Antimicrobial Resistance: How Can We Overcome the Problem?
Abstract
1. Introduction
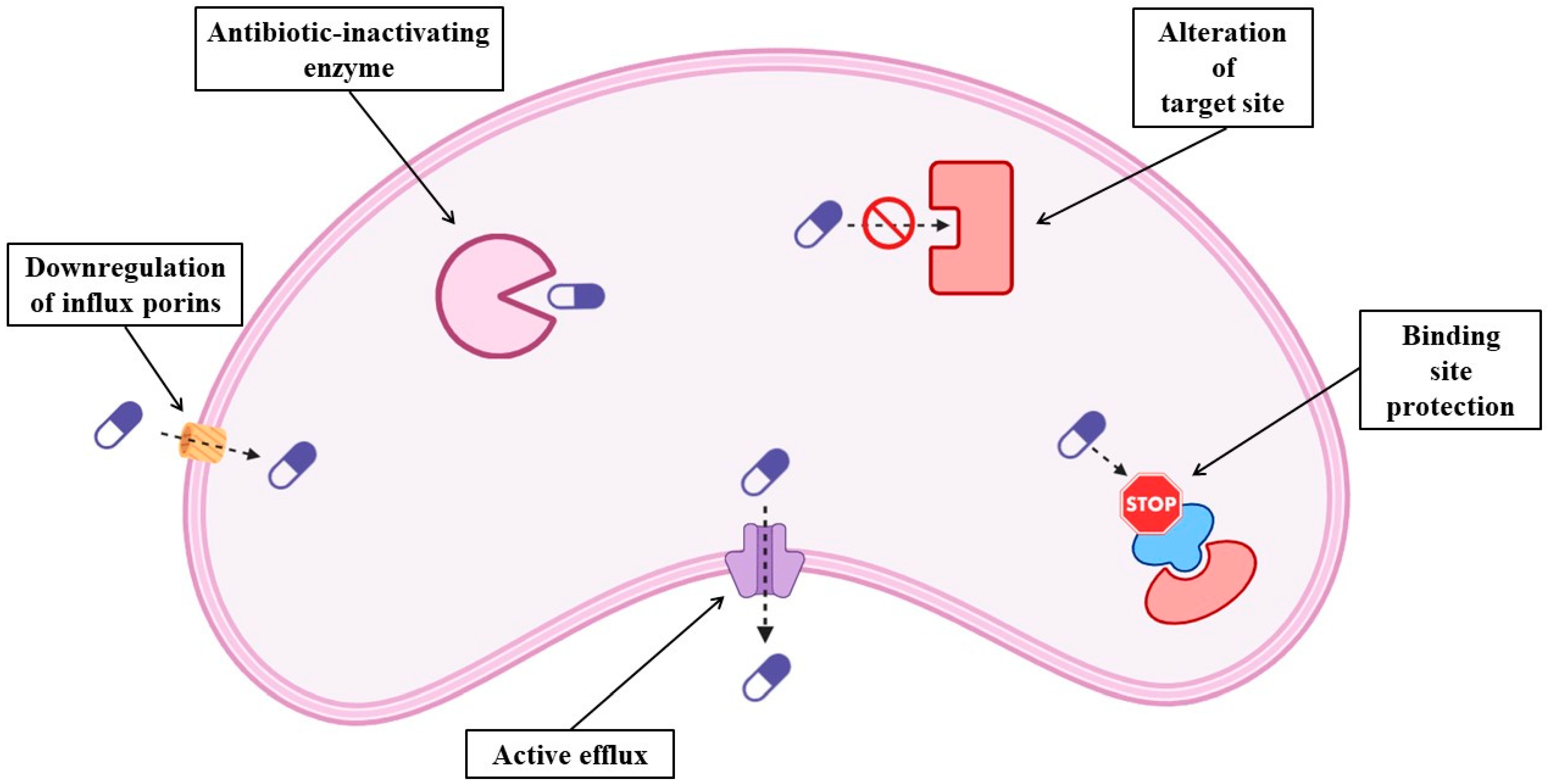
2. What Is Antimicrobial Resistance and How Is It Acquired?
3. Environmental Resistome and Its Monitoring
4. Strategies to Prevent Anti-Microbial Resistance
5. Alternative Treatments to Fight Anti-Microbial Resistance
5.1. Phage Therapy
5.2. Metal Nanoparticles in Biomedical Science
5.3. Antimicrobial Peptides
5.4. Artificial Intelligence-Assisted Generation of New Antimicrobial Molecules
6. Conclusions
- Epidemiological study of the environmental resistome. This approach is needed to establish policies to reduce the misuse of antibiotics or to eliminate antibiotics from wastewaters. Molecular biology tools, e.g., sequencing, have been used to survey viral emergence in wastewater [91,92]. We suggest a similar approach to surveying antibiotic resistance in wastewater.
