Paper-Based Biosensors for the Detection of Nucleic Acids from Pathogens
Abstract
1. Introduction
2. Detection Method
2.1. Structure-Recognition Biosensors
2.1.1. Aptamer-Based Sensors
2.1.2. Nucleic Acid-Modified Nanostructures
2.1.3. Nucleic Acid Immunoassays
2.1.4. Nucleic Acid Hybridization Sensors
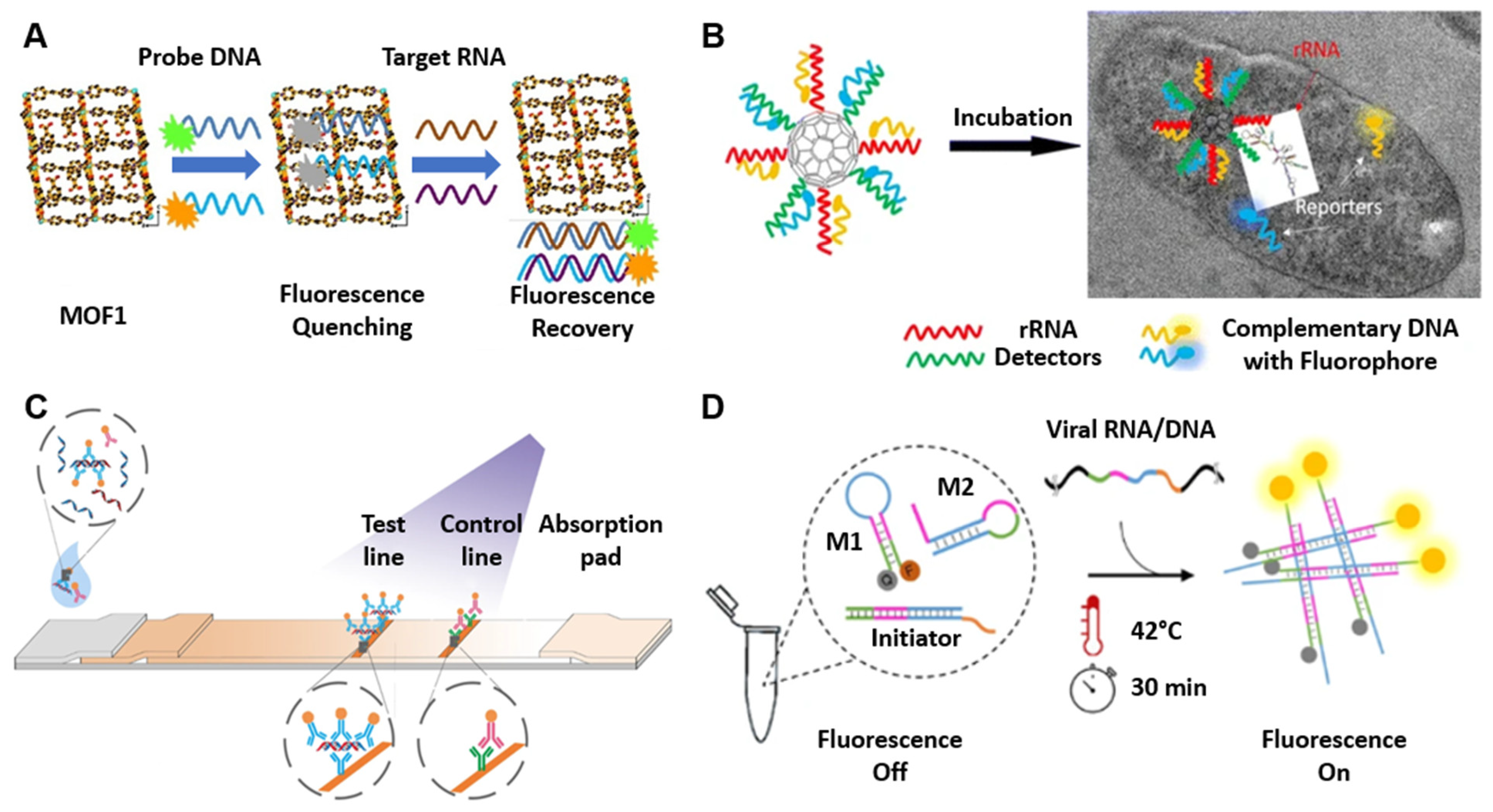
2.2. Oligonucleotide-Guided Biosensors
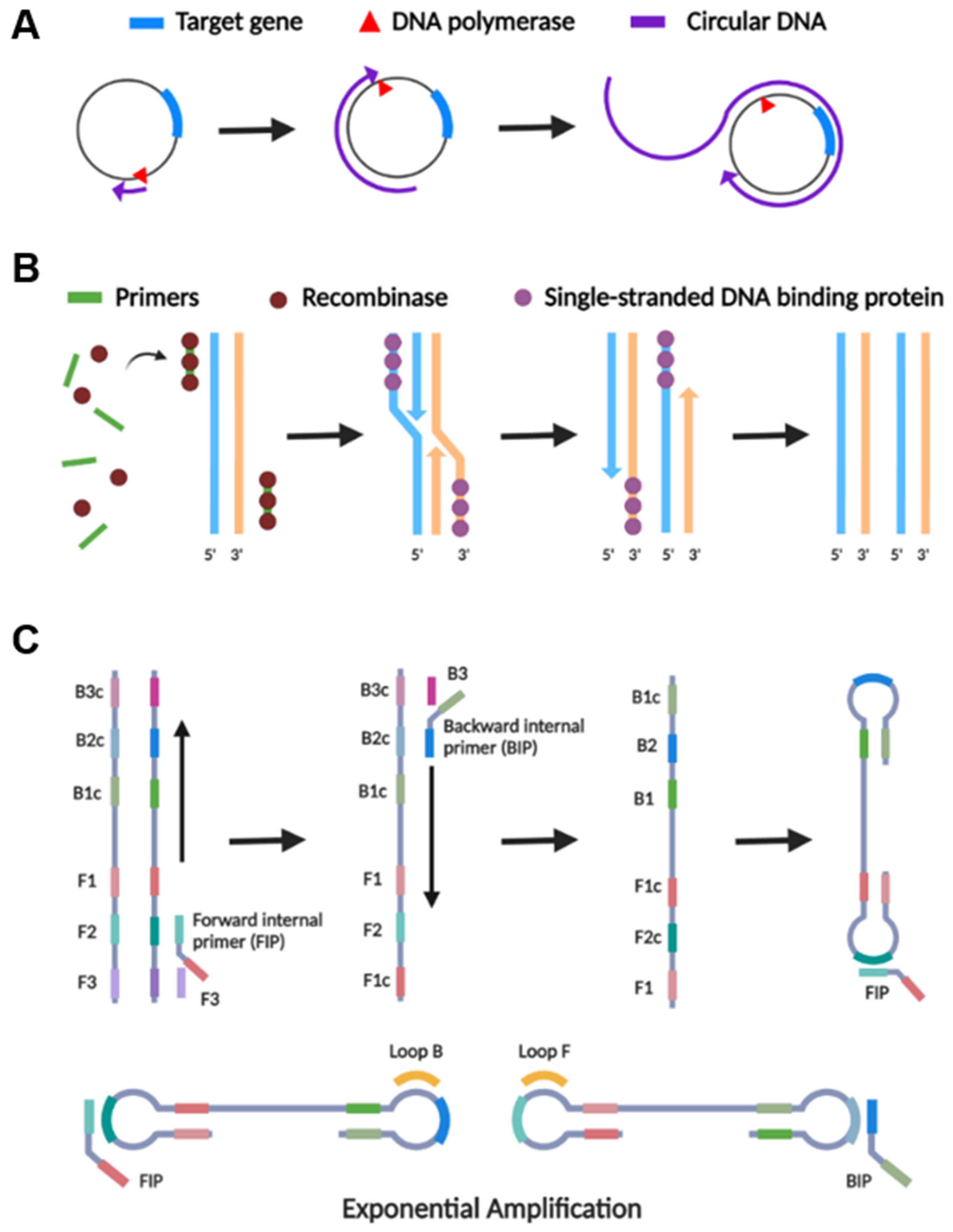
2.2.1. Rolling Circle Amplification (RCA)
2.2.2. Recombinase Polymerase Amplification (RPA)
2.2.3. Loop-Mediated Isothermal Amplification (LAMP)
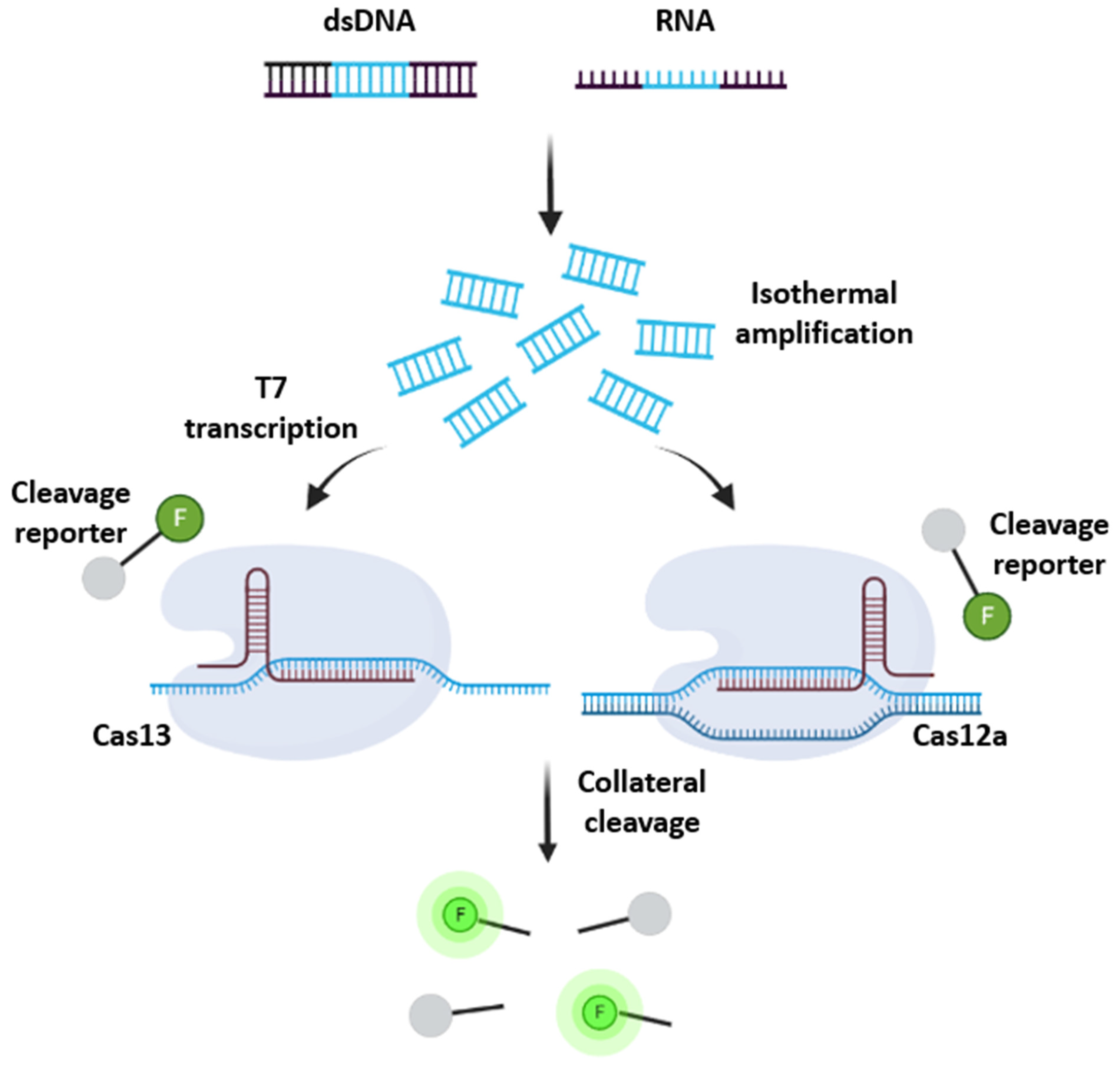
2.2.4. CRISPR/Cas
2.3. Critical Thinking in Choosing Detection Methods
3. Reporting Method
3.1. Electrochemical
3.2. Fluorometric
3.2.1. Fluorescent Dyes
3.2.2. Nanoparticles
3.2.3. Fluorescent Proteins/Organic Molecules
3.3. Colorimetric
3.3.1. pH
| Colorimetric Reporters | Mechanism | Color Change (Negative-Positive) | LoD (Copies a or CFU b per Reaction) | References |
|---|---|---|---|---|
| Phenol red | pH | Red-Yellow | 50–1000 a | [83,89,164] |
