Comprehensive Statistical Evaluation of Etest®, UMIC®, MicroScan and Disc Diffusion versus Standard Broth Microdilution: Workflow for an Accurate Detection of Colistin-Resistant and Mcr-Positive E. coli
Abstract
1. Introduction
2. Results
3. Discussion
4. Materials and Methods
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- European Centre for Disease Prevention and Control. Antimicrobial Consumption in the EU/EEA, Annual Epidemiological Report for 2018; European Centre for Disease Prevention and Control: Solna, Sweden, 2019.
- Poirel, L.; Jayol, A.; Nordmann, P. Polymyxins: Antibacterial Activity, Susceptibility Testing, and Resistance Mechanisms Encoded by Plasmids or Chromosomes. Clin. Microbiol. Rev. 2017, 30, 557–596. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.-Y.; Wang, Y.; Walsh, T.R.; Yi, L.-X.; Zhang, R.; Spencer, J.D.; Doi, Y.; Tian, G.; Dong, B.; Huang, X.; et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: A microbiological and molecular biological study. Lancet Infect. Dis. 2016, 16, 161–168. [Google Scholar] [CrossRef]
- Díaz-Jiménez, D.; García-Meniño, I.; Fernández, J.; García, V.; Mora, A. Chicken and turkey meat: Consumer exposure to multidrug-resistant Enterobacteriaceae including mcr-carriers, uropathogenic E. coli and high-risk lineages such as ST131. Int. J. Food Microbiol. 2020, 331, 108750. [Google Scholar] [CrossRef] [PubMed]
- García, V.; García-Meniño, I.; Mora, A.; Flament-Simon, S.C.; Díaz-Jiménez, D.; Blanco, J.E.; Alonso, M.P.; Blanco, J. Co-occurrence of mcr-1, mcr-4 and mcr-5 genes in multidrug-resistant ST10 Enterotoxigenic and Shiga toxin-producing Escherichia coli in Spain (2006–2017). Int. J. Antimicrob. Agents 2018, 52, 104–108. [Google Scholar] [CrossRef] [PubMed]
- García-Meniño, I.; García, V.; Mora, A.; Díaz-Jiménez, D.; Flament-Simon, S.C.; Alonso, M.P.; Blanco, J.E.; Blanco, M.; Blanco, J. Swine Enteric Colibacillosis in Spain: Pathogenic Potential of mcr-1 ST10 and ST131 E. coli Isolates. Front. Microbiol. 2018, 9, 2659. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Feng, Y.; Liu, L.; Wei, L.; Kang, M.; Zong, Z. Identification of novel mobile colistin resistance gene mcr-10. Emerg. Microbes Infect. 2020, 9, 508–516. [Google Scholar] [CrossRef]
