Going beyond the Control of Quorum-Sensing to Combat Biofilm Infections
Abstract
:1. Pathogens in Biofilms Are Well Protected against Antibiotics and the Immune System
2. Communication Mediated by Small Molecules Is Required for Biofilm Formation and Maintenance
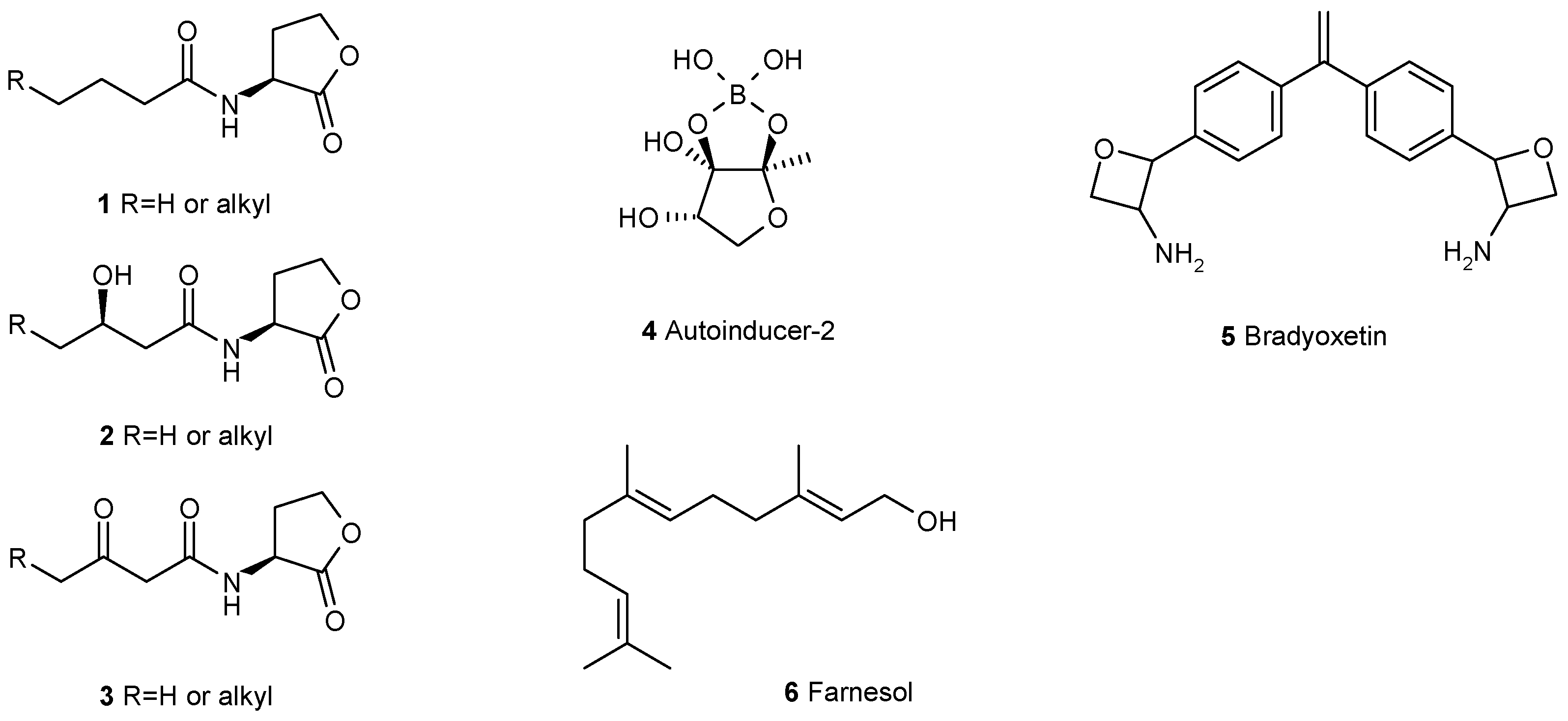
3. Interfering with the Communication between Microbial Cells Weakens Biofilms
3.1. Blocking Quorum Sensing
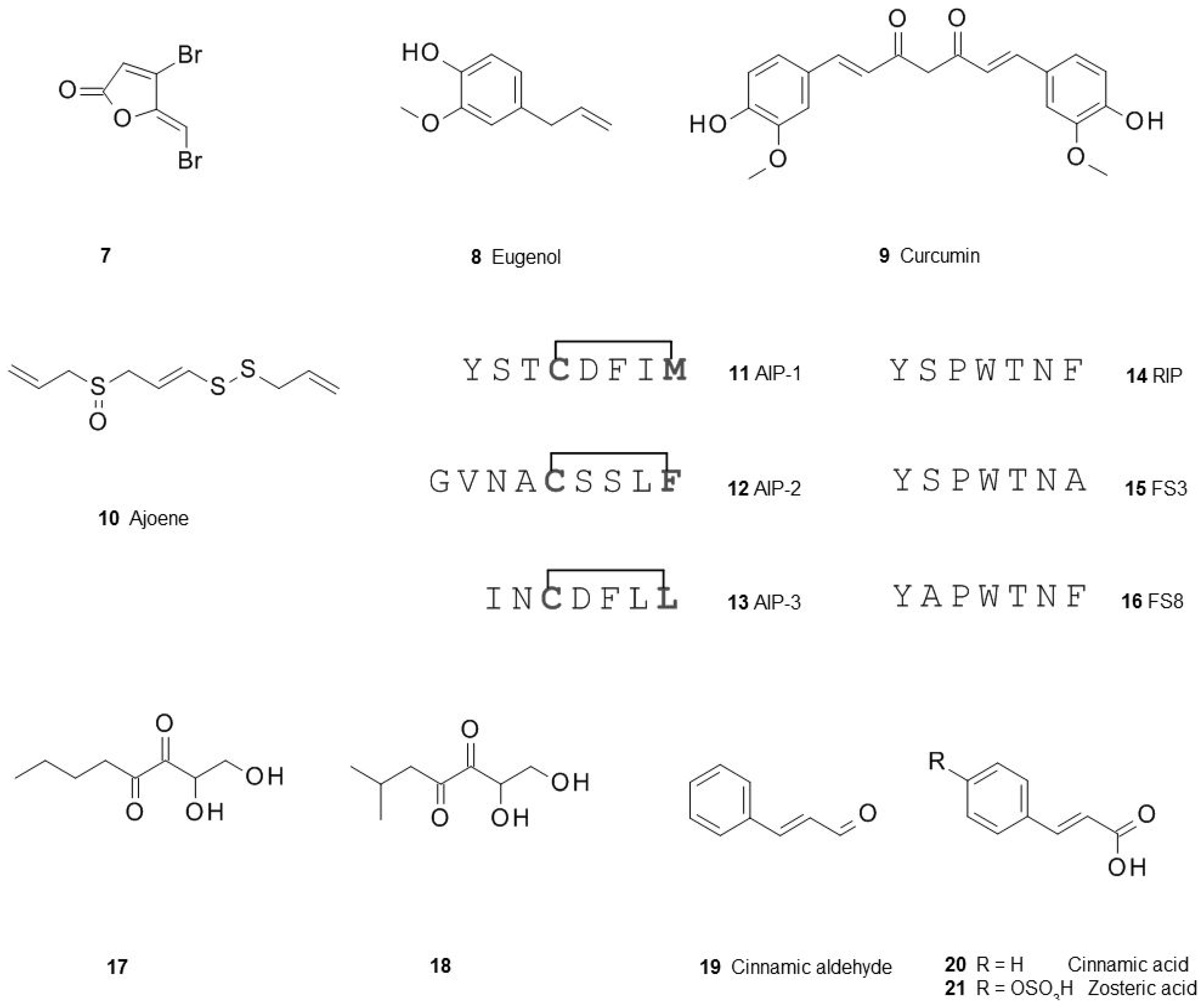
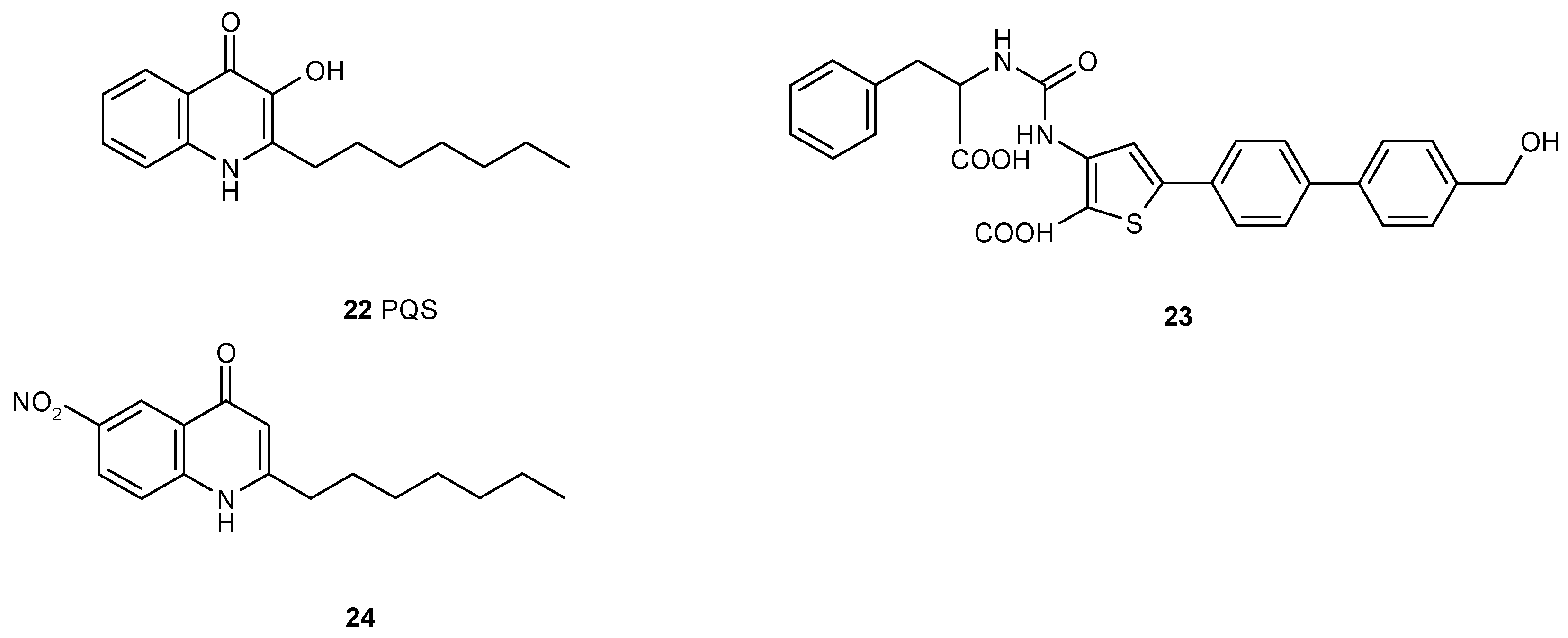
3.2. Multiple Interferences of Quorum Sensing Inhibitors with Biofilm Formation
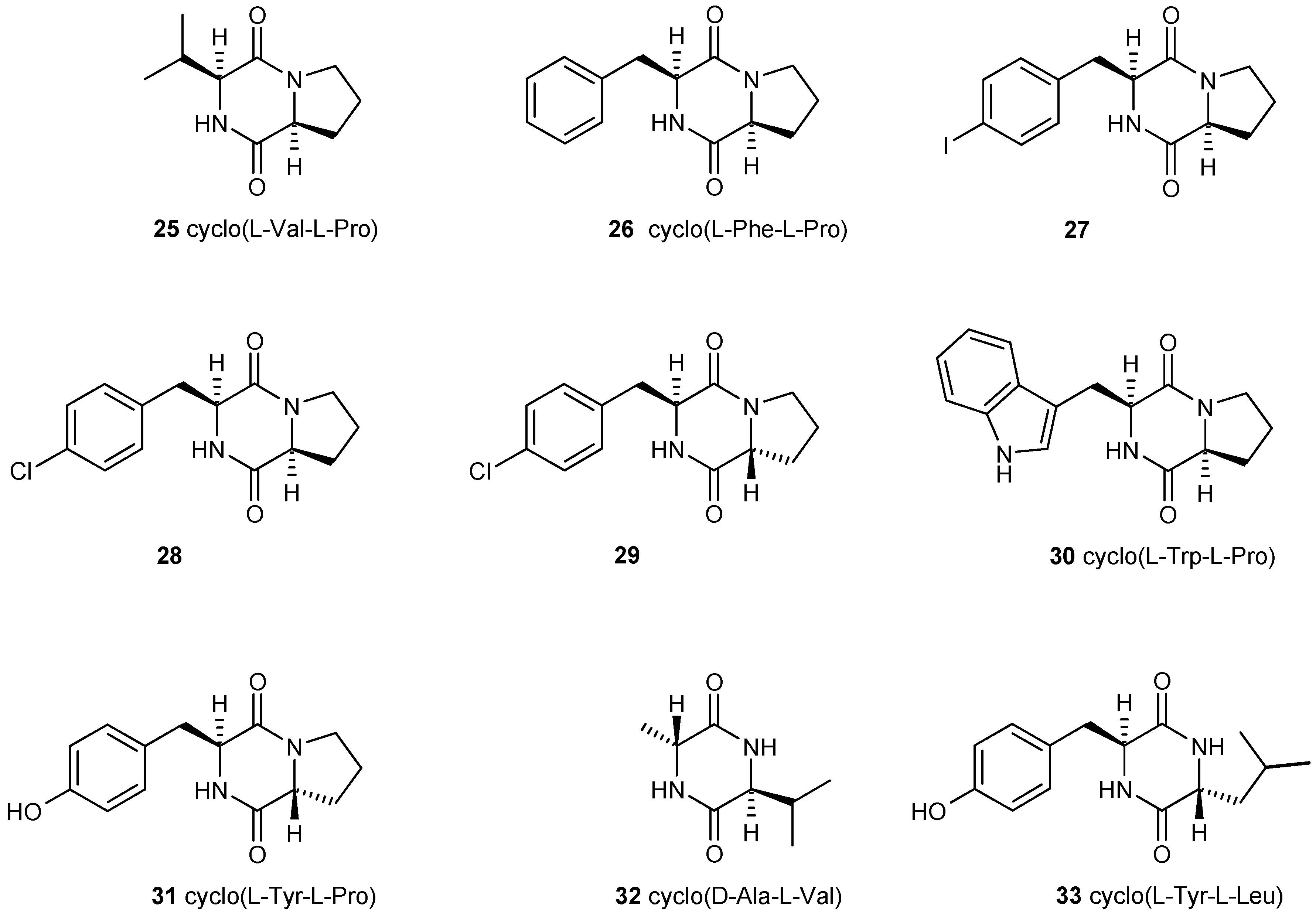
3.3. Dispersal of Biofilms
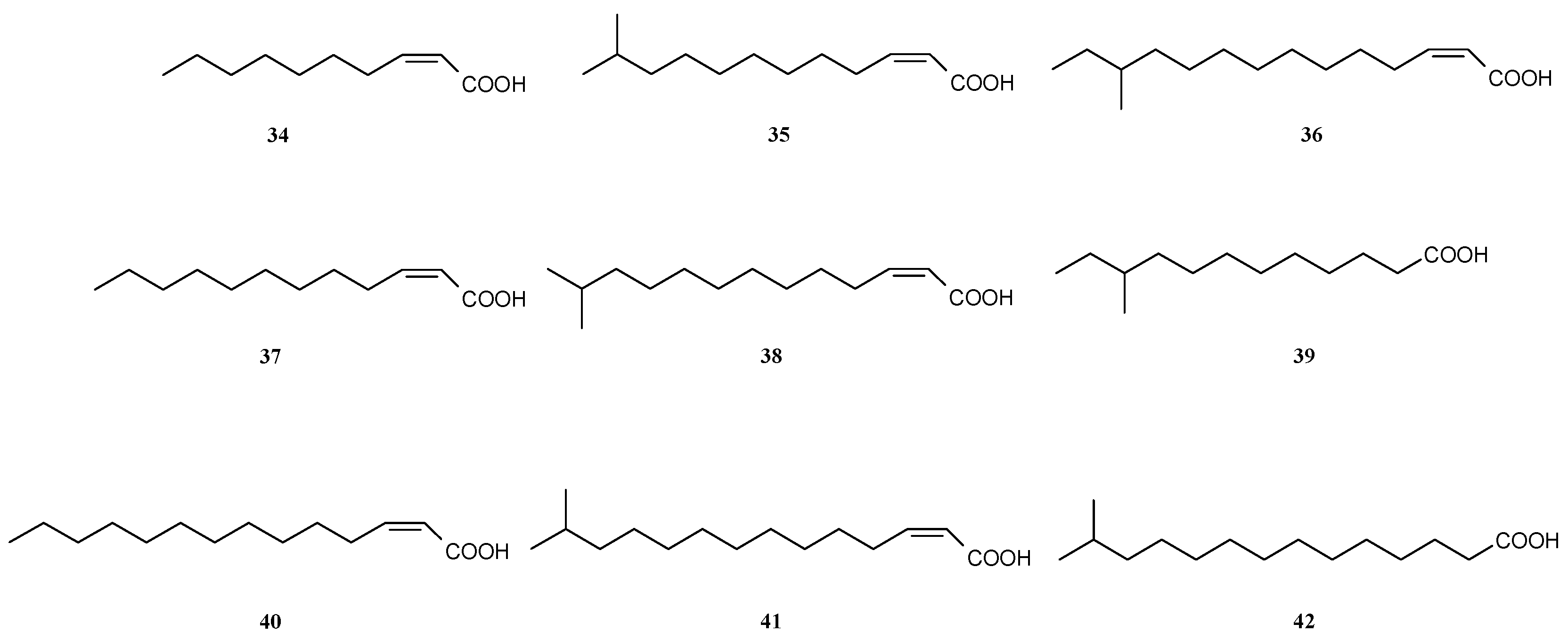
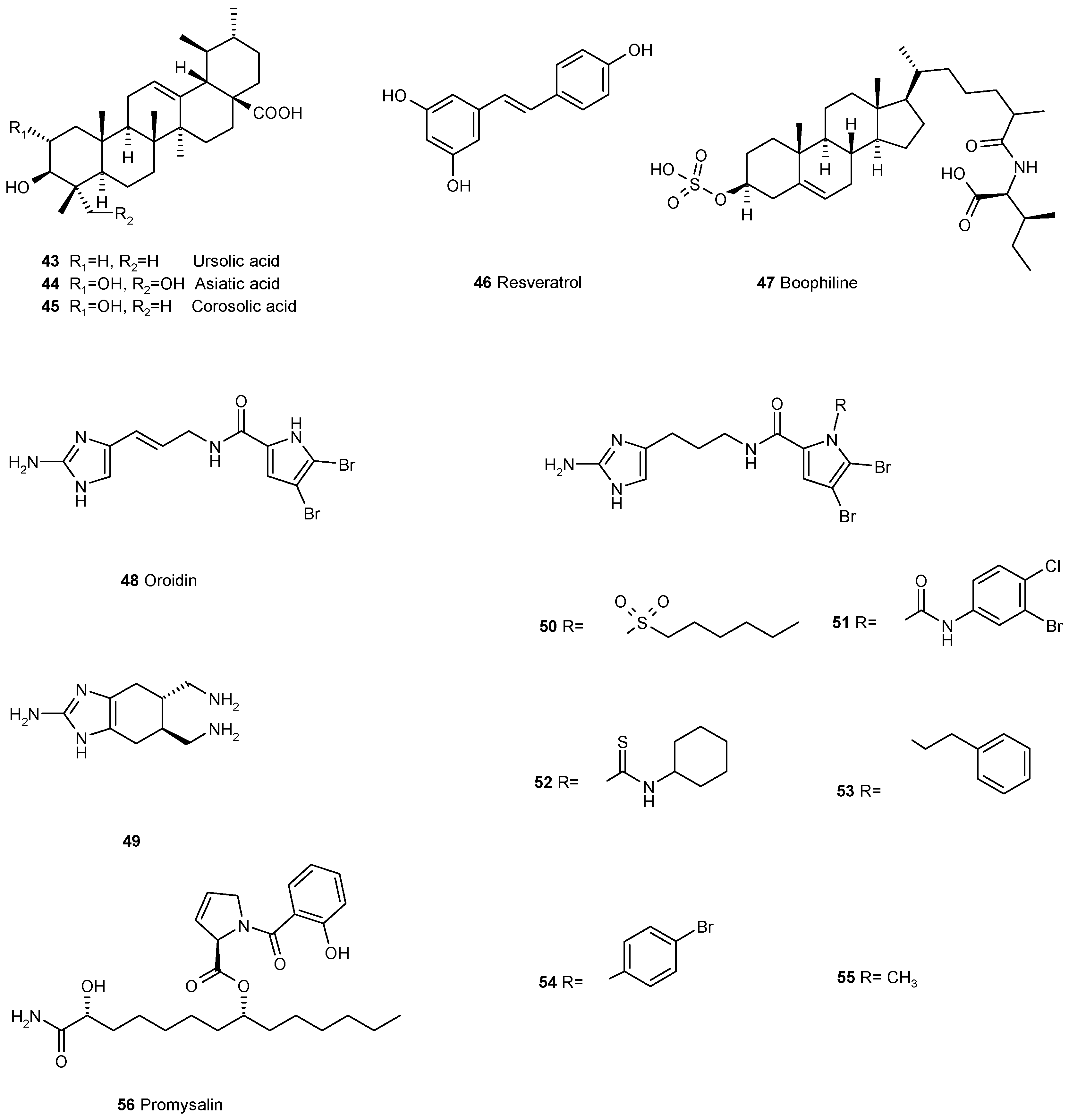
3.4. Other Ways of Biofilm Prevention or Dispersal
3.5. Combinations with Antibiotics
4. The Long Way to Go
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Costerton, J.W.; Cheng, K.J.; Geesey, G.G.; Ladd, T.I.; Nickel, J.C.; Dasgupta, M.; Marrie, T.J. Bacterial biofilms in nature and disease. Annu. Rev. Microbiol. 1987, 41, 435–464. [Google Scholar] [CrossRef] [PubMed]
- Singh, P.K.; Schaefer, A.L.; Parsek, M.R.; Moninger, T.O.; Welsh, M.J.; Greenberg, E.P. Quorum-sensing signals indicate that cystic fibrosis lungs are infected with bacterial biofilms. Nature 2000, 407, 762–764. [Google Scholar] [CrossRef] [PubMed]
- Wenzel, R.P. Health care-associated infections: Major issues in the early years of the 21st century. Clin. Infect. Dis. 2007, 45 (Suppl. 1), S85–S88. [Google Scholar] [CrossRef] [PubMed]
- Olsen, I. Biofilm-specific antibiotic tolerance and resistance. Eur. J. Clin. Microbiol. Infect. Dis. 2015, 34, 877–886. [Google Scholar] [CrossRef] [PubMed]
- Jolivet-Gougeon, A.; Bonnaure-Mallet, M. Biofilms as a mechanism of bacterial resistance. Drug Discov. Today Technol. 2014, 11, 49–56. [Google Scholar] [CrossRef] [PubMed]
- Cantón, R.; Morosini, M.I. Emergence and spread of antibiotic resistance following exposure to antibiotics. FEMS Microbiol. Rev. 2011, 35, 977–991. [Google Scholar] [CrossRef] [PubMed]
- Mah, T.-F.; Pitts, B.; Pellock, B.; Walker, G.C.; Stewart, P.S.; O’Toole, G.A. A genetic basis for Pseudomonas aeruginosa biofilm antibiotic resistance. Nature 2003, 426, 306–310. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Deng, Z.; Yan, A. Bacterial multidrug efflux pumps: Mechanisms, physiology and pharmacological exploitations. Biochem. Biophys. Res. Commun. 2014, 453, 254–267. [Google Scholar] [CrossRef] [PubMed]
- Waters, C.M.; Bassler, B.L. Quorum sensing: Cell-to-cell communication in bacteria. Annu. Rev. Cell Dev. Biol. 2005, 21, 319–346. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Ma, S. Small molecules modulating AHL-based quorum sensing to attenuate bacteria virulence and biofilms as promising antimicrobial drugs. Curr. Med. Chem. 2014, 21, 296–311. [Google Scholar] [CrossRef] [PubMed]
- Pereira, C.S.; Thompson, J.A.; Xavier, K.B. AI-2-mediated signalling in bacteria. FEMS Microbiol. Rev. 2013, 37, 156–181. [Google Scholar] [CrossRef] [PubMed]
- Loh, J.; Carlson, R.W.; York, W.S.; Stacey, G. Bradyoxetin, a unique chemical signal involved in symbiotic gene regulation. Proc. Natl. Acad. Sci. USA 2002, 99, 14446–14451. [Google Scholar] [CrossRef] [PubMed]
- Albuquerque, P.; Casadevall, A. Quorum sensing in fungi—A review. Med. Mycol. 2012, 50, 337–345. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Wu, J.; Tao, F.; Zhang, L.-H. Listening to a new language: DSF-based quorum sensing in Gram-negative bacteria. Chem. Rev. 2011, 111, 160–173. [Google Scholar] [CrossRef] [PubMed]
