S-Layer Protein-Based Biosensors
Abstract
1. Introduction
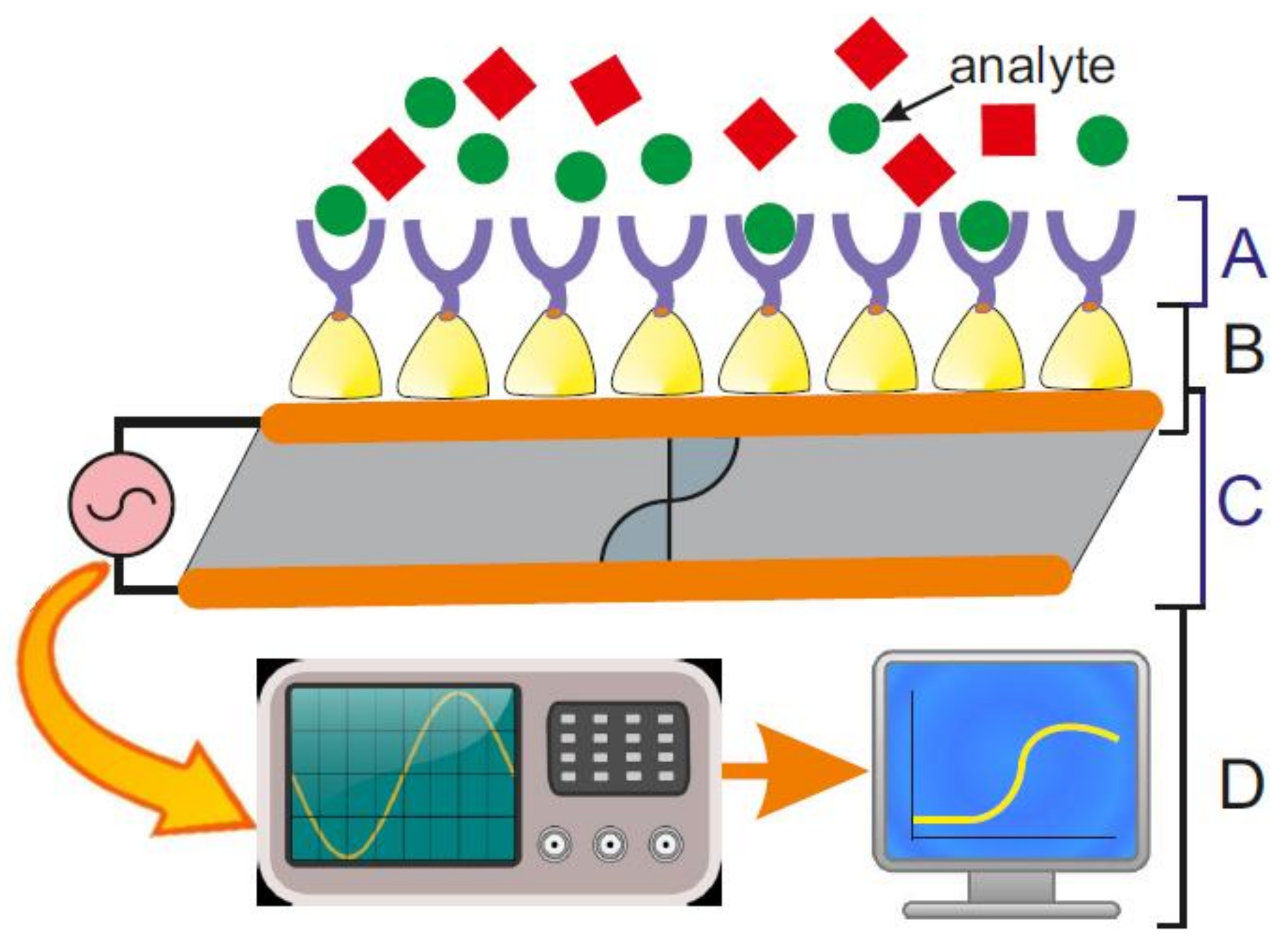
- (A)
- a biosensing element or bioreceptor, to which the analyte has a highly specific binding affinity;
- (B)
- an interface architecture, which provides an environment for the proper functioning of the biosensing element and where the specific biological event, which gives rise to a certain physical phenomenon takes place;
- (C)
- a transducer converting the physical phenomenon or chemical response resulting from the analyte’s interaction with the biological element (e.g., physicochemical, optical, piezoelectric, electrochemical, etc.) into electrical signals. The latter can be reproducibly measured, quantified and processed [3]; and,
- (D)
- an associated electronics comprising of signal amplifier, signal processor and an interface, like a display, which finally allows a user-friendly visualization and evaluation of the data [4].
2. Bacterial S-Layer Proteins
3. Modified S-Layers as Components in Biosensors
4. Genetically Engineered S-Layers as Components in Biosensors
5. S-Layer Lattices for Generation of Functional Lipid Membrane Platforms
6. Conclusions & Outlook
Acknowledgments
Conflicts of Interest
Abbreviations
| 2D | two-dimensional |
| αHL | α-hemolysin |
| AFM | atomic force microscopy |
| BLM | bilayer lipid membrane |
| CD133 | tumor marker |
| ChOx | cholesterol oxidase |
| CV | cyclovoltammetry |
| EBV | Epstein-Barr virus |
| EIS | electrochemical impedance spectroscopy |
| GOx | glucose oxidase |
| HepG2 | human liver carcinoma cells |
| IgG | immunoglobulin G |
| MCF-7 | human breast adenocarcinoma cell |
| nAChR | nicotinic acetylcholine receptor |
| NADH | nicotinamide adenine dinucleotide hydride |
| PGLa(-) | negatively charged analogue of peptidyl-glycylleucine-carboxyamide |
| PSA | prostate-specific antigen |
| QCM-D | quartz crystal microbalance with dissipation monitoring |
| RyR1 | ryanodine receptor/Ca2+ release channel |
| rSbpA/Lac | recombinant fusion protein comprising of SbpA and laccase |
| rSbpA/ZZ | recombinant fusion protein comprising of SbpA and two copies of the Fc-binding Z-domain (a synthetic analogue of the IgG-binding domain of protein A from Staphylococcus aureus) |
| SAM | self-assembled monolayer |
| SAW | surface acoustic waves |
| SbpA | S-layer protein of Lysinibacillus sphaericus CCM 2177 |
| SbsB | S-layer protein of Geobacillus stearothermophilus PV72/p2 |
| S-layer | two dimensional arrays of proteinaceous subunits forming surface layers on prokaryotic cells |
| SPR | surface plasmon resonance |
| SsLM | S-layer supported lipid membrane |
| SUM | S-layer ultrafiltration membrane |
| TEM | transmission electron microscopy |
| TIRFM | total internal reflection fluorescence microscopy |
| VDAC | voltage-dependent anion channel |
References
- Thévenot, D.R.; Toth, K.; Durst, R.A.; Wilson, G.S. Electrochemical biosensors: Recommended definitions and classification. Biosens. Bioelectron. 2001, 16, 121–131. [Google Scholar] [CrossRef]
- Dugas, V.; Elaissari, A.; Chevalier, Y. Surface sensitization techniques and recognition receptors immobilization on biosensors and microarrays. In Recognition Receptors in Biosensors; Springer: New York, NY, USA, 2010; pp. 47–134. [Google Scholar]
- Turner, A.P.F. Biosensors: Fundamentals and applications—Historic book now open access. Biosens. Bioelectron. 2015, 65. [Google Scholar] [CrossRef] [PubMed]
- Cavalcanti, A.; Shirinzadeh, B.; Zhang, M.; Kretly, L.C. Nanorobot hardware architecture for medical defense. Sensors 2008, 8, 2932–2958. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, J.J.; Montemagno, C.D. Bionanomechanical systems. Annu. Rev. Mater. Res. 2004, 34, 315–337. [Google Scholar] [CrossRef]
- Sackmann, E. Supported membranes: Scientific and practical applications. Science 1996, 271, 43–48. [Google Scholar] [CrossRef] [PubMed]
- Rädler, J.; Sackmann, E. Functionalization of solids by ultrathin soft polymer films and polymer/lipid film composites: Modeling of cell surfaces and cell recognition processes. Curr. Opin. Solid State Mater. Sci. 1997, 2, 330–336. [Google Scholar] [CrossRef]
- Dubey, M.; Weidner, T.; Gamble, L.J.; Castner, D.G. Structure and order of phosphonic acid-based self-assembled monolayers on Si(100). Langmuir 2010, 26, 14747–14754. [Google Scholar] [CrossRef] [PubMed]
- Lessel, M.; Bäumchen, O.; Klos, M.; Hähl, H.; Fetzer, R.; Paulus, M.; Seemann, R.; Jacobs, K. Self-assembled silane monolayers: An efficient step-by-step recipe for high-quality, low energy surfaces. Surf. Interface Anal. 2015, 47, 557–564. [Google Scholar] [CrossRef]
- Lvov, Y.; Decher, G.; Möhwald, H. Assembly, structural characterization, and thermal behavior of layer-by-layer deposited ultrathin films of poly(vinyl sulfate) and poly(allylamine). Langmuir 1993, 9, 481–486. [Google Scholar] [CrossRef]
- Vericat, C.; Vela, M.E.; Benitez, G.; Carro, P.; Salvarezza, R.C. Self-assembled monolayers of thiols and dithiols on gold: New challenges for a well-known system. Chem. Soc. Rev. 2010, 39, 1805–1834. [Google Scholar] [CrossRef] [PubMed]
- Sleytr, U.B.; Bayley, H.; Sara, M.; Breitwieser, A.; Küpcü, S.; Mader, C.; Weigert, S.; Unger, F.M.; Messner, P.; Jahn-Schmid, B.; et al. VI. Applications of S-layers. FEMS Microbiol. Rev. 1997, 20, 151–175. [Google Scholar] [CrossRef] [PubMed]
- Sleytr, U.B.; Egelseer, E.M.; Ilk, N.; Messner, P.; Schäffer, C.; Pum, D.; Schuster, B. Prokaryotic cell wall components: Structure and biochemistry nanobiotechnological applications of S-layers. In Prokaryotic Cell Wall Compounds: Structure and Biochemistry; Springer: Berlin/Heidelberg, Germany, 2010; pp. 459–481. [Google Scholar]
- Sleytr, U.B.; Egelseer, E.M.; Ilk, N.; Pum, D.; Schuster, B. S-Layers as a basic building block in a molecular construction kit. FEBS J. 2007, 274, 323–334. [Google Scholar] [CrossRef] [PubMed]
- Sleytr, U.B.; Schuster, B.; Egelseer, E.M.; Pum, D. S-layers: Principles and applications. FEMS Microbiol. Rev. 2014, 38, 823–864. [Google Scholar] [CrossRef] [PubMed]