- Wang, R.; Van Dorp, L.; Shaw, L.; Bradley, P.; Wang, Q.; Wang, X.; Jin, L.; Zhang, Q.; Liu, Y.; Rieux, A.; et al. The global distribution and spread of the mobilized colistin resistance gene mcr-1. Nat. Commun. 2018, 9, 1–9. [Google Scholar] [CrossRef]
- Chen, L.; Zhang, J.; Wang, J.; Butaye, P.; Kelly, P.; Li, M.; Yang, F.; Gong, J.; Yassin, A.K.; Guo, W.; et al. Newly identified colistin resistance genes, mcr-4 and mcr-5, from upper and lower alimentary tract of pigs and poultry in China. PLoS ONE 2018, 13, e0193957. [Google Scholar] [CrossRef]
- Lutgring, J.D.; Kim, A.; Campbell, D.; Karlsson, M.; Brown, A.C.; Burd, E.M. Evaluation of the MicroScan Colistin Well and Gradient Diffusion Strips for Colistin Susceptibility Testing in Enterobacteriaceae. J. Clin. Microbiol. 2019, 57, e01866-18. [Google Scholar] [CrossRef]
- Matuschek, E.; Åhman, J.; Webster, C.; Kahlmeter, G. Antimicrobial susceptibility testing of colistin—evaluation of seven commercial MIC products against standard broth microdilution for Escherichia coli, Klebsiella pneumoniae, Pseudomonas aeruginosa, and Acinetobacter spp. Clin. Microbiol. Infect. 2018, 24, 865–870. [Google Scholar] [CrossRef]
- Sekyere, J.O.; Sephofane, A.K.; Mbelle, N.M. Comparative Evaluation of CHROMagar COL-APSE, MicroScan Walkaway, ComASP Colistin, and Colistin MAC Test in Detecting Colistin-resistant Gram-Negative Bacteria. Sci. Rep. 2020, 10, 6221. [Google Scholar] [CrossRef] [PubMed]
- Pfennigwerth, N.; Kaminski, A.; Korte-Berwanger, M.; Pfeifer, Y.; Simon, M.; Werner, G.; Jantsch, J.; Marlinghaus, L.; Gatermann, S.G. Evaluation of six commercial products for colistin susceptibility testing in Enterobacterales. Clin. Microbiol. Infect. 2019, 25, 1385–1389. [Google Scholar] [CrossRef] [PubMed]
- Gales, A.C.; Reis, A.O.; Jones, R.N. Contemporary Assessment of Antimicrobial Susceptibility Testing Methods for Polymyxin B and Colistin: Review of Available Interpretative Criteria and Quality Control Guidelines. J. Clin. Microbiol. 2001, 39, 183–190. [Google Scholar] [CrossRef] [PubMed]
- CLSI. Verification of Commercial Microbial Identification and Antimicrobial Susceptibility Testing Systems, 1st ed.; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2015. [Google Scholar]
