Biofilms Exposed: Innovative Imaging and Therapeutic Platforms for Persistent Infections
Abstract
1. Introduction
Objectives of This Review
- Provide an overview of the current understanding of biology, structural organization, and clinical significance of biofilms in relation to different types of infections.
- Examine the shortcomings of traditional antimicrobial methods in terms of successfully eliminating biofilms.
- Describe the latest developments in biofilm-targeted treatment approaches, such as CRISPR-Cas-based interventions, bacteriophage therapy, enzyme-mediated methods, nanotechnology-based systems, and quorum-sensing inhibition.
- Highlight the translational significance of preclinical in vitro, ex vivo, and in vivo models that resemble clinical biofilms.
- Examine intelligent drug-delivery systems that respond to cues unique to biofilms.
- Determine the present obstacles, unmet research needs, and possible avenues for the creation of clinically feasible anti-biofilm treatments.
2. Classical Techniques in Biofilm Research: Historical Foundations and Limitations
2.1. Crystal Violet Assay
2.2. Colony-Forming Unit (CFU) Counts
2.3. Tube Method
2.4. Congo Red Agar Assay
2.5. Drip-Flow and Early Flow-Cell Systems
2.6. Biomass Dry-Weight Measurement
2.7. Light Microscopy
2.8. Scanning Electron Microscopy (SEM)
2.9. Transmission Electron Microscopy (TEM)
3. Cutting-Edge Technologies and Models in Biofilm Research
3.1. Advanced Microscopy and Imaging Techniques in Biofilm Research
3.1.1. Stimulated Emission Depletion (STED) Microscopy
3.1.2. Lattice Light-Sheet Microscopy (LLSM)
3.1.3. Stochastic Optical Reconstruction Microscopy (STORM)
3.1.4. Confocal Laser Scanning Microscopy (CLSM)
3.1.5. Atomic Force Microscopy (AFM)
3.1.6. Cryo-Electron Microscopy (Cryo-EM)
3.1.7. Optical Coherence Tomography (OCT)
3.1.8. Raman Spectroscopy Imaging (RSI)
3.1.9. Bioluminescence Imaging (BLI)
3.1.10. Digital Holographic Microscopy (DHM)
| Technique | Resolution nm | Live Imaging | Label-Free | Key Features | Typical Applications | Reference |
|---|---|---|---|---|---|---|
| Stochastic Optical Reconstruction (STED) Microscopy | ~30–50 | No | No | Super-resolution; EPS structure; nanoscale matrix mapping | Antibiotic penetration, EPS-targeted therapies | [87] |
| Lattice Light-Sheet Microscopy | ~300 lateral, ~500 axial | Yes | No | Low phototoxicity; fast volumetric imaging; dynamic events in 3D biofilms | Host–microbe interaction, live biofilm dynamics | [63] |
| Stochastic Optical Reconstruction Microscopy (STORM) | ~20–30 | No | No | Single-molecule localization; super-resolution of matrix, enzymes | Peptidoglycan mapping, maturation tracking | [88] |
| Confocal laser scanning electron microscopy (CLSM) | ~200–300 | Yes | No | 3D reconstructions; multichannel imaging; live/dead discrimination | Biofilm architecture, viability, drug diffusion studies | [89] |
| Atomic Force Microscopy (AFM) | ~1 (surface) | Yes (partial) | Yes | Nanoscale topography; mechanical properties; elasticity and adhesion | Matrix cohesion, drug-induced biomechanical changes | [90] |
| Cryo-Electron Microscopy (Cryo-EM) | ~0.3–1 | No | Yes | Near-atomic resolution; native-state imaging of molecular complexes | Structural biology, biofilm–phage/protein interactions | [91] |
| Optical Coherence Tomography (OCT) | ~1–15 µm | Yes | Yes | Deep penetration; real-time label-free cross-sections | Medical/industrial biofilms, flow-cell monitoring | [92] |
| Raman Spectroscopy Imaging | ~300–500 | Yes (slow) | Yes | Molecular composition; metabolic fingerprinting; antibiotic mapping | Chemical analysis, resistance zones, EPS structure | [93] |
| Bioluminescence Imaging (BLI) | ~1–5 mm (low) | Yes | Yes | Real-time in vivo imaging; metabolic activity monitoring | Infection progression, therapeutic efficacy | [83] |
| Digital Holographic Microscopy (DHM) | ~500 lateral | Yes | Yes | Label-free 3D imaging; real-time biomass/motility tracking | High-throughput screening, early-stage studies | [94] |
3.2. AI-Driven Biofilm Modeling: Transforming Research and Therapeutic Development
3.3. In Vivo Models for Biofilm Research
4. Biofilm Control Approach
4.1. Synthetic Biology and Genetic Engineering Approaches for Biofilm Control
4.2. Nanotechnology Approaches for Biofilm Control
| Nanomaterial | Pathogen Name | Mechanism | Material Base | Active Concentration Applied | MBIC | Reference |
|---|---|---|---|---|---|---|
| Chitosan-based nanogels | S. aureus, K. pneumoniae, P. aeruginosa | Chitosan’s cationic nature allows it to interact with negatively charged bacterial cell membranes, leading to increased permeability and disruption of biofilm formation. Additionally, chitosan can be functionalized with antimicrobial agents to enhance its efficacy. | Chitosan | 25 μg/mL | 200 μg/mL | [164] |
| Hyaluronic acid-based nanogels | Methicillin-resistant S. aureus (MRSA), P. aeruginosa | These nanogels can deliver nitric oxide (NO) and antimicrobial peptides (AMPs) directly into biofilms. NO disrupts biofilm structure, while AMPs kill the bacteria, resulting in synergistic antibiofilm activity. | Hyaluronic acid | 128 mg/mL | 512 mg/mL | [165] |
| Alginate-based nanogels | S. aureus, E. coli | Alginate nanogels can be loaded with agents like tannic acid and iron ions to induce photothermal therapy (PTT) and chemodynamic therapy (CDT), generating reactive oxygen species (ROS) that disrupt biofilms. | Alginate | 0.625 mg/L | 25 μg/mL | [166] |
| Polymeric nanoparticles | S. aureus, P. aeruginosa, E. coli | These nanoparticles can encapsulate antibiotics, enhancing their stability and penetration into biofilms. They can also be designed to release drugs in response to specific stimuli within the biofilm environment. | Caged guanidine groups | Aqueous dispersions of CGNs, GNs, and ICGNs (0.1 mg mL−1, 0.2 mL) | 0.4 mg mL−1, 0.2 mL | [167] |
| Liposomes | S. aureus, P. aeruginosa | Liposomes can fuse with bacterial membranes, facilitating the delivery of encapsulated antibiotics directly into the biofilm matrix, thereby increasing local drug concentration and efficacy. | Phospholipids | No lipid concentrations given | Biofilm inhibition inferred from relative viability assays | [168] |
| Dendrimers | S. aureus, P. aeruginosa | Dendrimers possess a highly branched structure with functional groups that can be tailored for antimicrobial activity. They can disrupt bacterial membranes and inhibit biofilm formation. | Quaternary ammonium-functionalized, metal ion complexed) | No quantitative biofilm inhibition thresholds provided | No quantitative biofilm inhibition thresholds provided | [169] |
| Peptide-based nanoparticles | S. aureus, P. aeruginosa | These nanoparticles utilize antimicrobial peptides that can insert into bacterial membranes, causing disruption and cell death, effectively preventing biofilm development. | Peptide dendrimer (Trp/Arg) | 20 μM, 40 μM, 80 μM | 16–32 mg/L | [170] |
| Gold nanoparticles (Au NPs) | Methicillin- Resistant S. aureus. | Photothermal activity of Au NPs induce the antimicrobial activity against drug-resistant bacteria species. | Gold nanoparticles (AuNPs) | 109.5 μg/mL | 165 μg/mL | |
| Silver nanoparticles (Ag NPs) | P. aeruginosa | Ag NPs generate ROS inside the cell membrane which led to cell wall rupture and cell death. | Silver nanoparticles (various types) | 1–200 µg/mL against P. aeruginosa | Not reported as a distinct value | [171] |
| Silver nanoparticles (Ag NPs) | P. aeruginosa, E. coli, K. pneumonae and S. aureus | Small size and spherical shape of silver nanoparticles responsible for antibacterial activity. | Silver nanoparticles (AgNPs) | 0.3 mg | 1.0 mg | [172] |
| Green synthesized Ag NPs. | Ampicillin-resistant K. pneumoniae, E. coli | Green Ag NPs interrupt the metabolic activities of bacteria by interacting with enzymes. | Iron oxide nanoparticles (Fe3O4 NPs) | 64 μg/mL | 64 μg/mL | [173] |
| Methionine capped-Au NPs | Gram-negative- A. baumannii and S. enterica Gram-positive- Methicillin- Resistant S. aureus and M. luteus. | Photothermal activity of Au NPs induce the antimicrobial activity against drug-resistant bacteria species. | Ultra-small gold nanoparticles (Au0/Au+) | 30 mg L | 70 mg L−1 | [174] |
| Gold nanorods | Methicillin- Resistant S. aureus | The presence of single-valent gold on the surface of nanoparticles induced the antimicrobial activity against both Gram-positive and gram- negative drug-resistant bacteria. | Gold nanorods (AuNRs) | 2× MIC | 8 μg mL−1 | [175] |
| Metallosurfactant-based cobalt oxide/hydroxide nanoparticles. | S. aureus | Presence of metallosurfactants enhanced the antibacterial activity of Co NPs by rupturing the cell wall of bacteria. | Chitosan-based nanoparticles (CSNPs) | 10 μg mL−1 | 10 μg mL−1 | [176] |
| Polyhydroxybutyrate-Co3O4 bio nanocomposites. | E. coli, S. aureus | Biosynthesized Co NPs have high thermal stability and improved structural properties which enhanced the bactericidal properties of Co NPs. | Zinc oxide nanoparticles (ZnO NPs) | 50 μg/mL | 50 μg/mL | [177] |
| Silver nanoparticles (AgNPs) synthesized using chemical reduction method | A. baumannii | Fe3O4 NPs developed antibacterial activity by interacting with the ATP associated mechanisms. | Silver nanoparticles (AgNPs) | 0.5 mg/mL | 0.5 mg/mL | [178] |
| Copper oxide nanoparticles (CuO NPs) synthesized using Azadirachta indica (neem) leaf extract | S. aureus | Magnetic nanoparticles generate artificial magnetic field inside the biofilm and enhance the drug penetration and antimicrobial activity inside bacterial cell. | Copper oxide nanoparticles (CuO NPs) | 32 μg/mL | 32 μg/mL | [179] |
| Aluminum oxide nanoparticles (Al2O3 NPs) | A. baumannii | Al2O3 NPs inhibit the EPS and reduce the formation of biofilm. | Al2O3 NPs | 120 µg mL−1 | 120 µg mL−1 | [180] |
| AgNPs_mPEG, AgNPs_mPEG_AK, and AK | P. aeruginosa | The antimicrobial activity of AgNPs relies on their ability to release Ag1 to the bacteria, as determined by the corrosion rate, which depends on the morphology (size and shape) of the NPs. Accordingly, it has also been reported that smaller AgNPs have more powerful bactericidal activity as the area of surface contact of AgNPs with microorganisms is increased in relation to larger AgNPs. | Selenium nanoparticles (SeNPs) | 50 μg/mL | 50 μg/mL | [181] |
| Liquid crystal nanoparticles (LCNPs) | P. aeruginosa S. aureus | The nanostructure enables the antibiotic to be released in a controlled and sustained manner, ensuring therapeutic concentrations for extended periods of time. By delivering the antibiotic directly into the biofilm, LCNPs raise the local concentration of the medication at the site of infection, improving its bactericidal activity. | Liquid crystal nanoparticles (LCNPs) | 2 μg/mL | 2 μg/mL | [182] |
| Nitric oxide (NO) releasing nanocomposite Activation: near-infrared (NIR) light irradiation | P. gingivalis F. nucleatum S. mutans | Photodynamic therapy (PDT): When exposed to near-infrared light, ICG produces reactive oxygen species (ROS), which destroy bacteria. Photothermal therapy (PTT): NIR irradiation causes localized heating, breaking biofilm structure and hastening bacterial elimination. Nitric oxide release: Heat from PTT causes the release of NO from SNO, which has antibacterial characteristics and aids in biofilm breakdown. | Bismuth sulfide (Bi2S3) nanoparticles and bis-N-nitroso compounds (BNN) | 128 μg/mL | 128 μg/mL | [183] |
| Silver nanoparticles (AgNPs) | S. aureus P. aeruginosa | Disrupt bacterial membranes. Generate reactive oxygen species (ROS). Interfere with DNA replication Penetrate biofilms due to their small size | silver nitrate (AgNO3) | 0.25 μg/mL to 2.0 μg/mL | 0.25 μg/mL to 2.0 μg/mL | [184] |
| Zinc oxide nanoparticles (ZnO NPs) | S. aureus, E. coli | ROS generation. Disruption of membrane integrity. Photocatalytic activity (especially under UV light) | Zinc oxide nanoparticles | 50 µg/mL | 50 µg/mL | [185] |
| Chitosan nanoparticles | S. aureus, P. aeruginosa | Electrostatic interaction with negatively charged bacterial membranes. Disruption of cell wall and membrane. Interference with nutrient transport and metabolism. | Chitosan–silver nanocomposite (CS–AgNPs) | 64 μg/mL | 64 μg/mL | [186] |
| Polymeric nanoparticles (e.g., PLGA-based) | P. aeruginosa, S. aureus | Sustained drug release. Targeted delivery to biofilm. Potential surface modification for improved interaction with biofilm. | PLGA (polylactic-co-glycolic acid). | 10–12.5 mg/mL | 102.5 mg/mL | [187] |
| Chitosan nanoparticles (CS NPs) | C. albicans | Chitosan and bCTL are positively charged, allowing for strong electrostatic interaction with the negatively charged fungal cell walls and biofilm matrix. Nanoparticles penetrate the extracellular matrix of the biofilm, destabilizing its structure. bCTL disrupts fungal cell membranes, increasing permeability and leading to cell death. | Chitosan and the antifungal crosslinker phytic acid | 140 ± 2.2 µg/mL | 140 ± 2.2 µg/mL | [188] |
| Copper nanoparticles (CuNPs) | E. coli, S. aureus, | Membrane damage. Oxidative stress via ROS. Metal ion toxicity | Green synthesized copper nanoparticles (CuNPs) | 30 µg/mL | MIC—89–91% inhibition | [189] |
| Titanium dioxide nanoparticles (TiO2 NPs) | Broad-spectrum (Gram-positive and Gram-negative bacteria) | Photocatalytic ROS generation under UV light. Oxidative damage to cellular structures | Titanium dioxide nanoparticles (TiO2 NPs) | 100 μg/mL | 100 μg/mL against P. aeruginosa | [190] |
| Graphene oxide (GO) and reduced graphene oxide (rGO) | E. coli, S. aureus, K. pneumoniae | Physical disruption (sharp edges pierce membranes). Oxidative stress, electron-transfer interactions. | Graphene oxide silver nanocomposite (GO-Ag) | 16 μg/mL | 16 μg/mL | [191] |
| Glycopeptide dendrimers | P. aeruginosa | Glycopeptide dendrimers bind exclusively to the P. aeruginosa lectin LecA via multivalent carbohydrate–lectin interactions, affecting biofilm attachment mechanisms. This binding disrupts the appropriate cell-cell and cell-surface interactions required for biofilm development. The multivalency considerably improves biofilm inhibition while also promoting the dispersal of produced biofilms without killing bacteria. | Tetravalent G2 dendrimers (GalAG2 and GalBG2) and Octavalent G3 dendrimers (GalAxG3 and GalBxG3). | G2—20 µM G3—≈9–13 µM | 13 µM | [192] |
| Nitric oxide (NO)-releasing alkyl-modified poly(amidoamin) (PAMAM) dendrimers | S. mutans | Disrupts bacterial signaling and weakens the biofilm matrix. The hydrophobic alkyl chains enhance bacterial membrane permeability, promoting antimicrobial effects. | Poly(amidoamine) (PAMAM) dendrimers | Not specified | Not reported | [193] |
| Biodegradable nanoemulsions (benzoyl peroxide (BPO) and ciprofloxacin (CIP)) | S. aureus P. aeruginosa | The nanoemulsions carry both benzoyl peroxide, which produces reactive oxygen species (ROS), and ciprofloxacin, a broad-spectrum antibiotic, straight to biofilms. The lipid-based nanoemulsion improves penetration into the biofilm matrix, resulting in sustained and localized drug release. This synergistic combination breaks bacterial membranes, lowers biofilm biomass, and efficiently kills bacteria in wound-associated biofilms. | BPO nanoparticles in a lemongrass oil–based nanoemulgel | 20 µM | 20 µM | [194] |
| Crosslinked nanoemulsions | S. aureus P. aeruginosa | The crosslinked nanoemulsions penetrate biofilms and deliver antibacterial essential oils in a regulated manner. Their crosslinked structure promotes wound stability and retention, hence increasing antibacterial activity. This disrupts bacterial membranes, reduces biofilm biomass, and accelerates wound healing without the use of synthetic medications. | Flavonoids, tannins | 31–125 µg/mL | 31–250 µg/mL | [195] |
| ZIF-8 (Zn-based MOF) | E. coli, S. aureus, MRSA | Releases Zn2+ ions, disrupts membranes, generates ROS | Zinc (Zn) | ~10–100 µg/mL | ~10–100 µg/mL | [196] |
| UiO-66 (Zr-based MOF) | S. aureus, P. aeruginosa | High-stability MOF; used for drug delivery and ROS generation | UiO-66-NH2 (and UiO-66) nanoparticles loaded with cefazolin | MIC measured but not numerically detailed | Only noted as improved | [197] |
| MOF-antibiotic hybrids | Drug-resistant E. coli, S. aureus, MRSA | Controlled antibiotic release enhances efficacy and reduces resistance development | Synthetic antimicrobial peptide (AMP) | 1000 µg/mL | 2000 µg/mL | [198] |
