Antibiofilm Activity of Heather and Manuka Honeys and Antivirulence Potential of Some of Their Constituents on the DsbA1 Enzyme of Pseudomonas aeruginosa
Abstract
1. Introduction
2. Results
2.1. Determination of the Time Required for Optimal Biofilm Formation
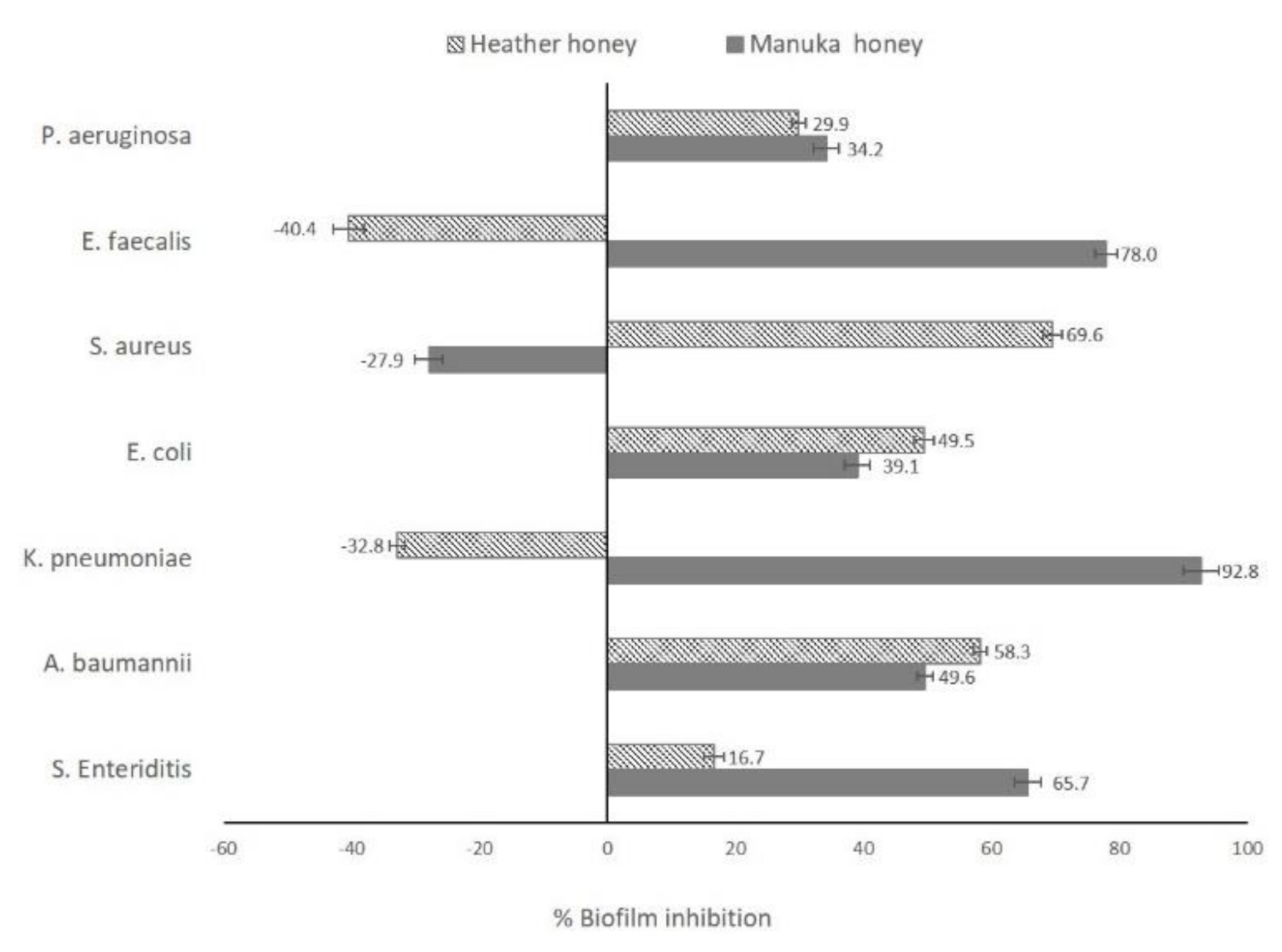
2.2. Antibiofilm Activity
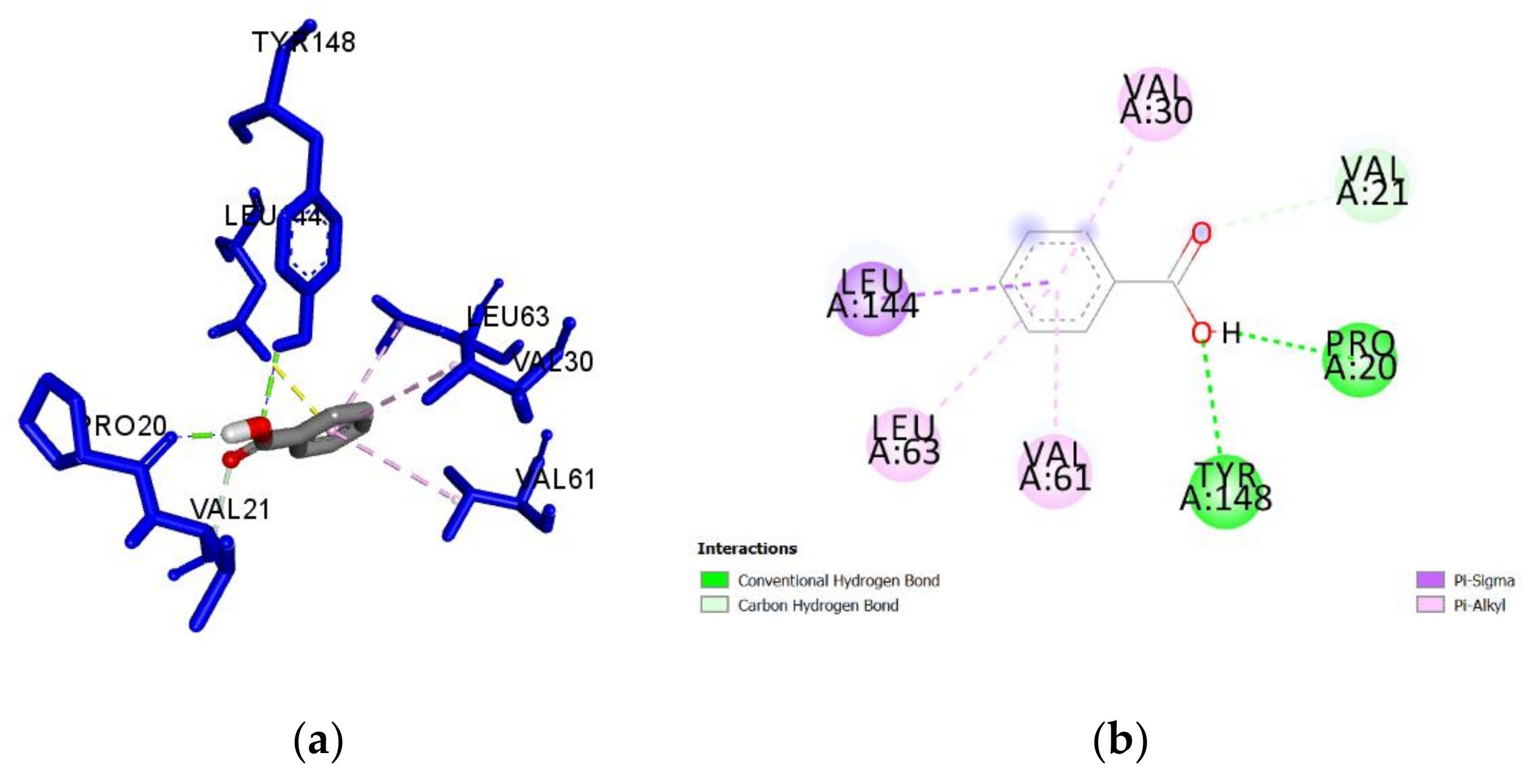
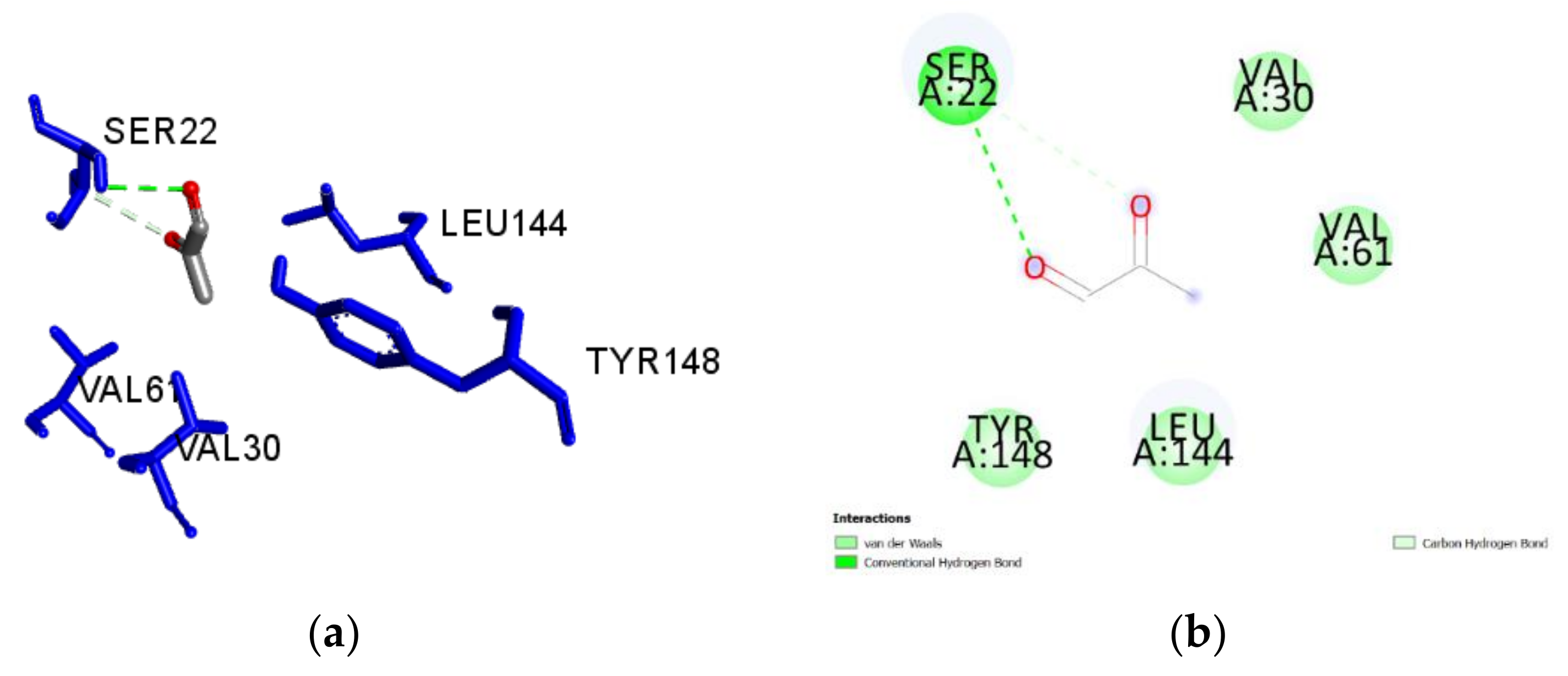
2.3. Molecular Docking of Honey Constituents against P. aeruginosa DsbA1
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains, Culture Conditions, and Inoculum Preparation
4.2. Honey Sources and Preparation of Samples
4.3. Antibiofilm Assay
4.4. Molecular Docking Experiment
4.4.1. Protein Preparation
4.4.2. Ligand Preparation
4.4.3. Grid Box Preparation and Docking Studies
4.4.4. Protein-Ligand Interactions and Predictive Inhibition
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Arzanlou, M.; Chai, W.C.; Venter, H. Intrinsic, adaptive and acquired antimicrobial resistance in Gram-negative bacteria. Essays Biochem. 2017, 61, 49–59. [Google Scholar] [CrossRef]
- Rossiter, S.E.; Fletcher, M.H.; Wuest, W.M. Natural Products as Platforms to Overcome Antibiotic Resistance. Chem. Rev. 2017, 117, 12415–12474. [Google Scholar] [CrossRef] [PubMed]
- Chandra, H.; Bishnoi, P.; Yadav, A.; Patni, B.; Mishra, A.P.; Nautiyal, A.R. Antimicrobial Resistance and the Alternative Resources with Special Emphasis on Plant-Based Antimicrobials—A Review. Plants 2017, 6, 16. [Google Scholar] [CrossRef] [PubMed]
- Dickey, S.W.; Cheung, G.Y.C.; Otto, M. Different drugs for bad bugs: Antivirulence strategies in the age of antibiotic resistance. Nat. Rev. Drug Discov. 2017, 16, 457–471. [Google Scholar] [CrossRef]
- Heras, B.; Scanlon, M.J.; Martin, J.L. Targeting virulence not viability in the search for future antibacterials. Br. J. Clin. Pharmacol. 2015, 79, 208–215. [Google Scholar] [CrossRef] [PubMed]
- Shouldice, S.R.; Heras, B.; Walden, P.M.; Totsika, M.; Schembri, M.A.; Martin, J.L. Structure and Function of DsbA, a Key Bacterial Oxidative Folding Catalyst. Antioxidants Redox Signal. 2011, 14, 1729–1760. [Google Scholar] [CrossRef]
- Yu, J. DsbA: A protein-folding catalyst contributing to bacterial virulence. Microbes Infect. 1999, 1, 1221–1228. [Google Scholar] [CrossRef]
- Lee, Y.; Kim, Y.; Yeom, S.; Kim, S.; Park, S.; Jeon, C.O.; Park, W. The role of disulfide bond isomerase A (DsbA) of Escherichia coli O157:H7 in biofilm formation and virulence. FEMS Microbiol. Lett. 2008, 278, 213–222. [Google Scholar] [CrossRef]