- Preventive actions by developing protection strategies to combat antibiotic resistance (stringent hygiene measures in hospitals, enforcing restrictions for narrow-spectrum antibiotics, etc.)
- Curative actions by developing alternatives to current antibiotics. In Table 1, we summarize the advantages and inconveniences of these alternative strategies to fight antimicrobial resistance.
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AMR | Antimicrobial resistance |
| GLASS | Global Antimicrobial Resistance and Use Surveillance System |
| HGT | Horizontal gene transfer |
| MGE | Mobile genetic element |
| ARG | Antibiotic resistance gene |
| WWTP | Wastewater treatment plant |
| LMIC | Low-to-middle-income country |
| PPE | Personal protective equipment |
| NP | Nanoparticle |
| AMP | Antimicrobial peptide |
| AI | Artificial intelligence |
| ROS | Reactive oxygen species |
| Ag+ | Silver ions |
| TiO2 | Titanium dioxide |
| LPS | Lipopolysaccharide |
| DMCT | Demeclocycline hydrochloride |
| MDR | Multidrug-resistant |
| FDA | Food and drug administration |
| MOA | Mechanism of action |
| CADD | Computer aided drug design |
| ML | Machine learning |
References
- WHO. Global Action Plan on Antimicrobial Resistance; World Health Organization: Geneva, Switzerland, 2015; ISBN 9789241509763. [Google Scholar]
- Lin, J.; Nishino, K.; Roberts, M.C.; Tolmasky, M.; Aminov, R.I.; Zhang, L. Mechanisms of Antibiotic Resistance. Front. Microbiol. 2015, 6, 2013–2015. [Google Scholar] [CrossRef] [PubMed]
- Naghavi, M.; Vollset, S.E.; Ikuta, K.S.; Swetschinski, L.R.; Gray, A.P.; Wool, E.E.; Robles Aguilar, G.; Mestrovic, T.; Smith, G.; Han, C.; et al. Global Burden of Bacterial Antimicrobial Resistance 1990–2021: A Systematic Analysis with Forecasts to 2050. Lancet 2024, 404, 1199–1226. [Google Scholar] [CrossRef] [PubMed]
- WHO. Global Antimicrobial Resistance and Use Surveillance System (GLASS) Report 2022; World Health Organization: Geneva, Switzerland, 2022; ISBN 9789240062702. [Google Scholar]
- Aslam, B.; Wang, W.; Arshad, M.I.; Khurshid, M.; Muzammil, S.; Rasool, M.H.; Nisar, M.A.; Alvi, R.F.; Aslam, M.A.; Qamar, M.U.; et al. Antibiotic Resistance: A Rundown of a Global Crisis. Infect. Drug Resist. 2018, 11, 1645–1658. [Google Scholar] [CrossRef]
- Perry, J.A.; Westman, E.L.; Wright, G.D. The Antibiotic Resistome: What’s New? Curr. Opin. Microbiol. 2014, 21, 45–50. [Google Scholar] [CrossRef]
- Sabtu, N.; Enoch, D.A.; Brown, N.M. Antibiotic Resistance: What, Why, Where, When and How? Br. Med. Bull. 2015, 116, 105–113. [Google Scholar] [CrossRef] [PubMed]
- Roca, I.; Akova, M.; Baquero, F.; Carlet, J.; Cavaleri, M.; Coenen, S.; Cohen, J.; Findlay, D.; Gyssens, I.; Heure, O.E.; et al. The Global Threat of Antimicrobial Resistance: Science for Intervention. New Microbes New Infect. 2015, 6, 22–29. [Google Scholar] [CrossRef]
- Munita, J.M.; Arias, C.A. Mechanisms of Antibiotic Resistance. In Virulence Mechanisms of Bacterial Pathogens; Wiley: Hoboken, NJ, USA, 2016; pp. 481–511. [Google Scholar] [CrossRef]
- Ventola, C.L. The Antibiotic Resistance Crisis: Part 1: Causes and Threats. Pharm. Ther. 2015, 40, 277–283. [Google Scholar]
- Darby, E.M.; Trampari, E.; Siasat, P.; Gaya, M.S.; Alav, I.; Webber, M.A.; Blair, J.M.A. Molecular Mechanisms of Antibiotic Resistance Revisited. Nat. Rev. Microbiol. 2023, 21, 280–295. [Google Scholar] [CrossRef]
- Von Wintersdorff, C.J.H.; Penders, J.; Van Niekerk, J.M.; Mills, N.D.; Majumder, S.; Van Alphen, L.B.; Savelkoul, P.H.M.; Wolffs, P.F.G. Dissemination of Antimicrobial Resistance in Microbial Ecosystems through Horizontal Gene Transfer. Front. Microbiol. 2016, 7, 173. [Google Scholar] [CrossRef]
- Wachino, J. Horizontal Gene Transfer Systems for Spread of Antibiotic Resistance in Gram-Negative Bacteria. Microbiol. Immunol. 2025, 69, 367–376. [Google Scholar] [CrossRef]
- Rodríguez-Beltrán, J.; DelaFuente, J.; León-Sampedro, R.; MacLean, R.C.; San Millán, Á. Beyond Horizontal Gene Transfer: The Role of Plasmids in Bacterial Evolution. Nat. Rev. Microbiol. 2021, 19, 347–359. [Google Scholar] [CrossRef]
- Holmes, A.H.; Moore, L.S.P.; Sundsfjord, A.; Steinbakk, M.; Regmi, S.; Karkey, A.; Guerin, P.J.; Piddock, L.J.V. Understanding the Mechanisms and Drivers of Antimicrobial Resistance. Lancet 2016, 387, 176–187. [Google Scholar] [CrossRef]
- WHO World Health Organization. Promoting Antimicrobial Stewardship to Tackle Antimicrobial Resistance. Available online: https://www.who.int/europe/activities/promoting-antimicrobial-stewardship-to-tackle-antimicrobial-resistance (accessed on 8 January 2026).