- Sleytr, U.B.; Schuster, B.; Egelseer, E.M.; Pum, D.; Horejs, C.M.; Tscheliessnig, R.; Ilk, N. Nanobiotechnology with S-layer proteins as building blocks. Prog. Mol. Biol. Transl. Sci. 2011, 103, 277–352. [Google Scholar] [PubMed]
- Heibel, C.; Ruehe, J.; Knoll, W. Tethered Membranes on Solid Supports; American Chemical Society, Polymer Preprints, Division of Polymer Chemistry: Blackburg, VA, USA, 1997; pp. 956–957. [Google Scholar]
- Knoll, W.; Naumann, R.; Friedrich, M.; Robertson, J.W.F.; Lösche, M.; Heinrich, F.; McGillivray, D.J.; Schuster, B.; Gufler, P.C.; Pum, D.; et al. Solid supported lipid membranes: New concepts for the biomimetic functionalization of solid surfaces. Biointerphases 2008, 3, FA125–FA135. [Google Scholar] [CrossRef] [PubMed]
- Sinner, E.K.; Ritz, S.; Naumann, R.; Schiller, S.; Knoll, W. Self-assembled tethered bimolecular lipid membranes. Adv. Clin. Chem. 2009, 49, 159–179. [Google Scholar] [PubMed]
- Knoll, W.; Köper, I.; Naumann, R.; Sinner, E.K. Tethered bimolecular lipid membranes-A novel model membrane platform. Electrochim. Acta 2008, 53, 6680–6689. [Google Scholar] [CrossRef]
- Sinner, E.K.; Knoll, W. Functional tethered membranes. Curr. Opin. Chem. Biol. 2001, 5, 705–711. [Google Scholar] [CrossRef]
- Jackman, J.A.; Knoll, W.; Cho, N.J. Biotechnology applications of tethered lipid bilayer membranes. Materials 2012, 5, 2637–2657. [Google Scholar] [CrossRef]
- Schiller, S.M.; Naumann, R.; Lovejoy, K.; Kunz, H.; Knoll, W. Archaea analogue thiolipids for tethered bilayer lipid membranes on ultrasmooth gold surfaces. Angew. Chem. Int. Ed. 2003, 42, 208–211. [Google Scholar] [CrossRef] [PubMed]
- Schiller, S.M.; Reisinger-Friebis, A.; Götz, H.; Hawker, C.J.; Frank, C.W.; Naumann, R.; Knoll, W. Biomimetic lipoglycopolymer membranes: Photochemical surface attachment of supramolecular architectures with defined orientation. Angew. Chem. Int. Ed. 2009, 48, 6896–6899. [Google Scholar] [CrossRef] [PubMed]
- Grieshaber, D.; MacKenzie, R.; Vörös, J.; Reimhult, E. Electrochemical biosensors—Sensor principles and architectures. Sensors 2008, 8, 1400–1458. [Google Scholar] [CrossRef] [PubMed]
- Cooper, C.L.; Dubin, P.L.; Kayitmazer, A.B.; Turksen, S. Polyelectrolyte-protein complexes. Curr. Opin. Colloid Interface Sci. 2005, 10, 52–78. [Google Scholar] [CrossRef]
- Halai, R.; Cooper, M. Label-free technologies: Which technique to use and what to watch out for! Methods Pharmacol. Toxicol. 2015, 53, 3–15. [Google Scholar]
- Hoa, X.D.; Kirk, A.G.; Tabrizian, M. Towards integrated and sensitive surface plasmon resonance biosensors: A review of recent progress. Biosens. Bioelectron. 2007, 23, 151–160. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Tang, W.; Wang, D.; Wu, X.; Li, N.; Liu, F. Amplified QCM-D biosensor for protein based on aptamer-functionalized gold nanoparticles. Biosens. Bioelectron. 2010, 26, 575–579. [Google Scholar] [CrossRef] [PubMed]
- Poitras, C.; Tufenkji, N. A QCM-D-based biosensor for E. coli O157:H7 highlighting the relevance of the dissipation slope as a transduction signal. Biosens. Bioelectron. 2009, 24, 2137–2142. [Google Scholar] [CrossRef] [PubMed]
- Tang, W.; Wang, D.; Xu, Y.; Li, N.; Liu, F. A self-assembled DNA nanostructure-amplified quartz crystal microbalance with dissipation biosensing platform for nucleic acids. Chem. Commun. 2012, 48, 6678–6680. [Google Scholar] [CrossRef] [PubMed]
- Yakovleva, M.E.; Safina, G.R.; Danielsson, B. A study of glycoprotein-lectin interactions using quartz crystal microbalance. Anal. Chim. Acta 2010, 668, 80–85. [Google Scholar] [CrossRef] [PubMed]
- Chang, B.Y.; Park, S.M. Electrochemical impedance spectroscopy. Annu. Rev. Anal. Chem. 2010, 3, 207–229. [Google Scholar] [CrossRef] [PubMed]
- Daniels, J.S.; Pourmand, N. Label-free impedance biosensors: Opportunities and challenges. Electroanalysis 2007, 19, 1239–1257. [Google Scholar] [CrossRef] [PubMed]
- Kang, X.; Wang, J.; Wu, H.; Aksay, I.A.; Liu, J.; Lin, Y. Glucose Oxidase-graphene-chitosan modified electrode for direct electrochemistry and glucose sensing. Biosens. Bioelectron. 2009, 25, 901–905. [Google Scholar] [CrossRef] [PubMed]
- Katz, E.; Willner, I. Probing Biomolecular Interactions at Conductive and Semiconductive Surfaces by Impedance Spectroscopy: Routes to Impedimetric Immunosensors, DNA-Sensors, and Enzyme Biosensors. Electroanalysis 2003, 15, 913–947. [Google Scholar] [CrossRef]
- Damiati, S.; Küpcü, S.; Peacock, M.; Eilenberger, C.; Zamzami, M.; Qadri, I.; Choudhry, H.; Sleytr, U.B.; Schuster, B. Acoustic and hybrid 3D-printed electrochemical biosensors for the real-time immunodetection of liver cancer cells (HepG2). Biosens. Bioelectron. 2017, 94, 500–506. [Google Scholar] [CrossRef] [PubMed]
- Damiati, S.; Peacock, M.; Leonhardt, S.; Damiati, L.; Baghdadi, M.; Becker, H.; Kodzius, R.; Schuster, B. Embedded Disposable Functionalized Electrochemical Biosensor with a 3D-Printed Flow Cell for Detection of Hepatic Oval Cells (HOCs). Genes 2018, 9, 89. [Google Scholar] [CrossRef] [PubMed]
- Damiati, S.; Peacock, M.; Sleytr, U.B.; Schuster, B. Bioinspired Diagnostic Sensor Based on Functional Nanostructures of S-Proteins to Target the Folate Receptors in Breast Cancer Cells. Sensor. Actuators B Chem. 2018, in press. [Google Scholar] [CrossRef]
- Sapsford, K.E.; Ligler, F.S. Real-time analysis of protein adsorption to a variety of thin films. Biosens. Bioelectron. 2004, 19, 1045–1055. [Google Scholar] [CrossRef] [PubMed]
- Schuster, B.; Sleytr, U.B. Biomimetic interfaces based on S-layer proteins, lipid membranes and functional biomolecules. J. R. Soc. Interface 2014, 11, 20140232. [Google Scholar] [CrossRef] [PubMed]
- Albers, S.V.; Meyer, B.H. The archaeal cell envelope. Nat. Rev. Microbiol. 2011, 9, 414–426. [Google Scholar] [CrossRef] [PubMed]
- Sleytr, U.B.; Sára, M. Bacterial and archaeal S-layer proteins: Structure-function relationships and their biotechnological applications. Trends Biotechnol. 1997, 15, 20–26. [Google Scholar] [CrossRef]