| Phenolphthalein | pH | Pink-Colorless | 1.225 b | [165] |
| Chemosensor-L | pH | Yellow-Colorless | 0.5 b | [166,167] |
| MColorpHast | pH | Green/brown-Yellow | 3.36 × 104 a | [168] |
| Crystal violet | DNA-binding | Colorless-Purple | 4413 a | [169] |
| Methylene blue | DNA-binding | Colorless-Blue | 25 b | [170] |
| Fuchsin | DNA-binding | Colorless-Purple | 1–300 b | [171,172] |
| Gold nanoparticles | Aggregation | Colorless-Red | 1.806 × 105–1.806 × 109 a | [173,175] |
| Silver nanoparticles | Aggregation | Colorless-Yellow | 1.55 × 1010–2.30 × 1010 a | [176] |
| Polystyrene latex microspheres | Aggregation | Colorless-Red/ Colorless-Blue | 1.806 × 1011 a | [177,178] |
| Hydroxynaphthol blue | Mg2+ depletion | Violet-Blue | 2.07 × 104 a | [179] |
3.3.2. DNA Binding
3.3.3. Aggregation
3.3.4. Mg2+ Depletion
3.4. Critical Thinking in Choosing Reporting Methods
4. Device Formats
4.1. Lateral Flow Assays (LFAs)
4.2. Microfluidic Paper-Based Analytical Devices (µPADs)
4.3. Integrated Design of Paper-Based Biosensors
