Similarities in Virulence and Extended Spectrum Beta-Lactamase Gene Profiles among Cefotaxime-Resistant Escherichia coli Wastewater and Clinical Isolates
Abstract
1. Introduction
2. Results and Discussion
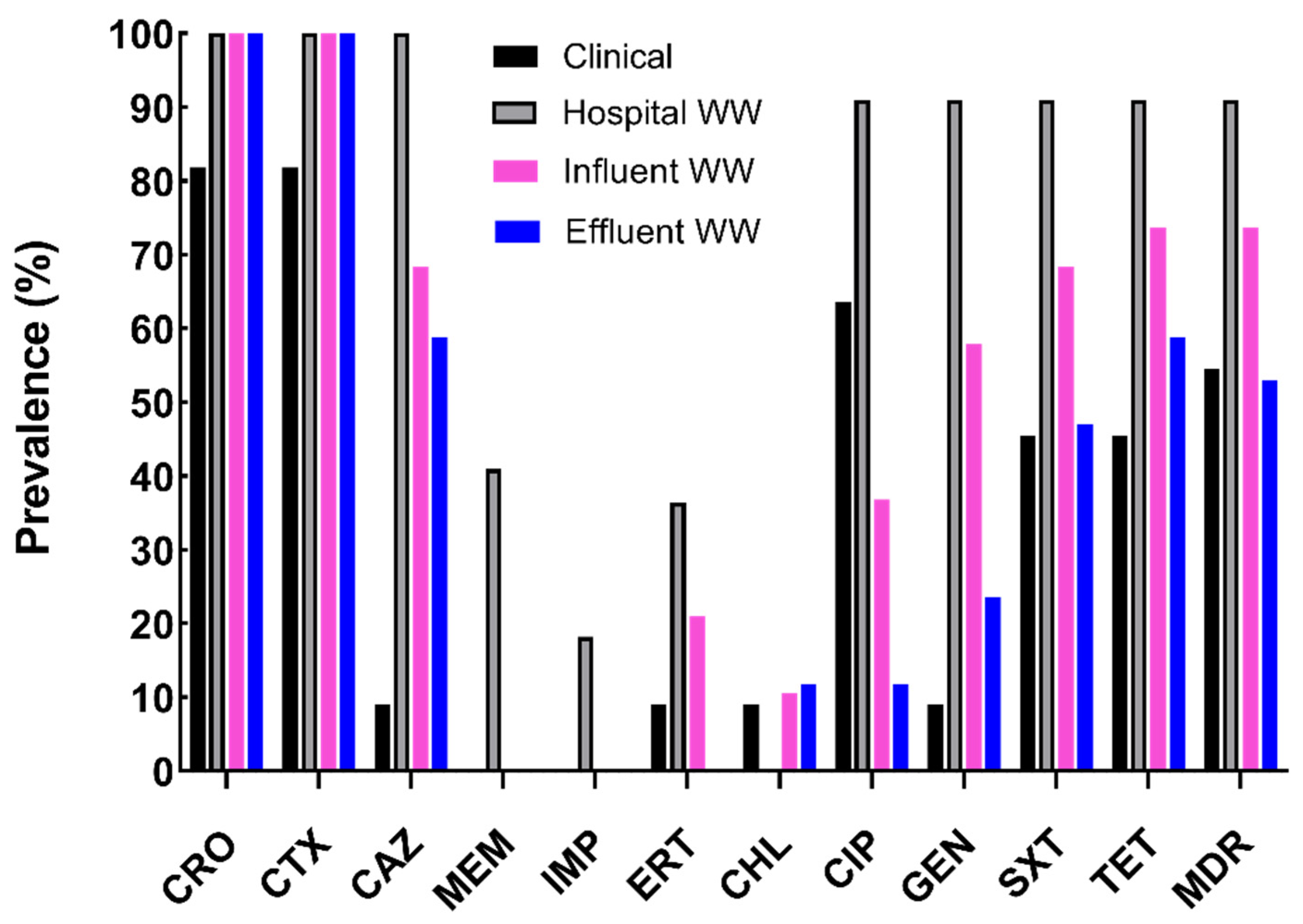
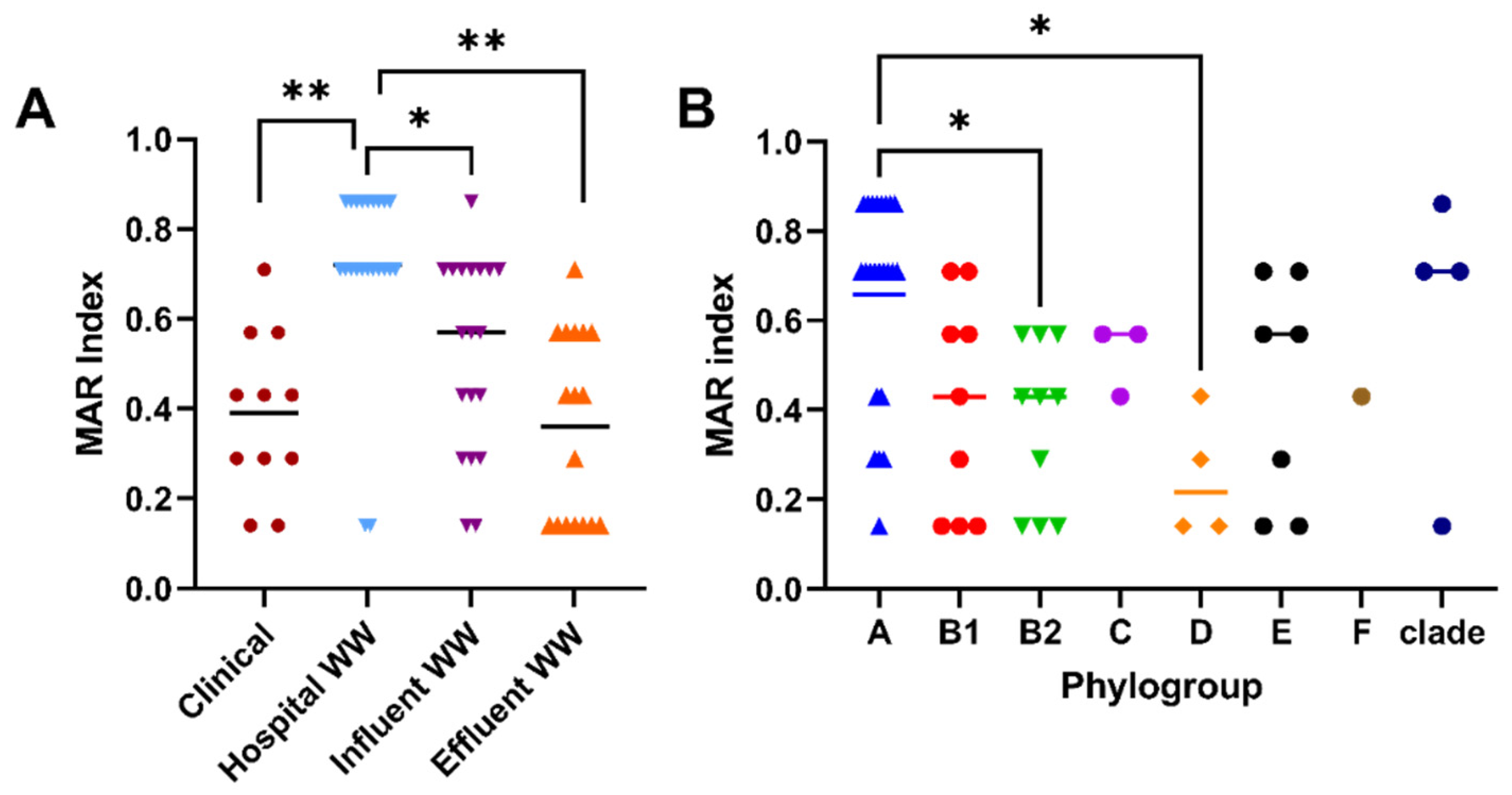
2.1. Characterization of Antibiotic Resistance Profiles of Cefotaxime-Resistant E. coli
2.2. Prevalence of β-Lactamases among CTX-Resistant Populations
2.3. Whole-Genome Sequence Analysis of Carbapenemase-Producing E. coli Isolate
2.4. Prevalence of Virulence Genes among CTX-Resistant Populations
3. Materials and Methods
3.1. Acquisition of CTX-Resistant E. coli Isolates
3.2. Antibiotic Susceptibility Testing
3.3. Plasmid Isolation and Transformation
3.4. Identification of Beta-Lactamases and Virulence Genes
3.5. Phylogroup Identification
3.6. Whole Genome Sequencing and Analysis
3.7. Statistics
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- CDC. Antibiotic Resistance Threats in the United States 2019; Department of Health and Human Services: Atlanta, GA, USA, 2019. [CrossRef]
- McEwen, S.A.; Collignon, P.J. Antimicrobial Resistance: A One Health Perspective. Microbiol. Spectr. 2018, 6, 2. [Google Scholar] [CrossRef] [PubMed]
- Al-Jassim, N.; Ansari, M.I.; Harb, M.; Hong, P.Y. Removal of bacterial contaminants and antibiotic resistance genes by conventional wastewater treatment processes in Saudi Arabia: Is the treated wastewater safe to reuse for agricultural irrigation? Water Res. 2015, 73, 277–290. [Google Scholar] [CrossRef] [PubMed]
- Sharma, V.K.; Johnson, N.; Cizmas, L.; McDonald, T.J.; Kim, H. A review of the influence of treatment strategies on antibiotic resistant bacteria and antibiotic resistance genes. Chemosphere 2016, 150, 702–714. [Google Scholar] [CrossRef] [PubMed]
