Engineered TtgR-Based Whole-Cell Biosensors for Quantitative and Selective Monitoring of Bioactive Compounds
Abstract
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. Construction of Plasmids and Escherichia coli Cell-Based Biosensors
2.3. Biosensor Assay
2.4. In Silico Docking Experiments
2.5. Application of Escherichia coli Cell-Based Biosensors in Quercetin and Resveratrol Quantification
2.6. Statistical Analysis
3. Results
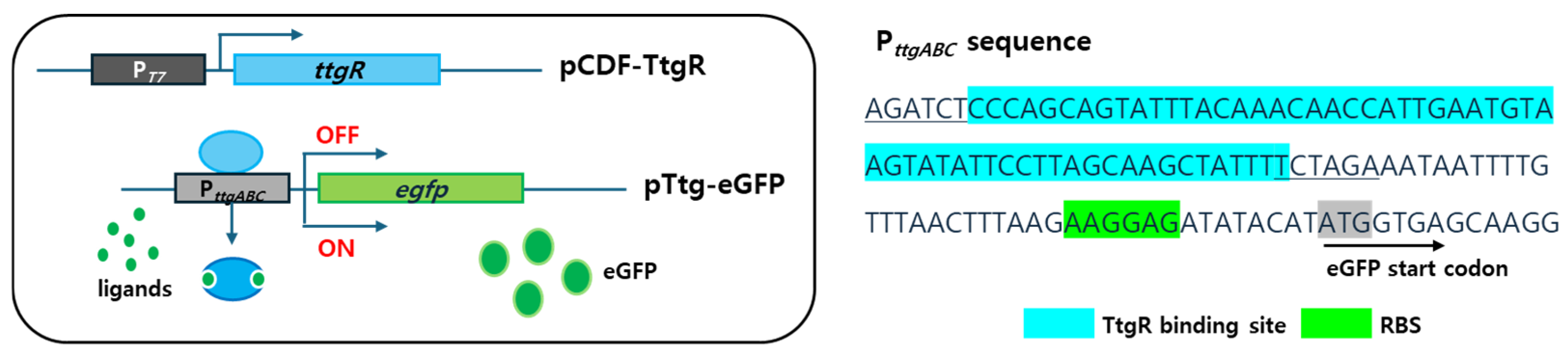
3.1. Sensing Mechanism of Escherichia coli Cell-Based Biosensors Employing the ttg Operon System
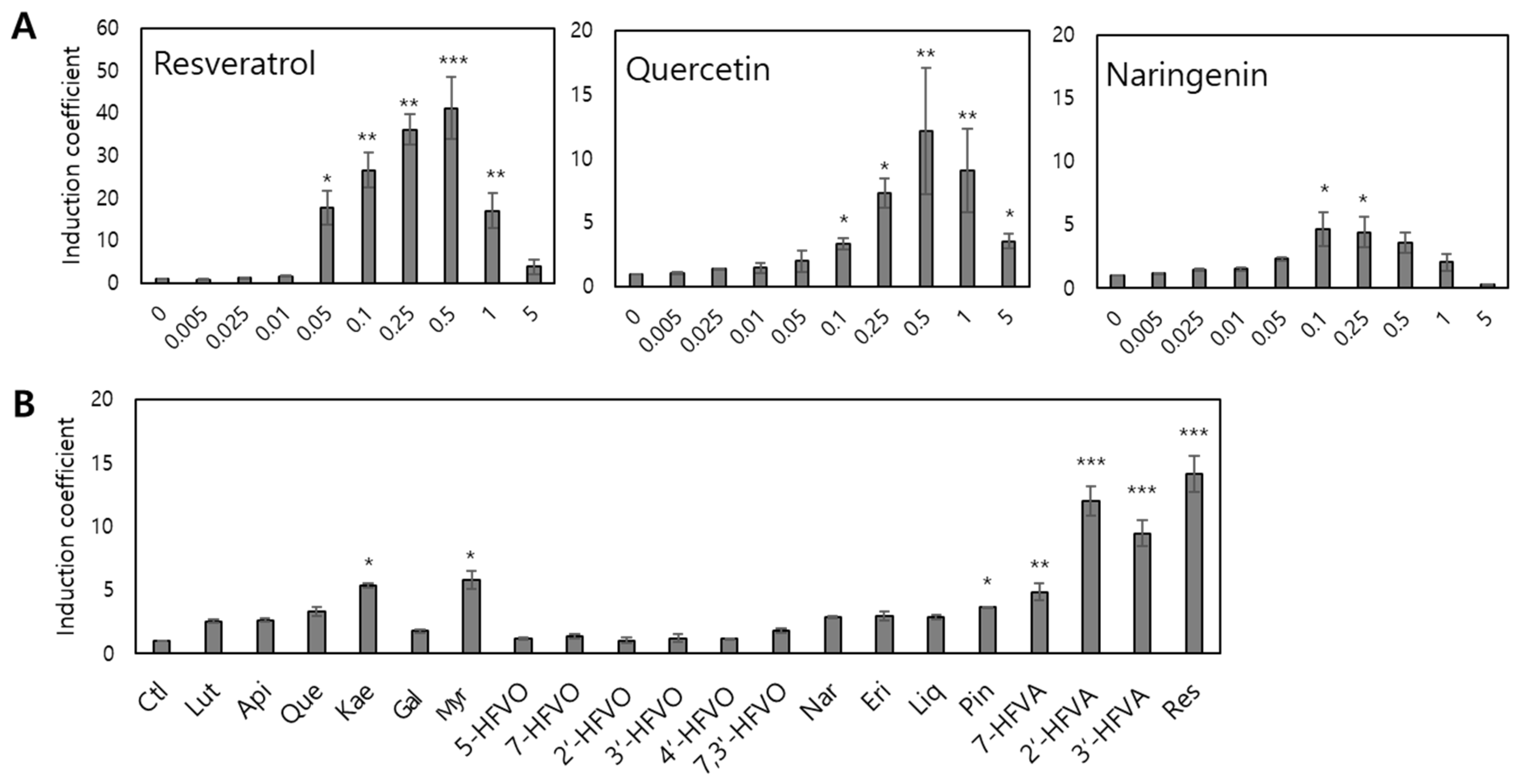
3.2. Characteristics of Escherichia coli Cell-Based Biosensors Employing the TtgR Wild-Type
3.3. Genetic Engineering of TtgR
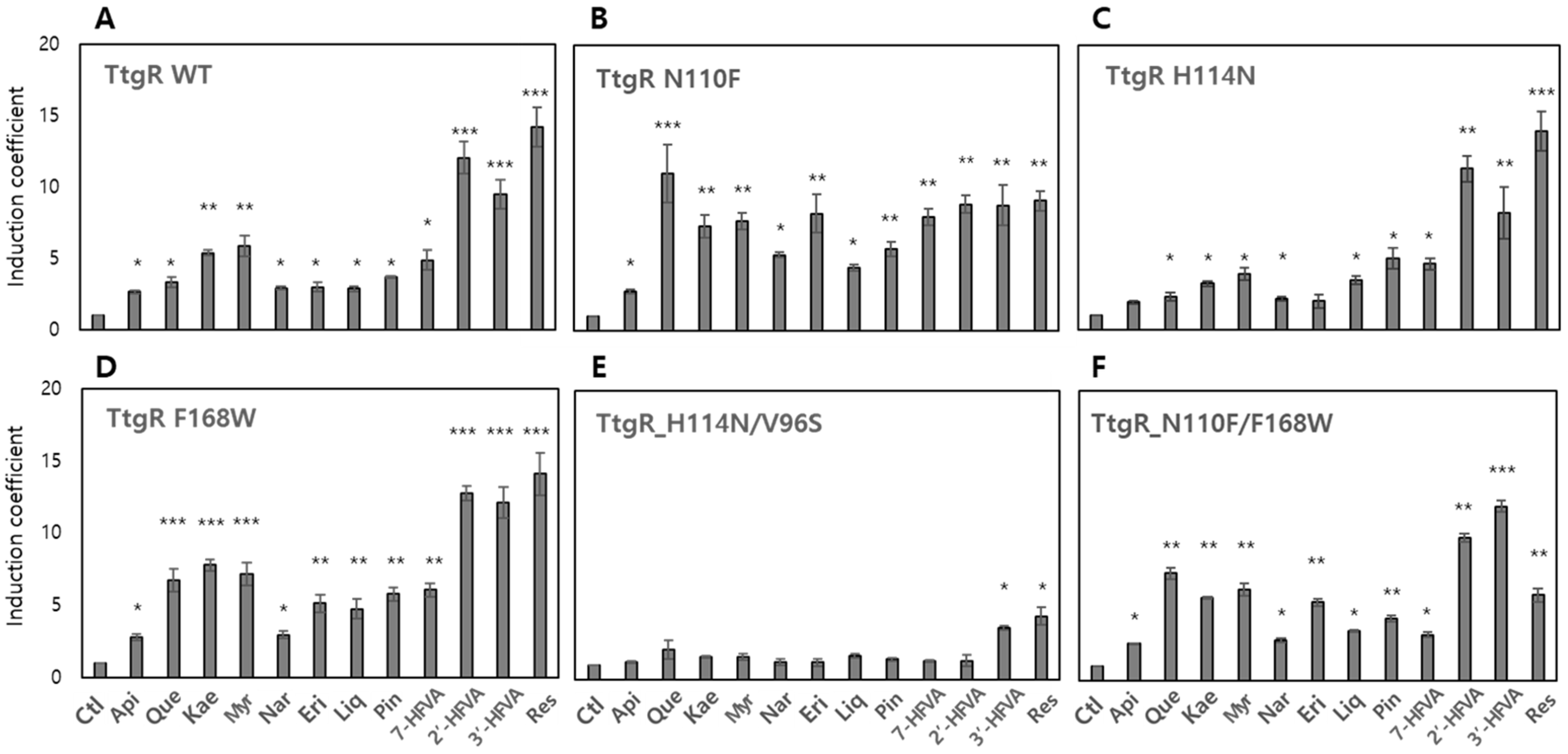
3.4. Effects of Engineered TtgRs on the Biosensors
3.4.1. Biosensor-TtgRs with Mutated Asn110
3.4.2. Biosensor-TtgRs with Mutated His114
3.4.3. Biosensor-TtgRs with Mutated Val96 and Ile141
3.4.4. Biosensor-TtgRs with Phe168 and Double Mutations
3.5. Computational Analysis of TtgR–Ligand Interactions
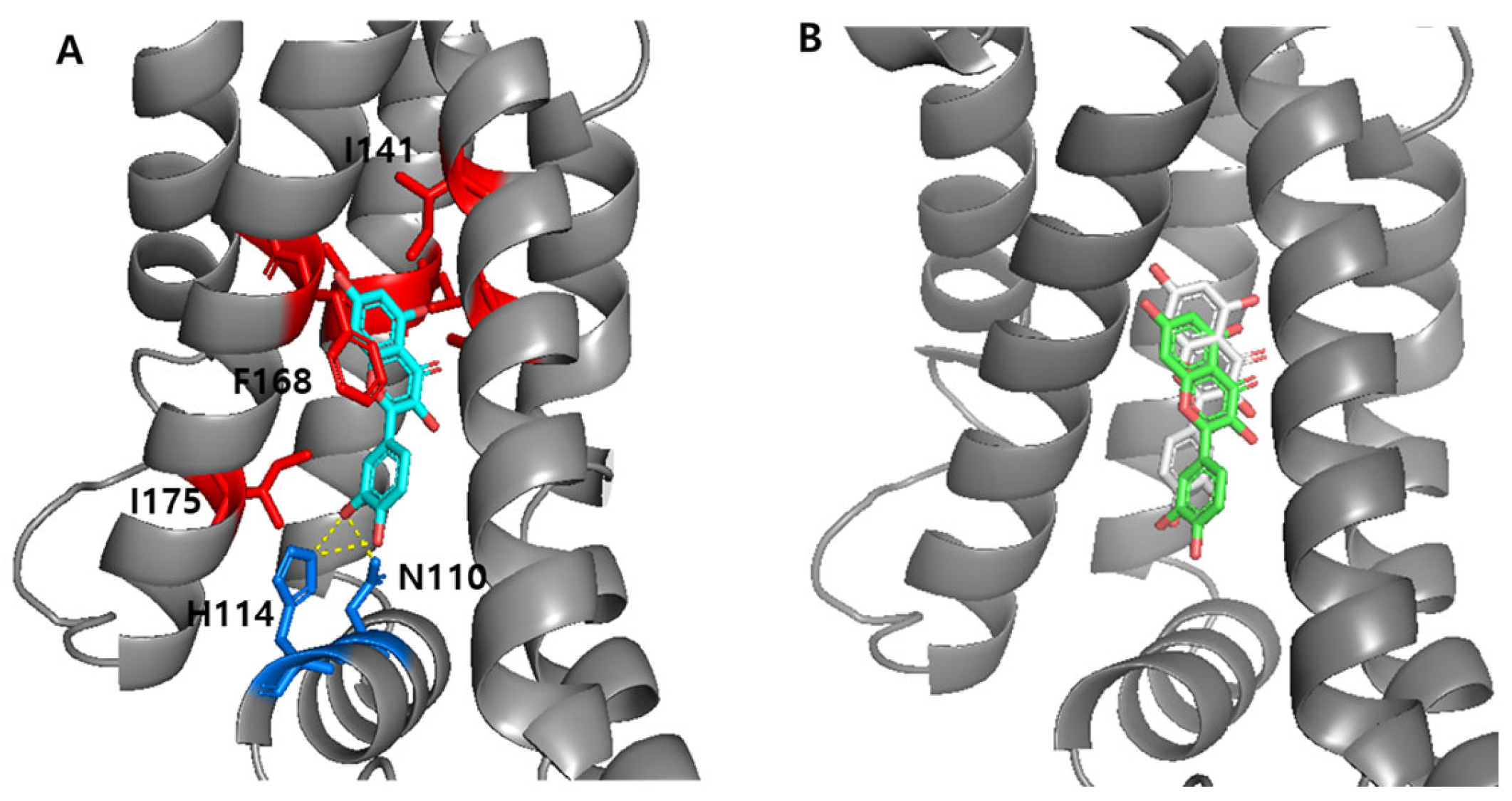
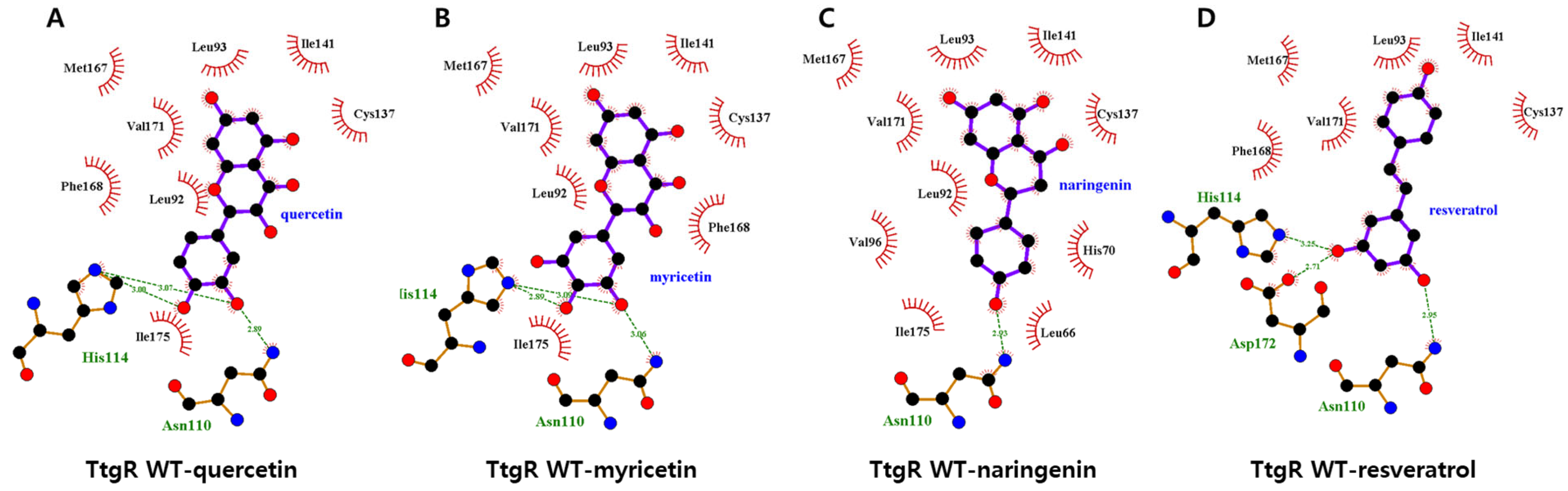
3.5.1. Interaction Between the TtgR Wild-Type and Ligands
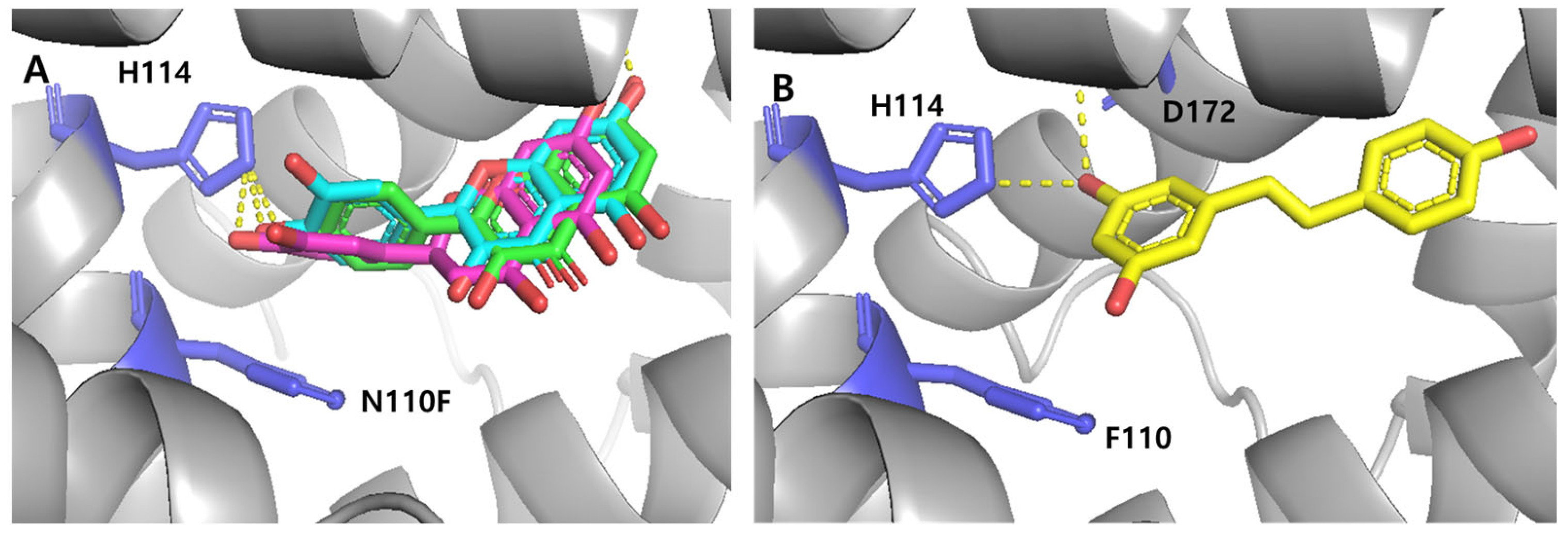
3.5.2. Engineered TtgRs with Asn110 Mutations
3.5.3. Engineered TtgRs with His114 and Phe168 Mutations
3.6. Quantification of Quercetin and Resveratrol Using the TtgR-Based Biosensors
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Sun, J.; Deng, Z.; Yan, A. Bacterial multidrug efflux pumps: Mechanisms, physiology and pharmacological exploitations. Biochem. Biophys. Res. Commun. 2014, 453, 254–267. [Google Scholar] [CrossRef] [PubMed]
- Daniels, C.; Daddaoua, A.; Lu, D.; Zhang, X.; Ramos, J.-L. Domain cross-talk during effector binding to the multidrug binding TTGR regulator. J. Biol. Chem. 2010, 285, 21372–21381. [Google Scholar] [CrossRef] [PubMed]
