Abstract
Animal and food sources are seen as a potential transmission pathway of multi-drug resistance (MDR) micro-organisms to humans. Escherichia. coli is frequently used as an indicator of fecal contamination in the food industry and known as a reservoir of antimicrobial resistance genes (ARGs). Microbial contamination as a major outcome for the poultry and egg industry and is a serious public health problem. In the present study we performed the quantification of β-glucoronidase positive E. coli in 60 fecal samples of breeding and laying hens collected in Portugal in 2019. Phylogenetic and pathotypic characterization, antimicrobial susceptibility, and detection of resistant extended-spectrum β-lactamase (ESBL) genes were assessed. The phylogenetic and pathogenic characterization and detection of ESBL genes were assessed by real-time PCR and antimicrobial susceptibility was evaluated using the disk diffusion method. Overall, E. coli quantification was 6.03 log CFU/g in breeding hens and 6.02 log CFU/g in laying hens. The most frequent phylogroups were B1. None of the isolates was classified as diarrheagenic E. coli (DEC). In total, 57% of the isolates showed MDR and 3.8% were positive for ESBL. Our study highlights that consumers may be exposed to MDR E. coli, presenting a major hazard to food safety and a risk to public health.
1. Introduction
Animal food products, such as eggs, meat, and milk, are abundant in proteins essential for the body’s maintenance, repair, and growth [1]. Poultry is among the most reported carriers of foodborne pathogens [2]. Hughes et al. [3] also asserted that poultry meat, red meat, and eggs are recognized as major vectors for the transmission of pathogens.
E. coli is a typical inhabitant of the gut of warm-blooded animals and is used frequently as an indicator bacterium of fecal contamination in the food industry. E. coli is a non-spore-forming, Gram-negative rod, usually motile by peritrichous flagella that is a member of the Enterobacteriaceae [4]. Many monitoring programs include E. coli because they are established markers of fecal contamination, ubiquitous in food-producing animals, easy to cultivate, and readily acquire resistance mechanisms to combat agents with activity against Gram-negative organisms [5]. They are also known reservoirs of ARGs that can be transferred horizontally to and from other closely related bacteria [6]. E. coli is considered a good indicator of the selective pressure imposed by antimicrobial use in food animals and has been hypothesized to be a potential predictor of emerging resistance in pathogenic bacteria that cannot be recovered from meat or animal samples Furthermore, annual trends indicate a possible correlation between Salmonella spp. and E. coli resistance [7]. The reason for using E. coli as an indicator is that it appears only at low background levels in the environment but possesses high survival rates [8].
Microorganisms from animal, environmental, and human sources normally contaminate raw foods [9]. The initial number of living microorganisms, including pathogens, will be substantially reduced when properly processed. However, the prevalence of pathogenic microorganisms and deterioration in ready-to-eat (RTE) foods can substantially increase through post-processing handling activities, the duration of exposure at points of sale and storage conditions [10].
At slaughter, resistant strains from the gut readily soil poultry carcasses, and as a result, poultry meat is often contaminated with resistant E. coli, likewise eggs become contaminated during laying [11,12,13,14,15,16,17,18]. Hence, resistant fecal E. coli from poultry can infect humans both directly and via food. These resistant bacteria may colonize the human intestinal tract and may also transfer resistance genes to human endogenous flora [19]. In the case of eggs, microbial contamination has a major outcome for the poultry industry and contaminated eggs are a serious public health problem worldwide. The importance of these diseases in humans can range from mild symptoms to life-threatening situations [20]. Egg and its products are an important component source of necessary nutrients. Eggs can act as a vector in the transmission of food poisoning microorganisms. Many investigations have already reported contamination of eggs with Salmonella spp., Listeria monocytogenes [21], Campylobacter spp. [22], and E. coli [23], and if the appropriate treatment does not occur, these pathogens can reach consumers’ homes and become a food safety problem.
Over the past 50 years, the use of antibiotics combined with strict biosecurity and hygiene measures has helped the poultry industry grow, preventing the negative impacts of many avian diseases caused by previously referred microorganisms [24]. The use of antibiotics to control gastrointestinal infections can lead to a change in the intestinal microbiota of hens, which can influence their immunity and health [25].
Scientific evidence suggests that the use of antimicrobials in animal production may promote bacterial resistance in treated animals [26]. Bacterial resistance of E. coli to antibiotics has been the subject of several studies in recent years [4,27]. Bacterial resistance to animal antibiotics is a public health problem. Antibiotic abuse and associated selection pressure led to decreased therapeutic efficacy and created populations of antibiotic-resistant microorganisms. Antibiotic resistance can spread over time, despite the suspension of antibiotic use. Several studies have suggested that antimicrobial resistance (AMR) bacteria and their AMR determinants can be transmitted from food animals to humans by direct contact and/or through animal products [28,29].
The use of antibiotics for growth promotion purposes is prohibited in the European Union. In intensive production systems, animals are exposed to a high risk of infection, as they live under stressful conditions and are driven to increase productivity. In these systems, the frequent application of antibiotics are perfect circumstances for bacterial strains to develop and resist antibiotics [30,31,32]. In Portugal, antibiotics used for application in animals, authorized for the treatment of infections, are oxytetracycline (OCT), amoxicillin (AMX), tylosin (TYL), colistin (CL), doxycycline (DOX), ampicillin (AMP), tiamulin (TIA), sulfadiazine (SFD), and enrofloxacin (ENR) [33].
E. coli strains have been classified based on genetic substructures associated with different phylogenies in different phylogroups that present different phenotypic and genotypic characteristics [34]. The PCR-based assay developed by Clermont et al. [35] is intended for the classification of E. coli strains into the major phylogroups A, B1, B2, and D; however, this method could only validate 80–85% of all E. coli phylogroups and it is sometimes necessary to use more alternatives [36,37]. A modification was made to the triplex method by adding one gene, resulting in a quadruple PCR [38]. Five strains or clades (I–V) were also found in E. coli strains, of which clade I is currently included in the phylogenetic grouping, making eight groups: A, B1, B2, C, D, E, F, and clade I [39]. Studies have shown that strains associated with virulent extraintestinal infection generally belong to phylogroups B2, D, or E and that commensal isolates of E. coli are generally affiliated with groups A and B1 [40,41].
Although this microorganism is considered a commensal, there are strains that can cause diarrheal diseases [42]. The virulence attributes have been used to differentiate pathogenic strains of E. coli and divided into diarrheal pathogens causing diarrhea (DEC) and extraintestinal E. coli (ExPEC) [43] based on the site of infection. There are six classic pathotypes of DEC: enteropathogenic (EPEC), shiga toxin–producing (STEC), enteroaggregative (EAEC), enterotoxigenic (ETEC), enteroinvasive (EIEC), and diffusely adherent E. coli (DAEC) [44]. Two additional E. coli pathotypes, belonging to ExPEC, are responsible for extraintestinal infections: uropathogenic E. coli (UPEC) causing urinary tract infections and neonatal meningitis associated E. coli (NMEC) [45,46]. Avian pathogenic E. coli (APEC) are a member of DEC, closely related to EPEC, which are frequently assigned to specific phylogenetic groups along with human UPEC and NMEC that cause disease outside the intestine [47,48]. During the last decades, the emergence of AMR bacteria has been enormously announced worldwide. In relation to an extensive use of β-lactam antibiotics in both clinical and nonclinical settings, a great diversity of β-lactamase types has consequently emerged [49]. In this context, ESBL constitute a mechanism of resistance of great clinical relevance that is spreading not only in humans but also among domestic animals [50]. ESBL-producing Enterobacteriaceae have been recognized as highly prevalent in food-producing animals and derived food, in the Mediterranean countries [51,52,53,54,55].
This study was conducted to investigate the prevalence and characterization of β-glucoronidase positive E. coli in fecal samples collected from breeding and laying hens in Portugal. The isolates were characterized by their phylogroups and a search for virulence genes was performed (pathotypes). Antimicrobial resistance assays and ESBL-associated genes detection were also performed. The main objectives were to increase knowledge regarding E. coli carriage in hens and to raise aware of the risks that consumers may be exposed to with the spread of MDR strains, especially if good hygiene practices are not completely fulfilled, causing a public health and/or food safety problem.
