Antimicrobial Resistance Profiles of Human Commensal Neisseria Species
Abstract
1. Introduction
2. Results
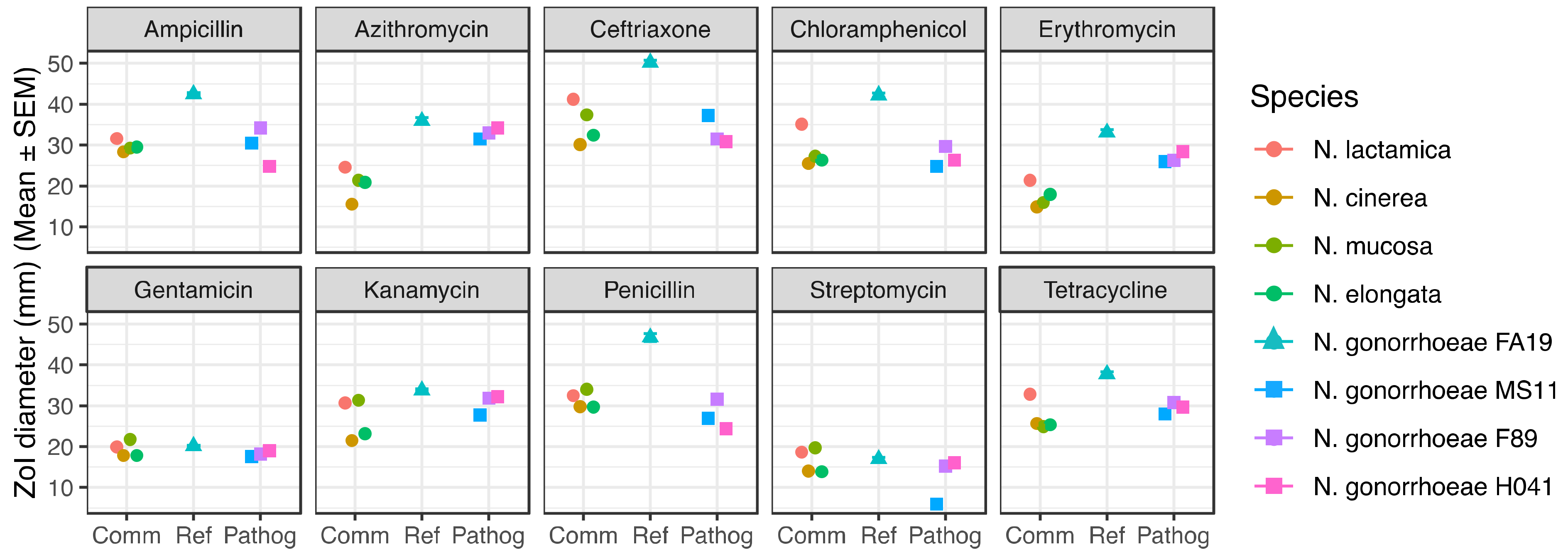
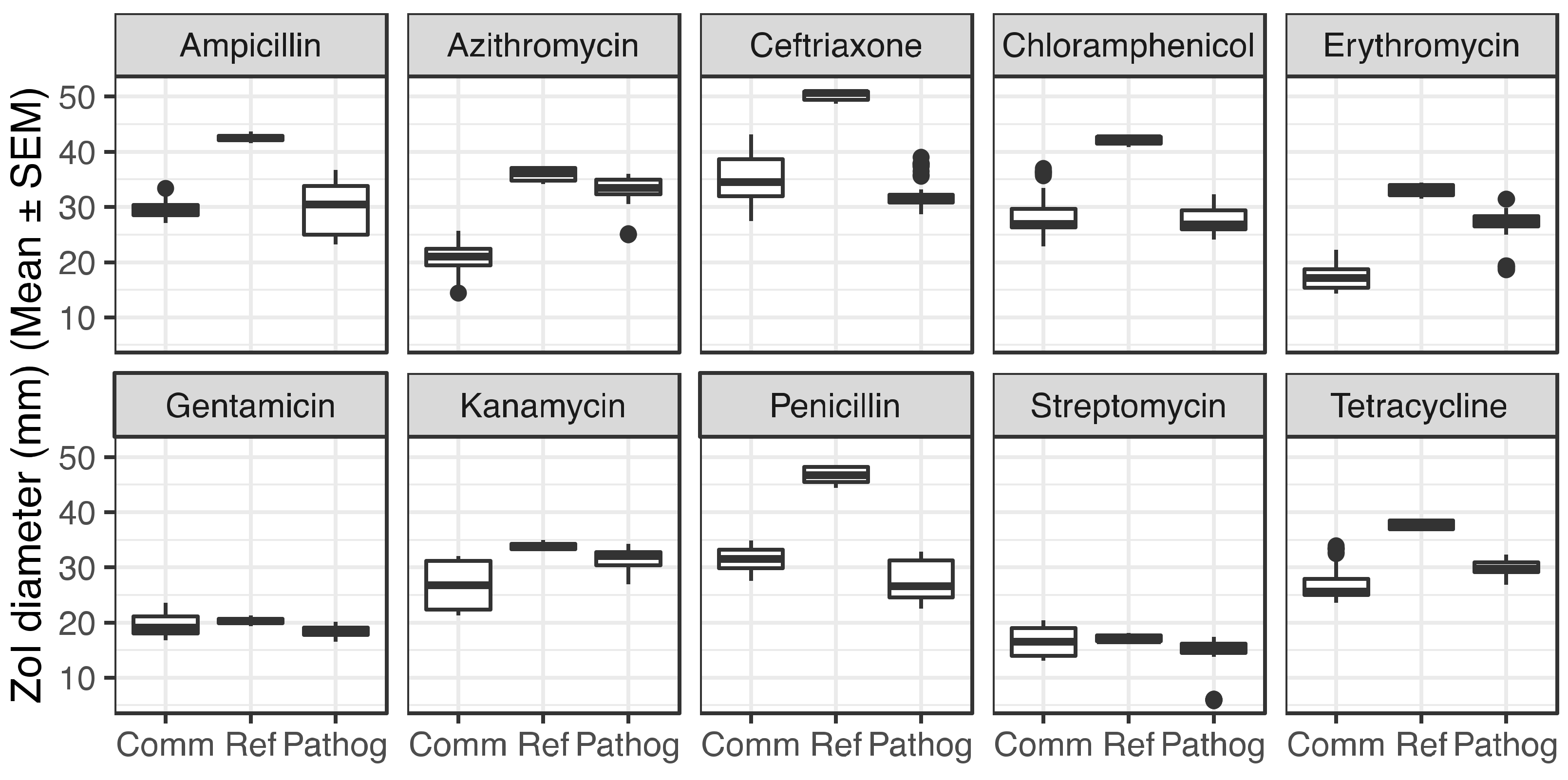
2.1. Commensal Neisseria Display Increased Resistance Levels to Several Antibiotics, Detected by Disk Diffusion Assays
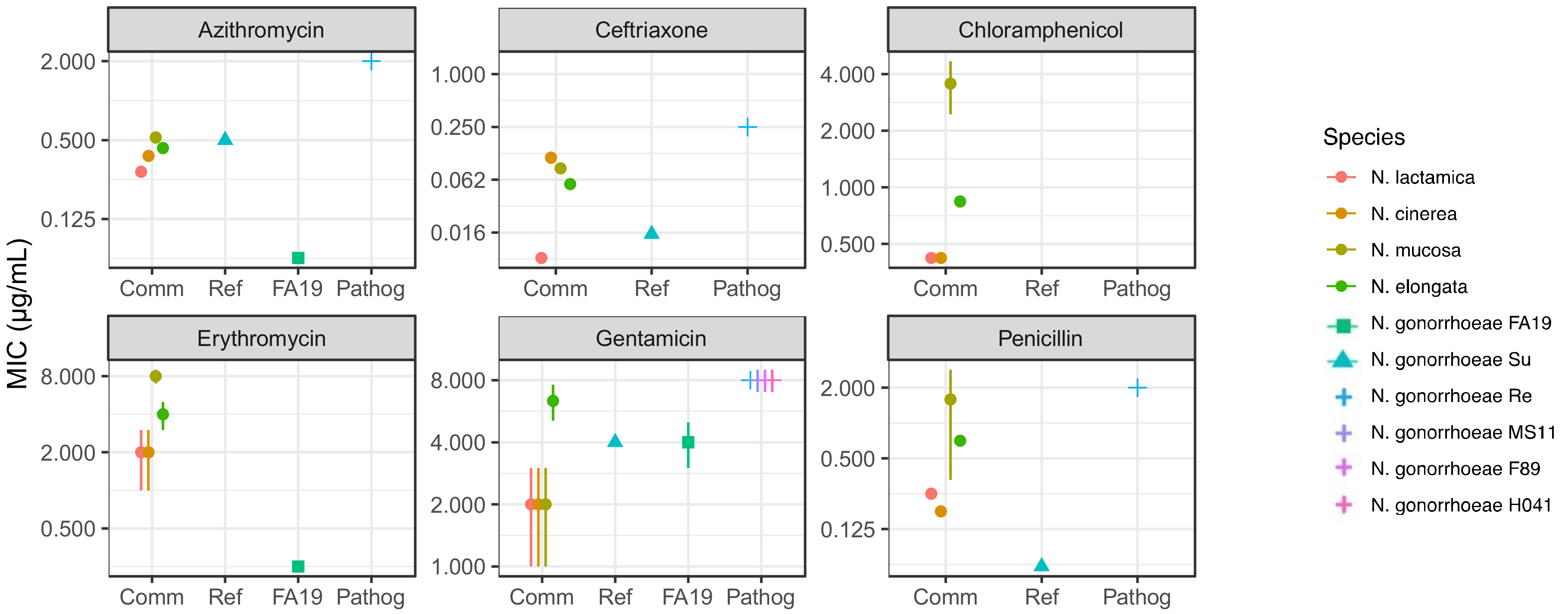