- Blanco, P.; Hernando-Amado, S.; Reales-Calderon, J.A.; Corona, F.; Lira, F.; Alcalde-Rico, M.; Bernardini, A.; Sanchez, M.B.; Martinez, J.L. Bacterial multidrug efflux pumps: Much more than antibiotic resistance determinants. Microorganisms 2016, 4, 14. [Google Scholar] [CrossRef] [PubMed]
- Grkovic, S.; Brown, M.H.; Roberts, N.J.; Paulsen, I.T.; Skurray, R.A. QacR is a repressor protein that regulates expression of theStaphylococcus aureus multidrug efflux pump QacA. J. Biol. Chem. 1998, 273, 18665–18673. [Google Scholar] [CrossRef]
- Terán, W.; Felipe, A.; Segura, A.; Rojas, A.; Ramos, J.-L.; Gallegos, M.A.-T. Antibiotic-dependent induction of Pseudomonas putida DOT-T1E TtgABC efflux pump is mediated by the drug binding repressor TtgR. Antimicrob. Agents Chemother. 2003, 47, 3067–3072. [Google Scholar] [CrossRef]
- Klyachko, K.A.; Schuldiner, S.; Neyfakh, A.A. Mutations affecting substrate specificity of the Bacillus subtilis multidrug transporter Bmr. J. Bacteriol. 1997, 179, 2189–2193. [Google Scholar] [CrossRef]
- Alguel, Y.; Meng, C.; Terán, W.; Krell, T.; Ramos, J.L.; Gallegos, M.-T.; Zhang, X. Crystal structures of multidrug binding protein TtgR in complex with antibiotics and plant antimicrobials. J. Mol. Biol. 2007, 369, 829–840. [Google Scholar] [CrossRef]
- Terán, W.; Krell, T.; Ramos, J.L.; Gallegos, M.-T. Effector-repressor interactions, binding of a single effector molecule to the operator-bound TtgR homodimer mediates derepression. J. Biol. Chem. 2006, 281, 7102–7109. [Google Scholar] [CrossRef]
- Wu, Y.; Zhang, S.; York, D.M.; Wang, L. Adsorption of Flavonoids in a Transcriptional Regulator TtgR: Relative Binding Free Energies and Intermolecular Interactions. J. Phys. Chem. B 2024, 128, 6529–6541. [Google Scholar] [CrossRef]
- Dias, M.C.; Pinto, D.C.; Silva, A.M. Plant flavonoids: Chemical characteristics and biological activity. Molecules 2021, 26, 5377. [Google Scholar] [CrossRef]
- Pourcel, L.; Routaboul, J.-M.; Cheynier, V.; Lepiniec, L.; Debeaujon, I. Flavonoid oxidation in plants: From biochemical properties to physiological functions. Trends Plant Sci. 2007, 12, 29–36. [Google Scholar] [CrossRef]