2. Results
2.1. Quantification of β-Glucoronidase Positive E. coli from Hens Fecal Samples
The microbiological load of E. coli in the 60 fecal samples was calculated in TBX, TBX supplemented with ampicillin (TBXamp) or with enrofloxacin (TBXenro) and the average of the results obtained were 6.03 log CFU/g in TBX, 4.77 log CFU/g in TBXamp, and 3.49 log CFU/g in TBXenro. Results of E. coli enumeration in the three different agar and type of hens are in shown in Table 1.

Table 1.
Microbiological load of E. coli in TBX, TBX supplemented with ampicillin (TBXamp) or with enrofloxacin (TBXenro), for breeding hens and egg laying feces samples (mean ± standard deviation).
From the analysis of Table 1, it is possible to verify that there are only significant differences (p < 0.05) between laying hens and breeding hens feces samples in the Log TBXenro parameter, noting that this is significantly higher in the breeding hens.
2.2. Determination of the E. coli Phylogenetic and Patotypes Groups
Seventy nine isolates were selected for further studies as shown in Table 2.

Table 2.
Number of samples of laying and breeding hens analyzed, and number of isolates of E. coli resistant to ampicillin or enrofloxacin selected for further characterization.
The most predominant phylogroup was B1 (75%) followed by phylogroup A (19%) (Table 3).

Table 3.
Percentages of phylogroup detected in isolates from hens feces.
Phylogroup B1 was the most predominant with more than 80% of the isolates from breeding hens and 60.7% in egg laying hens, followed by phylogroup A. Only isolates from egg laying hens were identified as phylogroup D (3.6%) or as not belonging to any phylogroup (3.6%), requiring the performance of multilocus sequence typing (MLST) that was not made. Isolates belonging to phylogroup B2, C, F, or clade I were not detected.
Concerning pathotypes, none of the isolates studied was classified as DEC.
2.3. Susceptibility to Antimicrobials
An overview of the antimicrobial susceptibility of the 79 E. coli isolates from breeding and egg laying hens is given in Table 4.

Table 4.
Susceptibility to antibiotics of E. coli isolated from feces of egg laying and breeding hens according to EUCAST [56] and CLSI [57] parameters.
AMP was the antibiotic to which the largest number of isolates showed resistance, with 81% (64 isolates) of the resistant isolates; in decreasing order of resistance of the isolates: NAL with 65.8% (52 isolates), TET with 62.0% (49 isolates), CIP with 59.5% (47 isolates), SULF with 44.3% (35 isolates), TMP with 31.6% (25 isolates), CHL with 11.4% (9 isolates) and AZM, and CAZ and GRM with 3.8% (3 isolates). As for CTX and MEM, none of the 79 isolates from hens’ feces showed resistance, making them the only two antibiotics to which the 79 isolates were 100% sensitive.
Since the 79 hens’ feces isolates were derived from two distinct functions: breeders and layers, we decided to verify the distribution of susceptibility to the different antimicrobials as a function of the two hens’ functions (Figure 1). Excluding that for Azithromycin, a greater percentage of resistant isolates is observed in breeding hens.
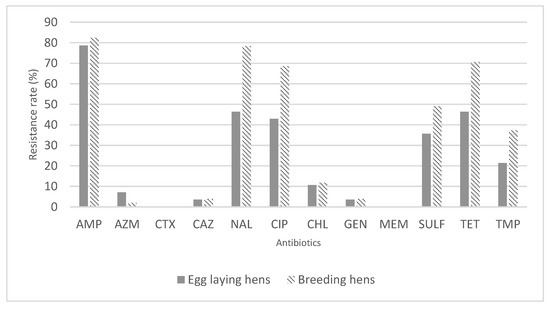
Figure 1.
Percentage of resistance to the antimicrobials under study, for feces isolates of E. coli from egg laying hens or breeding hens.
2.4. Multiresistant Isolates
The analysis of the resistance profiles of the 79 E. coli isolates showed that none of the isolates under study was susceptible to all the tested antimicrobial groups; 18 (22.8%) were resistant to only one group, 16 (3.6%) were resistant to two groups, and 45 (57%) were MDR (resistant to three or more groups of antimicrobials).
The largest number of isolates (19 isolates) exhibited resistance to five different categories of antibiotics; moreover, tree isolates exhibited resistance to six groups. The PEN, FLU, TET, MA, and SULF profile was the most frequent with 42.2% (n = 19). Table 5 summarizes the multiple MDR patterns exhibited by the 45 isolates.

Table 5.
Distribution of multi resistant isolates of E. coli and number of categories for which they exhibited resistance.
When we looked at the results of E. coli MDR isolates, we observed that 36 isolates (80%) belong to breeding hens and only 9 belong to laying hens.
Further statistical analysis was carried in order to evaluate the existence of significant differences between ENRO- and AMP-resistant E. coli isolates for laying hens and breeding hens. In fact, on average, the number of antibiotic resistance observed in E. coli isolates for laying hens is significantly lower (p < 0.05) than E. coli isolates for breeding hens. No significant differences (p > 0.05) were observed between ENRO- and AMP-resistant E. coli isolates number of antibiotic resistance. The MDR isolates of E. coli within the phylogroups identified was also analyzed, and significant differences were observed between phylogroups A and B1, verifying that phylogroup A isolates presented a significant lower number of antibiotic resistance when compared with phylogroup B1 (p < 0.05).
2.5. Detection of ESBL Resistance Genes
From the isolates that showed resistance to CAZ, we verified which resistance genes were present for ESBL. In tree isolates (rate of 3.8% in analyzed samples) genes blaTem and blaCTX-M were detected. More information about the three isolates can be found in Table 6.

Table 6.
Characteristics of isolates positive for resistant ESBL genes.
3. Discussion
E. coli isolates from breeding hens were more resistant to AMP (83.3%) than to ENRO (69.2%). Likewise, results from laying hens also demonstrate that the isolates were more resistant to AMP (75.1%) than to ENRO (47%). Of the total samples, isolates were more resistant to AMP (79.3%) than ENRO (58.4%), obtaining a higher number of isolates resistant to AMP than resistant to ENRO. The use of both antimicrobials is allowed for disease treatments in animals and their spread in farms may lead to an increase in microbial resistance in the population. Seventy-nine E. coli strains were isolated from the fecal samples of laying hens. E. coli strains are now classified into eight phylogroups: A, B1, B2, C, D, E, F, and clade I [41]. Studies have shown that isolates belonging to the A and B1 phylogroups are commensals, while those that belong to the B2, D, and E groups are the extraintestinal pathogenic strains [58,59,60]. Obeng et al. [61] determined the phylogenetic groups of E. coli isolates from the feces of intensively farmed and free-range poultry from South Australia. They found that the predominant phylogenetic groups were phylogroup B1 with 39.4% and phylogroup A with 32.3%. In a more recent study, Hayashi et al. [62], revealed that the 70 E. coli isolates from hens’ samples in Japan majorly belonged to group B1 (25.7%) and group A (14.3%). Similar to our results, these studies also demonstrate the predominance of B1 and A in samples from hens. In the study by Projahn et al. [63], broiler breeder lots and the corresponding eggs were analyzed. Of the eggs tested, 0.9% (n = 560) were contaminated on the outer surface of the shell. Additional analysis showed a relationship between the species found in the eggs and those isolated from the corresponding lots of origin, which demonstrates a pseudo-vertical transfer of Enterobacteriaceae to the hatchery. Isolates of the four positive eggs of flock were all found to be E. coli of the phylogroup B1. This study demonstrates the contamination of eggshells through E. coli contaminated feces from egg laying hens, presenting a risk to the health of consumers. It is also possible that they constitute a possible source of contamination for the chicks, given the detection of their presence in the feces of breeding hens, thus representing a risk for chicken and egg consumers. Furthermore, isolates belonging to phylogroups D and E that have been associated with virulent extraintestinal infection were found. Adefioye et al. [64] concluded in their study that most of the human isolates from fecal samples of apparently healthy individuals belonged to phylogroup B1. They only found a few isolates belonging to B2 and D phylogroups and concluded that these isolates were mostly commensals, which as a result of antibiotic exposure and other environmental and genetic factors, may revert to being pathogenic [65].