- Rodrigues-Oliveira, T.; Belmok, A.; Vasconcellos, D.; Schuster, B.; Kyaw, C.M. Archaeal S-Layers: Overview and Current State of the Art. Front. Microbiol. 2017, 8, 2597. [Google Scholar] [CrossRef] [PubMed]
- Messner, P.; Schäffer, C.; Egelseer, E.M.; Sleytr, U.B. Occurrence, Structure, Chemistry, Genetics, Morphogenesis, and Function of S-Layers. In Prokaryotic Cell Wall Compounds—Structure and Biochemistry; König, H., Claus, H., Varma, A., Eds.; Springer: Heidelberg, Germany, 2010; pp. 53–109. [Google Scholar]
- Whitman, W.B.; Coleman, D.C.; Wiebe, W.J. Prokaryotes: The unseen majority. Proc. Natl. Acad. Sci. USA 1998, 95, 6578–6583. [Google Scholar] [CrossRef] [PubMed]
- Sleytr, U.B. Heterologous reattachment of regular arrays of glycoproteins on bacterial surfaces. Nature 1975, 257, 400–402. [Google Scholar] [CrossRef] [PubMed]
- Sleytr, U.B.; Huber, C.; Ilk, N.; Pum, D.; Schuster, B.; Egelseer, E.M. S-Layers as a tool kit for nanobiotechnological applications. FEMS Microbiol. Lett. 2007, 267, 131–144. [Google Scholar] [CrossRef] [PubMed]
- Sleytr, U.B. Regular arrays of macromolecules on bacterial cell walls: Structure, chemistry, assembly, and function. Int. Rev. Cytol. 1978, 53, 1–64. [Google Scholar] [PubMed]
- Liew, P.W.Y.; Jong, B.C.; Najimudin, N. Hypothetical protein Avin_16040 as the S-layer protein of Azotobacter vinelandii and its involvement in plant root surface attachment. Appl. Environ. Microbiol. 2015, 81, 7484–7495. [Google Scholar] [CrossRef] [PubMed]
- Sleytr, U.B.; Messner, P. Self-assembly of crystalline bacterial cell surface layers (S-layers). In Electron Microscopy of Subcellular Dynamics; Plattner, H., Ed.; CRC Press: Boca Raton, FL, USA, 1989; pp. 13–31. [Google Scholar]
- De Yoreo, J.J.; Chung, S.; Friddle, R.W. In situ atomic force microscopy as a tool for investigating interactions and assembly dynamics in biomolecular and biomineral systems. Adv. Funct. Mater. 2013, 23, 2525–2538. [Google Scholar] [CrossRef]
- Sleytr, U.B. Curiosity and Passion for Science and Art; World Scientific: Singapore, 2016. [Google Scholar]
- Pavkov-Keller, T.; Howorka, S.; Keller, W. The structure of bacterial S-layer proteins. In Molecular Assembly in Natural and Engineered Systems; Howorka, S., Ed.; Elsevier Academic Press Inc.: Burlington, NJ, USA, 2011; Volume 103, pp. 73–130. [Google Scholar]
- Sleytr, U.B.; Beveridge, T.J. Bacterial S-layers. Trends Microbiol. 1999, 7, 253–260. [Google Scholar] [CrossRef]
- Picher, M.M.; Küpcü, S.; Huang, C.J.; Dostalek, J.; Pum, D.; Sleytr, U.B.; Ertl, P. Nanobiotechnology advanced antifouling surfaces for the continuous electrochemical monitoring of glucose in whole blood using a lab-on-a-chip. Lab Chip 2013, 13, 1780–1789. [Google Scholar] [CrossRef] [PubMed]
- Sára, M.; Pum, D.; Sleytr, U.B. Permeability and charge-dependent adsorption properties of the S-layer lattice from Bacillus coagulans E38-66. J. Bacteriol. 1992, 174, 3487–3493. [Google Scholar] [CrossRef] [PubMed]
- Sára, M.; Sleytr, U.B. S-Layer Proteins. J. Bacteriol. 2000, 182, 859–868. [Google Scholar] [CrossRef] [PubMed]
- Sleytr, U.B.; Messner, P. Crystalline bacterial cell surface layers (S-layers). In Encyclopedia of Microbiology, 3rd ed.; Schaechter, M., Ed.; Academic Press/Elsevier Science: San Diego, CA, USA, 2009; Volume 1, pp. 89–98. [Google Scholar]
- Sleytr, U.B.; Messner, P.; Pum, D.; Sára, M. Crystalline bacterial cell surface layers (S-layers): From supramolecular cell structure to biomimetics and nanotechnology. Angew. Chem. Int. Ed. 1999, 38, 1034–1054. [Google Scholar] [CrossRef]
- Sleytr, U.B.; Sára, M.; Pum, D.; Schuster, B.; Messner, P.; Schäffer, C. Self-assembly protein systems: Microbial S-layers. In Biopolymers. Polyamides and Complex Proteinaceous Materials I; Steinbüchel, A., Fahnestock, S.R., Eds.; Wiley-VCH: Weinheim, Germany, 2002; pp. 285–338. [Google Scholar]
- Fagan, R.P.; Fairweather, N.F. Biogenesis and functions of bacterial S-layers. Nat. Rev. Microbiol. 2014, 12, 211–222. [Google Scholar] [CrossRef] [PubMed]
- Sleytr, U.B.; Egelseer, E.M.; Pum, D.; Schuster, B. S-Layers. In NanoBiotechnology: Concepts, Methods and Perspectives; Niemeyer, C.M., Mirkin, C.A., Eds.; Wiley-VCH: Weinheim, Germany, 2004; pp. 77–92. [Google Scholar]
- Pum, D.; Messner, P.; Sleytr, U.B. Role of the S layer in morphogenesis and cell division of the archaebacterium Methanocorpusculum sinense. J. Bacteriol. 1991, 173, 6865–6873. [Google Scholar] [CrossRef] [PubMed]
- Engelhardt, H.; Peters, J. Structural Research on Surface Layers: A Focus on Stability, Surface Layer Homology Domains, and Surface Layer–Cell Wall Interactions. J. Struct. Biol. 1998, 124, 276–302. [Google Scholar] [CrossRef] [PubMed]
- Sotiropoulou, S.; Mark, S.S.; Angert, E.R.; Batt, C.A. Nanoporous S-layer protein lattices. A biological ion gate with calcium selectivity. J. Phys. Chem. C 2007, 111, 13232–13237. [Google Scholar] [CrossRef]
- Sleytr, U.B.; Györvary, E.; Pum, D. Crystallization of S-layer protein lattices on surfaces and interfaces. Prog. Organ. Coat. 2003, 47, 279–287. [Google Scholar] [CrossRef]
- Pum, D.; Sára, M.; Sleytr, U.B. Structure, surface charge, and self-assembly of the S-layer lattice from Bacillus coagulans E38-66. J. Bacteriol. 1989, 171, 5296–5303. [Google Scholar] [CrossRef] [PubMed]
- Pum, D.; Sleytr, U.B. Large-scale reconstruction of crystalline bacterial surface layer proteins at the air-water interface and on lipids. Thin Solid Films 1994, 244, 882–886. [Google Scholar] [CrossRef]
- Pum, D.; Weinhandl, M.; Hödl, C.; Sleytr, U.B. Large-scale recrystallization of the S-layer of Bacillus coagulans E38-66 at the air/water interface and on lipid films. J. Bacteriol. 1993, 175, 2762–2766. [Google Scholar] [CrossRef] [PubMed]
- Györvary, E.S.; Stein, O.; Pum, D.; Sleytr, U.B. Self-assembly and recrystallization of bacterial S-layer proteins at silicon supports imaged in real time by atomic force microscopy. J. Microsc. 2003, 212 Pt 3, 300–306. [Google Scholar] [CrossRef] [PubMed]
- Tang, J.; Ebner, A.; Ilk, N.; Lichtblau, H.; Huber, C.; Zhu, R.; Pum, D.; Leitner, M.; Pastushenko, V.; Gruber, H.J.; et al. High-affinity tags fused to S-layer proteins probed by atomic force microscopy. Langmuir 2008, 24, 1324–1329. [Google Scholar] [CrossRef] [PubMed]