| Ag-loaded nano-metal–organic framework (Ag@nanoMOF) | S. aureus E. coli | The nanoMOF emits silver ions (Ag+), disrupting bacterial membranes and biological functions. The MOF produces reactive oxygen species (ROS), which improves antibacterial and anti-biofilm properties. This dual approach effectively inhibits and eliminates biofilms. | Silver nanoparticles (chemically reduced) | 0.5 mg/mL | 0.5 mg/mL | [151] |
| Cu-MOF (IITI-3) | M. tuberculosis | The anti-mycobacterial activity of INH@IITI-3 demonstrated significant bacterial killing and altered the structural morphology of the bacteria. | Copper (Cu) | 50–500 µg/mL | 50–500 µg/mL | [199] |
| Outer-Membrane Vesicles (OMVs) | P. aeruginosa | OMVs carry β-lactamases (antibiotic resistance), deliver quorum-sensing signals (PQS), and neutralize AMPs. | Amoxicillin (AMX) | 100 µg/mL | 100 µg/mL | [200] |
| Outer-Membrane Vesicles (OMVs) | N. meningitidis | OMVs deliver lipooligosaccharides and porins; modulate immune response and mediate adhesion/invasion. | Zn-MOF (carrier alone) | 10 µg/mL | 10 µg/mL | [201] |
| Outer-Membrane Vesicles (OMVs) | H. pylori | OMVs contain VacA cytotoxin, promote inflammation, and facilitate biofilm formation. | Am-Zn-MOF (Amoxicillin-loaded) | 10 µg/mL | 10 µg/mL | [202] |
| Outer-Membrane Vesicles (OMVs) | B. ovatus | OMVs deliver inulin-degrading enzymes, aiding nutrient sharing and mutualistic gut ecology | LPS, adhesins, OmpA | 10 µg (spot) | 10 µg | [203] |
| Outer-Membrane Vesicles (OMVs) | B. fragilis | OMVs package glycosidases and proteases; aid in polysaccharide digestion and microbial cross-feeding. | Polysaccharide A, OmpA-like proteins, LPS variants | 0.2, 1, 2 µg/mL | 0.2, 1, 2 µg/mL | [204] |
| Polymeric CO-releasing micelles | P. aeruginosa | CO inhibits bacterial respiration and biofilm construction, increasing amikacin’s penetration and potency against bacteria. | Amikacin-loaded PLGA nanoparticles | 5 µg/mL | 5 µg/mL | [205] |
| P(PEGMA-b-DEAEMA) polymeric micelles | C. albicans and C. tropicalis | Promote ~70% biofilm removal and significantly reduce cell viability of both strains. | Ultra-small solid lipid nanoparticles (us-SLNs) | 32 µg/mL | >256 µg/mL | [206] |
| P(PEGMA-b-DEAEMA) polymeric micelles co-delivered with fluconazole (Flu) | C. albicans | Exhibit synergistic effects, leading to a 2.2-log reduction in cell viability. | Poly(ε-caprolactone)-based nanofiber mats loaded with chlorhexidine | 4 µg/mL | 8 µg/mL | [206] |
| Gentamicin-conjugated magnetic nanoparticles (MNPs-G) | S. aureus | Under optimized magnetic field exposure, MNPs-G achieve homogenous distribution over the biofilm, resulting in increased bacterial death. | Silica–gentamicin nanohybrids | ~6.26 µg/mL | ~6.26 µg/mL | [207] |
| Gentamicin-conjugated magnetic nanoparticles (MNPs-G) | E. coli and other ESKAPE pathogens | MNPs-G exhibit bactericidal activity comparable to free gentamicin in solution, effectively killing planktonic cells of various ESKAPE pathogens. | Gentamicin-loaded PLGA nanoparticles | 5 µg/mL | 10 µg/mL | [208] |
| Magnetic iron oxide nanoparticles (MIONPs) | S. aureus | Magnetically propagated MIONPs create artificial pathways within the biofilm matrix, allowing gentamicin to penetrate more deeply. | Iron oxide nanoparticles | 64 µg/mL | 64 µg/mL | [209] |
| TiO2 films (Sol–gel method) | D. geothermalis | Exposure to 360 nm UV light, TiO2’s photocatalytic activity produces reactive oxygen species (ROS), resulting in considerable biofilm reduction (>107 to <106 cells/cm2). | Titanium dioxide nanoparticles | 100 µg/mL | 100 µg/mL | [210] |
| TiO2 films (ALD method) | D. geothermalis | ALD-prepared TiO2 films generate ROS under UV light, disrupting adhesion structures and reducing biofilm density. | Titanium dioxide (TiO2) photocatalytic coatings | UV irradiation of the TiO2: 360 nm light, 20 Wh m−2 | not applicable | [211] |
| Sulfur-doped TiO2 films | D. geothermalis | Doping TiO2 with sulfur did not enhance the biofilm-destroying capacity under UV light exposure. | Photocatalytic TiO2 coatings, prepared via sol–gel and atomic layer deposition (ALD) | UV light exposure-20 W·h m−2 of 360 nm radiation | Not applicable | [212] |
| H2S-releasing polymeric nanoparticles | Sulfate-reducing bacteria (e.g., Desulfovibrio) | Controlled release of H2S modulates bacterial growth by altering redox balance; can inhibit overgrowth of SRB, impacting gut microbial homeostasis. | hydrogen sulfide (H2S) | Not reported | Not reported | [213] |
| Metal sulfide nanoparticles (FeS, MoS2) | F. nucleatum | May release H2S or interact with sulfur metabolism, influencing bacterial survival and virulence factors related to gut dysbiosis and colorectal cancer risk. | Multilayered magnetic nanoparticles (ML-MNPs) composed of an iron oxide core (Fe3O4), followed by inner silver (Ag) and ultrasmall MoS2 | Not reported | Not reported | [214] |
| Sulfur-doped TiO2 nanoparticles | A. baumannii | Enhanced photocatalytic activity produces reactive sulfur species affecting biofilm integrity and bacterial survival through oxidative stress mechanisms. | Green-synthesized anatase TiO2 nanoparticles (~47 nm) | ~7.81 µg/mL | ~7.81 µg/mL | [215] |
| AgPd0.38 nanocages | E. coli, S. aureus, P. aeruginosa | Generates surface-bound ROS via oxidase-like activity, leading to bacterial cell wall disruption and death. | AgPd0.38 | 62.5 µg/mL | 62.5 µg/mL | [216] |
| Mesoporous silica nanoparticles (nMS) loaded with nano-silver (nAg) and chlorhexidine (Chx) | S. mutans (primary cariogenic bacterium) and other oral biofilm bacteria | pH-responsive release of nano-silver and chlorhexidine in acidic biofilm microenvironments; suppresses acid production and biofilm formation. | nMS-nAg-Chx | 18.75 µg/mL | 18.75 µg/mL | [217] |
| Micro-nano hybrid multifunctional motor (MnO2-based) | S. aureus, P. aeruginosa, E. coli | Self-propulsion via oxygen microbubbles (H2O2 + MnO2 catalyst) enables deep biofilm penetration; generates hydroxyl radicals (OH) that disrupt biofilm matrix and kill bacteria. | Manganese dioxide (MnO2) | 1, 2, and 4 mg/mL | 2, 4, and 8 mg/mL | [218] |
4.3. Quorum-Sensing Inhibitors and Anti-Virulence Strategies: Disrupting Bacterial Communication
| Inhibitor (Quorum Sensing) | Target Mechanism | Bacteria | Virulence Suppression | Source | Active Inhibitory Concentration | Reference |
|---|---|---|---|---|---|---|
| CRISPR-Cas9 | Anti-QS Anti-biofilm | E. coli SE15 | Reduced biofilm formation. Downregulation of mqsR, pgaB, pgaC, csgE, and csgF | CRISPR–Cas9 genome editing (donor DNA for homologous recombination; generated ΔluxS mutants from clinical E. coli SE15). | none reported | [228] |
| CRISPR interference | Anti-QS Anti-biofilm | E. coli AK-117 | Reduced biofilm formation | CRISPRi (dCas9 + sgRNAs targeting luxS in E. coli AK-117) | not applicable for CRISPRi studies | [229] |
| Zingerone | Anti-QS Anti-biofilm | P. aeruginosa PAO1 P. aeruginosa clinical isolates. | Reduced biofilm, pyocyanin, hemolysin, elastase, proteases, rhamno lipid production. Reduced swarming, swimming, and twitching motility | Zingerone (phytochemical from ginger) | Not provided in this study | [230] |
| Zeaxanthin | Anti-QS Anti-biofilm | P. aeruginosa PAO1 | Reduced biofilm formation. Downregulated rhlA and lasB expression | Purified from the green alga Chlorella ellipsoidea | 53.5 µM | [231] |
| Solonamides analogs | Anti-QS Anti-toxin | S. aureus | Reduced RNAIII and hla expression. Marginally enhanced biofilm formation | Cyclodepsipeptide isolated from the marine bacterium Photobacterium halotolerans | 5 µg/mL | [232] |
| Flavonoids | Anti-QS | P. aeruginosa PA14 | Reduced pyocyanin production and swarming motility. rhlA transcription inhibition | Naturally produced plant metabolites | Quercetin (100 µM): LasR activity ≈ 20–25% of control. | [233] |
| AIP analogs | Anti-QS Anti-biofilm | S. epidermidis RP62A | Reduced biofilm formation (using non-native agonist of AgrC-type I) | Analogs were created via solid-phase peptide synthesis | AIP-I (native)-99 nM AIP-I D1AS6A-18 nM | [234] |
| Coumarin | Anti-QS Anti-biofilm | P. aeruginosa PAO1 and clinical isolates | 1. Reduced biofilm production, 2. Down-regulation of lasI, rhlI, rhlR, pqsB, pqsC, pqsH, ambBCDE, 3. Reduced protease and pyocyanin production 4. Reduced expression of T3SS secretion system-associated genes | Coumarin–hydroxamic acid conjugate | 3.6 µM | [235] |
| Fluoro-substituted Isothiocyanates | Anti-QS | P. aeruginosa | 1. Reduced pyocyanin production. 2. Reduced swarming motility. 3. Attenuated in vivo virulence of P. aeruginosa PAO1-UW toward C.elegans; 4. Attenuated P. aeruginosa P14 virulence in an ex vivo human skin burn wound model | Synthetic isothiocyanate- and maleimide-containing HSL-like small molecules | 3.6 ± 1.9 µM | [236] |
| Pyridoxal lactohydrazone | Anti-QS | P. aeruginosa PAO1 | Reduced swarming and twitching motility | Condensation of pyridoxal (vitamin B6) with lactic acid hydrazide | Sub-MIC concentrations (8 µg/mL and 32 µg/mL) | [237] |
| Glyceryl trinitrate | Anti-QS Anti-biofilm | P. aeruginosa PAO1 and clinical isolates | Reduced biofilm, pyocyanin and proteases production | Antivirulence compound | 0.25 mg/mL (¼ MIC; sub-inhibitory) | [238] |
| Terrein | Anti-QS Anti-biofilm | P. aeruginosa PAO1 | 1. Reduced elastase, pyocyanin, rhamnolipid, and biofilm production; 2. Attenuated in vivo virulence of P. aeruginosa PAO1 toward C. elegans and mice | Secondary metabolite isolated from the fungus Aspergillus terreus | No half-maximal inhibitory concentration calculated | [239] |
| 1,5-dihydropyrrol-2-ones analogs | Anti-QS | E. coli JB357 gfp reporter strain | QS inhibition | Thioether-linked dihydropyrrol-2-one analogs | 32 µM | [240] |
| Parthenolide | Anti-QS Anti-biofilm | P. aeruginosa PAO1 | Reduced pyocyanin, proteases, and biofilm production | Natural phytochemical | No | [241] |
| Diketopiperazine | Anti-QS Anti-biofilm | B. cenocepacia | 1. Reduced biofilm formation. 2. Reduced protease and siderophore production | Synthetic diketopiperazine analogs | 7.2 ± 0.2 µM | [242] |
| Lysionotin | Anti-toxin Anti-QS | S. aureus | 1. Downregulate fhla and gr expression. 2. Reducedα-hemolysin production | Natural flavonoid | No | [243] |
| Methyl Eugenol-A | Interferes with AHL-regulated functions | C. violaceum | Inhibits EPS production, biofilm formation, and flagellar movement | Natural phenolic found in clove oil | 0.2 mg/mL | [244] |
| Curcumin/10-undecenoic acid | Inhibit LuxS/AI-2 and LasI/LasR QS systems | P. aeruginosa, B. subtilis | Counteracts bacterial pathogenicity and virulence | Natural polyphenolic compound | MIC values: 62.5 μg/mL | [245] |
| Ajoene | Downregulates QS genes | P. aeruginosa | Inhibits virulence factors and biofilm formation | Bioassay-guided fractionation of garlic oil | 15 µM | [246] |
| Tanreqing (TRQ) formula | Inhibits upstream QS regulators (e.g., GacS/GacA, PprA/PprB) | P. aeruginosa | Inhibits virulence factors | Lab-prepared extract combining all five herbs (Huang Qin, Jin Yin Hua, Lian Qiao, Xiong Dan, Shan Yang Jiao) | Results are shown across TRQ dilutions (1/4, 1/8, 1/16 TRQ), not as IC50 values | [247] |
| Paecilomycone | Inhibits PQS and HHQ synthesis in PQS system | P. aeruginosa | Inhibits virulence factors | natural compound originally isolated from a fungal source-specifically Paecilomyces species | 96.5 µM | [248] |
| Synthetic quorum-sensing inhibitors | Mimicking or blocking natural quorum-sensing signals to disrupt communication pathways. | Ralstonia solanacearum species complex | Impaired coordination of virulence factor expression, leading to diminished disease symptoms in host plants. | N-sulfonyl homoserine lactone analogs | 1.66-4.91 µM | [249] |
| Ralfuranones | Interference with quorum-sensing signaling molecules, affecting the regulation of virulence factors. | Ralstonia solanacearum species complex | Decreased expression of virulence genes, resulting in reduced pathogenicity. | Derived from the 3-hydroxymethyl-2-methyl-4(1H)-quinolone | 23.7 µM | [250] |
| DSF analogs | Compete with natural DSF molecules, disrupting normal QS signaling. | S. maltophilia, P. aeruginosa | Altered biofilm dynamics and reduced antibiotic resistance, diminishing virulence. | Natural DSF comes from Xanthomonas, Burkholderia, Stenotrophomonas, etc. via biosynthesis. | 0.5 µM | [222] |
| Enzymatic degradation (e.g., DSF hydrolases) | Degrade DSF molecules, lowering their concentration and interfering with QS. | E. carotovora P. aeruginosa | Broad-spectrum reduction in QS-mediated virulence traits, including toxin production and biofilm formation. | Stenotrophomonas maltophilia | Not reported | [251] |
| Essential oils (e.g., clove, cinnamon, oregano) | Disrupt AHL-mediated signaling pathways, inhibiting signal synthesis and reception. | P. fluorescens, S. putrefaciens | Reduction in biofilm formation and extracellular enzyme production, leading to decreased spoilage activity. | Eugenol, β-caryophyllene | MIC: 0.2 mg/mL | [252] |
| Phenolic compounds (e.g., vanillin, eugenol) | Interfere with QS signal molecules and inhibit biofilm formation. | A. hydrophila, L. monocytogenes | Suppression of biofilm development and virulence factor expression, enhancing food safety. | Vanilla bean Clove, cinnamon, basil | MIC: 250 µg/mL MIC: 700 mg/L | [253] |
| Plant extracts (e.g., garlic, ginger) | Contain compounds that mimic or degrade QS signals, disrupting communication. | E. coli, S. enterica | Inhibition of QS-regulated behaviors, including toxin production and motility, reducing pathogenicity. | Allicin, ajoene and S-allyl cysteine | MIC: 0.325-0.625 mg/mL | [254] |
| Lactic acid bacteria metabolites | Produce bacteriocins and acids that interfere with QS systems. | S. thermophiles, L. bulgaricus | Decrease in QS activity, leading to reduced biofilm formation and spoilage potential. | Lactic acid, formic acid, folic acid, pyruvic acid, glutathione | Not reported | [255] |
| AHL analogs | Compete with natural AHLs for receptor binding, disrupting QS signaling. | P. aeruginosa, E. coli | Inhibition of biofilm formation and suppression of virulence gene expression. | Natural phenolic compound | 100 µM | [10] |
| Furanones | Interfere with QS signal reception and stability. | Vibrio spp., P. aeruginosa | Reduction in biofilm development and attenuation of virulence factors. | Synthetic brominated furanone derivative | 50 µM | [256] |
| Plant-derived compounds (e.g., garlic extract, cinnamaldehyde) | Inhibit QS signal synthesis and reception pathways. | E. coli, S. enterica | Decreased biofilm formation and prevention of VBNC state induction. | Allium sativum | 0.325–0.625 mg/mL | [257] |
| Enzymatic degraders (e.g., AHL lactonases, acylases) | Degrade QS signaling molecules, disrupting communication. | E. coli, S. enterica | Impaired biofilm maturation and reduced virulence expression. | AHL lactonase (AiiA) Bacillus sps | Not reported | [258] |
| Adenosine | Modulates QS pathways, potentially interfering with signal molecule synthesis or reception. | Cylospora | May reduce expression of virulence factors and biofilm formation, enhancing host immune response. | Endogenous nucleoside; found in all living cells | Not reported | [259] |
| AI-2, DSF family signals | Biofilm formation, motility, and antibiotic resistance | S. aureus, E. coli | Decrease in QS-regulated behaviors, including toxin production and biofilm formation. | Universally produced by many Gram-negative and Gram-positive bacteria | Not applicable | [80] |
| Probiotic-produced Autoinducers | Interfere with pathogenic QS signals by competitive binding or signal degradation, disrupting bacterial communication. | E. coli, S. enterica, C. difficile | Reduced biofilm formation and downregulation of virulence gene expression. | Lactobacillus, Bifidobacterium, E. coli Nissle 1917 | Not reported | [260] |
| Probiotic biofilm formation | Colonizes intestinal mucosa to outcompete pathogens for adhesion, indirectly suppressing QS-mediated virulence. | E. faecalis, S. aureus | Limits pathogen colonization and strengthens mucosal barrier function | Enterococcus faecalis, Lactobacillus spp., Bifidobacterium spp. | Not reported | [261] |
| Plant-derived QS inhibitors (e.g., flavonoids, phenolics) | Block QS signal synthesis or reception, degrade signaling molecules. | P. aeruginosa, Vibrio spp. | Inhibits biofilm development and virulence factor production. | Various plant sources | 50 µM | [262] |