- Chokshi, A.; Sifri, Z.; Cennimo, D.; Horng, H. Global Contributors to Antibiotic Resistance. J. Glob. Infect. Dis. 2019, 11, 36–42. [Google Scholar] [CrossRef]
- Denning, D.W.; Perlin, D.S.; Muldoon, E.G.; Colombo, A.L.; Chakrabarti, A.; Richardson, M.D.; Sorrell, T.C. Delivering on Antimicrobial Resistance Agenda Not Possible without Improving Fungal Diagnostic Capabilities. Emerg. Infect. Dis. 2017, 23, 177–183. [Google Scholar] [CrossRef] [PubMed]
- Marc, C.; Vrignaud, B.; Levieux, K.; Robine, A.; Guen, C.G.L.; Launay, E. Inappropriate Prescription of Antibiotics in Pediatric Practice: Analysis of the Prescriptions in Primary Care. J. Child Health Care 2016, 20, 530–536. [Google Scholar] [CrossRef]
- Almagor, J.; Temkin, E.; Benenson, I.; Fallach, N.; Carmeli, Y. The Impact of Antibiotic Use on Transmission of Resistant Bacteria in Hospitals: Insights from an Agent-Based Model. PLoS ONE 2018, 13, e0197111. [Google Scholar] [CrossRef] [PubMed]
- Jebri, S.; Rahmani, F.; Hmaied, F. Bacteriophages as Antibiotic Resistance Genes Carriers in Agro-food Systems. J. Appl. Microbiol. 2021, 130, 688–698. [Google Scholar] [CrossRef]
- Bhattacharjee, A.S.; Phan, D.; Zheng, C.; Ashworth, D.; Schmidt, M.; Men, Y.; Ferreira, J.F.S.; Muir, G.; Hasan, N.A.; Ibekwe, A.M. Dissemination of Antibiotic Resistance Genes through Soil-Plant-Earthworm Continuum in the Food Production Environment. Environ. Int. 2024, 183, 108374. [Google Scholar] [CrossRef]
- Nnorom, M.-A.; Saroj, D.; Avery, L.; Hough, R.; Guo, B. A Review of the Impact of Conductive Materials on Antibiotic Resistance Genes during the Anaerobic Digestion of Sewage Sludge and Animal Manure. J. Hazard. Mater. 2023, 446, 130628. [Google Scholar] [CrossRef]
- Zhang, J.; Yu, D.; Dian, L.; Hai, Y.; Xin, Y.; Wei, Y. Metagenomics Insights into the Profiles of Antibiotic Resistome in Combined Sewage Overflows from Reads to Metagenome Assembly Genomes. J. Hazard. Mater. 2022, 429, 128277. [Google Scholar] [CrossRef]
- Liu, B.; Yu, K.; Ahmed, I.; Gin, K.; Xi, B.; Wei, Z.; He, Y.; Zhang, B. Key Factors Driving the Fate of Antibiotic Resistance Genes and Controlling Strategies during Aerobic Composting of Animal Manure: A Review. Sci. Total Environ. 2021, 791, 148372. [Google Scholar] [CrossRef] [PubMed]
- Singer, A.C.; Xu, Q.; Keller, V.D.J. Translating Antibiotic Prescribing into Antibiotic Resistance in the Environment: A Hazard Characterisation Case Study. PLoS ONE 2019, 14, e0221568. [Google Scholar] [CrossRef]
- Bonetta, S.; Di Cesare, A.; Pignata, C.; Sabatino, R.; Macrì, M.; Corno, G.; Panizzolo, M.; Bonetta, S.; Carraro, E. Occurrence of Antibiotic-Resistant Bacteria and Resistance Genes in the Urban Water Cycle. Environ. Sci. Pollut. Res. 2023, 30, 35294–35306. [Google Scholar] [CrossRef]
- Larsson, D.G.J.; Flach, C.F. Antibiotic Resistance in the Environment. Nat. Rev. Microbiol. 2022, 20, 257–269. [Google Scholar] [CrossRef] [PubMed]
- Okoye, C.O.; Okeke, E.S.; Okoye, K.C.; Echude, D.; Andong, F.A.; Chukwudozie, K.I.; Okoye, H.U.; Ezeonyejiaku, C.D. Occurrence and Fate of Pharmaceuticals, Personal Care Products (PPCPs) and Pesticides in African Water Systems: A Need for Timely Intervention. Heliyon 2022, 8, e09143. [Google Scholar] [CrossRef]