- EUCAST. The European Committee on Antimicrobial Susceptibility Testing. Breakpoint Tables for Interpretation of MICs and Zone Diameters. Available online: https://www.eucast.org/fileadmin/src/media/PDFs/EUCAST_files/Breakpoint_tables/v_10.0_Breakpoint_Tables.pdf (accessed on 3 August 2020).
- Clinical and Laboratory Standards Institute (CLSI). Performance Standards for Antimicrobial Susceptibility Testing, 30th ed.; CLSI supplement M100; CLSI: Wayne, PA, USA, 2020; ISBN 978-1-68440-066-9. [Google Scholar]
- Lo-Ten-Foe, J.R.; De Smet, A.M.G.A.; Diederen, B.M.W.; Kluytmans, J.A.J.W.; Van Keulen, P.H.J. Comparative Evaluation of the VITEK 2, Disk Diffusion, Etest, Broth Microdilution, and Agar Dilution Susceptibility Testing Methods for Colistin in Clinical Isolates, Including Heteroresistant Enterobacter cloacae and Acinetobacter baumannii Strains. Antimicrob. Agents Chemother. 2007, 51, 3726–3730. [Google Scholar] [CrossRef] [PubMed]
- Maalej, S.; Meziou, M.; Rhimi, F.; Hammami, A. Comparison of disc diffusion, Etest and agar dilution for susceptibility testing of colistin against Enterobacteriaceae. Lett. Appl. Microbiol. 2011, 53, 546–551. [Google Scholar] [CrossRef]
- Chew, K.L.; La, M.-V.; Lin, R.T.P.; Teo, J.W.P. Colistin and Polymyxin B Susceptibility Testing for Carbapenem-Resistant and mcr -Positive Enterobacteriaceae: Comparison of Sensititre, MicroScan, Vitek 2, and Etest with Broth Microdilution. J. Clin. Microbiol. 2017, 55, 2609–2616. [Google Scholar] [CrossRef]
- Jayol, A.; Nordmann, P.; Lehours, P.; Poirel, L.; Dubois, V. Comparison of methods for detection of plasmid-mediated and chromosomally encoded colistin resistance in Enterobacteriaceae. Clin. Microbiol. Infect. 2018, 24, 175–179. [Google Scholar] [CrossRef]
- Zhang, H.; Zhao, D.-D.; Quan, J.; Hua, X.; Yu, Y. mcr-1 facilitated selection of high-level colistin-resistant mutants in Escherichia coli. Clin. Microbiol. Infect. 2019, 25, 517.e1–517.e4. [Google Scholar] [CrossRef]
- Zając, M.; Sztromwasser, P.; Bortolaia, V.; Leekitcharoenphon, P.; Cavaco, L.M.; Ziȩtek-Barszcz, A.; Hendriksen, R.S.; Wasyl, D. Occurrence and Characterization of mcr-1-Positive Escherichia coli Isolated from Food-Producing Animals in Poland, 2011–2016. Front. Microbiol. 2019, 10, 1753. [Google Scholar] [CrossRef]