| Immune modulation by probiotics | Stimulates host immune responses, producing cytokines that counteract infection and inflammation. | S. enterica, C. difficile | Supports intestinal barrier integrity and suppresses pathogen-induced inflammation. | Citrus fruits, green tea, various plants | 50–100 µM | [263] |
| Synthetic QS inhibitors | Engineered molecules or circuits designed to interfere with QS signaling pathways. | P. aeruginosa | Disruption of QS-mediated behaviors such as biofilm formation, virulence factor production, and antibiotic resistance. | Engineered small molecules or genetic circuits designed via synthetic biology | 10–100 µM for synthetic analogs | [264] |
| Probiotic interference | Probiotic strains engineered to produce QS inhibitors or compete with pathogens for QS signals. | E. coli, S. enterica, C. difficile | Reduction in pathogen colonization and virulence through competitive inhibition and signal interference. | Engineered or natural probiotic strains (Lactobacillus, Bifidobacterium, E. coli Nissle 1917) | Rarely specified | [265] |
| Phage-mediated QS disruption | Bacteriophages engineered to degrade QS signals or produce QS inhibitors. | P. aeruginosa | Decreased biofilm formation and virulence factor expression through targeted QS disruption. | Engineered bacteriophages designed to degrade QS signals or produce QS inhibitors | Not reported | [266] |
| Sulfonamide-based DSF analogs | Interfere with DSF receptor binding, disrupting DSF-mediated QS. | S. maltophilia, B. cepacia | Reduced biofilm formation and virulence factor production. | Chemically synthesized small molecules mimicking DSF structure | 5–50 µM | [267] |
| Retinoic acid (RA) | Modulates DSF-mediated QS, affecting biofilm formation and motility. | S. maltophilia | Decreased biofilm formation and motility, enhancing antimicrobial efficacy. | Naturally occurring metabolite of Vitamin A | 50 µM | [268] |
| Endogenous DSF analog | Mimics DSF, disrupting DSF-mediated QS. | S. maltophilia | Inhibition of biofilm formation and motility. | Naturally produced by bacteria or chemically synthesized to mimic DSF | Not reported | [269] |
| Coumarin–chalcone conjugate | Anti-QS Anti-biofilm | P. eruginosa | Decreased biofilm formation and virulence factor expression through targeted QS disruption. | Chemically synthesized conjugate combining coumarin and chalcone scaffolds | 20–50 µM | [270] |
| QteE | Sequesters LasR, increasing the threshold for QS activation. | P. aeruginosa | Delayed expression of QS controlled genes, reducing early activation of virulence factors. | Protein from Pseudomonas aeruginosa | Not reported | [271] |
| QscR | Binds to LasR and RhlR, preventing premature activation of QS. | P. aeruginosa | Suppressed expression of QS controlled genes, mitigating early virulence factor production. | Transcriptional regulator from Pseudomonas aeruginosa | 56–62 nM | [221] |
| QslA | Inhibits LasR and RhlR activity, modulating QS response. | P. aeruginosa | Reduced biofilm formation and virulence factor production. | Transcriptional regulator from Pseudomonas aeruginosa | Not reported | [272] |
| Tyramine | Quorum-sensing inhibition via interference with the CepIR system | B. cenocepacia. | Decreased biofilm formation, reduced motility, attenuated virulence in Caenorhabditis elegans model. | Naturally occurring biogenic amine | Not reported | [273] |
| QStatin | Specifically targets LuxO, a central response regulator in the quorum-sensing pathway of Vibrio species. QStatin binds selectively to phosphorylated LuxO, blocking its ability to activate small regulatory RNAs (sRNAs), which are essential for downstream quorum-sensing responses. | V. cholerae V. harveyi | Disrupts quorum-sensing-controlled gene expression Reduces virulence-related phenotypes Selective: QStatin does not inhibit general bacterial growth or non-target bacteria significantly | Small-molecule inhibitor designed to selectively target LuxO | 2–5 µM | [274] |
| Furanones | Competitive inhibition of AHL receptor proteins (LuxR-type regulators) | P. aeruginosa, E. coli, V. spp. | Reduces biofilm formation, motility, and production of virulence factors (e.g., toxins) | Synthetic halogenated furanone | 3–9 µM | [275] |
| Baicalein | Downregulates quorum-sensing-regulated gene expression | P. aeruginosa, S. aureus | Inhibits biofilm formation and reduces toxin production | Flavonoid from Scutellaria baicalensis and other plants | Not specified | [276] |
| Quercetin | Interferes with AI-2 signaling pathways | E. coli, S. spp. | Inhibits quorum-sensing gene expression, reduces swarming, and suppresses virulence | Naturally occurring flavonoid | 281 µM for biofilm inhibition | [227] |
| Vanillin | Blocks AHL signal molecule activity | C. violaceum, P. aeruginosa | Inhibits violacein pigment production and biofilm formation | Natural compound from vanilla beans | IC50 = 0.81 mM; MIC = 16 mM | [277] |
| Furanone C-30 | Inhibits LasR receptor in the las quorum-sensing system | P. aeruginosa | Reduces biofilm formation, elastase activity, and enhances tobramycin susceptibility (initially) | Synthetic brominated furanone | IC50: 3–9 µM | [278] |
| QS Inhibitor-loaded biomimetic nanoparticles | Inhibits AI-2 quorum-sensing system, disrupts cell communication + induces ferroptosis-like bacterial death via lipid peroxidation | Methicillin-Resistant S. aureus (MRSA) | Suppresses virulence gene expression, reduces biofilm formation, and promotes bacterial death in infected lung tissue | Natural compounds or synthetic analogs | 2–5 µM | [279] |
| L-mimosine | Inhibits quorum-sensing-regulatory small RNAs (sRNAs) specifically, qrr sRNAs in Vibrio species | V. harveyi E. coli P. aeruginosa | Reduces bioluminescence (in Vibrio), biofilm formation, motility, and toxin production | Mimosa pudica and Leucaena species | Not well reported | [280] |
| 2-(4-(acridin 9yl amino)phenyl)isoindoline-1,3-dione derivatives | Inhibits the Pseudomonas Quinolone Signal (PQS) quorum-sensing system | P. aeruginosa | Reduces virulence-related phenotypes | Acridine-based isoindoline derivatives | Not well reported | [281] |
4.4. Phage Therapy and Bacteriophage-Derived Enzymes: Precision Tools Against Biofilms
4.5. Host-Directed Therapies and Immune Modulation: Rewiring Defenses Against Biofilms
5. Translational Challenges and Future Directions in Biofilm Therapeutics
5.1. Regulatory Bottlenecks and Standardization Gaps
5.2. Ethical and Biosafety Considerations
5.3. Microbiome-Preserving Strategies and Precision Design
5.4. Towards Real-Time, Autonomous Therapies
5.5. Pathways to Clinical Translation
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| AFM | Atomic Force Microscopy |
| AHLs | N-acyl homoserine lactones |
| BLI | Bioluminescence Imaging |
| CF | cystic fibrosis |
| CFU | Colony-forming unit |
| CLSM | Confocal Laser Scanning Microscopy |
| CNNs | Convolutional Neural Networks |
| Cryo-EM | Cryo-Electron Microscopy |
| DFU | Diabetic foot ulcers |
| DHM | Digital Holographic Microscopy |
| EPS | Extracellular Polymeric Substance |
| GANs | Generative Adversarial Networks |
| GNNs | Graph Neural Networks |
| HDTs | Host-directed therapies |
| LLSM | Lattice Light-Sheet Microscopy |
| LSTM | Long Short-Term Memory |
| MBEC | Minimum biofilm eradication concentration |
| MOFs | Metal–organic frameworks |
| NET | Neutrophil extracellular trap |
| NPs | Engineered nanoparticles |
| OCT | Optical Coherence Tomography |
| PAS | Phage antibiotic synergy |
| PINNs | Physics-Informed Neural Networks |
| QS | Quorum sensing |
| QSIs | Quorum-sensing inhibitors |
| RF | Random Forests |
| RIPs | RNAIII-inhibiting peptides |
| RL | Reinforcement Learning |
| RSI | Raman Spectroscopy Imaging |
| SEM | Scanning Electron Microscopy |
| STED | Stimulated Emission Depletion Microscopy |
| STORM | Stochastic Optical Reconstruction Microscopy |
| SVMs | Support Vector Machines |
| TEM | Transmission Electron Microscopy |
| A. baumannii | Acinetobacter baumannii |
| A. hydrophila | Aeromonas hydrophila |
| B. cenocepacia | Burkholderia cenocepacia |
| B. cepacian | Burkholderia cepacian |
| B. fragilis | Bacteroides fragilis |
| B. ovatus | Bacteroides ovatus |
| C. albicans | Candida albicans |
| C. difficile | Clostridium difficile |
| C. glabrata | Candida glabrata |
| C. violaceum | Chromobacterium violaceum |
| E. carotovora | Erwinia carotovora |
| E. coli | Escherichia coli |
| F. nucleatum | Fusobacterium nucleatum |
| H. pylori | Helicobacter pylori |
| K. pneumoniae | Klebsiella pneumoniae |
| L. monocytogenes | Listeria monocytogenes |
| M. luteus | Micrococcus luteus |
| M. tuberculosis | Mycobacterium tuberculosis |
| N. meningitidis | Neisseria meningitidis |
| P. aeruginosa | Pseudomonas aeruginosa |
| P. fluorescens | Pseudomonas fluorescens |
| P. gingivalis | Porphyromonas gingivalis |
| S. aureus | Staphylococcus aureus |
| S. enterica | Salmonella enterica |
| S. fonticola | Serratia fonticola |
| S. maltophilia | Stenotrophomonas maltophilia |
| S. mutan | Streptococcus mutans |
| S. putrefaciens | Shewanella putrefaciens |
| S. suis | Streptococcus suis |
| V. cholerae | Vibrio cholerae |
| V. harveyi | Vibrio harveyi |
| X. campestris | Xanthomonas campestris |
References
- Alam, K.; Farraj, D.A.A.; Mah-e-Fatima, S.; Yameen, M.A.; Elshikh, M.S.; Alkufeidy, R.M.; Mustafa, A.E.-Z.M.A.; Bhasme, P.; Alshammari, M.K.; Alkubaisi, N.A.; et al. Anti-biofilm activity of plant derived extracts against infectious pathogen-Pseudomonas aeruginosa PAO1. J. Infect. Public Health 2020, 13, 1734–1741. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Yin, R.; Cheng, J.; Lin, J. Bacterial biofilm formation on biomaterials and approaches to its treatment and prevention. Int. J. Mol. Sci. 2023, 24, 11680. [Google Scholar] [CrossRef]
- Bhasme, P.; Wei, Q.; Xu, A.M.; Naqvi, S.T.A.; Wang, D.; Ma, L.Y.Z. Evaluation and characterization of the predicted diguanylate cyclase-encoding genes in. Microbiologyopen 2020, 9, e975. (In English) [Google Scholar] [CrossRef]
- Selim, H.M.R.M.; Gomaa, F.A.M.; Alshahrani, M.Y.; Morgan, R.N.; Aboshanab, K.M. Phage therapeutic delivery methods and clinical trials for combating clinically relevant pathogens. Ther. Deliv. 2025, 16, 247–269. [Google Scholar]
- Xu, A.; Wang, D.; Wang, Y.; Zhang, L.; Xie, Z.; Cui, Y.; Bhamse, P.; Yu, H.; Zhang, X.; Li, D.; et al. Mutations in surface-sensing receptor WspA lock the Wsp signal transduction system into a constitutively active state. Environ. Microbiol. 2024, 24, 247–269. [Google Scholar] [CrossRef]
- Hanke, M.L.; Kielian, T. Deciphering mechanisms of staphylococcal biofilm evasion of host immunity. Front. Cell. Infect. Microbiol. 2012, 2, 62. [Google Scholar] [CrossRef]
- Silva, A.; Silva, V.; López, M.; Rojo-Bezares, B.; Carvalho, J.A.; Castro, A.P.; Sáenz, Y.; Igrejas, G.; Poeta, P. Antimicrobial resistance, genetic lineages, and biofilm formation in Pseudomonas aeruginosa isolated from human infections: An emerging one health concern. Antibiotics 2023, 12, 1248. [Google Scholar] [CrossRef]
- Afrasiabi, S.; Partoazar, A. Targeting bacterial biofilm-related genes with nanoparticle-based strategies. Front. Microbiol. 2024, 15, 1387114. [Google Scholar]
- Azeredo, J.; García, P.; Drulis-Kawa, Z. Targeting biofilms using phages and their enzymes. Curr. Opin. Biotechnol. 2021, 68, 251–261. [Google Scholar] [CrossRef] [PubMed]
- Juszczuk-Kubiak, E. Molecular aspects of the functioning of pathogenic bacteria biofilm based on quorum sensing (QS) signal-response system and innovative non-antibiotic strategies for their elimination. Int. J. Mol. Sci. 2024, 25, 2655. [Google Scholar] [PubMed]
- Ding, Y.; Chen, S. A bibliographic outlook: Machine learning on biofilm. Res. Dir. Biotechnol. Des. 2025, 3, e2. [Google Scholar] [CrossRef]
- Seneviratne, C.J.; Suriyanarayanan, T.; Widyarman, A.S.; Lee, L.S.; Lau, M.; Ching, J.; Delaney, C.; Ramage, G. Multi-omics tools for studying microbial biofilms: Current perspectives and future directions. Crit. Rev. Microbiol. 2020, 46, 759–778. [Google Scholar] [CrossRef]
- Almatroudi, A. Investigating biofilms: Advanced methods for comprehending microbial behavior and antibiotic resistance. Front. Biosci. Landmark 2024, 29, 133. [Google Scholar]
- Grudlewska-Buda, K.; Skowron, K.; Gospodarek-Komkowska, E. Comparison of the intensity of biofilm formation by Listeria monocytogenes using classical culture-based method and digital droplet PCR. AMB Express 2020, 10, 75. [Google Scholar] [CrossRef] [PubMed]
- Haney, E.F.; Trimble, M.J.; Hancock, R.E. Microtiter plate assays to assess antibiofilm activity against bacteria. Nat. Protoc. 2021, 16, 2615–2632. [Google Scholar] [CrossRef]
- Walter, B.M.; Cartman, S.T.; Minton, N.P.; Butala, M.; Rupnik, M. The SOS response master regulator LexA is associated with sporulation, motility and biofilm formation in Clostridium difficile. PLoS ONE 2015, 10, e0144763. [Google Scholar]
- Rajamani, S.; Sandy, R.; Kota, K.; Lundh, L.; Gomba, G.; Recabo, K.; Duplantier, A.; Panchal, R.G. Robust biofilm assay for quantification and high throughput screening applications. J. Microbiol. Methods 2019, 159, 179–185. [Google Scholar] [CrossRef] [PubMed]
- Thieme, L.; Hartung, A.; Tramm, K.; Graf, J.; Spott, R.; Makarewicz, O.; Pletz, M.W. Adaptation of the start-growth-time method for high-throughput biofilm quantification. Front. Microbiol. 2021, 12, 631248. [Google Scholar]
- Ayon, N.J. High-throughput screening of natural product and synthetic molecule libraries for antibacterial drug discovery. Metabolites 2023, 13, 625. [Google Scholar] [CrossRef]
- Jia, J.; Xue, X.; Guan, Y.; Fan, X.; Wang, Z. Biofilm characteristics and transcriptomic profiling of Acinetobacter johnsonii defines signatures for planktonic and biofilm cells. Environ. Res. 2022, 213, 113714. [Google Scholar] [CrossRef]
- Gilbert, P.; Maira-Litran, T.; McBain, A.J.; Rickard, A.H.; Whyte, F.W. The physiology and collective recalcitrance of microbial biofilm communities. Adv. Microb. Physiol. 2002, 46, 203–256. [Google Scholar]
- Wu, Y.; Li, X.; Fu, X.; Huang, X.; Zhang, S.; Zhao, N.; Ma, X.; Saiding, Q.; Yang, M.; Tao, W. Innovative nanotechnology in drug delivery systems for advanced treatment of posterior segment ocular diseases. Adv. Sci. 2024, 11, 2403399. [Google Scholar]
- Kragh, K.N.; Alhede, M.; Kvich, L.; Bjarnsholt, T. Into the well—A close look at the complex structures of a microtiter biofilm and the crystal violet assay. Biofilm 2019, 1, 100006. [Google Scholar] [CrossRef] [PubMed]
- Amran, S.S.D.; Syaida, A.A.R.; Jalil, M.T.M.; Nor, N.H.M.; Yahya, M.F.Z.R. Preparation of Biofilm Assay Using 96-Well and 6-Well Microplates for Quantitative Measurement and Structural Characterization: A Review. Sci. Lett. 2024, 18, 121–134. [Google Scholar]
- Salcedo, F.; Pereyra, C.M.; Arruebarrena Di Palma, A.; Lamattina, L.; Creus, C.M. Methods for studying biofilms in Azospirillum and other PGPRs. In Handbook for Azospirillum: Technical Issues and Protocols; Springer: Berlin/Heidelberg, Germany, 2015; pp. 199–229. [Google Scholar]
- Cullimore, R. Monitoring Methodologies for Biofouling Events. In Microbiology of Well Biofouling; CRC Press: Boca Raton, FL, USA, 2018; pp. 77–136. [Google Scholar]
- Kırmusaoğlu, S. Biofilm and Screening Antibiofilm Activity of Agents. In Antimicrobials, Antibiotic Resistance, Antibiofilm Strategies and Activity Methods; IntechOpen: London, UK, 2019; p. 99. [Google Scholar]
- Nwaiwu, J. Development of Alternative Protocols for Antimicrobial Efficacy Testing of Antimicrobial Surfaces. Master’s Thesis, University of Waterloo, Waterloo, CN, Canada, 2020. [Google Scholar]
- Ogundero, A. The Theoretical and Experimental Exploration of the Use of Predatory Bacteria to Control Biofilms. Ph.D. Thesis, University of Glasgow, Glasgow, UK, 2023. [Google Scholar]
- Schmutzler, K.; Schmid, A.; Buehler, K. A three-step method for analysing bacterial biofilm formation under continuous medium flow. Appl. Microbiol. Biotechnol. 2015, 99, 6035–6047. [Google Scholar] [CrossRef] [PubMed]
- Goeres, D.M.; Walker, D.K.; Buckingham-Meyer, K.; Lorenz, L.; Summers, J.; Fritz, B.; Goveia, D.; Dickerman, G.; Schultz, J.; Parker, A.E. Development, standardization, and validation of a biofilm efficacy test: The single tube method. J. Microbiol. Methods 2019, 165, 105694. [Google Scholar] [CrossRef]
- Wilson, M.-M. In Vitro Study of Effectivity of Bacteriophages in the Prevention and Degradation of Biofilms Formed by Uropathogenic E. coli in Urinary Chatheters. Master’s Thesis, University of the Witwatersrand, Johannesburg, South Africa, 2018. [Google Scholar]
