Taxonomic Identification of Different Species of the Genus Aeromonas by Whole-Genome Sequencing and Use of Their Species-Specific β-Lactamases as Phylogenetic Markers
Abstract
1. Introduction
2. Results
2.1. Identification by MALDI-TOF
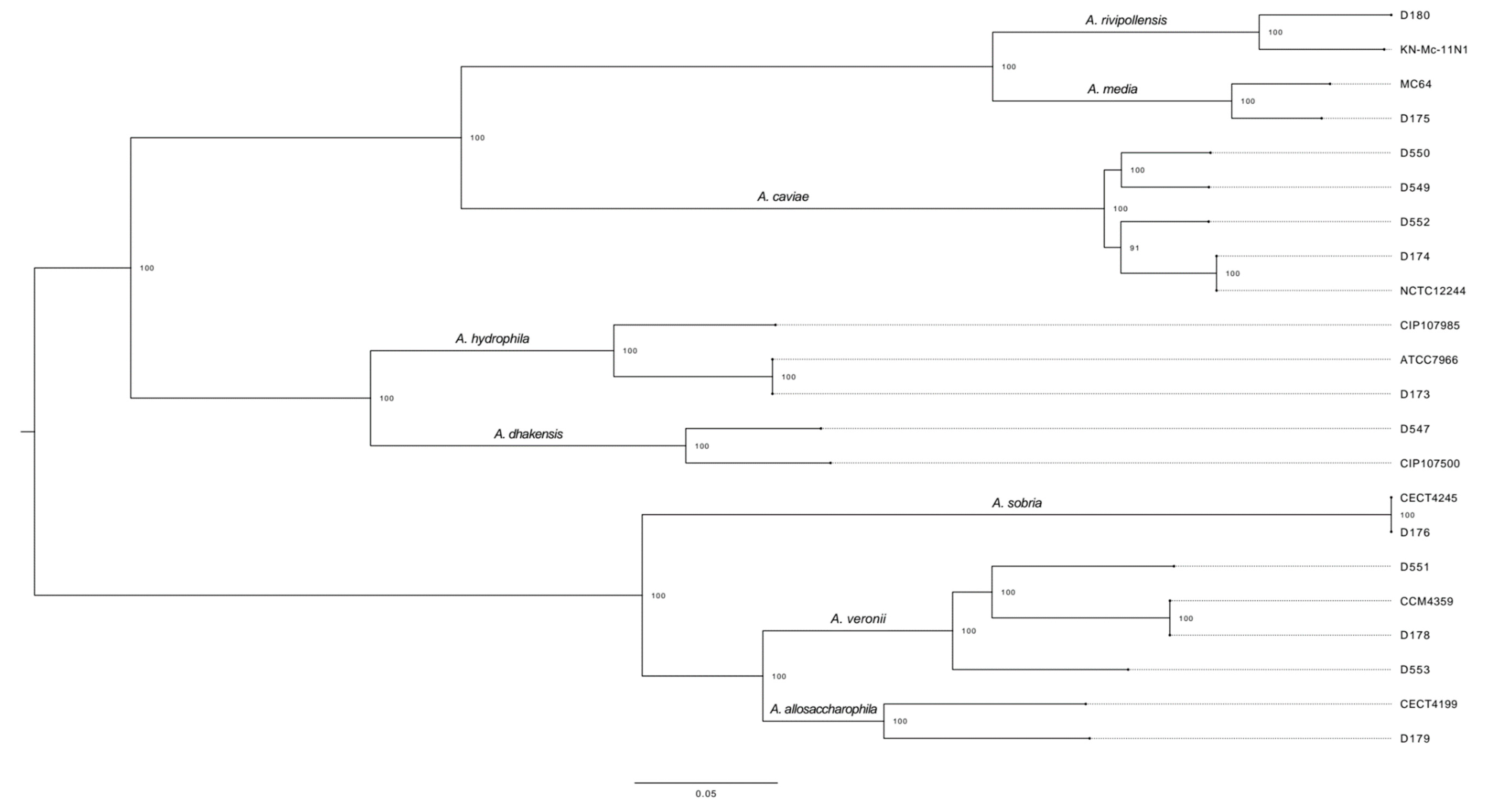
2.2. WGS Analysis: Core-Genome Alignment
2.3. Antimicrobial Susceptibility Testing
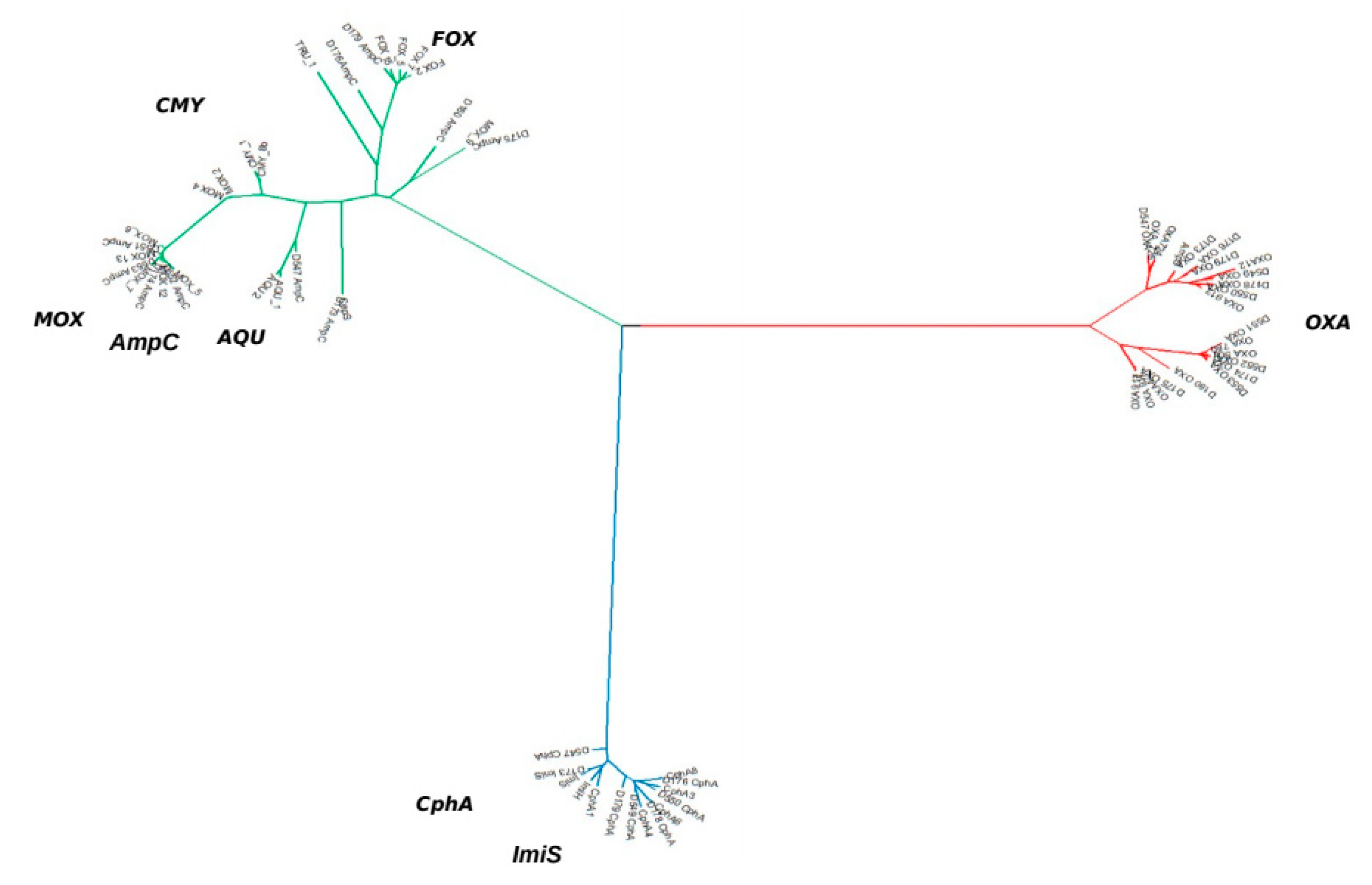
2.4. β-Lactamases
2.5. Characterisation of Other Mechanisms of Resistance to Antimicrobials
3. Discussion
4. Materials and Methods
4.1. Identification and Antimicrobial Susceptibility Testing
4.2. DNA Extraction and Whole-Genome Sequencing
4.3. Computational Analysis
4.3.1. Whole-Genome Comparison
4.3.2. Detection of Antimicrobial Resistance Genes
4.3.3. Antimicrobial Resistance Genes Phylogenetic Tree
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Sample Availability
References
- Martin-Carnahan, A.; Joseph, S.W. Order XII. Aeromonadales ord. nov. In Bergey’s Manual of Systematic Bacteriology, 2nd ed.; Brenner, D.J., Krieg, N.R., Staley, J.T., Garrity, G.M., Eds.; Williams & Wilkins: Philadelphia, PA, USA, 2005; pp. 556–578. [Google Scholar]
- Fernández-Bravo, A.; Figueras, M.J. An update on the genus Aeromonas: Taxonomy, epidemiology, and pathogenicity. Microorganisms 2020, 8, 129. [Google Scholar] [CrossRef]
- Parker, J.L.; Shaw, J.G. Aeromonas spp. clinical microbiology and disease. J. Infect. 2011, 62, 109–118. [Google Scholar] [CrossRef] [PubMed]
- Janda, J.M.; Abbott, S.L. The genus Aeromonas: Taxonomy, pathogenicity, and infection. Clin. Microbiol. 2010, 23, 35–73. [Google Scholar] [CrossRef]
- Gonçalves Pessoa, R.B.; de Oliveira, W.F.; Marques, D.S.C.; dos Santos Correia, M.T.; de Carvalho, E.V.M.M.; Coelho, L.C.B.B. The genus Aeromonas: A general approach. Microb. Pathog. 2019, 130, 81–94. [Google Scholar] [CrossRef]
- Abbott, S.L.; Cheung, W.K.W.; Janda, J.M. The genus Aeromonas: Biochemical characteristics, atypical reactions, and phenotypic identification schemes. J. Clin. Microbiol. 2003, 4, 2348–2357. [Google Scholar] [CrossRef] [PubMed]
- Navarro, A.; Martínez-Murcia, A. Phylogenetic analyses of the genus Aeromonas based on housekeeping gene sequencing and its influence on systematics. J. Appl. Microbiol. 2018, 125, 622–631. [Google Scholar] [CrossRef]