- Finley, R.L.; Collignon, P.; Larsson, D.G.J.; Mcewen, S.A.; Li, X.Z.; Gaze, W.H.; Reid-Smith, R.; Timinouni, M.; Graham, D.W.; Topp, E. The Scourge of Antibiotic Resistance: The Important Role of the Environment. Clin. Infect. Dis. 2013, 57, 704–710. [Google Scholar] [CrossRef] [PubMed]
- Ding, D.; Wang, B.; Zhang, X.; Zhang, J.; Zhang, H.; Liu, X.; Gao, Z.; Yu, Z. The Spread of Antibiotic Resistance to Humans and Potential Protection Strategies. Ecotoxicol. Environ. Saf. 2023, 254, 114734. [Google Scholar] [CrossRef]
- Zaghen, F.; Sora, V.M.; Meroni, G.; Laterza, G.; Martino, P.A.; Soggiu, A.; Bonizzi, L.; Zecconi, A. Epidemiology of Antimicrobial Resistance Genes in Staphyloccocus aureus Isolates from a Public Database in a One Health Perspective—Sample Characteristics and Isolates’ Sources. Antibiotics 2023, 12, 1225. [Google Scholar] [CrossRef]
- Sora, V.M.; Zaghen, F.; Zecconi, A. How to Improve Surveillance Program for Shiga Toxin-Producing E. Coli (STEC): Gap Analysis and Pilot Study. Pathogens 2024, 13, 511. [Google Scholar] [CrossRef]
- Li, B.; Yan, T. Next Generation Sequencing Reveals Limitation of QPCR Methods in Quantifying Emerging Antibiotic Resistance Genes (ARGs) in the Environment. Appl. Microbiol. Biotechnol. 2021, 105, 2925–2936. [Google Scholar] [CrossRef]
- Meroni, G.; Sora, V.M.; Martino, P.A.; Sbernini, A.; Laterza, G.; Zaghen, F.; Soggiu, A.; Zecconi, A. Epidemiology of Antimicrobial Resistance Genes in Streptococcus Agalactiae Sequences from a Public Database in a One Health Perspective. Antibiotics 2022, 11, 1236. [Google Scholar] [CrossRef]
- Gajdács, M.; Albericio, F. Antibiotic Resistance: From the Bench to Patients. Antibiotics 2019, 8, 129. [Google Scholar] [CrossRef]
- European Food Safety Authority (EFSA); European Centre for Disease Prevention and Control (ECDC). The European Union One Health 2022 Zoonoses Report. EFSA J. 2023, 21, e8442. [Google Scholar] [CrossRef]
- Nepal, G.; Bhatta, S. Self-Medication with Antibiotics in WHO Southeast Asian Region: A Systematic Review. Cureus 2018, 10, e2428. [Google Scholar] [CrossRef]
- Wittebole, X.; De Roock, S.; Opal, S.M. A Historical Overview of Bacteriophage Therapy as an Alternative to Antibiotics for the Treatment of Bacterial Pathogens. Virulence 2014, 5, 226–235. [Google Scholar] [CrossRef]
- Wright, A.; Hawkins, C.H.; Änggård, E.E.; Harper, D.R. A Controlled Clinical Trial of a Therapeutic Bacteriophage Preparation in Chronic Otitis Due to Antibiotic-Resistant Pseudomonas Aeruginosa; A Preliminary Report of Efficacy. Clin. Otolaryngol. 2009, 34, 349–357. [Google Scholar] [CrossRef]
- Ghannad, M.S.; Mohammadi, A. Bacteriophage: Time to Re-Evaluate the Potential of Phage Therapy as a Promising Agent to Control Multidrug-Resistant Bacteria. Iran. J. Basic Med. Sci. 2012, 15, 693–701. [Google Scholar]
- Westra, E.R.; Buckling, A.; Fineran, P.C. CRISPR–Cas Systems: Beyond Adaptive Immunity. Nat. Rev. Microbiol. 2014, 12, 317–326. [Google Scholar] [CrossRef] [PubMed]
- Oechslin, F. Resistance Development to Bacteriophages Occurring during Bacteriophage Therapy. Viruses 2018, 10, 351. [Google Scholar] [CrossRef]
- Lenneman, B.R.; Fernbach, J.; Loessner, M.J.; Lu, T.K.; Kilcher, S. Enhancing Phage Therapy through Synthetic Biology and Genome Engineering. Curr. Opin. Biotechnol. 2021, 68, 151–159. [Google Scholar] [CrossRef]