- Allkja, J. Development of Minimum Information Guidelines and Standardized in Vitro Methods for Biofilm Experiments. Ph.D. Thesis, Universidade do Porto, Porto, Portugal, 2021. [Google Scholar]
- Xiong, Z.; Tian, X.; Wang, G.; Song, X.; Xia, Y.; Zhang, H.; Ai, L. Development of a high-throughput screening method for exopolysaccharide-producing Streptococcus thermophilus based on Congo red. Food Res. Int. 2022, 162, 112094. [Google Scholar] [PubMed]
- Oladoye, P.O.; Bamigboye, M.O.; Ogunbiyi, O.D.; Akano, M.T. Toxicity and decontamination strategies of Congo red dye. Groundw. Sustain. Dev. 2022, 19, 100844. [Google Scholar] [CrossRef]
- Harja, M.; Buema, G.; Bucur, D. Recent advances in removal of Congo Red dye by adsorption using an industrial waste. Sci. Rep. 2022, 12, 6087. [Google Scholar] [CrossRef]
- Sudha, S.S.; Aranganathan, V. Antibiofilm analysis, synergistic potential and biocompatibility evaluation of a bacteriocin from Bacillus subtilis (MK733983). Indian J. Microbiol. 2024, 64, 1646–1663. [Google Scholar] [CrossRef] [PubMed]
- Papadopoulos, C. A Novel Experimental Platform for Monitoring and Imaging Bacterial Biofilm Growth in Porous Media Flows. Ph.D. Thesis, Institut National Polytechnique de Toulouse-INPT, Toulouse, France, 2023. [Google Scholar]
- Rabbitt, D.; Villapún, V.M.; Carter, L.N.; Man, K.; Lowther, M.; O’Kelly, P.; Knowles, A.J.; Mottura, A.; Tang, Y.T.; Luerti, L. Rethinking biomedical titanium alloy design: A review of challenges from biological and manufacturing perspectives. Adv. Healthc. Mater. 2025, 14, 2403129. [Google Scholar]
- Dhanasinghe, C.; Kao, C.M.; Liu, P.F.; Surampalli, R.Y.; Zhang, T.C.; Al-Hashimi, B.M. Advanced Techniques for Sampling, Quantification, and Characterization of Microplastics. In Microplastics in the Environment: Fate, Impacts, Removal, and Management; Wiley: Hoboken, NJ, USA, 2025; pp. 69–106. [Google Scholar]
- Jones, L.M. Microbiologically Influenced Corrosion: Development of a Model System to Investigate the Role of Biofilm Communities Within MIC and Their Control Using Industrial Biocides. Ph.D. Thesis, University of Southampton, Southampton, UK, 2025. [Google Scholar]
- Bourn, M.D.; Daly, L.F.; Huggett, J.F.; Braybrook, J.; Rivera, J.F. Evaluation of image analysis tools for the measurement of cellular morphology. Front. Cell Dev. Biol. 2025, 13, 1572212. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues, C.F.; Allkja, J.; Mendes, L.; Azevedo, A.S. Methods for the visualization of multispecies biofilms. In Multispecies Biofilms: Technologically Advanced Methods to Study Microbial Communities; Springer: Berlin/Heidelberg, Germany, 2022; pp. 35–78. [Google Scholar]
- Lee, D.; Lee, H.-S.; Lee, M.; Kang, M.; Kim, G.; Kim, T.Y.; Lee, N.Y.; Park, Y. Advances in imaging techniques for the study of individual bacteria and their pathophysiology. arXiv 2025, arXiv:2501.01592. [Google Scholar] [CrossRef]
- Relucenti, M.; Familiari, G.; Donfrancesco, O.; Taurino, M.; Li, X.; Chen, R.; Artini, M.; Papa, R.; Selan, L. Microscopy methods for biofilm imaging: Focus on SEM and VP-SEM pros and cons. Biology 2021, 10, 51. [Google Scholar] [CrossRef]
- Gerardi, D.; Bernardi, S.; Bruni, A.; Falisi, G.; Botticelli, G. Characterization and morphological methods for oral biofilm visualization: Where are we nowadays? AIMS Microbiol. 2024, 10, 391. [Google Scholar] [CrossRef] [PubMed]
- Choubey, S.; Deshpande, J. A Study on Exploring the Composition, Structure and Innovative Analytical Techniques Developed in Biomembrane Research. Biosci. Biotechnol. Res. Asia 2025, 22, 63. [Google Scholar] [CrossRef]
- Samuel, H.S.; Ekpan, F.-D.M. The use of Scanning Electron Microscopy (SEM) for medical application: A mini review. Eurasian J. Sci. Technol. 2024, 4, 289–294. [Google Scholar]
- McCutcheon, J.; Southam, G. Advanced biofilm staining techniques for TEM and SEM in geomicrobiology: Implications for visualizing EPS architecture, mineral nucleation, and microfossil generation. Chem. Geol. 2018, 498, 115–127. [Google Scholar] [CrossRef]
- Rohde, M. Bacterial ultrastructure. In Molecular Medical Microbiology; Elsevier: Amsterdam, The Netherlands, 2024; pp. 23–43. [Google Scholar]
- Van Hoogstraten, S.; Kuik, C.; Arts, J.; Cillero-Pastor, B. Molecular imaging of bacterial biofilms—A systematic review. Crit. Rev. Microbiol. 2024, 50, 971–992. [Google Scholar] [CrossRef]
- Millan-Solsona, R.; Brown, S.R.; Zhang, L.; Madugula, S.S.; Zhao, H.; Dumerer, B.; Bible, A.N.; Lavrik, N.V.; Vasudevan, R.K.; Biswas, A. Analysis of Biofilm Assembly by Large Area Automated AFM. NPJ Biofilms Microbiomes 2025, 11, 75. [Google Scholar]
- Das, M.; Khamaru, D.; Santra, S.; Gupta, R.; Das, S.; Banerjee, R. The Enigmatic World of Biofilm. In Biofilm Applications to Revolutionize Food Technology; Springer: Berlin/Heidelberg, Germany, 2025; pp. 1–26. [Google Scholar]
- Albalawi, T. Strategies for Detecting Bacterial Biofilms: Unveiling the Hidden World of Microbial Aggregates. Egypt. J. Soil Sci. 2024, 64, 1069–1096. [Google Scholar] [CrossRef]
- Zhang, Y.; Young, P.; Traini, D.; Li, M.; Ong, H.X.; Cheng, S. Challenges and current advances in in vitro biofilm characterization. Biotechnol. J. 2023, 18, 2300074. [Google Scholar] [CrossRef]
- Coppola, F.; Fratianni, F.; Bianco, V.; Wang, Z.; Pellegrini, M.; Coppola, R.; Nazzaro, F. New Methodologies as Opportunities in the Study of Bacterial Biofilms, Including Food-Related Applications. Microorganisms 2025, 13, 1062. [Google Scholar] [CrossRef]
- Müller, T.; Schumann, C.; Kraegeloh, A. STED microscopy and its applications: New insights into cellular processes on the nanoscale. ChemPhysChem 2012, 13, 1986–2000. [Google Scholar] [CrossRef]
- Blom, H.; Widengren, J. Stimulated emission depletion microscopy. Chem. Rev. 2017, 117, 7377–7427. [Google Scholar] [CrossRef] [PubMed]
- Di Martino, P. Extracellular polymeric substances, a key element in understanding biofilm phenotype. AIMS Microbiol. 2018, 4, 274. [Google Scholar] [CrossRef]
- Thomsen, H. Biophotonics Targeting Pharmaceutical Challenges. Ph.D. Thesis, University of Gothenburg, Gothenburg, Sweden, 2018. [Google Scholar]
- Kromer, C. Nanoscale Tools for Biosensing and Treatment of Bacterial Biofilms and Eukaryotic Cells. Ph.D. Thesis, Freie Universität Berlin, Berlin, Germany, 2024. [Google Scholar]
- Bharadwaj, A.; Kumar, A.; Mathew, S.P.; Mitra, R.; Bhattacharyya, J.; Jaganathan, B.G.; Boruah, B.R. Advancing cellular insights: Super-resolution STORM imaging of cytoskeletal structures in human stem and cancer cells. Biochem. Biophys. Rep. 2024, 39, 101798. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Zhang, M.; Wang, Y.; Donarski, E.; Gahlmann, A. Optically accessible microfluidic flow channels for noninvasive high-resolution biofilm imaging using lattice light sheet microscopy. J. Phys. Chem. B 2021, 125, 12187–12196. [Google Scholar] [CrossRef]
- Hassan, L.; Rukan, M.; Mohammad, E.; Syed, M.A. Recent Advances in Microbiology Techniques for Imaging Microbes and Their Components. In Microbiology in the Era of Artificial Intelligence; CRC Press: Boca Raton, FL, USA, 2025; pp. 103–135. [Google Scholar]
- Xu, J.; Ma, H.; Liu, Y. Stochastic optical reconstruction microscopy (STORM). Curr. Protoc. Cytom. 2017, 81, 12.46.1–12.46.27. [Google Scholar] [CrossRef]
- Bharadwaj, A.; Kumar, A.; Kalita, R.; Mitra, R.; Sharma, A.; Jaganathan, B.G.; Bhattacharyya, J.; Sarma, A.; Garcia, E.; Kumar, S. Super-resolved fluorescence imaging utilising accessible stochastic optical reconstruction microscopy (easySTORM) implemented on a low-cost, modular open-source (openFrame) microscope. Meas. Sci. Technol. 2024, 35, 125402. [Google Scholar]
- Liu, H.; Wang, X.; Wang, Z.; Shen, Y. Evaluation of bacterial biofilm, smear layer, and debris removal efficacy of a hydro-dynamic cavitation system with physiological saline using a new ex vivo model: A CLSM and SEM study. BMC Oral Health 2025, 25, 95. [Google Scholar] [CrossRef]
- Tartari, T.; Estrela, C.; de Araújo, L.B.B.; Graeff, M.S.Z.; de Andrade, F.B.; Duarte, M.A.H. Use of confocal laser scanning microscopy to evaluate the metal ion removal and destabilization of Enterococcus faecalis biofilms by EDTA and etidronic acid. Odontology 2025, 1–9. [Google Scholar] [CrossRef]
- Dhahri, S.; Marlière, C. Atomic Force Microsocopy: Key Unconventional Approach for Bacterial Nanotubes Characterization In Vivo. ACS Omega 2024, 9, 46950–46959. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Xue, J.; Zhu, H.; Chen, S.; Wang, Y.; Xiao, Z.; Luo, Y. Advances in biofilm characterization: Utilizing rheology and atomic force microscopy in foods and related fields. Adv. Compos. Hybrid Mater. 2024, 7, 143. [Google Scholar] [CrossRef]
- Medina-Ramirez, I.E.; Macías-Díaz, J.E.; Masuoka-Ito, D.; Zapien, J.A. Holotomography and atomic force microscopy: A powerful combination to enhance cancer, microbiology and nanotoxicology research. Discov. Nano 2024, 19, 64. [Google Scholar]
- Francius, G. Correlation between nanomechanical and chemical features of crosslinked hydrogels combining AFM with chemometric methods. In Proceedings of the AFM BioMed Conference, Barcelona, Spain, 8–11 April 2025; Volume 469, pp. 100–133. [Google Scholar]
- Heydari, S.; Liu, J. High-throughput cryo-electron tomography enables multiscale visualization of the inner life of microbes. Curr. Opin. Struct. Biol. 2025, 93, 103065. [Google Scholar]
- Viljoen, A.; Mathelié-Guinlet, M.; Ray, A.; Strohmeyer, N.; Oh, Y.J.; Hinterdorfer, P.; Müller, D.J.; Alsteens, D.; Dufrêne, Y.F. Force spectroscopy of single cells using atomic force microscopy. Nat. Rev. Methods Primers 2021, 1, 63. [Google Scholar]
- Mohammadi, A.T.; Naderi, N.; Kenari, P.B.; Mir, A.; Sabouri, F.; Amin, M.; Fatahi, G. Advancements in Pharmacology: Integrating Molecular Mechanisms, Therapeutics, and Drug Development; Nobel Sciences: Stockholm, Sweden, 2025. [Google Scholar]
- Li, K.; Li, D.; Gao, L.; Weng, Z.; Zheng, M.; He, Y.; Li, Z. Application of optical coherence tomography for morphological change of Staphylococcus during reproduction. In Proceedings of the 10th International Symposium on Advanced Optical Manufacturing and Testing Technologies: Intelligent Sensing Technologies and Applications, Chengdu, China, 20–24 June 2021; Volume 12075, SPIE. pp. 124–133. [Google Scholar]
- Lin, W.C. Development and Applications of Near-and Mid-infrared OCT. Ph.D. Thesis, University of Sheffield, Sheffield, UK, 2021. [Google Scholar]
- Qamar, A.; Kerdi, S.; Amin, N.; Zhang, X.; Vrouwenvelder, J.; Ghaffour, N. A deep neural networks framework for in-situ biofilm thickness detection and hydrodynamics tracing for filtration systems. Sep. Purif. Technol. 2022, 301, 121959. [Google Scholar] [CrossRef]
- Lima, C.; Ahmed, S.; Xu, Y.; Muhamadali, H.; Parry, C.; McGalliard, R.J.; Carrol, E.D.; Goodacre, R. Simultaneous Raman and infrared spectroscopy: A novel combination for studying bacterial infections at the single cell level. Chem. Sci. 2022, 13, 8171–8179. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Bian, Z.; Wang, Y. Biofilm formation and inhibition mediated by bacterial quorum sensing. Appl. Microbiol. Biotechnol. 2022, 106, 6365–6381. [Google Scholar] [CrossRef]
- Syed, A.J.; Anderson, J.C. Applications of bioluminescence in biotechnology and beyond. Chem. Soc. Rev. 2021, 50, 5668–5705. [Google Scholar] [CrossRef]
- Resendiz-Sharpe, A.; Vanhoffelen, E.; Velde, G.V. Bioluminescence Imaging, a Powerful Tool to Assess Fungal Burden in Live Mouse Models of Infection. In Antifungal Immunity: Methods and Protocols; Springer: Berlin/Heidelberg, Germany, 2023; pp. 197–210. [Google Scholar]
- Jiang, T.; Bai, X.; Li, M. Advances in the development of bacterial bioluminescence imaging. Annu. Rev. Anal. Chem. 2024, 17, 265–288. [Google Scholar] [CrossRef]
- Mihaylova, E.M. Imaging of Live Cells by Digital Holographic Microscopy. Photonics 2024, 11, 980. [Google Scholar] [CrossRef]
- Riekeles, M.L.I. Microbial Motility as a Universal Biosignature. Ph.D. Thesis, Technische Universitaet Berlin, Berlin, Germany, 2025. [Google Scholar]
- Buzalewicz, I.; Kaczorowska, A.; Fijałkowski, W.; Pietrowska, A.; Matczuk, A.K.; Podbielska, H.; Wieliczko, A.; Witkiewicz, W.; Jędruchniewicz, N. Quantifying the dynamics of bacterial biofilm formation on the surface of soft contact lens materials using digital holographic tomography to advance biofilm research. Int. J. Mol. Sci. 2024, 25, 2653. [Google Scholar] [CrossRef] [PubMed]
- Parthasarathi, S.; Chaudhury, A.; Swain, A.; Basu, J.K. Microscopy Insights as an Invaluable Tool for Studying Antimicrobial Interactions with Bacterial Membranes. ChemistrySelect 2025, 10, e02345. [Google Scholar] [CrossRef]
- Jeong, D.; Kim, M.J.; Park, Y.; Chung, J.; Kweon, H.-S.; Kang, N.-G.; Hwang, S.J.; Youn, S.H.; Hwang, B.K.; Kim, D. Visualizing extracellular vesicle biogenesis in gram-positive bacteria using super-resolution microscopy. BMC Biol. 2022, 20, 270. [Google Scholar] [CrossRef]
- Reichhardt, C.; Parsek, M.R. Confocal laser scanning microscopy for analysis of Pseudomonas aeruginosa biofilm architecture and matrix localization. Front. Microbiol. 2019, 10, 677. [Google Scholar] [CrossRef]
- Rotsch, C.; Radmacher, M. Drug-induced changes of cytoskeletal structure and mechanics in fibroblasts: An atomic force microscopy study. Biophys. J. 2000, 78, 520–535. [Google Scholar] [CrossRef]
- Liedtke, J.; Depelteau, J.S.; Briegel, A. How advances in cryo-electron tomography have contributed to our current view of bacterial cell biology. J. Struct. Biol. X 2022, 6, 100065. [Google Scholar] [CrossRef] [PubMed]
- Kore, S.M. Impact of Shear Flow on Marine Biofilms Using Hyperspectral Imaging. Ph.D. Thesis, Clemson University, Clemson, SC, USA, 2025. [Google Scholar]
- Zhang, P.; Feng, B.; Chen, Y.-P.; Dai, Y.-Z.; Guo, J.-S. In situ characterizations for EPS-involved microprocesses in biological wastewater treatment systems. Crit. Rev. Environ. Sci. Technol. 2019, 49, 917–946. [Google Scholar] [CrossRef]
- Pan, J.; Peng, Q.; Zhang, G.; Xie, Q.; Gong, X.; Qian, P.-Y.; Ma, C.; Zhang, G. Antifouling mechanism of natural product-based coatings investigated by digital holographic microscopy. J. Mater. Sci. Technol. 2021, 84, 200–207. [Google Scholar] [CrossRef]
- Bankar, N.; Bankar, M.; Shelke, Y.; Kale, Y.; Khedkar, S. The role of artificial intelligence in biofilm detection. Proc. AIP Conf. Proc. 2024, 3188, 080038. [Google Scholar]
- Sahana, M. AI-Driven Discovery of Nanostructures That Disrupt Antibiotic-Resistant Biofilms. Master’s Thesis, University of Mysore, Mysore, Indian, 2025. [Google Scholar]
- Chen, H.; Xia, A.; Yan, H.; Huang, Y.; Zhu, X.; Zhu, X.; Liao, Q. Mass transfer in heterogeneous biofilms: Key issues in biofilm reactors and AI-driven performance prediction. Environ. Sci. Ecotechnol. 2024, 22, 100480. [Google Scholar] [CrossRef]
- Zou, H.; Sopasakis, A.; Maillard, F.; Karlsson, E.; Duljas, J.; Silwer, S.; Ohlsson, P.; Hammer, E.C. Bacterial community characterization by deep learning aided image analysis in soil chips. Ecol. Inform. 2024, 81, 102562. [Google Scholar] [CrossRef]
- Liao, Y.; Zhao, J.; Bian, J.; Zhang, Z.; Xu, S.; Qin, Y.; Miao, S.; Li, R.; Liu, R.; Zhang, M. From mechanism to application: Decrypting light-regulated denitrifying microbiome through geometric deep learning. Imeta 2024, 3, e162. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Tabassum, N.; Toma, T.T.; Wang, Y.; Gahlmann, A.; Acton, S.T. 3D GAN image synthesis and dataset quality assessment for bacterial biofilm. Bioinformatics 2022, 38, 4598–4604. [Google Scholar] [CrossRef]
- Holicheva, A.A.; Kozlov, K.S.; Boiko, D.A.; Kamanin, M.S.; Provotorova, D.V.; Kolomoets, N.I.; Ananikov, V.P. Deep generative modeling of annotated bacterial biofilm images. NPJ Biofilms Microbiomes 2025, 11, 16. [Google Scholar] [CrossRef]
- Huang, Y. Machine Learning for Actionable Knowledge Discovery in Synthetic Biology. Doctoral Dissertation, Newcastle University, Newcastle upon Tyne, UK, 2023. [Google Scholar]
- Samue, F. Machine Learning Approaches to Anomaly Detection in Biomedical Data and Medical Imaging. Available online: https://www.researchgate.net/publication/385078064_Machine_Learning_Approaches_to_Anomaly_Detection_in_Biomedical_Data_and_Medical_Imaging?enrichId=rgreq-a8bbaa738c8ee17c0e6c5b61c3ab4afc-XXX&enrichSource=Y292ZXJQYWdlOzM4NTA3ODA2NDtBUzoxMTQzMTI4MTI4NDcyMj (accessed on 18 August 2025).