- Kumar, S.; Pandey, A.K. Chemistry and biological activities of flavonoids: An overview. Sci. World J. 2013, 2013, 162750. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Escamilla, A.M.; Fernandez-Ballester, G.; Morel, B.; Casares-Atienza, S.; Ramos, J.L. Molecular binding mechanism of TtgR repressor to antibiotics and antimicrobials. PLoS ONE 2015, 10, e0138469. [Google Scholar] [CrossRef]
- Paczkowski, J.E.; Mukherjee, S.; McCready, A.R.; Cong, J.-P.; Aquino, C.J.; Kim, H.; Henke, B.R.; Smith, C.D.; Bassler, B.L. Flavonoids suppress Pseudomonas aeruginosa virulence through allosteric inhibition of quorum-sensing receptors. J. Biol. Chem. 2017, 292, 4064–4076. [Google Scholar] [CrossRef]
- Espinosa-Urgel, M.; Serrano, L.; Ramos, J.L.; Fernández-Escamilla, A.M. Engineering biological approaches for detection of toxic compounds: A new microbial biosensor based on the Pseudomonas putida TtgR repressor. Mol. Biotechnol. 2015, 57, 558–564. [Google Scholar] [CrossRef]
- Rogers, J.K.; Guzman, C.D.; Taylor, N.D.; Raman, S.; Anderson, K.; Church, G.M. Synthetic biosensors for precise gene control and real-time monitoring of metabolites. Nucleic Acids Res. 2015, 43, 7648–7660. [Google Scholar] [CrossRef]
- Chen, W.; Zhang, X.; Xiong, D.; Jin, J.-M.; Tang, S.-Y. Engineering the effector specificity of regulatory proteins for the in vitro detection of biomarkers and pesticide residues. Appl. Microbiol. Biotechnol. 2019, 103, 3205–3213. [Google Scholar] [CrossRef] [PubMed]
- Ding, N.; Zhou, S.; Deng, Y. Transcription-factor-based biosensor engineering for applications in synthetic biology. ACS Synth. Biol. 2021, 10, 911–922. [Google Scholar] [CrossRef]
- Pham, C.; Stogios, P.J.; Savchenko, A.; Mahadevan, R. Advances in engineering and optimization of transcription factor-based biosensors for plug-and-play small molecule detection. Curr. Opin. Biotechnol. 2022, 76, 102753. [Google Scholar] [CrossRef]
- Jeon, Y.; Lee, Y.; Jang, G.; Kim, B.-G.; Yoon, Y. Design of Pb (II)-Specific E. coli-based biosensors by engineering regulatory proteins and host cells. Front. Microbiol. 2022, 13, 881050. [Google Scholar] [CrossRef]