- Monnet, V.; Juillard, V.; Gardan, R. Peptide conversations in Gram-positive bacteria. Crit. Rev. Microbiol. 2014, 8. [Google Scholar] [CrossRef] [PubMed]
- Novick, R.P.; Geisinger, E. Quorum sensing in staphylococci. Annu. Rev. Genet. 2008, 42, 541–564. [Google Scholar] [CrossRef] [PubMed]
- Pichlmaier, M.; Marwitz, V.; Kühn, C.; Niehaus, M.; Klein, G.; Bara, C.; Haverich, A.; Abraham, W.-R. High prevalence of asymptomatic bacterial colonization of rhythm management devices. Europace 2008, 10, 1067–1072. [Google Scholar] [CrossRef] [PubMed]
- Scutera, S.; Zucca, M.; Savoia, D. Novel approaches for the design and discovery of quorum-sensing inhibitors. Expert Opin. Drug Discov. 2014, 9, 353–366. [Google Scholar] [CrossRef] [PubMed]
- Jimenez, P.N.; Koch, G.; Thompson, J.A.; Xavier, K.B.; Cool, R.H.; Quax, W.J. The multiple signaling systems regulating virulence in Pseudomonas aeruginosa. Microbiol. Mol. Biol. Rev. 2012, 76, 46–65. [Google Scholar] [CrossRef] [PubMed]
- Hirakawa, H.; Tomita, H. Interference of bacterial cell-to-cell communication: A new concept of antimicrobial chemotherapy breaks antibiotic resistance. Front. Microbiol. 2013, 4. [Google Scholar] [CrossRef] [PubMed]
- Jiang, T.; Li, M. Quorum sensing inhibitors: A patent review. Expert Opin. Ther. Pat. 2013, 23, 867–894. [Google Scholar] [CrossRef] [PubMed]
- De Nys, R.; Wright, A.D.; König, G.M.; Sticher, O. New halogenated furanones from the marine red alga Delisea pulchra (cf. fimbriata). Tetrahedron 1993, 49, 11213–11220. [Google Scholar] [CrossRef]
- Estrela, A.B.; Heck, M.G.; Abraham, W.-R. Novel approaches to control biofilm infections. Curr. Med. Chem. 2009, 16, 1512–1530. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Zheng, H.; Tang, Y.; Yu, W.; Gong, Q. Eugenol inhibits quorum sensing at sub-inhibitory concentrations. Biotechnol. Lett. 2013, 35, 631–637. [Google Scholar] [CrossRef] [PubMed]
- Rudrappa, T.; Bais, H.P. Curcumin, a known phenolic from Curcuma longa, attenuates the virulence of Pseudomonas aeruginosa PAO1 in whole plant and animal pathogenicity models. J. Agric. Food Chem. 2008, 56, 1955–1962. [Google Scholar] [CrossRef] [PubMed]
- Jakobsen, T.H.; van Gennip, M.; Phipps, R.K.; Shanmugham, M.S.; Christensen, L.D.; Alhede, M.; Skindersoe, M.E.; Rasmussen, T.B.; Friedrich, K.; Uthe, F.; et al. Ajoene, a sulfur-rich molecule from garlic, inhibits genes controlled by quorum sensing. Antimicrob. Agents Chemother. 2012, 56, 2314–2325. [Google Scholar] [CrossRef] [PubMed]
- Gray, B.; Hall, P.; Gresham, H. Targeting agr- and agr-Like quorum sensing systems for development of common therapeutics to treat multiple gram-positive bacterial infections. Sensors 2013, 13, 5130–5166. [Google Scholar] [CrossRef] [PubMed]
- Balaban, N.; Cirioni, O.; Giacometti, A.; Ghiselli, R.; Braunstein, J.B.; Silvestri, C.; Mocchegiani, F.; Saba, V.; Scalise, G. Treatment of Staphylococcus aureus biofilm infection by the quorum-sensing inhibitor RIP. Antimicrob. Agents Chemother. 2007, 51, 2226–2229. [Google Scholar] [CrossRef] [PubMed]
- Cirioni, O.; Mocchegiani, F.; Cacciatore, I.; Vecchiet, J.; Silvestri, C.; Baldassarre, L.; Ucciferri, C.; Orsetti, E.; Castelli, P.; Provinciali, M.; et al. Quorum sensing inhibitor FS3-coated vascular graft enhances daptomycin efficacy in a rat model of staphylococcal infection. Peptides 2013, 40, 77–81. [Google Scholar] [CrossRef] [PubMed]
- Simonetti, O.; Cirioni, O.; Mocchegiani, F.; Cacciatore, I.; Silvestri, C.; Baldassarre, L.; Orlando, F.; Castelli, P.; Provinciali, M.; Vivarelli, M.; et al. The efficacy of the quorum sensing inhibitor FS8 and tigecycline in preventing prosthesis biofilm in an animal model of staphylococcal infection. Int. J. Mol. Sci. 2013, 14, 16321–16332. [Google Scholar] [CrossRef] [PubMed]
