Surface Functionalization by Hydrophobin-EPSPS Fusion Protein Allows for the Fast and Simple Detection of Glyphosate
Abstract
1. Introduction
2. Materials and Methods
2.1. Chemicals
2.2. Molecular Cloning of Ccg2 and Ccg2_GS_EcEPSPS
2.3. Protein Expression and Purification
2.4. Surface Functionalization and Contact Angle Measurement
2.5. Measurement of Enzyme Activity in Solution
2.6. Measurement of Enzyme Activity of the Immobilized Proteins
2.7. Statistics
3. Results
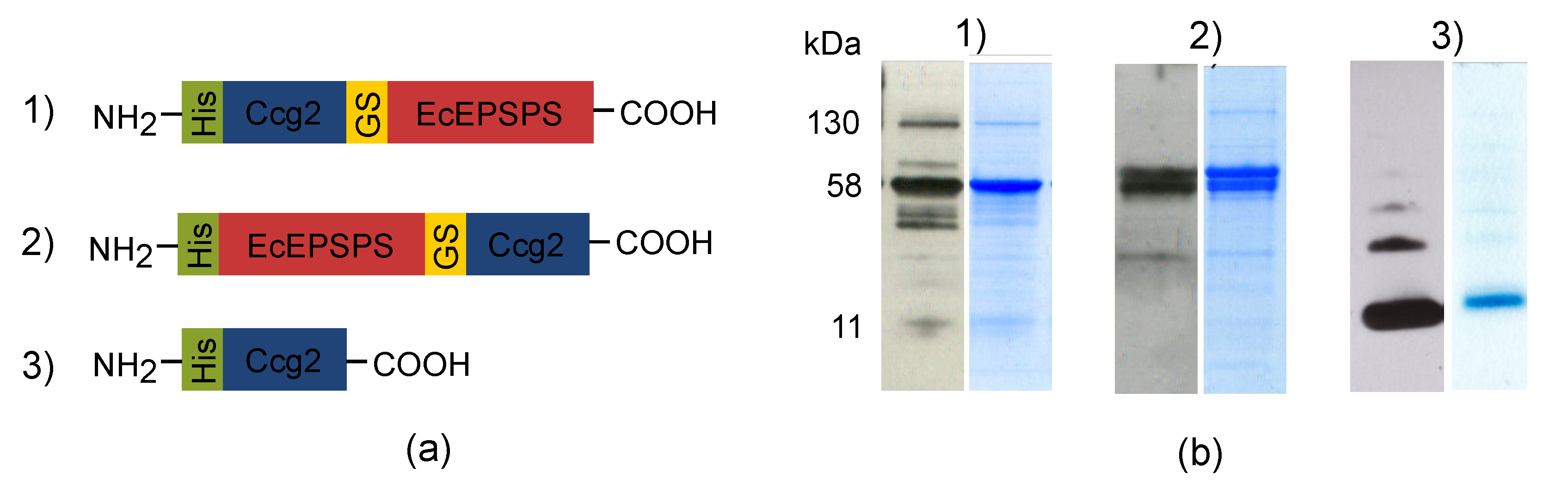
3.1. Cloning of Fusion Genes and Protein Purification
3.2. Self-Assembling Properties and Activity Measurement in Solution
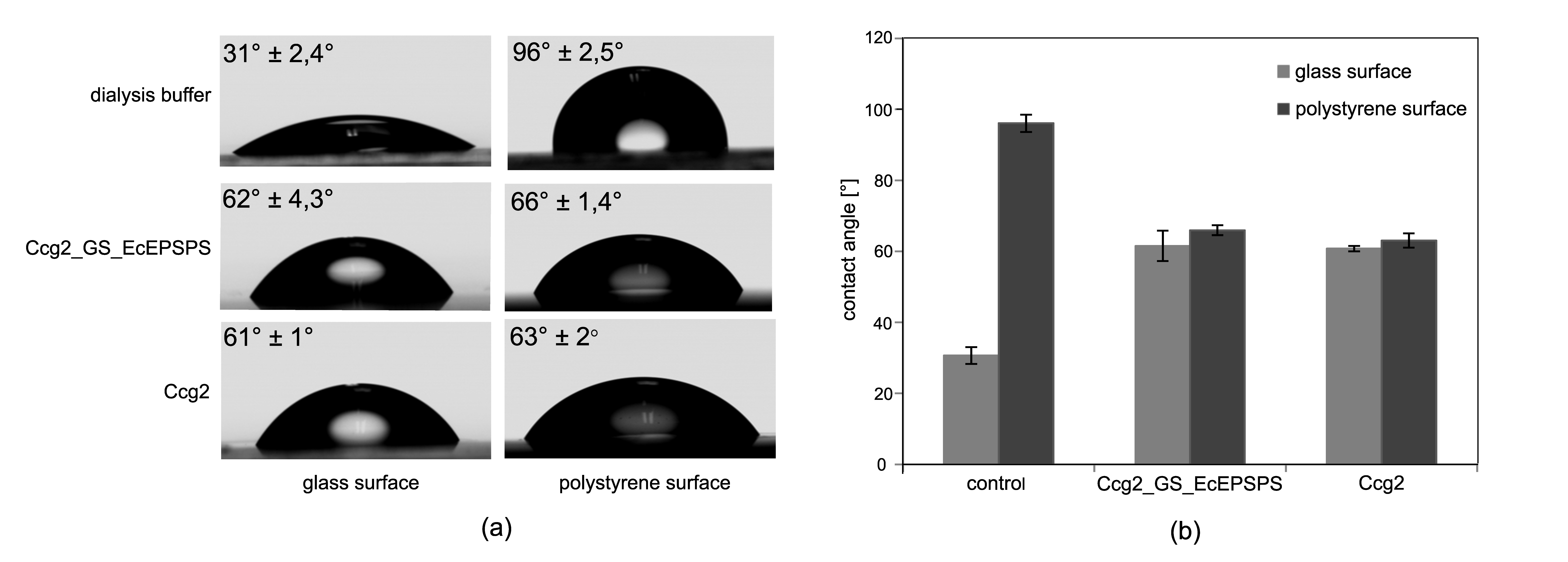
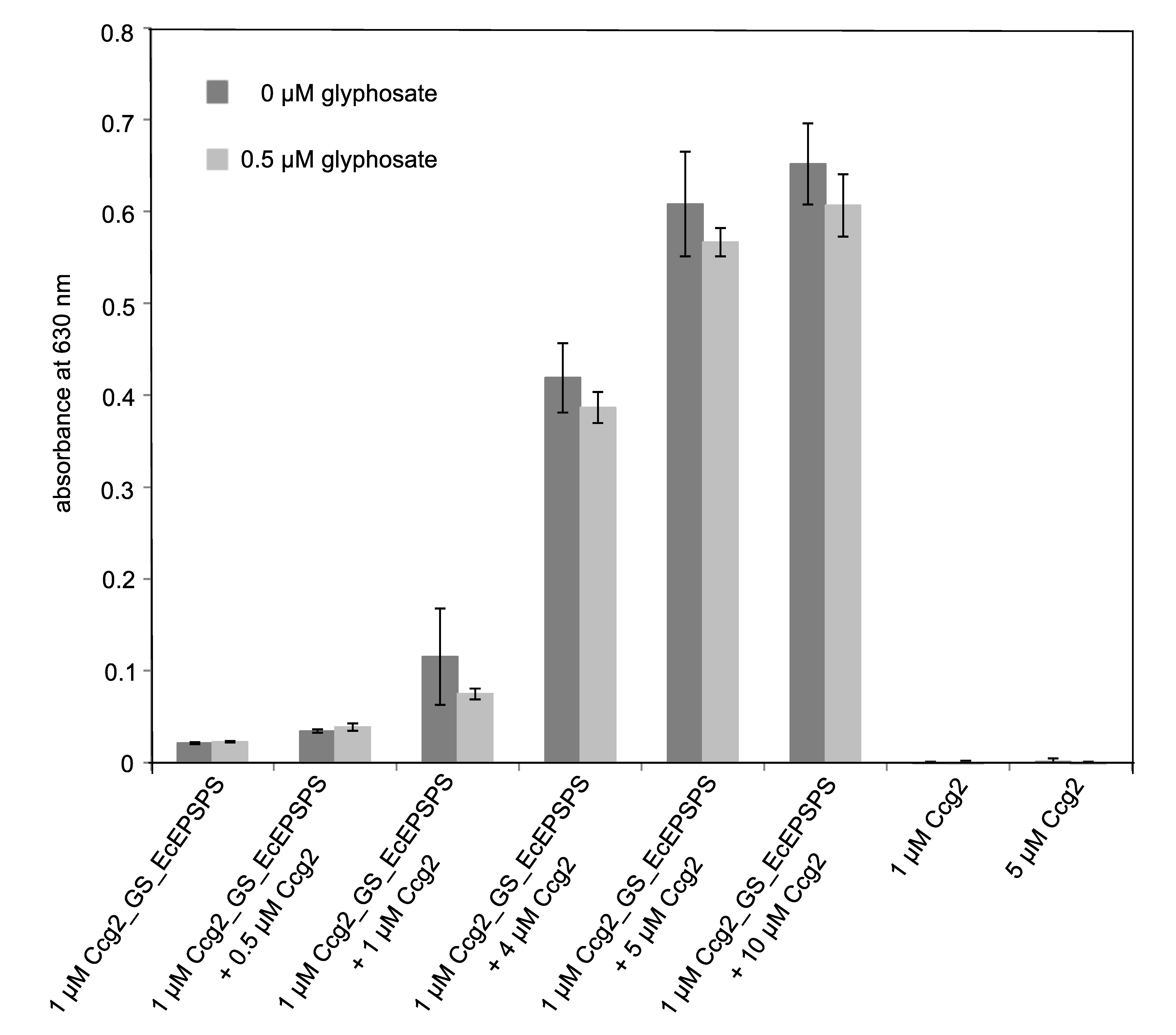
3.3. Surface Functionalization and Determination of Occupancy Ratio
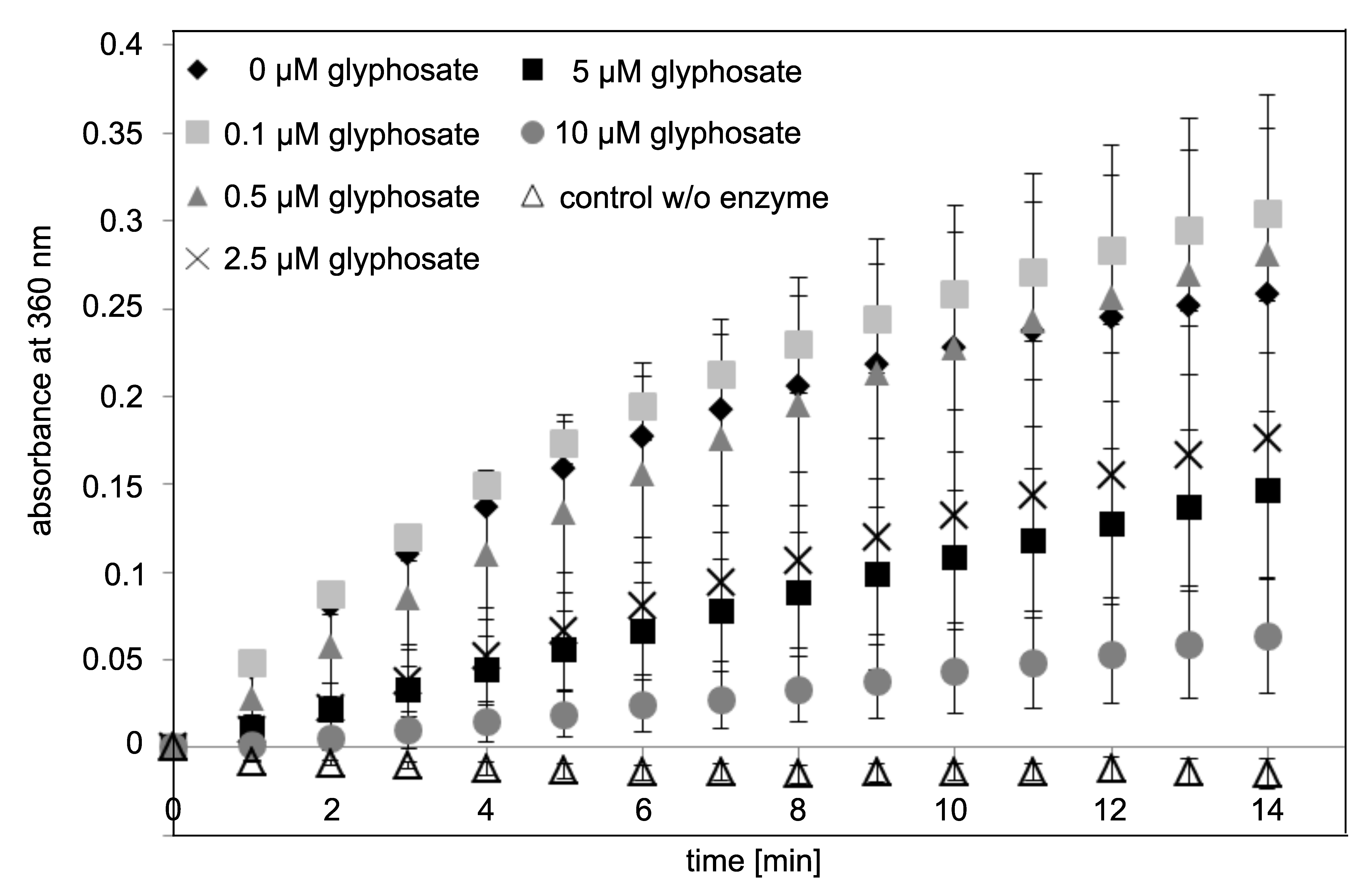
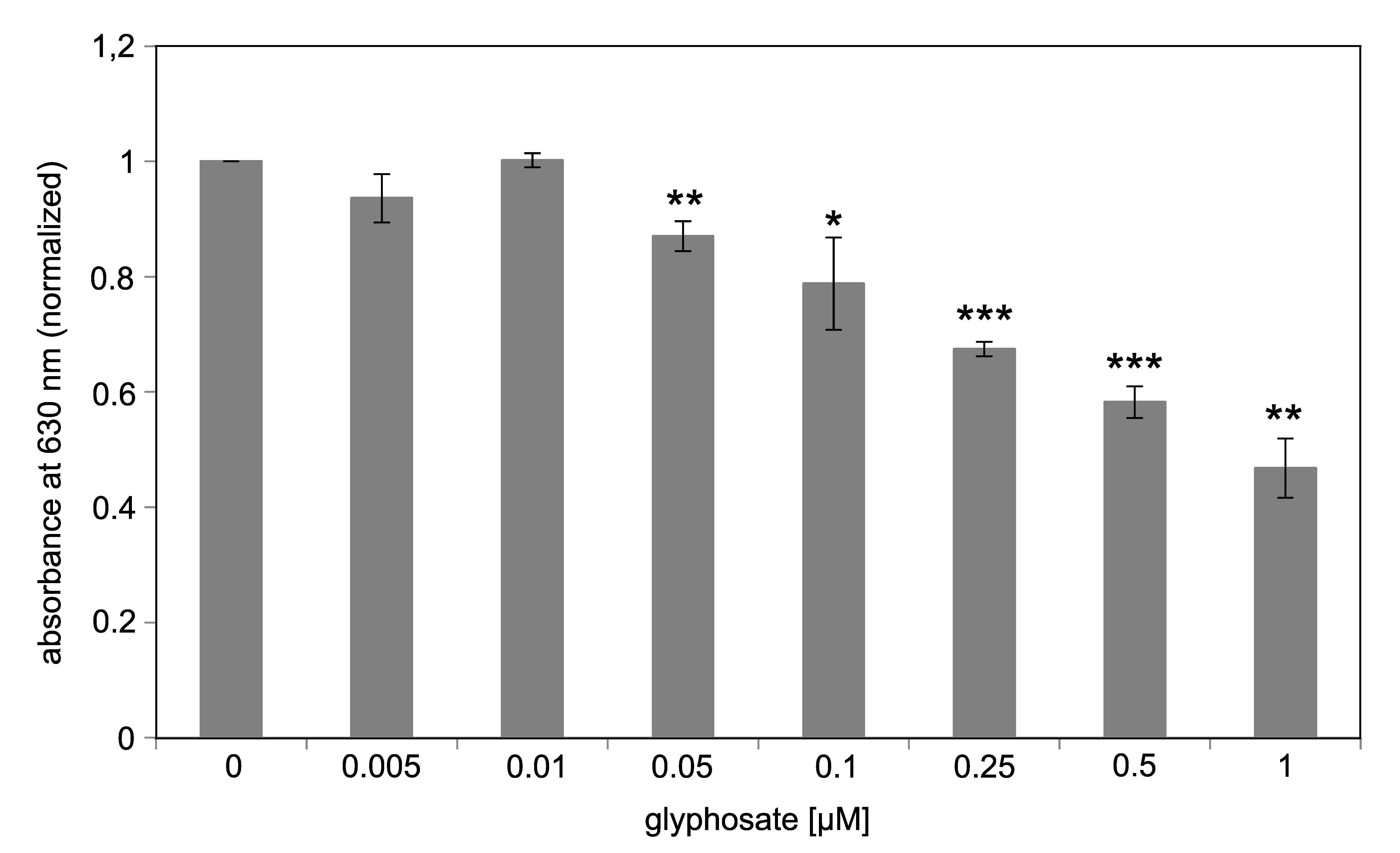
3.4. Concentration-Dependent Inhibition of EPSPS Enzymatic Activity by Glyphosate
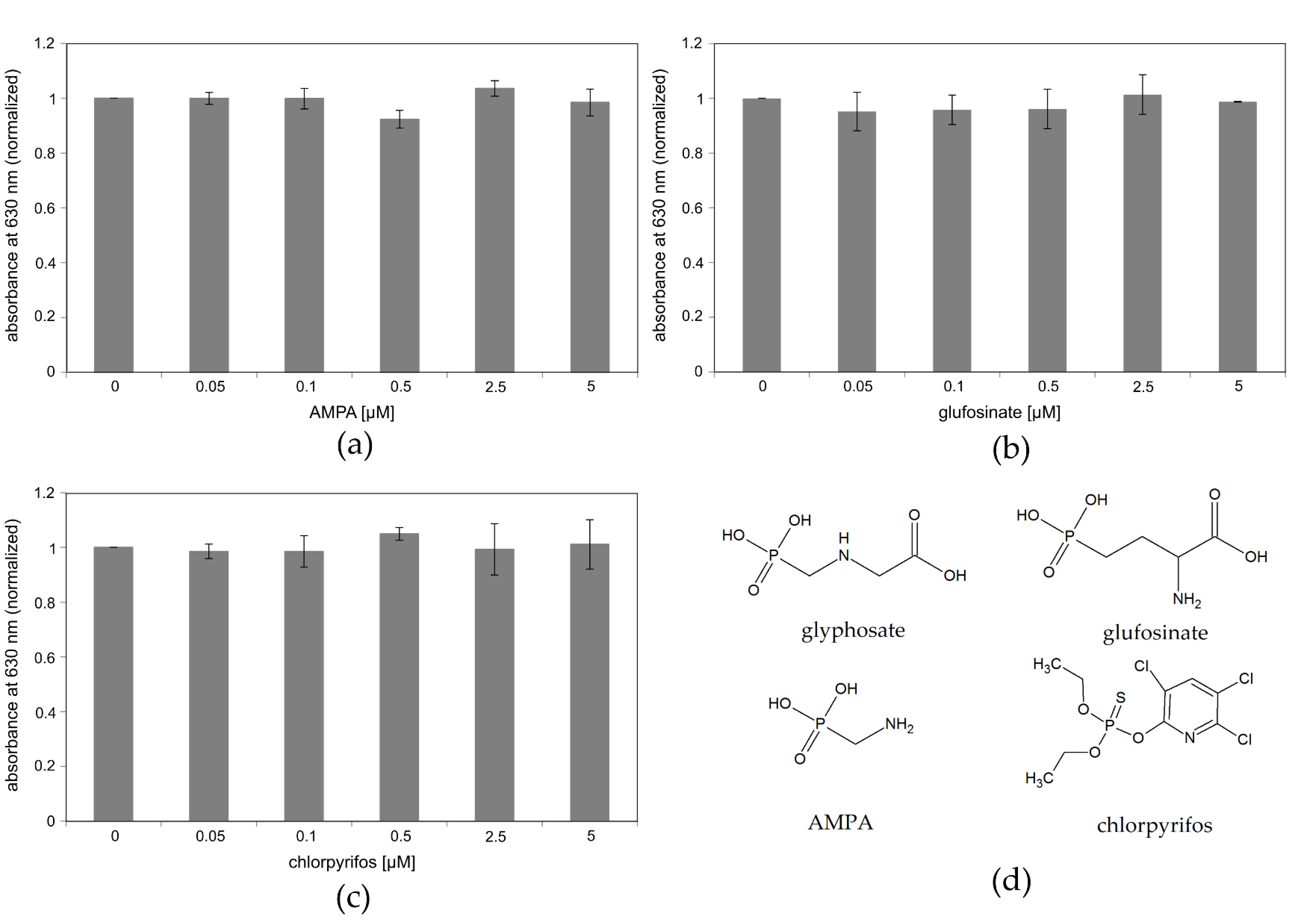
3.5. Cross Reactivity of the Assay
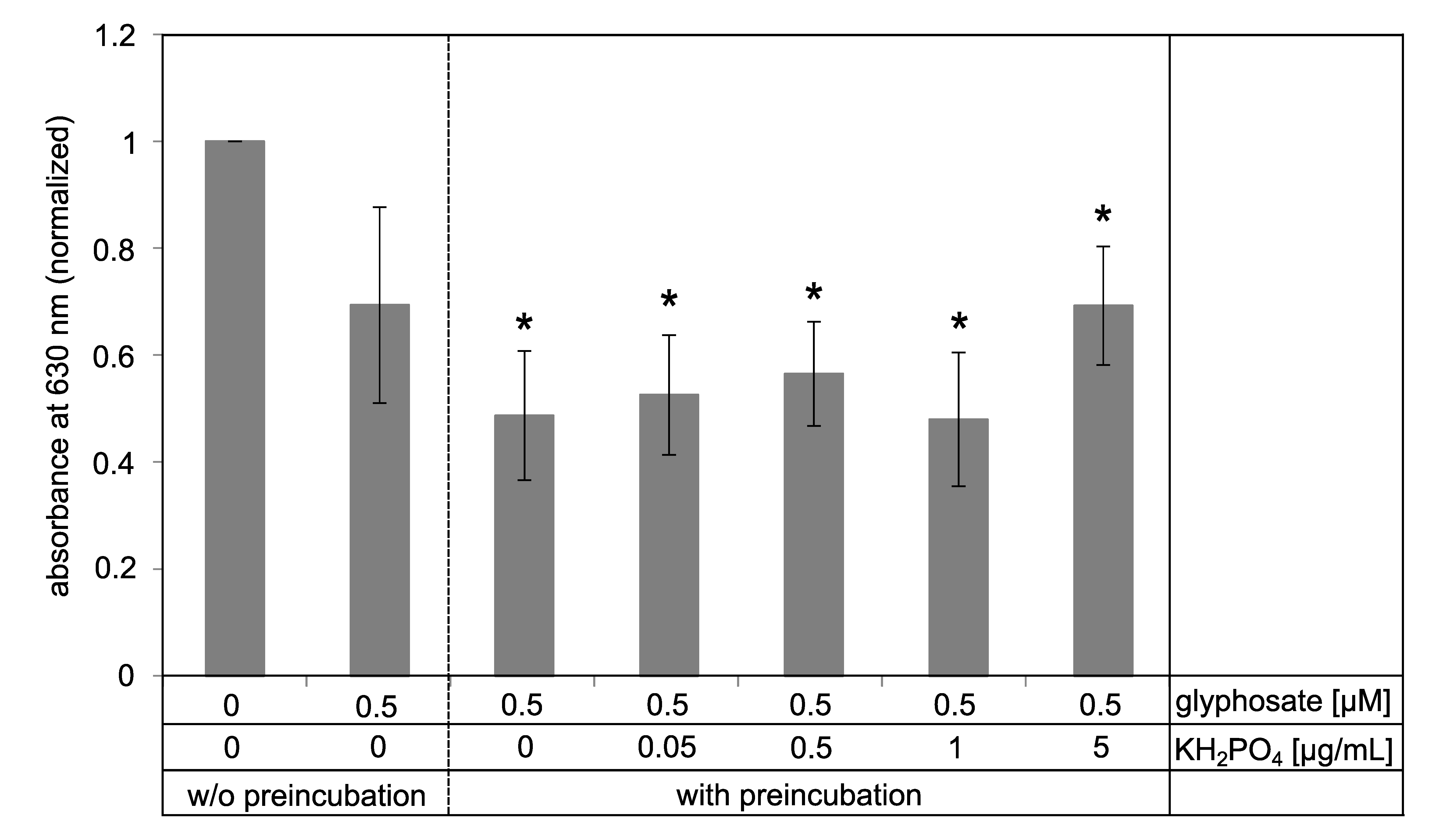