- Tang, J.; Ebner, A.; Badelt-Lichtblau, H.; Völlenkle, C.; Rankl, C.; Kraxberger, B.; Leitner, M.; Wildling, L.; Gruber, H.J.; Sleytr, U.B.; et al. Recognition imaging and highly ordered molecular templating of bacterial S-layer nanoarrays containing affinity-tags. Nano Lett. 2008, 8, 4312–4319. [Google Scholar] [CrossRef] [PubMed]
- Pum, D.; Sleytr, U.B. Anisotropic crystal growth of the S-layer of Bacillus sphaericus CCM 2177 at the air/water interface. Colloid Surf. A 1995, 102, 99–104. [Google Scholar] [CrossRef]
- Pum, D.; Sleytr, U.B. Monomolecular reassembly of a crystalline bacterial cell surface layer (S-layer) on untreated and modified silicon surfaces. Supramol. Sci. 1995, 2, 193–197. [Google Scholar] [CrossRef]
- Sleytr, U.B.; Sára, M.; Pum, D.; Schuster, B. Crystalline bacterial cell surface layers (S-layers): A versatile self-assembly system. In Supramolecular Polymers, 2nd ed.; Ciferri, A., Ed.; Taylor & Francis Group: Boca Raton, FL, USA, 2005; pp. 583–612. [Google Scholar]
- Rothbauer, M.; Küpcü, S.; Sticker, D.; Sleytr, U.B.; Ertl, P. Exploitation of S-layer anisotropy: PH-dependent nanolayer orientation for cellular micropatterning. ACS Nano 2013, 7, 8020–8030. [Google Scholar] [CrossRef] [PubMed]
- Weygand, M.; Kjaer, K.; Howes, P.B.; Wetzer, B.; Pum, D.; Sleytr, U.B.; Lösche, M. Structural reorganization of phospholipid headgroups upon recrystallization of an S-layer lattice. J. Phys. Chem. B 2002, 106, 5793–5799. [Google Scholar] [CrossRef]
- Weygand, M.; Schalke, M.; Howes, P.B.; Kjaer, K.; Friedmann, J.; Wetzer, B.; Pum, D.; Sleytr, U.B.; Lösche, M. Coupling of protein sheet crystals (S-layers) to phospholipid monolayers. J. Mater. Chem. 2000, 10, 141–148. [Google Scholar] [CrossRef]
- Weygand, M.; Wetzer, B.; Pum, D.; Sleytr, U.B.; Cuvillier, N.; Kjaer, K.; Howes, P.B.; Lösche, M. Bacterial S-layer protein coupling to lipids: X-ray reflectivity and grazing incidence diffraction studies. Biophys. J. 1999, 76, 458–468. [Google Scholar] [CrossRef]
- Comolli, L.R. Conformational Transitions Driving Lattice Growth at an S-Layer Boundary Resolved by cryo-TEM. Biophys. J. 2013, 104, 353A–353A. [Google Scholar] [CrossRef]
- Comolli, L.R.; Siegerist, C.E.; Shin, S.H.; Bertozzi, C.; Regan, W.; Zettl, A.; De Yoreo, J. Conformational Transitions at an S-Layer Growing Boundary Resolved by Cryo-TEM. Angew. Chem. Int. Ed. 2013, 52, 4829–4832. [Google Scholar] [CrossRef] [PubMed]
- Sleytr, U.B.; Messner, P.; Pum, D.; Sára, M. Kristalline Zelloberflächen-Schichten prokaryotischer Organismen (S Schichten): Von der supramolekularen Zellstruktur zur Biomimetik und Nanotechnologie. Angew. Chem. 1999, 111, 1098–1120. [Google Scholar] [CrossRef]
- Schuster, B.; Sleytr, U.B. Nanotechnology with S-layer proteins. In Methods in Molecular Biology; Gerrard, J.A., Gerrard, J.A., Eds.; Humana Press: Totowa, NJ, USA, 2013; Volume 996, pp. 153–175. [Google Scholar]
- Györvary, E.S.; O’Riordan, A.; Quinn, A.J.; Redmond, G.; Pum, D.; Sleytr, U.B. Biomimetic nanostructure fabrication: Nonlithographic lateral patterning and self-assembly of functional bacterial S-layers at silicon supports. Nano Lett. 2003, 3, 315–319. [Google Scholar] [CrossRef]
- Schuster, B.; Sleytr, U.B. The effect of hydrostatic pressure on S layer-supported lipid membranes. Biochim. Biophys. Acta 2002, 1563, 29–34. [Google Scholar] [CrossRef]
- Schuster, B.; Sleytr, U.B. Biomimetic S-layer supported lipid membranes. Curr. Nanosci. 2006, 2, 143–152. [Google Scholar] [CrossRef]
- Ilk, N.; Egelseer, E.M.; Sleytr, U.B. S-layer fusion proteins-construction principles and applications. Curr. Opin. Biotechnol. 2011, 22, 824–831. [Google Scholar] [CrossRef] [PubMed]
- Moll, D.; Huber, C.; Schlegel, B.; Pum, D.; Sleytr, U.B.; Sára, M. S-layer-streptavidin fusion proteins as template for nanopatterned molecular arrays. Proc. Natl. Acad. Sci. USA 2002, 99, 14646–14651. [Google Scholar] [CrossRef] [PubMed]
- Ilk, N.; Schumi, C.T.; Bohle, B.; Egelseer, E.M.; Sleytr, U.B. Expression of an endotoxin-free S-layer/allergen fusion protein in gram-positive Bacillus subtilis 1012 for the potential application as vaccines for immunotherapy of atopic allergy. Microb. Cell Fact. 2011, 10, 6. [Google Scholar] [CrossRef] [PubMed]
- Ilk, N.; Küpcü, S.; Moncayo, G.; Klimt, S.; Ecker, R.C.; Hofer-Warbinek, R.; Egelseer, E.M.; Sleytr, U.B.; Sára, M. A functional chimaeric S-layer-enhanced green fluorescent protein to follow the uptake of S-layer-coated liposomes into eukaryotic cells. Biochem. J. 2004, 379, 441–448. [Google Scholar] [CrossRef] [PubMed]
- Sára, M.; Egelseer, E.M.; Huber, C.; Ilk, N.; Pleschberger, M.; Pum, D.; Sleytr, U.B. S-layer proteins: Potential application in nano(bio)technology. In Microbial Bionanotechnology: Biological Self-Assembly Systems and Biopolymer-Based Nanostructures; Rehm, B., Ed.; Horizon Bioscience: Auckland, New Zealand, 2006; pp. 307–338. [Google Scholar]
- Schuster, B. Biomimetic design of nano-patterned membranes. Nanobiotechnology 2005, 1, 153–164. [Google Scholar] [CrossRef]
- Schuster, B.; Kepplinger, C.; Sleytr, U.B. Biomimetic S-layer stabilized lipid membranes. In Biomimetics in Biophysics: Model Systems, Experimental Techniques and Computation; Toca-Herrera, J.L., Ed.; Research Signpost: Kerala, India, 2010; pp. 1–12. [Google Scholar]
- Schuster, B.; Pum, D.; Sleytr, U.B. S-layer stabilized lipid membranes. Biointerphases 2008, 3, FA3–FA11. [Google Scholar] [CrossRef] [PubMed]
- Schuster, B.; Sleytr, U.B. 2D-Protein Crystals (S-Layers) as Support for Lipid Membranes. In Advances in Planar Lipid Bilayers and Liposomes; Tien, T.H., Ottova, A., Eds.; Elsevier Science: Amsterdam, The Netherlands, 2005; Volume 1, pp. 247–293. [Google Scholar]
- Schuster, B.; Sleytr, U.B. Composite S-layer lipid structures. J. Struct. Biol. 2009, 168, 207–216. [Google Scholar] [CrossRef] [PubMed]
