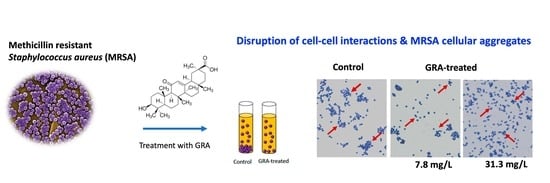
18β-Glycyrrhetinic Acid Induces Metabolic Changes and Reduces Staphylococcus aureus Bacterial Cell-to-Cell Interactions
Abstract
1. Introduction
2. Results
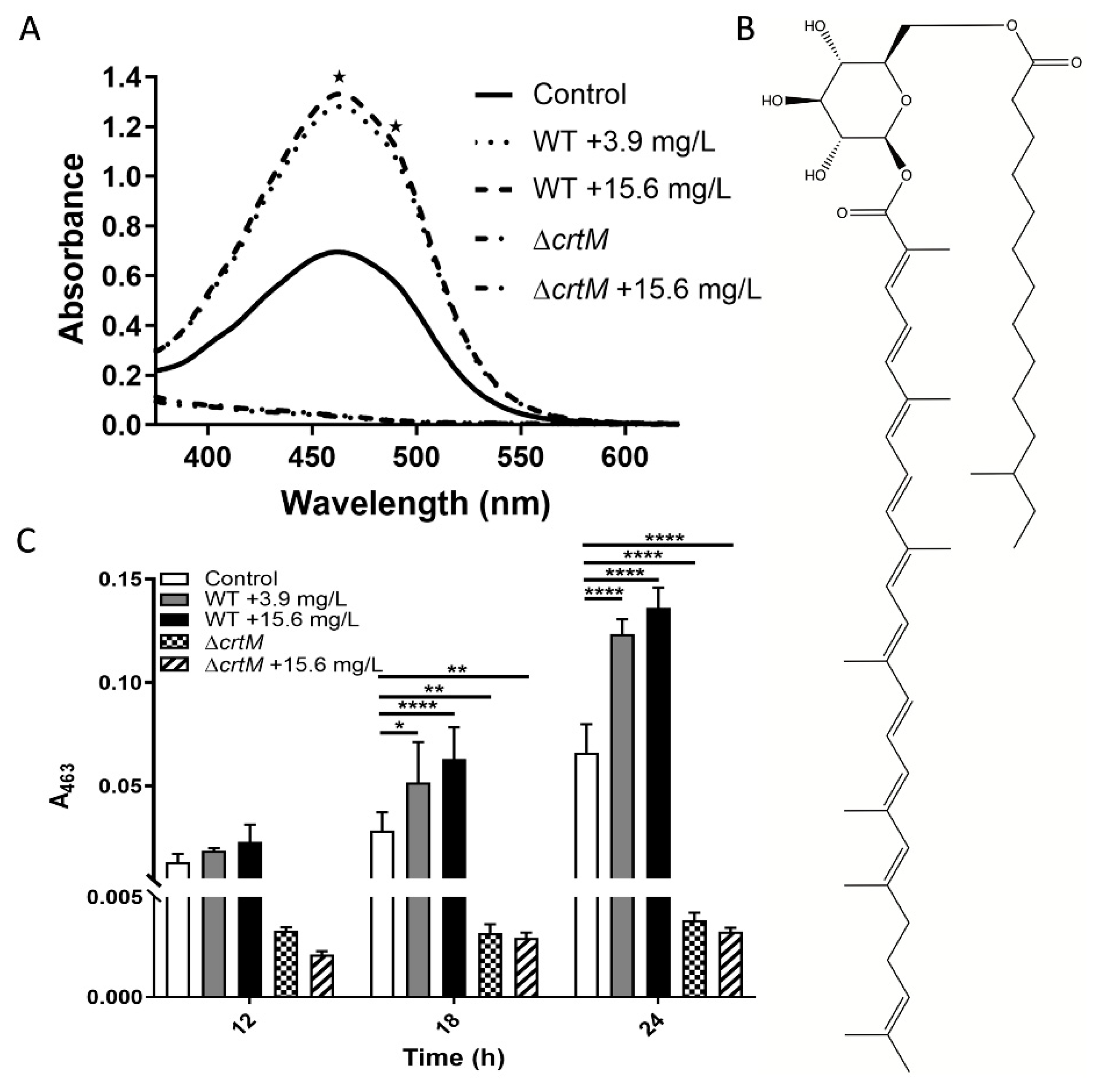
2.1. Increased Pigment Production upon GRA Treatment
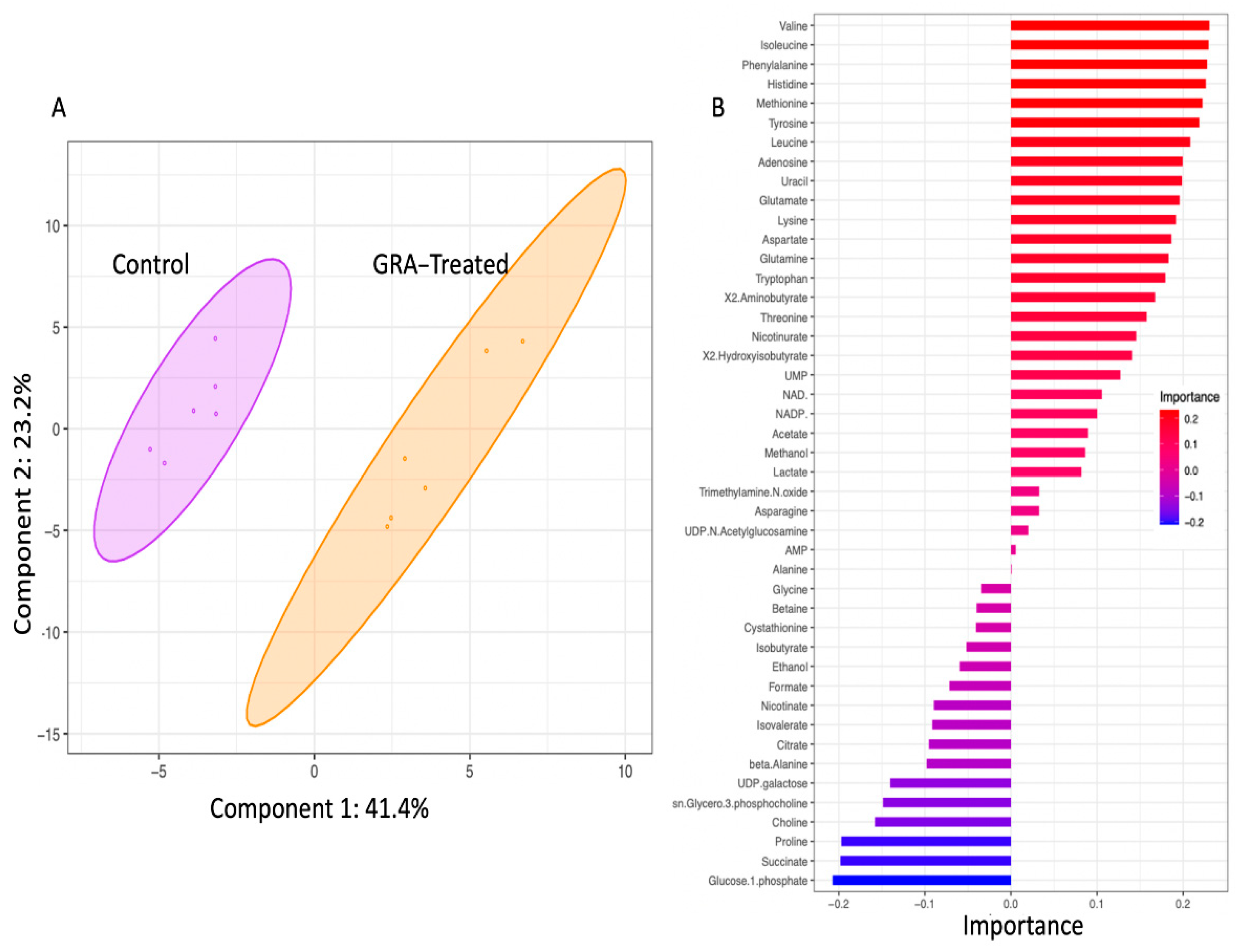
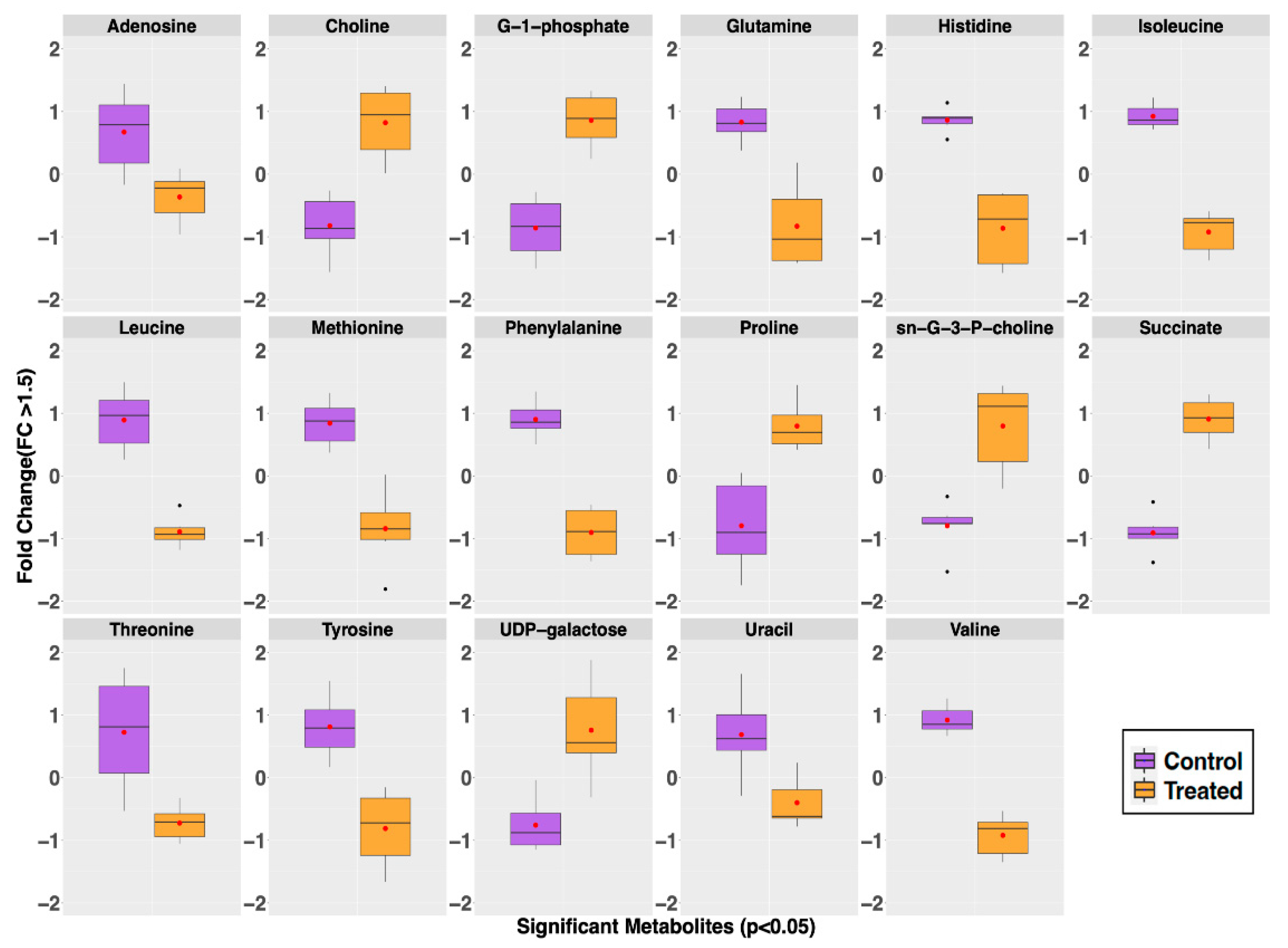
2.2. Metabolite Profiles of S. aureus Treated with GRA
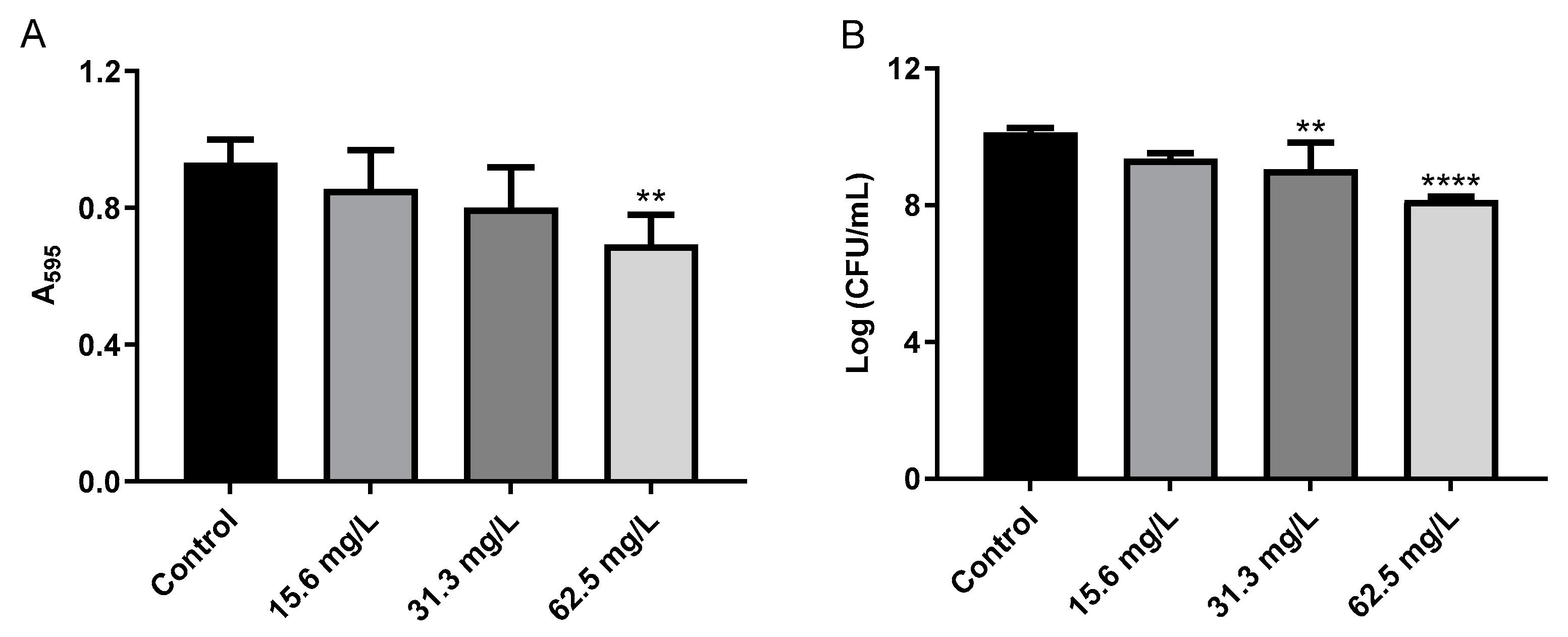
2.3. Effect of GRA on Biofilms and Cell-Surface Adhesion
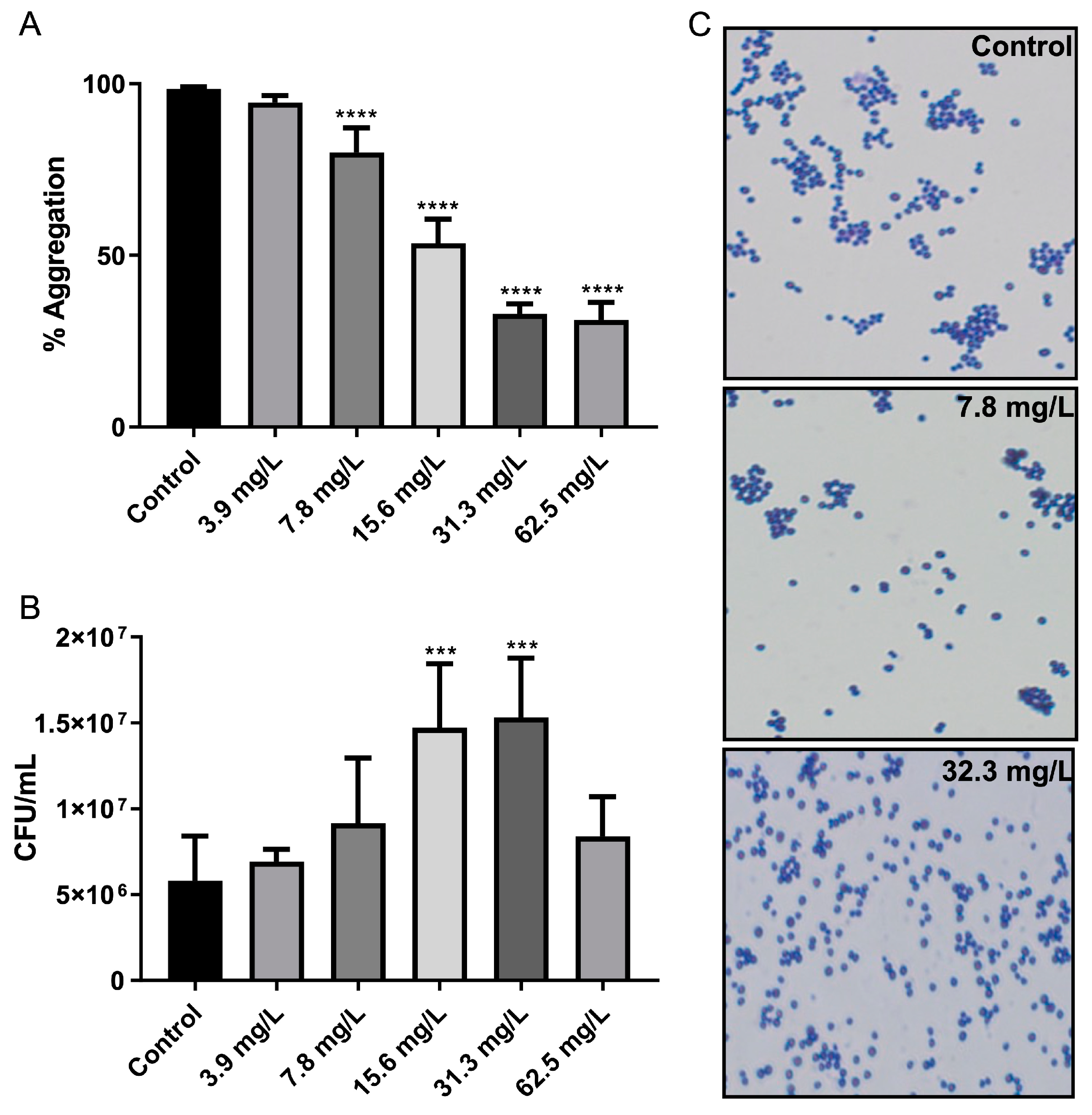
2.4. Bacterial Cell–Cell Aggregation
2.5. Treatment with GRA Reduces Expression of the Staphylococcal Alpha-Hemolysin (hla) Gene
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains
4.2. Stock Preparation of 18-β-Glycyrrhetinic Acid
4.3. Pigmentation Assay
4.4. Metabolic Sample of WT Untreated and Treated with GRA
4.5. 1D 1H NMR Analysis
4.6. NMR Statistical Analysis
4.7. Biofilm Activity Assay
4.8. Cell-Surface Adhesion and Disruption
4.9. Bacteria Cell–Cell Aggregation and Microscopy
4.10. RNA Extraction and Quantitation
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Gualerzi, C.O.B.L.; Fabbretti, A.; Pon, C.L. (Eds.) Antibiotics: Targets, Mechanisms, and Resistance, 1st ed.; Wiley-VCH: Hoboken, NJ, USA, 2013. [Google Scholar]
- Newman, D.J.; Cragg, G.M. Natural products as sources of new drugs over the 30 years from 1981 to 2010. J. Nat. Prod. 2012, 75, 311–335. [Google Scholar] [CrossRef] [PubMed]
- Asl, M.N.; Hosseinzadeh, H. Review of pharmacological effects of Glycyrrhiza sp. and its bioactive compounds. Phytother. Res. 2008, 22, 709–724. [Google Scholar] [CrossRef] [PubMed]
- Gupta, V.K.; Fatima, A.; Faridi, U.; Negi, A.S.; Shanker, K.; Kumar, J.K.; Rahuja, N.; Luqman, S.; Sisodia, B.S.; Saikia, D.; et al. Antimicrobial potential of Glycyrrhiza glabra roots. J. Ethnopharmacol. 2008, 116, 377–380. [Google Scholar] [CrossRef]
- Li, J.Y.; Cao, H.Y.; Liu, P.; Cheng, G.H.; Sun, M.Y. Glycyrrhizic acid in the treatment of liver diseases: Literature review. Biomed. Res. Int. 2014, 2014, 872139. [Google Scholar] [CrossRef]