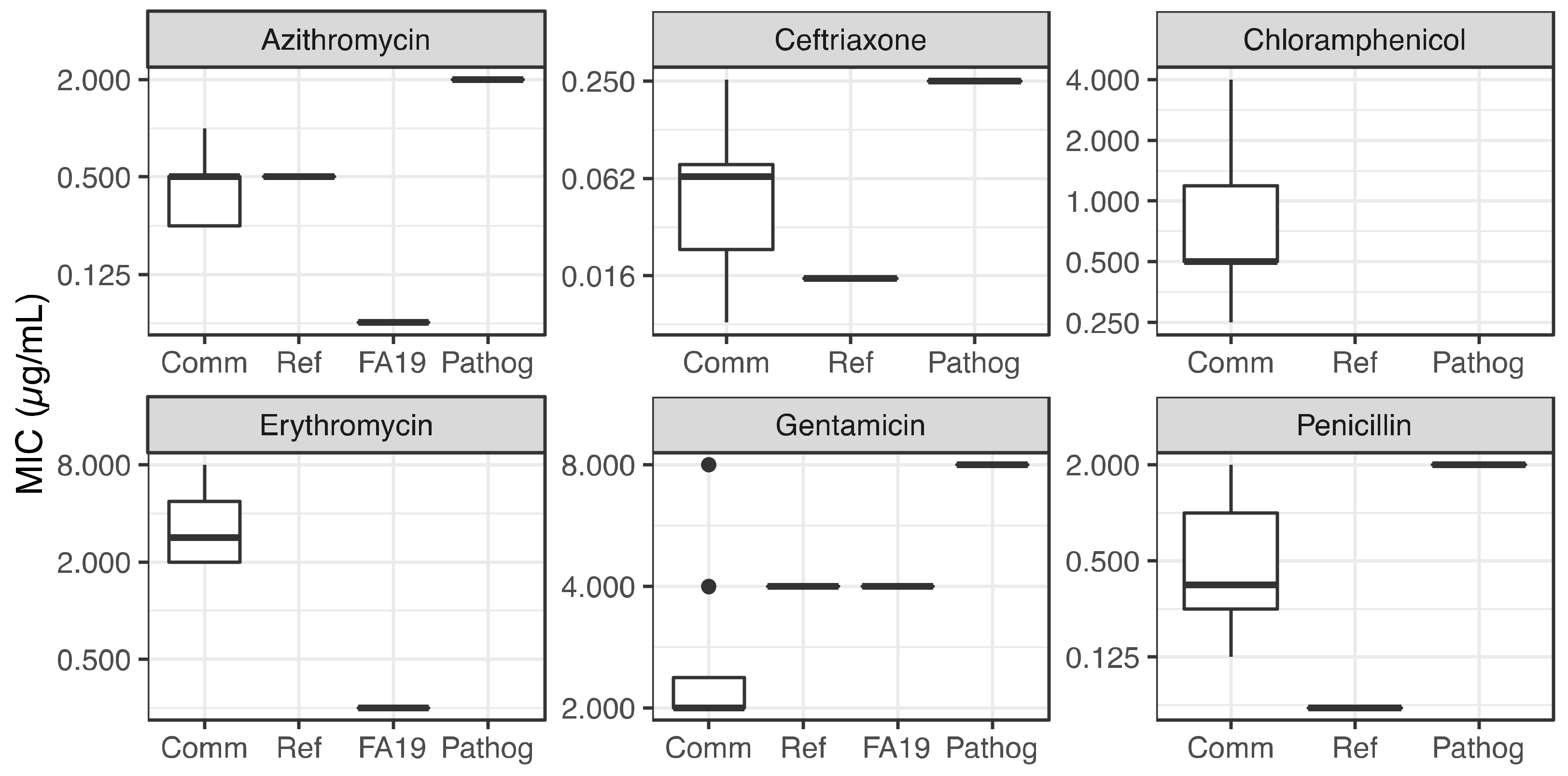
2.2. Commensal Neisseria Display Increased Resistance Levels to Several Antibiotics, Detected by Minimal Inhibitory Concentrations
2.3. Mutations in Penicillin-Binding Protein 2 (PBP2) and other Protein Sequences Can Partially Explain Resistance in Commensal Species
3. Discussion
4. Materials and Methods
4.1. Bacterial Strains
4.2. Disk Diffusion Assays
4.3. Minimal Inhibitory Concentrations
4.4. Penicillin-Binding Protein 2 (PBP2) Sequence Alignment
4.5. Data Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Centers for Disease Control and Prevention. Sexually Transmitted Disease Surveillance 2017; U.S. Department of Health and Human Services: Atlanta, GA, USA, 2018.