5. Conclusions and Prospects
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Abayasekara, L.M.; Perera, J.; Chandrasekharan, V.; Gnanam, V.S.; Udunuwara, N.A.; Liyanage, D.S.; Bulathsinhala, N.E.; Adikary, S.; Aluthmuhandiram, J.V.S.; Thanaseelan, C.S.; et al. Detection of Bacterial Pathogens from Clinical Specimens Using Conventional Microbial Culture and 16S Metagenomics: A Comparative Study. BMC Infect. Dis. 2017, 17, 631. [Google Scholar] [CrossRef]
- Kok, J.; Blyth, C.C.; Foo, H.; Patterson, J.; Taylor, J.; McPhie, K.; Ratnamohan, V.M.; Iredell, J.R.; Dwyer, D.E. Comparison of a Rapid Antigen Test with Nucleic Acid Testing during Cocirculation of Pandemic Influenza A/H1N1 2009 and Seasonal Influenza A/H3N2. J. Clin. Microbiol. 2010, 48, 290–291. [Google Scholar] [CrossRef]
- Craw, P.; Balachandran, W. Isothermal Nucleic Acid Amplification Technologies for Point-of-Care Diagnostics: A Critical Review. Lab Chip 2012, 12, 2469–2486. [Google Scholar] [CrossRef]