- Wang, L.; Yang, R.; Yuan, B.; Liu, Y.; Liu, C. The antiviral and antimicrobial activities of licorice, a widely-used Chinese herb. Acta Pharm. Sin. B 2015, 5, 310–315. [Google Scholar] [CrossRef] [PubMed]
- Ploeger, B.; Mensinga, T.; Sips, A.; Seinen, W.; Meulenbelt, J.; DeJongh, J. The pharmacokinetics of glycyrrhizic acid evaluated by physiologically based pharmacokinetic modeling. Drug Metab. Rev. 2001, 33, 125–147. [Google Scholar] [CrossRef] [PubMed]
- Long, D.R.; Mead, J.; Hendricks, J.M.; Hardy, M.E.; Voyich, J.M. 18beta-Glycyrrhetinic acid inhibits methicillin-resistant Staphylococcus aureus survival and attenuates virulence gene expression. Antimicrob. Agents Chemother. 2013, 57, 241–247. [Google Scholar] [CrossRef]
- Lowy, F.D. Staphylococcus aureus infections. N. Engl. J. Med. 1998, 339, 520–532. [Google Scholar] [CrossRef]
- Boucher, H.W.; Talbot, G.H.; Bradley, J.S.; Edwards, J.E.; Gilbert, D.; Rice, L.B.; Scheld, M.; Spellberg, B.; Bartlett, J. Bad bugs, no drugs: No ESKAPE! An update from the Infectious Diseases Society of America. Clin. Infect. Dis. 2009, 48, 1–12. [Google Scholar] [CrossRef]
- Rice, L.B. Federal funding for the study of antimicrobial resistance in nosocomial pathogens: No ESKAPE. J. Infect. Dis. 2008, 197, 1079–1081. [Google Scholar] [CrossRef]
- Archer, N.K.; Mazaitis, M.J.; Costerton, J.W.; Leid, J.G.; Powers, M.E.; Shirtliff, M.E. Staphylococcus aureus biofilms: Properties, regulation, and roles in human disease. Virulence 2011, 2, 445–459. [Google Scholar] [CrossRef] [PubMed]
- de Breij, A.; Karnaoukh, T.G.; Schrumpf, J.; Hiemstra, P.S.; Nibbering, P.H.; van Dissel, J.T.; de Visser, P.C. The licorice pentacyclic triterpenoid component 18beta-glycyrrhetinic acid enhances the activity of antibiotics against strains of methicillin-resistant Staphylococcus Aureus. Eur. J. Clin. Microbiol. Infect. Dis. 2016, 35, 555–562. [Google Scholar] [CrossRef] [PubMed]
- Tounta, V.L.Y.; Cheyne, A.; Larrouy-Maumus, G. Metabolomics in Infectious Diseases and Drug Discovery. Mol. Omics 2021, 17, 376–393. [Google Scholar] [CrossRef] [PubMed]
- Kok, M.; Maton, L.; van der Peet, M.; Hankemeier, T.; van Hasselt, C.J.G. Unraveling antimicrobial resistance using metabolomics. Drug Discov. Today 2022, 27, 1774–1783. [Google Scholar] [CrossRef]