3.6. Pre-Incubation of the Functionalized Surface with Glyphosate Solution
4. Discussion
4.1. Contact Angle Measurements
4.2. Activity Measurement and Glyphosate Inhibition
4.3. Practicability of the Assay
5. Conclusions
6. Patents
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Benachour, N.; Séralini, G.-E. Glyphosate Formulations Induce Apoptosis and Necrosis in Human Umbilical, Embryonic, and Placental Cells. Chem. Res. Toxicol. 2009, 22, 97–105. [Google Scholar] [CrossRef] [PubMed]
- Guyton, K.Z.; Loomis, D.; Grosse, Y.; El Ghissassi, F.; Benbrahim-Tallaa, L.; Guha, N.; Scoccianti, C.; Mattock, H.; Straif, K. Carcinogenicity of tetrachlorvinphos, parathion, malathion, diazinon, and glyphosate. Lancet Oncol. 2015, 16, 490–491. [Google Scholar] [CrossRef]
- Tarazona, J.V.; Court-Marques, D.; Tiramani, M.; Reich, H.; Pfeil, R.; Istace, F.; Crivellente, F. Glyphosate toxicity and carcinogenicity: A review of the scientific basis of the European Union assessment and its differences with IARC. Arch. Toxicol. 2017, 91, 2723–2743. [Google Scholar] [CrossRef] [PubMed]