- Küpfer, M.; Kuhnert, P.; Korczak, B.M.; Peduzzi, R.; Demarta, A. Genetic relationships of Aeromonas strains inferred from 16S rRNA, gyrB and rpoB gene sequences. Int. J. Syst. Evol. Microbiol. 2006, 56, 2743–2751. [Google Scholar] [CrossRef]
- Yáñez, M.A.; Catalán, V.; Apráiz, D.; Figueras, M.J.; Martínez-Murcia, A.J. Phylogenetic analysis of members of the genus Aeromonas based on gyrB gene sequences. Int. J. Syst. Evol. Microbiol. 2003, 53, 875–883. [Google Scholar] [CrossRef]
- Zhou, Y.; Yu, L.; Nan, Z.; Zhang, P.; Kan, B.; Yan, D.; Su, J. Taxonomy, virulence genes and antimicrobial resistance of Aeromonas isolated from extra-intestinal and intestinal infections. BMC Infect. Dis. 2019, 9, 158. [Google Scholar] [CrossRef]
- De Melo, B.S.T.; Mendes-Marques, C.L.; de Lima Campos, T.; de Almeida, A.M.P.; Leal, N.C.; Xavier, D.E. High-resolution genome-wide analysis is essential for the identification of ambiguous Aeromonas strains. FEMS Microbiol. Lett. 2019, 366, fnz245. [Google Scholar] [CrossRef] [PubMed]
- Schuetz, A.N. Emerging agents of gastroenteritis: Aeromonas, plesiomonas, and the diarrheagenic pathotypes of Escherichia coli. Semin. Diagn. Pathol. 2019, 36, 187–192. [Google Scholar] [CrossRef]
- Shin, H.B.; Yoon, J.; Lee, Y.; Kim, M.S.; Lee, K. Comparison of MALDI-TOF MS, Housekeeping gene sequencing, and 16s rRNA gene sequencing for identification of Aeromonas clinical isolates. Yonsei Med. J. 2015, 56, 550–555. [Google Scholar] [CrossRef] [PubMed]
- Ruiz de Alegría-Puig, C.; Aguirre-Quiñonero, A.; Agüero-Balbín, J.; Roiz-Mesones, M.P.; Martínez-Martínez, L. Correlation between MALDI-TOF Vitek-MSTM system and conventional identification methods of gastrointestinal infection causing bacteria. Rev. Esp. Quimioter. 2016, 29, 265–268. [Google Scholar] [PubMed]
- Vávrová, A.; Balážová, T.; Sedláček, I.; Tvrzová, L.; Šedo, O. Evaluation of the MALDI-TOF MS profiling for identification of newly described Aeromonas spp. Folia Microbiol. 2015, 60, 375–383. [Google Scholar] [CrossRef]
- Sinclair, H.A.; Heney, C.; Sidjabat, H.E.; George, N.M.; Bergh, H.; Anuj, S.N.; Nimmo, G.R.; Paterson, D.L. Genotypic and phenotypic identification of Aeromonas species and CphA-mediated carbapenem resistance in Queensland, Australia. Diagn. Microbiol. Infect. Dis. 2016, 85, 98–101. [Google Scholar] [CrossRef]