- Doud, M.B.; Robertson, J.M.; Strathdee, S.A. Optimizing Phage Therapy with Artificial Intelligence: A Perspective. Front. Cell Infect. Microbiol. 2025, 15, 1611857. [Google Scholar] [CrossRef]
- Boeckaerts, D.; Stock, M.; Ferriol-González, C.; Oteo-Iglesias, J.; Sanjuán, R.; Domingo-Calap, P.; De Baets, B.; Briers, Y. Prediction of Klebsiella Phage-Host Specificity at the Strain Level. Nat. Commun. 2024, 15, 4355. [Google Scholar] [CrossRef] [PubMed]
- Gaborieau, B.; Vaysset, H.; Tesson, F.; Charachon, I.; Dib, N.; Bernier, J.; Dequidt, T.; Georjon, H.; Clermont, O.; Hersen, P.; et al. Prediction of Strain Level Phage-Host Interactions across the Escherichia Genus Using Only Genomic Information. Nat. Microbiol. 2024, 9, 2847–2861. [Google Scholar] [CrossRef]
- Nelson, D.; Loomis, L.; Fischetti, V.A. Prevention and Elimination of Upper Respiratory Colonization of Mice by Group A Streptococci by Using a Bacteriophage Lytic Enzyme. Proc. Natl. Acad. Sci. USA 2001, 98, 4107–4112. [Google Scholar] [CrossRef] [PubMed]
- Vázquez, R.; García, E.; García, P. Phage Lysins for Fighting Bacterial Respiratory Infections: A New Generation of Antimicrobials. Front. Immunol. 2018, 9, 2252. [Google Scholar] [CrossRef]
- Nelson, D.; Schuch, R.; Chahales, P.; Zhu, S.; Fischetti, V.A. PlyC: A Multimeric Bacteriophage Lysin. Proc. Natl. Acad. Sci. USA 2006, 103, 10765–10770. [Google Scholar] [CrossRef]
- Briers, Y.; Walmagh, M.; Van Puyenbroeck, V.; Cornelissen, A.; Cenens, W.; Aertsen, A.; Oliveira, H. Engineered Endolysin-Based “Artilysins” to Combat Multidrug-Resistant Gram-Negative Pathogens. mBio 2014, 5, 1128. [Google Scholar] [CrossRef] [PubMed]
- Young, R.; Phage, J.J.G. Therapy Redux—What Is to Be Done?: Bacteriophage Biology Should Move beyond a Model System to Support Human Health. Science 2015, 176, 139–148. [Google Scholar] [CrossRef]
- Khan, I.; Saeed, K.; Khan, I. Nanoparticles: Properties, Applications and Toxicities. Arab. J. Chem. 2019, 12, 908–931. [Google Scholar] [CrossRef]
- Shin, W.K.; Cho, J.; Kannan, A.G.; Lee, Y.S.; Kim, D.W. Cross-Linked Composite Gel Polymer Electrolyte Using Mesoporous Methacrylate-Functionalized SiO2 Nanoparticles for Lithium-Ion Polymer Batteries. Sci. Rep. 2016, 6, 26332. [Google Scholar] [CrossRef]
- García-Rodríguez, A.; Vila, L.; Cortés, C.; Hernández, A.; Marcos, R. Effects of Differently Shaped TiO2NPs (Nanospheres, Nanorods and Nanowires) on the in Vitro Model (Caco-2/HT29) of the Intestinal Barrier. Part. Fibre Toxicol. 2018, 15, 33. [Google Scholar] [CrossRef]
- Cheon, J.Y.; Kim, S.J.; Rhee, Y.H.; Kwon, O.H.; Park, W.H. Shape-Dependent Antimicrobial Activities of Silver Nanoparticles. Int. J. Nanomed. 2019, 14, 2773–2780. [Google Scholar] [CrossRef]
- Kumar, M.; Sarma, D.K.; Shubham, S.; Kumawat, M.; Verma, V.; Nina, P.B.; Parasannanavar, D.J.; Kumar, S.; Singh, B.; Tiwari, R.R. Futuristic Non-Antibiotic Therapies to Combat Antibiotic Resistance: A Review. Front. Microbiol. 2021, 12, 609459. [Google Scholar] [CrossRef]
- Yuan, Q.; He, Y.; Mao, B.; Zuo, P.; Wu, W.; Huang, Y.; Javed, H.; Hu, N. Nano-Metal Oxides Naturally Attenuate Antibiotic Resistance in Wastewater: Killing Antibiotic Resistant Bacteria by Dissolution and Decreasing Antibiotic Tolerance by Attachment. NanoImpact 2020, 18, 100225. [Google Scholar] [CrossRef]