- Pillonetto, M.; Mazzetti, A.; Becker, G.N.; Siebra, C.A.; Arend, L.N.; Barth, A.L. Low level of polymyxin resistance among nonclonal mcr-1–positive Escherichia coli from human sources in Brazil. Diagn. Microbiol. Infect. Dis. 2019, 93, 140–142. [Google Scholar] [CrossRef]
- Manges, A.R.; Geum, H.M.; Guo, A.; Edens, T.J.; Fibke, C.D.; Pitout, J.D.D. Global Extraintestinal Pathogenic Escherichia coli (ExPEC) Lineages. Clin. Microbiol. Rev. 2019, 32. [Google Scholar] [CrossRef] [PubMed]
- Teo, J.W.P.; Kalisvar, M.; Venkatachalam, I.; Ng, O.T.; Lin, R.T.P.; Octavia, S. mcr-3 and mcr-4 Variants in Carbapenemase-Producing Clinical Enterobacteriaceae Do Not Confer Phenotypic Polymyxin Resistance. J. Clin. Microbiol. 2017, 56, e01562-17. [Google Scholar] [CrossRef] [PubMed]
- Díaz-Jiménez, D.; García-Meniño, I.; Herrera, A.; Lestón, L.; Mora, A. Microbiological risk assessment of Turkey and chicken meat for consumer: Significant differences regarding multidrug resistance, mcr or presence of hybrid aEPEC/ExPEC pathotypes of E. coli. Food Control. 2020, 2020. [Google Scholar] [CrossRef]
- García-Meniño, I.; Díaz-Jiménez, D.; García, V.; De Toro, M.; Flament-Simon, S.C.; Blanco, J.; Mora, A. Genomic Characterization of Prevalent mcr-1, mcr-4, and mcr-5 Escherichia coli Within Swine Enteric Colibacillosis in Spain. Front. Microbiol. 2019, 10, 2469. [Google Scholar] [CrossRef]
- Humphries, R.M.; Ambler, J.; Mitchell, S.L.; Castanheira, M.; Dingle, T.; Hindler, J.A.; Koeth, L.; Sei, K.; Hardy, D.; Zimmer, B.; et al. CLSI Methods Development and Standardization Working Group Best Practices for Evaluation of Antimicrobial Susceptibility Tests. J. Clin. Microbiol. 2018, 56. [Google Scholar] [CrossRef] [PubMed]
- Hunter, J.D. Matplotlib: A 2D Graphics Environment. Comput. Sci. Eng. 2007, 9, 90–95. [Google Scholar] [CrossRef]


| Methods (Cut-OFF Values) | No. of Isolates (%) Categorized as Resistant | No. of Isolates (%; CI) Exhibiting EA | No. of Isolates (%; CI) Exhibiting CA | No. of Isolates (%; CI) Exhibiting ME | No. of Isolates (%; CI) Exhibiting VME | Sensitivity (%) N = 62 | Specificity (%) N = 88 |
|---|---|---|---|---|---|---|---|
| disc diffusion (≤11 mm) | 54 (36) | NA | 134 (89.3; 84.4–94.2) | 4 (2.7; 0.1–5.3) | 12 (8; 3.7–12.3) | 50 (80.9) | 84 (95.4) |