- Morozumi, M.; Chiba, N.; Igarashi, Y.; Mitsuhashi, N.; Wajima, T.; Iwata, S.; Ubukata, K. Direct Identification of Streptococcus Agalactiae and Capsular Type by Real-Time PCR in Vaginal Swabs from Pregnant Women. J. Infect. Chemother. 2015, 21, 34–38. [Google Scholar] [CrossRef]
- Park, J.S.; Cho, D.-H.; Yang, J.H.; Kim, M.Y.; Shin, S.M.; Kim, E.-C.; Park, S.S.; Seong, M.-W. Usefulness of a Rapid Real-Time PCR Assay in Prenatal Screening for Group B Streptococcus Colonization. Ann. Lab. Med. 2012, 33, 39–44. [Google Scholar] [CrossRef]
- Guiducci, S.; Moriondo, M.; Nieddu, F.; Ricci, S.; De Vitis, E.; Casini, A.; Poggi, G.M.; Indolfi, G.; Resti, M.; Azzari, C. Culture and Real-Time Polymerase Chain Reaction Sensitivity in the Diagnosis of Invasive Meningococcal Disease: Does Culture Miss Less Severe Cases? PLoS ONE 2019, 14, e0212922. [Google Scholar] [CrossRef]
- CDC. Overview of Testing for SARS-CoV-2, the Virus That Causes COVID-19. Available online: https://www.cdc.gov/coronavirus/2019-ncov/hcp/testing-overview.html (accessed on 20 February 2022).