- Wi, T.; Lahra, M.M.; Ndowa, F.; Bala, M.; Dillon, J.R.; Ramon-Pardo, P.; Eremin, S.R.; Bolan, G.; Unemo, M. Antimicrobial resistance in Neisseria gonorrhoeae: Global surveillance and a call for international collaborative action. PLoS Med. 2017, 14, e1002344. [Google Scholar] [CrossRef]
- Centers for Diseases Control and Prevention. Sexually Transmitted Disease Surveillance 2016; U.S. Department of Health and Human Services: Atlanta, GA, USA, 2017.
- Kirkcaldy, R.D.; Kidd, S.; Weinstock, H.S.; Papp, J.R.; Bolan, G.A. Trends in antimicrobial resistance in Neisseria gonorrhoeae in the USA: The Gonococcal Isolate Surveillance Project (GISP), January 2006–June 2012. Sex Transm. Infect. 2013, 89 (Suppl. 4), 5–10. [Google Scholar] [CrossRef]
- Golparian, D.; Harris, S.R.; Sanchez-Buso, L.; Hoffmann, S.; Shafer, W.M.; Bentley, S.D.; Jensen, J.S.; Unemo, M. Genomic evolution of Neisseria gonorrhoeae since the preantibiotic era (1928–2013): Antimicrobial use/misuse selects for resistance and drives evolution. BMC Genom. 2020, 21, 116. [Google Scholar] [CrossRef]
- Unemo, M.; Del Rio, C.; Shafer, W.M. Antimicrobial Resistance Expressed by Neisseria gonorrhoeae: A Major Global Public Health Problem in the 21st Century. Microbiol. Spectr. 2016, 4. [Google Scholar] [CrossRef]
- Carnicer-Pont, D.; Smithson, A.; Fina-Homar, E.; Bastida, M.T. Gonococcus Antimicrobial Resistance Surveillance Working Group. First cases of Neisseria gonorrhoeae resistant to ceftriaxone in Catalonia, Spain, May 2011. Enferm. Infecc. Microbiol. Clin. 2012, 30, 218–219. [Google Scholar] [CrossRef] [PubMed]
- Monfort, L.; Caro, V.; Devaux, Z.; Delannoy, A.S.; Brisse, S.; Sednaoui, P. First Neisseria gonorrhoeae genotyping analysis in France: Identification of a strain cluster with reduced susceptibility to Ceftriaxone. J. Clin. Microbiol. 2009, 47, 3540–3545. [Google Scholar] [CrossRef] [PubMed]
- Tapsall, J. Multidrug-resistant Neisseria gonorrhoeae. CMAJ 2009, 180, 268–269. [Google Scholar] [CrossRef] [PubMed]
- Ohnishi, M.; Golparian, D.; Shimuta, K.; Saika, T.; Hoshina, S.; Iwasaku, K.; Nakayama, S.; Kitawaki, J.; Unemo, M. Is Neisseria gonorrhoeae initiating a future era of untreatable gonorrhea? Detailed characterization of the first strain with high-level resistance to ceftriaxone. Antimicrob. Agents Chemother. 2011, 55, 3538–3545. [Google Scholar] [CrossRef] [PubMed]
- Golparian, D.; Shafer, W.M.; Ohnishi, M.; Unemo, M. Importance of multidrug efflux pumps in the antimicrobial resistance property of clinical multidrug-resistant isolates of Neisseria gonorrhoeae. Antimicrob. Agents Chemother. 2014, 58, 3556–3559. [Google Scholar] [CrossRef]
- Unemo, M.; Golparian, D.; Hellmark, B. First three Neisseria gonorrhoeae isolates with high-level resistance to azithromycin in Sweden: A threat to currently available dual-antimicrobial regimens for treatment of gonorrhea? Antimicrob. Agents Chemother. 2014, 58, 624–625. [Google Scholar] [CrossRef] [PubMed]
- Unemo, M.; Shafer, W.M. Antimicrobial resistance in Neisseria gonorrhoeae in the 21st century: Past, evolution, and future. Clin. Microbiol. Rev. 2014, 27, 587–613. [Google Scholar] [CrossRef]
- Rotman, E.; Seifert, H.S. The genetics of Neisseria species. Annu. Rev. Genet. 2014, 48, 405–431. [Google Scholar] [CrossRef]
- Sparling, P.F. Genetic transformation of Neisseria gonorrhoeae to streptomycin resistance. J. Bacteriol. 1966, 92, 1364–1371. [Google Scholar] [CrossRef]
- Hamilton, H.L.; Dillard, J.P. Natural transformation of Neisseria gonorrhoeae: From DNA donation to homologous recombination. Mol. Microbiol. 2006, 59, 376–385. [Google Scholar] [CrossRef] [PubMed]
