Crumbling the Castle: Targeting DNABII Proteins for Collapsing Bacterial Biofilms as a Therapeutic Approach to Treat Disease and Combat Antimicrobial Resistance
Abstract
:1. Introduction
2. Bacterial Biofilms and Antimicrobial Resistance
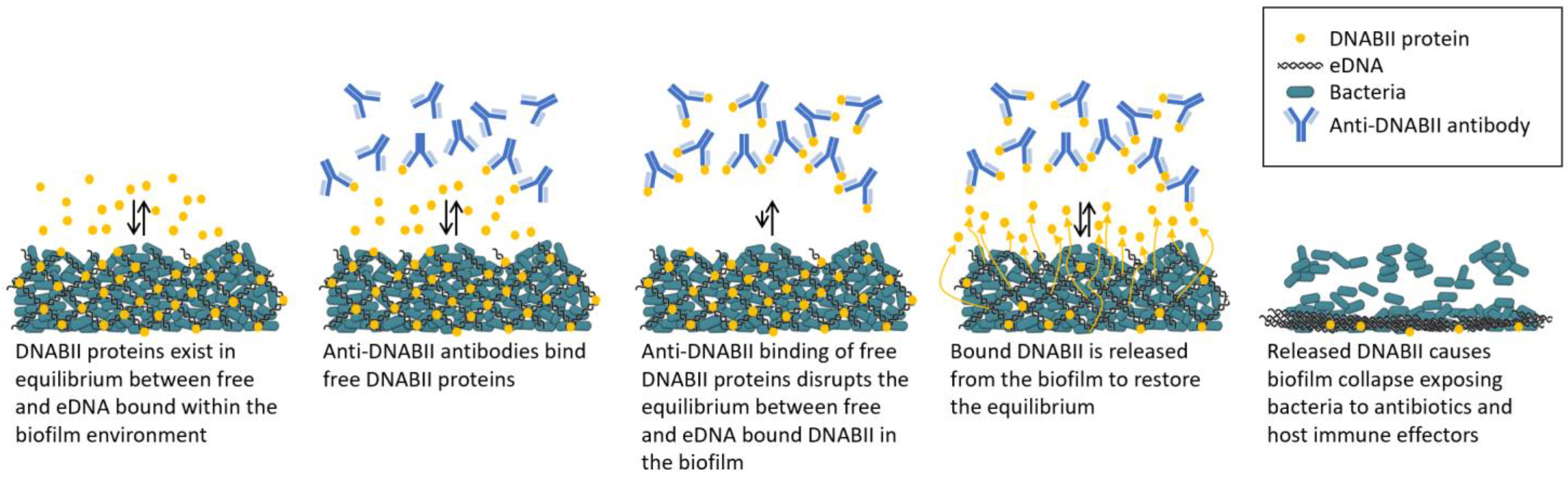
3. Targeting DNA-Binding Tip Regions of DNABII Proteins to Rapidly Collapse Biofilms
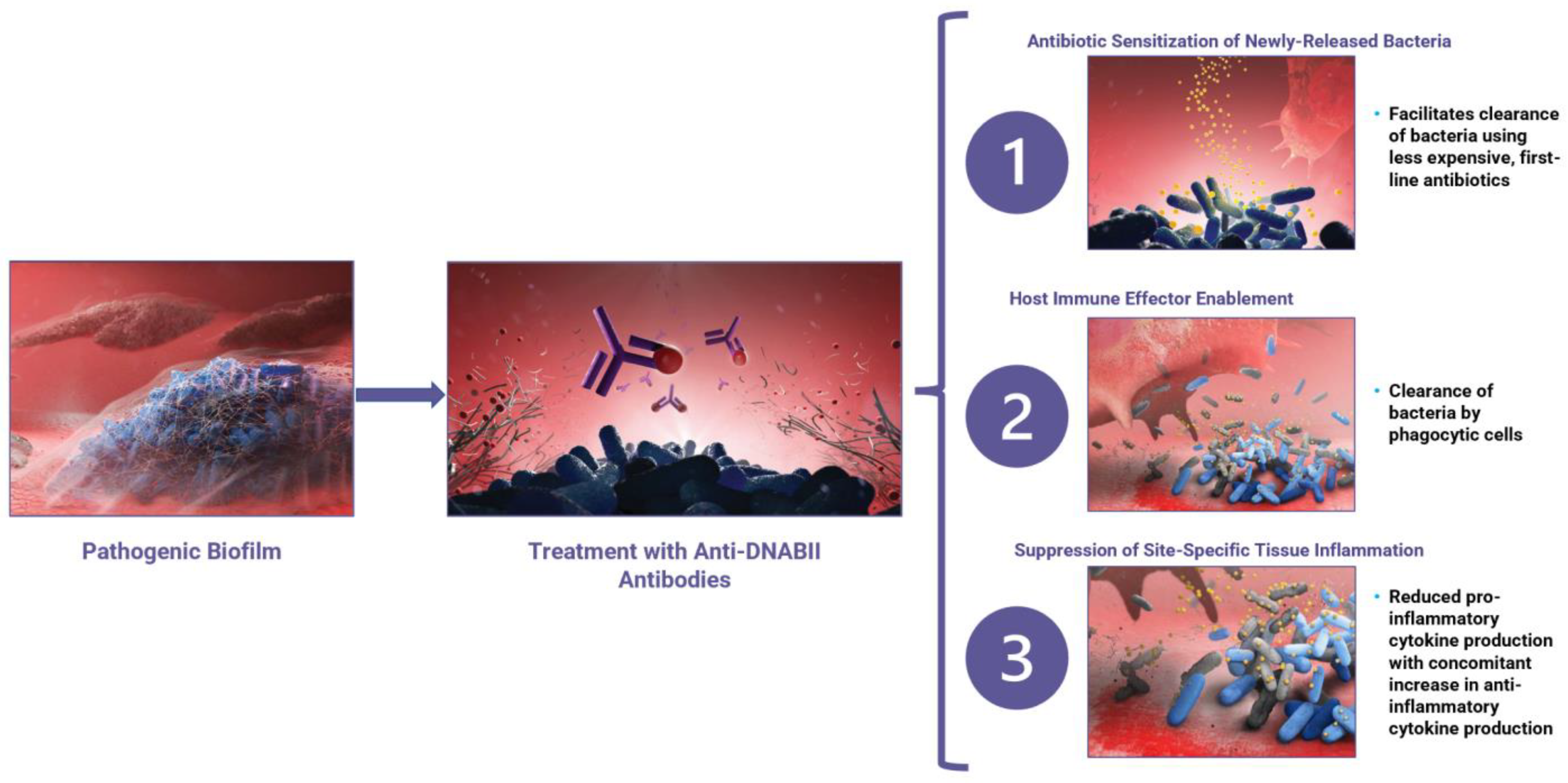
4. Tripartite Mode of Action
4.1. Mode of Action 1: Anti-DNABII-Antibody-Induced Rapid Biofilm Collapse Sensitizes Bacteria to Antibiotics
4.2. Mode of Action 2: Anti-DNABII-Antibody-Induced Rapid Biofilm Collapse Enhances Innate Immune System Clearance of Bacteria
4.3. Mode of Action 3: Anti-DNABII Antibody Treatment Promotes Establishment of an Anti-Inflammatory State
5. Clinical Benefits of Anti-DNABII Antibodies
6. CMTX-101: A Nontraditional Treatment for Attacking Bacterial Biofilms
- Promote the appropriate use of antibacterial agents: When CMTX-101 disrupts biofilms, NRel bacteria are released with increased sensitivity to multiple antibiotic classes. This enables the effective use of older antibacterial agents, whose use is currently limited by AMR, and reduces the pressure to use newer antibacterial agents as first- or second-line therapies.
- Improve patient outcomes: Biofilms reduce the access of antibacterial agents, as well as macrophages and neutrophils, to bacteria; thus, treatment generally requires higher doses and longer courses of antibiotic therapy to achieve a clinical cure. CMTX-101-mediated biofilm collapse sensitizes bacteria to antibiotic action, enhances bacterial clearance by neutrophils and macrophages, and promotes the establishment of an anti-inflammatory environment, increasing the likelihood of improved outcomes when treating bacterial biofilm infections and enabling shorter courses of antibacterial therapy.
- Reduce microbial resistance: The mechanism by which CMTX-101 disrupts biofilms suggests a low potential for resistance development due to two factors. First, the DNABII proteins are the only nucleoid-associated proteins that can stabilize the biofilm [52]; however, they are also essential proteins for intracellular DNA-processing where mutations in ihf or hu alleles severely reduce bacterial fitness [67,68]. Second, the mechanism of biofilm disruption by CMTX-101 does not exert direct selective pressure on bacterial survival, further reducing the probability of resistance development.
