MIPs and Aptamers for Recognition of Proteins in Biomimetic Sensing
Abstract
:1. Introduction
- molecular recognition of the analyte by the aptamer or MIP, which is determined by the corroborative effect of the composition/sequence and spatial arrangement of the respective biomimetic elements.
- generation of a measurable signal as result of the interaction of the target-loaded biomimetic recognition element with the transducer.
2. Preparation and Performance of MIPs for the Recognition of Proteins
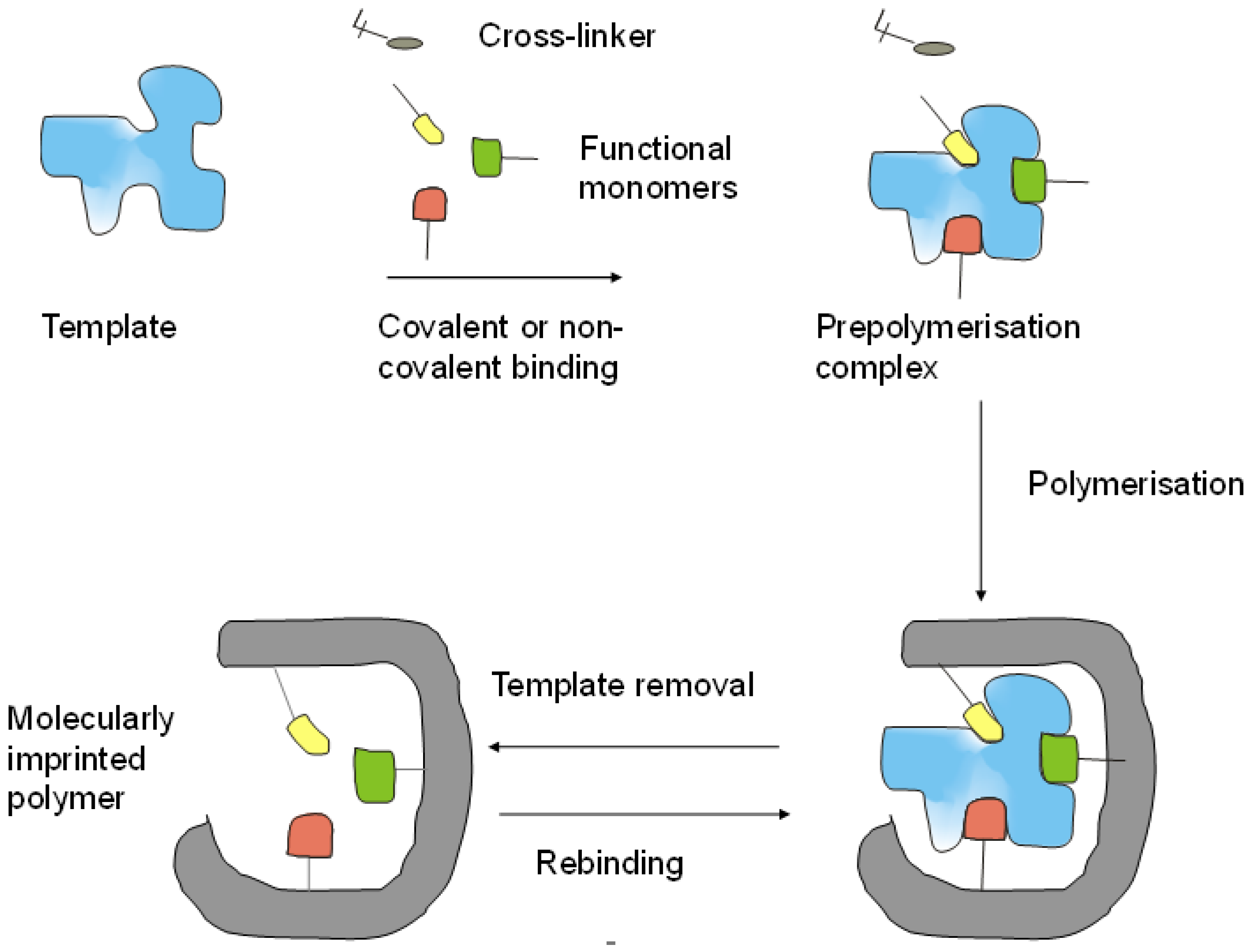
2.1. Building Blocks of MIPs
2.2. Preparation of MIPs for Proteins
2.2.1. Bulk Imprinting of Proteins
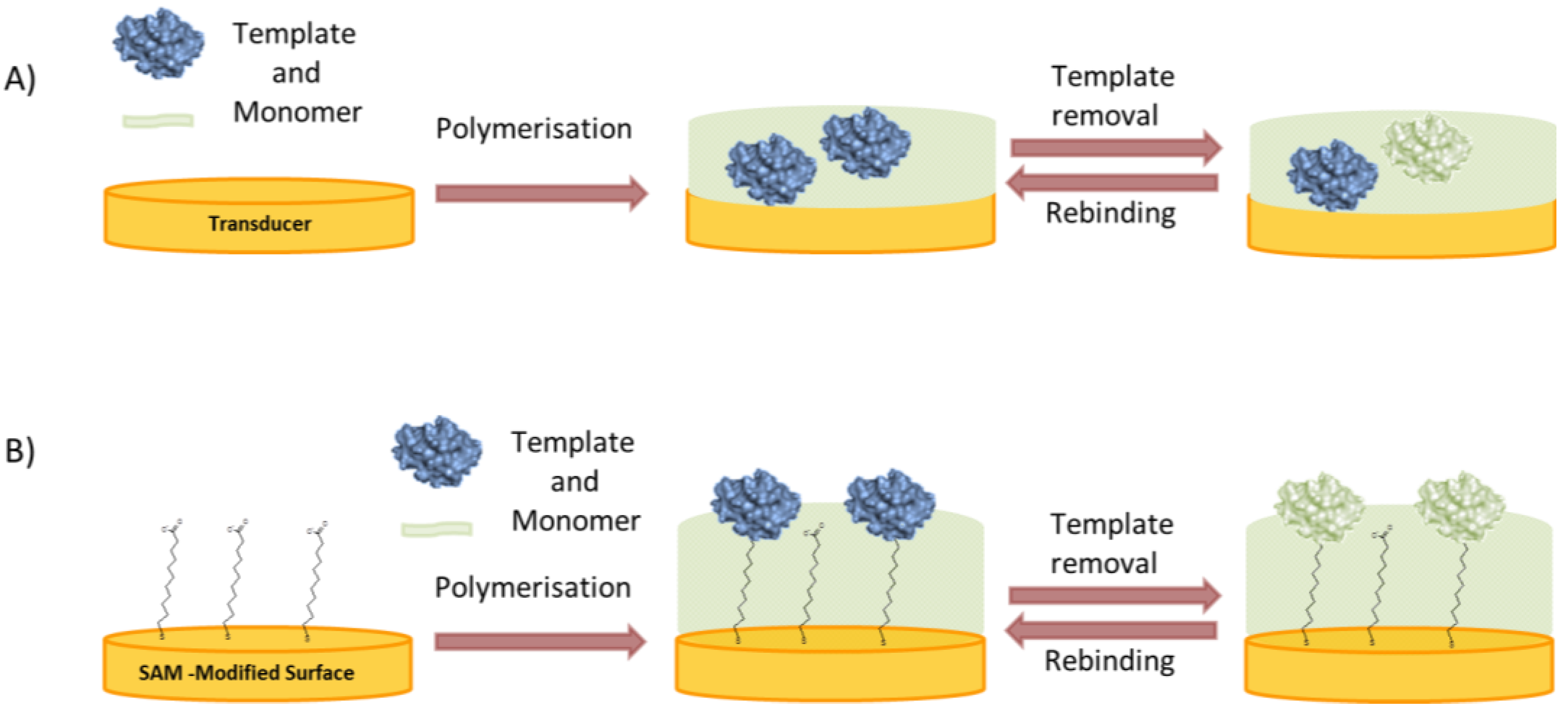
2.2.2. Surface Imprinting
- In the “bottom-up” concept the protein is immobilized directly on the substrate where the polymer is to be deposited (Figure 5B). Various interactions can be exploited to anchor the protein to the surface in an oriented manner, thereby ensuring the formation of uniform binding pockets. Glycoproteins can be immobilized by boronic acids [56,57] enzymes with their inhibitors or substrates [58,59] and, in general, any reversible receptor can be used as anchor, e.g., aptamers [60].