- Thompson, J.A.; Oliveira, R.A.; Djukovic, A.; Ubeda, C.; Xavier, K.B. Manipulation of the quorum sensing signal AI-2 affects the antibiotic-treated gut microbiota. Cell Rep. 2015, 10, 1861–1871. [Google Scholar] [CrossRef] [PubMed]
- Roy, V.; Smith, J.A.I.; Wang, J.; Stewart, J.E.; Bentley, W.E.; Sintim, H.O. Synthetic analogs tailor native AI-2 signaling across bacterial species. J. Am. Chem. Soc. 2010, 132, 11141–11150. [Google Scholar] [CrossRef] [PubMed]
- Brackman, G.; Celen, S.; Hillaert, U.; van Calenbergh, S.; Cos, P.; Maes, L.; Nelis, H.J.; Coenye, T. Structure-activity relationship of cinnamaldehyde analogs as inhibitors of AI-2 based quorum sensing and their effect on virulence of Vibrio spp. PLoS ONE 2011, 6, e16084. [Google Scholar] [CrossRef] [PubMed]
- Brackman, G.; Defoirdt, T.; Miyamoto, C.; Bossier, P.; Calenbergh, S.V.; Nelis, H.; Coenyie, T. Cinnamaldehyde and cinnamaldehyde derivatives reduce virulence in Vibrio spp. by decreasing the DNA-binding activity of the quorum sensing response regulator LuxR. BMC Microbiol. 2008, 8, 149–162. [Google Scholar] [CrossRef] [PubMed]
- Lu, C.; Kirsch, B.; Zimmer, C.; de Jong, J.C.; Henn, C.; Maurer, C.K.; Müsken, M.; Häussler, S.; Steinbach, A.; Hartmann, R.W. Discovery of antagonists of PqsR, a key player in 2-alkyl-4-quinolone-dependent quorum sensing in Pseudomonas aeruginosa. Chem. Biol. 2012, 19, 381–390. [Google Scholar] [CrossRef] [PubMed]
- Sahner, J.H.; Empting, M.; Kamal, A.; Weidel, E.; Groh, M.; Börger, C.; Hartmann, R.W. Exploring the chemical space of ureidothiophene-2-carboxylic acids as inhibitors of the quorum sensing enzyme PqsD from Pseudomonas aeruginosa. Eur. J. Med. Chem. 2015, 96, 14–21. [Google Scholar] [CrossRef] [PubMed]
- Defoirdt, T.; Brackman, G.; Coenye, T. Quorum sensing inhibitors: How strong is the evidence? Trends Microbiol. 2013, 21, 619–624. [Google Scholar] [CrossRef] [PubMed]
- De Carvalho, M.P.; Abraham, W.-R. Antimicrobial and biofilm inhibiting diketopiperazines. Curr. Med. Chem. 2012, 19, 3564–3577. [Google Scholar] [CrossRef] [PubMed]
- Holden, M.T.G.; Chhabra, S.R.; de Nys, R.; Stead, P.; Bainton, N.J.; Hill, P.J.; Manefeld, M.; Kumar, N.; Labatte, M.; England, D.; et al. Quorum-sensing cross talk: Isolation and chemical characterization of cyclic dipeptides from Pseudomonas aeruginosa and other Gram-negative bacteria. Mol. Microbiol. 1999, 33, 1254–1266. [Google Scholar] [CrossRef] [PubMed]
- Campbell, J.; Lin, Q.; Geske, G.D.; Blackwell, H.E. New and unexpected insights into the modulation of LuxR-type quorum sensing by cyclic dipeptides. ACS Chem. Biol. 2009, 4, 1051–1059. [Google Scholar] [CrossRef] [PubMed]
- Ryan, R.P.; Dow, J.M. Diffusible signals and interspecies communication in bacteria. Microbiology 2008, 154, 1845–1858. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Wang, W.; Xu, S.X.; Magarvey, N.A.; McCormick, J.K. Lactobacillus reuteri-produced cyclic dipeptides quench agr-mediated expression of toxic shock syndrome toxin-1 in staphylococci. Proc. Natl. Acad. Sci. USA 2011, 108, 3360–3365. [Google Scholar] [CrossRef] [PubMed]
- Scopel, M.; Abraham, W.-R.; Henriques, A.T.; Macedo, A.J. Dipeptide cis-cyclo(leucyl-tyrosyl) produced by sponge associated Penicillium sp.F37 inhibits biofilm formation of the pathogenic Staphylococcus epidermidis. Bioorganic Med. Chem. Lett. 2013, 23, 624–626. [Google Scholar] [CrossRef] [PubMed]