- Ezeuko, A.S.; Ojemaye, M.O.; Okoh, O.O.; Okoh, A.I. Potentials of Metallic Nanoparticles for the Removal of Antibiotic Resistant Bacteria and Antibiotic Resistance Genes from Wastewater: A Critical Review. J. Water Process Eng. 2021, 41, 102041. [Google Scholar] [CrossRef]
- El-Sesy, M.E.; Othman, S.A. Promising Antibacterial Activities of Anethole and Green-Synthesized Magnetite Nanoparticles against Multiple Antibiotic-Resistant Bacteria. Water Sci. Technol. 2023, 87, 729–747. [Google Scholar] [CrossRef]
- Bruna, T.; Maldonado-Bravo, F.; Jara, P.; Caro, N. Silver Nanoparticles and Their Antibacterial Applications. Int. J. Mol. Sci. 2021, 22, 7202. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Wang, M.; Cui, Y.; Zhang, C. Can We Arrest the Evolution of Antibiotic Resistance? The Differences between the Effects of Silver Nanoparticles and Silver Ions. Environ. Sci. Technol. 2022, 56, 5090–5101. [Google Scholar] [CrossRef] [PubMed]
- Maklakova, M.; Villarreal-Gómez, L.J.; Nefedova, E.; Shkil, N.; Luna Vázquez-Gómez, R.; Pestryakov, A.; Bogdanchikova, N. Potential Antibiotic Resurgence: Consecutive Silver Nanoparticle Applications Gradually Increase Bacterial Susceptibility to Antibiotics. ACS Omega 2025, 10, 4624–4635. [Google Scholar] [CrossRef] [PubMed]
- Roy, A.S.; Parveen, A.; Koppalkar, A.R.; Prasad, M.V.N.A. Effect of Nano-Titanium Dioxide with Different Antibiotics against Methicillin-Resistant Staphylococcus Aureus. J. Biomater. Nanobiotechnol. 2010, 1, 37–41. [Google Scholar] [CrossRef]
- Metz-Boutigue, M.H.; Goumon, Y.; Strub, J.M.; Lugardon, K.; Aunis, D. Antimicrobial Chromogranins and Proenkephalin-A—Derived Peptides. Ann. N. Y. Acad. Sci. 2003, 992, 168–178. [Google Scholar] [CrossRef]
- Scavello, F.; Kharouf, N.; Lavalle, P.; Haikel, Y.; Schneider, F.; Metz-Boutigue, M.H. The Antimicrobial Peptides Secreted by the Chromaffin Cells of the Adrenal Medulla Link the Neuroendocrine and Immune Systems: From Basic to Clinical Studies. Front. Immunol. 2022, 13, 977175. [Google Scholar] [CrossRef]
- Zhang, J.; Li, W.; Chen, J.; Qi, W.; Wang, F.; Zhou, Y. Impact of Biofilm Formation and Detachment on the Transmission of Bacterial Antibiotic Resistance in Drinking Water Distribution Systems. Chemosphere 2018, 203, 368–380. [Google Scholar] [CrossRef]
- Lei, J.; Sun, L.C.; Huang, S.; Zhu, C.; Li, P.; He, J.; Mackey, V.; Coy, D.H.; He, Q.Y. The Antimicrobial Peptides and Their Potential Clinical Applications. Am. J. Transl. Res. 2019, 11, 3919–3931. [Google Scholar] [PubMed]
- Kang, H.K.; Kim, C.; Seo, C.H.; Park, Y. The Therapeutic Applications of Antimicrobial Peptides (AMPs): A Patent Review. J. Microbiol. 2017, 55, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Mwangi, J.; Hao, X.; Lai, R.; Zhang, Z.Y. Antimicrobial Peptides: New Hope in the War against Multidrug Resistance. Zool. Res. 2019, 40, 488–505. [Google Scholar] [CrossRef]
- Santos-Júnior, C.D.; Torres, M.D.T.; Duan, Y.; Rodríguez Del Río, Á.; Schmidt, T.S.B.; Chong, H.; Fullam, A.; Kuhn, M.; Zhu, C.; Houseman, A.; et al. Discovery of Antimicrobial Peptides in the Global Microbiome with Machine Learning. Cell 2024, 187, 3761–3778.e16. [Google Scholar] [CrossRef] [PubMed]
- Bucataru, C.; Ciobanasu, C. Antimicrobial Peptides: Opportunities and Challenges in Overcoming Resistance. Microbiol. Res. 2024, 286, 127822. [Google Scholar] [CrossRef]