- Decrease the spread of infections caused by multidrug-resistant organisms: CMTX-101 is active against biofilms formed by all bacterial pathogens examined to date, including ESKAPEE and multidrug-resistant (MDR) pathogens. We have specifically observed that biofilms formed from MRSA and tobramycin-resistant P. aeruginosa are rapidly disrupted by CMTX-101 (unpublished data). NRel bacteria following CMTX-101 treatment are sensitized to the actions of many antibacterial agents, allowing the increased effectiveness of first-line antibiotics and the reservation of second- and third-line antibiotics for the most serious infections with determined genotypic resistance. Hence, effective killing with first-line antibiotics used in combination with CMTX-101 may reduce the probability of further development and spread of bacterial resistance.
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sharma, P.; Towse, A. New Drugs to Tackle Antimicrobial Resistance: Analysis of EU Policy Options. Available online: https://www.ohe.org/publications/new-drugs-tackle-antimicrobial-resistance-analysis-eu-policy-options (accessed on 27 November 2021).
- Fact Sheet: Antimicrobial Resistance. Available online: https://www.who.int/news-room/fact-sheets/detail/antimicrobial-resistance#:~:text=Antimicrobial%20Resistance%20(AMR)%20occurs%20when%20bacteria,%20viruses,%20fungi,risk%20of%20disease%20spread,%20severe%20illness%20and%20death (accessed on 19 October 2021).
- CDC’s Antibiotic Resistance Threats in the United States. 2019. Available online: https://www.cdc.gov/drugresistance/biggest-threats.html (accessed on 19 October 2021).
- Nelson, R.E.; Hatfield, K.M.; Wolford, H.; Samore, M.H.; Scott, R.D.; Reddy, S.C.; Olubajo, B.; Paul, P.; Jernigan, J.A.; Baggs, J. National Estimates of Healthcare Costs Associated with Multidrug-Resistant Bacterial Infections Among Hospitalized Patients in the United States. Clin. Infect. Dis. 2021, 72, S17–S26. [Google Scholar] [CrossRef]
- ECDC/EMEA Joint Technical Report. The Bacterial Challenge: Time to React. A Call to Narrow the Gap between Multidrug-Resistant Bacteria in the EU and the Development of New Antibacterial Agents. Available online: https://www.fdanews.com/ext/resources/files/archives/e/EMEA-576176-2009.pdf (accessed on 27 November 2021).
- O’Neil, J. Review on Antimicrobial Resistance Antimicrobial Resistance: Tackling a Crisis for the Health and Wealth of Nations. Available online: https://amr-review.org/sites/default/files/AMR%20Review%20Paper%20-%20Tackling%20a%20crisis%20for%20the%20health%20and%20wealth%20of%20nations_1.pdf (accessed on 27 November 2021).
- de Kraker, M.E.; Stewardson, A.J.; Harbarth, S. Will 10 Million People Die a Year due to Antimicrobial Resistance by 2050? PLoS Med. 2016, 13, e1002184. [Google Scholar] [CrossRef] [Green Version]
- Generating Antibiotic Incentives Now. Available online: https://www.fda.gov/media/110982/download (accessed on 27 November 2021).