Tenaillon et al. [34] referred that genetic diversity of E. coli exhibits host taxonomic and environmental components. This can be illustrated by the prevalence of the four main phylogenetic groups in various human and animal populations. In humans, group A strains are predominant (40.5%), followed by B2 strains (25.5%), while B1 and D strains (17% each) are less common. In animals, there is a predominance of B1 strains (41%), followed by A strains (22%), B2 (21%) and, to a lesser extent, D strains (16%). Our results corroborate this statement since the most predominant phylogroups found in hens were B1 and A.
The presence of antibiotic-resistant foodborne pathogens in food can lead to gastrointestinal disturbances in humans [66]. On the other hand, antibiotic-resistant pathogens can transfer the gene to other microorganisms, resulting in the spread of AMR pathogens [67,68]. There are not many studies available concerning AMR in hens’ fecal samples. Two studies used a similar number of isolates from the same source. In the study by Langata et al. [69], AMR patterns among 85 resistant hen fecal isolates in Kenya were characterized and Abassi et al. [70] characterized 83 E. coli fecal isolates recovered from hens, in Tunisia. NAL was used in the three studies; Portugal has the higher number of resistant isolates (65.8%) while Kenya only has 18.8%. In the case of TET, the three studies presented a higher number of isolates being resistant between 90% in Tunisia and 42% in Kenya. Resistance to CTX was found in Tunisia and the number of isolates resistant to CAZ were similar in Tunisia and Portugal, and were not tested in Kenya. CIP was tested in Portugal and Kenya with 60% of Portuguese isolates being resistant and only 1.2% in Kenya. The differences found can be related to different use of antibiotics in agriculture and chicken or egg production.
In Portugal, the Directorate General for Food and Veterinary (DGAV) [33] controls the use of antibiotics and reports those authorized for the treatment of infections. Our results demonstrate antimicrobial-resistant isolates in hens’ feces that are not present in these reports, that is, that are not permitted for animal use.
When analyzing the results of antimicrobial susceptibility differentiating the types of hens between breeding and egg laying hens, we found that isolates from the feces of breeding hens showed a higher percentage of resistance to a greater number of antibiotics. Excluding that for Azithromycin, a greater percentage of resistant isolates was observed in breeding hens. This can be related to prophylactic use of antibiotics in poultry production.
Based on these results, it is not possible to make any correlation between the resistances and phylogroups.
Knowledge about MDR load and resistance patterns in isolates extracted from food-producing animals is imperative to design targeted interventions to limit antibiotic use. The use of commensal intestinal E. coli as a marker for the presence of resistance in bacterial flora is a critical component of MDR surveillance programs in food producing and wild animals [71]. In Liu et al. [72], fecal samples were obtained from six broiler fattening farms in China. They describe that the MDR of E. coli isolates was 91%. According to Koju et al. [73], hen caecum samples were collected from slaughterhouses/stores in Nepal and it was found that 71% showed resistance to at least three categories of antimicrobials. Comparing the MDR value obtained in our study (57%) with the values from those reports, we found that the number of the MDR of E. coli isolates from hen samples in Portugal is not as high as in other countries, despite this, it is still a worrying reality. The monitoring and treatment of drug-resistant bacteria in the poultry industry will be a long and difficult task, and one which will require a collaborative effort and should include aspects of chick breeding, the breeding environment, and feed additives.
The most common pattern of antibiotic resistance is the one that conjugates penicillins, fluoroquinolones, tetracyclines, macrolides, and sulfonamides. When we compare these data with those published by DGAV, we find that three of the categories (PEN, TET, and MA) present in the most common pattern presented by our isolates, correspond to the three classes of antibiotics most commercialized in Portugal.
Isolates from breeding hens showed higher AMR and more MDR isolates when compared to isolates from laying hens. Once again, these results demonstrate the relationship between the prophylactic use of antibiotics and poultry production.
According to the European Food Safety Authority (EFSA), the prevalence of presumptive ESBL E. coli producers in the different animal species and their products varies within the EU countries [74]. In Denmark, isolates of CTX-M producing E. coli from healthy egg laying hens (not exposed to antimicrobial agents) were found in stool samples collected from the ground and cloacal swabs, so there is a possibility to find ESBL–E. coli between eggshells, which can be contaminated by contact with feces [75]. In the present study, the blaTEM and blaCTX-M genes were detected in three E. coli isolates (5%) and could be considered as potential ESBL producers. Both the blaTEM and blaCTX-M genes have been detected among food producing meat at retail and broiler products in recent studies in Portugal [76,77]. Machado et al. [76] identified that 60% of uncooked hen carcasses (n = 20) and 10% of feces from healthy hens (n = 20) were positive for blaTEM and blaCTX-M genes. Clemente et al. [77] examined the existence of ESBL in meat collected at retail stores in Portugal and found a prevalence of 30.3% in poultry meat, while this was 11.8% and 10.5% for beef and pork, respectively. The results of these studies demonstrate the importance of monitoring the ESBL genes in E. coli isolates and how their study is essential for food chain safety and human health. In fact, and interesting to point out, is that the three isolates harboring ESBL genes were from three different farms, from distinct types of hens (breeding and laying) and from two different phylogroups (B1 and E) and all showing MDR. Mahmud et al. [78] have already described a high prevalence of MDR in ESBL-producing E. coli and 71% of ESBL-producing E. coli isolates were MDR. The fact that isolates came from different places indicates the potential spread of microorganisms highly resistant to antimicrobials that can reach consumers’ homes, thus changing their environmental microbiota, and if food becomes contaminated, it could accumulate and proliferate in the intestine, where genetic transfer can occur.
4. Conclusions
This research work complements and updates previous studies carried out in Portugal on MDR E. coli from food products of animal origin.
In the present study, we found E. coli isolates belonging mostly to phylogroup B1 and A, which are reported as commensals, but also to phylogroup E, classified as extraintestinal pathogenic strains. None of the isolates studied was classified as DEC; however, as the isolation agar used was TBX only β-glucoronidase positive E. coli were selected. In total, 81% of isolates tested were resistant to at least one antimicrobial and a greater percentage of resistant isolates was observed in breeding hens; 57% of total isolates were MDR and again, a greater percentage of MDR isolates was observed in breeding hens. Three resistant isolates were considered as potential ESBL producers and once again, two out of three were from breeding hens.
The work shows that E. coli can exhibit multiple resistance to various antimicrobials, posing a major risk to food safety and public health.
This study also reinforces previous reports that ESBL-producing E. coli has become one of the leading indicators for estimating MDR burden in animals and other sectors from a unique health perspective.
5. Materials and Methods
5.1. Sampling and Bacterial Isolation
Sixty fecal samples from breeding (n = 31) and egg laying (n = 29) hens were collected from the Centro and Lisboa and Vale do Tejo regions, between March and May 2019 from 36 farms, by Portuguese Food Authorities under the control programs for Salmonella. Laboratory procedures for the isolation and identification of E. coli followed the protocols defined by ISO 16649-2:2001 [79]. Briefly, 25 g of each fecal sample was mixed with 225 mL of peptone-buffered water (BPW) (Bio-Rad, Hercules, CA, USA). Decimal dilutions were prepared up to 10−5 with Tryptone Salt Broth (Bio-Rad) and 1 mL of the 10−4 and 10−5 dilutions were plated in Tryptone Bile X-Glucuronide (TBX) agar (Bio-Rad), 10−3 to 10−5 dilutions were plated in TBX agar supplemented with 100 μg/mL of ampicillin (TBXamp) (Sigma-Aldrich, St. Louis, MO, USA) and 10−2 to 10−5 dilutions were plated in TBX agar supplemented with 10 μg/mL of enrofloxacin (TBXenro) (Sigma-Aldrich). Ampicillin was chosen as a selective agent because it has a broad spectrum of action, particularly against Gram-negative bacteria, and is rapidly absorbed and eliminated in poultry. Enrofloxacin is applied in birds for the treatment of infections of the digestive and respiratory tract, belongs to the group of fluoroquinolones, is bactericidal, and is used against Gram-negative bacteria. Plates were incubated at 44 °C for 18 to 24 h. After the incubation period, all plates were analyzed, and the number of colonies was counted for each dilution applied. A colony from TBXamp and from TBXenro was then sub-cultured onto a slant tube containing Heart Infusion Agar (HIA) (Biogerm, Moreira, Portugal), placed again at 35 °C for 18 ± 2 h. Subsequently, these tubes were stored at 4 °C agar.