- Bocian-Ostrzycka, K.M.; Grzeszczuk, M.J.; Banaś, A.M.; Jagusztyn-Krynicka, E.K. Bacterial thiol oxidoreductases—From basic research to new antibacterial strategies. Appl. Microbiol. Biotechnol. 2017, 101, 3977–3989. [Google Scholar] [CrossRef]
- Lasica, A.M.; Jagusztyn-Krynicka, E.K. The role of Dsb proteins of Gram-negative bacteria in the process of pathogenesis. FEMS Microbiol. Rev. 2007, 31, 626–636. [Google Scholar] [CrossRef]
- Totsika, M.; Vagenas, D.; Paxman, J.J.; Wang, G.; Dhouib, R.; Sharma, P.; Martin, J.L.; Scanlon, M.J.; Heras, B. Inhibition of Diverse DsbA Enzymes in Multi-DsbA Encoding Pathogens. Antioxid. Redox Signal. 2018, 29, 653–666. [Google Scholar] [CrossRef] [PubMed]
- Reardon-Robinson, M.E.; Ton-That, H. Disulfide-Bond-Forming Pathways in Gram-Positive Bacteria. J. Bacteriol. 2015, 198, 746–754. [Google Scholar] [CrossRef]
- Kumar, A.; Alam, A.; Rani, M.; Ehtesham, N.Z.; Hasnain, S.E. Biofilms: Survival and defense strategy for pathogens. Int. J. Med. Microbiol. 2017, 307, 481–489. [Google Scholar] [CrossRef]
- Muhammad, M.H.; Idris, A.L.; Fan, X.; Guo, Y.; Yu, Y.; Jin, X.; Qiu, J.; Guan, X.; Huang, T. Beyond Risk: Bacterial Biofilms and Their Regulating Approaches. Front. Microbiol. 2020, 11, 928. [Google Scholar] [CrossRef]
- Lee, K.; Yoon, S.S. Pseudomonas aeruginosa Biofilm, a Programmed Bacterial Life for Fitness. J. Microbiol. Biotechnol. 2017, 27, 1053–1064. [Google Scholar] [CrossRef] [PubMed]
- Pang, Z.; Raudonis, R.; Glick, B.R.; Lin, T.-J.; Cheng, Z. Antibiotic resistance in Pseudomonas aeruginosa: Mechanisms and alternative therapeutic strategies. Biotechnol. Adv. 2019, 37, 177–192. [Google Scholar] [CrossRef] [PubMed]
- Clinton, A.; Carter, T. Chronic Wound Biofilms: Pathogenesis and Potential Therapies. Lab. Med. 2015, 46, 277–284. [Google Scholar] [CrossRef] [PubMed]
- Serra, R.; Grande, R.; Butrico, L.; Rossi, A.; Settimio, U.F.; Caroleo, B.; Amato, B.; Gallelli, L.; De Franciscis, S. Chronic wound infections: The role of Pseudomonas aeruginosa and Staphylococcus aureus. Expert Rev. Anti-Infect. Ther. 2015, 13, 605–613. [Google Scholar] [CrossRef]
- Ha, U.-H.; Wang, Y.; Jin, S. DsbA of Pseudomonas aeruginosa Is Essential for Multiple Virulence Factors. Infect. Immun. 2003, 71, 1590–1595. [Google Scholar] [CrossRef]
- Mohanty, B.; Rimmer, K.; McMahon, R.M.; Headey, S.J.; Vazirani, M.; Shouldice, S.R.; Coinçon, M.; Tay, S.; Morton, C.J.; Simpson, J.S.; et al. Fragment library screening identifies hits that bind to the non-catalytic surface of Pseudomonas aeruginosa DsbA1. PLoS ONE 2017, 12, e0173436. [Google Scholar] [CrossRef]
- Viuda-Martos, M.; Ruiz-Navajas, Y.; Fernández-López, J.; Pérez-Álvarez, J. Functional Properties of Honey, Propolis, and Royal Jelly. J. Food Sci. 2008, 73, R117–R124. [Google Scholar] [CrossRef] [PubMed]