- Benagli, C.; Demarta, A.; Caminada, A.P.; Ziegler, D.; Petrini, O.; Tonolla, M. A Rapid MALDI-TOF MS Identification Database at Genospecies Level for Clinical and Environmental Aeromonas Strains. PLoS ONE 2012, 7, e48441. [Google Scholar] [CrossRef]
- Chen, P.L.; Lamy, B.; Ko, W.C. Aeromonas dhakensis, an increasingly recognized human pathogen. Front. Microbiol. 2016, 7, 793. [Google Scholar] [CrossRef]
- Wu, C.J.; Chen, P.L.; Hsueh, P.R.; Chang, M.C.; Tsai, P.J.; Shih, H.I.; Wang, H.C.; Chou, P.H.; Ko, W.C. Clinical implications of species identification in monomicrobial Aeromonas bacteremia. PLoS ONE 2015, 10, e0117821. [Google Scholar] [CrossRef]
- Kitagawa, H.; Ohge, H.; Yu, L.; Kayama, S.; Hara, T.; Kashiyama, S.; Kajihara, T.; Hisatsune, J.; Sueda, T.; Sugai, M. Aeromonas dhakensis is not a rare cause of Aeromonas bacteremia in Hiroshima, Japan. J. Infect. Chemother. 2020, 26, 316–320. [Google Scholar] [CrossRef] [PubMed]
- Borriss, R.; Rueckert, C.; Blom, J.; Bezuidt, O.; Reva, O.; Klenk, H.P. Whole Genome Sequence Comparisons in Taxonomy. Methods Microbiol. 2011, 38, 409–436. [Google Scholar]
- Fosse, T.; Giraud-Morin, C.; Madinier, I. Phénotypes de résistance aux β-lactamines dans le genre Aeromonas. Pathol. Biol. 2003, 51, 290–296. [Google Scholar] [CrossRef]
- Chen, P.L.; Ko, W.C.; Wu, C.J. Complexity of β-lactamases among clinical Aeromonas isolates and its clinical implications. J. Microbio. Immunol. Infect. 2012, 45, 398–403. [Google Scholar] [CrossRef] [PubMed]
- Walsh, T.R.; Hall, L.; Macgowan, A.P.; Bennett, P.M. Sequence analysis of two chromosomally mediated inducible β-lactamases from Aeromonas sobria, strain 163a, one class d penicillinase, the other an AmpC cephalosporinase. J. Antimicrob. Chemother. 1995, 36, 41–52. [Google Scholar] [CrossRef]
- Juan, C.; Torrens, G.; González-Nicolau, M.; Oliver, A. Diversity and regulation of intrinsic β-lactamases from non-fermenting and other Gram-negative opportunistic pathogens. FEMS Microbiol. Rev. 2017, 41, 780–814. [Google Scholar] [CrossRef]
- Hughes, H.Y.; Conlan, S.P.; Lau, A.F.; Dekker, J.P.; Michelin, A.V.; Youn, J.H.; Henderson, D.K.; Frank, K.M.; Segre, J.A.; Palmore, T.N. Detection and whole-genome sequencing of carbapenemase- producing Aeromonas hydrophila isolates from routine perirectal surveillance culture. J. Clin. Microbiol. 2016, 54, 1167–1170. [Google Scholar] [CrossRef]