- Aubertheau, E.; Stalder, T.; Mondamert, L.; Ploy, M.C.; Dagot, C.; Labanowski, J. Impact of wastewater treatment plant discharge on the contamination of river biofilms by pharmaceuticals and antibiotic resistance. Sci. Total Environ. 2017, 579, 1387–1398. [Google Scholar] [CrossRef]
- Rizzo, L.; Manaia, C.; Merlin, C.; Schwartz, T.; Dagot, C.; Ploy, M.C.; Michael, I.; Fatta-Kassinos, D. Urban wastewater treatment plants as hotspots for antibiotic resistant bacteria and genes spread into the environment: A review. Sci. Total Environ. 2013, 447, 345–360. [Google Scholar] [CrossRef]
- Korzeniewska, E.; Korzeniewska, A.; Harnisz, M. Antibiotic resistant Escherichia coli in hospital and municipal sewage and their emission to the environment. Ecotoxicol. Environ. Saf. 2013, 91, 96–102. [Google Scholar] [CrossRef]
- Amador, P.P.; Fernandes, R.M.; Prudencio, M.C.; Barreto, M.P.; Duarte, I.M. Antibiotic resistance in wastewater: Occurrence and fate of Enterobacteriaceae producers of class A and class C beta-lactamases. J. Environ. Sci. Health 2015, 50, 26–39. [Google Scholar] [CrossRef]
- Beattie, R.E.; Skwor, T.; Hristova, K.R. Survivor microbial populations in post-chlorinated wastewater are strongly associated with untreated hospital sewage and include ceftazidime and meropenem resistant populations. Sci. Total Environ. 2020, 740, 140186. [Google Scholar] [CrossRef]
- Kappell, A.D.; DeNies, M.S.; Ahuja, N.H.; Ledeboer, N.A.; Newton, R.J.; Hristova, K.R. Detection of multi-drug resistant Escherichia coli in the urban waterways of Milwaukee, WI. Front. Microbiol. 2015, 6, 336. [Google Scholar] [CrossRef]
- Centers for Disease Control and Prevention. Outpatient Antibiotic Prescriptions—United States, 2018; CDC: Atlanta, GA, USA, 2018.