- Mandal, M.D.; Mandal, S. Honey: Its medicinal property and antibacterial activity. Asian Pac. J. Trop. Biomed. 2011, 1, 154–160. [Google Scholar] [CrossRef]
- Hussain, M.B. Role of Honey in Topical and Systemic Bacterial Infections. J. Altern. Complement. Med. 2018, 24, 15–24. [Google Scholar] [CrossRef]
- Temaru, E.; Shimura, S.; Amano, K.; Karasawa, T. Antibacterial activity of honey from stingless honeybees (Hymenoptera; Apidae; Meliponinae). Pol. J. Microbiol. 2007, 56, 281–285. [Google Scholar] [PubMed]
- Maddocks, S.E.; Jenkins, R.E. Honey: A sweet solution to the growing problem of antimicrobial resistance? Future Microbiol. 2013, 8, 1419–1429. [Google Scholar] [CrossRef]
- Minden-Birkenmaier, B.A.; Bowlin, G.L. Honey-Based Templates in Wound Healing and Tissue Engineering. Bioengineering 2018, 5, 46. [Google Scholar] [CrossRef]
- Da Silva, P.M.; Gauche, C.; Gonzaga, L.V.; Costa, A.C.O.; Fett, R. Honey: Chemical composition, stability and authenticity. Food Chem. 2016, 196, 309–323. [Google Scholar] [CrossRef]
- Albaridi, N.A. Antibacterial Potency of Honey. Int. J. Microbiol. 2019, 2019, 1–10. [Google Scholar] [CrossRef]
- Nolan, V.C.; Harrison, J.; Cox, J. Dissecting the Antimicrobial Composition of Honey. Antibiotics 2019, 8, 251. [Google Scholar] [CrossRef]
- Kwakman, P.H.S.; Zaat, S.A.J. Antibacterial components of honey. IUBMB Life 2012, 64, 48–55. [Google Scholar] [CrossRef]
- Fyfe, L.; Okoro, P.; Paterson, E.; Coyle, S.; McDougall, G.J. Compositional analysis of Scottish honeys with antimicrobial activity against antibiotic-resistant bacteria reveals novel antimicrobial components. LWT Food Sci. Technol. 2017, 79, 52–59. [Google Scholar] [CrossRef]
- Dezmirean, G.I.; Mărghitaş, L.A.; Bobiş, O.; Dezmirean, D.S.; Bonta, V.; Erler, S. Botanical Origin Causes Changes in Nutritional Profile and Antioxidant Activity of Fermented Products Obtained from Honey. J. Agric. Food Chem. 2012, 60, 8028–8035. [Google Scholar] [CrossRef] [PubMed]
- Karabagias, I.K.; Maia, M.; Karabagias, V.K.; Gatzias, I.; Badeka, A.V. Characterization of Eucalyptus, Chestnut and Heather Honeys from Portugal Using Multi-Parameter Analysis and Chemo-Calculus. Foods 2018, 7, 194. [Google Scholar] [CrossRef] [PubMed]
- Kavanagh, S.; Gunnoo, J.; Passos, T.M.; Stout, J.C.; White, B. Physicochemical properties and phenolic content of honey from different floral origins and from rural versus urban landscapes. Food Chem. 2019, 272, 66–75. [Google Scholar] [CrossRef]
- Mavric, E.; Wittmann, S.; Barth, G.; Henle, T. Identification and quantification of methylglyoxal as the dominant antibacterial constituent of Manuka (Leptospermum scoparium) honeys from New Zealand. Mol. Nutr. Food Res. 2008, 52, 483–489. [Google Scholar] [CrossRef]