- Bauermeister, A.; Mannochio-Russo, H.; Costa-Lotufo, L.V.; Jarmusch, A.K.; Dorrestein, P.C. Mass spectrometry-based metabolomics in microbiome investigations. Nat. Rev. Microbiol. 2022, 20, 143–160. [Google Scholar] [CrossRef]
- Moco, S. Studying Metabolism by NMR-Based metabolomics. Front. Mol. Biosci. 2022, 9, 882487. [Google Scholar] [CrossRef]
- Edison, A.S.; Colonna, M.; Gouveia, G.J.; Holderman, N.R.; Judge, M.T.; Shen, X.; Zhang, S. NMR: Unique Strengths that Enhance Modern Metabolomics Research. Anal. Chem. 2021, 93, 478–499. [Google Scholar] [CrossRef]
- Pelz, A.; Wieland, K.P.; Putzbach, K.; Hentschel, P.; Albert, K.; Gotz, F. Structure and biosynthesis of staphyloxanthin from Staphylococcus aureus. J. Biol. Chem. 2005, 280, 32493–32498. [Google Scholar] [CrossRef]
- Song, Y.; Liu, C.I.; Lin, F.Y.; No, J.H.; Hensler, M.; Liu, Y.L.; Jeng, W.Y.; Low, J.; Liu, G.Y.; Nizet, V.; et al. Inhibition of staphyloxanthin virulence factor biosynthesis in Staphylococcus aureus: In vitro, in vivo, and crystallographic results. J. Med. Chem. 2009, 52, 3869–3880. [Google Scholar] [CrossRef]
- Caiazza, N.C.; O’Toole, G.A. Alpha-toxin is required for biofilm formation by Staphylococcus aureus. J. Bacteriol. 2003, 185, 3214–3217. [Google Scholar] [CrossRef]
- Anderson, M.J.; Lin, Y.-C.; Gillman, A.N.; Parks, P.J.; Schlievert, P.M.; Perterson, M.L. Alpha toxin promotes Staphylococcus aureus mucosal biofilm formation. Front. Cell. Infect. Microbiol. 2012, 2, 64. [Google Scholar] [CrossRef] [PubMed]
- den Reijer, P.M.; Haisma, E.M.; Lemmens-den Toom, N.A.; Koning, R.A.; Demmers, J.A.A.; Dekkers, D.H.W.; Rijkers, E.; El Ghalbzouri, A.; Nibbering, P.H.; van Wamel, W. Detection of alpha-toxin and other virulence factors in biofilms of Staphylococcus aureus on polystyrene and a human epidermal model. PLoS ONE 2016, 11, e0145722. [Google Scholar] [CrossRef]
- Flemming, H.C.; Wingender, J. The biofilm matrix. Nat. Rev. Microbiol. 2010, 8, 623–633. [Google Scholar] [CrossRef] [PubMed]
- Foster, T.J. The MSCRAMM Family of Cell-Wall-Anchored Surface Proteins of Gram-positive Cocci. Trends Microbiol. 2019, 27, 927–941. [Google Scholar] [CrossRef]
- Foster, T.J. Surface proteins of Staphylococcus aureus. Microbiol. Spectr. 2019, 7. [Google Scholar] [CrossRef]