- Giesy, J.P.; Dobson, S.; Solomon, K.R. Ecotoxicological risk assessment for Roundup herbicide. In Reviews of Environmental Contamination and Toxicology; Ware, G.W., Ed.; Springer Verlag: New York, NY, USA, 2000; Volume 167, pp. 35–120. [Google Scholar]
- Zaller, J.G.; Heigl, F.; Ruess, L.; Grabmaier, A. Glyphosate herbicide affects belowground interactions between earthworms and symbiotic mycorrhizal fungi in a model ecosystem. Sci. Rep. 2014, 4, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Annett, R.; Habibi, H.R.; Hontela, A. Impact of glyphosate and glyphosate-based herbicides on the freshwater environment. J. Appl. Toxicol. 2014, 34, 458–479. [Google Scholar] [CrossRef] [PubMed]
- Glyphosate Testing Report: Findings in American Mothers’ Breast Milk, Urine and Water. Available online: https://www.momsacrossamerica.com/glyphosate_testing_results (accessed on 20 February 2019).
- Jansons, M.; Pugajeva, I.; Bartkevics, V. Occurrence of glyphosate in beer from the Latvian market. Food Addit. Contam. Part A 2018, 35, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Krüger, M.; Schledorn, P.; Schrödl, W.; Hoppe, H.-W.; Lutz, W.; Shehata, A.A. Detection of Glyphosate residues in animals and humans. Environ. Anal. Toxicol. 2014, 4, 1–5. [Google Scholar]
- Jensen, P.K.; Wujcik, C.E.; McGuire, M.K.; McGuire, M.A. Validation of reliable and selective methods for direct determination of glyphosate and aminomethylphosphonic acid in milk and urine using LC-MS/MS. J. Environ. Sci. Health Part B 2016, 51, 254–259. [Google Scholar] [CrossRef]
- Ehling, S.; Reddy, T.M. Analysis of Glyphosate and Aminomethylphosphonic Acid in Nutritional Ingredients and Milk by Derivatization with Fluorenylmethyloxycarbonyl Chloride and Liquid Chromatography–Mass Spectrometry. J. Agric. Food Chem. 2015, 63, 10562–10568. [Google Scholar] [CrossRef]
- Steinborn, A.; Alder, L.; Michalski, B.; Zomer, P.; Bendig, P.; Martinez, S.A.; Class, T.J.; Pinheiro, N.C.; Mol, H.G.J. Determination of Glyphosate Levels in Breast Milk Samples from Germany by LC-MS/MS and GC-MS/MS. J. Agric. Food Chem. 2016, 64, 1414–1421. [Google Scholar] [CrossRef]