- Brackman, G.; Hillaert, U.; Calenbergh, S.V.; Nelis, H.J.; Coenye, T. Use of quorum sensing inhibitors to interfere with biofilm formation and development in Burkholderia multivorans and Burkholderia cenocepacia. Res. Microbiol. 2009, 160, 144–151. [Google Scholar] [CrossRef] [PubMed]
- Song, J.; Choi, B.; Jin, E.J.; Yoon, Y.; Choi, K.H. Curcumin suppresses Streptococcus mutans adherence to human tooth surfaces and extracellular matrix proteins. Eur. J. Clin. Microbiol. Infect. Dis. 2012, 31, 1347–1352. [Google Scholar] [CrossRef] [PubMed]
- Hu, P.; Huang, P.; Chen, M.W. Curcumin reduces Streptococcus mutans biofilm formation by inhibiting sortase A activity. Arch. Oral Biol. 2013, 58, 1343–1348. [Google Scholar] [CrossRef] [PubMed]
- Arora, D.P.; Hossain, S.; Xu, Y.; Boon, E.M. Nitric oxide regulation of bacterial biofilms. Biochemistry 2015, 54, 3717–3728. [Google Scholar] [CrossRef] [PubMed]
- Barraud, N.; Hassett, D.J.; Hwang, S.H.; Rice, S.A.; Kjelleberg, S.; Webb, J.S. Involvement of nitric oxide in biofilm dispersal of Pseudomonas aeruginosa. J. Bacteriol. 2006, 188, 7344–7353. [Google Scholar] [CrossRef] [PubMed]
- Potter, A.J.; Kidd, S.P.; Edwards, J.L.; Falsetta, M.L.; Apicella, M.A.; Jennings, M.P.; McEwan, A.G. Thioredoxin reductase is essential for protection of Neisseria gonorrhoeae against killing by nitric oxide and for bacterial growth during interaction with cervical epithelial cells. J. Infect. Dis. 2009, 199, 227–235. [Google Scholar] [CrossRef] [PubMed]
- Schlag, S.; Nerz, C.; Birkenstock, T.A.; Altenberend, F.; Götz, F. Inhibition of staphylococcal biofilm formation by nitrite. J. Bacteriol. 2007, 189, 7911–7919. [Google Scholar] [CrossRef] [PubMed]
- Barraud, N.; Storey, M.V.; Moore, Z.P.; Webb, J.S.; Rice, S.A.; Kjelleberg, S. Nitric oxide-mediated dispersal in single- and multi-species biofilms of clinically and industrially relevant microorganisms. Microb. Biotechnol. 2009, 2, 370–378. [Google Scholar] [CrossRef] [PubMed]
- Wilken, M.; Huchzermeyer, B. Suppression of mycelia formation by NO produced endogenously in Candida tropicalis. Eur. J. Cell Biol. 1999, 78, 209–213. [Google Scholar] [CrossRef]
- Babaei, S.; Teichert-Kuliszewska, K.; Monge, J.C.; Mohamed, F.; Bendeck, M.P.; Stewart, D.J. Role of nitric oxide in the angiogenic response in vitro to basic fibroblast growth factor. Circ. Res. 1998, 82, 1007–1015. [Google Scholar] [CrossRef] [PubMed]
- Ridnour, L.A.; Thomas, D.D.; Mancardi, D.; Espey, M.G.; Miranda, K.M.; Paolocci, N.; Feelisch, M.; Fukuto, J.; Wink, D.A. The chemistry of nitrosative stress induced by nitric oxide and reactive nitrogen oxide species. Putting perspective on stressful biological situations. Biol. Chem. 2004, 385, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Miller, C. Inhaled nitric oxide. In Proceedings of the 36th European Cystic Fibrosis Conference, Lisbon, Portugal, 12–15 June 2013; Available online: http://www.hopkinscme.net/ofp/eCysticFibrosisReview/newsletters/2013/0913.html (accessed on 5 January 2016).
- Keefer, L.K. Fifty years of diazeniumdiolate research. From laboratory curiosity to broad-spectrum biomedical advances. ACS Chem. Biol. 2011, 6, 1147–1155. [Google Scholar] [CrossRef] [PubMed]
- Barraud, N.; Kardak, B.G.; Yepuri, N.R.; Howlin, R.P.; Webb, J.S.; Faust, S.N.; Kjelleberg, S.; Rice, S.A.; Kelso, M.J. Cephalosporin-3′-diazeniumdiolates: Targeted NO-donor prodrugs for dispersing bacterial biofilms. Angew. Chem. Int. Ed. 2012, 51, 9057–9060. [Google Scholar] [CrossRef] [PubMed]