- Gaupp, R.; Ledala, N.; Somerville, G.A. Staphylococcal response to oxidative stress. Front. Cell. Infect. Microbiol. 2012, 2, 33. [Google Scholar] [CrossRef]
- Liu, G.Y.; Essex, A.; Buchanan, J.T.; Datta, V.; Hoffman, H.M.; Bastian, J.F.; Fierer, J.; Nizet, V. Staphylococcus aureus golden pigment impairs neutrophil killing and promotes virulence through its antioxidant activity. J. Exp. Med. 2005, 202, 209–215. [Google Scholar] [CrossRef]
- Clauditz, A.; Resch, A.; Wieland, K.P.; Peschel, A.; Gotz, F. Staphyloxanthin plays a role in the fitness of Staphylococcus aureus and its ability to cope with oxidative stress. Infect. Immun. 2006, 74, 4950–4953. [Google Scholar] [CrossRef]
- Mishra, N.N.; Liu, G.Y.; Yeaman, M.R.; Nast, C.C.; Proctor, R.A.; McKinnell, J.; Bayer, A.S. Carotenoid-related alteration of cell membrane fluidity impacts Staphylococcus aureus susceptibility to host defense peptides. Antimicrob. Agents Chemother. 2011, 55, 526–531. [Google Scholar] [CrossRef]
- Sen, S.; Sirobhushanam, S.; Johnson, S.R.; Song, Y.; Tefft, R.; Gatto, C.; Wilkinson, B.J. Growth-Environment Dependent Modulation of Staphylococcus aureus Branched-Chain to Straight-Chain Fatty Acid Ratio and Incorporation of Unsaturated Fatty Acids. PLoS ONE 2016, 11, e0165300. [Google Scholar] [CrossRef]
- Kaneda, T. Iso-anteiso-fatty acids in bacteria: Biosynthesis, function, and taxonomic significance. Microbiol. Rev. 1991, 55, 288–302. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.M.; Rock, C.O. membrane lipid homeostasis in bacteria. Nat. Rev. Microbiol. 2008, 6, 222–233. [Google Scholar] [CrossRef] [PubMed]
- Kaiser, J.C.; Heinrichs, D.E. Branching out: Alterations in Bacterial Physiology and Virulence due to Branched Chain Amino Acid Deprivation. mBio 2018, 9, e01188-18. [Google Scholar] [CrossRef]
- Wargo, M.J. Homeostasis and catabolism of choline and glycine betaine: Lessons from Pseudomonas aeruginosa. Appl. Environ. Microbiol. 2013, 79, 2112–2120. [Google Scholar] [CrossRef] [PubMed]
- Casey, D.; Sleator, R.D. A genomic analysis of osmotolerance in Staphylococcus aureus. Gene 2021, 767, 145268. [Google Scholar] [CrossRef] [PubMed]