- Adegoke, A.A.; Madu, C.E.; Aiyegoro, O.A.; Stenstrom, T.A.; Okoh, A.I. Antibiogram and beta-lactamase genes among cefotaxime resistant E. coli from wastewater treatment plant. Antimicrob. Resist. Infect. Control. 2020, 9, 46. [Google Scholar] [CrossRef]
- Paterson, D.L.; Bonomo, R.A. Extended-spectrum beta-lactamases: A clinical update. Clin. Microbiol. Rev. 2005, 18, 657–686. [Google Scholar] [CrossRef] [PubMed]
- Silva, I.; Tacao, M.; Tavares, R.D.S.; Miranda, R.; Araujo, S.; Manaia, C.M.; Henriques, I. Fate of cefotaxime-resistant Enterobacteriaceae and ESBL-producers over a full-scale wastewater treatment process with UV disinfection. Sci. Total Environ. 2018, 639, 1028–1037. [Google Scholar] [CrossRef] [PubMed]
- Hutinel, M.; Fick, J.; Larsson, D.G.J.; Flach, C.F. Investigating the effects of municipal and hospital wastewaters on horizontal gene transfer. Environ. Pollut. 2021, 276, 116733. [Google Scholar] [CrossRef] [PubMed]
- Edberg, S.C.; Rice, E.W.; Karlin, R.J.; Allen, M.J. Escherichia coli: The best biological drinking water indicator for public health protection. Symp. Ser. Soc. Appl. Microbiol. 2000, 88, 106S–116S. [Google Scholar] [CrossRef] [PubMed]
- Matic, I.; Rayssiguier, C.; Radman, M. Interspecies gene exchange in bacteria: The role of SOS and mismatch repair systems in evolution of species. Cell 1995, 80, 507–515. [Google Scholar] [CrossRef]
- Beattie, R.E.; Bakke, E.; Konopek, N.; Thill, R.; Munson, E.; Hristova, K.R. Antimicrobial Resistance Traits of Escherichia coli Isolated from Dairy Manure and Freshwater Ecosystems Are Similar to One Another but Differ from Associated Clinical Isolates. Microorganisms 2020, 8, 747. [Google Scholar] [CrossRef]
- Bojar, B.; Sheridan, J.; Beattie, R.; Cahak, C.; Liedhegner, E.; Munoz-Price, L.S.; Hristova, K.R.; Skwor, T. Antibiotic resistance patterns of Escherichia coli isolates from the clinic through the wastewater pathway. Int. J. Hyg. Environ. Health 2021, 238, 113863. [Google Scholar] [CrossRef]
- Nelson, R.E.; Hatfield, K.M.; Wolford, H.; Samore, M.H.; Scott, R.D., II; Reddy, S.C.; Olubajo, B.; Paul, P.; Jernigan, J.A.; Baggs, J. National Estimates of Healthcare Costs Associated With Multidrug-Resistant Bacterial Infections Among Hospitalized Patients in the United States. Clin. Infect. Dis. 2021, 72, S17–S26. [Google Scholar] [CrossRef]
- Bezabih, Y.M.; Sabiiti, W.; Alamneh, E.; Bezabih, A.; Peterson, G.M.; Bezabhe, W.M.; Roujeinikova, A. The global prevalence and trend of human intestinal carriage of ESBL-producing Escherichia coli in the community. J. Antimicrob. Chemother. 2020, 76, 22–29. [Google Scholar] [CrossRef]
- Marano, R.B.M.; Fernandes, T.; Manaia, C.M.; Nunes, O.; Morrison, D.; Berendonk, T.U.; Kreuzinger, N.; Tenson, T.; Corno, G.; Fatta-Kassinos, D.; et al. A global multinational survey of cefotaxime-resistant coliforms in urban wastewater treatment plants. Environ. Int. 2020, 144, 106035. [Google Scholar] [CrossRef]
- Diwan, V.; Chandran, S.P.; Tamhankar, A.J.; Stalsby Lundborg, C.; Macaden, R. Identification of extended-spectrum beta-lactamase and quinolone resistance genes in Escherichia coli isolated from hospital wastewater from central India. J. Antimicrob. Chemother. 2012, 67, 857–859. [Google Scholar] [CrossRef] [PubMed]
- Paulshus, E.; Kuhn, I.; Mollby, R.; Colque, P.; O’Sullivan, K.; Midtvedt, T.; Lingaas, E.; Holmstad, R.; Sorum, H. Diversity and antibiotic resistance among Escherichia coli populations in hospital and community wastewater compared to wastewater at the receiving urban treatment plant. Water Res. 2019, 161, 232–241. [Google Scholar] [CrossRef] [PubMed]
- Clermont, O.; Christenson, J.K.; Denamur, E.; Gordon, D.M. The Clermont Escherichia coli phylo-typing method revisited: Improvement of specificity and detection of new phylo-groups. Environ. Microbiol. Rep. 2013, 5, 58–65. [Google Scholar] [CrossRef] [PubMed]
- Schreiber, H.L.t.; Conover, M.S.; Chou, W.C.; Hibbing, M.E.; Manson, A.L.; Dodson, K.W.; Hannan, T.J.; Roberts, P.L.; Stapleton, A.E.; Hooton, T.M.; et al. Bacterial virulence phenotypes of Escherichia coli and host susceptibility determine risk for urinary tract infections. Sci. Transl. Med. 2017, 9, eaaf1283. [Google Scholar] [CrossRef]
- Stoppe, N.C.; Silva, J.S.; Carlos, C.; Sato, M.I.Z.; Saraiva, A.M.; Ottoboni, L.M.M.; Torres, T.T. Worldwide Phylogenetic Group Patterns of Escherichia coli from Commensal Human and Wastewater Treatment Plant Isolates. Front. Microbiol. 2017, 8, 2512. [Google Scholar] [CrossRef]
- Sabate, M.; Prats, G.; Moreno, E.; Balleste, E.; Blanch, A.R.; Andreu, A. Virulence and antimicrobial resistance profiles among Escherichia coli strains isolated from human and animal wastewater. Res. Microbiol. 2008, 159, 288–293. [Google Scholar] [CrossRef]