- Galzitskaya, O.V. Creation of New Antimicrobial Peptides. Int. J. Mol. Sci. 2023, 24, 9451. [Google Scholar] [CrossRef]
- Mokoena, M.P. Lactic Acid Bacteria and Their Bacteriocins: Classification, Biosynthesis and Applications against Uropathogens: A Mini-Review. Molecules 2017, 22, 1255. [Google Scholar] [CrossRef]
- Wang, S.; Fang, Q.; Lu, Z.; Gao, Y.; Trembleau, L.; Ebel, R.; Andersen, J.H.; Philips, C.; Law, S.; Deng, H. Discovery and Biosynthetic Investigation of a New Antibacterial Dehydrated Non-Ribosomal Tripeptide. Angew. Chem. Int. Ed. 2021, 60, 3229–3237. [Google Scholar] [CrossRef]
- Galván Márquez, I.J.; McKay, B.; Wong, A.; Cheetham, J.J.; Bean, C.; Golshani, A.; Smith, M.L. Mode of Action of Nisin on Escherichia coli. Can. J. Microbiol. 2020, 66, 161–168. [Google Scholar] [CrossRef] [PubMed]
- Tong, Z.; Zhang, Y.; Ling, J.; Ma, J.; Huang, L.; Zhang, L. An in Vitro Study on the Effects of Nisin on the Antibacterial Activities of 18 Antibiotics against Enterococcus Faecalis. PLoS ONE 2014, 9, e89209. [Google Scholar] [CrossRef]
- Mathur, H.; Field, D.; Rea, M.C.; Cotter, P.D.; Hill, C.; Ross, R.P. Bacteriocin-Antimicrobial Synergy: A Medical and Food Perspective. Front. Microbiol. 2017, 8, 1205. [Google Scholar] [CrossRef]
- Li, S.; She, P.; Zhou, L.; Zeng, X.; Xu, L.; Liu, Y.; Chen, L.; Wu, Y. High-Throughput Identification of Antibacterials Against Pseudomonas Aeruginosa. Front. Microbiol. 2020, 11, 591426. [Google Scholar] [CrossRef]
- Zhao, X.; Zhen, Z.; Wang, X.; Guo, N. Synergy of a Combination of Nisin and Citric Acid against Staphylococcus Aureus and Listeria Monocytogenes. Food Addit. Contam. Part A 2017, 34, 2058–2068. [Google Scholar] [CrossRef]
- Zhang, Q.Y.; Yan, Z.B.; Meng, Y.M.; Hong, X.Y.; Shao, G.; Ma, J.J.; Cheng, X.R.; Liu, J.; Kang, J.; Fu, C.Y. Antimicrobial Peptides: Mechanism of Action, Activity and Clinical Potential. Mil. Med. Res. 2021, 8, 48. [Google Scholar] [CrossRef] [PubMed]
- Szymczak, P.; Zarzecki, W.; Wang, J.; Duan, Y.; Wang, J.; Coelho, L.P.; de la Fuente-Nunez, C.; Szczurek, E. AI-Driven Antimicrobial Peptide Discovery: Mining and Generation. Acc. Chem. Res. 2025, 58, 1831–1846. [Google Scholar] [CrossRef]
- DiMasi, J.A.; Grabowski, H.G.; Hansen, R.W. Innovation in the Pharmaceutical Industry: New Estimates of R&D Costs. J. Health Econ. 2016, 47, 20–33. [Google Scholar] [CrossRef] [PubMed]
- Lepore, C.; Silver, L.; Theuretzbacher, U.; Thomas, J.; Visi, D. The Small-Molecule Antibiotics Pipeline: 2014–2018. Nat. Rev. Drug Discov. 2019, 18, 739. [Google Scholar] [CrossRef]
- Torres, M.D.T.; de la Fuente-Nunez, C. Toward Computer-Made Artificial Antibiotics. Curr. Opin. Microbiol. 2019, 51, 30–38. [Google Scholar] [CrossRef] [PubMed]
- Schneider, G. Automating Drug Discovery. Nat. Rev. Drug Discov. 2018, 17, 97–113. [Google Scholar] [CrossRef]
- Melo, M.C.R.; Maasch, J.R.M.A.; de la Fuente-Nunez, C. Accelerating Antibiotic Discovery through Artificial Intelligence. Commun. Biol. 2021, 4, 1050. [Google Scholar] [CrossRef]
- Talat, A.; Khan, A.U. Artificial Intelligence as a Smart Approach to Develop Antimicrobial Drug Molecules: A Paradigm to Combat Drug-Resistant Infections. Drug Discov. Today 2023, 28, 103491. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, M.; Hinkley, T.; Tsuda, S.; Abul-Haija, Y.M.; McBurney, R.T.; Kulikov, V.; Mathieson, J.S.; Galiñanes Reyes, S.; Castro, M.D.; Cronin, L. Using Evolutionary Algorithms and Machine Learning to Explore Sequence Space for the Discovery of Antimicrobial Peptides. Chem 2018, 4, 533–543. [Google Scholar] [CrossRef]