- The preparation of MIPs for the recognition of proteins does not necessarily need the use of the whole macromolecule as template, but a representative fragment may suffice (epitope approach). Consequently, imprinting a peptide sequence, which represents a small and exposed part of the whole protein, was successfully applied for MIPs which recognize the parent protein [61,62,63,64,65]. This approach has several advantages, e.g., the polymerization can be performed in organic solvents, the cost of the peptide is lower than for the protein, it can be synthesized in a very pure form and also a well selected peptide sequence may increase selectivity. On the other hand, the epitopes are typically built up from amino acids located in different sections of the peptide chain held in spatial proximity by the protein’s tertiary structure. A linear epitope, suitable for imprinting, can only be found at the exposed termini of the protein. Therefore, larger fragments of the protein have been also used as the template during polymerization, e.g., the Fab-fragment for a MIP towards IgG [66].
2.3. Binding Performance of MIPs
3. Preparation and Performance of Aptamers for the Recognition of Proteins
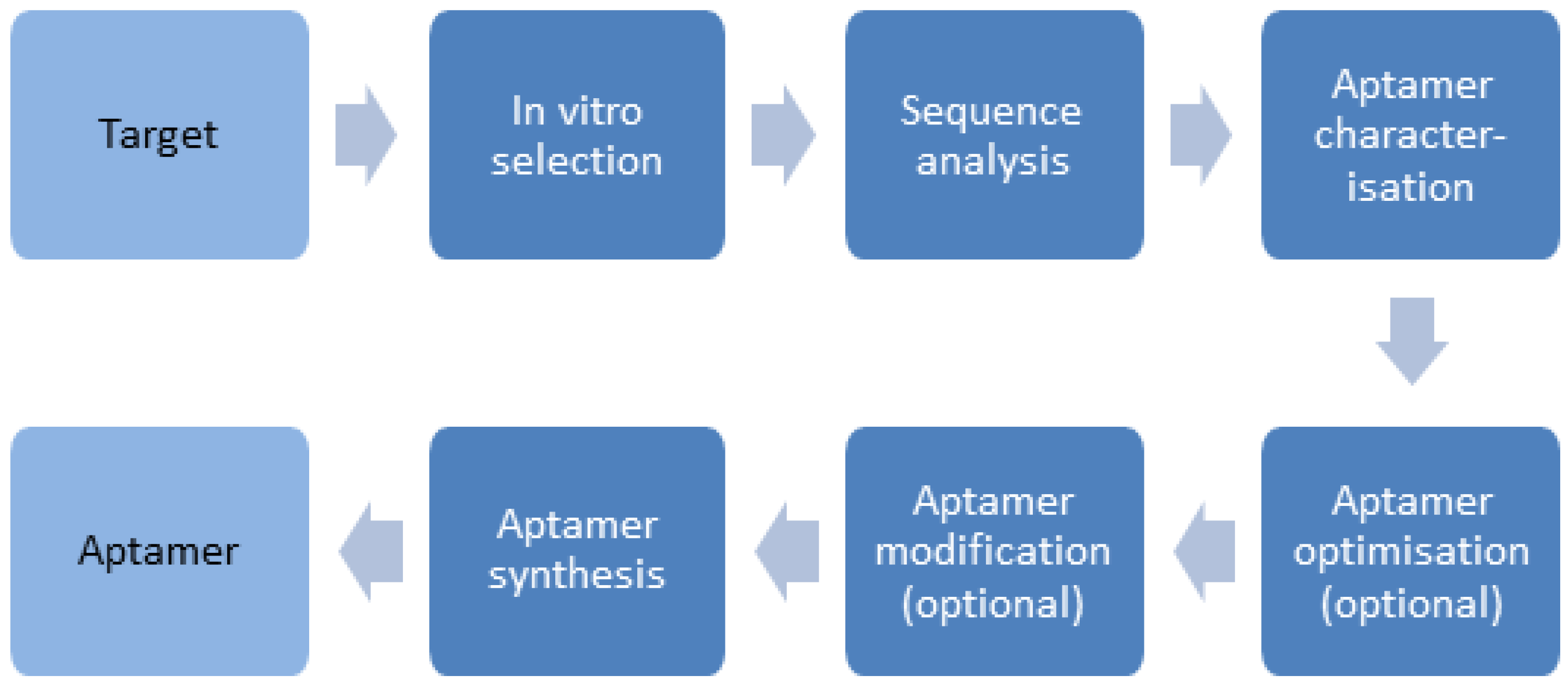
3.1. Preparation of Aptamers
3.2. Binding Performance of Aptamers
4. Catalytically Active MIPs and Aptamers
4.1. Catalytically Active MIPs
4.2. Catalytically Active Aptamers: Aptazymes and Riboswitches
5. Chemical Sensing Schemes
- Label-free evaluation of changes of the recognition layer upon binding of the target protein by QCM and SPR
- Direct indication of target binding
- Fluorescence quenching of the MIP layer upon protein binding
- Changes in the MIP film permeability for an electro-active electrochemical probe
6. Conclusions and Outlook
Acknowledgments
Conflicts of Interest
References
- Polyakov, M.V. Adsorption properties and structure of silica gel. Zhurnal Fizieskoj Khimii/Akad. SSSR 1931, 2, 799–805. [Google Scholar]
- Polyakov, M.V.; Kuleshina, L.; Neimark, I. On the dependence of silica gel adsorption properties on the character of its porosity. Zhurnal Fizieskoj Khimii/Akad. SSSR 1937, 10, 100–112. [Google Scholar]
- Dickey, F.H. The preparation of specific adsorbents. Proc. Natl. Acad. Sci. USA 1949, 35, 227–229. [Google Scholar] [CrossRef] [PubMed]
- Wulff, G.; Sarhan, A. Use of polymers with enzyme-analogous structures for resolution of racemates. Angew. Chem. Int. Ed. 1972, 11, 341–344. [Google Scholar]
- Wulff, G.; Vesper, W.; Grobe-Einsler, R.; Sarhan, A. Enzyme-analogue built polymers, 4. On the synthesis of polymers containing chiral cavities and their use for the resolution of racemates. Makromol. Chem. Macromol. Chem. Phys. 1977, 178, 2799–2816. [Google Scholar] [CrossRef]
- Arshady, R.; Mosbach, K. Synthesis of substrate-selective polymers by host-guest polymerization. Macromol. Chem. Phys. Makromol. Chem. 1981, 182, 687–692. [Google Scholar] [CrossRef]
- Glad, M.; Norrlow, O.; Sellergren, B.; Siegbahn, N.; Mosbach, K. Use of silane monomers for molecular imprinting and enzyme entrapment in polysiloxane-coated porous silica. J. Chromatogr. 1985, 347, 11–23. [Google Scholar] [CrossRef]
- Norrlow, O.; Glad, M.; Mosbach, K. Acrylic polymer preparations containing recognition sites obtained by imprinting with substrates. J. Chromatogr. 1984, 299, 29–41. [Google Scholar] [CrossRef]
- Uzun, L.; Turner, A.P.F. Molecularly-imprinted polymer sensors: Realising their potential. Biosens. Bioelectron. 2016, 76, 131–144. [Google Scholar] [CrossRef] [PubMed]
- Kempe, M.; Glad, M.; Mosbach, K. An approach towards surface imprinting using the enzyme ribonuclease A. J. Mol. Recognit. 1995, 8, 35–39. [Google Scholar] [CrossRef] [PubMed]
- Takeuchi, T.; Hishiya, T. Molecular imprinting of proteins emerging as a tool for protein recognition. Org. Biomol. Chem. 2008, 6, 2459–2467. [Google Scholar] [CrossRef] [PubMed]
- Whitcombe, M.J.; Chianella, I.; Larcombe, L.; Piletsky, S.A.; Noble, J.; Porter, R.; Horgan, A. The rational development of molecularly imprinted polymer-based sensors for protein detection. Chem. Soc. Rev. 2011, 40, 1547–1571. [Google Scholar] [CrossRef] [PubMed]
- Yang, K.; Zhang, L.; Liang, Z.; Zhang, Y. Protein-imprinted materials: Rational design, application and challenges. Anal. Bioanal. Chem. 2012, 403, 2173–2183. [Google Scholar] [CrossRef] [PubMed]
- Li, S.J.; Cao, S.S.; Whitcombe, M.J.; Piletsky, S.A. Size matters: Challenges in imprinting macromolecules. Prog. Polym. Sci. 2014, 39, 145–163. [Google Scholar] [CrossRef]
- Erdössy, J.; Horváth, V.; Yarman, A.; Scheller, F.W.; Gyurcsányi, R.E. Electrosynthesized molecularly imprinted polymers for protein recognition. Trends Anal. Chem. 2016, 79, 179–190. [Google Scholar] [CrossRef]
- Mipdatabase. Available online: http://www.mipdatabase.com (accessed on 15 July 2016).