| disc-diffusion (≤13 mm) | 76 (50.7) | NA | 136 (90.7; 86.1–95.3) | 14 (9.3; 4.7–13.9) | 0 (0; 0) | 62 (100) | 74 (84.1) |
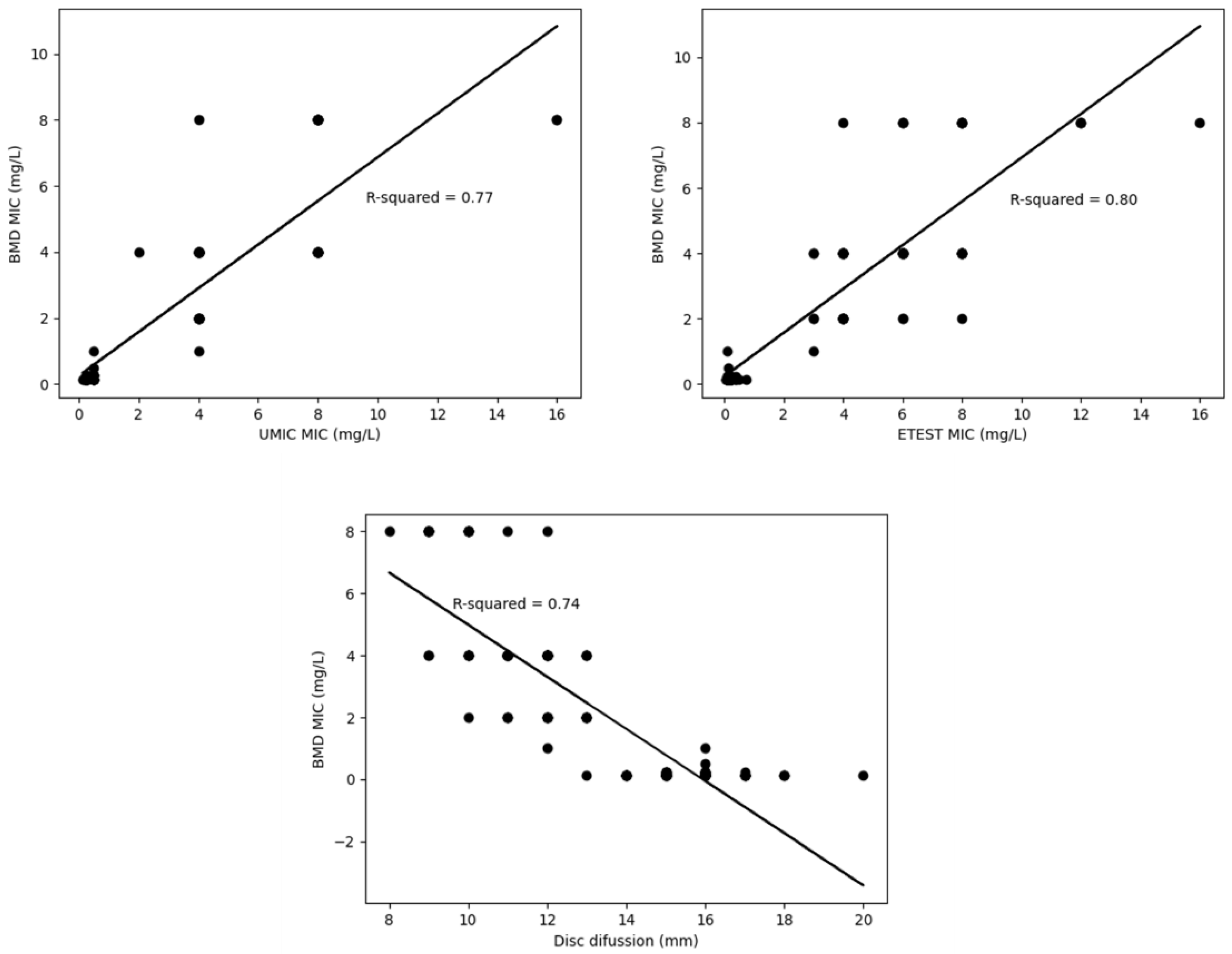
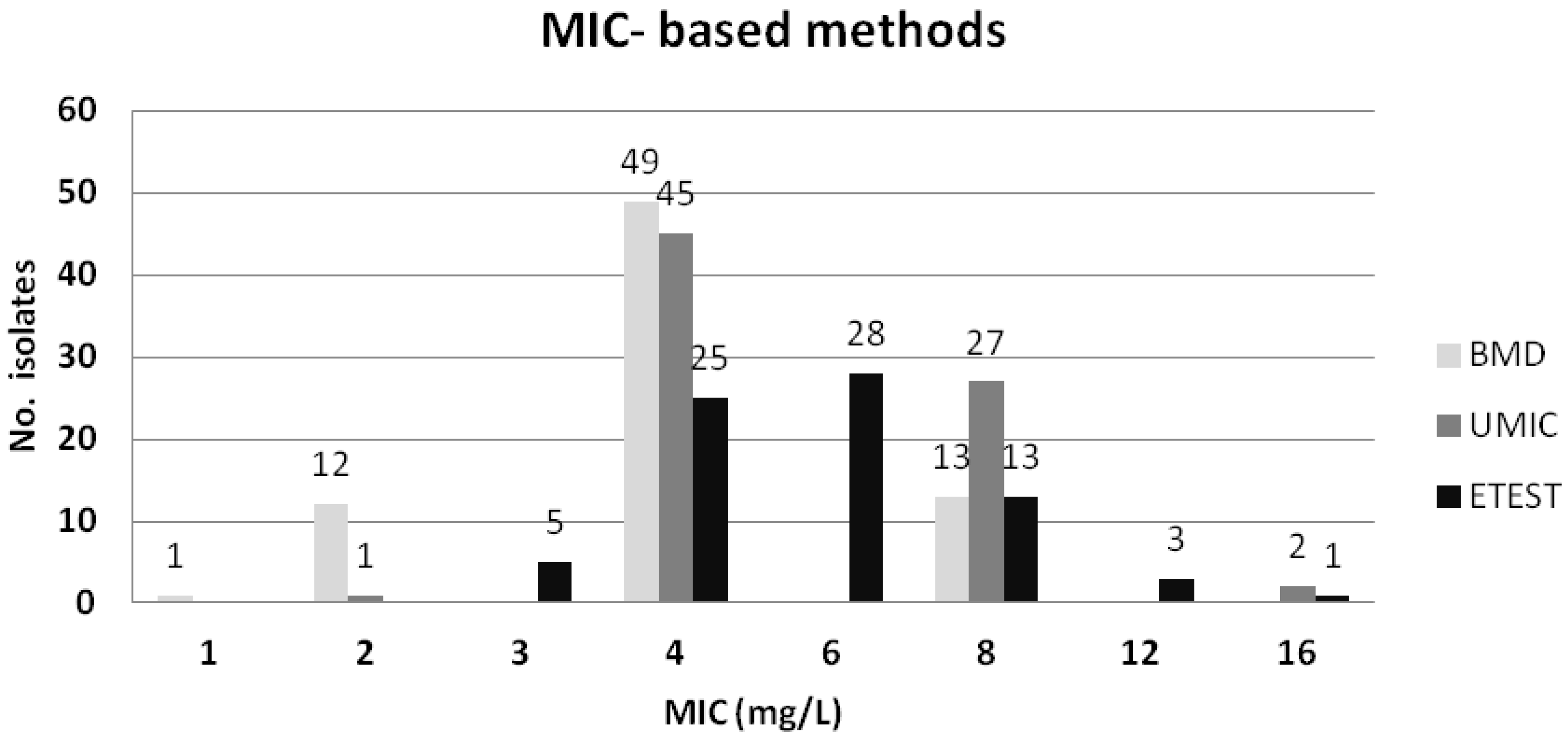
| UMIC® (>2 mg/L) | 74 (49.3) | 49 (32.7; 25.2–40.2) | 136 (90.7; 86.1–95.3) | 13 (8.7; 4.2–13.2) | 1 (0.7; 0–2) | 61 (98.4) | 75 (85.2) |
| Etest® (>2 mg/L) | 70 (46.7) | 55 (36.7; 29–44.4) | 137 (91.3; 86.6–95.8) | 13 (8.7; 4.2–13.2) | 0 (0; 0) | 62 (100) | 75 (85.2) |
| MicroScan (>2 mg/L) | 76 (50.7) | NA | 136 (90.7; 86.1–95.3) | 14 (9.3; 4.7–13.9) | 0 (0; 0) | 62 (100) | 74 (84.1) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
García-Meniño, I.; Lumbreras, P.; Valledor, P.; Díaz-Jiménez, D.; Lestón, L.; Fernández, J.; Mora, A. Comprehensive Statistical Evaluation of Etest®, UMIC®, MicroScan and Disc Diffusion versus Standard Broth Microdilution: Workflow for an Accurate Detection of Colistin-Resistant and Mcr-Positive E. coli. Antibiotics 2020, 9, 861. https://doi.org/10.3390/antibiotics9120861
García-Meniño I, Lumbreras P, Valledor P, Díaz-Jiménez D, Lestón L, Fernández J, Mora A. Comprehensive Statistical Evaluation of Etest®, UMIC®, MicroScan and Disc Diffusion versus Standard Broth Microdilution: Workflow for an Accurate Detection of Colistin-Resistant and Mcr-Positive E. coli. Antibiotics. 2020; 9(12):861. https://doi.org/10.3390/antibiotics9120861
Chicago/Turabian StyleGarcía-Meniño, Isidro, Pilar Lumbreras, Pablo Valledor, Dafne Díaz-Jiménez, Luz Lestón, Javier Fernández, and Azucena Mora. 2020. "Comprehensive Statistical Evaluation of Etest®, UMIC®, MicroScan and Disc Diffusion versus Standard Broth Microdilution: Workflow for an Accurate Detection of Colistin-Resistant and Mcr-Positive E. coli" Antibiotics 9, no. 12: 861. https://doi.org/10.3390/antibiotics9120861
APA StyleGarcía-Meniño, I., Lumbreras, P., Valledor, P., Díaz-Jiménez, D., Lestón, L., Fernández, J., & Mora, A. (2020). Comprehensive Statistical Evaluation of Etest®, UMIC®, MicroScan and Disc Diffusion versus Standard Broth Microdilution: Workflow for an Accurate Detection of Colistin-Resistant and Mcr-Positive E. coli. Antibiotics, 9(12), 861. https://doi.org/10.3390/antibiotics9120861