- Johnston, M.; McBride, M.; Dahiya, D.; Owusu-Apenten, R.; Nigam, P.S. Antibacterial activity of Manuka honey and its components: An overview. AIMS Microbiol. 2018, 4, 655–664. [Google Scholar] [CrossRef]
- Ferreres, F.; Andrade, P.; Tomás-Barberán, F.A. Natural Occurrence of Abscisic Acid in Heather Honey and Floral Nectar. J. Agric. Food Chem. 1996, 44, 2053–2056. [Google Scholar] [CrossRef]
- Salonen, A.; Virjamo, V.; Tammela, P.; Fauch, L.; Julkunen-Tiitto, R. Screening bioactivity and bioactive constituents of Nordic unifloral honeys. Food Chem. 2017, 237, 214–224. [Google Scholar] [CrossRef]
- Feás, X.; Iglesias, A.; Rodrigues, S.; Estevinho, L.M. Effect of Erica sp. Honey against Microorganisms of Clinical Importance: Study of the Factors Underlying this Biological Activity. Molecules 2013, 18, 4233–4246. [Google Scholar] [CrossRef]
- Fernandes, L.; Oliveira, A.; Henriques, M.; Rodrigues, M.E. Honey as a Strategy to Fight Candida tropicalis in Mixed-Biofilms with Pseudomonas aeruginosa. Antibiotics 2020, 9, 43. [Google Scholar] [CrossRef]
- Lu, J.; Turnbull, L.; Burke, C.M.; Liu, M.; Carter, D.A.; Schlothauer, R.C.; Whitchurch, C.B.; Harry, E.J. Manuka-type honeys can eradicate biofilms produced by Staphylococcus aureusstrains with different biofilm-forming abilities. PeerJ 2014, 2, e326. [Google Scholar] [CrossRef] [PubMed]
- Paramasivan, S.; Drilling, A.; Jardeleza, C.; Jervis-Bardy, J.; Vreugde, S.; Wormald, P.J. Methylglyoxal-augmented Manuka honey as a topical anti-Staphylococcus aureus biofilm agent: Safety and efficacy in an in vivo model. Int. Forum Allergy Rhinol. 2014, 4, 187–195. [Google Scholar] [CrossRef]
- Piotrowski, M.; Karpiński, P.; Pituch, H.; Van Belkum, A.; Obuch-Woszczatyński, P. Antimicrobial effects of Manuka honey on in vitro biofilm formation by Clostridium difficile. Eur. J. Clin. Microbiol. Infect. Dis. 2017, 36, 1661–1664. [Google Scholar] [CrossRef] [PubMed]
- Hannan, A.; Bajwa, A.E.; Riaz, S.; Arshad, U.; Saleem, S.; Bajwa, U.I. In vitro Salmonella typhi biofilm formation on gallstones and its disruption by Manuka honey. Pak. J. Pharm. Sci. 2018, 31, 129–135. [Google Scholar] [PubMed]
- Maddocks, S.E.; Jenkins, R.E.; Rowlands, R.S.; Purdy, K.J.; Cooper, R.A. Manuka honey inhibits adhesion and invasion of medically important wound bacteriain vitro. Future Microbiol. 2013, 8, 1523–1536. [Google Scholar] [CrossRef] [PubMed]
- Roberts, A.E.L.; Powell, L.C.; Pritchard, M.F.; Thomas, D.W.; Jenkins, R.E. Anti-pseudomonad Activity of Manuka Honey and Antibiotics in a Specialized ex vivo Model Simulating Cystic Fibrosis Lung Infection. Front. Microbiol. 2019, 10, 869. [Google Scholar] [CrossRef]