- Schuster, B.; Pum, D.; Sàra, M.; Sleytr, U.B. S-Layer Proteins as Key Components of a Versatile Molecular Construction Kit for Biomedical Nanotechnology. Mini-Rev. Med. Chem. 2006, 6, 909–920. [Google Scholar] [CrossRef] [PubMed]
- Sleytr, U.B.; Pum, D.; Sára, M.; Schuster, B. Molecular nanotechnology and nanobiotechnology with 2-D protein crystals. In Encylopedia of Nanoscience and Nanotechnology; Nalwa, H.S., Ed.; Academic Press: San Diego, CA, USA, 2004; Volume 5, pp. 693–702. [Google Scholar]
- Sleytr, U.B.; Schuster, B.; Pum, D. Nanotechnology and biomimetics with 2D protein crystals (S layers). IEEE Eng. Med. Biol. 2003, 22, 140–150. [Google Scholar] [CrossRef]
- Clark, L.C., Jr.; Lyons, C. Electrode systems for continuous monitoring in cardiovascular surgery. Ann. N. Y. Acad. Sci. 1962, 102, 29–45. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Zhao, X.L.; Li, Z.H.; Zhu, Z.G.; Qian, S.H.; Flewitt, A.J. Current and emerging technology for continuous glucose monitoring. Sensors 2017, 17, 182. [Google Scholar] [CrossRef] [PubMed]
- Justino, C.I.L.; Duarte, A.C.; Rocha-Santos, T.A.P. Critical overview on the application of sensors and biosensors for clinical analysis. Trends Anal. Chem. 2016, 85, 36–60. [Google Scholar] [CrossRef]
- Mehrotra, P. Biosensors and their applications—A review. J. Oral Biol. Craniofac. Res. 2016, 6, 153–159. [Google Scholar] [CrossRef] [PubMed]
- Shende, P.; Sahu, P.; Gaud, R. A technology roadmap of smart biosensors from conventional glucose monitoring systems. Ther. Deliv. 2017, 8, 411–423. [Google Scholar] [CrossRef] [PubMed]
- Hasanzadeh, M.; Shadjou, N. Electrochemical nanobiosensing in whole blood: Recent advances. Trends Anal. Chem. 2016, 80, 167–176. [Google Scholar] [CrossRef]
- Soleymani, J.; Perez-Guaita, D.; Hasanzadeh, M.; Shadjou, N.; Jouyban, A. Materials and methods of signal enhancement for spectroscopic whole blood analysis: Novel research overview. Trends Anal. Chem. 2017, 86, 122–142. [Google Scholar] [CrossRef]
- Schuster, B.; Gufler, P.C.; Pum, D.; Sleytr, U.B. Interplay of phospholipase A2 with S-layer-supported lipid monolayers. Langmuir 2003, 19, 3393–3397. [Google Scholar] [CrossRef]
- Sleytr, U.B.; Sára, M. Ultrafiltration membranes with uniform pores from crystalline bacterial cell envelope layers. Appl. Microbiol. Biotechnol. 1986, 25, 83–90. [Google Scholar] [CrossRef]
- Sára, M.; Küpcü, S.; Weiner, C.; Weigert, S.; Sleytr, U.B. Crystalline protein layers as isoporous molecular sieves and immobilisation and affinity matrices. In Immobilised Macromolecules: Application Potentials; Sleytr, U.B., Messner, P., Pum, D., Sára, M., Eds.; Springer-Verlag: London, UK, 1993; pp. 71–86. [Google Scholar]
- Sára, M.; Manigley, C.; Wolf, G.; Sleytr, U.B. Isoporous ultrafiltration membranes from bacterial cell envelope layers. J. Membr. Sci. 1988, 36, 179–186. [Google Scholar] [CrossRef]
- Sára, M.; Sleytr, U.B. Production and characteristics of ultrafiltration membranes with uniform pores from two-dimensional arrays of proteins. J. Membr. Sci. 1987, 33, 27–49. [Google Scholar] [CrossRef]
- Park, T.J.; Lee, S.J.; Park, J.P.; Yang, M.H.; Choi, J.H.; Lee, S.Y. Characterization of a bacterial self-assembly surface layer protein and its application as an electrical nanobiosensor. J. Nanosci. Nanotechnol. 2011, 11, 402–407. [Google Scholar] [CrossRef] [PubMed]
- Pum, D.; Neubauer, A.; Sleytr, U.B.; Pentzien, S.; Reetz, S.; Kautek, W. Physico-chemical properties of crystalline nanoscale enzyme-protein-metal layer composites in biosensors. Ber. Bunsenges. Phys. Chem. 1997, 101, 1686–1689. [Google Scholar] [CrossRef]
- Pum, D.; Sára, M.; Sleytr, U.B. Two-dimensional (glyco)protein crystals as patterning elements and immobilisation matrices for the development of biosensors. In Immobilised Macromolecules: Application Potentials; Sleytr, U.B., Messner, P., Pum, D., Sára, M., Eds.; Springer-Verlag: London, UK, 1993; pp. 141–160. [Google Scholar]
- Sára, M.; Küpcü, S.; Weiner, C.; Weigert, S.; Sleytr, U.B. S-layers as immobilization and affinity matrices. In Advances in Bacterial Paracrystalline Surface Layers; Beveridge, T.J., Koval, S.F., Eds.; Plenum Press: New York, NY, USA; London, UK, 1993; Volume 252. [Google Scholar]
- Neubauer, A.; Pum, D.; Sleytr, U.B. An amperometric glucose sensor based on isoporous crystalline protein membranes as immobilization matrix. Anal. Lett. 1993, 26, 1347–1360. [Google Scholar] [CrossRef]
- Neubauer, A.; Hödl, C.; Pum, D.; Sleytr, U.B. A multistep enzyme sensor for sucrose based on S-layer microparticles as immobilization matrix. Anal. Lett. 1994, 27, 849–865. [Google Scholar] [CrossRef]
- Neubauer, A.; Pum, D.; Sleytr, U.B. Fibre-optic glucose biosensor using enzyme membranes with 2-D crystalline structure. Biosens. Bioelectron. 1996, 11, 317–325. [Google Scholar] [CrossRef]
- Neubauer, A.; Pentzien, S.; Reetz, S.; Kautek, W.; Pum, D.; Sleytr, U.B. Pulsed-laser metal contacting of biosensors on the basis of crystalline enzyme-protein layer composites. Sens. Actuator B Chem. 1997, 40, 231–236. [Google Scholar] [CrossRef]
- Schuster, B.; Sleytr, U.B. Relevance of glycosylation of S-layer proteins for cell surface properties. Acta Biomater. 2015, 19, 149–157. [Google Scholar] [CrossRef] [PubMed]
- Ferraz, H.C.; Guimarães, J.A.; Alves, T.L.M.; Constantino, C.J.L. Monomolecular films of cholesterol oxidase and S-Layer proteins. Appl. Surf. Sci. 2011, 257, 6535–6539. [Google Scholar] [CrossRef]
- Guimarães, J.A.; Ferraz, H.C.; Alves, T.L.M. Langmuir-Blodgett films of cholesterol oxidase and S-layer proteins onto screen-printed electrodes. Appl. Surf. Sci. 2014, 298, 68–74. [Google Scholar] [CrossRef]
- Scheicher, S.R.; Kainz, B.; Köstler, S.; Suppan, M.; Bizzarri, A.; Pum, D.; Sleytr, U.B.; Ribitsch, V. Optical oxygen sensors based on Pt(II) porphyrin dye immobilized on S-layer protein matrices. Biosens. Bioelectron. 2009, 25, 797–802. [Google Scholar] [CrossRef] [PubMed]
- Weinert, U.; Vogel, M.; Reinemann, C.; Strehlitz, B.; Pollmann, K.; Raff, J. S-layer proteins as an immobilization matrix for aptamers on different sensor surfaces. Eng. Life Sci. 2015, 15, 710–720. [Google Scholar] [CrossRef]