- Ghatak, S.; Blom, J.; Das, S.; Sanjukta, R.; Puro, K.; Mawlong, M.; Shakuntala, I.; Sen, A.; Goesmann, A.; Kumar, A.; et al. Pan-genome analysis of Aeromonas hydrophila, Aeromonas veronii and Aeromonas caviae indicates phylogenomic diversity and greater pathogenic potential for Aeromonas hydrophila. Antonie van Leeuwenhoek Int. J. Gen. Mol. Microbiol. 2016, 109, 945–956. [Google Scholar] [CrossRef] [PubMed]
- Marti, E.; Balcazar, J.L. Aeromonas rivipollensis sp. Nov.; A novel species isolated from aquatic samples. J. Basic Microbiol. 2015, 55, 1435–1439. [Google Scholar] [CrossRef]
- Ebmeyer, S.; Kristiansson, E.; Joakim Larsson, D.G. The mobile FOX AmpC beta-lactamases originated in Aeromonas allosaccharophila. Int. J. Antimicrob. Agents 2019, 54, 798–802. [Google Scholar] [CrossRef]
- Ebmeyer, S.; Kristiansson, E.; Joakim Larsson, D.G. CMY-1/MOX-family AmpC β-lactamases MOX-1, MOX-2 and MOX-9 were mobilized independently from three Aeromonas species. J. Antimicrob. Chemother. 2019, 74, 1202–1206. [Google Scholar] [CrossRef]
- Niumsup, P.; Simm, A.M.; Nurmahomed, K.; Walsh, T.R.; Bennett, P.M.; Avison, M.B. Genetic linkage of the penicillinase gene, amp, and blrAB, encoding the regulator of β-lactamase expression in Aeromonas spp. J. Antimicrob. Chemother. 2003, 51, 1351–1358. [Google Scholar] [CrossRef]
- Walsh, T.R.; Stunt, R.A.; Nabi, J.A.; MacGowan, A.P.; Bennett, P.M. Distribution and expression of beta-lactamase genes among Aeromonas spp. J. Antimicrob. Chemother. 1997, 40, 171–178. [Google Scholar] [CrossRef]
- Perez-Perez, F.J.; Hanson, N.D. Detection of plasmid-mediated AmpC β-lactamase genes in clinical isolates by using multiplex PCR. J. Clin. Microbiol. 2002, 40, 2153–2162. [Google Scholar] [CrossRef] [PubMed]
- Jacoby, G.A. AmpC β-lactamases. Antimicrob Agents Chemother. 2009, 22, 161–182. [Google Scholar] [CrossRef] [PubMed]
- De Luca, F.; Giraud-Morin, C.; Rossolini, G.M.; Docquier, J.D.; Fosse, T. Genetic and biochemical characterization of TRU-1, the endogenous class C β-lactamase from Aeromonas enteropelogenes. Antimicrob. Agents Chemother. 2010, 54, 1547–1554. [Google Scholar] [CrossRef]
- Lamy, B.; Laurent, F.; Kodjo, A. Validation of a partial rpoB gene sequence as a tool for phylogenetic identification of aeromonads isolated from environmental sources. Can. J. Microbiol. 2010, 56, 217–228. [Google Scholar] [CrossRef] [PubMed]