- Shojaee, A. Bacteria Growth Modeling Using Long-Short-Term-Memory Networks. Master’s Thesis, University of Cincinnati, Cincinnati, OH, USA, 2021. [Google Scholar]
- Franco, N.R. Machine Learning for Precision Medicine: A Combination of Data-Driven and Physics Based Models. 2022. Available online: https://hdl.handle.net/10589/196604 (accessed on 18 August 2025).
- Millevoi, C. Physics-Informed Neural Network in Porous Media and Epidemiological Applications. Ph.D. Thesis, Università degli Studi di Padova, Padova, Italy, 2024. [Google Scholar]
- Guzmán-Soto, I.; McTiernan, C.; Gonzalez-Gomez, M.; Ross, A.; Gupta, K.; Suuronen, E.J.; Mah, T.-F.; Griffith, M.; Alarcon, E.I. Mimicking biofilm formation and development: Recent progress in in vitro and in vivo biofilm models. Iscience 2021, 24, 102443. [Google Scholar] [CrossRef]
- Highmore, C.; Melaugh, G.; Morris, R.; Parker, J.; Direito, S.; Romero, M.; Soukarieh, F.; Robertson, S.; Bamford, N. Translational challenges and opportunities in biofilm science: A BRIEF for the future. NPJ Biofilms Microbiomes 2022, 8, 68. [Google Scholar] [CrossRef]
- Jandl, B.; Dighe, S.; Gasche, C.; Makristathis, A.; Muttenthaler, M. Intestinal biofilms: Pathophysiological relevance, host defense, and therapeutic opportunities. Clin. Microbiol. Rev. 2024, 37, e0013323. [Google Scholar] [CrossRef]
- Sahoo, K.; Meshram, S.; Sahoo Jr, K. Biofilm Formation in Chronic Infections: A Comprehensive Review of Pathogenesis, Clinical Implications, and Novel Therapeutic Approaches. Cureus 2024, 16. [Google Scholar] [CrossRef]
- Bhattacharjee, B.; Ghosh, S.; Patra, D.; Haldar, J. Advancements in release-active antimicrobial biomaterials: A journey from release to relief. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 2022, 14, e1745. [Google Scholar] [CrossRef]
- Billings, C.J. Implantable Medical Devices for Local Drug Delivery and Tissue Regeneration to Combat Chronic Bacterial Infection. Doctoral Dissertation, University of Tennessee College of Veterinary, Knoxville, TN, USA, 2022. [Google Scholar]
- Mannering, N.; Narulla, R.; Lenane, B. In-Vitro and In-Vivo Models for the Study of Prosthetic Joint Infections. In Infection in Knee Replacement; Springer: Berlin/Heidelberg, Germany, 2021; pp. 41–47. [Google Scholar]
- van Agtmaal, J.L.; van Hoogstraten, S.W.; Arts, J.J. Prosthetic Joint Infection Research Models in NZW Rabbits: Opportunities for Standardization—A Systematic Review. J. Funct. Biomater. 2024, 15, 307. [Google Scholar] [CrossRef] [PubMed]
- Chen, V.; Burgess, J.L.; Verpile, R.; Tomic-Canic, M.; Pastar, I. Novel diagnostic technologies and therapeutic approaches targeting chronic wound biofilms and microbiota. Curr. Dermatol. Rep. 2022, 11, 60–72. [Google Scholar] [CrossRef]
- Saeed, S.; Martins-Green, M. Assessing animal models to study impaired and chronic wounds. Int. J. Mol. Sci. 2024, 25, 3837. [Google Scholar] [CrossRef]
- Miron, A.; Giurcaneanu, C.; Mihai, M.M.; Beiu, C.; Voiculescu, V.M.; Popescu, M.N.; Soare, E.; Popa, L.G. Antimicrobial biomaterials for chronic wound care. Pharmaceutics 2023, 15, 1606. [Google Scholar] [CrossRef] [PubMed]
- Reyne, N.; McCarron, A.; Cmielewski, P.; Parsons, D.; Donnelley, M. To bead or not to bead: A review of Pseudomonas aeruginosa lung infection models for cystic fibrosis. Front. Physiol. 2023, 14, 1104856. [Google Scholar] [CrossRef]
- Werneburg, G.T. Catheter-associated urinary tract infections: Current challenges and future prospects. Res. Rep. Urol. 2022, 14, 109–133. [Google Scholar] [CrossRef] [PubMed]
- Nissanka, M.C.; Dilhari, A.; Wijesinghe, G.K.; Weerasekera, M.M. Advances in experimental bladder models: Bridging the gap between in vitro and in vivo approaches for investigating urinary tract infections. BMC Urol. 2024, 24, 206. [Google Scholar] [CrossRef]
- Jakubovics, N.S.; Goodman, S.D.; Mashburn-Warren, L.; Stafford, G.P.; Cieplik, F. The dental plaque biofilm matrix. Periodontol. 2000 2021, 86, 32–56. [Google Scholar] [CrossRef]
- Haque, M.M.; Yerex, K.; Kelekis-Cholakis, A.; Duan, K. Advances in novel therapeutic approaches for periodontal diseases. BMC Oral Health 2022, 22, 492. [Google Scholar] [CrossRef]
- Barnes, A.M.; Frank, K.L.; Dale, J.L.; Manias, D.A.; Powers, J.L.; Dunny, G.M. Enterococcus faecalis colonizes and forms persistent biofilm microcolonies on undamaged endothelial surfaces in a rabbit endovascular infection model. FEMS Microbes 2021, 2, xtab014. [Google Scholar] [CrossRef]
- Schwarz, C.; Töre, Y.; Hoesker, V.; Ameling, S.; Grün, K.; Völker, U.; Schulze, P.C.; Franz, M.; Faber, C.; Schaumburg, F. Host-pathogen interactions of clinical S. aureus isolates to induce infective endocarditis. Virulence 2021, 12, 2073–2087. [Google Scholar] [CrossRef]
- Motta, J.-P.; Wallace, J.L.; Buret, A.G.; Deraison, C.; Vergnolle, N. Gastrointestinal biofilms in health and disease. Nat. Rev. Gastroenterol. Hepatol. 2021, 18, 314–334. [Google Scholar] [CrossRef]
- Duff, A.F.; Jurcisek, J.A.; Kurbatfinski, N.; Chiang, T.; Goodman, S.D.; Bakaletz, L.O.; Bailey, M.T. Oral and middle ear delivery of otitis media standard of care antibiotics, but not biofilm-targeted antibodies, alter chinchilla nasopharyngeal and fecal microbiomes. Npj Biofilms Microbiomes 2024, 10, 10. [Google Scholar] [CrossRef] [PubMed]
- Empitu, M.A.; Kadariswantiningsih, I.N.; Shakri, N.M. Pharmacological strategies for targeting biofilms in otorhinolaryngologic infections and overcoming antimicrobial resistance. Biomed. Rep. 2025, 22, 95. [Google Scholar] [CrossRef] [PubMed]
- Gill, R.P.K.; Gantchev, J.; Martínez Villarreal, A.; Ramchatesingh, B.; Netchiporouk, E.; Akilov, O.E.; Ødum, N.; Gniadecki, R.; Koralov, S.B.; Litvinov, I.V. Understanding cell lines, patient-derived xenograft and genetically engineered mouse models used to study cutaneous T-cell lymphoma. Cells 2022, 11, 593. [Google Scholar] [CrossRef] [PubMed]
- Lloyd, M.G.; Moffat, J.F. Humanized severe combined immunodeficient (SCID) mouse models for varicella-zoster virus pathogenesis. In Varicella-zoster Virus: Genetics, Pathogenesis and Immunity; Springer: Berlin/Heidelberg, Germany, 2022; pp. 135–161. [Google Scholar]
- Mohamed, H.S.; Houmed Aboubaker, M.; Dumont, Y.; Didelot, M.-N.; Michon, A.-L.; Galal, L.; Jean-Pierre, H.; Godreuil, S. Multidrug-Resistant Enterobacterales in Community-Acquired Urinary Tract Infections in Djibouti, Republic of Djibouti. Antibiotics 2022, 11, 1740. [Google Scholar] [CrossRef]
- Meneely, P.M.; Dahlberg, C.L.; Rose, J.K. Working with Worms: Caenorhabditis elegans as a Model Organism. Curr. Protoc. Essent. Lab. Tech. 2019, 19, e35. [Google Scholar] [CrossRef]
- Atkinson, S.; Goldstone, R.J.; Joshua, G.W.P.; Chang, C.Y.; Patrick, H.L.; Cámara, M.; Wren, B.W.; Williams, P. Biofilm Development on by Is Facilitated by Quorum Sensing-Dependent Repression of Type III Secretion. PLoS Pathog. 2011, 7, e1001250. (In English) [Google Scholar] [CrossRef]
- Javanmard, Z.; Pourhajibagher, M.; Bahador, A. Advancing anti-biofilm strategies: Innovations to combat biofilm-related challenges and enhance efficacy. J. Basic Microbiol. 2024, 64, e2400271. [Google Scholar] [CrossRef]
- Singh, R. Revolutionizing Antimicrobial Therapies Through Biofilm-Targeted Nanomedicine. Curr. Pharm. Res. 2025, 1, 78–97. [Google Scholar] [CrossRef]
- Martínez-García, E.; de Lorenzo, V. Pseudomonas putida as a synthetic biology chassis and a metabolic engineering platform. Curr. Opin. Biotechnol. 2024, 85, 103025. [Google Scholar] [CrossRef]
- Arora, A. Crispri-mediated gene silencing in biofilm cycle and quorum sensing. In Gene Editing in Plants: Crispr-Cas and Its Applications; Springer: Berlin/Heidelberg, Germany, 2024; pp. 139–178. [Google Scholar]
- Zhang, Y.; Zhang, T.; Xiao, X.; Kawalek, A.; Ou, J.; Ren, A.; Sun, W.; de Bakker, V.; Liu, Y.; Li, Y. Genome-wide CRISPRi-seq identified ferredoxin-NADP reductase FprB as a synergistic target for gallium therapy in Pseudomonas aeruginosa. bioRxiv 2024. [Google Scholar]
- Zuberi, A.; Ahmad, N.; Ahmad, H.; Saeed, M.; Ahmad, I. Beyond antibiotics: CRISPR/Cas9 triumph over biofilm-associated antibiotic resistance infections. Front. Cell. Infect. Microbiol. 2024, 14, 1408569. [Google Scholar] [CrossRef] [PubMed]
- Ghaznavi, G.; Vosough, P.; Ghasemian, A.; Tabar, M.M.M.; Tayebi, L.; Taghizadeh, S.; Savardashtaki, A. Engineering bacteriophages for targeted superbug eradication. Mol. Biol. Rep. 2025, 52, 221. [Google Scholar] [CrossRef]
- Chen, Y.C.; Destouches, L.; Cook, A.; Fedorec, A.J. Synthetic microbial ecology: Engineering habitats for modular consortia. J. Appl. Microbiol. 2024, 135, lxae158. [Google Scholar] [CrossRef]
- Kozaeva, E.; Eida, A.A.; Gunady, E.F.; Dangl, J.L.; Conway, J.M.; Brophy, J.A. Roots of synthetic ecology: Microbes that foster plant resilience in the changing climate. Curr. Opin. Biotechnol. 2024, 88, 103172. [Google Scholar] [CrossRef]
- Xu, B.; Su, Q.; Yang, Y.; Huang, S.; Yang, Y.; Shi, X.; Choo, K.-H.; Ng, H.Y.; Lee, C.-H. Quorum quenching in membrane bioreactors for fouling retardation: Complexity provides opportunities. Environ. Sci. Technol. 2024, 58, 13171–13193. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, S.; Ray, R.R. Nanomaterials, Phytocompounds. In Biofilm Associated Livestock Diseases and their Management; Springer Nature: Berlin/Heidelberg, Germany, 2025; p. 233. [Google Scholar]
- Stojanov, S.; Berlec, A. Smart bionanomaterials for treatment and diagnosis of inflammatory bowel disease. Nanotechnol. Rev. 2024, 13, 20240057. [Google Scholar] [CrossRef]
- Chang, X.; Liu, X.; Wang, X.; Ma, L.; Liang, J.; Li, Y. Recent Advances in Spatiotemporal Manipulation of Engineered Bacteria for Precision Cancer Therapy. Int. J. Nanomed. 2025, 20, 5859–5872. [Google Scholar] [CrossRef]
- Elazzazy, A.M.; Baeshen, M.N.; Alasmi, K.M.; Alqurashi, S.I.; Desouky, S.E.; Khattab, S.M. Where Biology Meets Engineering: Scaling up microbial nutraceuticals to bridge nutrition, therapeutics, and global impact. Microorganisms 2025, 13, 566. [Google Scholar] [CrossRef]
- Dash, A.; Ragavendran, C. Innovative Approaches to Combating Dental Biofilms: Nanotechnology and its Applications. Biocatal. Agric. Biotechnol. 2024, 61, 103406. [Google Scholar] [CrossRef]
- Waheed, S.; Li, Z.; Zhang, F.; Chiarini, A.; Armato, U.; Wu, J. Engineering nano-drug biointerface to overcome biological barriers toward precision drug delivery. J. Nanobiotechnol. 2022, 20, 395. [Google Scholar] [CrossRef]
- Mohanta, Y.K.; Chakrabartty, I.; Mishra, A.K.; Chopra, H.; Mahanta, S.; Avula, S.K.; Patowary, K.; Ahmed, R.; Mishra, B.; Mohanta, T.K. Nanotechnology in combating biofilm: A smart and promising therapeutic strategy. Front. Microbiol. 2023, 13, 1028086. [Google Scholar] [CrossRef]
- Bharti, S.; Kumar, A. Nanotechnology in Targeted Delivery of Antimicrobials and Overcoming Resistance. BioNanoScience 2025, 15, 1–22. [Google Scholar] [CrossRef]
- Arenas-Vivo, A.; Amariei, G.; Aguado, S.; Rosal, R.; Horcajada, P. An Ag-loaded photoactive nano-metal organic framework as a promising biofilm treatment. Acta Biomater. 2019, 97, 490–500. [Google Scholar] [CrossRef] [PubMed]
- Arenas-Vivo, A.; Celis Arias, V.; Amariei, G.; Rosal, R.; Izquierdo-Barba, I.; Hidalgo, T.; Vallet-Regí, M.; Beltrán, H.I.; Loera-Serna, S.; Horcajada, P. Antiadherent AgBDC metal–organic framework coating for escherichia coli biofilm inhibition. Pharmaceutics 2023, 15, 301. [Google Scholar] [CrossRef]
- Sadeghi Mohammadi, S.; Vaezi, Z.; Naderi-Manesh, H. Improvement of anti-biofilm activities via co-delivery of curcumin and gentamicin in lipid-polymer hybrid nanoparticle. J. Biomater. Sci. Polym. Ed. 2022, 33, 174–196. [Google Scholar] [CrossRef] [PubMed]
- Ahsan, A.; Thomas, N.; Barnes, T.J.; Subramaniam, S.; Loh, T.C.; Joyce, P.; Prestidge, C.A. Lipid nanocarriers-enabled delivery of antibiotics and antimicrobial adjuvants to overcome bacterial biofilms. Pharmaceutics 2024, 16, 396. [Google Scholar] [CrossRef]
- Rubio-Canalejas, A.; Baelo, A.; Herbera, S.; Blanco-Cabra, N.; Vukomanovic, M.; Torrents, E. 3D spatial organization and improved antibiotic treatment of a Pseudomonas aeruginosa–Staphylococcus aureus wound biofilm by nanoparticle enzyme delivery. Front. Microbiol. 2022, 13, 959156. [Google Scholar] [CrossRef]
- Das, D.; Lin, C.-W.; Chuang, H.-S. LAMP-based point-of-care biosensors for rapid pathogen detection. Biosensors 2022, 12, 1068. [Google Scholar] [CrossRef]
- Kameswaran, S.; Gujjala, S.; Zhang, S.; Kondeti, S.; Mahalingam, S.; Bangeppagari, M.; Bellemkonda, R. Quenching and quorum sensing in bacterial bio-films. Res. Microbiol. 2024, 175, 104085. [Google Scholar] [CrossRef] [PubMed]
- Almatroudi, A. Biofilm resilience: Molecular mechanisms driving antibiotic resistance in clinical contexts. Biology 2025, 14, 165. [Google Scholar] [CrossRef]
- Polinarski, M.A.; Beal, A.L.; Silva, F.E.; Bernardi-Wenzel, J.; Burin, G.R.; de Muniz, G.I.; Alves, H.J. New perspectives of using chitosan, silver, and chitosan–silver nanoparticles against multidrug-resistant bacteria. Part. Part. Syst. Charact. 2021, 38, 2100009. [Google Scholar] [CrossRef]
- Laib, I.; Mohammed, H.A.; Laouini, S.E.; Bouafia, A.; Abdullah, M.M.; Al-Lohedan, H.A.; Al-Essa, Q.; Trzepieciński, T. Cutting-edge Nano-therapeutics: Silver nanoparticles loaded with ciprofloxacin for powerful antidiabetic, antioxidant, anti-inflammatory, and antibiotic action against resistant pathogenic bacteria. Int. J. Food Sci. Technol. 2025, 60, vvaf024. [Google Scholar] [CrossRef]