- Duffy, E.M.; Buurman, E.T.; Chiang, S.L.; Cohen, N.R.; Uria-Nickelsen, M.; Alm, R.A. The CARB-X portfolio of nontraditional antibacterial products. ACS Infect. Dis. 2021, 7, 2043–2049. [Google Scholar] [CrossRef]
- Davies, D. Understanding biofilm resistance to antibacterial agents. Nat. Rev. Drug Discov. 2003, 2, 114–122. [Google Scholar] [CrossRef]
- Mah, T.F.; O’Toole, G.A. Mechanisms of biofilm resistance to antimicrobial agents. Trends Microbiol. 2001, 9, 34–39. [Google Scholar] [CrossRef]
- Vestby, L.K.; Gronseth, T.; Simm, R.; Nesse, L.L. Bacterial biofilm and its role in the pathogenesis of disease. Antibiotics 2020, 9, 59. [Google Scholar] [CrossRef] [Green Version]
- Lebeaux, D.; Ghigo, J.M.; Beloin, C. Biofilm-related infections: Bridging the gap between clinical management and fundamental aspects of recalcitrance toward antibiotics. Microbiol. Mol. Biol. Rev. 2014, 78, 510–543. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mah, T.F. Biofilm-specific antibiotic resistance. Future Microbiol. 2012, 7, 1061–1072. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nickel, J.C.; Ruseska, I.; Wright, J.B.; Costerton, J.W. Tobramycin resistance of Pseudomonas aeruginosa cells growing as a biofilm on urinary catheter material. Antimicrob. Agents Chemother. 1985, 27, 619–624. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sharma, D.; Misba, L.; Khan, A.U. Antibiotics versus biofilm: An emerging battleground in microbial communities. Antimicrob. Resist. Infect. Control 2019, 8, 76. [Google Scholar] [CrossRef] [PubMed]
- Boisvert, A.A.; Cheng, M.P.; Sheppard, D.C.; Nguyen, D. Microbial biofilms in pulmonary and critical care diseases. Ann. Am. Thorac. Soc. 2016, 13, 1615–1623. [Google Scholar] [CrossRef] [Green Version]
- Ciofu, O.; Rojo-Molinero, E.; Macia, M.D.; Oliver, A. Antibiotic treatment of biofilm infections. APMIS 2017, 125, 304–319. [Google Scholar] [CrossRef]
- Gunn, J.S.; Bakaletz, L.O.; Wozniak, D.J. What’s on the outside matters: The role of the extracellular polymeric substance of Gram-negative biofilms in evading host immunity and as a target for therapeutic intervention. J. Biol. Chem. 2016, 291, 12538–12546. [Google Scholar] [CrossRef] [Green Version]
- Koo, H.; Allan, R.N.; Howlin, R.P.; Stoodley, P.; Hall-Stoodley, L. Targeting microbial biofilms: Current and prospective therapeutic strategies. Nat. Rev. Microbiol. 2017, 15, 740–755. [Google Scholar] [CrossRef]
- Rabin, N.; Zheng, Y.; Opoku-Temeng, C.; Du, Y.; Bonsu, E.; Sintim, H.O. Biofilm formation mechanisms and targets for developing antibiofilm agents. Future Med. Chem. 2015, 7, 493–512. [Google Scholar] [CrossRef]
- Roy, R.; Tiwari, M.; Donelli, G.; Tiwari, V. Strategies for combating bacterial biofilms: A focus on anti-biofilm agents and their mechanisms of action. Virulence 2018, 9, 522–554. [Google Scholar] [CrossRef] [PubMed]
- Flemming, H.C.; Wingender, J. The biofilm matrix. Nat. Rev. Microbiol. 2010, 8, 623–633. [Google Scholar] [CrossRef] [PubMed]
- Santos, A.; Galdino, A.C.M.; Mello, T.P.; Ramos, L.S.; Branquinha, M.H.; Bolognese, A.M.; Columbano Neto, J.; Roudbary, M. What are the advantages of living in a community? A microbial biofilm perspective! Memórias Inst. Oswaldo Cruz 2018, 113, e180212. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yan, J.; Bassler, B.L. Surviving as a community: Antibiotic tolerance and persistence in bacterial biofilms. Cell Host Microbe 2019, 26, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Flemming, H.C.; Neu, T.R.; Wozniak, D.J. The EPS matrix: The “house of biofilm cells”. J. Bacteriol. 2007, 189, 7945–7947. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fong, J.N.C.; Yildiz, F.H. Biofilm matrix proteins. Microbiol. Spectr. 2015, 3, MB-0004-2014. [Google Scholar] [CrossRef] [Green Version]