- Rebello, R.C.; Regua-Mangia, A.H. Potential enterovirulence and antimicrobial resistance in Escherichia coli isolates from aquatic environments in Rio de Janeiro, Brazil. Sci. Total Environ. 2014, 490, 19–27. [Google Scholar] [CrossRef]
- Gundogdu, A.; Jennison, A.V.; Smith, H.V.; Stratton, H.; Katouli, M. Extended-spectrum beta-lactamase producing Escherichia coli in hospital wastewaters and sewage treatment plants in Queensland, Australia. Can. J. Microbiol. 2013, 59, 737–745. [Google Scholar] [CrossRef]
- Woerther, P.L.; Burdet, C.; Chachaty, E.; Andremont, A. Trends in Human Fecal Carriage of Extended-Spectrum beta-Lactamases in the Community: Toward the Globalization of CTX-M. Clin. Microbiol. Rev. 2013, 26, 744–758. [Google Scholar] [CrossRef]
- Rhee, C.; Kadri, S.S.; Dekker, J.P.; Danner, R.L.; Chen, H.C.; Fram, D.; Zhang, F.; Wang, R.; Klompas, M.; Program, C.D.C.P.E. Prevalence of Antibiotic-Resistant Pathogens in Culture-Proven Sepsis and Outcomes Associated With Inadequate and Broad-Spectrum Empiric Antibiotic Use. JAMA Netw. Open 2020, 3, e202899. [Google Scholar] [CrossRef]
- Ruppe, E.; Lixandru, B.; Cojocaru, R.; Buke, C.; Paramythiotou, E.; Angebault, C.; Visseaux, C.; Djuikoue, I.; Erdem, E.; Burduniuc, O.; et al. Relative fecal abundance of extended-spectrum-beta-lactamase-producing Escherichia coli strains and their occurrence in urinary tract infections in women. Antimicrob. Agents Chemother. 2013, 57, 4512–4517. [Google Scholar] [CrossRef] [PubMed]
- Dorado-Garcia, A.; Smid, J.H.; van Pelt, W.; Bonten, M.J.M.; Fluit, A.C.; van den Bunt, G.; Wagenaar, J.A.; Hordijk, J.; Dierikx, C.M.; Veldman, K.T.; et al. Molecular relatedness of ESBL/AmpC-producing Escherichia coli from humans, animals, food and the environment: A pooled analysis. J. Antimicrob. Chemother. 2018, 73, 339–347. [Google Scholar] [CrossRef]
- Overdevest, I.; Willemsen, I.; Rijnsburger, M.; Eustace, A.; Xu, L.; Hawkey, P.; Heck, M.; Savelkoul, P.; Vandenbroucke-Grauls, C.; van der Zwaluw, K.; et al. Extended-Spectrum beta-Lactamase Genes of Escherichia coli in Chicken Meat and Humans, the Netherlands. Emerg. Infect. Dis. 2011, 17, 1216–1222. [Google Scholar] [CrossRef] [PubMed]
- Canton, R.; Gonzalez-Alba, J.M.; Galan, J.C. CTX-M Enzymes: Origin and Diffusion. Front. Microbiol. 2012, 3, 110. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, H.; Hayashi, W.; Iimura, M.; Taniguchi, Y.; Soga, E.; Matsuo, N.; Kawamura, K.; Arakawa, Y.; Nagano, Y.; Nagano, N.; et al. Wastewater as a Probable Environmental Reservoir of Extended-Spectrum-β-Lactamase Genes: Detection of Chimeric β-Lactamases CTX-M-64 and CTX-M-123. Appl. Environ. Microbiol. 2019, 85, e01740-19. [Google Scholar] [CrossRef]
- Castanheira, M.; Farrell, S.E.; Deshpande, L.M.; Mendes, R.E.; Jones, R.N. Prevalence of β-lactamase-encoding genes among Enterobacteriaceae bacteremia isolates collected in 26 U.S. hospitals: Report from the SENTRY Antimicrobial Surveillance Program (2010). Antimicrob. Agents Chemother. 2013, 57, 3012–3020. [Google Scholar] [CrossRef]
- Denisuik, A.J.; Lagacé-Wiens, P.R.; Pitout, J.D.; Mulvey, M.R.; Simner, P.J.; Tailor, F.; Karlowsky, J.A.; Hoban, D.J.; Adam, H.J.; Zhanel, G.G. Molecular epidemiology of extended-spectrum β-lactamase-, AmpC β-lactamase- and carbapenemase-producing Escherichia coli and Klebsiella pneumoniae isolated from Canadian hospitals over a 5 year period: CANWARD 2007-11. J. Antimicrob. Chemother. 2013, 68 (Suppl. 1), i57–i65. [Google Scholar] [CrossRef]
- Pai, H.; Choi, E.H.; Lee, H.J.; Hong, J.Y.; Jacoby, G.A. Identification of CTX-M-14 extended-spectrum beta-lactamase in clinical isolates of Shigella sonnei, Escherichia coli, and Klebsiella pneumoniae in Korea. J. Clin. Microbiol. 2001, 39, 3747–3749. [Google Scholar] [CrossRef]
- Constantinides, B.; Chau, K.K.; Quan, T.P.; Rodger, G.; Andersson, M.I.; Jeffery, K.; Lipworth, S.; Gweon, H.S.; Peniket, A.; Pike, G.; et al. Genomic surveillance of Escherichia coli and Klebsiella spp. in hospital sink drains and patients. Microb. Genom. 2020, 6. [Google Scholar] [CrossRef]
- Borgogna, T.R.; Borgogna, J.-L.; Mielke, J.A.; Brown, C.J.; Top, E.M.; Botts, R.T.; Cummings, D.E. High Diversity of CTX-M Extended-Spectrum β-Lactamases in Municipal Wastewater and Urban Wetlands. Microb. Drug Resist. 2016, 22, 312–320. [Google Scholar] [CrossRef]
- Bevan, E.R.; Jones, A.M.; Hawkey, P.M. Global epidemiology of CTX-M β-lactamases: Temporal and geographical shifts in genotype. J. Antimicrob. Chemother. 2017, 72, 2145–2155. [Google Scholar] [CrossRef] [PubMed]
- Branger, C.; Zamfir, O.; Geoffroy, S.; Laurans, G.; Arlet, G.; Thien, H.V.; Gouriou, S.; Picard, B.; Denamur, E. Genetic background of Escherichia coli and extended-spectrum beta-lactamase type. Emerg. Infect. Dis. 2005, 11, 54–61. [Google Scholar] [CrossRef]
