. In vitro anti-Helicobacter pylori activity of the probiotic strain Bacillus subtilis 3 is due to secretion of antibiotics. Antimicrob. Agents Chemother. 2001, 45, 3156–3161. [Google Scholar] [CrossRef]
- Pinchuk, I.V.; Bressollier, P.; Sorokulova, I.B.; Verneuil, B.; Urdaci, M.C. Amicoumacin antibiotic production and genetic diversity of Bacillus subtilis strains isolated from different habitats. Res. Microbiol. 2002, 153, 269–276. [Google Scholar] [CrossRef]
- Gao, X.-Y.; Liu, Y.; Miao, L.-L.; Li, E.-W.; Hou, T.-T.; Liu, Z.-P. Mechanism of anti-Vibrio activity of marine probiotic strain Bacillus pumilus H2, and characterization of the active substance. AMB Express 2017, 7, 23. [Google Scholar] [CrossRef] [PubMed]
- Zidour, M.; Chevalier, M.; Belguesmia, Y.; Cudennec, B.; Grard, T.; Drider, D.; Souissi, S.; Flahaut, C. Isolation and Characterization of Bacteria Colonizing Acartia tonsa Copepod Eggs and Displaying Antagonist Effects against Vibrio anguillarum, Vibrio alginolyticus and Other Pathogenic Strains. Front. Microbiol. 2017, 8, 1919. [Google Scholar] [CrossRef] [PubMed]
- Lefevre, M.; Racedo, S.M.; Ripert, G.; Housez, B.; Cazaubiel, M.; Maudet, C.; Jüsten, P.; Marteau, P.; Urdaci, M.C. Probiotic strain Bacillus subtilis CU1 stimulates immune system of elderly during common infectious disease period: A randomized, double-blind placebo-controlled study. Immun. Ageing 2015, 12, 24. [Google Scholar] [CrossRef]
- Li, Y.; Li, Z.; Yamanaka, K.; Xu, Y.; Zhang, W.; Vlamakis, H.; Kolter, R.; Moore, B.S.; Qian, P.-Y. Directed natural product biosynthesis gene cluster capture and expression in the model bacterium Bacillus subtilis. Sci. Rep. 2015, 5, 9383. [Google Scholar] [CrossRef]
- Reimer, D.; Pos, K.M.; Thines, M.; Grün, P.; Bode, H.B. A natural prodrug activation mechanism in nonribosomal peptide synthesis. Nat. Chem. Biol. 2011, 7, 888–890. [Google Scholar] [CrossRef]
- Park, H.B.; Perez, C.E.; Perry, E.K.; Crawford, J.M. Activating and Attenuating the Amicoumacin Antibiotics. Molecules 2016, 21, 824. [Google Scholar] [CrossRef]
- Niu, H.; Wang, J.; Zhuang, W.; Liu, D.; Chen, Y.; Zhu, C.; Ying, H. Comparative transcriptomic and proteomic analysis of Arthrobacter sp. CGMCC 3584 responding to dissolved oxygen for cAMP production. Sci. Rep. 2018, 8, 1246. [Google Scholar] [CrossRef]
- Mahesh, S.; Tang, K.-C.; Raj, M. Amide Bond Activation of Biological Molecules. Molecules 2018, 23, 2615. [Google Scholar] [CrossRef] [PubMed]
- Gao, C.; Lavey, B.J.; Lo, C.-H.L.; Datta, A.; Wentworth, P.; Janda, K.D. Direct Selection for Catalysis from Combinatorial Antibody Libraries Using a Boronic Acid Probe: Primary Amide Bond Hydrolysis. J. Am. Che. Soc. 1998, 120, 2211–2217. [Google Scholar] [CrossRef]
- Robinson, B.A.; Tester, J.W. Kinetics of alkaline hydrolysis of organic esters and amides in neutrally-buffered solution. Int. J. Chem. Kinet. 1990, 22, 431–448. [Google Scholar] [CrossRef]
- Martin, M.T.; Angeles, T.S.; Sugasawara, R.; Aman, N.I.; Napper, A.D.; Darsley, M.J.; Sanchez, R.I.; Booth, P.; Titmas, R.C. Antibody-Catalyzed Hydrolysis of an Unsubstituted Amide. J. Am. Chem. Soc. 1994, 116, 6508–6512. [Google Scholar] [CrossRef]
- Suzuki, T.; Nagasawa, T.; Enomoto, M.; Kuwahara, S. Stereoselective total synthesis of amicoumacin C. Tetrahedron 2015, 71, 1992–1997. [Google Scholar] [CrossRef]
- Tsukaguchi, S.; Enomoto, M.; Towada, R.; Ogura, Y.; Kuwahara, S. Unified Total Synthesis of Hetiamacins A–D. Eur. J. Org. Chem. 2019, 2019, 6110–6116. [Google Scholar] [CrossRef]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- Truong, D.T.; Franzosa, E.A.; Tickle, T.L.; Scholz, M.; Weingart, G.; Pasolli, E.; Tett, A.; Huttenhower, C.; Segata, N. MetaPhlAn2 for enhanced metagenomic taxonomic profiling. Nat. Methods 2015, 12, 902–903. [Google Scholar] [CrossRef]
- Madeira, F.; Park, Y.M.; Lee, J.; Buso, N.; Gur, T.; Madhusoodanan, N.; Basutkar, P.; Tivey, A.R.N.; Potter, S.C.; Finn, R.D.; et al. The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic. Acids Res. 2019, 47, W636–W641. [Google Scholar] [CrossRef]
- Okonechnikov, K.; Golosova, O.; Fursov, M.; Ugene Team. Unipro UGENE: A unified bioinformatics toolkit. Bioinformatics 2012, 28, 1166–1167. [Google Scholar] [CrossRef]
- Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef] [PubMed]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Terekhov, S.S.; Nazarov, A.S.; Mokrushina, Y.A.; Baranova, M.N.; Potapova, N.A.; Malakhova, M.V.; Ilina, E.N.; Smirnov, I.V.; Gabibov, A.G. Deep Functional Profiling Facilitates the Evaluation of the Antibacterial Potential of the Antibiotic Amicoumacin. Antibiotics 2020, 9, 157. https://doi.org/10.3390/antibiotics9040157
Terekhov SS, Nazarov AS, Mokrushina YA, Baranova MN, Potapova NA, Malakhova MV, Ilina EN, Smirnov IV, Gabibov AG. Deep Functional Profiling Facilitates the Evaluation of the Antibacterial Potential of the Antibiotic Amicoumacin. Antibiotics. 2020; 9(4):157. https://doi.org/10.3390/antibiotics9040157
Chicago/Turabian StyleTerekhov, Stanislav S., Anton S. Nazarov, Yuliana A. Mokrushina, Margarita N. Baranova, Nadezhda A. Potapova, Maja V. Malakhova, Elena N. Ilina, Ivan V. Smirnov, and Alexander G. Gabibov. 2020. "Deep Functional Profiling Facilitates the Evaluation of the Antibacterial Potential of the Antibiotic Amicoumacin" Antibiotics 9, no. 4: 157. https://doi.org/10.3390/antibiotics9040157
APA StyleTerekhov, S. S., Nazarov, A. S., Mokrushina, Y. A., Baranova, M. N., Potapova, N. A., Malakhova, M. V., Ilina, E. N., Smirnov, I. V., & Gabibov, A. G. (2020). Deep Functional Profiling Facilitates the Evaluation of the Antibacterial Potential of the Antibiotic Amicoumacin. Antibiotics, 9(4), 157. https://doi.org/10.3390/antibiotics9040157