- McGuire, M.K.; Price, W.J.; Shafii, B.; Carrothers, J.M.; A Lackey, K.; A Goldstein, D.; Jensen, P.K.; Vicini, J.L. Glyphosate and aminomethylphosphonic acid are not detectable in human milk. Am. J. Clin. Nutr. 2016, 103, 1285–1290. [Google Scholar] [CrossRef] [PubMed]
- Richard, S.; Moslemi, S.; Sipahutar, H.; Benachour, N.; Séralini, G.-E. Differential Effects of Glyphosate and Roundup on Human Placental Cells and Aromatase. Environ. Health Perspect. 2005, 113, 716–720. [Google Scholar] [CrossRef] [PubMed]
- Szekacs, I.; Farkas, E.; Gemes, B.L.; Takacs, E.; Szekacs, A.; Horvath, R. Integrin targeting of glyphosate and its cell adhesion modulation effects on osteoblastic MC3T3-E1 cells revealed by label-free optical biosensing. Sci. Rep. 2018, 8, 17401. [Google Scholar] [CrossRef] [PubMed]
- IARC Monographs Volume 112: Evaluation of Five Organophosphate Insecticides and Herbicides. Available online: https://www.iarc.fr/wp-content/uploads/2018/07/MonographVolume112-1.pdf (accessed on 29 July 2019).