- Kutilova, I.; Medvecky, M.; Leekitcharoenphon, P.; Munk, P.; Masarikova, M.; Davidova-Gerzova, L.; Jamborova, I.; Bortolaia, V.; Pamp, S.J.; Dolejska, M. Extended-spectrum beta-lactamase-producing Escherichia coli and antimicrobial resistance in municipal and hospital wastewaters in Czech Republic: Culture-based and metagenomic approaches. Environ. Res. 2021, 193, 110487. [Google Scholar] [CrossRef] [PubMed]
- Nzima, B.; Adegoke, A.A.; Ofon, U.A.; Al-Dahmoshi, H.O.M.; Saki, M.; Ndubuisi-Nnaji, U.U.; Inyang, C.U. Resistotyping and extended-spectrum beta-lactamase genes among Escherichia coli from wastewater treatment plants and recipient surface water for reuse in South Africa. New Microbes. New Infect. 2020, 38, 100803. [Google Scholar] [CrossRef] [PubMed]
- Martínez, J.L.; Coque, T.M.; Baquero, F. What is a resistance gene? Ranking risk in resistomes. Nat. Rev. Microbiol. 2015, 13, 116–123. [Google Scholar] [CrossRef]
- Wang, J.; Chu, L.; Wojnárovits, L.; Takács, E. Occurrence and fate of antibiotics, antibiotic resistant genes (ARGs) and antibiotic resistant bacteria (ARB) in municipal wastewater treatment plant: An overview. Sci. Total Environ. 2020, 744, 140997. [Google Scholar] [CrossRef]
- Gillings, M.R.; Gaze, W.H.; Pruden, A.; Smalla, K.; Tiedje, J.M.; Zhu, Y.-G. Using the class 1 integron-integrase gene as a proxy for anthropogenic pollution. ISME J. 2015, 9, 1269–1279. [Google Scholar] [CrossRef]
- Bonomo, R.A.; Burd, E.M.; Conly, J.; Limbago, B.M.; Poirel, L.; Segre, J.A.; Westblade, L.F. Carbapenemase-Producing Organisms: A Global Scourge. Clin. Infect. Dis. 2018, 66, 1290–1297. [Google Scholar] [CrossRef]
- Logan, L.K.; Weinstein, R.A. The Epidemiology of Carbapenem-Resistant Enterobacteriaceae: The Impact and Evolution of a Global Menace. J. Infect. Dis. 2017, 215, S28–S36. [Google Scholar] [CrossRef]
- Bettelheim, K.A. The sources of "OH" serotypes of Escherichia coli. J. Hyg. 1978, 80, 83–113. [Google Scholar] [CrossRef]
- Dong, D.; Mi, Z.; Li, D.; Gao, M.; Jia, N.; Li, M.; Tong, Y.; Zhang, X.; Zhu, Y. Novel IncR/IncP6 Hybrid Plasmid pCRE3-KPC Recovered from a Clinical KPC-2-Producing Citrobacter braakii Isolate. mSphere 2020, 5, e00891-19. [Google Scholar] [CrossRef]
- Stoesser, N.; Sheppard, A.E.; Peirano, G.; Anson, L.W.; Pankhurst, L.; Sebra, R.; Phan, H.T.T.; Kasarskis, A.; Mathers, A.J.; Peto, T.E.A.; et al. Genomic epidemiology of global Klebsiella pneumoniae carbapenemase (KPC)-producing Escherichia coli. Sci. Rep. 2017, 7, 5917. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Fernandez, A.; Villa, L.; Bibbolino, G.; Bressan, A.; Trancassini, M.; Pietropaolo, V.; Venditti, M.; Antonelli, G.; Carattoli, A. Novel Insights and Features of the NDM-5-Producing Escherichia coli Sequence Type 167 High-Risk Clone. mSphere 2020, 5, e00269-20. [Google Scholar] [CrossRef] [PubMed]
- Alav, I.; Sutton, J.M.; Rahman, K.M. Role of bacterial efflux pumps in biofilm formation. J. Antimicrob. Chemother. 2018, 73, 2003–2020. [Google Scholar] [CrossRef] [PubMed]
- Du, D.; Wang-Kan, X.; Neuberger, A.; van Veen, H.W.; Pos, K.M.; Piddock, L.J.V.; Luisi, B.F. Multidrug efflux pumps: Structure, function and regulation. Nat. Rev. Microbiol. 2018, 16, 523–539. [Google Scholar] [CrossRef] [PubMed]
- Yao, Y.; Lazaro-Perona, F.; Falgenhauer, L.; Valverde, A.; Imirzalioglu, C.; Dominguez, L.; Cantón, R.; Mingorance, J.; Chakraborty, T. Insights into a Novel bla(KPC-2)-Encoding IncP-6 Plasmid Reveal Carbapenem-Resistance Circulation in Several Enterobacteriaceae Species from Wastewater and a Hospital Source in Spain. Front. Microbiol. 2017, 8, 1143. [Google Scholar] [CrossRef] [PubMed]
- Yeo, C.C.; Tham, J.M.; Kwong, S.M.; Yiin, S.; Poh, C.L. Tn5563, a transposon encoding putative mercuric ion transport proteins located on plasmid pRA2 of Pseudomonas alcaligenes. FEMS Microbiol. Lett. 1998, 165, 253–260. [Google Scholar] [CrossRef][Green Version]
- Rozwandowicz, M.; Brouwer, M.S.M.; Fischer, J.; Wagenaar, J.A.; Gonzalez-Zorn, B.; Guerra, B.; Mevius, D.J.; Hordijk, J. Plasmids carrying antimicrobial resistance genes in Enterobacteriaceae. J. Antimicrob. Chemother. 2018, 73, 1121–1137. [Google Scholar] [CrossRef]
- Koraimann, G. Spread and Persistence of Virulence and Antibiotic Resistance Genes: A Ride on the F Plasmid Conjugation Module. EcoSal Plus 2018, 8. [Google Scholar] [CrossRef]
- Rinkel, M.; Hubert, J.C.; Roux, B.; Lett, M.C. Identification of a new transposon Tn5403 in a Klebsiella pneumoniae strain isolated from a polluted aquatic environment. Curr. Microbiol. 1994, 29, 249–254. [Google Scholar] [CrossRef]
- Mshana, S.E.; Imirzalioglu, C.; Hossain, H.; Hain, T.; Domann, E.; Chakraborty, T. Conjugative IncFI plasmids carrying CTX-M-15 among Escherichia coli ESBL producing isolates at a University hospital in Germany. BMC Infect. Dis. 2009, 9, 97. [Google Scholar] [CrossRef] [PubMed]