- Mahfouz, N.; Ferreira, I.; Beisken, S.; von Haeseler, A.; Posch, A.E. Large-scale assessment of antimicrobial resistance marker databases for genetic phenotype prediction: A systematic review. J. Antimicrob. Chemother. 2020, 75, 3099–3108. [Google Scholar] [CrossRef]
- Moura, Q.; Fernandes, M.R.; Cerdeira, L.; Santos, A.C.M.; de Souza, T.A.; Ienne, S.; Pignatari, A.C.C.; Gales, A.C.; Silva, R.M.; Lincopan, N. Draft genome sequence of a multidrug-resistant Aeromonas hydrophila ST508 strain carrying rmtD and bla CTX-M-131 isolated from a bloodstream infection. J. Glob. Antimicrob. Resist. 2017, 10, 289–290. [Google Scholar] [CrossRef] [PubMed]
- Lukkana, M.; Wongtavatchai, J.; Chuanchuen, R. Class 1 integrons in Aeromonas hydrophila isolates from farmed nile tilapia (Oreochromis nilotica). J. Vet. Med. Sci. 2012, 74, 435–440. [Google Scholar] [CrossRef]
- Adesoji, A.T.; Ogunjobi, A.A.; Olatoye, I.O. Genotypic Characterization of Aminoglycoside Resistance Genes from Bacteria Isolates in Selected Municipal Drinking Water Distribution Sources in Southwestern Nigeria. Ethiop. J. Health Sci. 2019, 29, 321. [Google Scholar]
- Machuca, J.; Agüero, J.; Miró, E.; del Carmen Conejo, M.; Oteo, J.; Bou, G.; González-López, J.J.; Oliver, A.; Navarro, F.; Pascual, Á.; et al. Prevalence of quinolone resistance mechanisms in Enterobacteriaceae producing acquired AmpC β-lactamases and/or carbapenemases in Spain. Enferm. Infecc. Microbiol. Clin. 2017, 38, 485–490. [Google Scholar] [CrossRef]
- Kadlec, K.; von Czapiewski, E.; Kaspar, H.; Wallmann, J.; Michael, G.B.; Steinacker, U.; Schwarz, S. Molecular basis of sulfonamide and trimethoprim resistance in fish-pathogenic Aeromonas isolates. Appl. Environ. Microbiol. 2011, 77, 7147–7150. [Google Scholar] [CrossRef]
- EUCAST. Breakpoint Tables for Interpretation of MICs and Zone Diameters, Version 10.0. 2020. Available online: http://www.eucast.org (accessed on 18 March 2021).
- Clinical and Laboratory Standards Institute (CLSI). Performance Standards for Antimicrobial Susceptibility Testing, 29th ed.; CLSI supplement M100; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2019. [Google Scholar]