- Duffin, P.M.; Seifert, H.S. DNA uptake sequence-mediated enhancement of transformation in Neisseria gonorrhoeae is strain dependent. J. Bacteriol. 2010, 192, 4436–4444. [Google Scholar] [CrossRef] [PubMed]
- Mell, J.C.; Redfield, R.J. Natural competence and the evolution of DNA uptake specificity. J. Bacteriol. 2014, 196, 1471–1483. [Google Scholar] [CrossRef] [PubMed]
- Marri, P.R.; Paniscus, M.; Weyand, N.J.; Rendon, M.A.; Calton, C.M.; Hernandez, D.R.; Higashi, D.L.; Sodergren, E.; Weinstock, G.M.; Rounsley, S.D.; et al. Genome sequencing reveals widespread virulence gene exchange among human Neisseria species. PLoS ONE 2010, 5, e11835. [Google Scholar] [CrossRef]
- Bennett, J.S.; Jolley, K.A.; Earle, S.G.; Corton, C.; Bentley, S.D.; Parkhill, J.; Maiden, M.C. A genomic approach to bacterial taxonomy: An examination and proposed reclassification of species within the genus Neisseria. Microbiology 2012, 158, 1570–1580. [Google Scholar] [CrossRef] [PubMed]
- Diallo, K.; MacLennan, J.; Harrison, O.B.; Msefula, C.; Sow, S.O.; Daugla, D.M.; Johnson, E.; Trotter, C.; MacLennan, C.A.; Parkhill, J.; et al. Genomic characterization of novel Neisseria species. Sci. Rep. 2019, 9, 13742. [Google Scholar] [CrossRef]
- McKenna, W.R.; Mickelsen, P.A.; Sparling, P.F.; Dyer, D.W. Iron uptake from lactoferrin and transferrin by Neisseria gonorrhoeae. Infect. Immun. 1988, 56, 785–791. [Google Scholar] [CrossRef]
- Clinical and Laboratory Standards Institute. Performance Standards for Antimicrobial Susceptibility Testing; Twenty-Third Informational Supplement; Clinical and Laboratory Standards Institute: Annapolis Junction, MD, USA, 2013. [Google Scholar]
- Bala, M.; Singh, V.; Philipova, I.; Bhargava, A.; Chandra Joshi, N.; Unemo, M. Gentamicin in vitro activity and tentative gentamicin interpretation criteria for the CLSI and calibrated dichotomous sensitivity disc diffusion methods for Neisseria gonorrhoeae. J. Antimicrob. Chemother. 2016, 71, 1856–1859. [Google Scholar] [CrossRef]
- Kirkcaldy, R.D.; Harvey, A.; Papp, J.R.; Del Rio, C.; Soge, O.O.; Holmes, K.K.; Hook, E.W., 3rd; Kubin, G.; Riedel, S.; Zenilman, J.; et al. Neisseria gonorrhoeae Antimicrobial Susceptibility Surveillance—The Gonococcal Isolate Surveillance Project, 27 Sites, United States, 2014. MMWR Surveill. Summ. 2016, 65, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Zarantonelli, L.; Borthagaray, G.; Lee, E.H.; Veal, W.; Shafer, W.M. Decreased susceptibility to azithromycin and erythromycin mediated by a novel mtrR promoter mutation in Neisseria gonorrhoeae. J. Antimicrob. Chemother. 2001, 47, 651–654. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.; Duncan, M.; Tomberg, J.; Davies, C.; Unemo, M.; Nicholas, R.A. Genetics of chromosomally mediated intermediate resistance to ceftriaxone and cefixime in Neisseria gonorrhoeae. Antimicrob. Agents Chemother. 2009, 53, 3744–3751. [Google Scholar] [CrossRef]
- Demczuk, W.; Sidhu, S.; Unemo, M.; Whiley, D.M.; Allen, V.G.; Dillon, J.R.; Cole, M.; Seah, C.; Trembizki, E.; Trees, D.L.; et al. Neisseria gonorrhoeae Sequence Typing for Antimicrobial Resistance, a Novel Antimicrobial Resistance Multilocus Typing Scheme for Tracking Global Dissemination of N. gonorrhoeae Strains. J. Clin. Microbiol. 2017, 55, 1454–1468. [Google Scholar] [CrossRef]
- Ito, M.; Deguchi, T.; Mizutani, K.S.; Yasuda, M.; Yokoi, S.; Ito, S.; Takahashi, Y.; Ishihara, S.; Kawamura, Y.; Ezaki, T. Emergence and spread of Neisseria gonorrhoeae clinical isolates harboring mosaic-like structure of penicillin-binding protein 2 in Central Japan. Antimicrob. Agents Chemother. 2005, 49, 137–143. [Google Scholar] [CrossRef] [PubMed]