- Kriegeskorte, A.; Grubmuller, S.; Huber, C.; Kahl, B.C.; von Eiff, C.; Proctor, R.A.; Peters, G.; Eisenreich, W.; Becker, K. Staphylococcus aureus small colony variants show common metabolic features in central metabolism irrespective of the underlying auxotrophism. Front. Cell. Infect. Microbiol. 2014, 4, 141. [Google Scholar] [CrossRef] [PubMed]
- Dorries, K.; Schlueter, R.; Lalk, M. Impact of antibiotics with various target sites on the metabolome of Staphylococcus Aureus. Antimicrob. Agents Chemother. 2014, 58, 7151–7163. [Google Scholar] [CrossRef]
- Jurtshuk, P.J. Bacterial Metabolism. In Medical Microbiology, 4th ed.; Baron, S., Ed.; University of Texas Medical Branch at Galveston: Galveston, TX, USA, 1996. [Google Scholar]
- Kim, H.K.P.Y.; Kim, H.N.; Choi, B.H.; Jeong, H.G.; Lee, D.G.; and Hahm, K.S. Antimicrobial mechanism of β-glycyrrhetinic acid isolated from licorice, Glycyrrhiza glabra. Biotechnol. Lett. 2002, 24, 1899–1902. [Google Scholar] [CrossRef]
- Verstraeten, S.; Catteau, L.; Boukricha, L.; Quetin-Leclercq, J.; Mingeot-Leclercq, M.P. Effect of Ursolic and Oleanolic Acids on Lipid Membranes: Studies on MRSA and Models of Membranes. Antibiotics 2021, 10, 1381. [Google Scholar] [CrossRef]
- de Leon, L.; Beltran, B.; Moujir, L. Antimicrobial activity of 6-oxophenolic triterpenoids. Mode of action against Bacillus subtilis. Planta Med. 2005, 71, 313–319. [Google Scholar] [CrossRef]
- Cohen, B.E. Functional linkage between genes that regulate osmotic stress responses and multidrug resistance transporters: Challenges and opportunities for antibiotic discovery. Antimicrob. Agents Chemother. 2014, 58, 640–646. [Google Scholar] [CrossRef] [PubMed]
- Schwan, W.R.; Wetzel, K.J. Osmolyte transport in Staphylococcus aureus and the role in pathogenesis. World J. Clin. Infect. Dis. 2016, 6, 22–27. [Google Scholar] [CrossRef] [PubMed]
- Chung, P.Y.; Navaratnam, P.; Chung, L.Y. Synergistic antimicrobial activity between pentacyclic triterpenoids and antibiotics against Staphylococcus aureus strains. Ann. Clin. Microbiol. Antimicrob. 2011, 10, 25. [Google Scholar] [CrossRef] [PubMed]