- Ellington, A.D.; Szostak, J.W. In vitro selection of RNA molecules that bind specific ligands. Nature 1990, 346, 818–822. [Google Scholar] [CrossRef] [PubMed]
- Robertson, D.L.; Joyce, G.F. Selection in vitro of an RNA enzyme that specifically cleaves single-stranded DNA. Nature 1990, 344, 467–468. [Google Scholar] [CrossRef] [PubMed]
- Tuerk, C.; Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 1990, 249, 505–510. [Google Scholar] [CrossRef] [PubMed]
- Bock, L.C.; Griffin, L.C.; Latham, J.A.; Vermaas, E.H.; Toole, J.J. Selection of single-stranded DNA molecules that bind and inhibit human thrombin. Nature 1992, 355, 564–566. [Google Scholar] [CrossRef] [PubMed]
- Cohen, B.A.; Colas, P.; Brent, R. An artificial cell-cycle inhibitor isolated from a combinatorial library. Proc. Natl. Acad. Sci. USA 1998, 95, 14272–14277. [Google Scholar] [CrossRef] [PubMed]
- Crawford, M.; Woodman, R.; Ko Ferrigno, P. Peptide aptamers: Tools for biology and drug discovery. Brief. Funct. Genom. Proteom. 2003, 2, 72–79. [Google Scholar] [CrossRef]
- Hoppe-Seyler, F.; Crnkovic-Mertens, I.; Tomai, E.; Butz, K. Peptide aptamers: Specific inhibitors of protein function. Curr. Mol. Med. 2004, 4, 529–538. [Google Scholar] [CrossRef] [PubMed]
- Reverdatto, S.; Burz, D.S.; Shekhtman, A. Peptide aptamers: Development and applications. Curr. Top. Med. Chem. 2015, 15, 1082–1101. [Google Scholar] [CrossRef] [PubMed]
- Breaker, R.R. Natural and engineered nucleic acids as tools to explore biology. Nature 2004, 432, 838–845. [Google Scholar] [CrossRef] [PubMed]
- Feigon, J.; Dieckmann, T.; Smith, F.W. Aptamer structures from a to zeta. Chem. Biol. 1996, 3, 611–617. [Google Scholar] [CrossRef]
- Klussmann, S. The Aptamer Handbook: Functional Oligonucleotides and Their Applications; Wiley: Weinheim, Germany, 2006. [Google Scholar]
- Ellington, A.D.; Szostak, J.W. Selection in vitro of single-stranded DNA molecules that fold into specific ligand-binding structures. Nature 1992, 355, 850–852. [Google Scholar] [CrossRef] [PubMed]
- Boiziau, C.; Toulme, J.J. A method to select chemically modified aptamers directly. Antisense Nucleic Acid Drug Dev. 2001, 11, 379–385. [Google Scholar] [CrossRef] [PubMed]
- Connell, G.J.; Illangesekare, M.; Yarus, M. Three small ribooligonucleotides with specific arginine sites. Biochemistry 1993, 32, 5497–5502. [Google Scholar] [CrossRef] [PubMed]
- Famulok, M. Molecular recognition of amino-acids by RNA-aptamers—An L-citrulline binding rna motif and its evolution into an L-arginine binder. J. Am. Chem. Soc. 1994, 116, 1698–1706. [Google Scholar] [CrossRef]
- Wallis, M.G.; von Ahsen, U.; Schroeder, R.; Famulok, M. A novel rna motif for neomycin recognition. Chem. Biol. 1995, 2, 543–552. [Google Scholar] [CrossRef]
- Wochner, A.; Menger, M.; Orgel, D.; Cech, B.; Rimmele, M.; Erdmann, V.A.; Glokler, J. A DNA aptamer with high affinity and specificity for therapeutic anthracyclines. Anal. Biochem. 2008, 373, 34–42. [Google Scholar] [CrossRef] [PubMed]
- Mendonsa, S.D.; Bowser, M.T. In vitro selection of aptamers with affinity for neuropeptide y using capillary electrophoresis. J. Am. Chem. Soc. 2005, 127, 9382–9383. [Google Scholar] [CrossRef] [PubMed]
- Green, L.S.; Jellinek, D.; Jenison, R.; Ostman, A.; Heldin, C.H.; Janjic, N. Inhibitory DNA ligands to platelet-derived growth factor b-chain. Biochemistry 1996, 35, 14413–14424. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Zhou, J.; Luo, F.; Mohammed, A.B.; Zhang, X.L. Aptamer from whole-bacterium selex as new therapeutic reagent against virulent mycobacterium tuberculosis. Biochem. Biophys. Res. Commun. 2007, 357, 743–748. [Google Scholar] [CrossRef] [PubMed]
- Hamula, C.L.; Zhang, H.; Guan, L.L.; Li, X.F.; Le, X.C. Selection of aptamers against live bacterial cells. Anal. Chem. 2008, 80, 7812–7819. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Jiang, H.; Liu, F. In vitro selection of novel rna ligands that bind human cytomegalovirus and block viral infection. RNA 2000, 6, 571–583. [Google Scholar] [CrossRef] [PubMed]
- Shangguan, D.; Li, Y.; Tang, Z.W.; Cao, Z.H.C.; Chen, H.W.; Mallikaratchy, P.; Sefah, K.; Yang, C.Y.J.; Tan, W.H. Aptamers evolved from live cells as effective molecular probes for cancer study. Proc. Natl. Acad. Sci. USA 2006, 103, 11838–11843. [Google Scholar] [CrossRef] [PubMed]
- Tang, Z.W.; Parekh, P.; Turner, P.; Moyer, R.W.; Tan, W.H. Generating aptamers for recognition of virus-infected cells. Clin. Chem. 2009, 55, 813–822. [Google Scholar] [CrossRef] [PubMed]
- Nimjee, S.M.; Rusconi, C.P.; Sullenger, B.A. Aptamers: An emerging class of therapeutics. Annu. Rev. Med. 2005, 56, 555–583. [Google Scholar] [CrossRef] [PubMed]
- Saffhill, R.; Schneider-Bernloehr, H.; Orgel, L.E.; Spiegelman, S. In vitro selection of bacteriophage q-beta ribonucleic acid variants resistant to ethidium bromide. J. Mol. Biol. 1970, 51, 531–539. [Google Scholar] [CrossRef]
- Scopus. Available online: https://www.scopus.com (accessed on 15 July 2016).