- Bhattacharya, P.; Sengupta, I.; Dey, A.; Chakraborty, S.; Neogi, S. Antibacterial effect of ciprofloxacin loaded reduced graphene oxide nanosheets against Pseudomonas aeruginosa strain. Colloid Interface Sci. Commun. 2021, 40, 100344. [Google Scholar] [CrossRef]
- Yun, J.; Kang, J.; Sharipov, M.; Ryu, W. Recent progress in hydrogel microneedle sensors based on electrochemical and optical sensing. Appl. Spectrosc. Rev. 2025, 1–41. [Google Scholar] [CrossRef]
- Li, B.; Mao, J.; Wu, J.; Mao, K.; Jia, Y.; Chen, F.; Liu, J. Nano–Bio Interactions: Biofilm-Targeted Antibacterial Nanomaterials. Small 2024, 20, 2306135. [Google Scholar] [CrossRef]
- Ma, S.; Moser, D.; Han, F.; Leonhard, M.; Schneider-Stickler, B.; Tan, Y. Preparation and antibiofilm studies of curcumin loaded chitosan nanoparticles against polymicrobial biofilms of Candida albicans and Staphylococcus aureus. Carbohydr. Polym. 2020, 241, 116254. [Google Scholar] [CrossRef]
- Kłodzińska, S.N.; Wan, F.; Jumaa, H.; Sternberg, C.; Rades, T.; Nielsen, H.M. Utilizing nanoparticles for improving anti-biofilm effects of azithromycin: A head-to-head comparison of modified hyaluronic acid nanogels and coated poly (lactic-co-glycolic acid) nanoparticles. J. Colloid Interface Sci. 2019, 555, 595–606. [Google Scholar] [CrossRef]
- Deacon, J.; Abdelghany, S.M.; Quinn, D.J.; Schmid, D.; Megaw, J.; Donnelly, R.F.; Jones, D.S.; Kissenpfennig, A.; Elborn, J.S.; Gilmore, B.F. Antimicrobial efficacy of tobramycin polymeric nanoparticles for Pseudomonas aeruginosa infections in cystic fibrosis: Formulation, characterisation and functionalisation with dornase alfa (DNase). J. Control. Release 2015, 198, 55–61. [Google Scholar] [CrossRef]
- Wang, C.; Zhao, W.; Cao, B.; Wang, Z.; Zhou, Q.; Lu, S.; Lu, L.; Zhan, M.; Hu, X. Biofilm-responsive polymeric nanoparticles with self-adaptive deep penetration for in vivo photothermal treatment of implant infection. Chem. Mater. 2020, 32, 7725–7738. [Google Scholar] [CrossRef]
- Ferreira, M.; Pinto, S.N.; Aires-da-Silva, F.; Bettencourt, A.; Aguiar, S.I.; Gaspar, M.M. Liposomes as a Nanoplatform to Improve the Delivery of Antibiotics into Staphylococcus aureus Biofilms. Pharmaceutics 2021, 13, 321. [Google Scholar] [CrossRef] [PubMed]
- Alfei, S.; Caviglia, D. Prevention and eradication of biofilm by dendrimers: A possibility still little explored. Pharmaceutics 2022, 14, 2016. [Google Scholar] [CrossRef] [PubMed]
- Osiro, K.O.; Hashemi, N.; Brango-Vanegas, J.; Oliveira, S.M.; Franco, O.L. Emerging peptide-based technology for biofilm control. Expert Opin. Biol. Ther. 2024, 24, 1311–1315. [Google Scholar] [CrossRef]
- de Lacerda Coriolano, D.; de Souza, J.B.; Bueno, E.V.; Medeiros, S.M.d.F.R.d.S.; Cavalcanti, I.D.L.; Cavalcanti, I.M.F. Antibacterial and antibiofilm potential of silver nanoparticles against antibiotic-sensitive and multidrug-resistant Pseudomonas aeruginosa strains. Braz. J. Microbiol. 2021, 52, 267–278. [Google Scholar] [CrossRef]
- Neihaya, H.; Zaman, H. Investigating the effect of biosynthesized silver nanoparticles as antibiofilm on bacterial clinical isolates. Microb. Pathog. 2018, 116, 200–208. [Google Scholar] [CrossRef]
- Hamida, R.S.; Ali, M.A.; Goda, D.A.; Khalil, M.I.; Redhwan, A. Cytotoxic effect of green silver nanoparticles against ampicillin-resistant Klebsiella pneumoniae. RSC Adv. 2020, 10, 21136–21146. [Google Scholar] [CrossRef]
- Žalnėravičius, R.; Mikalauskaitė, A.; Niaura, G.; Paškevičius, A.; Jagminas, A. Ultra-small methionine-capped Au0/Au+ nanoparticles as efficient drug against the antibiotic-resistant bacteria. Mater. Sci. Eng. C 2019, 102, 646–652. [Google Scholar] [CrossRef]
- Chen, J.; Dai, T.; Yu, J.; Dai, X.; Chen, R.; Wu, J.; Li, N.; Fan, L.; Mao, Z.; Sheng, G. Integration of antimicrobial peptides and gold nanorods for bimodal antibacterial applications. Biomater. Sci. 2020, 8, 4447–4457. [Google Scholar] [CrossRef]
- Dogra, V.; Kaur, G.; Jindal, S.; Kumar, R.; Kumar, S.; Singhal, N.K. Bactericidal effects of metallosurfactants based cobalt oxide/hydroxide nanoparticles against Staphylococcus aureus. Sci. Total Environ. 2019, 681, 350–364. [Google Scholar] [CrossRef]
- Safaei, M.; Taran, M.; Jamshidy, L.; Imani, M.M.; Mozaffari, H.R.; Sharifi, R.; Golshah, A.; Moradpoor, H. Optimum synthesis of polyhydroxybutyrate-Co3O4 bionanocomposite with the highest antibacterial activity against multidrug resistant bacteria. Int. J. Biol. Macromol. 2020, 158, 477–485. [Google Scholar] [CrossRef] [PubMed]
- Gabrielyan, L.; Hakobyan, L.; Hovhannisyan, A.; Trchounian, A. Effects of iron oxide (Fe3O4) nanoparticles on Escherichia coli antibiotic-resistant strains. J. Appl. Microbiol. 2019, 126, 1108–1116. [Google Scholar] [CrossRef] [PubMed]
- Arias, L.S.; Pessan, J.P.; de Souza Neto, F.N.; Lima, B.H.R.; de Camargo, E.R.; Ramage, G.; Delbem, A.C.B.; Monteiro, D.R. Novel nanocarrier of miconazole based on chitosan-coated iron oxide nanoparticles as a nanotherapy to fight Candida biofilms. Colloids Surf. B Biointerfaces 2020, 192, 111080. [Google Scholar] [CrossRef] [PubMed]
- Muzammil, S.; Khurshid, M.; Nawaz, I.; Siddique, M.H.; Zubair, M.; Nisar, M.A.; Imran, M.; Hayat, S. Aluminium oxide nanoparticles inhibit EPS production, adhesion and biofilm formation by multidrug resistant Acinetobacter baumannii. Biofouling 2020, 36, 492–504. [Google Scholar] [CrossRef]
- Palau Gauthier, M.; Muñoz, E.; Gustà, M.F.; Larrosa, M.N.; Gomis, X.; Gilabert, J.; Almirante Gragera, B.; Puntes, V.; Texidó, R.; Gavaldà, J. In Vitro Antibacterial Activity of Silver Nanoparticles Conjugated with Amikacin and Combined with Hyperthermia against Drug-Resistant and Biofilm-Producing Strains. Microbiol. Spectr. 2023, 11, e0028023. [Google Scholar]
- Thorn, C.R.; Kopecki, Z.; Wignall, A.; Kral, A.; Prestidge, C.A.; Thomas, N. Liquid crystal nanoparticle platform for increased efficacy of cationic antimicrobials against biofilm infections. Nanomed. Nanotechnol. Biol. Med. 2022, 42, 102536. [Google Scholar] [CrossRef]
- Wu, X.; Qi, M.; Liu, C.; Yang, Q.; Li, S.; Shi, F.; Sun, X.; Wang, L.; Li, C.; Dong, B. Near-infrared light-triggered nitric oxide nanocomposites for photodynamic/photothermal complementary therapy against periodontal biofilm in an animal model. Theranostics 2023, 13, 2350. [Google Scholar] [CrossRef]
- Kim, M.-H. Nanoparticle-based therapies for wound biofilm infection: Opportunities and challenges. IEEE Trans. Nanobiosci. 2016, 15, 294–304. [Google Scholar] [CrossRef]
- Elabbasy, M.T.; El Bayomi, R.M.; Abdelkarim, E.A.; Hafez, A.E.-S.E.; Othman, M.S.; Ghoniem, M.E.; Samak, M.A.; Alshammari, M.H.; Almarshadi, F.A.; Elsamahy, T. Antibacterial and Antibiofilm Activity of Green-Synthesized Zinc Oxide Nanoparticles Against Multidrug-Resistant Escherichia coli Isolated from Retail Fish. Molecules 2025, 30, 768. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Zeng, Y.; Liu, J.; Wang, L.; Yang, M.; Zhou, J. Antimicrobial and anti-biofilm effects of dihydroartemisinin-loaded chitosan nanoparticles against methicillin-resistant Staphylococcus aureus. Microb. Pathog. 2025, 199, 107208. [Google Scholar] [CrossRef]
- Rostamnejad, D.; Esnaashari, F.; Zahmatkesh, H.; Rasti, B.; Zamani, H. Diclofenac-loaded PLGA nanoparticles downregulate LasI/R quorum sensing genes in pathogenic P. aeruginosa isolates. Arch. Microbiol. 2024, 206, 112. [Google Scholar] [CrossRef]
- Liu, X.; Xu, S.; Chen, X.; Kang, S.; Liu, J.; Jiang, Q.; Zhang, S.; Hao, L.; Ran, H.; Huang, W. Synergistic effect of bovine cateslytin-loaded nanoparticles combined with ultrasound against Candida albicans biofilm. Future Med. Chem. 2023, 15, 43–55. [Google Scholar] [CrossRef] [PubMed]
- Ali, A.; Ovais, M.; Cui, X.; Rui, Y.; Chen, C. Safety assessment of nanomaterials for antimicrobial applications. Chem. Res. Toxicol. 2020, 33, 1082–1109. [Google Scholar] [CrossRef]
- Bereanu, A.-S.; Vintilă, B.I.; Bereanu, R.; Codru, I.R.; Hașegan, A.; Olteanu, C.; Săceleanu, V.; Sava, M. TiO2 Nanocomposite Coatings and Inactivation of Carbapenemase-Producing Klebsiella Pneumoniae Biofilm—Opportunities and Challenges. Microorganisms 2024, 12, 684. [Google Scholar] [CrossRef]
- Maksimova, Y.; Pyankova, E.; Nesterova, L.; Maksimov, A. Effects of Graphene Derivatives and Near-Infrared Laser Irradiation on E. coli Biofilms and Stress Response Gene Expression. Int. J. Mol. Sci. 2025, 26, 4728. [Google Scholar] [CrossRef]
- Reymond, J.-L.; Bergmann, M.; Darbre, T. Glycopeptide dendrimers as Pseudomonas aeruginosa biofilm inhibitors. Chem. Soc. Rev. 2013, 42, 4814–4822. [Google Scholar] [CrossRef]
- Backlund, C.J.; Worley, B.V.; Schoenfisch, M.H. Anti-biofilm action of nitric oxide-releasing alkyl-modified poly (amidoamine) dendrimers against Streptococcus mutans. Acta Biomater. 2016, 29, 198–205. [Google Scholar] [CrossRef]
- Nabawy, A.; Makabenta, J.M.; Schmidt-Malan, S.; Park, J.; Li, C.-H.; Huang, R.; Fedeli, S.; Chattopadhyay, A.N.; Patel, R.; Rotello, V.M. Dual antimicrobial-loaded biodegradable nanoemulsions for synergistic treatment of wound biofilms. J. Control. Release 2022, 347, 379–388. [Google Scholar] [CrossRef] [PubMed]
- Li, C.-H.; Landis, R.F.; Makabenta, J.M.; Nabawy, A.; Tronchet, T.; Archambault, D.; Liu, Y.; Huang, R.; Golan, M.; Cui, W. Nanotherapeutics using all-natural materials. Effective treatment of wound biofilm infections using crosslinked nanoemulsions. Mater. Horiz. 2021, 8, 1776–1782. [Google Scholar] [CrossRef]
- Hu, W.; Ouyang, Q.; Jiang, C.; Huang, S.; Alireza, N.-E.; Guo, D.; Liu, J.; Peng, Y. Biomedical Metal–Organic framework materials on antimicrobial therapy: Perspectives and challenges. Mater. Today Chem. 2024, 41, 102300. [Google Scholar] [CrossRef]
- Dastneshan, A.; Rahiminezhad, S.; Mezajin, M.N.; Jevinani, H.N.; Akbarzadeh, I.; Abdihaji, M.; Qahremani, R.; Jahanbakhshi, M.; Lalami, Z.A.; Heydari, H. Cefazolin encapsulated UIO-66-NH2 nanoparticles enhance the antibacterial activity and biofilm inhibition against drug-resistant S. aureus: In vitro and in vivo studies. Chem. Eng. J. 2023, 455, 140544. [Google Scholar] [CrossRef]
- Hamza, A. Sensing Platform based on Metal Organic Framework for the Quantitative Determination of Antibiotic Remains in Animal Derived Food. Chemistry Department COMSATS university Islamabad Lahore Campus. Doctoral Dissertation, COMSATS University Islamabad, Islamabad, Pakistan, 2024. [Google Scholar]
- Kumar, P.; Behera, A.; Tiwari, P.; Karthik, S.; Biswas, M.; Sonawane, A.; Mobin, S.M. Exploring the antimicrobial potential of isoniazid loaded Cu-based metal–organic frameworks as a novel strategy for effective killing of Mycobacterium tuberculosis. J. Mater. Chem. B 2023, 11, 10929–10940. [Google Scholar] [CrossRef] [PubMed]
- Gurunathan, S.; Thangaraj, P.; Das, J.; Kim, J.-H. Antibacterial and antibiofilm effects of Pseudomonas aeruginosa derived outer membrane vesicles against Streptococcus mutans. Heliyon 2023, 9, e22606. [Google Scholar] [CrossRef]
- Martins, P.; Machado, D.; Theizen, T.H.; Guarnieri, J.P.O.; Bernardes, B.G.; Gomide, G.P.; Corat, M.A.F.; Abbehausen, C.; Módena, J.L.P.; Melo, C.F.O.R. Outer membrane vesicles from Neisseria meningitidis (Proteossome) used for nanostructured Zika virus vaccine production. Sci. Rep. 2018, 8, 8290. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, A.A.Q.; Besio, R.; Xiao, L.; Forlino, A. Outer membrane vesicles (OMVs) as biomedical tools and their relevance as immune-modulating agents against H. pylori infections: Current status and future prospects. Int. J. Mol. Sci. 2023, 24, 8542. [Google Scholar] [CrossRef]
- Xerri, N.L.; Payne, S.M. Bacteroides thetaiotaomicron outer membrane vesicles modulate virulence of Shigella flexneri. MBio 2022, 13, e02360-22. [Google Scholar] [CrossRef]
- de Freitas, M.C.R.; de Almeida, P.E.; Vieira, W.V.; Ferreira-Machado, A.B.; Resende, J.A.; da Silva, V.L.; Diniz, C.G. Inflammatory modulation and outer membrane vesicles (OMV) production associated to Bacteroides fragilis response to subinhibitory concentrations of metronidazole during experimental infection. Anaerobe 2022, 73, 102504. [Google Scholar] [CrossRef]
- Zhou, Q.; Wang, T.; Li, K.; Zhang, S.; Wang, K.; Hong, W.; Liu, R.; Li, P. Biofilm microenvironment-responsive polymeric CO releasing micelles for enhanced amikacin efficacy. J. Control. Release 2023, 357, 561–571. [Google Scholar] [CrossRef]
- Albayaty, Y.N.; Thomas, N.; Ramirez-Garcia, P.D.; Davis, T.P.; Quinn, J.F.; Whittaker, M.R.; Prestidge, C.A. Polymeric micelles with anti-virulence activity against Candida albicans in a single-and dual-species biofilm. Drug Deliv. Transl. Res. 2021, 11, 1586–1597. [Google Scholar] [CrossRef] [PubMed]
- Quan, K.; Zhang, Z.; Ren, Y.; Busscher, H.J.; Van Der Mei, H.C.; Peterson, B.W. Homogeneous distribution of magnetic, antimicrobial-carrying nanoparticles through an infectious biofilm enhances biofilm-killing efficacy. ACS Biomater. Sci. Eng. 2019, 6, 205–212. [Google Scholar] [CrossRef]
- El-Khawaga, A.M.; Ayman, M.; Hafez, O.; Shalaby, R.E. Photocatalytic, antimicrobial and antibiofilm activities of MgFe2O4 magnetic nanoparticles. Sci. Rep. 2024, 14, 12877. [Google Scholar] [CrossRef] [PubMed]