- Ghataora, J.S.; Gebhard, S.; Reeksting, B.J. Chimeric MerR-family regulators and logic elements for the design of metal sensitive genetic circuits in Bacillus subtilis. ACS Synth. Biol. 2023, 12, 735–749. [Google Scholar] [CrossRef]
- Li, M.; Chen, Z.; Huo, Y.-X. Application evaluation and performance-directed improvement of the native and engineered biosensors. ACS Sens. 2024, 9, 5002–5024. [Google Scholar] [CrossRef]
- Zhou, G.J.; Zhang, F. Applications and tuning strategies for transcription factor-based metabolite biosensors. Biosensors 2023, 13, 428. [Google Scholar] [CrossRef] [PubMed]
- de Carvalho Matias, E.; Bezerra, K.; Costa, A.L.; Clemente Junior, W.; Oliveira, J.; Ribeiro Junior, L.; Galvão, D.; Fulco, U. Quantum biochemical analysis of the TtgR regulator and effectors. Sci. Rep. 2024, 14, 8519. [Google Scholar] [CrossRef] [PubMed]
- Yoon, Y.; Kim, S.; Chae, Y.; Kim, S.W.; Kang, Y.; An, G.; Jeong, S.-W.; An, Y.-J. Simultaneous detection of bioavailable arsenic and cadmium in contaminated soils using dual-sensing bioreporters. Appl. Microbiol. Biotechnol. 2016, 100, 3713–3722. [Google Scholar] [CrossRef]
- DeLano, W.L.; Bromberg, S. PyMOL User’s Guide; DeLano Scientific LLC: San Francisco, CA, USA, 2004; p. 629. [Google Scholar]
- Morris, G.M.; Huey, R.; Lindstrom, W.; Sanner, M.F.; Belew, R.K.; Goodsell, D.S.; Olson, A.J. AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. J. Comput. Chem. 2009, 30, 2785–2791. [Google Scholar] [CrossRef]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera—A visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef] [PubMed]
- Wallace, A.C.; Laskowski, R.A.; Thornton, J.M. LIGPLOT: A program to generate schematic diagrams of protein-ligand interactions. Protein Eng. Des. Sel. 1995, 8, 127–134. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2013; Available online: https://www.gbif.org/tool/81287/r-a-language-and-environment-for-statistical-computing (accessed on 24 June 2025).