- Stewart, P.S. Antimicrobial tolerance in biofilms. Microbiol. Spectr. 2015, 3, MB-0010-2014. [Google Scholar] [CrossRef] [Green Version]
- Romling, U.; Balsalobre, C. Biofilm infections, their resilience to therapy and innovative treatment strategies. J. Intern. Med. 2012, 272, 541–561. [Google Scholar] [CrossRef]
- Ghasemi, M.; Hense, B.A.; Eberl, H.J.; Kuttler, C. Simulation-based exploration of quorum sensing triggered resistance of biofilms to antibiotics. Bull. Math. Biol. 2018, 80, 1736–1775. [Google Scholar] [CrossRef] [PubMed]
- Orazi, G.; O’Toole, G.A. “It Takes a Village”: Mechanisms underlying antimicrobial recalcitrance of polymicrobial biofilms. J. Bacteriol. 2020, 202, e00530. [Google Scholar] [CrossRef]
- Stewart, P.S.; Costerton, J.W. Antibiotic resistance of bacteria in biofilms. Lancet 2001, 358, 135–138. [Google Scholar] [CrossRef]
- Whitchurch, C.B.; Tolker-Nielsen, T.; Ragas, P.C.; Mattick, J.S. Extracellular DNA required for bacterial biofilm formation. Science 2002, 295, 1487. [Google Scholar] [CrossRef] [PubMed]
- Gloag, E.S.; Turnbull, L.; Huang, A.; Vallotton, P.; Wang, H.; Nolan, L.M.; Mililli, L.; Hunt, C.; Lu, J.; Osvath, S.R.; et al. Self-organization of bacterial biofilms is facilitated by extracellular DNA. Proc. Natl. Acad. Sci. USA 2013, 110, 11541–11546. [Google Scholar] [CrossRef] [Green Version]
- Lewenza, S. Extracellular DNA-induced antimicrobial peptide resistance mechanisms in Pseudomonas aeruginosa. Front. Microbiol. 2013, 4, 21. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Molin, S.; Tolker-Nielsen, T. Gene transfer occurs with enhanced efficiency in biofilms and induces enhanced stabilisation of the biofilm structure. Curr. Opin. Biotechnol. 2003, 14, 255–261. [Google Scholar] [CrossRef]
- Mulcahy, H.; Charron-Mazenod, L.; Lewenza, S. Extracellular DNA chelates cations and induces antibiotic resistance in Pseudomonas aeruginosa biofilms. PLoS Pathog. 2008, 4, e1000213. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pinchuk, G.E.; Ammons, C.; Culley, D.E.; Li, S.M.; McLean, J.S.; Romine, M.F.; Nealson, K.H.; Fredrickson, J.K.; Beliaev, A.S. Utilization of DNA as a sole source of phosphorus, carbon, and energy by Shewanella spp.: Ecological and physiological implications for dissimilatory metal reduction. Appl. Environ. Microbiol. 2008, 74, 1198–1208. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Novotny, L.A.; Amer, A.O.; Brockson, M.E.; Goodman, S.D.; Bakaletz, L.O. Structural stability of Burkholderia cenocepacia biofilms is reliant on eDNA structure and presence of a bacterial nucleic acid binding protein. PLoS ONE 2013, 8, e67629. [Google Scholar] [CrossRef]
- Buzzo, J.R.; Devaraj, A.; Gloag, E.S.; Jurcisek, J.A.; Robledo-Avila, F.; Kesler, T.; Wilbanks, K.; Mashburn-Warren, L.; Balu, S.; Wickham, J.; et al. Z-form extracellular DNA is a structural component of the bacterial biofilm matrix. Cell 2021, 184, 5740–5758. [Google Scholar] [CrossRef] [PubMed]
- Kamashev, D.; Rouviere-Yaniv, J. The histone-like protein HU binds specifically to DNA recombination and repair intermediates. EMBO J. 2000, 19, 6527–6535. [Google Scholar] [CrossRef] [Green Version]
- Rice, P.A.; Yang, S.; Mizuuchi, K.; Nash, H.A. Crystal structure of an IHF-DNA complex: A protein-induced DNA U-turn. Cell 1996, 87, 1295–1306. [Google Scholar] [CrossRef] [Green Version]