Confirmation of characteristic colonies was carried out by lactose fermentation and indol production: from the HIA a tube containing phenol red lactose broth (Sigma-Aldrich, Missouri, EUA) and a tube containing tryptophan broth (Biogerm) were inoculated and incubated at 35–37 °C for 24 h. The positive test consists of a color change from red to yellow, indicating a pH change to acidic in the lactose broth and the formation of a ring after the addition of Kovacs reagent (Sigma-Aldrich) in tryptophan broth.
5.2. Phylogenetic Grouping and Determination of E. coli Pathotypes
The phylogenetic group (A, B1, B2 D, E, F, and clade I) of E. coli isolates was determined by a specific multiplex PCR designed by Clermont et al. [38]. DNA extraction was performed using 1000 μL of presumptive E. coli grown overnight at 37 °C on Tryptic Soy Broth (TSB) tubes (Bio-Rad) were centrifuged for 10 min at 13,000 rpm (Bio-Rad: Model 16 Microcentrifuge), the supernatant was discarded, and the pellet was resuspended in 1000 μL of DNase-free ultra-pure water and vortexed for 2 min using BR-2000 vortexer (Bio-Rad) and the cells were lysed by boiling for 15 min using a digital heat block (VWR, Pensilvânia, EUA). The cell debris were removed by centrifugation at 10,000 rpm for 5 min using a MiniSpin microcentrifuge (ThermoFisher Scientific, Waltham, MA, USA). The supernatant was used as a template in the PCR assay in a final volume of 20 µL, containing PCR master mix multiplex PCR NZYTaq 2x Green Master mix (NZYTech, Lisbon, Portugal), and the primer set arpA (2 μM), chuA (1 μM), yjaA (1 μM), and TspE4.C2 (1 μM) (Eurofins Genomics, Porto, Portugal) as described in Table 7. For the PCR reaction, we considered the number of samples to be validated, positive controls, and negative control. E. coli O111 (arpA+; TspE4.C2+), E. coli O157:H7 (arpA+; chuA+), and E. coli K12 (arpA+; yjaA+) were used as positive controls. As a negative control, in place of the template, DNase-free water was added in the same amount. The PCR conditions were as follows: 95 °C, 3 min; 39 cycles of 95 °C, 30 s; 58 °C, 30 s; 72 °C, 30 s; and 72 °C, 5 min. The results were visualized using UV light (Syngene® Cambridge, UK).

Table 7.
List of primers used for the determination of phylogenetic groups.
According to the presence or absence of arpA, chuA, yjaA, and TspE4.C2 genes, a phylogroup was assigned to each isolate, as previously described by Clermont et al. [38] Table 8.

Table 8.
Assignment of phylogroups of E. coli isolates based on the presence of genes arpA, chuA, yjaA, and TspE4.C2.
For the distinction of phylogroups, which could be group A or C, E or D, or E, or clade I, two more PCR were performed.
For the distinction of phylogroups A or C, 1 µL of template was used for PCR in a final volume of 20 µL, containing MgCl2 (1.5 mM), Taq buffer (1x), and trpA (0.25 μM) (Eurofins Genomics) as described in Table 9, Taq (1U), (NZYTech) dNTPs (0.25 μM) (NZYTech) and DNase-free water. DNA extraction was performed as previously described. For the PCR reation, the number of samples to be validated for this phylogenetic group was considered, a positive and a negative control. E. coli FV19459 (trpA+) was used as a positive control. As a negative control, in place of the template, DNase-free water was added in the same amount. The PCR conditions were as follows: 95 °C, 5 min; 30 cycles of 95 °C, 30 s; 59 °C, 30 s; 72 °C, 6 s; and 72 °C, 5 min. The results were visualized using UV light (Syngene®).

Table 9.
List of primers used for the distinction of phylogroups A or C.
To distinguish between E or D, or E or I clade, 2.4 µL of template was used for PCR in a final volume of 20 µL, containing PCR master mix multiplex PCR NZTYtaq 2x Green Master Mix (NZYTech), arpA (2 μM) (Eurofins Genomics) as described in Table 10 and DNase-free water were used. DNA extraction was performed as previously described. For the PCR reaction, were considered the number of samples to be validated, a positive and a negative control. E. coli O157:H7 (arpA+) was used as a positive control. As a negative control, in place of the template, DNase-free water was added in the same amount. The PCR conditions were as follows: 95 °C 5 min; 30 cycles of 95 °C, 30 s; 57 °C, 30 s; 72 °C, 9 s; and 72 °C, 5 min. The results were visualized using UV light (Syngene®).

Table 10.
List of primers used for the distinction of phylogroups E or D, or E or I clade.
For the elaboration and determination of the pathotypes (ETEC, EIEC, EAEC, EPEC, EHEC/STEC) the primers (Eurofins Genomics) designated by Schmidt et al. [80], Aranda et al. [81] and ISO/TS 13136 [82] were used. DNA extraction was performed as previously described. As positive controls E. coli IH2859f (eae+; bfp+), E. coli LMV_E_37 (eae+; bfp+), E. coli LMV_E_38 (est+), E. coli LMV_E_39 (12 et+), E. coli LMV_E_40 (ipaH+), E. coli LMV_E_41 (aggr+; cvd432+), E. coli O157.34 (eae+; stx1+; stx2+), E. coli O157.156 (eae+; stx1+; stx2+) and E. coli O157.157 (eae+; stx1+; stx2+) were used. As a negative control, in place of the template, DNase-free water was added in the same amount. All gene amplifications were obtained by multiplex PCR. For all assays, the master mix used was the multiplex PCR NZTYtaq 2x Green Master mix (NZYTech). For all assays, the total reaction volume was 20 μL. The list of primers used to determine each pathotype, their concentrations, amplification conditions, and volume per reaction are described in Table 11 and Table 12. The results were visualized using UV light (Syngene®).

Table 11.
List of primers used for the determination of pathotypes.

Table 12.
List of concentrations of each primer, amplification conditions.
5.3. Antimicrobial Susceptibility of E. coli Isolates
Antimicrobial susceptibility was tested by disk-diffusion method on Mueller–Hinton agar (Mha) (Bio-Rad). Briefly, a suspension of E. coli in saline solution (NaCl 0.85%) with a 0.5 McFarland turbidity was prepared. A sterile swab was dipped into this suspension, swirled well against the walls of the tube to remove excess solution, and used to inoculated by streaking Mha (Bio-Rad) plate. The disks of the antimicrobials were placed onto the surface and lightly pressed. Plates were incubated at 37 °C for 20 ± 4 h. At the end of incubation, the inhibition zone diameters were measured with a ruler. The antibiotics tested were as follows: AMP (10 μg) (Bio-Rad), CTX (5 μg) (Bio-Rad), CAZ (10 μg) (Bio-Rad), MEM (10 μg) (Oxoid, Basingstoke, UK), NAL (30 μg) (Oxoid), CIP (5 μg) (Oxoid), GEN (10 μg) (Bio-Rad), AZM (15 μg) (Oxoid), TE (30 μg) (Oxoid), TMP (5 μg) (Bio-Rad), CHL (30 μg) (Bio-Rad), and SULF (300 μg) (Oxoid). The breakpoints for AMP, CTX, CAZ, MEM, NAL, CIP, GEN, AZM, TE, TMP and CHL fitted the susceptibility profile according to EUCAST [56] parameters. SULF fitted the susceptibility profile according to CLSI [57] since EUCAST does not show breakpoint values. Reference strains E. coli ATCC 25922, S. aureus ATCC 25923, and P. aeruginosa ATCC 27853 were adopted as control strains. The results of the susceptibility profile are presented in break intervals defined in two categories (S—susceptible and R—resistant).