- EUCAST Advice on Intrinsic Resistance and Exceptional Phenotypes v 3.2 (February, 2020, typographical errors corrected May, 2020). Available online: https://www.eucast.org/fileadmin/src/media/PDFs/EUCAST_files/Expert_Rules/2020/Intrinsic_Resistance_and_Unusual_Phenotypes_Tables_v3.2_20200225.pdf (accessed on 18 March 2021).
- Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 2014, 30, 2068–2069. [Google Scholar] [CrossRef]
- Page, A.J.; Cummins, C.A.; Hunt, M.; Wong, V.K.; Reuter, S.; Holden, M.T.G.; Fookes, M.; Falush, D.; Keane, J.A.; Parkhill, J. Roary: Rapid large-scale prokaryote pan genome analysis. Bioinformatics 2015, 31, 3691–3693. [Google Scholar] [CrossRef]
- Croucher, N.J.; Page, A.J.; Connor, T.R.; Delaney, A.J.; Keane, J.A.; Bentley, S.D.; Parkhill, J.; Harris, S.R. Rapid phylogenetic analysis of large samples of recombinant bacterial whole genome sequences using Gubbins. Nucleic Acids Res. 2014. [Google Scholar] [CrossRef] [PubMed]
- Yoon, S.H.; Ha, S.M.; Lim, J.M.; Kwon, S.J.; Chun, J. A large-scale evaluation of algorithms to calculate average nucleotide identity. Antonie van Leeuwenhoek. 2017, 110, 1281–1286. [Google Scholar] [CrossRef] [PubMed]
- Alcock, B.P.; Raphenya, A.R.; Lau, T.T.Y.; Tsang, K.K.; Bouchard, M.; Edalatmand, A.; Huynh, W.; Nguyen, A.L.V.; Cheng, A.A.; Liu, S.; et al. CARD 2020: Antibiotic resistome surveillance with the comprehensive antibiotic resistance database. Nucleic Acids Res. 2020, 48, 517–525. [Google Scholar] [CrossRef]
- Bortolaia, V.; Kaas, R.F.; Ruppe, E.; Roberts, M.C.; Schwarz, S.; Cattoir, V.; Philippon, A.; Allesoe, R.L.; Rebelo, A.R.; Florensa, A.R.; et al. ResFinder 4.0 for predictions of phenotypes from genotypes. J. Antimicrob. Chemother. 2020, 75, 3491–3500. [Google Scholar] [CrossRef] [PubMed]
- Clark, K.; Karsch-Mizrachi, I.; Lipman, D.J.; Ostell, J.; Sayers, E.W. GenBank. Nucleic Acids Res. 2016, 4, D67–D72. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
| LAB ID | Routine Laboratory Identification | MALDI-TOF Identification | Score |
|---|---|---|---|
| D180 ** | A. media | A. media DSM 4881T HAM | 2.22 |
| D175 | A. media ATCC 33907 (CECT4232) | A. media DSM 4881T HAM | 2.38 |
| D174 | A. caviae ATCC 15468 (CECT838) | A. caviae CECT838T DSM | 2.40 |
| D549 | A. caviae | A. caviae CECT838T DSM | 2.43 |
| D550 | A. caviae | A. caviae CECT838T DSM | 2.41 |
| D552 | A. caviae | A. caviae CECT838T DSM | 2.31 |