- Kung, C.-T.; Hou, C.-Y.; Wang, Y.-N.; Fu, L.-M. Microfluidic Paper-Based Analytical Devices for Environmental Analysis of Soil, Air, Ecology and River Water. Sens. Actuators B Chem. 2019, 301, 126855. [Google Scholar] [CrossRef]
- Fu, L.-M.; Wang, Y.-N. Detection Methods and Applications of Microfluidic Paper-Based Analytical Devices. TrAC Trends Anal. Chem. 2018, 107, 196–211. [Google Scholar] [CrossRef]
- Qi, Y.; Ma, J.; Chen, X.; Xiu, F.-R.; Chen, Y.; Lu, Y. Practical Aptamer-Based Assay of Heavy Metal Mercury Ion in Contaminated Environmental Samples: Convenience and Sensitivity. Anal. Bioanal. Chem. 2020, 412, 439–448. [Google Scholar] [CrossRef]
- Wang, Y.; Lv, M.; Chen, Z.; Deng, Z.; Liu, N.; Fan, J.; Zhang, W. A Fluorescence Resonance Energy Transfer Probe Based on DNA-Modified Upconversion and Gold Nanoparticles for Detection of Lead Ions. Front. Chem. 2020, 8, 238. [Google Scholar] [CrossRef]
- Chang, Z.; Zhu, B.; Liu, J.; Zhu, X.; Xu, M.; Travas-Sejdic, J. Electrochemical Aptasensor for 17β-Estradiol Using Disposable Laser Scribed Graphene Electrodes. Biosens. Bioelectron. 2021, 185, 113247. [Google Scholar] [CrossRef]
- Alnajrani, M.N.; Alsager, O.A. Lateral Flow Aptasensor for Progesterone: Competitive Target Recognition and Displacement of Short Complementary Sequences. Anal. Biochem. 2019, 587, 113461. [Google Scholar] [CrossRef]
- Chen, X.; Zhou, C.; Guo, X. Ultrasensitive Detection and Binding Mechanism of Cocaine in an Aptamer-Based Single-Molecule Device. Chin. J. Chem. 2019, 37, 897–902. [Google Scholar] [CrossRef]
- Nodoushan, S.M.; Nasirizadeh, N.; Sedighian, H.; Kachuei, R.; Azimzadeh-Taft, M.; Fooladi, A.A.I. Detection of Staphylococcal Enterotoxin A (SEA) Using a Sensitive Nanomaterial-Based Electrochemical Aptasensor. Diam. Relat. Mater. 2022, 127, 109042. [Google Scholar] [CrossRef]
- Ziu, I.; Laryea, E.T.; Alashkar, F.; Wu, C.G.; Martic, S. A Dip-and-Read Optical Aptasensor for Detection of Tau Protein. Anal. Bioanal. Chem. 2020, 412, 1193–1201. [Google Scholar] [CrossRef]
- Huang, Z.; Yu, X.; Yang, Q.; Zhao, Y.; Wu, W. Aptasensors for Staphylococcus aureus Risk Assessment in Food. Front. Microbiol. 2021, 12, 714265. [Google Scholar] [CrossRef]
- Cennamo, N.; Pasquardini, L.; Arcadio, F.; Lunelli, L.; Vanzetti, L.; Carafa, V.; Altucci, L.; Zeni, L. SARS-CoV-2 Spike Protein Detection through a Plasmonic D-Shaped Plastic Optical Fiber Aptasensor. Talanta 2021, 233, 122532. [Google Scholar] [CrossRef]
- Tang, X.-L.; Hua, Y.; Guan, Q.; Yuan, C.-H. Improved Detection of Deeply Invasive Candidiasis with DNA Aptamers Specific Binding to (1→3)-β-D-Glucans from Candida Albicans. Eur. J. Clin. Microbiol. Infect. Dis. 2016, 35, 587–595. [Google Scholar] [CrossRef]
- Guo, W.; Zhang, C.; Ma, T.; Liu, X.; Chen, Z.; Li, S.; Deng, Y. Advances in Aptamer Screening and Aptasensors’ Detection of Heavy Metal Ions. J. Nanobiotechnology 2021, 19, 166. [Google Scholar] [CrossRef]