- Signorell, A.; Aho, K.; Alfons, A.; Anderegg, N.; Aragon, T.; Arppe, A.; Baddeley, A.; Barton, K.; Bolker, B.; Borchers, H.W. DescTools: Tools for Descriptive Statistics; R Package Version 0.99; R Core Team: Vienna, Austria, 2019; Volume 28, p. 47. [Google Scholar]
- Nishikawa, K.K.; Hoppe, N.; Smith, R.; Bingman, C.; Raman, S. Epistasis shapes the fitness landscape of an allosteric specificity switch. Nat. Commun. 2021, 12, 5562. [Google Scholar] [CrossRef]
- Song, K.; Ji, H.; Lee, J.; Yoon, Y. Microbial Transcription Factor-Based Biosensors: Innovations from Design to Applications in Synthetic Biology. Biosensors 2025, 15, 221. [Google Scholar] [CrossRef] [PubMed]
- Ding, N.; Yuan, Z.; Zhang, X.; Chen, J.; Zhou, S.; Deng, Y. Programmable cross-ribosome-binding sites to fine-tune the dynamic range of transcription factor-based biosensor. Nucleic Acids Res. 2020, 48, 10602–10613. [Google Scholar] [CrossRef]
- Akher, F.B.; Ebrahimi, A.; Mostafavi, N. Characterization of π-stacking interactions between aromatic amino acids and quercetagetin. J. Mol. Struct. 2017, 1128, 13–20. [Google Scholar] [CrossRef]
- Shahidi, F.; Dissanayaka, C.S. Phenolic-protein interactions: Insight from in-silico analyses–a review. Food Prod. Process. Nutr. 2023, 5, 2. [Google Scholar] [CrossRef]
- Siedler, S.; Stahlhut, S.G.; Malla, S.; Maury, J.; Neves, A.R. Novel biosensors based on flavonoid-responsive transcriptional regulators introduced into Escherichia coli. Metab. Eng. 2014, 21, 2–8. [Google Scholar] [CrossRef]
- Lee, W.; Kim, H.; Jang, G.; Kim, B.-G.; Yoon, Y. Antimony sensing whole-cell bioreporters derived from ArsR genetic engineering. Appl. Microbiol. Biotechnol. 2020, 104, 2691–2699. [Google Scholar] [CrossRef]
- Kim, Y.; Jeon, Y.; Jang, G.; Kim, B.-G.; Yoon, Y. A novel Escherichia coli cell–based bioreporter for quantification of salicylic acid in cosmetics. Appl. Microbiol. Biotechnol. 2024, 108, 148. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Jang, G.; Kim, B.-G.; Yoon, Y. Modulation of the metal (loid) specificity of whole-cell bioreporters by genetic engineering of ZntR metal-binding loops. J. Microbiol. Biotechnol. 2020, 30, 681. [Google Scholar] [CrossRef]