- Sun, B.; Slomberg, D.L.; Chudasama, S.L.; Lu, Y.; Schoenfisch, M.H. Nitric oxide-releasing dendrimers as antibacterial agents. Biomacromolecules 2012, 13, 3343–3354. [Google Scholar] [CrossRef] [PubMed]
- Nablo, B.J.; Rothrock, A.R.; Schoenfisch, M.H. Nitric oxide-releasing sol-gels as antibacterial coatings for orthopedic implants. Biomaterials 2005, 26, 917–924. [Google Scholar] [CrossRef] [PubMed]
- Barraud, N.; Kelso, M.J.; Rice, S.A.; Kjelleberg, S. Nitric oxide: A key mediator of biofilm dispersal with applications in infectious diseases. Curr. Pharm. Des. 2015, 21, 31–42. [Google Scholar] [CrossRef] [PubMed]
- Davies, D.G.; Marques, C.N.H. A fatty acid messenger is responsible for inducing dispersion in microbial biofilms. J. Bacteriol. 2009, 191, 1393–1403. [Google Scholar] [CrossRef] [PubMed]
- Ryan, R.P.; McCarthy, Y.; Watt, S.A.; Niehaus, K.; Dow, J.M. Intraspecies signaling involving the diffusible signal factor BDSF (cis-2-dodecenoic acid) influences virulence in Burkholderia cenocepacia. J. Bacteriol. 2009, 191, 5013–5019. [Google Scholar] [CrossRef] [PubMed]
- Udine, C.; Brackman, G.; Bazzini, S.; Buroni, S.; van Acker, H.; Pasca, M.R.; Riccardi, G.; Coenye, T. Phenotypic and genotypic characterisation of Burkholderia cenocepacia J2315 mutants affected in homoserine lactone and diffusible signal factor-based quorum sensing systems suggests interplay between both types of systems. PLoS ONE 2013, 8, e55112. [Google Scholar] [CrossRef] [PubMed]
- Rahmani-Badi, A.; Sepehr, S.; Fallahi, H.; Heidari-Keshel, S. Dissection of the cis-2-decenoic acid signaling network in Pseudomonas aeruginosa using microarray technique. Front. Microbiol. 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Huang, T.P.; Wong, A.C. Extracellular fatty acids facilitate flagella-independent translocation by Stenotrophomonas maltophilia. Res. Microbiol. 2007, 158, 702–711. [Google Scholar] [CrossRef] [PubMed]
- Ryan, R.P.; Fouhy, Y.; Garcia, B.F.; Watt, S.A.; Niehaus, K.; Yang, L.; Tolker-Nielsen, T.; Dow, J.M. Interspecies signalling via the Stenotrophomonas maltophilia diffusible signal factor influences biofilm formation and polymyxin tolerance in Pseudomonas aeruginosa. Mol. Microbiol. 2008, 68, 75–86. [Google Scholar] [CrossRef] [PubMed]
- Inoue, T.; Shingaki, R.; Fukui, K. Inhibition of swarming motility of Pseudomonas aeruginosa by branched-chain fatty acids. FEMS Microbiol. Lett. 2008, 281, 81–86. [Google Scholar] [CrossRef] [PubMed]
- Boon, C.; Deng, Y.; Wang, L.-H.; He, Y.; Xu, J.-L.; Fan, Y.; Pan, S.Q.; Zhang, L.-H. A novel DSF-like signal from Burkholderia cenocepacia interferes with Candida albicans morphological transition. ISME J. 2008, 2, 27–36. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Boon, C.; Chen, S.; Lim, A.; Zhang, L.-H. Cis-2-dodecenoic acid signal modulates virulence of Pseudomonas aeruginosa through interference with quorum sensing systems and T3SS. BMC Microbiol. 2013, 13. [Google Scholar] [CrossRef] [PubMed]
- Amari, D.T.; Marques, C.N.H.; Davies, D.G. The putative enoyl-coenzyme A hydratase DspI is required for production of the Pseudomonas aeruginosa biofilm dispersion autoinducer cis-2-decenoic acid. J. Bacteriol. 2013, 195, 4600–4610. [Google Scholar] [CrossRef] [PubMed]
- Bi, H.; Christensen, Q.H.; Feng, Y.; Wang, H.; Cronan, J.E. The Burkholderia cenocepacia BDSF quorum sensing fatty acid is synthesized by a bifunctional crotonase homologue having both dehydratase and thioesterase activities. Mol. Microbiol. 2012, 83, 840–855. [Google Scholar] [CrossRef] [PubMed]