5.4. Multiresistant Isolates of E. coli Assessment
According to Magiorakos et al. [83], to characterize MDR bacteria it is necessary that the bacteria show an acquired non-susceptibility to at least one agent in three or more antimicrobial groups. The antimicrobial groups considered were penicilins, cephalosporins, carbapenems, fluoroquinolones, aminoglycosides, macrolides, tetracyclines, miscellaneous agents, and sulfonamides. Isolates that showed resistance to at least three different antimicrobial groups were classified as MDR.
5.5. Detection of Extended-Spectrum β-Lactamase Resistance Genes
The isolates that exhibited resistance to CAZ were analyzed by multiplex PCR using primers described in Table 13, targeting genes blaCTX-M, blaTEM, and blaSHV [84,85,86]. The DNA template was obtained as described in 4.2; 2 µL of the DNA suspension was used for PCR in a final volume of 20 µL, containing 1.5 mM MgCl2, 1x Taq buffer, 0.25 μM primers, 1.25 U Taq (NZYTech), 0.25 mM dNTPs (NZYTech), and DNase-free water. The PCR conditions were as follows, according to Oliveira et al. [87]: 95 °C, 5 min; 30 cycles of 94 °C, 30 s; 60 °C, 30 s; 72 °C, 30 s; and 72 °C, 5 min. A 2% agarose gel (NZYTech) in 1x TAE (NZYTech) was prepared containing 5 µL/100 mL of GreenSafe Premium (NZYTech). The gel was loaded with the reaction products: 4 μL of 6x NZYDNA loading dye (NZYTech) was added to the amplicons that did not contain a loading dye. E. coli R02 (blaCTX-M+, blaTEM+); E. coli H1015 (blaSHV-12+); E. coli H1043 (blaCTX-M-I+); E. coli H1046 (blaCTX-M-II+, blaTEM+), and E. coli H995 (blaCTX-M-IX+, blaTEM+) were used as positive controls. As a negative control, in place of the template, DNase-free water was added in the same amount. Each corresponding well was loaded with 8 μL of the reaction obtained after amplification and with 5 μL V-marker for ESBL (NZYTech) and finally the gel ran in 1x TAE (NZYTech) at 90 volts ± 10 volts for 1 h and visualized using UV light (Syngene® GeneFlash system).

Table 13.
List of primers used for detection of extended-spectrum β-lactamase resistance genes.
5.6. Data Analysis
Descriptive statistics (mean, standard deviation (SD)) were generated for all of the variables in the dataset. Analysis of variance was performed on microbiological load of E. coli in TBX, TBX supplemented with ampicillin (TBXamp), or with enrofloxacin (TBXenro), for breeding hens and egg laying hens’ feces’ samples data, using the IBM SPSS Statistics 23.0 for Windows [88]. The analysis was carried out using a t-test of independent samples with a variable grouping of sample sources (breeding hens and egg laying hens). All statements of significance were based on testing at the p < 0.05 level.
Author Contributions
Conceptualization, G.A., I.M.A. and L.R.; methodology, G.A., M.C. and S.P.; writing—original draft preparation, A.R.B. and G.A.; writing—review and editing, A.R.B., M.C., I.M.A., L.R. and G.A.; supervision, G.A.; project administration, G.A. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The data presented in this study are available on request from the corresponding author.
Acknowledgments
The authors would like to thank CECA-ICETA UIDB/00211/2020 for their support. The authors are deeply grateful to Stefano Morabito from The European Union Reference Laboratory (EURL) for Escherichia coli and to Jorge Blanco from Laboratorio de Referencia de E. coli (LREC), Facultad de Veterinaria de Lugo, Spain for providing the strains that were used as controls in each PCR reaction batch. The authors would also like to thank Bio-Rad Laboratories for their support in providing some of the antibiotics used in this study.
Conflicts of Interest
The authors declare no conflict of interest.
References
- McAfee, A.J.; McSorley, E.M.; Cuskelly, G.J.; Moss, B.W.; Wallace, J.M.W.; Bonham, M.P.; Fearon, A.M. Red meat consumption: An overview of the risks and benefits. Meat Sci. 2010, 84, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Abebe, E.; Gugsa, G.; Ahmed, M. Review on Major Food-Borne Zoonotic Bacterial Pathogens. J. Trop Med. 2020, 2020, 4674235. [Google Scholar] [CrossRef]
- Hughes, C.; Gillespie, I.A.; O’Brien, S.J. Foodborne transmission of infectious intestinal disease in England and Wales, 1992–2003. Food Control 2007, 18, 766–772. [Google Scholar] [CrossRef]
- Algammal, A.M.; Hetta, H.F.; Batiha, G.E.; Hozzein, W.N.; El Kazzaz, W.M.; Hashem, H.R. Virulence-determinants and antibiotic-resistance genes of MDR-E. coli isolated from secondary infections following FMD-outbreak in cattle. Sci. Rep. 2020, 10, 19779. [Google Scholar] [CrossRef] [PubMed]
- Tyson, G.H.; Li, C.; Hsu, C.H.; Jones, S.B.; McDermott, P.F. Diverse fluoroquinolone resistance plasmids from retail meat E. coli in the United States. Front. Microbiol. 2019, 10, 2826. [Google Scholar] [CrossRef]
- Poole, T.L.; Callaway, T.R.; Norman, K.N.; Scott, H.M.; Loneragan, G.H.; Ison, S.A. Transferability of antimicrobial resistance from multidrug-resistant Escherichia coli isolated from cattle in the USA to E. coli and Salmonella Newport recipients. J. Glob. Antimicrob. Resist. 2017, 11, 123–132. [Google Scholar] [CrossRef]
- Tadesse, D.A.; Zhao, S.; Tong, E.; Ayers, S.; Singh, A.; Bartholomew, M.J.; McDermott, P.F. Antimicrobial Drug Resistance in Escherichia coli from humans and food animals, United States, 1950–2002. Emerg. Infect. Dis. 2012, 18, 741–749. [Google Scholar] [CrossRef]
- van Elsas, J.; Semenov, A.; Costa, R. Survival of Escherichia coli in the environment: Fundamental and public health aspects. ISME J. 2011, 5, 173–183. [Google Scholar] [CrossRef]
- Adzitey, F. Incidence and antimicrobial susceptibility of Escherichia coli isolated from beef (meat muscle, liver and kidney) samples in Wa Abattoir, Ghana. Cogent. Food Agric. 2020, 6, 2–10. [Google Scholar] [CrossRef]
- Gupta, P.; Adhikari, A. Novel approaches to environmental monitoring and control of Listeria monocytogenes in food production facilities. Foods 2022, 11, 1760. [Google Scholar] [CrossRef]
- Caudry, S.D.; Stanisich, V.A. Incidence of antibiotic-resistant Escherichia coli associated with frozen chicken carcasses and characterization of conjugative R plasmids derived from such strains. Antimicrob. Agents Chemother. 1979, 16, 701–709. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Nazer, A.H. Transmissible drug resistance in Escherichia coli isolated from poultry and their carcasses in Iran. Cornell Vet. 1980, 70, 365–371. [Google Scholar] [PubMed]
- Bensink, J.C.; Botham, F.P. Antibiotic resistant coliform bacilli, isolated from freshly slaughtered poultry and from chilled poultry at retail outlets. Aust. Vet. J. 1983, 60, 80–83. [Google Scholar] [CrossRef] [PubMed]