- Hjerten, S.; Liao, J.L.; Nakazato, K.; Wang, Y.; Zamaratskaia, G.; Zhang, H.X. Gels mimicking antibodies in their selective recognition of proteins. Chromatographia 1997, 44, 227–234. [Google Scholar] [CrossRef]
- Tong, D.; Hetenyi, C.; Bikadi, Z.; Gao, J.P.; Hjerten, S. Some studies of the chromatographic properties of gels (‘artificial antibodies/receptors’) for selective adsorption of proteins. Chromatographia 2001, 54, 7–14. [Google Scholar] [CrossRef]
- Bosserdt, M.; Erdőssy, J.; Lautner, G.; Witt, J.; Kohler, K.; Gajovic-Eichelmann, N.; Yarman, A.; Wittstock, G.; Scheller, F.W.; Gyurcsányi, R.E. Microelectrospotting as a new method for electrosynthesis of surface-imprinted polymer microarrays for protein recognition. Biosens. Bioelectron. 2015, 73, 123–129. [Google Scholar] [CrossRef] [PubMed]
- Ambrosini, S.; Beyazit, S.; Haupt, K.; Bui, B.T.S. Solid-phase synthesis of molecularly imprinted nanoparticles for protein recognition. Chem. Commun. 2013, 49, 6746–6748. [Google Scholar] [CrossRef] [PubMed]
- Pluhar, B.; Mizaikoff, B. Advanced evaluation strategies for protein-imprinted polymer nanobeads. Macromol. Biosci. 2015, 15, 1507–1511. [Google Scholar] [CrossRef] [PubMed]
- Zhou, T.C.; Zhang, K.; Kamra, T.; Bulow, L.; Ye, L. Preparation of protein imprinted polymer beads by pickering emulsion polymerization. J. Mater. Chem. B 2015, 3, 1254–1260. [Google Scholar] [CrossRef]
- Hayden, O.; Lieberzeit, P.A.; Blaas, D.; Dickert, F.L. Artificial antibodies for bioanalyte detection-sensing viruses and proteins. Adv. Funct. Mater. 2006, 16, 1269–1278. [Google Scholar] [CrossRef]
- Lin, H.Y.; Hsu, C.Y.; Thomas, J.L.; Wang, S.E.; Chen, H.C.; Chou, T.C. The microcontact imprinting of proteins: The effect of cross-linking monomers for lysozyme, ribonuclease a and myoglobin. Biosens. Bioelectron. 2006, 22, 534–543. [Google Scholar] [CrossRef] [PubMed]
- Sener, G.; Ozgur, E.; Rad, A.Y.; Uzun, L.; Say, R.; Denizli, A. Rapid real-time detection of procalcitonin using a microcontact imprinted surface plasmon resonance biosensor. Analyst 2013, 138, 6422–6428. [Google Scholar] [CrossRef] [PubMed]
- Bognar, J.; Szucs, J.; Dorko, Z.; Horvath, V.; Gyurcsanyi, R.E. Nanosphere lithography as a versatile method to generate surface-imprinted polymer films for selective protein recognition. Adv. Funct. Mater. 2013, 23, 4703–4709. [Google Scholar] [CrossRef]
- Menaker, A.; Syritski, V.; Reut, J.; Opik, A.; Horvath, V.; Gyurcsanyi, R.E. Electrosynthesized surface-imprinted conducting polymer microrods for selective protein recognition. Adv. Mater. 2009, 21, 2271–2275. [Google Scholar] [CrossRef]
- Lautner, G.; Kaev, J.; Reut, J.; Opik, A.; Rappich, J.; Syritski, V.; Gyurcsányi, R.E. Selective artificial receptors based on micropatterned surface-imprinted polymers for label-free detection of proteins by spr imaging. Adv. Funct. Mater. 2011, 21, 591–597. [Google Scholar] [CrossRef]
- Liu, S.Q.; Bakovic, L.; Chen, A.C. Specific binding of glycoproteins with poly(aniline boronic acid) thin film. J. Electroanal. Chem. 2006, 591, 210–216. [Google Scholar] [CrossRef]
- Wang, S.S.; Ye, J.; Bie, Z.J.; Liu, Z. Affinity-tunable specific recognition of glycoproteins via boronate affinity-based controllable oriented surface imprinting. Chem. Sci. 2014, 5, 1135–1140. [Google Scholar] [CrossRef]
- Dechtrirat, D.; Gajovic-Eichelmann, N.; Bier, F.F.; Scheller, F.W. Hybrid material for protein sensing based on electrosynthesized mip on a mannose terminated selfassembled monolayer. Adv. Funct. Mater. 2014, 24, 2233–2239. [Google Scholar] [CrossRef]
- Jetzschmann, K.J.; Jágerszki, G.; Dechtrirat, D.; Yarman, A.; Gajovic-Eichelmann, N.; Gilsing, H.D.; Schulz, B.; Gyurcsányi, R.E.; Scheller, F.W. Vectorially imprinted hybrid nanofilm for acetylcholinesterase recognition. Adv. Funct. Mater. 2015, 25, 5178–5183. [Google Scholar] [CrossRef]
- Jolly, P.; Tamboli, V.; Harniman, R.L.; Estrela, P.; Allender, C.J.; Bowen, J.L. Aptamer-mip hybrid receptor for highly sensitive electrochemical detection of prostate specific antigen. Biosens. Bioelectron. 2016, 75, 188–195. [Google Scholar] [CrossRef] [PubMed]
- Bossi, A.M.; Sharma, P.S.; Montana, L.; Zoccatelli, G.; Laub, O.; Levi, R. Fingerprint-imprinted polymer: Rational selection of peptide epitope templates for the determination of proteins by molecularly imprinted polymers. Anal. Chem. 2012, 84, 4036–4041. [Google Scholar] [CrossRef] [PubMed]
- Cenci, L.; Anesi, A.; Busato, M.; Guella, G.; Bossi, A.M. Molecularly imprinted polymers coupled to matrix assisted laser desorption ionization mass spectrometry for femtomoles detection of cardiac troponin I peptides. J. Mol. Recognit. 2016, 29, 41–50. [Google Scholar] [CrossRef] [PubMed]
- Dechtrirat, D.; Jetzschmann, K.J.; Stocklein, W.F.M.; Scheller, F.W.; Gajovic-Eichelmann, N. Protein rebinding to a surface-confined imprint. Adv. Funct. Mater. 2012, 22, 5231–5237. [Google Scholar] [CrossRef]
- Nishino, H.; Huang, C.S.; Shea, K.J. Selective protein capture by epitope imprinting. Angew. Chem. Int. Ed. 2006, 45, 2392–2396. [Google Scholar] [CrossRef] [PubMed]
- Rachkov, A.; Minoura, N. Recognition of oxytocin and oxytocin-related peptides in aqueous media using a molecularly imprinted polymer synthesized by the epitope approach. J. Chromatogr. A 2000, 889, 111–118. [Google Scholar] [CrossRef]
- Erturk, G.; Uzun, L.; Tumer, M.A.; Say, R.; Denizli, A. Fab fragments imprinted spr biosensor for real-time human immunoglobulin g detection. Biosens. Bioelectron. 2011, 28, 97–104. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.D.; Dong, J.; Ming, H.M.; Ai, S.Y. Sensing of glycoprotein via a biomimetic sensor based on molecularly imprinted polymers and graphene-au nanoparticles. Analyst 2013, 138, 1219–1225. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.J.; Zhang, Z.H.; Xie, D.; Cai, R.; Chen, X.; Liu, Y.N.; Yao, S.Z. Surface-imprinting sensor based on carbon nanotubes/graphene composite for determination of bovine serum albumin. Electroanalysis 2012, 24, 2109–2116. [Google Scholar] [CrossRef]