- Evaristo, F.F.; Albuquerque, M.R.; dos Santos, H.S.; Bandeira, P.N.; Avila Fdo, N.; da Silva, B.R.; Vasconcelos, A.A.; Rabelo Ede, M.; Nascimento-Neto, L.G.; Arruda, F.V.; et al. Antimicrobial effect of the triterpene 3beta,6beta,16beta-trihydroxylup-20(29)-ene on planktonic cells and biofilms from Gram positive and Gram negative bacteria. Biomed. Res. Int. 2014, 2014, 729358. [Google Scholar] [CrossRef]
- Yu, J.L.D.; Lee, S. Destabilizing effect of glycyrrhetinic acid on pre-formed biofilms of Streptococcus mutans. J. Korean Acad. Oral Health 2016, 40, 38–42. [Google Scholar] [CrossRef][Green Version]
- Qian, Y.; Xia, L.; Wei, L.; Li, D.; Jiang, W. Artesunate inhibits Staphylococcus aureus biofilm formation by reducing alpha-toxin synthesis. Arch. Microbiol. 2021, 203, 707–717. [Google Scholar] [CrossRef]
- Jiang, W.; Li, B.; Zheng, X.; Liu, X.; Pan, X.; Qing, R.; Cen, Y.; Zheng, J.; Zhou, H. Artesunate has its enhancement on antibacterial activity of beta-lactams via increasing the antibiotic accumulation within methicillin-resistant Staphylococcus aureus (MRSA). J. Antibiot. 2013, 66, 339–345. [Google Scholar] [CrossRef]
- Haaber, J.; Cohn, M.T.; Frees, D.; Andersen, T.J.; Ingmer, H. Planktonic aggregates of Staphylococcus aureus protect against common antibiotics. PLoS ONE 2012, 7, e41075. [Google Scholar] [CrossRef]
- Hook, J.L.; Islam, M.N.; Parker, D.; Prince, A.S.; Bhattacharya, S.; Bhattacharya, J. Disruption of staphylococcal aggregation protects against lethal lung injury. J. Clin. Investig. 2018, 128, 1074–1086. [Google Scholar] [CrossRef]
- Fleming, D.; Rumbaugh, K. The Consequences of Biofilm Dispersal on the Host. Sci. Rep. 2018, 8, 10738. [Google Scholar] [CrossRef]
- Wille, J.; Coenye, T. Biofilm dispersion: The key to biofilm eradication or opening Pandora’s box? Biofilm 2020, 2, 100027. [Google Scholar] [CrossRef] [PubMed]
- Diep, B.A.; Gill, S.R.; Chang, R.F.; Phan, T.H.; Chen, J.H.; Davidson, M.G.; Lin, F.; Lin, J.; Carleton, H.A.; Mongodin, E.F.; et al. Complete genome sequence of USA300, an epidemic clone of community-acquired meticillin-resistant Staphylococcus aureus. Lancet 2006, 367, 731–739. [Google Scholar] [CrossRef]