- Brannigan, J.A.; Tirodimos, I.A.; Zhang, Q.Y.; Dowson, C.G.; Spratt, B.G. Insertion of an extra amino acid is the main cause of the low affinity of penicillin-binding protein 2 in penicillin-resistant strains of Neisseria gonorrhoeae. Mol. Microbiol. 1990, 4, 913–919. [Google Scholar] [CrossRef]
- Ameyama, S.; Onodera, S.; Takahata, M.; Minami, S.; Maki, N.; Endo, K.; Goto, H.; Suzuki, H.; Oishi, Y. Mosaic-like structure of penicillin-binding protein 2 Gene (penA) in clinical isolates of Neisseria gonorrhoeae with reduced susceptibility to cefixime. Antimicrob. Agents Chemother. 2002, 46, 3744–3749. [Google Scholar] [CrossRef]
- Pandori, M.; Barry, P.M.; Wu, A.; Ren, A.; Whittington, W.L.; Liska, S.; Klausner, J.D. Mosaic penicillin-binding protein 2 in Neisseria gonorrhoeae isolates collected in 2008 in San Francisco, California. Antimicrob. Agents Chemother. 2009, 53, 4032–4034. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Lipman, D.J. Protein database searches for multiple alignments. Proc. Natl. Acad. Sci. USA 1990, 87, 5509–5513. [Google Scholar] [CrossRef]
- Sievers, F.; Wilm, A.; Dineen, D.; Gibson, T.J.; Karplus, K.; Li, W.; Lopez, R.; McWilliam, H.; Remmert, M.; Soding, J.; et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 2011, 7, 539. [Google Scholar] [CrossRef] [PubMed]
- Fedarovich, A.; Cook, E.; Tomberg, J.; Nicholas, R.A.; Davies, C. Structural effect of the Asp345a insertion in penicillin-binding protein 2 from penicillin-resistant strains of Neisseria gonorrhoeae. Biochemistry 2014, 53, 7596–7603. [Google Scholar] [CrossRef]
- Unemo, M.; Golparian, D.; Nicholas, R.; Ohnishi, M.; Gallay, A.; Sednaoui, P. High-level cefixime- and ceftriaxone-resistant Neisseria gonorrhoeae in France: Novel penA mosaic allele in a successful international clone causes treatment failure. Antimicrob. Agents Chemother. 2012, 56, 1273–1280. [Google Scholar] [CrossRef] [PubMed]
- Bharat, A.; Demczuk, W.; Martin, I.; Mulvey, M.R. Effect of Variants of Penicillin-Binding Protein 2 on Cephalosporin and Carbapenem Susceptibilities in Neisseria gonorrhoeae. Antimicrob. Agents Chemother. 2015, 59, 5003–5006. [Google Scholar] [CrossRef] [PubMed]
- Takahata, S.; Senju, N.; Osaki, Y.; Yoshida, T.; Ida, T. Amino acid substitutions in mosaic penicillin-binding protein 2 associated with reduced susceptibility to cefixime in clinical isolates of Neisseria gonorrhoeae. Antimicrob. Agents Chemother. 2006, 50, 3638–3645. [Google Scholar] [CrossRef]
- Fiore, M.A.; Raisman, J.C.; Wong, N.H.; Hudson, A.O.; Wadsworth, C.B. Exploration of the Neisseria Resistome Reveals Resistance Mechanisms in Commensals That May Be Acquired by N. gonorrhoeae through Horizontal Gene Transfer. Antibiotics 2020, 9, 656. [Google Scholar] [CrossRef]
- Liu, G.; Tang, C.M.; Exley, R.M. Non-pathogenic Neisseria: Members of an abundant, multi-habitat, diverse genus. Microbiology 2015, 161, 1297–1312. [Google Scholar] [CrossRef]
- Workowski, K.A.; Bolan, G. Sexually Transmitted Diseases Treatment Guidelines, 2015. MMWR Recomm. Rep. 2015, 64, 60–68. [Google Scholar] [PubMed]
- Barbee, L.A.; Soge, O.O.; Morgan, J.; Leclair, A.; Bass, T.; Werth, B.J.; Hughes, J.P.; Golden, M.R. Gentamicin Alone Is Inadequate to Eradicate Neisseria gonorrhoeae From the Pharynx. Clin. Infect. Dis. 2020, 71, 1877–1882. [Google Scholar] [CrossRef]