- Ryan, R.P.; Dow, J.M. Communication with a growing family: Diffusible signal factor (DSF) signaling in bacteria. Trends Microbiol. 2011, 19, 145–152. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Schmid, N.; Wang, C.; Wang, J.; Pessi, G.; Wu, D.; Lee, J.; Aguilar, C.; Ahrens, C.H.; Chang, C.; et al. Cis-2-dodecenoic acid receptor RpfR links quorum-sensing signal perception with regulation of virulence through cyclic dimeric guanosine monophosphate turnover. Proc. Natl. Acad. Sci. USA. 2012, 109, 15479–15484. [Google Scholar] [CrossRef] [PubMed]
- Huedo, P.; Yero, D.; Martínez-Servat, S.; Estibariz, I.; Planell, R.; Martínez, P.; Ruyra, A.; Roher, N.; Roca, I.; Vila, J.; et al. Two different rpf clusters distributed among a population of Stenotrophomonas maltophilia clinical strains display differential diffusible signal factor production and virulence regulation. J. Bacteriol. 2014, 196, 2431–2442. [Google Scholar] [CrossRef] [PubMed]
- Marques, C.N.H.; Morozov, A.; Planzos, P.; Zelaya, H.M. The fatty acid signaling molecule cis-2-decenoic acid increases metabolic activity and reverts persister cells to an antimicrobial-susceptible state. Appl. Environ. Microbiol. 2014, 80, 6976–6991. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez, M.; Choi, M.H.; Tian, B.; Xu, J.; Rho, J.K.; Kim, M.O.; Cho, Y.-H.; Yoon, S.C. Simultaneous inhibition of rhamnolipid and polyhydroxyalkanoic acid synthesis and biofilm formation in Pseudomonas aeruginosa by 2-bromoalkanoic acids: Effect of inhibitor alkyl-chain-length. PLoS ONE 2013, 8, e73986. [Google Scholar] [CrossRef] [PubMed]
- Su, Z.; Peng, L.; Worthington, R.J.; Melander, C. Evaluation of 4,5-disubstituted-2-aminoimidazole-triazole conjugates for antibiofilm/antibiotic resensitization activity against MRSA and Acinetobacter baumannii. ChemMedChem 2011, 6, 2243–2251. [Google Scholar] [CrossRef] [PubMed]
- Ren, D.; Zuo, R.; Barrios, A.F.G.; Bedzyk, L.A.; Eldridge, G.R.; Pasmore, M.E.; Wood, T.K. Differential gene expression for investigation of Escherichia coli biofilm inhibition by plant extract ursolic acid. Appl. Environ. Microbiol. 2005, 71, 4022–4034. [Google Scholar] [CrossRef] [PubMed]
- Qin, N.; Tan, X.; Jiao, Y.; Liu, L.; Zhao, W.; Yang, S.; Jia, A. RNA-Seq-based transcriptome analysis of methicillin-resistant Staphylococcus aureus biofilm inhibition by ursolic acid and resveratrol. Sci. Rep. 2014, 4, 5467. [Google Scholar] [CrossRef] [PubMed]
- Garo, E.; Eldridge, G.R.; Goering, M.G.; Pulcini, E.D.; Hamilton, M.A.; Costerton, J.W.; James, G.A. Asiatic acid and corosolic acid enhance the susceptibility of Pseudomonas aeruginosa biofilms to tobramycin. Antimicrob. Agents Chemother. 2007, 51, 1813–1817. [Google Scholar] [CrossRef] [PubMed]
- Potterat, O.; Hostettmann, K.; Höltzel, A.; Jung, G.; Diehl, P.A.; Petrini, O. Boophiline, an antimicrobial sterol amide from the cattle tick Boophilus microplus. Helv. Chim. Acta 1997, 80, 2066–2072. [Google Scholar] [CrossRef]
- Zimmer, K.R.; Macedo, A.J.; Giordani, R.B.; Conceição, J.M.; Nicastro, G.G.; Boechat, A.L.; Baldini, R.L.; Abraham, W.-R.; Termignoni, C. A steroidal molecule present in the egg wax of the tick Rhipicephalus (Boophilus) microplus inhibits bacterial biofilms. Environ. Microbiol. 2013, 15, 2008–2018. [Google Scholar] [CrossRef] [PubMed]
- Forenza, S.; Minale, L.; Riccio, R.; Fattorusso, E. New bromo-pyrrole derivatives from the sponge Agelas oroides. J. Chem. Soc. D 1971, 18, 1129–1130. [Google Scholar] [CrossRef]
- Kelly, S.R.; Jensen, P.R.; Henkel, T.P.; Fenical, W.; Pawlik, J.R. Effects of Caribbean sponge extracts on bacterial attachment. Aquat. Microb. Ecol. 2003, 31, 175–182. [Google Scholar] [CrossRef]