- Diep, B.A.; Sensabaugh, G.F.; Somboonna, N.; Carleton, H.A.; Perdreau-Remington, F. Widespread skin and soft-tissue infections due to two methicillin-resistant Staphylococcus aureus strains harboring the genes for Panton-Valentine leucocidin. J. Clin. Microbiol. 2004, 42, 2080–2084. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Klevens, R.M.; Morrison, M.A.; Nadle, J.; Petit, S.; Gershman, K.; Ray, S.; Harrison, L.H.; Lynfield, R.; Dumyati, G.; Townes, J.M.; et al. Invasive methicillin-resistant Staphylococcus aureus infections in the United States. JAMA 2007, 298, 1763–1771. [Google Scholar] [CrossRef]
- Ammons, M.C.; Tripet, B.P.; Carlson, R.P.; Kirker, K.R.; Gross, M.A.; Stanisich, J.J.; Copié, V. Quantitative NMR metabolite profiling of methicillin-resistant and methicillin-susceptible Staphylococcus aureus discriminates between biofilm and planktonic phenotypes. J. Proteome Res. 2014, 13, 2973–2985. [Google Scholar] [CrossRef]
- Hwang, T.L.; Shaka, A.J. Water Suppression That Works. Excitation Sculpting Using Arbitrary Wave-Forms and Pulsed-Field Gradients. J. Magn. Reson. Ser. A 1995, 112, 275–279. [Google Scholar] [CrossRef]
- Fuchs, A.L.; Schiller, S.M.; Keegan, W.J.; Ammons, M.C.B.; Eilers, B.; Tripet, B.; Copié, V. Quantitative (1)H NMR Metabolomics Reveal Distinct Metabolic Adaptations in Human Macrophages Following Differential Activation. Metabolites 2019, 9, 248. [Google Scholar] [CrossRef]
- Weljie, A.M.; Newton, J.; Mercier, P.; Carlson, E.; Slupsky, C.M. Targeted profiling: Quantitative analysis of 1H NMR metabolomics data. Anal. Chem. 2006, 78, 4430–4442. [Google Scholar] [CrossRef]
- Wu, X.H.; Yu, H.L.; Ba, Z.Y.; Chen, J.Y.; Sun, H.G.; Han, B.Z. Sampling methods for NMR-based metabolomics of Staphylococcus Aureus. Biotechnol. J. 2010, 5, 75–84. [Google Scholar] [CrossRef]
- Chong, J.; Soufan, O.; Li, C.; Caraus, I.; Li, S.; Bourque, G.; Wishart, D.S.; Xia, J. MetaboAnalyst 4.0: Towards more transparent and integrative metabolomics analysis. Nucleic Acids Res. 2018, 46, W486–W494. [Google Scholar] [CrossRef]
- Kirker, K.R.; Secor, P.R.; James, G.A.; Fleckman, P.; Olerud, J.E.; Stewart, P.S. Loss of viability and induction of apoptosis in human keratinocytes exposed to Staphylococcus aureus biofilms in vitro. Wound Repair Regen. 2009, 17, 690–699. [Google Scholar] [CrossRef] [PubMed]