- Cai, D.; Ren, L.; Zhao, H.; Xu, C.; Zhang, L.; Yu, Y.; Wang, H.; Lan, Y.; Roberts, M.F.; Chuang, J.H.; et al. A molecular-imprint nanosensor for ultrasensitive detection of proteins. Nat. Nanotechnol. 2010, 5, 597–601. [Google Scholar] [CrossRef] [PubMed]
- Cieplak, M.; Szwabinska, K.; Sosnowska, M.; Bikram, K.C.C.; Borowicz, P.; Noworyta, K.; D’Souza, F.; Kutner, W. Selective electrochemical sensing of human serum albumin by semi-covalent molecular imprinting. Biosens. Bioelectron. 2015, 74, 960–966. [Google Scholar] [CrossRef] [PubMed]
- Karimian, N.; Turner, A.P.F.; Tiwari, A. Electrochemical evaluation of troponin T imprinted polymer receptor. Biosens. Bioelectron. 2014, 59, 160–165. [Google Scholar] [CrossRef] [PubMed]
- Silva, B.V.M.; Rodriguez, B.A.G.; Sales, G.F.; Sotomayor, M.D.T.; Dutra, R.F. An ultrasensitive human cardiac troponin t graphene screen-printed electrode based on electropolymerized-molecularly imprinted conducting polymer. Biosens. Bioelectron. 2016, 77, 978–985. [Google Scholar] [CrossRef] [PubMed]
- Klug, S.J.; Famulok, M. All you wanted to know about selex. Mol. Biol. Rep. 1994, 20, 97–107. [Google Scholar] [CrossRef] [PubMed]
- Szeitner, Z.; András, J.; Gyurcsányi, R.E.; Mészáros, T. Is less more? Lessons from aptamer selection strategies. J. Pharm. Biomed. Anal. 2014, 101, 58–65. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Zhu, Y.; Xue, F.; Mei, Z.; Yao, L.; Wang, X.; Zheng, L.; Liu, J.; Liu, G.; Peng, C.; et al. Recent trends in selex technique and its application to food safety monitoring. Mikrochim. Acta 2014, 181, 479–491. [Google Scholar] [CrossRef] [PubMed]
- Yuce, M.; Ullah, N.; Budak, H. Trends in aptamer selection methods and applications. Analyst 2015, 140, 5379–5399. [Google Scholar] [CrossRef] [PubMed]
- Balogh, Z.; Lautner, G.; Bardoczy, V.; Komorowska, B.; Gyurcsányi, R.E.; Mészáros, T. Selection and versatile application of virus-specific aptamers. FASEB J. 2010, 24, 4187–4195. [Google Scholar] [CrossRef] [PubMed]
- Lautner, G.; Balogh, Z.; Bardoczy, V.; Meszáros, T.; Gyurcsányi, R.E. Aptamer-based biochips for label-free detection of plant virus coat proteins by spr imaging. Analyst 2010, 135, 918–926. [Google Scholar] [CrossRef] [PubMed]
- Drolet, D.W.; Jenison, R.D.; Smith, D.E.; Pratt, D.; Hicke, B.J. A high throughput platform for systematic evolution of ligands by exponential enrichment (selex). Comb. Chem. High Throughput Screen. 1999, 2, 271–278. [Google Scholar] [PubMed]
- Cox, J.C.; Ellington, A.D. Automated selection of anti-protein aptamers. Bioorg. Med. Chem. 2001, 9, 2525–2531. [Google Scholar] [CrossRef]
- Eulberg, D.; Buchner, K.; Maasch, C.; Klussmann, S. Development of an automated in vitro selection protocol to obtain rna-based aptamers: Identification of a biostable substance p antagonist. Nucleic Acids Res. 2005, 33, e45. [Google Scholar] [CrossRef] [PubMed]
- Wochner, A.; Cech, B.; Menger, M.; Erdmann, V.A.; Glokler, J. Semi-automated selection of DNA aptamers using magnetic particle handling. Biotechniques 2007, 43, 344–353. [Google Scholar] [CrossRef] [PubMed]
- Glokler, J.; Schutze, T.; Konthur, Z. Automation in the high-throughput selection of random combinatorial libraries—Different approaches for select applications. Molecules 2010, 15, 2478–2490. [Google Scholar] [CrossRef] [PubMed]
- Hunniger, T.; Wessels, H.; Fischer, C.; Paschke-Kratzin, A.; Fischer, M. Just in time-selection: A rapid semiautomated selex of DNA aptamers using magnetic separation and beaming. Anal. Chem. 2014, 86, 10940–10947. [Google Scholar] [CrossRef] [PubMed]
- Hamedani, N.S.; Muller, J. Capillary electrophoresis for the selection of DNA aptamers recognizing activated protein c. Methods Mol. Biol. 2016, 1380, 61–75. [Google Scholar] [PubMed]
- Daniels, D.A.; Chen, H.; Hicke, B.J.; Swiderek, K.M.; Gold, L. A tenascin-c aptamer identified by tumor cell selex: Systematic evolution of ligands by exponential enrichment. Proc. Natl. Acad. Sci. USA 2003, 100, 15416–15421. [Google Scholar] [CrossRef] [PubMed]
- Ohuchi, S. Cell-selex technology. BioRes. Open Access 2012, 1, 265–272. [Google Scholar] [CrossRef] [PubMed]
- Hung, L.Y.; Wang, C.H.; Hsu, K.F.; Chou, C.Y.; Lee, G.B. An on-chip cell-selex process for automatic selection of high-affinity aptamers specific to different histologically classified ovarian cancer cells. Lab Chip 2014, 14, 4017–4028. [Google Scholar] [CrossRef]
- Song, K.M.; Lee, S.; Ban, C. Aptamers and their biological applications. Sensors (Basel) 2012, 12, 612–631. [Google Scholar] [CrossRef] [PubMed]
- Tombelli, S.; Minunni, M.; Mascini, M. Aptamers-based assays for diagnostics, environmental and food analysis. Biomol. Eng. 2007, 24, 191–200. [Google Scholar] [CrossRef] [PubMed]
- Tsae, P.K.; DeRosa, M.C. Outlook for aptamers after twenty five years. Curr. Top. Med. Chem. 2015, 15, 1153–1159. [Google Scholar] [CrossRef] [PubMed]
- Szeitner, Z.; Lautner, G.; Nagy, S.K.; Gyurcsányi, R.E.; Mészáros, T. A rational approach for generating cardiac troponin I selective spiegelmers. Chem. Commun. 2014, 50, 6801–6804. [Google Scholar] [CrossRef] [PubMed]
- Pendergrast, P.S.; Marsh, H.N.; Grate, D.; Healy, J.M.; Stanton, M. Nucleic acid aptamers for target validation and therapeutic applications. J. Biomol. Tech. 2005, 16, 224–234. [Google Scholar] [PubMed]
- Kim, Y.S.; Gu, M.B. Advances in aptamer screening and small molecule aptasensors. Adv. Biochem. Eng. Biotechnol. 2014, 140, 29–67. [Google Scholar] [PubMed]
- Kim, Y.S.; Raston, N.H.; Gu, M.B. Aptamer-based nanobiosensors. Biosens. Bioelectron. 2016, 76, 2–19. [Google Scholar] [PubMed]
- MacKay, S.; Wishart, D.; Xing, J.Z.; Chen, J. Developing trends in aptamer-based biosensor devices and their applications. IEEE Trans. Biomed. Circuits Syst. 2014, 8, 4–14. [Google Scholar] [CrossRef] [PubMed]
- Van den Kieboom, C.H.; van der Beek, S.L.; Mészáros, T.; Gyurcsányi, R.E.; Ferwerda, G.; de Jonge, M.I. Aptasensors for viral diagnostics. TrAC Trends Anal. Chem. 2015, 74, 58–67. [Google Scholar] [CrossRef]