- Conroy, D.J.R.; Millner, P.A.; Stewart, D.I.; Pollmann, K. Biosensing for the environment and defence: Aqueous uranyl detection using bacterial surface layer proteins. Sensors 2010, 10, 4739–4755. [Google Scholar] [CrossRef] [PubMed]
- Lakatos, M.; Matys, S.; Raff, J.; Pompe, W.; Huber, C. Colorimetric As(V) detection based on S-layer functionalized gold nanoparticles. Talanta 2015, 144, 241–246. [Google Scholar] [CrossRef] [PubMed]
- Huber, C.; Liu, J.; Egelseer, E.M.; Moll, D.; Knoll, W.; Sleytr, U.B.; Sára, M. Heterotetramers formed by an S-layer-streptavidin fusion protein and core-streptavidin as nanoarrayed template for biochip development. Small 2006, 2, 142–150. [Google Scholar] [CrossRef] [PubMed]
- Pleschberger, M.; Neubauer, A.; Egelseer, E.M.; Weigert, S.; Lindner, B.; Sleytr, U.B.; Muyldermans, S.; Sára, M. Generation of a functional monomolecular protein lattice consisting of an S-layer fusion protein comprising the variable domain of a camel heavy chain antibody. Bioconjug. Chem. 2003, 14, 440–448. [Google Scholar] [CrossRef] [PubMed]
- Pleschberger, M.; Saerens, D.; Weigert, S.; Sleytr, U.B.; Muyldermans, S.; Sára, M.; Egelseer, E.M. An S-layer heavy chain camel antibody fusion protein for generation of a nanopatterned sensing layer to detect the prostate-specific antigen by surface plasmon resonance technology. Bioconjug. Chem. 2004, 15, 664–671. [Google Scholar] [CrossRef] [PubMed]
- Weber, V.; Ilk, N.; Völlenkle, C.; Weigert, S.; Sára, M.; Sleytr, U.B.; Falkenhagen, D. Extrakorporale Blutreinigung mit spezifischen Absorbentien auf Basis der S-Schicht-Technologie. In Medizin 2002-Aus Forschung und Praxis; Dézsy, J., Ed.; Dr. Peter Müller Verlag: Wien, Austria, 2002. [Google Scholar]
- Völlenkle, C.; Weigert, S.; Ilk, N.; Egelseer, E.; Weber, V.; Loth, F.; Falkenhagen, D.; Sleytr, U.B.; Sára, M. Construction of a functional S-layer fusion protein comprising an immunoglobulin G-binding domain for development of specific adsorbents for extracorporeal blood purification. Appl. Environ. Microbiol. 2004, 70, 1514–1521. [Google Scholar] [CrossRef] [PubMed]
- Tschiggerl, H.; Breitwieser, A.; de Roo, G.; Verwoerd, T.; Schäffer, C.; Sleytr, U.B. Exploitation of the S-layer self-assembly system for site directed immobilization of enzymes demonstrated for an extremophilic laminarinase from Pyrococcus furiosus. J. Biotechnol. 2008, 133, 403–411. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Tschiggerl, H.; Casey, J.L.; Parisi, K.; Foley, M.; Sleytr, U.B. Display of a peptide mimotope on a crystalline bacterial cell surface layer (S-layer) lattice for diagnosis of Epstein-Barr virus infection. Bioconjug. Chem. 2008, 19, 860–865. [Google Scholar] [CrossRef] [PubMed]
- Ferner-Ortner-Bleckmann, J.; Schrems, A.; Ilk, N.; Egelseer, E.M.; Sleytr, U.B.; Schuster, B. Multitechnique study on a recombinantly produced Bacillus halodurans laccase and an S-layer/laccase fusion protein. Biointerphases 2011, 6, 63–72. [Google Scholar] [CrossRef] [PubMed]
- Sprott, G.D.; Yeung, A.; Dicaire, C.J.; Yu, S.H.; Whitfield, D.M. Synthetic archaeosome vaccines containing triglycosylarchaeols can provide additive and long-lasting immune responses that are enhanced by archaetidylserine. Archaea 2012, 2012, 513231. [Google Scholar] [CrossRef] [PubMed]
- De Rosa, M. Archaeal lipids: Structural features and supramolecular organization. Thin Solid Films 1996, 284–285, 13–17. [Google Scholar] [CrossRef]
- Hanford, M.J.; Peeples, T.L. Archaeal tetraether lipids: Unique structures and applications. Appl. Biochem. Biotechnol. 2002, 97, 45–62. [Google Scholar] [CrossRef]
- Engelhardt, H. Are S-layers exoskeletons? The basic function of protein surface layers revisited. J. Struct. Biol. 2007, 160, 115–124. [Google Scholar] [CrossRef] [PubMed]
- Engelhardt, H. Mechanism of osmoprotection by archaeal S-layers: A theoretical study. J. Struct. Biol. 2007, 160, 190–199. [Google Scholar] [CrossRef] [PubMed]
- Schuster, B.; Pum, D.; Sleytr, U.B. Voltage clamp studies on S-layer-supported tetraether lipid membranes. Biochim. Biophys. Acta 1998, 1369, 51–60. [Google Scholar] [CrossRef]
- Schuster, B.; Gufler, P.C.; Pum, D.; Sleytr, U.B. S-layer proteins as supporting scaffoldings for functional lipid membranes. IEEE Trans. Nanobiosci. 2004, 3, 16–21. [Google Scholar] [CrossRef]
- Schuster, B.; Sleytr, U.B. S-layer-supported lipid membranes. J. Biotechnol. 2000, 74, 233–254. [Google Scholar] [CrossRef]
- Schuster, B.; Sleytr, U.B. Single channel recordings of α-hemolysin reconstituted in S-layer-supported lipid bilayers. Bioelectrochemistry 2002, 55, 5–7. [Google Scholar] [CrossRef]
- Wetzer, B.; Pfandler, A.; Györvary, E.; Pum, D.; Lösche, M.; Sleytr, U.B. S-layer reconstitution at phospholipid monolayers. Langmuir 1998, 14, 6899–6906. [Google Scholar] [CrossRef]
- Schrems, A.; Kibrom, A.; Küpcü, S.; Kiene, E.; Sleytr, U.B.; Schuster, B. Bilayer lipid membrane formation on a chemically modified S-layer lattice. Langmuir 2011, 27, 3731–3738. [Google Scholar] [CrossRef] [PubMed]
- Schrems, A.; Larisch, V.D.; Stanetty, C.; Dutter, K.; Damiati, S.; Sleytr, U.B.; Schuster, B. Liposome fusion on proteinaceous S-layer lattices triggered via β-diketone ligand-europium(iii) complex formation. Soft Matter 2011, 7, 5514–5518. [Google Scholar] [CrossRef]
- Kepplinger, C.; Ilk, N.; Sleytr, U.B.; Schuster, B. Intact lipid vesicles reversibly tethered to a bacterial S-layer protein lattice. Soft Matter 2009, 5, 325–333. [Google Scholar] [CrossRef]
- Krivanek, R.; Rybar, P.; Küpcü, S.; Sleytr, U.B.; Hianik, T. Affinity interactions on a liposome surface detected by ultrasound velocimetry. Bioelectrochemistry 2002, 55, 57–59. [Google Scholar] [CrossRef]
- Mader, C.; Küpcü, S.; Sleytr, U.B.; Sára, M. S-layer-coated liposomes as a versatile system for entrapping and binding target molecules. Biochim. Biophys. Acta 2000, 1463, 142–150. [Google Scholar] [CrossRef]
- Badelt-Lichtblau, H.; Kainz, B.; Völlenkle, C.; Egelseer, E.M.; Sleytr, U.B.; Pum, D.; Ilk, N. Genetic engineering of the S-layer protein SbpA of Lysinibacillus sphaericus CCM 2177 for the generation of functionalized nanoarrays. Bioconjug. Chem. 2009, 20, 895–903. [Google Scholar] [CrossRef] [PubMed]