- Ojeniyi, A.A. Direct transmission of Escherichia coli from poultry to humans. Epidemiol. Infect. 1989, 103, 513–522. [Google Scholar] [CrossRef]
- Chaslus-Dancla, E.; Lafont, J.P. IncH plasmids in Escherichia coli strains isolated from broiler chicken carcasses. Appl. Environ. Microbiol. 1985, 49, 1016–1018. [Google Scholar] [CrossRef]
- Jayaratne, A.H.; Collins-Thompson, D.L.; Trevors, J.T. Occurrence of aminoglycoside phosphotransferase subclass I and II structural genes among Enterobacteriaceae spp. Isolated from meat samples. Appl. Microbiol. Biotechnol. 1990, 33, 547–552. [Google Scholar] [CrossRef]
- Turtura, G.C.; Massa, S.; Ghazvinizadeh, H. Antibiotic resistance among coliform bacteria isolated from carcasses of commercially slaughtered chickens. Int. J. Food Microbiol. 1990, 11, 351–354. [Google Scholar] [CrossRef]
- Lakhotia, R.L.; Stephens, J.F. Drug resistance and R factors among enterobacteria isolated from eggs. Poult. Sci. 1973, 52, 1955–1962. [Google Scholar] [CrossRef]
- van den Bogaard, A.E.; London, N.; Driessen, C.; Stobberingh, E.E. Antibiotic resistance of faecal Escherichia coli in poultry, poultry farmers and poultry slaughterers. J. Antimicrob. Chemother. 2001, 47, 763–771. [Google Scholar] [CrossRef]
- Kaneko, K.I.; Hayashidani, H.; Ohtomo, Y. Bacterial contamination of ready-to-eat foods and fresh products in retail shops and food factories. J. Food Prot. 1999, 62, 644–649. [Google Scholar] [CrossRef]
- Guzmán-Gómez, G.; Valdovinos, M.; Díaz, E.; Montaño, J.; Valle, M.; Vitela, M.; Quezada, S. Frequency of Salmonella and Listeria monocytogenes in Five Commercial Brands of Chicken Eggs Using a Combined Method of Enrichment and Nested-PCR. J. Food Prot. 2013, 76, 429–434. [Google Scholar] [CrossRef] [PubMed]
- Gharbi, M.; Béjaoui, A.; Ben Hamda, C.; Alaya, N.; Hamrouni, S.; Bessoussa, G.; Ghram, A.; Maaroufi, A. Campylobacter spp. In Eggs and Laying Hens in the North-East of Tunisia: High Prevalence and Multidrug-Resistance Phenotypes. Vet. Sci. 2022, 9, 108. [Google Scholar] [CrossRef] [PubMed]
- Kapena, M.S.; Muma, J.B.; Mubita, C.M.; Munyeme, M. Antimicrobial resistance of Escherichia coli and Salmonella in raw retail table eggs in Lusaka, Zambia. Vet. World 2020, 13, 2528–2533. [Google Scholar] [CrossRef] [PubMed]
- Mehdi, Y.; Montminy, M.P.L.; Gaucher, M.L.; Chorfi, Y.; Suresh, G.; Rouissi, T.; Brar, S.K.; Côté, C.; Ramirez, A.A.; Godbout, S. Use of antibiotics in broiler production: Global impacts and alternatives. Anim. Nutr. 2018, 4, 170–178. [Google Scholar] [CrossRef]
- Singh, P.; Karimi, A.; Devendra, K.; Waldroup, P.W.; Cho, K.K.; Kwon, Y.M. Influence of penicillin on microbial diversity of the cecal microbiota in broiler chickens. Poult. Sci. 2013, 92, 272–276. [Google Scholar] [CrossRef]
- O’Brien, T.F. Emergence, spread, and environmental effect of antimicrobial resistance: How use of an antimicrobial anywhere can increase resistance to any antimicrobial anywhere else. Clin. Infect. Dis. 2002, 34, S78–S84. [Google Scholar] [CrossRef]
- Wolny-Koładka, K.; Zdaniewicz, M. Antibiotic Resistance of Escherichia coli Isolated from Processing of Brewery Waste with the Addition of Bulking Agents. Sustainability 2021, 13, 10174. [Google Scholar] [CrossRef]
- Kluytmans, J.A.J.W.; Overdevest, I.T.M.A. Extended-spectrum beta-lactamase-producing Escherichia coli from retail chicken meat and humans: Comparison of strains, plasmids, resistance genes, and virulence factors. Clin. Infect. Dis. 2013, 56, 478–487. [Google Scholar] [CrossRef]
- Zachary, R.; Stromberg, Z.R.; Johnson, J.R.; Fairbrother, J.M.; Kilbourne, J.; Van Goor, A.; Curtiss, R.; Mellata, M. Evaluation of Escherichia coli isolates from healthy chickens to determine their potential risk to poultry and human health. PloS ONE 2017, 12, e0180599. [Google Scholar] [CrossRef]
- Apata, D.F. Antibiotic Resistance in Poultry. Int. J. Poult. Sci. 2009, 8, 404–408. [Google Scholar] [CrossRef]
- Barton, M.D. Antibiotic use in animal feed and its impact on human healt. Nutr. Res. Rev. 2000, 13, 279–299. [Google Scholar] [CrossRef] [PubMed]
- Bond, V.; Jewell, J. The Impacts of Antibiotic Use in Animals on Human Health and Animal Welfare; Investor Briefing No. 17; Business Benchmark on Farm Animal Welfare, 2014; pp. 1–9. Available online: https://www.bbfaw.com/media/1070/briefing-17-impacts-of-antibiotic-use-in-animals-on-human-health-and-animal-welfare.pdf (accessed on 23 November 2022).
- DGAV. Available online: https://www.dgav.pt/wp-content/uploads/2021/09/ESVAC-RELATORIO-2017.pdf (accessed on 23 November 2022).
- Tenaillon, O.; Skurnik, D.; Picard, B.; Denamur, E. The population genetics of commensal Escherichia coli. Nat. Rev. Microbiol. 2010, 8, 207–217. [Google Scholar] [CrossRef] [PubMed]
- Clermont, O.; Bonacorsi, S.; Bingen, E. Rapid and simple determination of the Escherichia coli phylogenetic group. Appl. Environ. Microbiol. 2000, 66, 4555–4558. [Google Scholar] [CrossRef] [PubMed]
- Doumith, M.; Day, M.J.; Hope, R.; Wain, J.N.; Woodford, N. Improved multiplex PCR strategy for rapid assignment of the four major Escherichia coli phylogenetic groups. J. Clin. Microbiol. 2012, 50, 3108–3110. [Google Scholar] [CrossRef]
- Katongole, P.; Kisawuzi, D.B.; Bbosa, H.K.; Kateete, D.P.; Najjuka, C.F. Phylogenetic groups and antimicrobial susceptibility patterns of uropathogenic Escherichia coli clinical isolates from patients at Mulago National Referral Hospital, Kampala, Uganda. F1000Research 2019, 8, 1828. [Google Scholar] [CrossRef]
- Clermont, O.; Christenson, J.K.; Denamur, E. The Clermont Escherichia coli phylotyping method revisited: Improvement of specificity and detection of new phylogroups. Environ. Microbiol. Rep. 2013, 5, 58–65. [Google Scholar] [CrossRef]
- Clermont, O.; Gordon, D.; Denamur, E. Guide to the various phylogenetic classification schemes for Escherichia coli and the correspondence among schemes. Microbiology 2015, 161, 980–988. [Google Scholar] [CrossRef]
- Santos, Y.P.A.; Flores, M.E.B.; Camacho, S.P.D.; Beltrán, M.J.U.; Campos, C.A.E.; Unda, J.R.P. Association of phylogenetic distribution and presence of integrons with multidrug resistance in Escherichia coli clinical isolates from children with diarrhoea. J. Infect. Public Health 2020, 13, 767–772. [Google Scholar] [CrossRef]
- Barzan, M.; Rad, M.; Tabar, G.R.H.; Azizzadeh, M. Phylogenetic analysis of Escherichia coli isolates from healthy and diarrhoeic calves in Mashhad, Iran. Bulg. J. Vet. Med. 2017, 20, 11–18. [Google Scholar] [CrossRef]