- Devaraj, A.; Buzzo, J.R.; Mashburn-Warren, L.; Gloag, E.S.; Novotny, L.A.; Stoodley, P.; Bakaletz, L.O.; Goodman, S.D. The extracellular DNA lattice of bacterial biofilms is structurally related to Holliday junction recombination intermediates. Proc. Natl. Acad. Sci. USA 2019, 116, 25068–25077. [Google Scholar] [CrossRef] [Green Version]
- Devaraj, A.; Justice, S.S.; Bakaletz, L.O.; Goodman, S.D. DNABII proteins play a central role in UPEC biofilm structure. Mol. Microbiol. 2015, 96, 1119–1135. [Google Scholar] [CrossRef] [Green Version]
- Goodman, S.D.; Obergfell, K.P.; Jurcisek, J.A.; Novotny, L.A.; Downey, J.S.; Ayala, E.A.; Tjokro, N.; Li, B.; Justice, S.S.; Bakaletz, L.O. Biofilms can be dispersed by focusing the immune system on a common family of bacterial nucleoid-associated proteins. Mucosal. Immunol. 2011, 4, 625–637. [Google Scholar] [CrossRef] [Green Version]
- Gustave, J.E.; Jurcisek, J.A.; McCoy, K.S.; Goodman, S.D.; Bakaletz, L.O. Targeting bacterial integration host factor to disrupt biofilms associated with cystic fibrosis. J. Cyst. Fibros. 2013, 12, 384–389. [Google Scholar] [CrossRef] [Green Version]
- Idicula, W.K.; Jurcisek, J.A.; Cass, N.D.; Ali, S.; Goodman, S.D.; Elmaraghy, C.A.; Jatana, K.R.; Bakaletz, L.O. Identification of biofilms in post-tympanostomy tube otorrhea. Laryngoscope 2016, 126, 1946–1951. [Google Scholar] [CrossRef] [PubMed]
- Novotny, L.A.; Goodman, S.D.; Bakaletz, L.O. Redirecting the immune response towards immunoprotective domains of a DNABII protein resolves experimental otitis media. NPJ Vaccines 2019, 4, 43. [Google Scholar] [CrossRef] [PubMed]
- Brockson, M.E.; Novotny, L.A.; Mokrzan, E.M.; Malhotra, S.; Jurcisek, J.A.; Akbar, R.; Devaraj, A.; Goodman, S.D.; Bakaletz, L.O. Evaluation of the kinetics and mechanism of action of anti-integration host factor-mediated disruption of bacterial biofilms. Mol. Microbiol. 2014, 93, 1246–1258. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Novotny, L.A.; Jurcisek, J.A.; Goodman, S.D.; Bakaletz, L.O. Monoclonal antibodies against DNA-binding tips of DNABII proteins disrupt biofilms in vitro and induce bacterial clearance in vivo. EBioMedicine 2016, 10, 33–44. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brandstetter, K.A.; Jurcisek, J.A.; Goodman, S.D.; Bakaletz, L.O.; Das, S. Antibodies directed against integration host factor mediate biofilm clearance from Nasopore. Laryngoscope 2013, 123, 2626–2632. [Google Scholar] [CrossRef] [Green Version]
- Devaraj, A.; Buzzo, J.; Rocco, C.J.; Bakaletz, L.O.; Goodman, S.D. The DNABII family of proteins is comprised of the only nucleoid associated proteins required for nontypeable Haemophilus influenzae biofilm structure. Microbiologyopen 2018, 7, e00563. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Martyn, L.; Sethia, R.; Chon, R.; Novotny, L.; Goodman, S.D.; Elmaraghy, C.; Bakaletz, L.O. Antibodies against the DNABII protein integration host factor (IHF) inhibit sinus implant biofilms. Laryngoscope 2020, 130, 1364–1371. [Google Scholar] [CrossRef] [PubMed]
- Mokrzan, E.M.; Ahearn, C.P.; Buzzo, J.R.; Novotny, L.A.; Zhang, Y.; Goodman, S.D.; Bakaletz, L.O. Nontypeable Haemophilus influenzae newly released (NRel) from biofilms by antibody-mediated dispersal versus antibody-mediated disruption are phenotypically distinct. Biofilm 2020, 2, 100039. [Google Scholar] [CrossRef]
- Novotny, L.A.; Goodman, S.D.; Bakaletz, L.O. Targeting a bacterial DNABII protein with a chimeric peptide immunogen or humanised monoclonal antibody to prevent or treat recalcitrant biofilm-mediated infections. EBioMedicine 2020, 59, 102867. [Google Scholar] [CrossRef]