- Kim, N.M.; Sinnott, R.W.; Sandoval, N.R. Transcription factor-based biosensors and inducible systems in non-model bacteria: Current progress and future directions. Curr. Opin. Biotechnol. 2020, 64, 39–46. [Google Scholar] [CrossRef] [PubMed]

| Name | Description | Reference | |
|---|---|---|---|
| Plasmids | pET-21a(+) | pBR322 ori, Amp r | Novagen |
| pCDF-Duet | CloDE13 ori, Str r | Novagen | |
| pTtg-eGFP | pET-21a(+) carrying PttgABC:egfp | This study | |
| pCDF-TtgR WT | pCDF-Duet carrying ttgR WT | This study | |
| pCDF-TtgR mut | pCDF-Duet carrying ttgR mutants: N110F, N110L, N110Y, H114N, H114A, V96S, I141S, I141L, F168W, N110Y/F168W, V96S/H114N | This study | |
| E. coli strains | E. coli BL21 (DE3) | F− ompT hsdSB(rB−mB−)gal dcm lon (DE3) | Stratagene |
| biosensor-TtgR WT | E. coli BL21 carrying pCDF-TtgR WT and pTtg-eGFP | This study | |
| biosensor-TtgR muts | E. coli BL21 carrying pCDF-TtgR muts and pTtg-eGFP | This study |
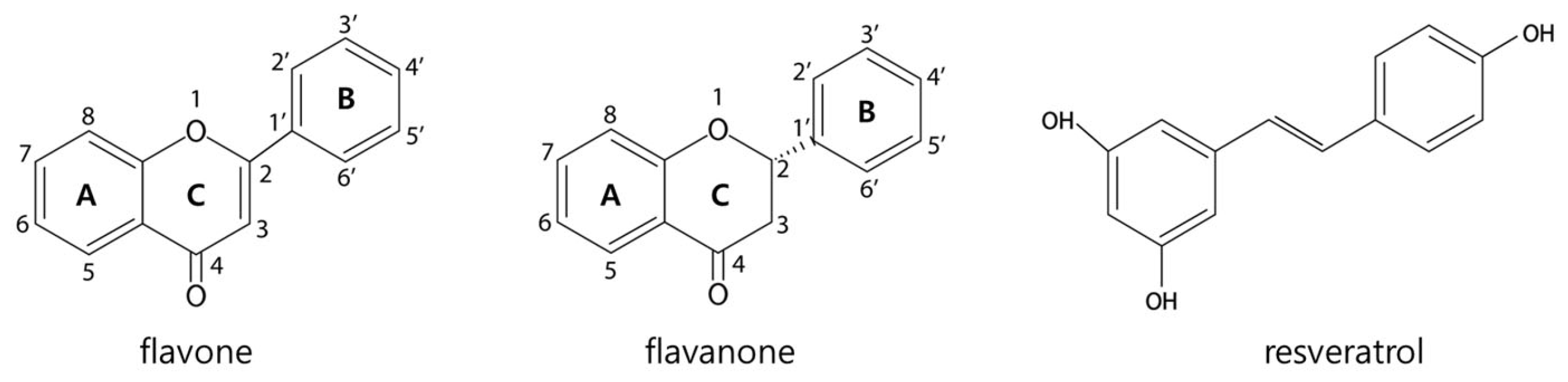
| Abbreviation | Full Name | 3 | 5 | 6 | 7 | 8 | 2′ | 3′ | 4′ | 5′ | 6′ | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Flavone | Lut | Luteolin | H | OH | H | OH | H | H | OH | OH | H | H |
| Api | Apigenin | H | OH | H | OH | H | H | H | OH | H | H | |
| Que | Quercetin | OH | OH | H | OH | H | H | OH | OH | H | H | |
| Kae | Kaempferol | OH | OH | H | OH | H | H | H | OH | H | H | |
| Myr | Myricetin | OH | OH | H | OH | H | H | OH | OH | OH | H | |
| Gal | Galangin | OH | OH | H | OH | H | H | H | H | H | H | |
| 4′-HFV | 4′-hydroxyflavone | H | H | H | H | H | H | H | OH | H | H | |
| 3′-HFV | 3′-hydroxyflavone | H | H | H | H | H | H | OH | H | H | H | |
| 2′-HFV | 2′-hydroxyflavone | H | H | H | H | H | OH | H | H | H | H | |
| 5-HFV | 5-hydroxyflavone | H | OH | H | H | H | H | H | H | H | H | |
| 7-HFV | 7-hydroxyflavone | H | H | H | OH | H | H | H | H | H | H | |
| 7,3′-DHFV | 7,3′-dihydroxyflavone | H | H | H | OH | H | H | OH | H | H | H | |