- Cassat, J.E.; Lee, C.Y.; Smeltzer, M.S. Investigation of biofilm formation in clinical isolates of Staphylococcus aureus. Methods Mol. Biol. 2007, 391, 127–144. [Google Scholar] [CrossRef] [PubMed]
- Djordjevic, D.; Wiedmann, M.; McLandsborough, L.A. Microtiter plate assay for assessment of Listeria monocytogenes biofilm formation. Appl. Environ. Microbiol. 2002, 68, 2950–2958. [Google Scholar] [CrossRef] [PubMed]
- Geoghegan, J.A.; Corrigan, R.M.; Gruszka, D.T.; Speziale, P.; O’Gara, J.P.; Potts, J.R.; Foster, T.J. Role of surface protein SasG in biofilm formation by Staphylococcus aureus. J. Bacteriol. 2010, 192, 5663–5673. [Google Scholar] [CrossRef] [PubMed]
- Voyich, J.M.; Braughton, K.R.; Sturdevant, D.E.; Whitney, A.R.; Said-Salim, B.; Porcella, S.F.; Long, R.D.; Dorward, D.W.; Gardner, D.J.; Kreiswirth, B.N.; et al. Insights into mechanisms used by Staphylococcus aureus to avoid destruction by human neutrophils. J. Immunol. 2005, 175, 3907–3919. [Google Scholar] [CrossRef] [PubMed]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Weaver, A.J., Jr.; Borgogna, T.R.; O’Shea-Stone, G.; Peters, T.R.; Copié, V.; Voyich, J.; Teintze, M. 18β-Glycyrrhetinic Acid Induces Metabolic Changes and Reduces Staphylococcus aureus Bacterial Cell-to-Cell Interactions. Antibiotics 2022, 11, 781. https://doi.org/10.3390/antibiotics11060781
Weaver AJ Jr., Borgogna TR, O’Shea-Stone G, Peters TR, Copié V, Voyich J, Teintze M. 18β-Glycyrrhetinic Acid Induces Metabolic Changes and Reduces Staphylococcus aureus Bacterial Cell-to-Cell Interactions. Antibiotics. 2022; 11(6):781. https://doi.org/10.3390/antibiotics11060781
Chicago/Turabian StyleWeaver, Alan J., Jr., Timothy R. Borgogna, Galen O’Shea-Stone, Tami R. Peters, Valérie Copié, Jovanka Voyich, and Martin Teintze. 2022. "18β-Glycyrrhetinic Acid Induces Metabolic Changes and Reduces Staphylococcus aureus Bacterial Cell-to-Cell Interactions" Antibiotics 11, no. 6: 781. https://doi.org/10.3390/antibiotics11060781
APA StyleWeaver, A. J., Jr., Borgogna, T. R., O’Shea-Stone, G., Peters, T. R., Copié, V., Voyich, J., & Teintze, M. (2022). 18β-Glycyrrhetinic Acid Induces Metabolic Changes and Reduces Staphylococcus aureus Bacterial Cell-to-Cell Interactions. Antibiotics, 11(6), 781. https://doi.org/10.3390/antibiotics11060781