- Esfahani, M.B.; Khodavandi, A.; Alizadeh, F.; Bahador, N. Antibacterial and anti-biofilm activities of microbial synthesized silver and magnetic iron oxide nanoparticles against Pseudomonas aeruginosa. IEEE Trans. NanoBioscience 2023, 22, 956–966. [Google Scholar] [CrossRef]
- Raulio, M.; Pore, V.; Areva, S.; Ritala, M.; Leskelä, M.; Lindén, M.; Rosenholm, J.B.; Lounatmaa, K.; Salkinoja-Salonen, M. Destruction of Deinococcus geothermalis biofilm by photocatalytic ALD and sol-gel TiO2 surfaces. J. Ind. Microbiol. Biotechnol. 2006, 33, 261–268. [Google Scholar] [CrossRef]
- Tao, L. Synthesis and Characterizations of Antimicrobial Coating by Atomic Layer Deposition Technique for Personal Protective Equipment Applications. Doctoral Dissertation, University of British Columbia, Vancouver, BC, Canada, 2022. [Google Scholar]
- Xin, H.; Liu, Y.; Xiao, Y.; Wen, M.; Sheng, L.; Jia, Z. Design and nanoengineering of photoactive antimicrobials for bioapplications: From fundamentals to advanced strategies. Adv. Funct. Mater. 2024, 34, 2402607. [Google Scholar] [CrossRef]
- Buret, A.G.; Allain, T.; Motta, J.-P.; Wallace, J.L. Effects of hydrogen sulfide on the microbiome: From toxicity to therapy. Antioxid. Redox Signal. 2022, 36, 211–219. [Google Scholar] [CrossRef]
- Sethulekshmi, A.; Saritha, A.; Joseph, K.; Aprem, A.S.; Sisupal, S.B. MoS2 based nanomaterials: Advanced antibacterial agents for future. J. Control. Release 2022, 348, 158–185. [Google Scholar] [CrossRef]
- Tariq, F.; Hussain, R.; Noreen, Z.; Javed, A.; Shah, A.; Mahmood, A.; Sajjad, M.; Bokhari, H.; ur Rahman, S. Enhanced antibacterial activity of visible light activated sulfur-doped TiO2 nanoparticles against Vibrio cholerae. Mater. Sci. Semicond. Process. 2022, 147, 106731. [Google Scholar] [CrossRef]
- Gao, F.; Shao, T.; Yu, Y.; Xiong, Y.; Yang, L. Surface-bound reactive oxygen species generating nanozymes for selective antibacterial action. Nat. Commun. 2021, 12, 745. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Fang, L.; Zhou, H.; Wang, M.; Zheng, H.; Wang, Y.; Weir, M.D.; Masri, R.; Oates, T.W.; Cheng, L. Silica nanoparticles containing nano-silver and chlorhexidine respond to pH to suppress biofilm acids and modulate biofilms toward a non-cariogenic composition. Dent. Mater. 2024, 40, 179–189. [Google Scholar] [CrossRef]
- Ji, H.; Hu, H.; Tang, Q.; Kang, X.; Liu, X.; Zhao, L.; Jing, R.; Wu, M.; Li, G.; Zhou, X. Precisely controlled and deeply penetrated micro-nano hybrid multifunctional motors with enhanced antibacterial activity against refractory biofilm infections. J. Hazard. Mater. 2022, 436, 129210. [Google Scholar] [CrossRef]
- Hemmati, J.; Nazari, M.; Abolhasani, F.S.; Ahmadi, A.; Asghari, B. In vitro investigation of relationship between quorum-sensing system genes, biofilm forming ability, and drug resistance in clinical isolates of Pseudomonas aeruginosa. BMC Microbiol. 2024, 24, 99. [Google Scholar] [CrossRef] [PubMed]
- Neil, B.; Cheney, G.L.; Rosenzweig, J.A.; Sha, J.; Chopra, A.K. Antimicrobial resistance in aeromonads and new therapies targeting quorum sensing. Appl. Microbiol. Biotechnol. 2024, 108, 205. [Google Scholar] [CrossRef]
- Kachhadiya, D.K.; Georrge, J.J. Quorum Sensing and its Inhibition in Pseudomonas aeruginosa: Molecular Targets and Mode of Action. Curr. Drug Targets 2025, 26, 613–631. [Google Scholar] [CrossRef] [PubMed]
- He, Y.-W.; Deng, Y.; Miao, Y.; Chatterjee, S.; Tran, T.M.; Tian, J.; Lindow, S. DSF-family quorum sensing signal-mediated intraspecies, interspecies, and inter-kingdom communication. Trends Microbiol. 2023, 31, 36–50. [Google Scholar] [CrossRef] [PubMed]
- Tuan, D.A.; Uyen, P.V.N.; Masak, J. Hybrid quorum sensing and machine learning systems for adaptive synthetic biology: Toward autonomous gene regulation and precision therapies. Preprint 2024. [Google Scholar] [CrossRef]
- Tu, Y.; Li, H.; Huo, J.; Gou, L.; Wen, X.; Yu, X.; Zhang, X.; Zeng, J.; Li, Y. Disrupting the bacterial language: Quorum quenching and its applications. Crit. Rev. Microbiol. 2025, 1–15. [Google Scholar] [CrossRef]
- Efremenko, E.; Aslanli, A.; Domnin, M.; Stepanov, N.; Senko, O. Enzymes with lactonase activity against fungal quorum molecules as effective antifungals. Biomolecules 2024, 14, 383. [Google Scholar] [CrossRef]
- Bravo, M.; Conchillo-Solé, Ò.; Coves, X.; García-Navarro, A.; Gómez, A.-C.; Márquez-Martínez, M.; Ferrer-Miralles, N.; Daura, X.; Gibert, I.; Yero, D. An acyl-homoserine lactone acylase found in Stenotrophomonas maltophilia exhibits both quorum quenching activity and the ability to degrade penicillin antibiotics. Sci. Rep. 2025, 15, 8557. [Google Scholar] [CrossRef]
- Zhang, C.; Li, C.; Aziz, T.; Alharbi, M.; Cui, H.; Lin, L. Screening of E. coli O157: H7 AI-2 QS inhibitors and their inhibitory effect on biofilm formation in combination with disinfectants. Food Biosci. 2024, 58, 103821. [Google Scholar] [CrossRef]
- Kang, S.; Kim, J.; Hur, J.K.; Lee, S.-S. CRISPR-based genome editing of clinically important Escherichia coli SE15 isolated from indwelling urinary catheters of patients. J. Med. Microbiol. 2017, 66, 18–25. [Google Scholar] [CrossRef]
- Arruda-Silva, F.; Bianchetto-Aguilera, F.; Gasperini, S.; Polletti, S.; Cosentino, E.; Tamassia, N.; Cassatella, M.A. Human neutrophils produce CCL23 in response to various TLR-agonists and TNFα. Front. Cell. Infect. Microbiol. 2017, 7, 176. [Google Scholar] [CrossRef]
- Kumar, L.; Chhibber, S.; Kumar, R.; Kumar, M.; Harjai, K. Zingerone silences quorum sensing and attenuates virulence of Pseudomonas aeruginosa. Fitoterapia 2015, 102, 84–95. [Google Scholar] [CrossRef]
- Gökalsın, B.; Aksoydan, B.; Erman, B.; Sesal, N.C. Reducing virulence and biofilm of Pseudomonas aeruginosa by potential quorum sensing inhibitor carotenoid: Zeaxanthin. Microb. Ecol. 2017, 74, 466–473. [Google Scholar] [CrossRef]
- Baldry, M.; Kitir, B.; Frøkiær, H.; Christensen, S.B.; Taverne, N.; Meijerink, M.; Franzyk, H.; Olsen, C.A.; Wells, J.M.; Ingmer, H. The agr inhibitors solonamide B and analogues alter immune responses to Staphylococccus aureus but do not exhibit adverse effects on immune cell functions. PLoS ONE 2016, 11, e0145618. [Google Scholar] [CrossRef] [PubMed]
- Paczkowski, J.E.; Mukherjee, S.; McCready, A.R.; Cong, J.-P.; Aquino, C.J.; Kim, H.; Henke, B.R.; Smith, C.D.; Bassler, B.L. Flavonoids suppress Pseudomonas aeruginosa virulence through allosteric inhibition of quorum-sensing receptors. J. Biol. Chem. 2017, 292, 4064–4076. [Google Scholar] [CrossRef] [PubMed]
- Yang, T.; Tal-Gan, Y.; Paharik, A.E.; Horswill, A.R.; Blackwell, H.E. Structure–function analyses of a Staphylococcus epidermidis autoinducing peptide reveals motifs critical for AgrC-type receptor modulation. ACS Chem. Biol. 2016, 11, 1982–1991. [Google Scholar] [CrossRef]
- Zhang, Y.; Sass, A.; Van Acker, H.; Wille, J.; Verhasselt, B.; Van Nieuwerburgh, F.; Kaever, V.; Crabbé, A.; Coenye, T. Coumarin reduces virulence and biofilm formation in Pseudomonas aeruginosa by affecting quorum sensing, type III secretion and C-di-GMP levels. Front. Microbiol. 2018, 9, 1952. [Google Scholar] [CrossRef] [PubMed]
- Amara, N.; Gregor, R.; Rayo, J.; Dandela, R.; Daniel, E.; Liubin, N.; Willems, H.M.E.; Ben-Zvi, A.; Krom, B.P.; Meijler, M.M. Fine-Tuning Covalent Inhibition of Bacterial Quorum Sensing. ChemBioChem 2016, 17, 825–835. [Google Scholar] [CrossRef]
- Heidari, A.; Noshiranzadeh, N.; Haghi, F.; Bikas, R. Inhibition of quorum sensing related virulence factors of Pseudomonas aeruginosa by pyridoxal lactohydrazone. Microb. Pathog. 2017, 112, 103–110. [Google Scholar] [CrossRef] [PubMed]
- Abbas, H.A.; Shaldam, M.A. Glyceryl trinitrate is a novel inhibitor of quorum sensing in Pseudomonas aeruginosa. Afr. Health Sci. 2016, 16, 1109–1117. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.; Park, J.-S.; Choi, H.-Y.; Yoon, S.S.; Kim, W.-G. Terrein is an inhibitor of quorum sensing and c-di-GMP in Pseudomonas aeruginosa: A connection between quorum sensing and c-di-GMP. Sci. Rep. 2018, 8, 8617. [Google Scholar] [CrossRef]
- Goh, W.-K.; Gardner, C.R.; Sekhar, K.V.C.; Biswas, N.N.; Nizalapur, S.; Rice, S.A.; Willcox, M.; Black, D.S.; Kumar, N. Synthesis, quorum sensing inhibition and docking studies of 1, 5-dihydropyrrol-2-ones. Bioorganic Med. Chem. 2015, 23, 7366–7377. [Google Scholar] [CrossRef]
- Kalia, M.; Yadav, V.K.; Singh, P.K.; Sharma, D.; Narvi, S.S.; Agarwal, V. Exploring the impact of parthenolide as anti-quorum sensing and anti-biofilm agent against Pseudomonas aeruginosa. Life Sci. 2018, 199, 96–103. [Google Scholar] [CrossRef]
- Scoffone, V.C.; Chiarelli, L.R.; Makarov, V.; Brackman, G.; Israyilova, A.; Azzalin, A.; Forneris, F.; Riabova, O.; Savina, S.; Coenye, T. Discovery of new diketopiperazines inhibiting Burkholderia cenocepacia quorum sensing in vitro and in vivo. Sci. Rep. 2016, 6, 32487. [Google Scholar] [CrossRef]
- Teng, Z.; Shi, D.; Liu, H.; Shen, Z.; Zha, Y.; Li, W.; Deng, X.; Wang, J. Lysionotin attenuates Staphylococcus aureus pathogenicity by inhibiting α-toxin expression. Appl. Microbiol. Biotechnol. 2017, 101, 6697–6703. [Google Scholar] [CrossRef]
- Ren, S.; Wang, B.; Qiu, X.; Wang, S.; Huang, G.; Wang, Y. Unveiling Antibacterial and Antibiofilm Mechanisms of Methyleugenol: Implications for Ecomaterial Functionalization. ACS EST Eng. 2024, 4, 1206–1217. [Google Scholar] [CrossRef]
- Vieira, T.F.; Leitão, M.M.; Cerqueira, N.M.; Sousa, S.F.; Borges, A.; Simões, M. Montelukast and cefoperazone act as antiquorum sensing and antibiofilm agents against Pseudomonas aeruginosa. J. Appl. Microbiol. 2024, 135, lxae088. [Google Scholar] [CrossRef]
- Jakobsen, T.H.; van Gennip, M.; Phipps, R.K.; Shanmugham, M.S.; Christensen, L.D.; Alhede, M.; Skindersoe, M.E.; Rasmussen, T.B.; Friedrich, K.; Uthe, F. Ajoene, a sulfur-rich molecule from garlic, inhibits genes controlled by quorum sensing. Antimicrob. Agents Chemother. 2012, 56, 2314–2325. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.; Wei, Q.; Tong, Q.; Cui, K.; He, G.; Lin, L.; Ma, L.Z.; Cornelis, P.; Wang, Y. Traditional Chinese medicine Tanreqing inhibits quorum sensing systems in Pseudomonas aeruginosa. Front. Microbiol. 2020, 11, 517462. [Google Scholar] [CrossRef] [PubMed]
- Whiteley, M.; Diggle, S.P.; Greenberg, E.P. Bacterial quorum sensing: The progress and promise of an emerging research area. Nature 2017, 551, 313. [Google Scholar] [CrossRef]
- Kai, K. The phc quorum-sensing system in Ralstonia solanacearum species complex. Annu. Rev. Microbiol. 2023, 77, 213–231. [Google Scholar] [CrossRef]
- Yoshihara, A.; Shimatani, M.; Sakata, M.; Takemura, C.; Senuma, W.; Hikichi, Y.; Kai, K. Quorum sensing inhibition attenuates the virulence of the plant pathogen Ralstonia solanacearum species complex. ACS Chem. Biol. 2020, 15, 3050–3059. [Google Scholar] [CrossRef]
- Wang, H.; Liao, L.; Chen, S.; Zhang, L.-H. A quorum quenching bacterial isolate contains multiple substrate-inducible genes conferring degradation of diffusible signal factor. Appl. Environ. Microbiol. 2020, 86, e02930-19. [Google Scholar] [CrossRef]
- Nazzaro, F.; Fratianni, F.; d’Acierno, A.; Coppola, R.; Ayala-Zavala, F.J.; da Cruz, A.G.; De Feo, V. Essential oils and microbial communication. In Essential Oils-Oils of Nature; IntechOpen: Rijeka, Croatia, 2019. [Google Scholar]
- Ribeiro, T.A.N.; Dos Santos, G.A.; Dos Santos, C.T.; Soares, D.C.F.; Saraiva, M.F.; Leal, D.H.S.; Sachs, D. Eugenol as a promising antibiofilm and anti-quorum sensing agent: A systematic review. Microb. Pathog. 2024, 196, 106937. [Google Scholar] [CrossRef]
- Kameswaran, S.; Ramesh, B. Natural QSIs for Biofilm Control in Pathogenic Bacteria. In Quorum Quenching; Elsevier: Amsterdam, The Netherlands, 2023. [Google Scholar]
- Mokoena, M.P.; Omatola, C.A.; Olaniran, A.O. Applications of lactic acid bacteria and their bacteriocins against food spoilage microorganisms and foodborne pathogens. Molecules 2021, 26, 7055. [Google Scholar] [CrossRef]
- Sadik, A.; Viswaswar, J.P.; Rajamoney, A.; Rekha, A.; Raj, D.M.; Prakashan, D.; Vasudevan, M.; Visakh, J.; Renuka, D.; Hely, S. Mitigation of quorum sensing mediated virulence factors of Pseudomonas aeruginosa: The role of Meldrum’s acid activated furan. Front. Microbiol. 2024, 14, 1272240. [Google Scholar] [CrossRef] [PubMed]
- Angelini, P. Plant-derived antimicrobials and their crucial role in combating antimicrobial resistance. Antibiotics 2024, 13, 746. [Google Scholar] [CrossRef]
- Davares, A.K.; Arsene, M.M.; Viktorovna, P.I.; Vyacheslavovna, Y.N.; Vladimirovna, Z.A.; Aleksandrovna, V.E.; Nikolayevich, S.A.; Nadezhda, S.; Anatolievna, G.O.; Nikolaevna, S.I. Quorum-sensing inhibitors from probiotics as a strategy to combat bacterial cell-to-cell communication involved in food spoilage and food safety. Fermentation 2022, 8, 711. [Google Scholar] [CrossRef]
- Antonioli, L.; Haskó, G. May be adenosine an immuno-quorum-sensing signal? Purinergic Signal. 2022, 18, 205–209. [Google Scholar] [CrossRef] [PubMed]
- Deng, Z.; Luo, X.M.; Liu, J.; Wang, H. Quorum sensing, biofilm, and intestinal mucosal barrier: Involvement the role of probiotic. Front. Cell. Infect. Microbiol. 2020, 10, 538077. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Zhang, Z.; Li, J.; Qin, G. New strategies for biocontrol of bacterial toxins and virulence: Focusing on quorum-sensing interference and biofilm inhibition. Toxins 2023, 15, 570. [Google Scholar] [CrossRef]