- Lu, J.; Cokcetin, N.N.; Burke, C.M.; Turnbull, L.; Liu, M.; Carter, D.; Whitchurch, C.B.; Harry, E.J. Honey can inhibit and eliminate biofilms produced by Pseudomonas aeruginosa. Sci. Rep. 2019, 9, 18160. [Google Scholar] [CrossRef]
- Abbas, H.A.; Gad, A.I. Eradication of biofilms formed by bacteria isolated from diabetic foot infections by potential antibiofilm agents alone and in combination with ciprofloxacin. Afr. J. Microbiol. Res. 2014, 8, 3882–3892. [Google Scholar]
- Halstead, F.D.; Webber, M.A.; Rauf, M.; Burt, R.; Dryden, M.; Oppenheim, B.A. In vitro activity of an engineered honey, medical-grade honeys, and antimicrobial wound dressings against biofilm-producing clinical bacterial isolates. J. Wound Care 2016, 25, 93–102. [Google Scholar] [CrossRef]
- Xu, Z.; Liang, Y.; Lin, S.; Chen, D.; Li, B.; Li, L.; Deng, Y. Crystal Violet and XTT Assays on Staphylococcus aureus Biofilm Quantification. Curr. Microbiol. 2016, 73, 474–482. [Google Scholar] [CrossRef]
- Sindi, A.; Chawn, M.V.B.; Hernandez, M.E.; Green, K.; Islam, K.; Locher, C.; Hammer, K. Anti-biofilm effects and characterisation of the hydrogen peroxide activity of a range of Western Australian honeys compared to Manuka and multifloral honeys. Sci. Rep. 2019, 9, 17666. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Ding, Y.; Chen, W.; Zhang, P.; Chen, Y.; Lv, X. Thein vitrostudy of ursolic acid and oleanolic acid inhibiting cariogenic microorganisms as well as biofilm. Oral Dis. 2012, 19, 494–500. [Google Scholar] [CrossRef]
- Kurek, A.; Markowska, K.; Grudniak, A.M.; Janiszowska, W.; Wolska, K.I. The effect of oleanolic and ursolic acids on the hemolytic properties and biofilm formation of Listeria monocytogenes. Pol. J. Microbiol. 2014, 63, 21–25. [Google Scholar] [CrossRef] [PubMed]
- Lawrence, J.A.; Huang, Z.; Rathinavelu, S.; Hu, J.-F.; Garo, E.; Ellis, M.; Norman, V.L.; Buckle, R.; Williams, R.B.; Starks, C.M.; et al. Optimized plant compound with potent anti-biofilm activity across gram-negative species. Bioorg. Med. Chem. 2020, 28, 115229. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.D. Grape Products and Oral Health. J. Nutr. 2009, 139, 1818S–1823S. [Google Scholar] [CrossRef]
- Smith, R.P.; Paxman, J.; Scanlon, M.J.; Heras, B. Targeting Bacterial Dsb Proteins for the Development of Anti-Virulence Agents. Molecules 2016, 21, 811. [Google Scholar] [CrossRef]
- Mirza, Z.R.M.H.; Hasan, T.; Seidel, V.; Yu, J. Geraniol as a novel antivirulence agent against bacillary dysentery-causing Shigella sonnei. Virulence 2018, 9, 450–455. [Google Scholar] [CrossRef] [PubMed]