- Hong, K.L.; Sooter, L.J. Single-Stranded DNA Aptamers against Pathogens and Toxins: Identification and Biosensing Applications. Biomed Res. Int. 2015, 2015, 419318. [Google Scholar] [CrossRef]
- Kim, D.-M.; Go, M.-J.; Lee, J.; Na, D.; Yoo, S.-M. Recent Advances in Micro/Nanomaterial-Based Aptamer Selection Strategies. Molecules 2021, 26, 5187. [Google Scholar] [CrossRef] [PubMed]
- Park, K.S. Nucleic Acid Aptamer-Based Methods for Diagnosis of Infections. Biosens. Bioelectron. 2018, 102, 179–188. [Google Scholar] [CrossRef] [PubMed]
- Pavlov, V.; Shlyahovsky, B.; Willner, I. Fluorescence Detection of DNA by the Catalytic Activation of an Aptamer/Thrombin Complex. J. Am. Chem. Soc. 2005, 127, 6522–6523. [Google Scholar] [CrossRef]
- Park, K.S.; Lee, C.Y.; Park, H.G. Target DNA Induced Switches of DNA Polymerase Activity. Chem. Commun. 2015, 51, 9942–9945. [Google Scholar] [CrossRef]
- Stifel, J.; Spöring, M.; Hartig, J.S. Expanding the Toolbox of Synthetic Riboswitches with Guanine-Dependent Aptazymes. Synth. Biol. 2019, 4, ysy022. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Yao, C.; Wang, Y.; Zheng, W. Visual Diagnostic of Helicobacter Pylori Based on a Cascade Amplification of PCR and G-Quadruplex DNAzyme as a Color Label. J. Microbiol. Methods 2018, 146, 46–50. [Google Scholar] [CrossRef]
- Anantharaj, A.; Das, S.J.; Sharanabasava, P.; Lodha, R.; Kabra, S.K.; Sharma, T.K.; Medigeshi, G.R. Visual Detection of SARS-CoV-2 RNA by Conventional PCR-Induced Generation of DNAzyme Sensor. Front. Mol. Biosci. 2020, 7, 586254. [Google Scholar] [CrossRef]
- Zhou, H.-C. Introduction to Metal–Organic Frameworks. Chem. Rev. 2012, 112, 673–674. [Google Scholar] [CrossRef]
- Wang, H.-S.; Liu, H.-L.; Wang, K.; Ding, Y.; Xu, J.-J.; Xia, X.-H.; Chen, H.-Y. Insight into the Unique Fluorescence Quenching Property of Metal-Organic Frameworks upon DNA Binding. Anal. Chem. 2017, 89, 11366–11371. [Google Scholar] [CrossRef]
- Xie, B.-P.; Qiu, G.-H.; Hu, P.-P.; Liang, Z.; Liang, Y.-M.; Sun, B.; Bai, L.-P.; Jiang, Z.-H.; Chen, J.-X. Simultaneous Detection of Dengue and Zika Virus RNA Sequences with a Three-Dimensional Cu-Based Zwitterionic Metal–Organic Framework, Comparison of Single and Synchronous Fluorescence Analysis. Sens. Actuators B Chem. 2018, 254, 1133–1140. [Google Scholar] [CrossRef]
- Cheng, Q.; Parvin, B. Buckyballs Conjugated with Nucleic Acid Sequences Identifies Microorganisms in Live Cell Assays. J. Nanobiotechnology 2017, 15, 78. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Q.; Parvin, B. Rapid Identification of a Subset of Foodborne Bacteria in Live-Cell Assays. Appl. Microbiol. Biotechnol. 2020, 104, 10571–10584. [Google Scholar] [CrossRef] [PubMed]
- Chauhan, V.M.; Elsutohy, M.M.; McClure, C.P.; Irving, W.L.; Roddis, N.; Aylott, J.W. Gold–Oligonucleotide Nanoconstructs Engineered to Detect Conserved Enteroviral Nucleic Acid Sequences. Biosensors 2021, 11, 238. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.-H.; Yu, K.-Y.; Huang, S.-P.; Chuang, H.-W.; Lin, W.-Z.; Cherng, J.-H.; Hung, Y.-W.; Yeh, M.-K.; Hong, P.-D.; Liu, C.-C. Development of a Nucleic Acid Lateral Flow Immunoassay for the Detection of Human Polyomavirus BK. Diagnostics 2020, 10, 403. [Google Scholar] [CrossRef]