- Arya, S.; Usha, R. Bioprospecting and Exploration of Phytochemicals as Quorum Sensing Inhibitors against Cariogenic Dental Biofilm. J. Pure Appl. Microbiol. 2024, 18, 100–117. [Google Scholar] [CrossRef]
- Raheem, A.; Liang, L.; Zhang, G.; Cui, S. Modulatory effects of probiotics during pathogenic infections with emphasis on immune regulation. Front. Immunol. 2021, 12, 616713. [Google Scholar] [CrossRef]
- Stephens, K.; Bentley, W.E. Synthetic biology for manipulating quorum sensing in microbial consortia. Trends Microbiol. 2020, 28, 633–643. [Google Scholar] [CrossRef]
- Gunaratnam, S.; Millette, M.; McFarland, L.V.; DuPont, H.L.; Lacroix, M. Potential role of probiotics in reducing Clostridioides difficile virulence: Interference with quorum sensing systems. Microb. Pathog. 2021, 153, 104798. [Google Scholar] [CrossRef]
- Wang, Y.; Dai, J.; Wang, X.; Wang, Y.; Tang, F. Mechanisms of interactions between bacteria and bacteriophage mediate by quorum sensing systems. Appl. Microbiol. Biotechnol. 2022, 106, 2299–2310. [Google Scholar] [CrossRef] [PubMed]
- Huedo, P.; Coves, X.; Daura, X.; Gibert, I.; Yero, D. Quorum sensing signaling and quenching in the multidrug-resistant pathogen Stenotrophomonas maltophilia. Front. Cell. Infect. Microbiol. 2018, 8, 122. [Google Scholar] [CrossRef]
- Wijesinghe, G.; Nobbs, A.; Bandara, H. The diffusible signaling factor family in microbial signaling: A current perspective. Crit. Rev. Microbiol. 2025, 1–16. [Google Scholar] [CrossRef]
- Horgan, C.; Baccari, C.; O’Driscoll, M.; Lindow, S.E.; O’Sullivan, T.P. BDSF Analogues Inhibit Quorum Sensing-Regulated Biofilm Production in Xylella fastidiosa. Microorganisms 2024, 12, 2496. [Google Scholar] [CrossRef]
- Zhang, Y.; Bhasme, P.; Reddy, D.S.; Liu, D.J.; Yu, Z.X.; Zhao, T.H.; Zheng, Y.Q.; Kumar, A.; Yu, H.Y.; Ma, L.Z. Dual functions: A coumarin-chalcone conjugate inhibits cyclic-di-GMP and quorum-sensing signaling to reduce biofilm formation and virulence of pathogens. MLife 2023, 2, 283–294. (In English) [Google Scholar] [CrossRef]
- Smith, P.; Schuster, M. Antiactivators prevent self-sensing in Pseudomonas aeruginosa quorum sensing. Proc. Natl. Acad. Sci. USA 2022, 119, e2201242119. [Google Scholar] [CrossRef]
- Ren, Y.; Zhu, R.; You, X.; Li, D.; Guo, M.; Fei, B.; Liu, Y.; Yang, X.; Liu, X.; Li, Y. Quercetin: A promising virulence inhibitor of Pseudomonas aeruginosa LasB in vitro. Appl. Microbiol. Biotechnol. 2024, 108, 57. [Google Scholar] [CrossRef]
- Yin, L.; Wang, Y.; Xiang, S.; Xu, K.; Wang, B.; Jia, A.-Q. Tyramine, one quorum sensing inhibitor, reduces pathogenicity and restores tetracycline susceptibility in Burkholderia cenocepacia. Biochem. Pharmacol. 2023, 218, 115906. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.S.; Jang, S.Y.; Bang, Y.-j.; Hwang, J.; Koo, Y.; Jang, K.K.; Lim, D.; Kim, M.H.; Choi, S.H. QStatin, a selective inhibitor of quorum sensing in Vibrio species. MBio 2018, 9, e02262-17. [Google Scholar] [CrossRef]
- Kalia, V.C.; Patel, S.K.; Kang, Y.C.; Lee, J.-K. Quorum sensing inhibitors as antipathogens: Biotechnological applications. Biotechnol. Adv. 2019, 37, 68–90. [Google Scholar] [CrossRef]
- Zeng, L.; Lin, F.; Ling, B. Effect of traditional Chinese medicine monomers interfering with quorum-sensing on virulence factors of extensively drug-resistant Acinetobacter baumannii. Front. Pharmacol. 2023, 14, 1135180. [Google Scholar] [CrossRef] [PubMed]
- Woods, K.E.; Akhter, S.; Rodriguez, B.; Townsend, K.A.; Smith, N.; Smith, B.; Wambua, A.; Craddock, V.; Abisado-Duque, R.G.; Santa, E.E. Characterization of natural product inhibitors of quorum sensing reveals competitive inhibition of Pseudomonas aeruginosa RhlR by ortho-vanillin. Microbiol. Spectr. 2024, 12, e0068124. [Google Scholar] [CrossRef] [PubMed]
- Bové, M.; Bao, X.; Sass, A.; Crabbé, A.; Coenye, T. The quorum-sensing inhibitor furanone C-30 rapidly loses its tobramycin-potentiating activity against Pseudomonas aeruginosa biofilms during experimental evolution. Antimicrob. Agents Chemother. 2021, 65, e0041321. [Google Scholar] [CrossRef]
- Hu, H.; Hua, S.Y.; Lin, X.; Lu, F.; Zhang, W.; Zhou, L.; Cui, J.; Wang, R.; Xia, J.; Xu, F. Hybrid biomimetic membrane coated particles-mediated bacterial ferroptosis for acute MRSA pneumonia. ACS Nano 2023, 17, 11692–11712. [Google Scholar] [CrossRef]
- Jakobsen, T.H.; Warming, A.N.; Vejborg, R.M.; Moscoso, J.A.; Stegger, M.; Lorenzen, F.; Rybtke, M.; Andersen, J.B.; Petersen, R.; Andersen, P.S.; et al. A Broad Range Quorum Sensing Inhibitor Working Through sRNA Inhibition. Sci. Rep. 2017, 7, 9857. [Google Scholar] [CrossRef]
- Mane, S.G.; Katagi, K.S.; Bhasme, P.; Pattar, S.; Wei, Q.; Joshi, S.D. Design, synthesis, antibiofilm, quorum sensing inhibition, anticancer and docking studies of novel 2-(4-acridine-9-ylamino)isoindoline-1,3-dione. Chem. Data Collect. 2019, 20, 100198. [Google Scholar] [CrossRef]
- São-José, C.; Costa, A.R.; Melo, L.D. Bacteriophages and their lytic enzymes as alternative antibacterial therapies in the age of antibiotic resistance. Front. Microbiol. 2022, 13, 884176. [Google Scholar] [CrossRef] [PubMed]
- Hassannia, M.; Naderifar, M.; Salamy, S.; Akbarizadeh, M.R.; Mohebi, S.; Moghadam, M.T. Engineered phage enzymes against drug-resistant pathogens: A review on advances and applications. Bioprocess Biosyst. Eng. 2024, 47, 301–312. [Google Scholar] [CrossRef]
- Olatunji, A.O.; Olaboye, J.A.; Maha, C.C.; Kolawole, T.O.; Abdul, S. Next-Generation strategies to combat antimicrobial resistance: Integrating genomics, CRISPR, and novel therapeutics for effective treatment. Eng. Sci. Technol. J. 2024, 5, 2284–2303. [Google Scholar] [CrossRef]
- Gliźniewicz, M.; Miłek, D.; Olszewska, P.; Czajkowski, A.; Serwin, N.; Cecerska-Heryć, E.; Dołęgowska, B.; Grygorcewicz, B. Advances in bacteriophage-mediated strategies for combating polymicrobial biofilms. Front. Microbiol. 2024, 14, 1320345. [Google Scholar] [CrossRef]
- Peng, J.; Guo, C.; Yang, C.; Zhang, L.; Yang, F.; Huang, X.; Yu, Y.; Zhang, T.; Peng, J. Phage therapy for bone and joint infections: A comprehensive exploration of challenges, dynamics, and therapeutic prospects. J. Glob. Antimicrob. Resist. 2024, 39, 12–21. [Google Scholar] [CrossRef]
- Srivastava, A.; Verma, N.; Kumar, V.; Apoorva, P.; Agarwal, V. Biofilm inhibition/eradication: Exploring strategies and confronting challenges in combatting biofilm. Arch. Microbiol. 2024, 206, 212. [Google Scholar] [CrossRef]
- Panthi, V.K.; Fairfull-Smith, K.E.; Islam, N. Liposomal drug delivery strategies to eradicate bacterial biofilms: Challenges, recent advances, and future perspectives. Int. J. Pharm. 2024, 655, 124046. [Google Scholar] [CrossRef]
- Lordelo, R.; Branco, R.; Gama, F.; Morais, P.V. Assessment of antimicrobial resistance, biofilm formation, and surface modification potential in hospital strains of Pseudomonas aeruginosa and Klebsiella pneumoniae. Heliyon 2024, 10, e30464. [Google Scholar] [CrossRef] [PubMed]
- Aerts, E. AI-Guided Detection of Antibiotic-Resistantbacteria Using Resistance Genes. Master’s Thesis, Chalmers University of Technology, Gothenburg, Sweden, 2024. [Google Scholar]
- Ikpe, F.; Williams, T.; Orok, E.; Ikpe, A. Antimicrobial resistance: Use of phage therapy in the management of resistant infections. Mol. Biol. Rep. 2024, 51, 925. [Google Scholar] [CrossRef]
- Wilson, C.; Thomson, E.C. Resilience to emerging infectious diseases and the importance of scientific innovation. Future Healthc. J. 2024, 11, 100023. [Google Scholar] [CrossRef]
- Gholap, A.D.; Khuspe, P.R.; Pardeshi, S.R.; Uddin, M.J.; Das, U.; Hatvate, N.T.; Rojekar, S.; Giram, P.; Khalid, M.; Choonara, Y.E. Achieving Optimal Health With Host-Directed Therapies (HDTs) in Infectious Diseases—A New Horizon. Adv. Ther. 2025, 8, 2400169. [Google Scholar] [CrossRef]
- Mahajan, S.; Rana, P.S.; Ferguson, D.L.; Wozniak, D.J. Unraveling the Complexity: Biofilms and Their Interactions with Myeloid Immune Cells. In Biofilm Matrix; Springer: Berlin/Heidelberg, Germany, 2024; pp. 311–336. [Google Scholar]
- Sun, D.; Chang, Q.; Lu, F. Immunomodulation in diabetic wounds healing: The intersection of macrophage reprogramming and immunotherapeutic hydrogels. J. Tissue Eng. 2024, 15, 20417314241265202. [Google Scholar] [CrossRef]
- do Nascimento Júnior, J.X.; Sola-Penna, M.; Zancan, P. Clotrimazole reverses macrophage M2 polarization by disrupting the PI3K/AKT/mTOR pathway. Biochem. Biophys. Res. Commun. 2024, 696, 149455. [Google Scholar] [CrossRef]
- Genito, C.J.; Darwitz, B.P.; Reber, C.P.; Moorman, N.J.; Graves, C.L.; Monteith, A.J.; Thurlow, L.R. mTOR signaling is required for phagocyte free radical production, GLUT1 expression, and control of Staphylococcus aureus infection. Mbio 2024, 15, e00862-24. [Google Scholar] [CrossRef]
- Rasquel-Oliveira, F.S.; Ribeiro, J.M.; Martelossi-Cebinelli, G.; Costa, F.B.; Nakazato, G.; Casagrande, R.; Verri, W.A. Staphylococcus aureus in inflammation and pain: Update on pathologic mechanisms. Pathogens 2025, 14, 185. [Google Scholar] [CrossRef] [PubMed]
- Nasri, M.; Ritter, M.U.; Mir, P.; Dannenmann, B.; Kaufmann, M.M.; Arreba-Tutusaus, P.; Xu, Y.; Borbaran-Bravo, N.; Klimiankou, M.; Lengerke, C. CRISPR-Cas9n-mediated ELANE promoter editing for gene therapy of severe congenital neutropenia. Mol. Ther. 2024, 32, 1628–1642. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Long, S.; Wang, H.; Wang, Y. Biofilm therapy for chronic wounds. Int. Wound J. 2024, 21, e14667. [Google Scholar] [CrossRef]
- Qu, S.; Ji, Y.; Fan, L.; Yan, T.; Zhu, G.; Song, H.; Yang, K.; Han, X. Light-Enhanced Hypoxia-Responsive Gene Editing for Hypoxia-Resistant Photodynamic and Immunotherapy. Adv. Healthc. Mater. 2024, 13, 2302615. [Google Scholar] [CrossRef]
- Tran, H.H.; Watkins, A.; Oh, M.J.; Babeer, A.; Schaer, T.P.; Steager, E.; Koo, H. Targeting biofilm infections in humans using small scale robotics. Trends Biotechnol. 2024, 42, 479–495. [Google Scholar] [CrossRef]
- Resnik, D.B. Biosafety, biosecurity, and bioethics. Monash Bioeth. Rev. 2024, 42, 137–167. [Google Scholar] [CrossRef]
- Singh, V.; Rastogi, M. Future and Challenges of Microbiome Engineering. In Microbiome Engineering; CRC Press: Boca Raton, FL, USA, 2024; pp. 263–280. [Google Scholar]
- Wiersma, B. Assessment of Biofilm Matrix Disruption as Treatment for Pseudomonas aeruginosa Lung Infections. Master’s Thesis, Utrecht University, Utrecht, The Netherlands, 2024. [Google Scholar]
- Fang, X.; Li, P.; Luo, C.; Wu, C.; Shi, X.; Lan, J.; Zhao, F.; Wang, F.; Zhang, K.; Yang, C. Epiphytic bacterial consortia drive growth regulation in potato under methyl jasmonate elicitation: A leaf surface multi-omics perspective. Plant Physiol. Biochem. 2025, 222, 109737. [Google Scholar] [CrossRef] [PubMed]
- Ramachandran, G.; Palanisamy, B.; Chackaravarthy, G.; Chelliah, C.K.; Rajivgandhi, G.; Quero, F.; Natesan, M. Diagnosis of Physical Stimuli Response Enhances the Anti-Quorum Sensing Agents in Controlling Bacterial Biofilm Formation. In Nanoscience and Nanotechnology for Smart Prevention, Diagnostics and Therapeutics: Fundamentals to Applications; Wiley: Hoboken, NJ, USA, 2024; pp. 91–115. [Google Scholar]
- Ma, Z.; Zuo, T.; Frey, N.; Rangrez, A.Y. A systematic framework for understanding the microbiome in human health and disease: From basic principles to clinical translation. Signal Transduct. Target. Ther. 2024, 9, 237. [Google Scholar] [CrossRef] [PubMed]
- Odoom, A.; Osman, A.-H.; Dzuvor, C.K. Recent advances in immunotherapeutic and vaccine-based approaches for the treatment of drug-resistant bacterial infections. ACS Infect. Dis. 2025, 11, 1366–1402. [Google Scholar] [CrossRef] [PubMed]
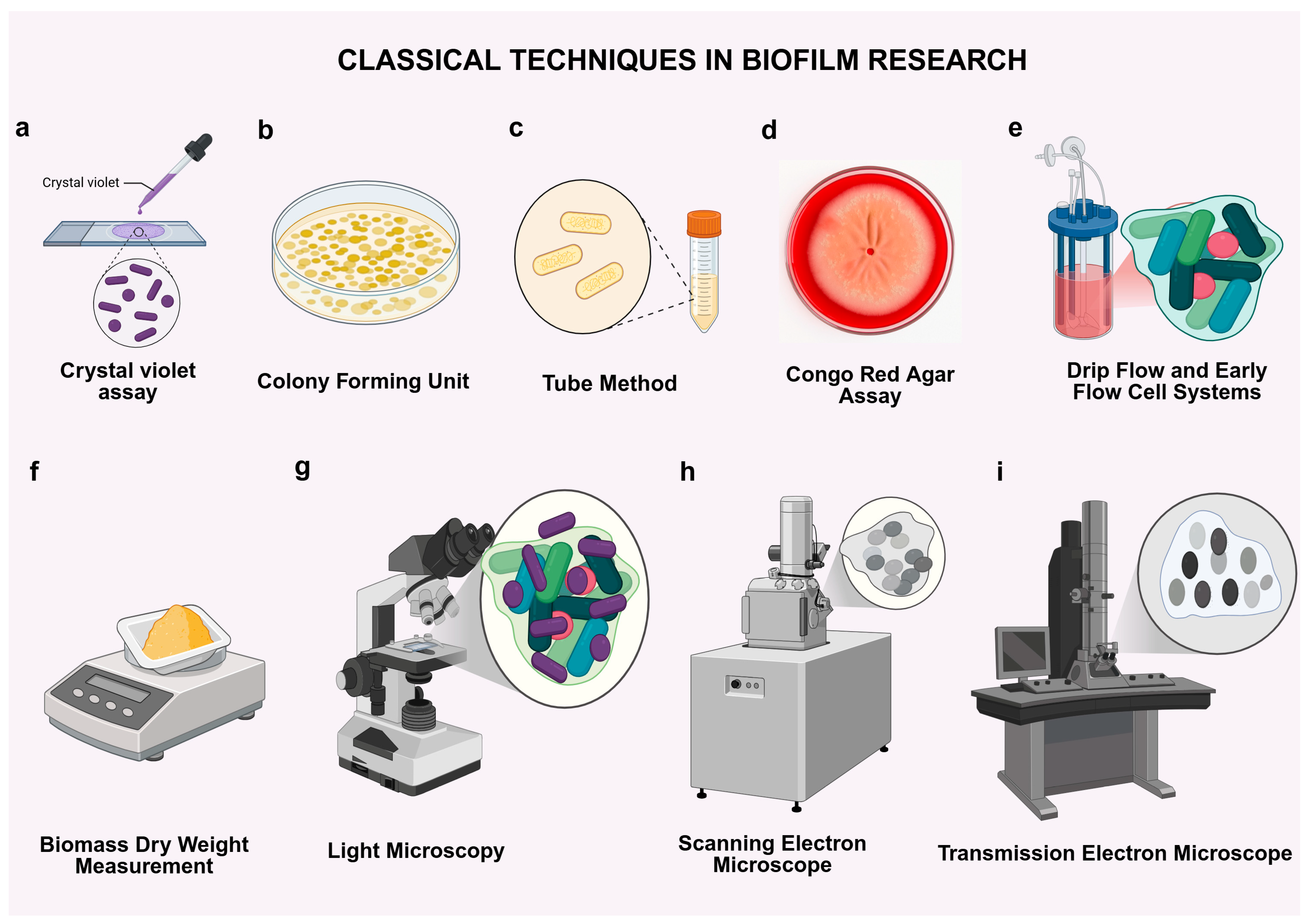

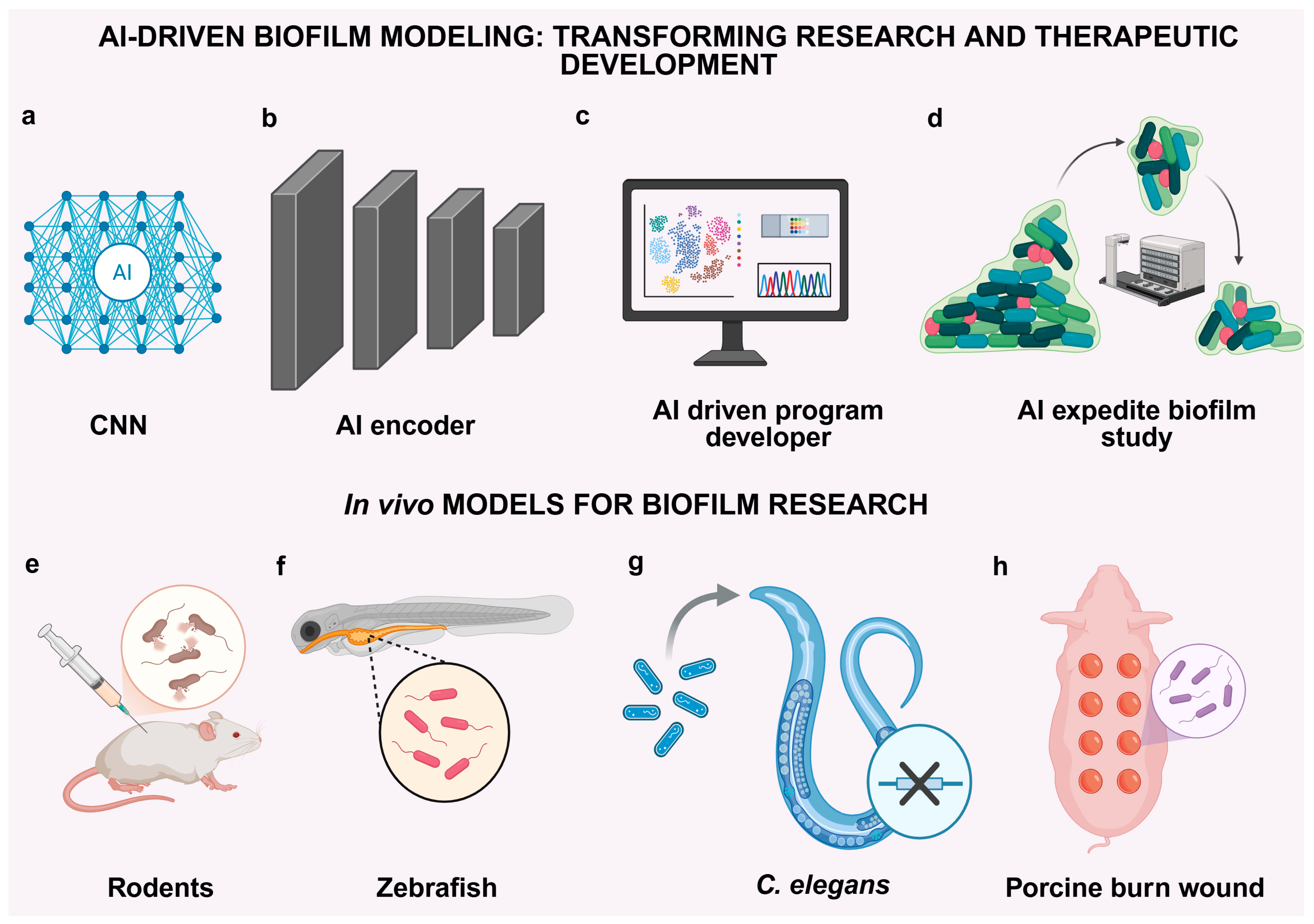

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Haval, M.; Unakal, C.; Ghagane, S.C.; Pandit, B.R.; Daniel, E.; Siewdass, P.; Ekimeri, K.; Rajamanickam, V.; Justiz-Vaillant, A.; Lootawan, K.-A.A.; et al. Biofilms Exposed: Innovative Imaging and Therapeutic Platforms for Persistent Infections. Antibiotics 2025, 14, 865. https://doi.org/10.3390/antibiotics14090865
Haval M, Unakal C, Ghagane SC, Pandit BR, Daniel E, Siewdass P, Ekimeri K, Rajamanickam V, Justiz-Vaillant A, Lootawan K-AA, et al. Biofilms Exposed: Innovative Imaging and Therapeutic Platforms for Persistent Infections. Antibiotics. 2025; 14(9):865. https://doi.org/10.3390/antibiotics14090865
Chicago/Turabian StyleHaval, Manasi, Chandrashekhar Unakal, Shridhar C. Ghagane, Bijay Raj Pandit, Esther Daniel, Parbatee Siewdass, Kingsley Ekimeri, Vijayanandh Rajamanickam, Angel Justiz-Vaillant, Kathy-Ann A. Lootawan, and et al. 2025. "Biofilms Exposed: Innovative Imaging and Therapeutic Platforms for Persistent Infections" Antibiotics 14, no. 9: 865. https://doi.org/10.3390/antibiotics14090865
APA StyleHaval, M., Unakal, C., Ghagane, S. C., Pandit, B. R., Daniel, E., Siewdass, P., Ekimeri, K., Rajamanickam, V., Justiz-Vaillant, A., Lootawan, K.-A. A., Oliveira, F. M. D., Bashetti, N., Naqvi, T. A., Shettar, A., & Bhasme, P. (2025). Biofilms Exposed: Innovative Imaging and Therapeutic Platforms for Persistent Infections. Antibiotics, 14(9), 865. https://doi.org/10.3390/antibiotics14090865