- Machado, A.M.; Miguel, M.G.; Vilas-Boas, M.; Figueiredo, A.C. Honey Volatiles as a Fingerprint for Botanical Origin—A Review on their Occurrence on Monofloral Honeys. Molecules 2020, 25, 374. [Google Scholar] [CrossRef] [PubMed]
- Tan, S.T.; Holland, P.T.; Wilkins, A.L.; Molan, P.C. Extractives from New Zealand honeys. 1. White clover, manuka and kanuka unifloral honeys. J. Agric. Food Chem. 1988, 36, 453–460. [Google Scholar] [CrossRef]
- Prithiviraj, B.; Bais, H.P.; Weir, T.; Suresh, B.; Najarro, E.H.; Dayakar, B.V.; Schweizer, H.P.; Vivanco, J.M. Down Regulation of Virulence Factors of Pseudomonas aeruginosa by Salicylic Acid Attenuates Its Virulence on Arabidopsis thaliana and Caenorhabditis elegans. Infect. Immun. 2005, 73, 5319–5328. [Google Scholar] [CrossRef]
- Woodward, M.J.; Sojka, M.; Sprigings, K.A.; Humphrey, T.J. The role of SEF14 and SEF17 fimbriae in the adherence of Salmonella enterica serotype Enteritidis to inanimate surfaces. J. Med. Microbiol. 2000, 49, 481–487. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Morris, G.M.; Huey, R.; Lindstrom, W.; Sanner, M.F.; Belew, R.K.; Goodsell, D.S.; Olson, A.J. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J. Comput. Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef] [PubMed]
- Weston, R.J.; Mitchell, K.R.; Allen, K.L. Antibacterial phenolic components of New Zealand manuka honey. Food Chem. 1999, 64, 295–301. [Google Scholar] [CrossRef]
- Yao, L.; Datta, N.; Tomás-Barberán, F.A.; Ferreres, F.; Martos, I.; Singanusong, R. Flavonoids, phenolic acids and abscisic acid in Australian and New Zealand Leptospermum honeys. Food Chem. 2003, 81, 159–168. [Google Scholar] [CrossRef]
- Daher, S.; Gülaçar, F.O. Identification of New Aromatic Compounds in the New Zealand Manuka Honey by Gas Chromatography-Mass Spectrometry. E-J. Chem. 2010, 7, 7. [Google Scholar] [CrossRef]
- Khalil, M.; Alam, N.; Moniruzzaman, M.; Sulaiman, S.; Gan, S.H. Phenolic Acid Composition and Antioxidant Properties of Malaysian Honeys. J. Food Sci. 2011, 76, C921–C928. [Google Scholar] [CrossRef]
- Oelschlaegel, S.; Gruner, M.; Wang, P.-N.; Boettcher, A.; Koelling-Speer, I.; Speer, K. Classification and Characterization of Manuka Honeys Based on Phenolic Compounds and Methylglyoxal. J. Agric. Food Chem. 2012, 60, 7229–7237. [Google Scholar] [CrossRef]
- Sergiel, I.; Pohl, P.; Biesaga, M. Characterisation of honeys according to their content of phenolic compounds using high performance liquid chromatography/tandem mass spectrometry. Food Chem. 2014, 145, 404–408. [Google Scholar] [CrossRef]