- Sakamoto, S.; Putalun, W.; Vimolmangkang, S.; Phoolcharoen, W.; Shoyama, Y.; Tanaka, H.; Morimoto, S. Enzyme-Linked Immunosorbent Assay for the Quantitative/Qualitative Analysis of Plant Secondary Metabolites. J. Nat. Med. 2018, 72, 32–42. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; He, S.; Wang, X.; Yan, Y.; Liu, J.; Wu, S.; Liu, S.; Lei, Y.; Chen, M.; Li, L.; et al. Rapid Lateral Flow Immunoassay for the Fluorescence Detection of SARS-CoV-2 RNA. Nat. Biomed. Eng. 2020, 4, 1150–1158. [Google Scholar] [CrossRef] [PubMed]
- Yu, X.; Zhang, S.; Guo, W.; Li, B.; Yang, Y.; Xie, B.; Li, K.; Zhang, L. Recent Advances on Functional Nucleic-Acid Biosensors. Sensors 2021, 21, 7109. [Google Scholar] [CrossRef]
- Abolhasan, R.; Mehdizadeh, A.; Rashidi, M.R.; Aghebati-Maleki, L.; Yousefi, M. Application of Hairpin DNA-Based Biosensors with Various Signal Amplification Strategies in Clinical Diagnosis. Biosens. Bioelectron. 2019, 129, 164–174. [Google Scholar] [CrossRef]
- Demidov, V.V. A New Use for Old Stuff: DNA Hairpins in DNA Amplification. Trends Biotechnol. 2002, 20, 189–190. [Google Scholar] [CrossRef]
- Iwe, I.A.; Li, W.; Li, Z.; Huang, J. Hairpin DNA-Mediated Isothermal Amplification (HDMIA) Techniques for Nucleic Acid Testing. Talanta 2021, 226, 122146. [Google Scholar] [CrossRef]
- Jung, Y.; Song, J.; Park, H.G. Ultrasensitive Nucleic Acid Detection Based on Phosphorothioated Hairpin-Assisted Isothermal Amplification. Sci. Rep. 2021, 11, 8399. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Duan, R.; Duan, Z.; Huang, F.; Xia, F. Fluorescence Signal Amplification Strategies Based on DNA Nanotechnology for MiRNA Detection. Chem. Res. Chin. Univ. 2020, 36, 194–202. [Google Scholar] [CrossRef]
- Evanko, D. Hybridization Chain Reaction. Nat. Methods 2004, 1, 186. [Google Scholar] [CrossRef]
- Zheng, J.; Ji, X.; Du, M.; Tian, S.; He, Z. Rational Construction of a DNA Nanomachine for HIV Nucleic Acid Ultrasensitive Sensing. Nanoscale 2018, 10, 17206–17211. [Google Scholar] [CrossRef] [PubMed]
- Mohammadniaei, M.; Zhang, M.; Ashley, J.; Christensen, U.B.; Friis-Hansen, L.J.; Gregersen, R.; Lisby, J.G.; Benfield, T.L.; Nielsen, F.E.; Henning Rasmussen, J.; et al. A Non-Enzymatic, Isothermal Strand Displacement and Amplification Assay for Rapid Detection of SARS-CoV-2 RNA. Nat. Commun. 2021, 12, 5089. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.; Tang, Y.; Jiang, Y.S.; Bhadra, S.; Du, Y.; Ellington, A.D.; Li, B. Strand-Exchange Nucleic Acid Circuitry with Enhanced Thermo-and Structure- Buffering Abilities Turns Gene Diagnostics Ultra-Reliable and Environmental Compatible. Sci. Rep. 2016, 6, 36605. [Google Scholar] [CrossRef]