- Harmer, C.J.; Moran, R.A.; Hall, R.M. Movement of IS26-associated antibiotic resistance genes occurs via a translocatable unit that includes a single IS26 and preferentially inserts adjacent to another IS26. mBio 2014, 5, e01801–e01814. [Google Scholar] [CrossRef] [PubMed]
- Varani, A.; He, S.; Siguier, P.; Ross, K.; Chandler, M. The IS6 family, a clinically important group of insertion sequences including IS26. Mob. DNA-Uk 2021, 12, 11. [Google Scholar] [CrossRef]
- Russo, T.A.; Olson, R.; MacDonald, U.; Metzger, D.; Maltese, L.M.; Drake, E.J.; Gulick, A.M.; Camilli, A. Aerobactin Mediates Virulence and Accounts for Increased Siderophore Production under Iron-Limiting Conditions by Hypervirulent (Hypermucoviscous) Klebsiella pneumoniae. Infect. Immun. 2014, 82, 2356–2367. [Google Scholar] [CrossRef]
- Soto, S.M.; Smithson, A.; Horcajada, J.P.; Martinez, J.A.; Mensa, J.P.; Vila, J. Implication of biofilm formation in the persistence of urinary tract infection caused by uropathogenic Escherichia coli. Clin. Microbiol. Infect. 2006, 12, 1034–1036. [Google Scholar] [CrossRef]
- Li, C.; Pan, D.; Li, M.; Wang, Y.; Song, L.; Yu, D.; Zuo, Y.; Wang, K.; Liu, Y.; Wei, Z.; et al. Aerobactin-Mediated Iron Acquisition Enhances Biofilm Formation, Oxidative Stress Resistance, and Virulence of Yersinia pseudotuberculosis. Front. Microbiol. 2021, 12. [Google Scholar] [CrossRef]
- Cosentino, S.; Voldby Larsen, M.; Møller Aarestrup, F.; Lund, O. PathogenFinder—Distinguishing Friend from Foe Using Bacterial Whole Genome Sequence Data. PLoS ONE 2013, 8, e77302. [Google Scholar] [CrossRef]
- Tarchouna, M.; Ferjani, A.; Ben-Selma, W.; Boukadida, J. Distribution of uropathogenic virulence genes in Escherichia coli isolated from patients with urinary tract infection. Int. J. Infect. Dis. 2013, 17, e450–e453. [Google Scholar] [CrossRef]
- Sarowska, J.; Futoma-Koloch, B.; Jama-Kmiecik, A.; Frej-Madrzak, M.; Ksiazczyk, M.; Bugla-PLoSkonska, G.; Choroszy-Krol, I. Virulence factors, prevalence and potential transmission of extraintestinal pathogenic Escherichia coli isolated from different sources: Recent reports. Gut. Pathog. 2019, 11, 10. [Google Scholar] [CrossRef]
- Johnson, J.R.; Kuskowski, M.A.; Gajewski, A.; Sahm, D.F.; Karlowsky, J.A. Virulence characteristics and phylogenetic background of multidrug-resistant and antimicrobial-susceptible clinical isolates of Escherichia coli from across the United States, 2000–2001. J. Infect. Dis. 2004, 190, 1739–1744. [Google Scholar] [CrossRef][Green Version]
- Johnson, J.R.; Moseley, S.L.; Roberts, P.L.; Stamm, W.E. Aerobactin and other virulence factor genes among strains of Escherichia coli causing urosepsis: Association with patient characteristics. Infect. Immun. 1988, 56, 405–412. [Google Scholar] [CrossRef] [PubMed]
- Calhau, V.; Mendes, C.; Pena, A.; Mendonca, N.; Da Silva, G.J. Virulence and plasmidic resistance determinants of Escherichia coli isolated from municipal and hospital wastewater treatment plants. J. Water Health 2015, 13, 311–318. [Google Scholar] [CrossRef]
- Picard, B.; Garcia, J.S.; Gouriou, S.; Duriez, P.; Brahimi, N.; Bingen, E.; Elion, J.; Denamur, E. The link between phylogeny and virulence in Escherichia coli extraintestinal infection. Infect. Immun. 1999, 67, 546–553. [Google Scholar] [CrossRef] [PubMed]
- Qin, X.; Hu, F.; Wu, S.; Ye, X.; Zhu, D.; Zhang, Y.; Wang, M. Comparison of adhesin genes and antimicrobial susceptibilities between uropathogenic and intestinal commensal Escherichia coli strains. PLoS ONE 2013, 8, e61169. [Google Scholar] [CrossRef] [PubMed]
- Zhi, S.; Stothard, P.; Banting, G.; Scott, C.; Huntley, K.; Ryu, K.; Otto, S.; Ashbolt, N.; Checkley, S.; Dong, T.; et al. Characterization of water treatment-resistant and multidrug-resistant urinary pathogenic Escherichia coli in treated wastewater. Water Res. 2020, 182, 115827. [Google Scholar] [CrossRef] [PubMed]
- CLSI. Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria that Grow Aerobically, Approved Standard, CLSI Document M07-A10, 10th ed.; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2015. [Google Scholar]
- CLSI. Performance Standards for Antimicrobial Susceptibility Testing, Twenty-Fifth Informational Supplement, CLSI Document M100-S25; Clinical and Laboratory Standards Insititute: Wayne, PA, USA, 2015. [Google Scholar]
- Chen, I.M.A.; Chu, K.; Palaniappan, K.; Ratner, A.; Huang, J.; Huntemann, M.; Hajek, P.; Ritter, S.; Varghese, N.; Seshadri, R.; et al. The IMG/M data management and analysis system v.6.0: New tools and advanced capabilities. Nucleic Acids Res. 2021, 49, D751–D763. [Google Scholar] [CrossRef]
- Mukherjee, S.; Stamatis, D.; Bertsch, J.; Ovchinnikova, G.; Sundaramurthi, J.C.; Lee, J.; Kandimalla, M.; Chen, I.M.A.; Kyrpides, N.C.; Reddy, T.B.K. Genomes OnLine Database (GOLD) v.8: Overview and updates. Nucleic Acids Res. 2021, 49, D723–D733. [Google Scholar] [CrossRef]