- Sanchís, J.; Kantiani, L.; Llorca, M.; Rubio, F.; Ginebreda, A.; Fraile, J.; Garrido, T.; Farré, M. Determination of glyphosate in groundwater samples using an ultrasensitive immunoassay and confirmation by on-line solid-phase extraction followed by liquid chromatography coupled to tandem mass spectrometry. Anal. Bioanal. Chem. 2012, 402, 2335–2345. [Google Scholar] [CrossRef] [PubMed]
- Okada, E.; Coggan, T.; Anumol, T.; Clarke, B.; Allinson, G. A simple and rapid direct injection method for the determination of glyphosate and AMPA in environmental water samples. Anal. Bioanal. Chem. 2019, 411, 715–724. [Google Scholar] [CrossRef] [PubMed]
- Eberbach, P.L.; Douglas, L.A. Method for the determination of glyphosate and (aminomethyl)phosphonic acid in soil using electron capture gas chromatography. J. Agric. Food Chem. 1991, 39, 1776–1780. [Google Scholar] [CrossRef]
- Rubio, F.; Veldhuis, L.J.; Clegg, B.S.; Fleeker, J.R.; Hall, J.C. Comparison of a Direct ELISA and an HPLC Method for Glyphosate Determinations in Water. J. Agric. Food Chem. 2003, 51, 691–696. [Google Scholar] [CrossRef]
- Reynoso, E.C.; Torres, E.; Bettazzi, F.; Palchetti, I. Trends and Perspectives in Immunosensors for Determination of Currently-Used Pesticides: The Case of Glyphosate, Organophosphates, and Neonicotinoids. Biosensors 2019, 9, 20. [Google Scholar] [CrossRef]
- Jan, M.R.; Shah, J.; Muhammad, M.; Ara, B.; Muhammad, D.M. Glyphosate herbicide residue determination in samples of environmental importance using spectrophotometric method. J. Hazard. Mater. 2009, 169, 742–745. [Google Scholar] [CrossRef]
- Sharma, M.; Schmid, M.; Rothballer, M.; Hause, G.; Zuccaro, A.; Imani, J.; Kampfer, P.; Domann, E.; Schafer, P.; Hartmann, A.; et al. Detection and identification of bacteria intimately associated with fungi of the order Sebacinales. Cell. Microbiol. 2008, 10, 2235–2246. [Google Scholar] [CrossRef]
- De Almeida, L.; Chigome, S.; Torto, N.; Frost, C.; Pletschke, B. A novel colorimetric sensor strip for the detection of glyphosate in water. Sens. Actuators B Chem. 2015, 206, 357–363. [Google Scholar] [CrossRef]
- Steinrücken, H.; Amrhein, N. The herbicide glyphosate is a potent inhibitor of 5-enolpyruvylshikimic acid-3-phosphate synthase. Biochem. Biophys. Res. Commun. 1980, 94, 1207–1212. [Google Scholar] [CrossRef]
- Amrhein, N.; Schab, J.; Steinrücken, H.C. The mode of action of the herbicide glyphosate. Naturwissenschaften 1980, 67, 356–357. [Google Scholar] [CrossRef]
- Rogers, S.G.; A Brand, L.; Holder, S.B.; Sharps, E.S.; Brackin, M.J. Amplification of the aroA gene from Escherichia coli results in tolerance to the herbicide glyphosate. Appl. Environ. Microbiol. 1983, 46, 37–43. [Google Scholar] [PubMed]
- Herrmann, K.M.; Weaver, L.M. The shikimate pathway. Annu. Rev. Plant. Biol. 1999, 50, 473–503. [Google Scholar] [CrossRef] [PubMed]
- Levin, J.G.; Sprinson, D.B. The enzymatic formation and isolation of 3- enolpyruvylshikimate 5-phosphate. J. Biol. Chem. 1964, 239, 1142–1151. [Google Scholar] [PubMed]
- Steinrücken, H.; Amrhein, N. 5-Enolpyruvylshikimate-3-phosphate synthase of Klebsiella pneumoniae. 1. Purification and properties. Eur. J. Biochem. 1984, 143, 341–349. [Google Scholar] [CrossRef]
- Alibhai, M.F.; Stallings, W.C. Closing down on glyphosate inhibition—With a new structure for drug discovery. Proc. Natl. Acad. Sci. USA 2001, 98, 2944–2946. [Google Scholar] [CrossRef]
- Schönbrunn, E.; Eschenburg, S.; Shuttleworth, W.A.; Schloss, J.V.; Amrhein, N.; Evans, J.N.S.; Kabsch, W. Interaction of the herbicide glyphosate with its target enzyme 5-enolpyruvylshikimate 3-phosphate synthase in atomic detail. Proc. Natl. Acad. Sci. USA 2001, 98, 1376–1380. [Google Scholar] [CrossRef]
- Itaya, K.; Ui, M. A new micromethod for the colorimetric determination of inorganic phosphate. Clin. Chim. Acta 1966, 14, 361–366. [Google Scholar] [CrossRef]
- Van Veldhoven, P.P.; Mannaerts, G.P. Inorganic and organic phosphate measurements in the nanomolar range. Anal. Biochem. 1987, 161, 45–48. [Google Scholar] [CrossRef]
- Milin, S. Comparaison de deux méthodes spectrophotométriques de dosage de l’acide phosphorique. Application à des sols et des végétaux. Cah. Tech. INRA 2012, 77, 1–15. [Google Scholar]
- Baykov, A.; Evtushenko, O.; Avaeva, S. A malachite green procedure for orthophosphate determination and its use in alkaline phosphatase-based enzyme immunoassay. Anal. Biochem. 1988, 171, 266–270. [Google Scholar] [CrossRef]
- Aguiar-Oliveira, E.; Maugeri, F. Thermal stability of the immobilized fructosyltransferase from Rhodotorula sp. Braz. J. Chem. Eng. 2011, 28, 363–372. [Google Scholar] [CrossRef]
- Hanefeld, U.; Gardossi, L.; Magner, E. Understanding enzyme immobilisation. Chem. Soc. Rev. 2009, 38, 453–468. [Google Scholar] [CrossRef] [PubMed]
- Boundy, J.A.; Smiley, K.L.; Swanson, C.L.; Hofreiter, B.T. Exoenzymic activity of alpha-amylase immobilized on a phenol-formaldehyde resin. Carbohydr. Res. 1976, 48, 239–244. [Google Scholar] [CrossRef]
- Brena, B.; González-Pombo, P.; Batista-Viera, F. Chapter 2 Immobilization of Enzymes: A Literature Survey. In Immobilization of Enzymes and Cells, 3rd ed.; Guisan, J.M., Ed.; Springer Science+Business: New York, NY, USA, 2013; Volume 1051, pp. 15–31. [Google Scholar]