- Howorka, S.; Sára, M.; Wang, Y.; Kuen, B.; Sleytr, U.B.; Lubitz, W.; Bayley, H. Surface-accessible residues in the monomeric and assembled forms of a bacterial surface layer protein. J. Biol. Chem. 2000, 275, 37876–37886. [Google Scholar] [CrossRef] [PubMed]
- Tang, J.; Badelt-Lichtblau, H.; Ebner, A.; Preiner, J.; Kraxberger, B.; Gruber, H.J.; Sleytr, U.B.; Ilk, N.; Hinterdorfer, P. Fabrication of highly ordered gold nanoparticle arrays templated by crystalline lattices of bacterial S-layer protein. ChemPhysChem 2008, 9, 2317–2320. [Google Scholar] [CrossRef] [PubMed]
- Kiene, E. Potential and Limitations of S-Layer as Support for Planar Lipid Bilayers. Ph.D. Thesis, University of Natural Resources and Life Sciences, Vienna, Austria, 2011. [Google Scholar]
- Tang, J.; Ebner, A.; Kraxberger, B.; Badelt-Lichtblau, H.; Gruber, H.J.; Sleytr, U.B.; Ilk, N.; Hinterdorfer, P. Mapping short affinity tags on bacterial S-layer with an antibody. ChemPhysChem 2010, 11, 2323–2326. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.; Groves, J.T. Engineering supported membranes for cell biology. Med. Biol. Eng. Comput. 2010, 48, 955–963. [Google Scholar] [CrossRef] [PubMed]
- Györvary, E.; Wetzer, B.; Sleytr, U.B.; Sinner, A.; Offenhäusser, A.; Knoll, W. Lateral diffusion of lipids in silane-, dextran-, and S-layer-supported mono- and bilayers. Langmuir 1999, 15, 1337–1347. [Google Scholar] [CrossRef]
- Nikolelis, D.P.; Hianik, T.; Krull, U.J. Biosensors based on thin lipid films and liposomes. Electroanalysis 1999, 11, 7–15. [Google Scholar] [CrossRef]
- Gufler, P.C.; Pum, D.; Sleytr, U.B.; Schuster, B. Highly robust lipid membranes on crystalline S-layer supports investigated by electrochemical impedance spectroscopy. Biochim. Biophys. Acta 2004, 1661, 154–165. [Google Scholar] [CrossRef] [PubMed]
- Schuster, B.; Pum, D.; Sára, M.; Braha, O.; Bayley, H.; Sleytr, U.B. S-layer ultrafiltration membranes: A new support for stabilizing functionalized lipid membranes. Langmuir 2001, 17, 499–503. [Google Scholar] [CrossRef]
- Schuster, B.; Weigert, S.; Pum, D.; Sára, M.; Sleytr, U.B. New Method for Generating Tetraether Lipid Membranes on Porous Supports. Langmuir 2003, 19, 2392–2397. [Google Scholar] [CrossRef]
- Sára, M.; Sleytr, U.B. Membrane Biotechnology: Two-dimensional Protein Crystals for Ultrafiltration Purposes. In Biotechnology; Rehm, H.J., Ed.; VCH: Weinheim, Germany, 1988; Volume 6b, pp. 615–636. [Google Scholar]
- Küpcü, S.; Sára, M.; Sleytr, U.B. Influence of covalent attachment of low molecular weight substances on the rejection and adsorption properties of crystalline proteinaceous ultrafiltration membranes. Desalination 1993, 90, 65–76. [Google Scholar] [CrossRef]
- Weigert, S.; Sára, M. Ultrafiltration membranes prepared from crystalline bacterial cell surface layers as model systems for studying the influence of surface properties on protein adsorption. J. Membr. Sci. 1996, 121, 185–196. [Google Scholar] [CrossRef]
- Weigert, S.; Sára, M. Surface modification of an ultrafiltration membrane with crystalline structure and studies on interactions with selected protein molecules. J. Membr. Sci. 1995, 106, 147–159. [Google Scholar] [CrossRef]
- Lohner, K.; Prossnigg, F. Biological activity and structural aspects of PGLa interaction with membrane mimetic systems. Biochim. Biophys. Acta 2009, 1788, 1656–1666. [Google Scholar] [CrossRef] [PubMed]
- Schuster, B.; Sleytr, U.B. Nanotechnology with S-Layer Proteins. In Protein Nanotechnology: Protocols, Instrumentation and Applications, 2nd ed.; Gerrard, J.A., Ed.; Humana Press, Springer Science+Business Media: New York, NY, USA, 2013; Volume 996, pp. 153–175. [Google Scholar]
- Schrems, A.; Larisch, V.D.; Sleytr, U.B.; Hohenegger, M.; Lohner, K.; Schuster, B. Insertion of an Anionic Analogue of the Antimicrobial Peptide PGLa in Lipid Architectures including S-Layer Supported Lipid Bilayers. Curr. Nanosci. 2013, 9, 262–270. [Google Scholar] [CrossRef]
- Chang, W.K.; Wimley, W.C.; Searson, P.C.; Hristova, K.; Merzlyakov, M. Characterization of antimicrobial peptide activity by electrochemical impedance spectroscopy. Biochim. Biophys. Acta 2008, 1778, 2430–2436. [Google Scholar] [CrossRef] [PubMed]
- Wimley, W.C.; Hristova, K. Antimicrobial Peptides; Successes, Challenges and Unanswered Questions. J. Membr. Biol. 2011, 239, 27–34. [Google Scholar] [CrossRef] [PubMed]
- Puiu, M.; Bala, C. Peptide-based biosensors: From self-assembled interfaces to molecular probes in electrochemical assays. Bioelectrochemistry 2018, 120, 66–75. [Google Scholar] [CrossRef] [PubMed]
- Schuster, B.; Sleytr, U.B. Tailor-made crystalline structures of truncated S-layer proteins on heteropolysaccharides. Soft Matter 2009, 5, 334–341. [Google Scholar] [CrossRef]
- Trojanowicz, M. Miniaturized biochemical sensing devices based on planar lipid membranes. Fresenius J. Anal. Chem. 2001, 372, 246–260. [Google Scholar] [CrossRef]
- Larisch, V.-D. Characterization of the Ryanodine Receptor 1 in Model Lipid Membranes. Master’s Thesis, University of Natural Resources and Life Sciences, Vienna, Austria, 2012. [Google Scholar]
- Damiati, S.; Zayni, S.; Schrems, A.; Kiene, E.; Sleytr, U.B.; Chopineau, J.; Schuster, B.; Sinner, E.K. Inspired and stabilized by nature: Ribosomal synthesis of the human voltage gated ion channel (VDAC) into 2D-protein-tethered lipid interfaces. Biomater. Sci. 2015, 3, 1406–1413. [Google Scholar] [CrossRef] [PubMed]
- Colombini, M. VDAC: The channel at the interface between mitochondria and the cytosol. Mol. Cell. Biochem. 2004, 256–257, 107–115. [Google Scholar] [CrossRef] [PubMed]
- Colombini, M. VDAC structure, selectivity, and dynamics. Biochim. Biophys. Acta 2012, 1818, 1457–1465. [Google Scholar] [CrossRef] [PubMed]
- Colombini, M. The VDAC channel: Molecular basis for selectivity. Biochim. Biophys. Acta 2016, 1863, 2498–2502. [Google Scholar] [CrossRef] [PubMed]