| Flavanone | Nar | Naringenin | H | OH | H | OH | H | H | H | OH | H | H |
| Eri | Eriodictyol | H | OH | H | OH | H | H | OH | OH | H | H | |
| Liq | Liquiritigenin | H | H | H | OH | H | H | H | OH | H | H | |
| Pin | Pinocembrin | H | OH | H | OH | H | H | H | H | H | H | |
| 3′-HFVA | 3′-hydroxyflavanone | H | H | H | H | H | H | OH | H | H | H | |
| 2′-HFVA | 2′-hydroxyflavanone | H | H | H | H | H | OH | H | H | H | H | |
| 7-HFVA | 7-hydroxyflavanone | H | H | H | OH | H | H | H | H | H | H | |
| Stilbene | Res | Resveratrol |
| Methods | Samples | Tested Conc. (mM) | Measured Conc. (mM) | Accuracy (%) |
|---|---|---|---|---|
| HPLC | Resveratrol | 10 | 10.12 ± 0.59 | 98.8 |
| 15 | 15.34 ± 0.48 | 97.8 | ||
| Quercetin | 10 | 9.8 ± 0.20 | 98.0 | |
| 15 | 15.62 ± 0.93 | 96.0 | ||
| Biosensor-TtgR WT | Resveratrol | 10 | 9.55 ± 0.64 | 95.5 |
| 15 | 14.24 ± 0.86 | 94.9 | ||
| Biosensor-TtgR N110F | Quercetin | 10 | 9.36 ± 0.25 | 93.6 |
| 15 | 14.42 ± 0.52 | 96.1 |
| TtgRs | I.C of Main Ligands | Detection Limit (mM) | Basal eGFP Signal | Proposed Mechanistic Reasons |
|---|---|---|---|---|
| WT | Resveratrol (12.4) | 0.0037 | Low | Native binding pocket optimized for resveratrol; strong DNA-binding repression |
| 2′-HFVA (10.9) | ||||
| Quercetin (2.9) | ||||
| N110F | Quercetin (11.0) | 0.0063 | Low–Moderate | Aromatic Phe substitution enhances π–π stacking with quercetin; increases conformational flexibility |
| Resveratrol (9.1) | ||||
| 2′-HFVA (8.8) | ||||
| N110Y | Resveratrol (9.3) | - | Moderate | Bulkier Tyr may improve aromatic interactions but also cause steric hindrance |
| 3′-HFVA (8.2) | ||||
| Quercetin (2.1) | ||||
| H114N/V96S | Resveratrol (4.4) | - | Moderate | Loss of aromaticity reduces π–π interactions; possible altered pocket shape |
| 3′-HFVA (3.6) | ||||
| Quercetin (2.1) | ||||
| N110L | 2′-HFVA (8.5) | - | Low | Hydrophobic Leu substitution maintains repression; minimal effect on ligand accommodation |
| Resveratrol (7.2) | ||||
| Quercetin (3.1) | ||||
| N110F/F168W | 2′-HFVA (12.0) | - | Low | Similar polarity to WT; minor structural change |