- Bélanger, L.A.; Garenaux, J.; Harel, M.; Boulianne, E.; Dozois, C. Escherichia coli from animal reservoirs as a potential source of human extraintestinal pathogenic Escherichia coli. FEMS Immunol. Med. Microbiol. 2011, 62, 1–10. [Google Scholar] [CrossRef]
- Croxall, G.; Hale, J.; Weston, V.; Manning, G.; Cheetham, P.; Achtman, M.; McNally, A. Molecular epidemiology of extraintestinal pathogenic Escherichia coli isolates from a regional cohort of elderly patients highlights the prevalence of ST131 strains with increased antimicrobial resistance in both community and hospital care settings. J. Antimicrob. Chemother. 2011, 66, 2501–2508. [Google Scholar] [CrossRef] [PubMed]
- Kaper, J.B.; Nataro, J.P.; Mobley, H.L.T. Pathogenic Escherichia coli. Nat. Rev. Microbiol. 2004, 2, 123–140. [Google Scholar] [CrossRef] [PubMed]
- Bekal, S.; Brousseau, R.; Masson, L.; Préfontaine, G.; Fairbrother, J.M.; Harel, J. Rapid identification of Escherichia coli pathotypes by virulence gene detection with DNA microarray. J. Clin. Microbiol. 2003, 41, 2113–2125. [Google Scholar] [CrossRef] [PubMed]
- Caine, L.A.; Nwodo, U.U.; Okoh, A.I.; Ndip, R.N.; Green, E. Occurrence of Virulence Genes Associated with Diarrheagenic Escherichia coli Isolated from Raw Cow’s Milk from Two Commercial Dairy Farms in the Eastern Cape Province, South Africa. Int. J. Environ. Res. Public Health 2014, 11, 11950–11963. [Google Scholar] [CrossRef] [PubMed]
- Murase, T.; Ozaki, H. Relationship between phylogenetic groups of Escherichia coli and Pathogenicity among Isolates from chickens with Colibacillosis and healthy chickens. Poult Sci. 2022, 101, 102007. [Google Scholar] [CrossRef] [PubMed]
- Pavlovic, M.; Luze, A.; Konrad, R.; Berger, A.; Sing, A.; Busch, U.; Huber, I. Development of a duplex real-time PCR for differentiation between E. coli and Shigella spp. J. Appl. Microbiol. 2011, 110, 1245–1251. [Google Scholar] [CrossRef] [PubMed]
- Zenati, F.; Barguigua, A.; Nayme, K.; Benbelaïd, F.; Khadir, A.; Bellahsene, C.; Bendahou, M.; Hafida, H.; Timinouni, M. Characterization of uropathogenic ESBL-producing Escherichia coli isolated from hospitalized patients in western Algeria. J. Infect. Dev. Ctries. 2019, 13, 291–302. [Google Scholar] [CrossRef]
- Alonso, C.A.; Zarazaga, M.; Ben Sallem, R.; Jouini, A.; Ben Slama, K.; Torres, C. Antibiotic resistance in Escherichia coli in husbandry animals: The African perspective. Lett. Appl. Microbiol. 2007, 64, 318–334. [Google Scholar] [CrossRef]
- Riaño, I.; Moreno, M.A.; Teshager, T.; Sáenz, Y.; Dominguez, L.; Torres, C. Detection and characterization of extended-spectrum beta-lactamases in Salmonella enterica strains of healthy food animals in Spain. J. Antimicrob. Chemother. 2006, 58, 844–847. [Google Scholar] [CrossRef]
- Meguenni, N.; Le Devendec, L.; Jouy, E.; Le Corvec, M.; Bounar-Kechih, S.; Bakour, R.; Kempf, I. First description of an extended-spectrum cephalosporin- and fluoroquinolone- resistant avian pathogenic Escherichia coli clone in Algeria. Avian Dis. 2015, 59, 20–23. [Google Scholar] [CrossRef]
- Belmahdi, M.; Bakour, S.; Al Bayssari, C.; Touati, A.; Rolain, J.M. Molecular Characterization of extended-spectrum β-lactamase- and plasmid AmpC-producing Escherichia coli strains isolated from broilers in Béjaïa, Algeria. J. Glob. Antimicrob. Resist. 2016, 6, 108–112. [Google Scholar] [CrossRef] [PubMed]
- Maamar, E.; Hammami, S.; Alonso, C.A.; Dakhli, N.; Abbassi, M.S.; Ferjani, S.; Hamzaoui, Z.; Saidani, M.; Torres, C.; Boutiba-Ben Boubaker, I. High prevalence of extended-spectrum and plasmidic AmpC beta-lactamase-producing Escherichia coli from poultry in Tunisia. Int. J. Food Microbiol. 2016, 231, 69–75. [Google Scholar] [CrossRef] [PubMed]
- Casella, T.; Nogueira, M.C.L.; Saras, E.; Haenni, M.; Madec, J.Y. High prevalence of ESBLs in retail chicken meat despite reduced use of antimicrobials in chicken production, France. Int. J. Food Microbiol. 2017, 257, 271–275. [Google Scholar] [CrossRef] [PubMed]
- The European Committee on Antimicrobial Susceptibility Testing. Breakpoint Tables for Interpretation of MICs and Zone Diameters. Version 12.0. 2022. Available online: http://www.eucast.org (accessed on 10 October 2022).
- Clinical and Laboratory Standards Institute (CLSI). Performance Standards for Antimicrobial Susceptibility Testing, 32nd ed.; CLSI Supplement M100; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2022; ISSN1 978-1-68440-134-5. ISSN2 978-1-68440-135-2. [Google Scholar]
- Clermont, O.; Dixit, O.; Vangchhia, B.; Condamine, B.; Dion, S.; Bridier-Nahmias, A.; Denamur, E.; Gordon, D. Characterization and rapid identification of phy- logroup G in Escherichia coli, a lineage with high virulence and antibiotic resistance potential. Environ. Microbiol 2019, 21, 3107–3117. [Google Scholar] [CrossRef]
- Bingen, E.; Picard, B.; Brahimi, N.; Mathy, S.; Desjardins, P.; Elion, J.; Denamur, E. Phylogenetic analysis of Escherichia coli strains causing neonatal meningitis suggests horizontal gene transfer from a predominant pool of highly virulent B2 group strains. J. Infect. Dis. 1998, 177, 642–650. [Google Scholar] [CrossRef]
- Picard, B.; Garcia, J.S.; Gouriou, S.; Duriez, P.; Brahimi, N.; Bingen, E.; Elion, J.; Denamur, E. The link between phylogeny and virulence in Escherichia coli extraintestinal infection. Infect. Immun. 1999, 67, 546–553. [Google Scholar] [CrossRef]
- Obeng, A.S.; Rickard, H.; Ndi, O.; Sexton, M.; Barton, M. Antibiotic resistance, phylogenetic grouping and virulence potential of Escherichia coli isolated from the faeces of intensively farmed and free range poultry. Vet. Microbiol. 2012, 154, 305–315. [Google Scholar] [CrossRef]
- Hayashi, W.; Ohsaki, Y.; Taniguchi, Y.; Koide, S.; Kawamura, K.; Suzuki, M.; Kimura, K.; Wachino, J.; Nagano, Y.; Arakawa, Y.; et al. High prevalence of blaCTX-M-14 among genetically diverse Escherichia coli recovered from retail raw chicken meat portions in Japan, Int. J. Food Microbiol. 2018, 284, 98–104. [Google Scholar] [CrossRef]
- Projahn, M.; Daehre, K.; Roesler, U.; Friese, A. Extended-spectrum-beta-lactamase- and plasmid-encoded cephamycinase-producing Enterobacteria in the broiler hatchery as a potential mode of pseudo-vertical transmission. Appl. Environ. Microbiol. 2016, 83, e02364-16. [Google Scholar] [CrossRef]
- Adefioye, O.J.; Weinreich, J.; Rödiger, S.; Schierack, P.; Olowe, O.A. Phylogenetic Characterization and Multilocus Sequence Typing of Extended-Spectrum Beta Lactamase-Producing Escherichia coli from Food-Producing Animals, Beef, and Humans in Southwest Nigeria. Microb. Drug Resist. 2021, 27, 111–120. [Google Scholar] [CrossRef]