| D547 * | A. caviae | A. hydrophila CECT 839T DSM | 2.34 |
| D173 | A. hydrophila ATCC 7966 (CECT839) | A. hydrophila CECT 839T DSM | 2.47 |
| D176 | A. sobria ATCC 43979 (CECT4245) | A. sobria CECT 4245T DSM | 2.45 |
| D178 | A. veronii ATCC 35624 (CECT4257) | A. veronii CECT5761T DSM | 2.26 |
| D551 | A. veronii | A. veronii DSM 7386T HAM | 2.38 |
| D553 | A. veronii | A. veronii CECT4257T DSM | 2.30 |
| D179 | A. allosaccharophila ATCC 51208 (CECT4911) | A. veronii DSM 11576THAM | 2.28 |
| LAB ID | WGS Identification | AMP | AMC | PRL | TPZ | CFZ | FOX | CXM | CTX | CAZ | ATM | FEP | IMP | ETP | NAL | CIP | SXT |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| D180 | A. rivipollensis | 6 | 7 | 24 | 24 | 14 | 33 | 38 | 30 | 37 | 46 | 42 | 32 | 32 | 33 | 36 | 29 |
| D175 | A. media | 6 | 12 | 22 | 25 | 7 | 16 | 26 | 30 | 29 | S | 36 | 23 | 27 | 31 | 34 | 19 |
| D174 | A. caviae | 6 | 12 | 22 | 26 | 9 | 17 | 28 | 35 | 31 | 40 | 36 | 23 | 24 | 31 | 37 | 22 |
| D549 | A. caviae | 6 | 16 | 25 | 31 | 19 | 33 | 38 | 36 | 34 | 45 | 40 | 27 | 30 | 6 | 34 | 29 |
| D550 | A. caviae | 6 | 17 | 27 | 26 | 28 | 34 | 38 | 39 | 35 | 42 | 40 | 21 | 26 | 6 | 26 | S |
| D552 | A. caviae | 6 | 12 | 28 | 28 | 7 | 17 | 24 | 32 | 26 | 42 | 36 | 30 | 22 | 6 | 27 | 22 |
| D173 | A. hydrophila | 27 | 30 | 29 | 32 | 6 | 11 | 30 | 29 | 28 | 36 | 34 | 34 | 32 | 32 | 30 | 28 |
| D547 | A. dhakensis | 6 | 11 | 23 | 31 | 6 | 6 | 25 | 16 | 31 | 52 | 40 | 25 | 27 | 34 | 39 | 34 |
| D178 | A. veronii | 6 | 12 | 20 | 23 | 15 | 27 | 30 | 30 | 33 | 34 | 32 | 18 | 22 | 31 | 37 | 22 |
| D551 | A. veronii | 6 | 26 | 32 | 32 | 13 | 29 | 36 | 36 | 36 | 44 | 40 | 35 | S | 6 | 29 | 6 |
| D553 | A. veronii | 6 | 15 | 25 | 28 | 8 | 22 | 34 | 38 | 32 | 42 | 40 | 28 | 22 | 6 | 31 | 22 |
| D179 | A. allosaccharophila | 6 | 16 | 20 | 23 | 6 | 20 | 32 | 6 | 32 | 44 | 36 | 21 | 23 | 34 | 40 | 21 |
| Species | ResFinder (% ID) | CARD (% ID) | ||||
|---|---|---|---|---|---|---|
| Class B | Class C | Class D | Class B | Class C | Class D | |
| A. rivipollensis (n = 1) | blaCMY-1 (95.21) blaCMY-8b (94.87) | blaOXA-427 (100) | blaMOX-9 (99.74–99.48) | blaOXA-427 (100) | ||
| A. media (n = 2) | blaCMY-8b (96.26) | blaOXA-427 (100–99.12) | blaMOX-9 (100) | blaOXA-427 (100–99.12) | ||