- Rocco, C.J.; Bakaletz, L.O.; Goodman, S.D. Targeting the HUbeta protein prevents Porphyromonas gingivalis from entering into preexisting biofilms. J. Bacteriol. 2018, 200, e00790. [Google Scholar] [CrossRef] [Green Version]
- Rocco, C.J.; Davey, M.E.; Bakaletz, L.O.; Goodman, S.D. Natural antigenic differences in the functionally equivalent extracellular DNABII proteins of bacterial biofilms provide a means for targeted biofilm therapeutics. Mol. Oral Microbiol. 2017, 32, 118–130. [Google Scholar] [CrossRef] [PubMed]
- Freire, M.O.; Devaraj, A.; Young, A.; Navarro, J.B.; Downey, J.S.; Chen, C.; Bakaletz, L.O.; Zadeh, H.H.; Goodman, S.D. A bacterial-biofilm-induced oral osteolytic infection can be successfully treated by immuno-targeting an extracellular nucleoid-associated protein. Mol. Oral Microbiol. 2017, 32, 74–88. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Novotny, L.A.; Chiang, T.; Goodman, S.D.; Elmaraghy, C.A.; Bakaletz, L.O. Humanized anti-DNABII Fab fragments plus ofloxacin eradicated biofilms in experimental otitis media. Laryngoscope 2021, 131, E2698–E2704. [Google Scholar] [CrossRef]
- Rood, K.M.; Buhimschi, I.A.; Jurcisek, J.A.; Summerfield, T.L.; Zhao, G.; Ackerman, W.E.; Wang, W.; Rumpf, R.W.; Thung, S.F.; Bakaletz, L.O.; et al. Skin microbiota in obese women at risk for surgical site infection after Cesarean delivery. Sci. Rep. 2018, 8, 8756. [Google Scholar] [CrossRef]
- Kurbatfinski, N.; Goodman, S.D.; Bakaletz, L.O. A humanized monoclonal antibody potentiates killing by antibiotics of diverse biofilm-forming respiratory pathogens. Antimicrob. Agents Chemother. 2022, 66, e01877-21. [Google Scholar] [CrossRef] [PubMed]
- Deveraj, A.; González, J.F.; Eichar, B.; Thilliez, G.; Kingsley, R.A.; Baker, S.; Allard, M.W.; Bakaletz, L.O.; Gunn, J.S.; Goodman, S.D. Enhanced biofilm and extracellular matrix production by chronic carriage versus acute isolates of Salmonella typhi. PLoS Pathog. 2021, 17, e1009209. [Google Scholar] [CrossRef]
- Donlan, R.M.; Costerton, J.W. Biofilms: Survival mechanisms of clinically relevant microorganisms. Clin. Microbiol. Rev. 2002, 15, 167–193. [Google Scholar] [CrossRef] [Green Version]
- Ammons, M.C. Anti-biofilm strategies and the need for innovations in wound care. Recent Pat. Antiinfect. Drug Discov. 2010, 5, 10–17. [Google Scholar] [CrossRef]
- Fronzo, C. An early, biofilm-focused wound management approach. J. Wound Care 2019, 28, 524–526. [Google Scholar] [CrossRef] [Green Version]
- Peel, T.N.; de Steiger, R. How to manage treatment failure in prosthetic joint infection. Clin. Microbiol. Infect. 2020, 26, 1473–1480. [Google Scholar] [CrossRef]
- Ferrandiz, M.J.; Carreno, D.; Ayora, S.; de la Campa, A.G. HU of Streptococcus pneumoniae is essential for the preservation of DNA supercoiling. Front. Microbiol. 2018, 9, 493. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Leite, B.; Werle, C.H.; Carmo, C.P.D.; Nobrega, D.B.; Milanez, G.P.; Culler, H.F.; Sircili, M.P.; Alvarez-Martinez, C.E.; Brocchi, M. Integration host factor is important for biofilm formation by Salmonella enterica Enteritidis. Pathog. Dis. 2017, 75, ftx074. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Watnick, P.; Kolter, R. Biofilm, city of microbes. J. Bacteriol. 2000, 182, 2675–2679. [Google Scholar] [CrossRef] [PubMed] [Green Version]

| Pathogen | IHF | HU | ||
|---|---|---|---|---|
| α-Tip 20-Mer Peptide (% Identical) | β-Tip 20-Mer Peptide (% Identical) | α-Tip 20-Mer Peptide (% Identical) | β-Tip 20-Mer Peptide (% Identical) | |
| WHO Priority 1: Critical | ||||
| Acinetobacter baumannii | 90 | 85 | 80 | 80 |
| Pseudomonas aeruginosa | 85 | 90 | 75 | 78 |