- Wattam, A.R.; Davis, J.J.; Assaf, R.; Boisvert, S.; Brettin, T.; Bun, C.; Conrad, N.; Dietrich, E.M.; Disz, T.; Gabbard, J.L.; et al. Improvements to PATRIC, the all-bacterial Bioinformatics Database and Analysis Resource Center. Nucleic Acids Res. 2017, 45, D535–D542. [Google Scholar] [CrossRef]
- Bortolaia, V.; Kaas, R.S.; Ruppe, E.; Roberts, M.C.; Schwarz, S.; Cattoir, V.; Philippon, A.; Allesoe, R.L.; Rebelo, A.R.; Florensa, A.F.; et al. ResFinder 4.0 for predictions of phenotypes from genotypes. J. Antimicrob. Chemother. 2020, 75, 3491–3500. [Google Scholar] [CrossRef]
- McArthur, A.G.; Waglechner, N.; Nizam, F.; Yan, A.; Azad, M.A.; Baylay, A.J.; Bhullar, K.; Canova, M.J.; De Pascale, G.; Ejim, L.; et al. The comprehensive antibiotic resistance database. Antimicrob. Agents Chemother. 2013, 57, 3348–3357. [Google Scholar] [CrossRef]
- Carattoli, A.; Zankari, E.; García-Fernández, A.; Voldby Larsen, M.; Lund, O.; Villa, L.; Møller Aarestrup, F.; Hasman, H. In silico detection and typing of plasmids using PlasmidFinder and plasmid multilocus sequence typing. Antimicrob. Agents Chemother. 2014, 58, 3895–3903. [Google Scholar] [CrossRef] [PubMed]
- Siguier, P.; Perochon, J.; Lestrade, L.; Mahillon, J.; Chandler, M. ISfinder: The reference centre for bacterial insertion sequences. Nucleic Acids Res. 2006, 34, D32–D36. [Google Scholar] [CrossRef] [PubMed]
- Johansson, M.H.K.; Bortolaia, V.; Tansirichaiya, S.; Aarestrup, F.M.; Roberts, A.P.; Petersen, T.N. Detection of mobile genetic elements associated with antibiotic resistance in Salmonella enterica using a newly developed web tool: MobileElementFinder. J. Antimicrob. Chemother. 2021, 76, 101–109. [Google Scholar] [CrossRef] [PubMed]
- Joensen, K.G.; Tetzschner, A.M.M.; Iguchi, A.; Aarestrup, F.M.; Scheutz, F. Rapid and Easy In Silico Serotyping of Escherichia coli Isolates by Use of Whole-Genome Sequencing Data. J. Clin. Microbiol. 2015, 53, 2410–2426. [Google Scholar] [CrossRef]
- Li, X.; Xie, Y.; Liu, M.; Tai, C.; Sun, J.; Deng, Z.; Ou, H.-Y. oriTfinder: A web-based tool for the identification of origin of transfers in DNA sequences of bacterial mobile genetic elements. Nucleic Acids Res. 2018, 46, W229–W234. [Google Scholar] [CrossRef]
- WHO. Global Action Plan on Antimicrobial Resistance; WHO: Geneva, Switzerland, 2015.
- Eckert, C.; Gautier, V.; Arlet, G. DNA sequence analysis of the genetic environment of various blaCTX-M genes. J. Antimicrob. Chemother. 2006, 57, 14–23. [Google Scholar] [CrossRef]
- Yagi, T.; Kurokawa, H.; Shibata, N.; Shibayama, K.; Arakawa, Y. A preliminary survey of extended-spectrum β-lactamases (ESBLs) in clinical isolates of Klebsiella pneumoniae and Escherichia coli in Japan. FEMS Microbiol. Lett. 2000, 184, 53–56. [Google Scholar] [CrossRef]
- McNeece, G.; Naughton, V.; Woodward, M.J.; Dooley, J.S.; Naughton, P.J. Array based detection of antibiotic resistance genes in Gram negative bacteria isolated from retail poultry meat in the UK and Ireland. Int. J. Food Microbiol. 2014, 179, 24–32. [Google Scholar] [CrossRef]
- Dallenne, C.; Da Costa, A.; Decre, D.; Favier, C.; Arlet, G. Development of a set of multiplex PCR assays for the detection of genes encoding important beta-lactamases in Enterobacteriaceae. J. Antimicrob. Chemother. 2010, 65, 490–495. [Google Scholar] [CrossRef]
- Colom, K.; Perez, J.; Alonso, R.; Fernandez-Aranguiz, A.; Larino, E.; Cisterna, R. Simple and reliable multiplex PCR assay for detection of blaTEM, bla(SHV) and blaOXA-1 genes in Enterobacteriaceae. FEMS Microbiol. Lett. 2003, 223, 147–151. [Google Scholar] [CrossRef]
- Levesque, C.; Piche, L.; Larose, C.; Roy, P.H. PCR mapping of integrons reveals several novel combinations of resistance genes. Antimicrob. Agents Chemother. 1995, 39, 185–191. [Google Scholar] [CrossRef] [PubMed]


| Source | N | Prevalence (%) | |||||||
|---|---|---|---|---|---|---|---|---|---|
| CTX-M-1 Subgroup | CTX-M-2 Subgroup | CTX-M-9 Subgroup a | TEM | OXA | SHV | KPC b | int1 | ||
| Clinical | 11 | 7 (64) | 0 | 1 (9) | 11 (100) | 5 (45) | 0 | 0 | 7 (64) |
| Hospital WW | 22 | 21 (95) | 0 | 0 | 13 (59) | 19 (86) | 0 | 9 (41) | 17 (77) |
| Urban Influent | 15 | 11 (73) | 0 | 4 (27) | 11 (73) | 10 (67) | 0 | 0 | 8 (53) |
| Treated effluent | 16 | 13 (81) | 0 | 5 (31) | 12 (75) | 13 (81) | 0 | 0 | 9 (56) |
| Total | 64 | 52 (81) | 0 | 10 (16) | 47 (73) | 47 (73) | 0 | 9 (14) | 41 (64) |
| CAZ | CRO | CTX | ERT | ESBL | blaCTX-M-1 | blaCTX-M-9 | blaOXA | blaTEM | blaKPC | |
|---|---|---|---|---|---|---|---|---|---|---|
| CAZ | 1 | |||||||||
| CRO | 0.1176 0.3548 | 1 | ||||||||
| CTX | 0.1176 0.3548 | 1.0000 * 0.0000 | 1 | |||||||
| ERT | 0.2925 * 0.0190 | −0.0949 0.4557 | −0.0949 0.4557 | 1 | ||||||
| ESBL | −0.3945 * 0.0013 | 0.2361 0.0604 | 0.2361 0.0604 | −0.3190 * 0.0102 | 1 | |||||
| blaCTX-M-1 | 0.1732 0.1711 | 0.4616 * 0.0001 | 0.4616 * 0.0001 | 0.0066 0.9585 | 0.2707 * 0.0305 | 1 | ||||