- Žula, A.; Kikelj, D.; Ilaš, J. 2-Aminoimidazoles in medicinal chemistry. Mini Rev. Med. Chem. 2013, 13, 1921–1943. [Google Scholar] [CrossRef] [PubMed]
- Richards, J.J.; Reyes, S.; Stowe, S.D.; Tucker, A.T.; Ballard, T.E.; Mathies, L.D.; Cavanagh, J.; Melander, C. Amide isosteres of oroidin: Assessment of antibiofilm activity and C. elegans toxicity. J. Med. Chem. 2009, 52, 4582–4585. [Google Scholar] [CrossRef] [PubMed]
- Ballard, T.E.; Richards, J.J.; Aquino, A.; Reed, C.S.; Melander, C. Antibiofilm activity of a diverse oroidin library generated through reductive acylation. J. Org. Chem. 2009, 74, 1755–1758. [Google Scholar] [CrossRef] [PubMed]
- Hodnik, Ž.; Łoś, J.M.; Žula, A.; Zidar, N.; Jakopin, Ž.; Łoś, M.; Dolenc, M.S.; Ilaš, J.; Węgrzyn, G.; Mašič, L.P.; et al. Inhibition of biofilm formation by conformationally constrained indole-based analogues of the marine alkaloid oroidin. Bioorganic Med. Chem. Lett. 2014, 24, 2530–2534. [Google Scholar] [CrossRef] [PubMed]
- Wright, C.J.; Wu, H.; Melander, R.J.; Melander, C.; Lamont, R.J. Disruption of heterotypic community development by Porphyromonas gingivalis with small molecule inhibitors. Mol. Oral Microbiol. 2014, 29, 185–193. [Google Scholar] [CrossRef] [PubMed]
- Polo, A.; Foladori, P.; Ponti, B.; Bettinetti, R.; Gambino, M.; Villa, F.; Cappitelli, F. Evaluation of zosteric acid for mitigating biofilm formation of Pseudomonas putida isolated from a membrane bioreactor system. Int. J. Mol. Sci. 2014, 15, 9497–9518. [Google Scholar] [CrossRef] [PubMed]
- Villa, F.; Pitts, B.; Stewart, P.S.; Giussani, B.; Roncoroni, S.; Albanese, D.; Giordano, C.; Tunesi, M.; Cappitelli, F. Efficacy of zosteric acid sodium salt on the yeast biofilm model Candida albicans. Microb. Ecol. 2011, 62, 584–598. [Google Scholar] [CrossRef] [PubMed]
- Villa, F.; Remelli, W.; Forlani, F.; Vitali, A.; Cappitelli, F. Altered expression level of Escherichia coli proteins in response to treatment with the antifouling agent zosteric acid sodium salt. Environ. Microbiol. 2012, 14, 1753–1761. [Google Scholar] [CrossRef] [PubMed]
- Cattò, C.; Dell’Orto, S.; Villa, F.; Villa, S.; Gelain, A.; Vitali, A.; Marzano, V.; Baroni, S.; Forlani, F.; Cappitelli, F. Unravelling the structural and molecular basis responsible for the anti-biofilm activity of zosteric acid. PLoS ONE 2015, 10, e0131519. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Estrada-de los Santos, P.; Matthijs, S.; Xie, G.L.; Busson, R.; Cornelis, P.; Rozenski, J.; de Mot, R. Promysalin, a salicylate-containing Pseudomonas putida antibiotic, promotes surface colonization and selectively targets other Pseudomonas. Chem. Biol. 2011, 18, 1320–1330. [Google Scholar] [CrossRef] [PubMed]
- Steele, A.D.; Knouse, K.W.; Keohane, C.E.; Wuest, W.M. Total synthesis and biological investigation of (−)-promysalin. J. Am. Chem. Soc. 2015, 137, 7314–7317. [Google Scholar] [CrossRef] [PubMed]
- Clatworthy, A.E.; Pierson, E.; Hung, D.T. Targeting virulence: A new paradigm for antimicrobial therapy. Nat. Chem. Biol. 2007, 3, 541–548. [Google Scholar] [CrossRef] [PubMed]
- Romilly, C.; Lays, C.; Tomasini, A.; Caldelari, I.; Benito, Y.; Hammann, P.; Geissmann, T.; Boisset, S.; Romby, P.; Vandenesch, F. A non-coding RNA promotes bacterial persistence and decreases virulence by regulating a regulator in Staphylococcus aureus. PLoS Pathog. 2014, 10, e1003979. [Google Scholar] [CrossRef] [PubMed]
- Rasmussen, T.B.; Skindersoe, M.E.; Bjarnsholt, T.; Phipps, R.K.; Christensen, K.B.; Jensen, P.O.; Andersen, J.B.; Koch, B.; Larsen, T.O.; Hentzer, M.; et al. Identity and effects of quorum-sensing inhibitors produced by Penicillium. species. Microbiology 2005, 151, 1325–1340. [Google Scholar] [CrossRef] [PubMed]