- Rückriemen, J.; Schwarzenbolz, U.; Adam, S.; Henle, T. Identification and Quantitation of 2-Acetyl-1-pyrroline in Manuka Honey (Leptospermum scoparium). J. Agric. Food Chem. 2015, 63, 8488–8492. [Google Scholar] [CrossRef]
- Grainger, M.N.; Manley-Harris, M.; Lane, J.R.; Field, R.J. Kinetics of conversion of dihydroxyacetone to methylglyoxal in New Zealand mānuka honey: Part I—Honey systems. Food Chem. 2016, 202, 484–491. [Google Scholar] [CrossRef]
- Álvarez-Suarez, J.M.; Giampieri, F.; Cordero, M.; Gasparrini, M.; Forbes-Hernández, T.Y.; Mazzoni, L.; Afrin, S.; Beltrán-Ayala, P.; González-Paramás, A.M.; Santos-Buelga, C.; et al. Activation of AMPK/Nrf2 signalling by Manuka honey protects human dermal fibroblasts against oxidative damage by improving antioxidant response and mitochondrial function promoting wound healing. J. Funct. Foods 2016, 25, 38–49. [Google Scholar] [CrossRef]
- Daniels, B.J.; Prijic, G.; Meidinger, S.; Loomes, K.M.; Stephens, J.M.; Schlothauer, R.C.; Furkert, D.P.; Brimble, M.A. Isolation, Structural Elucidation, and Synthesis of Lepteridine from Mānuka (Leptospermum scoparium) Honey. J. Agric. Food Chem. 2016, 64, 5079–5084. [Google Scholar] [CrossRef]
- Russell, K.M.; Molan, P.C.; Wilkins, A.L.; Holland, P.T. Identification of some antibacterial constituents of New Zealand manuka honey. J. Agric. Food Chem. 1990, 38, 10–13. [Google Scholar] [CrossRef]
- Yin, Y.; Chen, H.; Chen, L.; Bie, X.; Ding, T.; Zhang, X.; Wu, B.; Shen, C.; Zhang, R. Determination of three coriaria lactones in honey by ultra high performance liquid chromatography-high resolution mass spectrometry. Chin. J. Chromatogr. 2015, 33, 711. [Google Scholar] [CrossRef] [PubMed]
- Allinger, N.L. ChemInform Abstract: Conformational Analysis. 130. MM2. A Hydrocarbon Force Field Utilizing V1 And V2 Torsional Terms. Chem. Informationsdienst 1978, 9, 130. [Google Scholar] [CrossRef]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2009, 31, 455–461. [Google Scholar] [CrossRef]
- Gasteiger, J.; Marsili, M. Iterative partial equalization of orbital electronegativity—A rapid access to atomic charges. Tetrahedron 1980, 36, 3219–3228. [Google Scholar] [CrossRef]
- García-Sosa, A.T.; Hetényi, C.; Maran, U. Drug efficiency indices for improvement of molecular docking scoring functions. J. Comput. Chem. 2010, 31, 174–184. [Google Scholar] [CrossRef]
- Mullai, V.; Menon, T. Bactericidal Activity of Different Types of Honey against Clinical and Environmental Isolates of Pseudomonas aeruginosa. J. Altern. Complement. Med. 2007, 13, 439–442. [Google Scholar] [CrossRef]



| Ligand (Origin) | Docking Score (kcal/mol) | Ligand Efficiency | Interacting Residues | Distance (Å) | Category | Type |