- Ozalp, V.C.; Kavruk, M.; Dilek, O.; Bayrac, A.T. Aptamers: Molecular tools for medical diagnosis. Curr. Top. Med. Chem. 2015, 15, 1125–1137. [Google Scholar] [CrossRef] [PubMed]
- Strehlitz, B.; Reinemann, C.; Linkorn, S.; Stoltenburg, R. Aptamers for pharmaceuticals and their application in environmental analytics. Bioanal. Rev. 2012, 4, 1–30. [Google Scholar] [CrossRef] [PubMed]
- Ray, P.; Viles, K.D.; Soule, E.E.; Woodruff, R.S. Application of aptamers for targeted therapeutics. Arch. Immunol. Ther. Exp. (Warsz.) 2013, 61, 255–271. [Google Scholar] [CrossRef] [PubMed]
- Wochner, A.; Menger, M.; Rimmele, M. Characterisation of aptamers for therapeutic studies. Exp. Opin. Drug Discov. 2007, 2, 1205–1224. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Rossi, J.J. The therapeutic potential of cell-internalizing aptamers. Curr. Top. Med. Chem. 2009, 9, 1144–1157. [Google Scholar] [CrossRef] [PubMed]
- Zhu, G.; Ye, M.; Donovan, M.J.; Song, E.; Zhao, Z.; Tan, W. Nucleic acid aptamers: An emerging frontier in cancer therapy. Chem. Commun. (Camb.) 2012, 48, 10472–10480. [Google Scholar] [CrossRef] [PubMed]
- Gijs, M.; Aerts, A.; Impens, N.; Baatout, S.; Luxen, A. Aptamers as radiopharmaceuticals for nuclear imaging and therapy. Nucl. Med. Biol. 2016, 43, 253–271. [Google Scholar] [CrossRef] [PubMed]
- Jin, C.; Zheng, J.; Li, C.; Qiu, L.; Zhang, X.; Tan, W. Aptamers selected by cell-selex for molecular imaging. J. Mol. Evol. 2015, 81, 162–171. [Google Scholar] [CrossRef] [PubMed]
- Blank, M.; Blind, M. Aptamers as tools for target validation. Curr. Opin. Chem. Biol. 2005, 9, 336–342. [Google Scholar] [CrossRef] [PubMed]
- Guo, K.T.; SchAfer, R.; Paul, A.; Gerber, A.; Ziemer, G.; Wendel, H.P. A new technique for the isolation and surface immobilization of mesenchymal stem cells from whole bone marrow using high-specific DNA aptamers. Stem Cells 2006, 24, 2220–2231. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Cao, G.; Ding, H.; Zhang, D.; Yang, G.; Liu, N.; Fan, M.; Shen, B.; Shao, N. Screening of functional antidotes of rna aptamers against bovine thrombin. FEBS Lett. 2004, 562, 125–128. [Google Scholar] [CrossRef]
- Rusconi, C.P.; Roberts, J.D.; Pitoc, G.A.; Nimjee, S.M.; White, R.R.; Quick, G.; Scardino, E.; Fay, W.P.; Sullenger, B.A. Antidote-mediated control of an anticoagulant aptamer in vivo. Nat. Biotechnol. 2004, 22, 1423–1428. [Google Scholar] [CrossRef] [PubMed]
- Vater, A.; Klussmann, S. Turning mirror-image oligonucleotides into drugs: The evolution of spiegelmer(®) therapeutics. Drug Discov. Today 2015, 20, 147–155. [Google Scholar] [CrossRef] [PubMed]
- Vater, A.; Klussmann, S. Toward third-generation aptamers: Spiegelmers and their therapeutic prospects. Curr. Opin. Drug Discov. Dev. 2003, 6, 253–261. [Google Scholar]
- Brody, E.N.; Gold, L.; Lawn, R.M.; Walker, J.J.; Zichi, D. High-content affinity-based proteomics: Unlocking protein biomarker discovery. Exp. Rev. Mol. Diagn. 2010, 10, 1013–1022. [Google Scholar] [CrossRef] [PubMed]
- Ochsner, U.A.; Green, L.S.; Gold, L.; Janjic, N. Systematic selection of modified aptamer pairs for diagnostic sandwich assays. Biotechniques 2014, 56, 125–128, 130, 132–133. [Google Scholar] [CrossRef] [PubMed]
- Webber, J.; Stone, T.C.; Katilius, E.; Smith, B.C.; Gordon, B.; Mason, M.D.; Tabi, Z.; Brewis, I.A.; Clayton, A. Proteomics analysis of cancer exosomes using a novel modified aptamer-based array (somascan) platform. Mol. Cell. Proteom. 2014, 13, 1050–1064. [Google Scholar] [CrossRef] [PubMed]
- Tolle, F.; Brandle, G.M.; Matzner, D.; Mayer, G. A versatile approach towards nucleobase-modified aptamers. Angew. Chem. Int. Ed. Engl. 2015, 54, 10971–10974. [Google Scholar] [CrossRef] [PubMed]
- Tolle, F.; Rosenthal, M.; Pfeiffer, F.; Mayer, G. Click reaction on solid phase enables high fidelity synthesis of nucleobase-modified DNA. Bioconjug. Chem. 2016, 27, 500–503. [Google Scholar] [CrossRef] [PubMed]
- Wulff, G.; Liu, J.Q. Design of biomimetic catalysts by molecular imprinting in synthetic polymers: The role of transition state stabilization. Acc. Chem. Res. 2012, 45, 239–247. [Google Scholar] [CrossRef] [PubMed]
- Robinson, D.K.; Mosbach, K. Molecular imprinting of a transition-state analog leads to a polymer exhibiting esterolytic activity. J. Chem. Soc. Chem. Commun. 1989, 969–970. [Google Scholar] [CrossRef]
- Liu, J.Q.; Wulff, G. Molecularly imprinted polymers with strong carboxypeptidase a-like activity: Combination of an amidinium function with a zinc-ion binding site in transition-state imprinted cavities. Angew. Chem. Int. Ed. 2004, 43, 1287–1290. [Google Scholar] [CrossRef] [PubMed]
- Lakshmi, D.; Bossi, A.; Whitcombe, M.J.; Chianella, I.; Fowler, S.A.; Subrahmanyam, S.; Piletska, E.V.; Piletsky, S.A. Electrochemical sensor for catechol and dopamine based on a catalytic molecularly imprinted polymer-conducting polymer hybrid recognition element. Anal. Chem. 2009, 81, 3576–3584. [Google Scholar] [CrossRef] [PubMed]
- Gu, Y.; Yan, X.Y.; Li, C.; Zheng, B.; Li, Y.R.; Liu, W.L.; Zhang, Z.Q.; Yang, M. Biomimetic sensor based on molecularly imprinted polymer with nitroreductase-like activity for metronidazole detection. Biosens. Bioelectron. 2016, 77, 393–399. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Yin, Y.Z.; Liu, Y.; Bai, X.L.; Zhang, Z.M.; Xu, J.Y.; Shen, J.C.; Liu, J.Q. Incorporation of glutathione peroxidase active site into polymer based on imprinting strategy. Biosens. Bioelectron. 2009, 25, 657–660. [Google Scholar] [CrossRef] [PubMed]
- Diaz-Diaz, G.; Dineiro, Y.; Menendez, M.I.; Blanco-Lopez, M.C.; Lobo-Castanon, M.J.; Miranda-Ordieres, A.J.; Tunon-Blanco, P. Molecularly imprinted catalytic polymers with biomimetic chloroperoxidase activity. Polymer 2011, 52, 2468–2473. [Google Scholar] [CrossRef]
- Sode, K.; Ohta, S.; Yanai, Y.; Yamazaki, T. Construction of a molecular imprinting catalyst using target analogue template and its application for an amperometric fructosylamine sensor. Biosens. Bioelectron. 2003, 18, 1485–1490. [Google Scholar] [CrossRef]