- Bengtsson-Palme, J.; Kristiansson, E.; Larsson, D.G.J. Environmental factors influencing the development and spread of antibiotic resistance. FEMS Microbiol. Rev. 2018, 42, fux053. [Google Scholar] [CrossRef] [PubMed]
- Hossain, T.; Rafiq, K.; Islam, Z.; Chowdhury, S.; Islam, P.; Haque, Z.; Samad, M.A.; Sani, A.A.; Ferdous, M.R.A.; Islam, R.; et al. A survey on knowledge, attitude, and practices of large-animal farmers towards antimicrobial use, resistance, and residues in Mymensingh division of Bangladesh. Antibiotics 2022, 11, 442. [Google Scholar] [CrossRef] [PubMed]
- Javadi, A.; Khatibi, S.A. Effect of commercial probiotic (Protexin®) on growth, survival and microbial quality of shrimp (Litopenaeus vannamei). Nutr. Food Sci. 2017, 47, 204–216. [Google Scholar] [CrossRef]
- Hailu, W.; Helmy, Y.A.; Carney-Knisely, G.; Kauffman, M.; Fraga, D.; Rajashekara, G. Prevalence and Antimicrobial Resistance Profiles of Foodborne Pathogens Isolated from Dairy Cattle and Poultry Manure Amended Farms in Northeastern Ohio, the United States. Antibiotics 2021, 10, 1450. [Google Scholar] [CrossRef] [PubMed]
- Langata, L.M.; Maingi, J.M.; Musonye, H.A.; Kiiru, J.; Nyamache, A.K. Antimicrobial resistance genes in Salmonella and Escherichia coli isolates from chicken droppings in Nairobi, Kenya. BMC Res. Notes 2019, 12, 22. [Google Scholar] [CrossRef]
- Abbassi, M.S.; Kilani, H.; Abid, I.; Sáenz, Y.; Hynds, P.; Lengliz, S.; Chehida, N.B.; Boubaker, I.B. Genetic background of antimicrobial resistance in multiantimicrobial-resistant Escherichia coli isolates from feces of healthy broiler chickens in Tunisia. BioMed Res. Int. 2021, 2021, 1269849. [Google Scholar] [CrossRef]
- European Food Safety Authority. Scientific Opinion on the public health risks of bacterial strains producing extended-spectrum β-lactamases and/or AmpC β-lactamases in food and food-producing animals. EFSA J. 2011, 9, 2322. [Google Scholar] [CrossRef]
- Liu, C.; Wang, P.; Dai, Y.; Liu, Y.; Song, Y.; Yu, L.; Feng, C.; Liu, M.; Xie, Z.; Shang, Y.; et al. Longitudinal monitoring of multidrug resistance in Escherichia coli on broiler chicken fattening farms in Shandong, China. Poult. Sci. 2021, 100, 100887. [Google Scholar] [CrossRef]
- Koju, P.; Shrestha, R.; Shrestha, A.; Tamrakar, S.; Rai, A.; Shrestha, P.; Madhup, S.K.; Katuwal, N.; Shrestha, A.; Shrestha, A.; et al. Antimicrobial resistance in E. coli isolated from chicken cecum samples and factors contributing to antimicrobial resistance in Nepal. Trop. Med. Infect. Dis. 2022, 7, 249. [Google Scholar] [CrossRef]
- Huneau-Salaun, A.; Michel, V.; Huonnic, D.; Balaine, L.; Le Bouquin, S. Factors influencing bacterial eggshell contamination in conventional cages, furnished cages and free-range systems for laying hens under commercial conditions. Br. Poult. Sci. 2010, 51, 163–169. [Google Scholar] [CrossRef]
- Egea, P.; López-Cerero, L.; Navarro, M.D. Assessment of the presence of extended-spectrum beta-lactamase-producing Escherichia coli in eggshells and ready-to-eat products. Eur. J. Clin. Microbiol. Infect. Dis. 2011, 30, 1045–1047. [Google Scholar] [CrossRef]
- Machado, E.; Coque, T.M.; Cantón, R.; Sousa, J.C.; Peixe, L. Antibiotic resistance integrons and extended-spectrum {beta}-lactamases among Enterobacteriaceae isolates recovered from chickens and swine in Portugal. J. Antimicrob. Chemother. 2008, 62, 296–302. [Google Scholar] [CrossRef] [PubMed]
- Clemente, L.; Leão, C.; Moura, L.; Albuquerque, T.; Amaro, A. Prevalence and characterization of ESBL/AmpC producing Escherichia coli from fresh meat in Portugal. Antibiotics 2021, 10, 1333. [Google Scholar] [CrossRef] [PubMed]
- Mahmud, Z.H.; Kabir, M.H.; Ali, S.; Moniruzzaman, M.; Imran, K.M.; Nafiz, T.N.; Islam, M.S.; Hussain, A.; Hakim, S.A.I.; Worth, M.; et al. Extended-Spectrum Beta-Lactamase-Producing Escherichia coli in Drinking Water Samples from a Forcibly Displaced, Densely Populated Community Setting in Bangladesh. Front. Public Health 2020, 8, 228. [Google Scholar] [CrossRef]
- ISO 16649-2; Microbiology of Food and Animal Feeding Stuffs—Horizontal Method for the Enumeration of β-glucoronidase-positive Escherichia coli- Part 2: Colony—Count Technique at 44 °C Using 5-bromo-4-chloro-3-indolyl β-D-glucuronide. ISO: Geneva, Switzerland, 2001.
- Schmidt, H.; Knop, C.; Franke, S.; Aleksic, S.; Heesemann, J.; Karch, H. Development of PCR for screening of enteroaggregative Escherichia coli. J. Clin. Microbiol. 1995, 33, 701–705. [Google Scholar] [CrossRef] [PubMed]
- Aranda, K.R.; Fabbricotti, S.H.; Fagundes-Neto, U.; Scaletsky, I.C. Single multiplex assay to identify simultaneously enteropathogenic, enteroaggregative, enterotoxigenic, enteroinvasive and Shiga toxin-producing Escherichia coli strains in Brazilian children. FEMS Microbiol. Lett. 2007, 267, 145–150. [Google Scholar] [CrossRef]
- ISO/TS 13136; Microbiology of Food and Animal Feed—Real-Time Polymerase Chain Reaction (PCR)-Based Method for the Detection of Food-Borne Pathogens—Horizontal Method for the Detection of Shiga Toxin-Producing Escherichia coli (STEC) and the Determination of O157, O111, O26, O103 and O145 Serogroups. ISO: Geneva, Switzerland, 2012.
- Magiorakos, A.P.; Srinivasan, A.; Carey, R.B.; Carmeli, Y.; Falagas, M.E.; Giske, C.G.; Harbarth, S.; Hindler, J.F.; Kahlmeter, G.; Olsson-Liljequist, B.; et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 2012, 18, 268–281. [Google Scholar] [CrossRef]
- Arlet, G.; Rouveau, M.; Philippon, A. Substitution of alanine for aspartate at position 179 in the SHV-6 extended-spectrum beta-lactamase. FEMS Microbiol. Lett. 1997, 152, 163–167. [Google Scholar] [CrossRef] [PubMed]
- Monstein, H.J.; Ostholm-Balkhed, A.; Nilsson, M.V.; Nilsson, M.; Dornbusch, K.; Nilsson, L.E. Multiplex PCR amplification assay for the detection of blaSHV, blaTEM and blaCTX-M genes in Enterobacteriaceae. APMIS 2007, 115, 1400–1408. [Google Scholar] [CrossRef]
- Dallenne, C.; Da Costa, A.; Decré, D.; Favier, C.; Arlet, G. Development of a set of multiplex PCR assays for the detection of genes encoding important beta-lactamases in Enterobacteriaceae. J. Antimicrob. Chemother. 2010, 65, 490–495. [Google Scholar] [CrossRef]
- Oliveira, R.; Castro, J.; Silva, S.; Oliveira, H.; Saavedra, M.J.; Azevedo, N.F.; Almeida, C. Exploring the Antibiotic Resistance Profile of Clinical Klebsiella pneumoniae Isolates in Portugal. Antibiotics 2022, 11, 1613. [Google Scholar] [CrossRef] [PubMed]
- IBM. IBM SPSS Statistics for Windows 2015; IBM Corperation: Armonk, NY, USA, 2015. [Google Scholar]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).