- McInnes, R.S.; McCallum, G.E.; Lamberte, L.E.; van Schaik, W. Horizontal transfer of antibiotic resistance genes in the human gut microbiome. Curr. Opin. Microbiol. 2020, 53, 35–43. [Google Scholar] [CrossRef]
- Von Wintersdorff, C.J.; Penders, J.; van Niekerk, J.M.; Mills, N.D.; Majumder, S.; van Alphen, L.B.; Savelkoul, P.H.; Wolffs, P.F. Dissemination of Antimicrobial Resistance in Microbial Ecosystems through Horizontal Gene Transfer. Front. Microbiol. 2016, 7, 173. [Google Scholar] [CrossRef]
- Chen, M.; Zhang, C.; Zhang, X.; Chen, M. Meningococcal Quinolone Resistance Originated from Several Commensal Neisseria Species. Antimicrob. Agents Chemother. 2020, 64. [Google Scholar] [CrossRef] [PubMed]
- Furuya, R.; Onoye, Y.; Kanayama, A.; Saika, T.; Iyoda, T.; Tatewaki, M.; Matsuzaki, K.; Kobayashi, I.; Tanaka, M. Antimicrobial resistance in clinical isolates of Neisseria subflava from the oral cavities of a Japanese population. J. Infect. Chemother. 2007, 13, 302–304. [Google Scholar] [CrossRef]
- Galimand, M.; Gerbaud, G.; Courvalin, P. Spectinomycin resistance in Neisseria spp. due to mutations in 16S rRNA. Antimicrob. Agents Chemother. 2000, 44, 1365–1366. [Google Scholar] [CrossRef]
- Rouquette-Loughlin, C.E.; Reimche, J.L.; Balthazar, J.T.; Dhulipala, V.; Gernert, K.M.; Kersh, E.N.; Pham, C.D.; Pettus, K.; Abrams, A.J.; Trees, D.L.; et al. Mechanistic Basis for Decreased Antimicrobial Susceptibility in a Clinical Isolate of Neisseria gonorrhoeae Possessing a Mosaic-Like mtr Efflux Pump Locus. mBio 2018, 9. [Google Scholar] [CrossRef]
- Apicella, M.A. Commensal Bacteria: Not Just Innocent Bystanders. mBio 2019, 10. [Google Scholar] [CrossRef] [PubMed]
- Aho, E.L.; Murphy, G.L.; Cannon, J.G. Distribution of specific DNA sequences among pathogenic and commensal Neisseria species. Infect. Immun. 1987, 55, 1009–1013. [Google Scholar] [CrossRef] [PubMed]
- Swanson, J.; Barrera, O.; Sola, J.; Boslego, J. Expression of outer membrane protein II by gonococci in experimental gonorrhea. J. Exp. Med. 1988, 168, 2121–2129. [Google Scholar] [CrossRef]
- Clinical and Laboratory Standards Institute. Performance Standards for Antimicrobial Disk Susceptibility Tests, 12th ed.; Approved Standard; Clinical and Laboratory Standards Institute: Annapolis Junction, MD, USA, 2018. [Google Scholar]
- Alonso, C.A.; Domínguez, C.; Heras, J.; Mata, E.; Pascual, V.; Torres, C.; Zarazaga, M. Antibiogramj: A tool for analysing images from disk diffusion tests. Comput. Methods Programs Biomed. 2017, 143, 159–169. [Google Scholar] [CrossRef]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 2012, 9, 671–675. [Google Scholar] [CrossRef] [PubMed]
- Clinical and Laboratory Standards Institute. Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria that Grow Aerobically, 10th ed.; Approved Standard; Clinical and Laboratory Standards Institute: Annapolis Junction, MD, USA, 2018. [Google Scholar]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2017. [Google Scholar]
- Wickham, H. ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016. [Google Scholar]

| Antibiotic | Df | Sum of Sq | Mean Sq | F-Value | Pr (>F) | |
|---|---|---|---|---|---|---|
| Ampicillin | group | 2 | 1362 | 680.8 | 52.66 | <1 × 10−4 |
| Residuals | 105 | 1358 | 12.9 | S | ||
| Azithromycin | group | 2 | 4040 | 2020.2 | 273.6 | <1 × 10−4 |
| Residuals | 102 | 753 | 7.4 | S | ||
| Ceftriaxone | group | 2 | 2614 | 1306.9 | 126.2 | <1 × 10−4 |
| Residuals | 105 | 1087 | 10.4 | S | ||
| Chloramphenicol | group | 2 | 1708.4 | 854.2 | 98.43 | <1 × 10−4 |
| Residuals | 105 | 911.2 | 8.7 | S | ||
| Erythromycin | group | 2 | 2829 | 1414.5 | 277.6 | <1 × 10−4 |