- Stokes, J.M.; Yang, K.; Swanson, K.; Jin, W.; Cubillos-Ruiz, A.; Donghia, N.M.; MacNair, C.R.; French, S.; Carfrae, L.A.; Bloom-Ackerman, Z.; et al. A Deep Learning Approach to Antibiotic Discovery. Cell 2020, 180, 688–702.e13. [Google Scholar] [CrossRef] [PubMed]
- El Soufi, G.; Di Jorio, L.; Gerber, Z.; Cluzel, N.; Van Assche, J.; Delafoy, D.; Olaso, R.; Daviaud, C.; Loustau, T.; Schwartz, C.; et al. Highly Efficient and Sensitive Membrane-Based Concentration Process Allows Quantification, Surveillance, and Sequencing of Viruses in Large Volumes of Wastewater. Water Res. 2024, 249, 120959. [Google Scholar] [CrossRef]
- Bertrand, I.; Challant, J.; Jeulin, H.; Hartard, C.; Mathieu, L.; Lopez, S.; Schvoerer, E.; Courtois, S.; Gantzer, C. Epidemiological Surveillance of SARS-CoV-2 by Genome Quantification in Wastewater Applied to a City in the Northeast of France: Comparison of Ultrafiltration- and Protein Precipitation-Based Methods. Int. J. Hyg. Environ. Health 2021, 233, 113692. [Google Scholar] [CrossRef]
| Alternative Strategy | Advantages | Disadvantages | References |
|---|---|---|---|
| Phage therapy | High specificity. Topical and intravenous treatment. | Narrow spectrum of action. Prohibited use for empirical treatment. | [36,37,38,39,40,41,42,43,44,45,46,47,48,49] |
| Nanoparticles | Broad-spectrum. Can be combined with antibiotics to enhance efficacy. Low probability of resistance build-up. | Long-term toxicity unknown. High costs of production. | [50,51,52,53,54,55,56,57,58,59,60,61] |
| Antimicrobial peptides | Broad-spectrum. Synthetic enhancement of stability and efficacy. Low probability of resistance build-up. | Susceptible to in vivo degradation. High costs of production. High concentrations may exert toxicity. | [62,63,64,65,66,67,68,69,70,71,72,73,74,75,76,77,78,79] |
| Artificial intelligence | Quick drug discovery. Resistance patterns prediction. Efficacy enhancement of already existing treatments. | Requires high computing power. Highly dependent on dataset quality and completeness. Large amount of data required to train models. | [80,81,82,83,84,85,86,87] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Sora, V.M.; Wallet, C.; Meroni, G.; Loustau, T.; Rohr, O.; Zecconi, A.; Schwartz, C. Antimicrobial Resistance: How Can We Overcome the Problem? Antibiotics 2026, 15, 82. https://doi.org/10.3390/antibiotics15010082
Sora VM, Wallet C, Meroni G, Loustau T, Rohr O, Zecconi A, Schwartz C. Antimicrobial Resistance: How Can We Overcome the Problem? Antibiotics. 2026; 15(1):82. https://doi.org/10.3390/antibiotics15010082
Chicago/Turabian StyleSora, Valerio Massimo, Clementine Wallet, Gabriele Meroni, Thomas Loustau, Olivier Rohr, Alfonso Zecconi, and Christian Schwartz. 2026. "Antimicrobial Resistance: How Can We Overcome the Problem?" Antibiotics 15, no. 1: 82. https://doi.org/10.3390/antibiotics15010082
APA StyleSora, V. M., Wallet, C., Meroni, G., Loustau, T., Rohr, O., Zecconi, A., & Schwartz, C. (2026). Antimicrobial Resistance: How Can We Overcome the Problem? Antibiotics, 15(1), 82. https://doi.org/10.3390/antibiotics15010082