- Zhang, J.; Lei, J.P.; Ju, H.X.; Wang, C.Y. Electrochemical sensor based on chlorohemin modified molecularly imprinted microgel for determination of 2,4-dichlorophenol. Anal. Chim. Acta 2013, 786, 16–21. [Google Scholar] [CrossRef] [PubMed]
- Antuna-Jimenez, D.; Blanco-Lopez, M.C.; Miranda-Ordieres, A.J.; Lobo-Castanon, M.J. Artificial enzyme with magnetic properties and peroxidase activity on indoleamine metabolite tumor marker. Polymer 2014, 55, 1113–1119. [Google Scholar] [CrossRef]
- Famulok, M.; Hartig, J.S.; Mayer, G. Functional aptamers and aptazymes in biotechnology, diagnostics, and therapy. Chem. Rev. 2007, 107, 3715–3743. [Google Scholar] [CrossRef] [PubMed]
- Tucker, B.J.; Breaker, R.R. Riboswitches as versatile gene control elements. Curr. Opin. Struct. Biol. 2005, 15, 342–348. [Google Scholar] [CrossRef] [PubMed]
- Vitreschak, A.G.; Rodionov, D.A.; Mironov, A.A.; Gelfand, M.S. Riboswitches: The oldest mechanism for the regulation of gene expression? Trends Genet. 2004, 20, 44–50. [Google Scholar] [CrossRef] [PubMed]
- Nudler, E.; Mironov, A.S. The riboswitch control of bacterial metabolism. Trends Biochem. Sci. 2004, 29, 11–17. [Google Scholar] [CrossRef] [PubMed]
- Bocobza, S.E.; Aharoni, A. Switching the light on plant riboswitches. Trends Plant Sci. 2008, 13, 526–533. [Google Scholar] [CrossRef] [PubMed]
- Borsuk, P.; Dzikowska, A.; Empel, J.; Grzelak, A.; Grzeskowiak, R.; Weglenski, P. Structure of the arginase coding gene and its transcript in aspergillus nidulans. Acta Biochim. Pol. 1999, 46, 391–403. [Google Scholar] [PubMed]
- Edelmann, C.M., Jr. Clinical quiz. Renal tubular acidosis (rta). Pediatr. Nephrol. 1989, 3, 111–112. [Google Scholar] [CrossRef] [PubMed]
- Burgstaller, P.; Jenne, A.; Blind, M. Aptamers and aptazymes: Accelerating small molecule drug discovery. Curr. Opin. Drug Discov. Dev. 2002, 5, 690–700. [Google Scholar]
- Famulok, M. Allosteric aptamers and aptazymes as probes for screening approaches. Curr. Opin. Mol. Ther. 2005, 7, 137–143. [Google Scholar] [PubMed]
- Fan, P.; Suri, A.K.; Fiala, R.; Live, D.; Patel, D.J. Molecular recognition in the fmn-rna aptamer complex. J. Mol. Biol. 1996, 258, 480–500. [Google Scholar] [CrossRef] [PubMed]
- Patel, D.J. Structural analysis of nucleic acid aptamers. Curr. Opin. Chem. Biol. 1997, 1, 32–46. [Google Scholar] [CrossRef]
- Mir, A.; Golden, B.L. Two active site divalent ions in the crystal structure of the hammerhead ribozyme bound to a transition state analogue. Biochemistry 2016, 55, 633–636. [Google Scholar] [CrossRef] [PubMed]
- Oberthur, D.; Achenbach, J.; Gabdulkhakov, A.; Buchner, K.; Maasch, C.; Falke, S.; Rehders, D.; Klussmann, S.; Betzel, C. Crystal structure of a mirror-image l-rna aptamer (spiegelmer) in complex with the natural l-protein target ccl2. Nat. Commun. 2015, 6, 6923. [Google Scholar] [CrossRef] [PubMed]
- Russo Krauss, I.; Merlino, A.; Giancola, C.; Randazzo, A.; Mazzarella, L.; Sica, F. Thrombin-aptamer recognition: A revealed ambiguity. Nucleic Acids Res. 2011, 39, 7858–7867. [Google Scholar] [CrossRef] [PubMed]
- Tang, J.; Breaker, R.R. Rational design of allosteric ribozymes. Chem. Biol. 1997, 4, 453–459. [Google Scholar] [CrossRef]
- Tang, J.; Breaker, R.R. Mechanism for allosteric inhibition of an atp-sensitive ribozyme. Nucleic Acids Res. 1998, 26, 4214–4221. [Google Scholar] [CrossRef] [PubMed]
- Scott, W.G.; Horan, L.H.; Martick, M. The hammerhead ribozyme: Structure, catalysis, and gene regulation. Prog. Mol. Biol. Transl. Sci. 2013, 120, 1–23. [Google Scholar] [PubMed]
- Araki, M.; Okuno, Y.; Hara, Y.; Sugiura, Y. Allosteric regulation of a ribozyme activity through ligand-induced conformational change. Nucleic Acids Res. 1998, 26, 3379–3384. [Google Scholar] [CrossRef] [PubMed]
- Soukup, G.A.; Breaker, R.R. Design of allosteric hammerhead ribozymes activated by ligand-induced structure stabilization. Structure 1999, 7, 783–791. [Google Scholar] [CrossRef]
- Srinivasan, J.; Cload, S.T.; Hamaguchi, N.; Kurz, J.; Keene, S.; Kurz, M.; Boomer, R.M.; Blanchard, J.; Epstein, D.; Wilson, C.; et al. Adp-specific sensors enable universal assay of protein kinase activity. Chem. Biol. 2004, 11, 499–508. [Google Scholar] [CrossRef] [PubMed]
- Hartig, J.S.; Famulok, M. Reporter ribozymes for real-time analysis of domain-specific interactions in biomolecules: Hiv-1 reverse transcriptase and the primer-template complex. Angew. Chem. Int. Ed. Engl. 2002, 41, 4263–4266. [Google Scholar] [CrossRef]
- Appel, B.; Marschall, T.; Strahl, A.; Muller, S. Kinetic characterization of hairpin ribozyme variants. Methods Mol. Biol. 2012, 848, 41–59. [Google Scholar] [PubMed]
- Najafi-Shoushtari, S.H.; Mayer, G.; Famulok, M. Sensing complex regulatory networks by conformationally controlled hairpin ribozymes. Nucleic Acids Res. 2004, 32, 3212–3219. [Google Scholar] [CrossRef] [PubMed]
- Kossen, K.; Vaish, N.K.; Jadhav, V.R.; Pasko, C.; Wang, H.; Jenison, R.; McSwiggen, J.A.; Polisky, B.; Seiwert, S.D. High-throughput ribozyme-based assays for detection of viral nucleic acids. Chem. Biol. 2004, 11, 807–815. [Google Scholar] [CrossRef] [PubMed]
- Robertson, M.P.; Ellington, A.D. Design and optimization of effector-activated ribozyme ligases. Nucleic Acids Res. 2000, 28, 1751–1759. [Google Scholar] [CrossRef] [PubMed]
- Helm, M.; Petermeier, M.; Ge, B.; Fiammengo, R.; Jaschke, A. Allosterically activated diels-alder catalysis by a ribozyme. J. Am. Chem. Soc. 2005, 127, 10492–10493. [Google Scholar] [CrossRef] [PubMed]
- Serganov, A.; Keiper, S.; Malinina, L.; Tereshko, V.; Skripkin, E.; Hobartner, C.; Polonskaia, A.; Phan, A.T.; Wombacher, R.; Micura, R.; et al. Structural basis for diels-alder ribozyme-catalyzed carbon-carbon bond formation. Nat. Struct. Mol. Biol. 2005, 12, 218–224. [Google Scholar] [CrossRef] [PubMed]
- Radi, A.-E.; Acero Sánchez, J.L.; Baldrich, E.; O’Sullivan, C.K. Reagentless, reusable, ultrasensitive electrochemical molecular beacon aptasensor. J. Am. Chem. Soc. 2006, 128, 117–124. [Google Scholar] [CrossRef] [PubMed]