- Sheldon, R.A. Enzyme Immobilization: The Quest for Optimum Performance. Adv. Synth. Catal. 2007, 349, 1289–1307. [Google Scholar] [CrossRef]
- Hektor, H.J.; Scholtmeijer, K. Hydrophobins: Proteins with potential. Curr. Opin. Biotechnol. 2005, 16, 434–439. [Google Scholar] [CrossRef] [PubMed]
- Bayry, J.; Aimanianda, V.; Guijarro, J.I.; Sunde, M.; Latgé, J.-P. Hydrophobins—Unique Fungal Proteins. PLoS Pathog. 2012, 8, e1002700. [Google Scholar] [CrossRef]
- Wessels, J.G.H. Developmental Regulation of Fungal Cell Wall Formation. Annu. Rev. Phytopathol. 1994, 32, 413–437. [Google Scholar] [CrossRef]
- Wösten, H.A.B.; de Vocht, M.L. Hydrophobins, the fungal coat unravelled. Biochim. Biophys. Acta. 2000, 1469, 79–86. [Google Scholar] [CrossRef]
- Wosten, H.; De Vries, O.; Wessels, J. Interfacial Self-Assembly of a Fungal Hydrophobin into a Hydrophobic Rodlet Layer. Plant Cell 1993, 5, 1567–1574. [Google Scholar] [CrossRef] [PubMed]
- Linder, M.B. Hydrophobins: Proteins that self assemble at interfaces. Curr. Opin. Colloid Interface Sci. 2009, 14, 356–363. [Google Scholar] [CrossRef]
- Lo, V.C.; Ren, Q.; Pham, C.L.L.; Morris, V.K.; Kwan, A.H.; Sunde, M. Fungal Hydrophobin Proteins Produce Self-Assembling Protein Films with Diverse Structure and Chemical Stability. Nanomaterials 2014, 4, 827–843. [Google Scholar] [CrossRef] [PubMed]
- Soikkeli, M.; Kurppa, K.; Kainlauri, M.; Arpiainen, S.; Paananen, A.; Gunnarsson, D.; Joensuu, J.J.; Laaksonen, P.; Prunnila, M.; Linder, M.B.; et al. Graphene Biosensor Programming with Genetically Engineered Fusion Protein Monolayers. ACS Appl. Mater. Interfaces 2016, 8, 8257–8264. [Google Scholar] [CrossRef] [PubMed]
- Hennig, S.; Rödel, G.; Ostermann, K. Hydrophobin-Based Surface Engineering for Sensitive and Robust Quantification of Yeast Pheromones. Sensors 2016, 16, 602. [Google Scholar] [CrossRef] [PubMed]
- Bell-Pedersen, D.; Loros, J.J.; Dunlap, J.C. The Neurospora circadian clock-controlled gene, ccg-2, is allelic to eas and encodes a fungal hydrophobin required for formation of the conidial rodlet layer. Genes Dev. 1992, 6, 2382–2394. [Google Scholar] [CrossRef] [PubMed]
- Lauter, F.R.; E Russo, V.; Yanofsky, C. Developmental and light regulation of eas, the structural gene for the rodlet protein of Neurospora. Genes Dev. 1992, 6, 2373–2381. [Google Scholar] [CrossRef] [PubMed]
- MacIndoe, I.; Kwan, A.H.; Ren, Q.; Morris, V.K.; Yang, W.; Mackay, J.P.; Sunde, M. Self-assembly of functional, amphipathic amyloid monolayers by the fungal hydrophobin EAS. Proc. Natl. Acad. Sci. USA 2012, 109, E804–E811. [Google Scholar] [CrossRef]
- Kwan, A.H.Y.; Winefield, R.D.; Sunde, M.; Matthews, J.M.; Haverkamp, R.G.; Templeton, M.D.; Mackay, J.P. Structural basis for rodlet assembly in fungal hydrophobins. Proc. Natl. Acad. Sci. USA 2006, 103, 3621–3626. [Google Scholar] [CrossRef] [PubMed]
- Sammons, R.D.; Meyer, J.; Hall, E.; Ostrander, E.; Schrader, S. A Simple Continuous Assay for EPSP Synthase in Plant Tissue. 2007 (Poster). Available online: https://www.cottoninc.com/wp-content/uploads/2017/03/11a-Industry-Sammons-NCWSS07-poster.pdf (accessed on 28 June 2016).
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Haider, S.R.; Reid, H.J.; Sharp, B.L. Tricine-SDS-PAGE. In Protein Electrophoresis: Methods and Protocols; Kurien, B.T., Scofield, R.H., Eds.; Humana Press: Totowa, NJ, USA, 2012; Volume 869, pp. 81–91. [Google Scholar]
- Laemmli, U.K. Cleavage of structural proteins during assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef] [PubMed]
- De Vocht, M.L.; Reviakine, I.; Wösten, H.A.B.; Brisson, A.; Wessels, J.G.H.; Robillard, G.T. Structural and Functional Role of the Disulfide Bridges in the Hydrophobin SC3. J. Boil. Chem. 2000, 275, 28428–28432. [Google Scholar] [CrossRef] [PubMed]
- Kershaw, M.J.; Thornton, C.R.; Wakley, G.E.; Talbot, N.J. Four conserved intramolecular disulphide linkages are required for secretion and cell wall localization of a hydrophobin during fungal morphogenesis. Mol. Microbiol. 2005, 56, 117–125. [Google Scholar] [CrossRef] [PubMed]
- Schägger, H. Tricine-SDS-PAGE. Nat. Protoc. 2006, 1, 16–23. [Google Scholar] [CrossRef] [PubMed]
- Law, K.-Y. Definitions for Hydrophilicity, Hydrophobicity, and Superhydrophobicity: Getting the Basics Right. J. Phys. Chem. Lett. 2014, 5, 686–688. [Google Scholar] [CrossRef] [PubMed]
- Boeuf, S.; Throm, T.; Gutt, B.; Strunk, T.; Hoffmann, M.; Seebach, E.; Mühlberg, L.; Brocher, J.; Gotterbarm, T.; Wenzel, W.; et al. Engineering hydrophobin DewA to generate surfaces that enhance adhesion of human but not bacterial cells. Acta Biomater. 2012, 8, 1037–1047. [Google Scholar] [CrossRef]
- Takatsuji, Y.; Yamasaki, R.; Iwanaga, A.; Lienemann, M.; Linder, M.B.; Haruyama, T. Solid-support immobilization of a “swing” fusion protein for enhanced glucose oxidase catalytic activity. Colloids Surf. B Biointerfaces 2013, 112, 186–191. [Google Scholar] [CrossRef] [PubMed]
- Fokina, O.; Fenchel, A.; Winandy, L.; Fischer, R. Immobilization of LccC Laccase from Aspergillus nidulans on Hard Surfaces via Fungal Hydrophobins. Appl. Environ. Microbiol. 2016, 82, 6395–6402. [Google Scholar] [CrossRef] [PubMed]