| A. caviae (n = 5) | blaMOX-5 (99.91) blaMOX-6 (99.91) blaMOX-7 (100) (n = 3) | blaOXA-427 (100–98.87) | blaMOX-5 (99.91) blaMOX-6 (99.91) blaMOX-7 (100) (n = 3) | blaOXA-427 (100–98.87) | ||
| A. hydrophila (n = 2) | blaImiS (99.61) | blaMOX-5 (91.67) | blaAmpS (100) | blaImiS (99.61) | blaCepS (100) | blaOXA-12 (98.74) |
| A. dhakensis (n = 2) | blaCphA2 (100) blaImiH (99.74) | blaCMY-8b (91.99–98.35) | blaAmpH (100) | blaCphA2 (100) blaImiH (99.74) | blaAQU-2 (100-93.7) | blaOXA-724 (100) |
| A. sobria (n = 2) | blaCphA8 (100) | blaFOX-2 (98.09) | blaAmpS (98.87) | blaCphA8 (100) | blaFOX-2 (98.09) | blaOXA-12 (97.48) |
| A. veronii (n = 4) | blaCphA3 (100) blaCphA4 (99.48) blaCphA6 (99.87) (n = 2) | blaAmpS (100) | blaCphA3 (100) blaCphA4 (99.48) blaCphA6 (99.87) (n = 2) | blaOXA-12 (99.12) | ||
| A. allosacharophila (n = 2) | blaCphA4 (99.48) | blaFOX-7 (100) blaFOX-2 (100) | blaAmpS (98.87) | blaCphA4 (99.48) | blaFOX-7 (100) blaFOX-2 (100) | blaOXA-12 (98.99–98.74) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bertran, X.; Rubio, M.; Gómez, L.; Llovet, T.; Muñoz, C.; Navarro, F.; Miro, E. Taxonomic Identification of Different Species of the Genus Aeromonas by Whole-Genome Sequencing and Use of Their Species-Specific β-Lactamases as Phylogenetic Markers. Antibiotics 2021, 10, 354. https://doi.org/10.3390/antibiotics10040354
Bertran X, Rubio M, Gómez L, Llovet T, Muñoz C, Navarro F, Miro E. Taxonomic Identification of Different Species of the Genus Aeromonas by Whole-Genome Sequencing and Use of Their Species-Specific β-Lactamases as Phylogenetic Markers. Antibiotics. 2021; 10(4):354. https://doi.org/10.3390/antibiotics10040354
Chicago/Turabian StyleBertran, Xavier, Marc Rubio, Laura Gómez, Teresa Llovet, Carme Muñoz, Ferran Navarro, and Elisenda Miro. 2021. "Taxonomic Identification of Different Species of the Genus Aeromonas by Whole-Genome Sequencing and Use of Their Species-Specific β-Lactamases as Phylogenetic Markers" Antibiotics 10, no. 4: 354. https://doi.org/10.3390/antibiotics10040354
APA StyleBertran, X., Rubio, M., Gómez, L., Llovet, T., Muñoz, C., Navarro, F., & Miro, E. (2021). Taxonomic Identification of Different Species of the Genus Aeromonas by Whole-Genome Sequencing and Use of Their Species-Specific β-Lactamases as Phylogenetic Markers. Antibiotics, 10(4), 354. https://doi.org/10.3390/antibiotics10040354