| Residuals | 105 | 535 | 5.1 | S | ||
| Gentamicin | group | 2 | 159.2 | 79.59 | 44.21 | <1 × 10−4 |
| Residuals | 357 | 642.8 | 1.8 | S | ||
| Kanamycin | group | 2 | 653.1 | 326.5 | 36.57 | <1 × 10−4 |
| Residuals | 105 | 937.5 | 8.9 | S | ||
| Penicillin | group | 2 | 2851.8 | 1425.9 | 162.2 | <1 × 10−4 |
| Residuals | 105 | 922.9 | 8.8 | S | ||
| Streptomycin | group | 2 | 147.7 | 73.83 | 7.482 | 9.17 × 10−4 |
| Residuals | 105 | 1036.1 | 9.87 | NS | ||
| Tetracycline | group | 2 | 818.1 | 409 | 83.72 | <1 × 10−4 |
| Residuals | 105 | 513 | 4.9 | S |
| Antibiotic | Ampicillin | Azithromycin | Ceftriaxone | Chloramphenicol | Erythromycin | |||||
| diff | p adj | diff | p adj | diff | p adj | diff | p adj | diff | p adj | |
| Reference to commensal | 12.807 | <1 × 10−4 | 15.295 | <1 × 10−4 | 14.866 | <1 × 10−4 | 13.645 | <1 × 10−4 | 15.51 | <1 × 10−4 |
| Pathogenic to commensal | −0.062 | 0.9962 | 12.631 | <1 × 10−4 | −3.268 | <1 × 10−4 | −1.014 | 0.23 | 9.566 | <1 × 10−4 |
| Pathogenic to reference | −12.869 | <1 × 10−4 | −2.665 | 0.0195 | −18.134 | <1 × 10−4 | −14.659 | <1 × 10−4 | −5.943 | <1 × 10−4 |
| Antibiotic | Gentamicin | Kanamycin | Penicillin | Streptomycin | Tetracycline | |||||
| diff | p adj | diff | p adj | diff | p adj | diff | p adj | diff | p adj | |
| Reference to commensal | 0.733 | 0.0033 | 7.108 | <1 × 10−4 | 15.206 | <1 × 10−4 | 0.389 | 0.941 | 10.592 | <1 × 10−4 |
| Pathogenic to commensal | −1.141 | <1 × 10−4 | 4.741 | <1 × 10−4 | −3.683 | <1 × 10−4 | −2.285 | 0.0021 | 2.743 | <1 × 10−4 |
| Pathogenic to reference | −1.874 | <1 × 10−4 | −2.367 | 0.0719 | −18.889 | <1 × 10−4 | −2.674 | 0.0486 | −7.85 | <1 × 10−4 |
| Antibiotic | Df | Sum of Sq | Mean Sq | F-Value | Pr (>F) | Sign lev * | |
|---|---|---|---|---|---|---|---|
| Azithromycin | group | 3 | 12.61 | 4.202 | 7.912 | 1.61 × 10−4 | NS |
| Residuals | 59 | 31.33 | 0.531 | ||||
| Ceftriaxone | group | 2 | 8.61 | 4.303 | 1.727 | 0.187 | NS |
| Residuals | 59 | 147 | 2.492 | ||||
| Erythromycin | group | 1 | 13.68 | 13.682 | 19.35 | 9.75 × 10−5 | S |
| Residuals | 35 | 24.75 | 0.707 | ||||
| Gentamicin | group | 3 | 39.59 | 13.196 | 44.52 | <1 × 10−4 | S |
| Residuals | 70 | 20.75 | 0.296 | ||||
| Penicillin | group | 2 | 13.16 | 6.582 | 3.619 | 0.043 | NS |
| Residuals | 23 | 41.83 | 1.819 |
| Antibiotic | Reference-Commensal | FA19—Commensal | ||
|---|---|---|---|---|
| diff | p adj | diff | p adj | |
| Penicillin | 2.083 | 0.303 | ||
| Azithromycin | 0.333 | 0.969 | −2.655 | 3.41 × 10−3 |
| Ceftriaxone | −1.593 | 0.579 | ||
| Erythromycin | NA | NA | −3.75 | 9.71 × 10−5 |
| Gentamicin | 0.583 | 0.717 | ||
| Chloramphenicol | NA | NA | NA | NA |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Goytia, M.; Thompson, S.T.; Jordan, S.V.L.; King, K.A. Antimicrobial Resistance Profiles of Human Commensal Neisseria Species. Antibiotics 2021, 10, 538. https://doi.org/10.3390/antibiotics10050538
Goytia M, Thompson ST, Jordan SVL, King KA. Antimicrobial Resistance Profiles of Human Commensal Neisseria Species. Antibiotics. 2021; 10(5):538. https://doi.org/10.3390/antibiotics10050538
Chicago/Turabian StyleGoytia, Maira, Symone T. Thompson, Skylar V. L. Jordan, and Kacey A. King. 2021. "Antimicrobial Resistance Profiles of Human Commensal Neisseria Species" Antibiotics 10, no. 5: 538. https://doi.org/10.3390/antibiotics10050538
APA StyleGoytia, M., Thompson, S. T., Jordan, S. V. L., & King, K. A. (2021). Antimicrobial Resistance Profiles of Human Commensal Neisseria Species. Antibiotics, 10(5), 538. https://doi.org/10.3390/antibiotics10050538