| blaCTX-M-9 | −0.4326 * 0.0004 | 0.0954 0.4532 | 0.0954 0.4532 | −0.1960 0.1205 | 0.0593 0.6417 | −0.2343 0.0624 | 1 | |||
| blaOXA | 0.1508 0.2342 | 0.0340 0.7897 | 0.0340 0.7897 | −0.0076 0.9542 | −0.1396 0.2713 | −0.0170 0.8940 | −0.1309 0.3024 | 1 | ||
| blaTEM | −0.2093 0.0969 | 0.0340 0.7897 | 0.0340 0.7897 | −0.2886 * 0.0207 | 0.1440 0.2563 | 0.0736 0.5630 | 0.1614 0.2027 | −0.2015 0.1103 | 1 | |
| blaKPC | 0.3346 * 0.0069 | 0.0897 0.4808 | 0.0897 0.4808 | 0.6497 * 0.0000 | −0.2505 * 0.0459 | 0.1943 0.1239 | −0.1741 0.1689 | 0.1415 0.2647 | −0.3673 * 0.0028 | 1 |
| Antibiotic Resistance Genes | Location | Accession # | Resistance Target |
|---|---|---|---|
| aac(6’)-Ib-cr | Plasmid | DQ303918 | Aminoglycoside and fluoroquinolone resistance |
| aadA5 | Plasmid | AF137361 | Aminoglycoside resistance |
| aph(3’’)-Ib | Plasmid | AF321551 | Aminoglycoside resistance |
| aph(6)-Id | Plasmid | M28829 | Aminoglycoside resistance |
| blaCTX-M-15 | Plasmid | AY044436 | Beta-lactam resistance |
| blaOXA-1 | Plasmid | HQ170510 | Beta-lactam resistance |
| blaKPC-2 | Plasmid | AY034847 | Beta-lactam resistance |
| blaTEM-350 | Plasmid | WP045286946 | Beta-lactam resistance |
| mdf(A) | Chromosomal | Y08743 | Macrolide resistance |
| mph(A) | Plasmid | D16251 | Macrolide resistance |
| catB3 | Plasmid | AJ009818 | Phenicol resistance |
| sul1 | Plasmid | U12338 | Sulphonamide resistance |
| tet(A) | Plasmid | AJ517790 | Tetracycline resistance |
| tet(B) | Chromosomal | AF32677 | Tetracycline resistance |
| dfrA17 | Plasmid | FJ460238 | Trimethoprim resistance |
| sitABCD | Plasmid | AY598030 | Disinfectant resistance |
| qacE | Plasmid | X68232 | Disinfectant resistance |
| bacA | Chromosomal | 3002986 | Peptide antibiotic resistance |
| mdtG | Chromosomal | 3001329 | MFS antibiotic efflux pump |
| mdtH | Chromosomal | 3001216 | MFS antibiotic efflux pump |
| mdtP | Chromosomal | 3003550 | MFS antibiotic efflux pump |
| mdtN | Chromosomal | 3003548 | MFS antibiotic efflux pump |
| tolC | Chromosomal | 3000237 | MFS antibiotic efflux pump |
| emrA | Chromosomal | 3000027 | MFS antibiotic efflux pump |
| emrB | Chromosomal | 3000074 | MFS antibiotic efflux pump |
| emrK | Chromosomal | 3000206 | MFS antibiotic efflux pump |
| emrY | Chromosomal | 3000254 | MFS antibiotic efflux pump |
| evgA | Chromosomal | 3000833 | MFS antibiotic efflux pump |
| H-NS | Chromosomal | 3000676 | MFS antibiotic efflux pump |
| ampH | Chromosomal | 3004612 | ampC-type beta-lactamase |
| ampC1 | Chromosomal | 3004611 | ampC-type beta-lactamase |
| cpxA | Chromosomal | 3000830 | RND Efflux pump |
| gadX | Chromosomal | 3000508 | RND Efflux pump |
| mdtE | Chromosomal | 3000795 | RND Efflux pump |
| mdtF | Chromosomal | 3000796 | RND efflux pump |
| acrA | Chromosomal | 3004043 | RND Efflux pump |
| acrB | Chromosomal | 3000216 | RND Efflux pump |
| acrD | Chromosomal | 3000491 | RND Efflux pump |
| baeS | Chromosomal | 3000829 | RND Efflux pump |
| baeR | Chromosomal | 3000828 | RND Efflux pump |
| marA | Chromosomal | 3000263 | RND Efflux pump |
| yojI | Chromosomal | 3003952 | Peptide antibiotic efflux pump |
| pmrF | Chromosomal | 3003578 | Phosphoethanolamine transferase |
| kdpE | Chromosomal | 3003841 | Aminoglycoside efflux pump |
| msbA | Chromosomal | 3003950 | Nitroimidazole efflux pump |
| Point mutations | PMID | ||
| parC AGC-> ATC | 8851598 | Quinolone resistance | |
| parE TCG -> GCG | 28598203 | Quinolone resistance | |
| gyrA TCG -> TTG | 8891148 | Quinolone resistance | |
| gyrA GAC-> AAC | 12654733 | Quinolone resistance |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liedhegner, E.; Bojar, B.; Beattie, R.E.; Cahak, C.; Hristova, K.R.; Skwor, T. Similarities in Virulence and Extended Spectrum Beta-Lactamase Gene Profiles among Cefotaxime-Resistant Escherichia coli Wastewater and Clinical Isolates. Antibiotics 2022, 11, 260. https://doi.org/10.3390/antibiotics11020260
Liedhegner E, Bojar B, Beattie RE, Cahak C, Hristova KR, Skwor T. Similarities in Virulence and Extended Spectrum Beta-Lactamase Gene Profiles among Cefotaxime-Resistant Escherichia coli Wastewater and Clinical Isolates. Antibiotics. 2022; 11(2):260. https://doi.org/10.3390/antibiotics11020260
Chicago/Turabian StyleLiedhegner, Elizabeth, Brandon Bojar, Rachelle E. Beattie, Caitlin Cahak, Krassimira R. Hristova, and Troy Skwor. 2022. "Similarities in Virulence and Extended Spectrum Beta-Lactamase Gene Profiles among Cefotaxime-Resistant Escherichia coli Wastewater and Clinical Isolates" Antibiotics 11, no. 2: 260. https://doi.org/10.3390/antibiotics11020260
APA StyleLiedhegner, E., Bojar, B., Beattie, R. E., Cahak, C., Hristova, K. R., & Skwor, T. (2022). Similarities in Virulence and Extended Spectrum Beta-Lactamase Gene Profiles among Cefotaxime-Resistant Escherichia coli Wastewater and Clinical Isolates. Antibiotics, 11(2), 260. https://doi.org/10.3390/antibiotics11020260