| Quercetin (7.4) | ||||
| Resveratrol (5.9) | ||||
| H114A | Resveratrol (7.0) | - | Moderate–High | Removal of imidazole side chain disrupts hydrogen bonding |
| 2′-HFVA (4.0) | ||||
| Quercetin (3.1) | ||||
| H114N | Resveratrol (11.8) | - | Moderate | Maintains hydrogen bonding potential but alters geometry |
| 2′-HFVA (8.3) | ||||
| Quercetin (4.6) | ||||
| V96F | Resveratrol (2.2) | - | Moderate | Bulky aromatic substitution changes hydrophobic packing near ligand site |
| 2′-HFVA (2.0) | ||||
| Quercetin (1.6) | ||||
| V96S | Resveratrol (5.8) | - | Moderate–High | Polar Ser alters hydrophobic microenvironment; possible reduced ligand affinity |
| 2′-HFVA (3.7) | ||||
| Quercetin (2.1) | ||||
| I141S | Resveratrol (6.8) | - | High | Polar substitution at hydrophobic site destabilizes closed conformation |
| Myricetin (5.5) | ||||
| Quercetin (3.2) | ||||
| F168W | Resveratrol (14.1) | - | Low–Moderate | Larger aromatic Trp modifies π–π stacking geometry and steric profile |
| 2′-HFVA (12.8) | ||||
| Quercetin (6.7) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Song, K.; Ji, H.; Lee, J.; Jang, G.; Yoon, Y. Engineered TtgR-Based Whole-Cell Biosensors for Quantitative and Selective Monitoring of Bioactive Compounds. Biosensors 2025, 15, 554. https://doi.org/10.3390/bios15080554
Song K, Ji H, Lee J, Jang G, Yoon Y. Engineered TtgR-Based Whole-Cell Biosensors for Quantitative and Selective Monitoring of Bioactive Compounds. Biosensors. 2025; 15(8):554. https://doi.org/10.3390/bios15080554
Chicago/Turabian StyleSong, Kyeongseok, Haekang Ji, Jiwon Lee, Geupil Jang, and Youngdae Yoon. 2025. "Engineered TtgR-Based Whole-Cell Biosensors for Quantitative and Selective Monitoring of Bioactive Compounds" Biosensors 15, no. 8: 554. https://doi.org/10.3390/bios15080554
APA StyleSong, K., Ji, H., Lee, J., Jang, G., & Yoon, Y. (2025). Engineered TtgR-Based Whole-Cell Biosensors for Quantitative and Selective Monitoring of Bioactive Compounds. Biosensors, 15(8), 554. https://doi.org/10.3390/bios15080554