- Wessels, J.G.H.; De Vries, O.M.H.; Ásgeirsdóttir, S.; Springer, J. The thn mutation of Schizophyllum commune, which suppresses formation of aerial hyphae, affects expression of the Sc3 hydrophobin gene. J. Gen. Microbiol. 1991, 137, 2439–2445. [Google Scholar] [CrossRef] [PubMed]
- De Vries, O.M.H.; Fekkes, M.P.; Wösten, H.A.B.; Wessels, J.G.H. Insoluble hydrophobin complexes in the walls of Schizophyllum commune and other filamentous fungi. Arch. Microbiol. 1993, 159, 330–335. [Google Scholar] [CrossRef]
- Wösten, H.A.B. Hydrophobins: Multipurpose Proteins. Annu. Rev. Microbiol. 2001, 55, 625–646. [Google Scholar] [CrossRef]
- Rubin, J.L.; Gaines, C.G.; Jensen, R.A. Glyphosate Inhibition of 5-Enolpyruvylshikimate 3-Phosphate Synthase from Suspension-Cultured Cells of Nicotiana silvestris. Plant Physiol. 1984, 75, 839–845. [Google Scholar] [CrossRef]
- Steinrücken, H.C.; Amrhein, N. 5-Enolpyruvylshikimate-3-phosphate synthase of Klebsiella pneumoniae. 2. Inhibition by glyphosate [N-(phosphononmethyl)glycine]. JBIC J. Boil. Inorg. Chem. 1984, 143, 351–357. [Google Scholar] [CrossRef]
- Boocock, M.R.; Coggins, J.R. Kinetics of 5-enolpyruvylshikimate-3-phosphate synthase inhibition by glyphosate. FEBS Lett. 1983, 154, 127–133. [Google Scholar] [CrossRef]
- Kwan, A.H.; MacIndoe, I.; Vukašin, P.V.; Morris, V.K.; Kass, I.; Gupte, R.; Mark, A.E.; Templeton, M.D.; Mackay, J.P.; Sunde, M. The Cys3–Cys4 Loop of the Hydrophobin EAS Is Not Required for Rodlet Formation and Surface Activity. J. Mol. Boil. 2008, 382, 708–720. [Google Scholar] [CrossRef]
- Rubio, F.; Glaze, T.; Lance, J.; Hutchinson, Z. Survey of Glyphosate in Domestic and Imported Beer and Wine. 2016 (Poster). Available online: https://www.abraxiskits.com/wp-content/uploads/2016/08/glyphosateinbeerwine.pdf (accessed on 22 February 2019).
- Clegg, B.S.; Stephenson, G.R.; Hall, J.C. Development of an Enzyme-Linked Immunosorbent Assay for the Detection of Glyphosate. J. Agric. Food Chem. 1999, 47, 5031–5037. [Google Scholar] [CrossRef]
- Lesueur, C.; Pfeffer, M.; Fuerhacker, M. Photodegradation of phosphonates in water. Chemosphere 2005, 59, 685–691. [Google Scholar] [CrossRef]
- Jaworski, E.G. Mode of action of N-phosphonomethylglycine. Inhibition of aromatic amino acid biosynthsis. J. Agric. Food Chem. 1972, 20, 1195–1198. [Google Scholar] [CrossRef]
- Grandcoin, A.; Piel, S.; Baurès, E. AminoMethylPhosphonic acid (AMPA) in natural waters: Its sources, behavior and environmental fate. Water Res. 2017, 117, 187–197. [Google Scholar] [CrossRef]
- Marzabadi, M.R.; Gruys, K.J.; Pansegrau, P.D.; Walker, M.C.; Yuen, H.K.; Sikorski, J.A. An EPSP Synthase Inhibitor Joining Shikimate 3-Phosphate with Glyphosate: Synthesis and Ligand Binding Studies. Biochemistry 1996, 35, 4199–4210. [Google Scholar] [CrossRef]
- Priestman, M.A.; Healy, M.L.; Becker, A.; Alberg, D.G.; Bartlett, P.A.; Lushington, G.H.; Schönbrunn, E. Interaction of phosphonate analogues of the tetrahedral reaction intermediate with 5-enolpyruvylshikimate-3-phosphate synthase in atomic detail. Biochemistry 2005, 44, 3241–3248. [Google Scholar] [CrossRef]
- Toy, A.D.F.; Uhing, E.H. Aminomethylenephosphinic acids, salts thereof, and process for their production. U.S. Patent 3,160,632, 8 December 1964. [Google Scholar]
- Mertens, M.; Höss, S.; Neumann, G.; Afzal, J.; Reichenbecher, W. Glyphosate, a chelating agent—Relevant for ecological risk assessment? Environ. Sci. Pollut. Res. 2018, 25, 5298–5317. [Google Scholar] [CrossRef]
- Anderson, K.S.; Sikorski, J.A.; Johnson, K.A. Evaluation of 5-enolpyruvylshikimate-3-phosphate synthase substrate and inhibitor binding by stopped-flow and equilibrium fluorescence measurements. Biochemistry 1988, 27, 1604–1610. [Google Scholar] [CrossRef]
- Ream, J.E.; Yuen, H.K.; Frazier, R.B.; Sikorski, J.A. EPSP synthase: Binding studies using isothermal titration microcalorimetry and equilibrium dialysis and their implications for ligand recognition and kinetic mechanism. Biochemistry 1992, 31, 5528–5534. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Döring, J.; Rettke, D.; Rödel, G.; Pompe, T.; Ostermann, K. Surface Functionalization by Hydrophobin-EPSPS Fusion Protein Allows for the Fast and Simple Detection of Glyphosate. Biosensors 2019, 9, 104. https://doi.org/10.3390/bios9030104
Döring J, Rettke D, Rödel G, Pompe T, Ostermann K. Surface Functionalization by Hydrophobin-EPSPS Fusion Protein Allows for the Fast and Simple Detection of Glyphosate. Biosensors. 2019; 9(3):104. https://doi.org/10.3390/bios9030104
Chicago/Turabian StyleDöring, Julia, David Rettke, Gerhard Rödel, Tilo Pompe, and Kai Ostermann. 2019. "Surface Functionalization by Hydrophobin-EPSPS Fusion Protein Allows for the Fast and Simple Detection of Glyphosate" Biosensors 9, no. 3: 104. https://doi.org/10.3390/bios9030104
APA StyleDöring, J., Rettke, D., Rödel, G., Pompe, T., & Ostermann, K. (2019). Surface Functionalization by Hydrophobin-EPSPS Fusion Protein Allows for the Fast and Simple Detection of Glyphosate. Biosensors, 9(3), 104. https://doi.org/10.3390/bios9030104