- Sepehr, S.; Rahmani-Badi, A.; Babaie-Naiej, H.; Soudi, M.R. Unsaturated fatty acid, cis-2-decenoic acid, in combination with disinfectants or antibiotics removes pre-established biofilms formed by food-related bacteria. PLoS ONE 2014, 9, e101677. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Lim, A.; Lee, J.; Chen, S.; An, S.; Dong, Y.-H.; Zhang, L.-H. Diffusible signal factor (DSF) quorum sensing signal and structurally related molecules enhance the antimicrobial efficacy of antibiotics against some bacterial pathogens. BMC Microbiol. 2014, 14. [Google Scholar] [CrossRef] [PubMed]
- Rahmani-Badi, A.; Sepehr, S.; Mohammadi, P.; Soudi, M.R.; Babaie-Naiej, H.; Fallahi, H. A combination of cis-2-decenoic acid and antibiotics eradicates pre-established catheter-associated biofilms. J. Med. Microbiol. 2014, 63, 1509–1516. [Google Scholar]
- García-Contreras, R.; Maeda, T.; Wood, T.K. Can resistance against quorum-sensing interference be selected? ISME J. 2015, 10, 4–10. [Google Scholar] [CrossRef] [PubMed]
- Johns, B.E.; Purdy, K.J.; Tucker, N.P.; Maddocks, S.E. Phenotypic and genotypic characteristics of small colony variants and their role in chronic infection. Microbiol. Insights 2015, 8, 15–23. [Google Scholar] [PubMed]
- García-Contreras, R.; Peréz-Eretza, B.; Jasso-Chávez, R.; Lira-Silva, E.; Roldán-Sánchez, J.A.; González-Valdez, A.; Soberón-Chávez, G.; Coria-Jiménez, R.; Martínez-Vázquez, M.; Alcaraz, L.D.; et al. High variability in quorum quenching and growth inhibition by furanone C-30 in Pseudomonas aeruginosa clinical isolates from cystic fibrosis patients. Pathog. Dis. 2015, 73, ftv040. [Google Scholar] [CrossRef] [PubMed]
- Grandclément, C.; Tannières, M.; Moréra, S.; Dessaux, Y.; Faure, D.D. Quorum quenching: Role in nature and applied developments. FEMS Microbiol. Rev. 2015. [Google Scholar] [CrossRef] [PubMed]
- Gopal, R.; Lee, J.H.; Kim, Y.G.; Kim, M.S.; Seo, C.H.; Park, Y. Anti-microbial, anti-biofilm activities and cell selectivity of the NRC-16 peptide derived from witch flounder, Glyptocephalus cynoglossus. Mar. Drugs 2013, 11, 1836–1852. [Google Scholar] [CrossRef] [PubMed]
- Jiang, F.; Deng, Y.; Yeh, C.K.; Sun, Y. Quaternized chitosans bind onto preexisting biofilms and eradicate pre-attached microorganisms. J. Mater. Chem. B Mater. Biol. Med. 2014, 2, 8518–8527. [Google Scholar] [CrossRef] [PubMed]
- Zupancic, S.; Kocbek, P.; Baumgartner, S.; Kristl, J. Contribution of nanotechnology to improved treatment of periodontal disease. Curr. Pharm. Des. 2015, 21, 3257–3271. [Google Scholar] [CrossRef] [PubMed]
- Rawson, M.; Haggard, W.; Jennings, J.A. Osteocompatibility of biofilm inhibitors. Open Orthop. J. 2014, 8, 442–449. [Google Scholar] [PubMed]
© 2016 by the author; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abraham, W.-R. Going beyond the Control of Quorum-Sensing to Combat Biofilm Infections. Antibiotics 2016, 5, 3. https://doi.org/10.3390/antibiotics5010003
Abraham W-R. Going beyond the Control of Quorum-Sensing to Combat Biofilm Infections. Antibiotics. 2016; 5(1):3. https://doi.org/10.3390/antibiotics5010003
Chicago/Turabian StyleAbraham, Wolf-Rainer. 2016. "Going beyond the Control of Quorum-Sensing to Combat Biofilm Infections" Antibiotics 5, no. 1: 3. https://doi.org/10.3390/antibiotics5010003
APA StyleAbraham, W.-R. (2016). Going beyond the Control of Quorum-Sensing to Combat Biofilm Infections. Antibiotics, 5(1), 3. https://doi.org/10.3390/antibiotics5010003