- Cao, Z.; Tan, W. Molecular aptamers for real-time protein-protein interaction study. Chem.—Eur. J. 2005, 11, 4502–4508. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, R.; Kumar, P.K.R. Molecular beacon aptamer fluoresces in the presence of tat protein of HIV-1. Genes Cells 2000, 5, 389–396. [Google Scholar] [CrossRef] [PubMed]
- Nutiu, R.; Li, Y. Structure-switching signaling aptamers. J. Am. Chem. Soc. 2003, 125, 4771–4778. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Lubin, A.A.; Heeger, A.J.; Plaxco, K.W. Label-free electronic detection of thrombin in blood serum by using an aptamer-based sensor. Angew. Chem. Int. Ed. 2005, 44, 5456–5459. [Google Scholar] [CrossRef] [PubMed]
- Cho, E.J.; Lee, J.-W.; Ellington, A.D. Applications of aptamers as sensors. Annu. Rev. Anal. Chem. 2009, 2, 241–264. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Wang, Z.; Li, X.-F.; Le, X.C. Ultrasensitive detection of proteins by amplification of affinity aptamers. Angew. Chem. Int. Ed. 2006, 45, 1576–1580. [Google Scholar] [CrossRef] [PubMed]
- Fischer, N.O.; Tarasow, T.M.; Tok, J.B.H. Protein detection via direct enzymatic amplification of short DNA aptamers. Anal. Biochem. 2008, 373, 121–128. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Fung, C.W.; Cho, E.J.; Ellington, A.D. Real-time rolling circle amplification for protein detection. Anal. Chem. 2007, 79, 3320–3329. [Google Scholar] [CrossRef] [PubMed]
- Fredriksson, S.; Gullberg, M.; Jarvius, J.; Olsson, C.; Pietras, K.; Gustafsdottir, S.M.; Ostman, A.; Landegren, U. Protein detection using proximity-dependent DNA ligation assays. Nat. Biotech. 2002, 20, 473–477. [Google Scholar] [CrossRef] [PubMed]
- Hu, X.; Li, G.; Huang, J.; Zhang, D.; Qiu, Y. Construction of self-reporting specific chemical sensors with high sensitivity. Adv. Mater. 2007, 19, 4327–4332. [Google Scholar] [CrossRef]
- Yoshimi, Y.; Ohdaira, R.; Iiyama, C.; Sakai, K. “gate effect” of thin layer of molecularly-imprinted poly(methacrylic acid-co-ethyleneglycol dimethacrylate). Sens. Actuators B Chem. 2001, 73, 49–53. [Google Scholar] [CrossRef]
- Sartori, L.R.; Santos, W.D.R.; Kubota, L.T.; Segatelli, M.G.; Tarley, C.R.T. Flow-based method for epinephrine determination using a solid reactor based on molecularly imprinted poly(fepp-maa-egdma). Mater. Sci. Eng. C-Mater. Biol. Appl. 2011, 31, 114–119. [Google Scholar] [CrossRef]
- Reddy, S.M.; Sette, G.; Phan, Q. Electrochemical probing of selective haemoglobin binding in hydrogel-based molecularly imprinted polymers. Electrochim. Acta 2011, 56, 9203–9208. [Google Scholar] [CrossRef]
- Chen, A.; Yang, S. Replacing antibodies with aptamers in lateral flow immunoassay. Biosens. Bioelectron. 2015, 71, 230–242. [Google Scholar] [CrossRef] [PubMed]
- Zhou, W.; Huang, P.J.; Ding, J.; Liu, J. Aptamer-based biosensors for biomedical diagnostics. Analyst 2014, 139, 2627–2640. [Google Scholar] [CrossRef] [PubMed]
- Tretjakov, A.; Syritski, V.; Reut, J.; Boroznjak, R.; Opik, A. Molecularly imprinted polymer film interfaced with surface acoustic wave technology as a sensing platform for label-free protein detection. Anal. Chim. Acta 2016, 902, 182–188. [Google Scholar] [CrossRef] [PubMed]
- Lv, Y.; Qin, Y.; Svec, F.; Tan, T. Molecularly imprinted plasmonic nanosensor for selective sers detection of protein biomarkers. Biosens. Bioelectron. 2016, 80, 433–441. [Google Scholar] [CrossRef] [PubMed]
- Moreira, F.T.C.; Sharma, S.; Dutra, R.A.F.; Noronha, J.P.C.; Cass, A.E.G.; Sales, M.G.F. Protein-responsive polymers for point-of-care detection of cardiac biomarker. Sens. Actuators B-Chem. 2014, 196, 123–132. [Google Scholar] [CrossRef]
- Bosserdt, M.; Gajovic-Eichelman, N.; Scheller, F.W. Modulation of direct electron transfer of cytochrome c by use of a molecularly imprinted thin film. Anal. Bioanal. Chem. 2013, 405, 6437–6444. [Google Scholar] [CrossRef] [PubMed]
- Zhao, W.T.; Chen, Z.H.; Xue, B.; Sun, L.Q.; Luo, A.Q. A biomimetic sensor for fast lysozyme detection. Adv. Mater. Res. 2011, 239–242, 283–287. [Google Scholar] [CrossRef]
- Sefah, K.; Phillips, J.A.; Xiong, X.; Meng, L.; Van Simaeys, D.; Chen, H.; Martin, J.; Tan, W. Nucleic acid aptamers for biosensors and bio-analytical applications. Analyst 2009, 134, 1765–1775. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.Q.; He, X.W.; Wang, Y.Z.; Li, W.Y.; Zhang, Y.K. Epitope imprinted polymer coating cdte quantum dots for specific recognition and direct fluorescent quantification of the target protein bovine serum albumin. Biosens. Bioelectron. 2014, 54, 266–272. [Google Scholar] [CrossRef] [PubMed]
- Deng, Q.; Wu, J.; Zhai, X.; Fang, G.; Wang, S. Highly selective fluorescent sensing of proteins based on a fluorescent molecularly imprinted nanosensor. Sensors 2013, 13, 12994–13004. [Google Scholar] [CrossRef] [PubMed]
- Feng, X.B.; Gan, N.; Zhou, J.; Li, T.H.; Cao, Y.T.; Hu, F.T.; Yu, H.W.; Jiang, Q.L. A novel dual-template molecularly imprinted electrochemiluminescence immunosensor array using ru(bpy)(3)(2+)-silica@poly-l-lysine-au composite nanoparticles as labels for near-simultaneous detection of tumor markers. Electrochim. Acta 2014, 139, 127–136. [Google Scholar] [CrossRef]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Menger, M.; Yarman, A.; Erdőssy, J.; Yildiz, H.B.; Gyurcsányi, R.E.; Scheller, F.W. MIPs and Aptamers for Recognition of Proteins in Biomimetic Sensing. Biosensors 2016, 6, 35. https://doi.org/10.3390/bios6030035
Menger M, Yarman A, Erdőssy J, Yildiz HB, Gyurcsányi RE, Scheller FW. MIPs and Aptamers for Recognition of Proteins in Biomimetic Sensing. Biosensors. 2016; 6(3):35. https://doi.org/10.3390/bios6030035
Chicago/Turabian StyleMenger, Marcus, Aysu Yarman, Júlia Erdőssy, Huseyin Bekir Yildiz, Róbert E. Gyurcsányi, and Frieder W. Scheller. 2016. "MIPs and Aptamers for Recognition of Proteins in Biomimetic Sensing" Biosensors 6, no. 3: 35. https://doi.org/10.3390/bios6030035
APA StyleMenger, M., Yarman, A., Erdőssy, J., Yildiz, H. B., Gyurcsányi, R. E., & Scheller, F. W. (2016). MIPs and Aptamers for Recognition of Proteins in Biomimetic Sensing. Biosensors, 6(3), 35. https://doi.org/10.3390/bios6030035