- Colombini, M.; Blachly-Dyson, E.; Forte, M. VDAC, a channel in the outer mitochondrial membrane. Ion. Channels 1996, 4, 169–202. [Google Scholar] [PubMed]
- Komarov, A.G.; Deng, D.; Craigen, W.J.; Colombini, M. New insights into the mechanism of permeation through large channels. Biophys. J. 2005, 89, 3950–3959. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Lee, A.C.; Xu, X.; Colombini, M. The role of pyridine dinucleotides in regulating the permeability of the mitochondrial outer membrane. J. Biol. Chem. 1996, 271, 26724–26731. [Google Scholar] [CrossRef] [PubMed]
- Rostovtseva, T.K.; Komarov, A.; Bezrukov, S.M.; Colombini, M. Dynamics of nucleotides in VDAC channels: Structure-specific noise generation. Biophys. J. 2002, 82, 193–205. [Google Scholar] [CrossRef]
- Zizi, M.; Forte, M.; Blachly-Dyson, E.; Colombini, M. NADH regulates the gating of VDAC, the mitochondrial outer membrane channel. J. Biol. Chem. 1994, 269, 1614–1616. [Google Scholar] [PubMed]
- Howard, A.D.; McAllister, G.; Feighner, S.D.; Liu, Q.; Nargund, R.P.; Van der Ploeg, L.H.T.; Patchett, A.A. Orphan G-protein-coupled receptors and natural ligand discovery. Trends Pharmacol. Sci. 2001, 22, 132–140. [Google Scholar] [CrossRef]
- Howard, A.D.; Wang, R.; Pong, S.S.; Mellin, T.N.; Strack, A.; Guan, X.M.; Zeng, Z.; Williams, D.L., Jr; Feighner, S.D.; Nunes, C.N.; et al. Identification of receptors for neuromedin U and its role in feeding. Nature 2000, 406, 70–74. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.H.; Han, H.K. Nanomedicines: Current status and future perspectives in aspect of drug delivery and pharmacokinetics. J. Pharm. Investig. 2018, 48, 43–60. [Google Scholar] [CrossRef]
- Sala, M.; Diab, R.; Elaissari, A.; Fessi, H. Lipid nanocarriers as skin drug delivery systems: Properties, mechanisms of skin interactions and medical applications. Int. J. Pharm. 2018, 535, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Dearling, J.L.J.; Packard, A.B. Molecular imaging in nanomedicine—A developmental tool and a clinical necessity. J. Control. Release 2017, 261, 23–30. [Google Scholar] [CrossRef] [PubMed]
- Küpcü, S.; Sára, M.; Sleytr, U.B. Liposomes coated with crystalline bacterial cell surface protein (S-layer) as immobilization structures for macromolecules. Biochim. Biophys. Acta 1995, 1235, 263–269. [Google Scholar] [CrossRef]
- Ücisik, M.H.; Küpcü, S.; Breitwieser, A.; Gelbmann, N.; Schuster, B.; Sleytr, U.B. S-layer fusion protein as a tool functionalizing emulsomes and CurcuEmulsomes for antibody binding and targeting. Colloids Surf. B Biointerfaces 2015, 128, 132–139. [Google Scholar] [CrossRef] [PubMed]
- Ücisik, M.H.; Küpcü, S.; Debreczeny, M.; Schuster, B.; Sleytr, U.B. S-layer coated emulsomes as potential nanocarriers. Small 2013, 9, 2895–2904. [Google Scholar] [CrossRef] [PubMed]
- Ücisik, M.H.; Sleytr, U.B.; Schuster, B. Emulsomes meet S-layer proteins: An emerging targeted drug delivery system. Curr. Pharm. Biotechnol. 2015, 16, 392–405. [Google Scholar] [CrossRef] [PubMed]

| Type | Reactive Group | Crosslinker | Targeted Group | References |
|---|---|---|---|---|
| Natural SPLs | ||||
| Electrostatic interaction | Negative charges on SLP | Zwitterionic or positively charged lipids | [79,80,145] | |
| Lectin-type like binding | S-layer homologous domain on SLP | Secondary cell wall polymer coupled to lipids | [18] | |
| Chemical modi-fication of SLPs | ||||
| Covalent bond | Carboxyl groups on SLP | Carbodiimide analogues | Primary amine group from lipids | [146,147,168] |
| Covalent bond | Primary amino groups on SLP | SMCC analogues | Thiol group from lipids | [148] |
| Covalent bond | Primary amino groups on SLP | SPDP/TCEP; insertion of thiol group in SLP | Maleimide group from lipids | Schuster et al. in preparation |
| Chemical binding of linker on SLPs | ||||
| Strong ligation | Streptavidin chemically coupled to SLP | Biotinylated lipids | [149,150] | |
| Genetically engineered SLPs | ||||
| Covalent bond | Thiol-group from introduced cysteine | Maleimide group from lipids | [151,152,153] | |
| Multiple chelation | Multiple histidines (His6-tag) on SLP | Nickel(II)-NTA from lipids | [154] | |
| Strong ligation | Streptavidin fused to SLP | Biotinylated lipids | [89] | |
| Strong ligation | Strep-tag fused to SLP | Streptavidin | Biotinylated lipids | [72,73,155] |
| Membrane-Active Peptide | Source | Remarks | References |
|---|---|---|---|
| gramicidin A(gA) | Bacillus brevis | linear pentadeca peptide | [161] |
| alamethicin(Ala) | Trichoderma viride | linear, 20 amino acids | [159] |
| valinomycin (Val) | several Streptomyces strains, e.g., S. tsusimaensis and S. fulvissimus | cyclic dodecadepsi peptide, 12 amino acids and esters | [141,159] |
| peptidyl-glycine-leucine-carboxyamide (PGLa) analogue | synthesized via protein chemistry | 20 amino acid; negatively charged analogue of PGLa | [168] |
| Transmembrane Protein | Source | Remarks | References |
|---|---|---|---|
| α-hemolysin (αHL) | exotoxin from Staphylococcus aureus | pore-forming; homo-heptamer | [144,160] |
| ryanodine receptor 1 (RyR1) | skeletal muscle cells | Ca2+-release channel; homotetramer | [174] |
| nicotinic acetylcholine receptor (nAChR) | plasma membranes of neurons; on postsynaptic side of the neuromuscular junction | ligand gated ion channel; 5 subunits | [154] |
| voltage-dependent anion channel (VDAC) | located on the outer mitochondrial membrane; also produced by cell-free expression | porin, voltage gated; ion channel monomeric but can cluster | [175] |
© 2018 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Schuster, B. S-Layer Protein-Based Biosensors. Biosensors 2018, 8, 40. https://doi.org/10.3390/bios8020040
Schuster B. S-Layer Protein-Based Biosensors. Biosensors. 2018; 8(2):40. https://doi.org/10.3390/bios8020040
Chicago/Turabian StyleSchuster, Bernhard. 2018. "S-Layer Protein-Based Biosensors" Biosensors 8, no. 2: 40. https://doi.org/10.3390/bios8020040
APA StyleSchuster, B. (2018). S-Layer Protein-Based Biosensors. Biosensors, 8(2), 40. https://doi.org/10.3390/bios8020040