| Enterobacteriaceae (Serratia marcescens) | 75 | 80 | 47 | 79 |
| WHO Priority 2: High | ||||
| Enterococcus faecium | 45 * | 50 * | 47 | 45 |
| Staphylococcus aureus | NA | NA | 78 | 67 |
| Helicobacter pylori‡ | ND | ND | NS ‡ | NS ‡ |
| Campylobacter spp. (C. jejuni) | 75 | 80 | 47 | NS |
| Salmonella spp. (S. enterica) | 75 | 80 | 47 | 45 |
| Neisseria gonorrhoeae | 80 | 65 | 45 | 56 |
| WHO Priority 3: Medium | ||||
| Streptococcus pneumoniae | NA | NA | 78 | 86 |
| Haemophilus influenzae | 100 | 100 | 47 | 100 |
| Shigella spp. (S. dysenteriae) | 90 | 80 | 47 | 45 |
| Other | ||||
| Burkholderia cenocepacia | 85 | 80 | 42 | 50 |
| Escherichia coli | 95 | 80 | 75 | 80 |
| Klebsiella pneumoniae | 95 | 80 | 80 | 80 |
| Enterococcus faecalis | 75 | 80 | 47 | 45 |
| Mycobacterium tuberculosis | 75 | 75 | 47 | 45 |
| Aggregatibacter actinomycetemcomitans | 90 | 80 | 90 | 80 |
| Moraxella catarrhalis | 80 | 85 | 53 | 55 |
| Biofilm | In Vitro | Ex Vivo | In Vivo | Source [Reference] |
|---|---|---|---|---|
| Nontypeable Haemophilus influenzae |  |  |  | Clinical isolate [49,51,61] |
| Streptococcus pneumoniae |  | Strain 1121 [45,61] | ||
| Moraxella catarrhalis |  | Strain 7169 [50] | ||
| Streptococcus mutans |  | Strain UA159 [45] | ||
| Staphylococcus epidermidis |  | Clinical strain 1618 [45] | ||
| Neisseria gonorrhoeae |  | Strain 1291 [45] | ||
| Burkholderia cenocepacia |  | Multiple strains [39,50,61] | ||
| Aggregatibacter actinomycetemcomitans |  |  | Clinical strain D7S-1 [58] | |
| Porphyromonas gingivalis |  | Strain 381 [57] | ||
| Streptococcus gordonii |  | Chalis CH1 [57] | ||
| Streptococcus mitis |  | ATCC 33399 [56] | ||
| Streptococcus cristatus |  | ATCC 49999 [56] | ||
| Salmonella typhi |  | Multiple strains [62] | ||
| Streptococcus oralis |  | ATCC 10557 [56] | ||
| Enterococcus faecium |  | Unpublished data | ||
| Staphylococcus aureus (MRSA, MSSA) |  |  | ATCC 29213 (MSSA) [45,61]; Unpublished data (MRSA) | |
| Klebsiella pneumoniae |  | Unpublished data | ||
| Acinetobacter baumannii |  | Strain 17978 [61] | ||
| Pseudomonas aeruginosa |  |  | ATCC 27853 [45]; Strain 142-1 [61];Unpublished data | |
| Enterobacter spp. |  | Unpublished data | ||
| Escherichia coli |  | Strain MG1655; Uropathogenic strain UTT89 [45] | ||
| Mycobacterium smegmatis, M. abscessus |  | Unpublished data | ||
| Polymicrobial |  | Cystic Fibrosis sputum [46] Cesarean section wound [60] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rogers, J.V.; Hall, V.L.; McOsker, C.C. Crumbling the Castle: Targeting DNABII Proteins for Collapsing Bacterial Biofilms as a Therapeutic Approach to Treat Disease and Combat Antimicrobial Resistance. Antibiotics 2022, 11, 104. https://doi.org/10.3390/antibiotics11010104
Rogers JV, Hall VL, McOsker CC. Crumbling the Castle: Targeting DNABII Proteins for Collapsing Bacterial Biofilms as a Therapeutic Approach to Treat Disease and Combat Antimicrobial Resistance. Antibiotics. 2022; 11(1):104. https://doi.org/10.3390/antibiotics11010104
Chicago/Turabian StyleRogers, James V., Veronica L. Hall, and Charles C. McOsker. 2022. "Crumbling the Castle: Targeting DNABII Proteins for Collapsing Bacterial Biofilms as a Therapeutic Approach to Treat Disease and Combat Antimicrobial Resistance" Antibiotics 11, no. 1: 104. https://doi.org/10.3390/antibiotics11010104
APA StyleRogers, J. V., Hall, V. L., & McOsker, C. C. (2022). Crumbling the Castle: Targeting DNABII Proteins for Collapsing Bacterial Biofilms as a Therapeutic Approach to Treat Disease and Combat Antimicrobial Resistance. Antibiotics, 11(1), 104. https://doi.org/10.3390/antibiotics11010104