|---|---|---|---|---|---|---|
| Benzoic acid (M, H) | −5.4 | 0.60 | Tyr148 | 2.408 | H-Bond | Conventional |
| Pro20 | 1.778 | H-Bond | Conventional | |||
| Leu144 | 3.456 | Hydrophobic | Pi-Sigma | |||
| Val30 | 5.365 | Hydrophobic | Pi-Alkyl | |||
| Val61 | 4.581 | Hydrophobic | Pi-Alkyl | |||
| Leu63 | 4.592 | Hydrophobic | Pi-Alkyl | |||
| 5-methyl-3-furan-carboxylic acid (M) | −5.1 | 0.57 | Tyr148 | 2.414 | H-Bond | Conventional |
| Pro20 | 1.953 | H-Bond | Conventional | |||
| Val61 | 4.650 | Hydrophobic | Alkyl | |||
| Leu63 | 4.838 | Hydrophobic | Alkyl | |||
| Leu144 | 4.654 | Hydrophobic | Alkyl | |||
| Val61 | 4.713 | Hydrophobic | Alkyl | |||
| Leu63 | 4.688 | Hydrophobic | Alkyl | |||
| Methyl-glyoxal (M) | −2.8 | 0.56 | Ser22 | 2.996 | H-Bond | Conventional |
| Ser22 | 3.512 | H-Bond | Carbon H-Bond | |||
| Ser22 | 3.509 | H-Bond | Carbon H-Bond | |||
| 5-hydroxy-methyl-2-furaldehyde (M) | −5.0 | 0.56 | Tyr148 | 2.343 | H-Bond | Conventional |
| Pro20 | 1.922 | H-Bond | Conventional | |||
| Ser22 | 3.508 | H-Bond | Carbon H-Bond | |||
| Ser22 | 3.487 | H-Bond | Carbon H-Bond | |||
| Val30 | 4.982 | Hydrophobic | Alkyl | |||
| Val61 | 4.529 | Hydrophobic | Alkyl | |||
| Leu63 | 4.728 | Hydrophobic | Alkyl | |||
| Leu144 | 4.541 | Hydrophobic | Alkyl |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shirlaw, O.; Billah, Z.; Attar, B.; Hughes, L.; Qasaymeh, R.M.; Seidel, V.; Efthimiou, G. Antibiofilm Activity of Heather and Manuka Honeys and Antivirulence Potential of Some of Their Constituents on the DsbA1 Enzyme of Pseudomonas aeruginosa. Antibiotics 2020, 9, 911. https://doi.org/10.3390/antibiotics9120911
Shirlaw O, Billah Z, Attar B, Hughes L, Qasaymeh RM, Seidel V, Efthimiou G. Antibiofilm Activity of Heather and Manuka Honeys and Antivirulence Potential of Some of Their Constituents on the DsbA1 Enzyme of Pseudomonas aeruginosa. Antibiotics. 2020; 9(12):911. https://doi.org/10.3390/antibiotics9120911
Chicago/Turabian StyleShirlaw, Oscar, Zara Billah, Baraa Attar, Lisa Hughes, Rana M. Qasaymeh, Veronique Seidel, and Georgios Efthimiou. 2020. "Antibiofilm Activity of Heather and Manuka Honeys and Antivirulence Potential of Some of Their Constituents on the DsbA1 Enzyme of Pseudomonas aeruginosa" Antibiotics 9, no. 12: 911. https://doi.org/10.3390/antibiotics9120911
APA StyleShirlaw, O., Billah, Z., Attar, B., Hughes, L., Qasaymeh, R. M., Seidel, V., & Efthimiou, G. (2020). Antibiofilm Activity of Heather and Manuka Honeys and Antivirulence Potential of Some of Their Constituents on the DsbA1 Enzyme of Pseudomonas aeruginosa. Antibiotics, 9(12), 911. https://doi.org/10.3390/antibiotics9120911