- Luo, Z.; Li, Y.; Zhang, P.; He, L.; Feng, Y.; Feng, Y.; Qian, C.; Tian, Y.; Duan, Y. Catalytic Hairpin Assembly as Cascade Nucleic Acid Circuits for Fluorescent Biosensor: Design, Evolution and Application. TrAC Trends Anal. Chem. 2022, 151, 116582. [Google Scholar] [CrossRef]
- Banér, J.; Nilsson, M.; Mendel-Hartvig, M.; Landegren, U. Signal Amplification of Padlock Probes by Rolling Circle Replication. Nucleic Acids Res. 1998, 26, 5073–5078. [Google Scholar] [CrossRef] [PubMed]
- Gu, L.; Yan, W.; Liu, L.; Wang, S.; Zhang, X.; Lyu, M. Research Progress on Rolling Circle Amplification (RCA)-Based Biomedical Sensing. Pharmaceuticals 2018, 11, 35. [Google Scholar] [CrossRef]
- De Felice, M.; de Falco, M.; Zappi, D.; Antonacci, A.; Scognamiglio, V. Isothermal Amplification-Assisted Diagnostics for COVID-19. Biosens. Bioelectron. 2022, 205, 114101. [Google Scholar] [CrossRef]
- Kang, T.; Lu, J.; Yu, T.; Long, Y.; Liu, G. Advances in Nucleic Acid Amplification Techniques (NAATs): COVID-19 Point-of-Care Diagnostics as an Example. Biosens. Bioelectron. 2022, 206, 114109. [Google Scholar] [CrossRef] [PubMed]
- Qi, L.; Hu, Q.; Kang, Q.; Bi, Y.; Jiang, Y.; Yu, L. Detection of Biomarkers in Blood Using Liquid Crystals Assisted with Aptamer-Target Recognition Triggered in Situ Rolling Circle Amplification on Magnetic Beads. Anal. Chem. 2019, 91, 11653–11660. [Google Scholar] [CrossRef] [PubMed]
- Mignardi, M.; Mezger, A.; Qian, X.; La Fleur, L.; Botling, J.; Larsson, C.; Nilsson, M. Oligonucleotide Gap-Fill Ligation for Mutation Detection and Sequencing in Situ. Nucleic Acids Res. 2015, 43, e151. [Google Scholar] [CrossRef]
- Jain, S.; Dandy, D.S.; Geiss, B.J.; Henry, C.S. Padlock Probe-Based Rolling Circle Amplification Lateral Flow Assay for Point-of-Need Nucleic Acid Detection. Analyst 2021, 146, 4340–4347. [Google Scholar] [CrossRef]
- Chaibun, T.; Puenpa, J.; Ngamdee, T.; Boonapatcharoen, N.; Athamanolap, P.; O’Mullane, A.P.; Vongpunsawad, S.; Poovorawan, Y.; Lee, S.Y.; Lertanantawong, B. Rapid Electrochemical Detection of Coronavirus SARS-CoV-2. Nat. Commun. 2021, 12, 802. [Google Scholar] [CrossRef] [PubMed]
- Glökler, J.; Lim, T.S.; Ida, J.; Frohme, M. Isothermal Amplifications—A Comprehensive Review on Current Methods. Crit. Rev. Biochem. Mol. Biol. 2021, 56, 543–586. [Google Scholar] [CrossRef]
- Leonardo, S.; Toldrà, A.; Campàs, M. Biosensors Based on Isothermal DNA Amplification for Bacterial Detection in Food Safety and Environmental Monitoring. Sensors 2021, 21, 602. [Google Scholar] [CrossRef]
- Inoue, J.; Shigemori, Y.; Mikawa, T. Improvements of Rolling Circle Amplification (RCA) Efficiency and Accuracy Using Thermus Thermophilus SSB Mutant Protein. Nucleic Acids Res. 2006, 34, e69. [Google Scholar] [CrossRef]
- Zhang, K.; Zhang, H.; Cao, H.; Jiang, Y.; Mao, K.; Yang, Z. Rolling Circle Amplification as an Efficient Analytical Tool for Rapid Detection of Contaminants in Aqueous Environments. Biosensors 2021, 11, 352. [Google Scholar] [CrossRef] [PubMed]
- Piepenburg, O.; Williams, C.H.; Stemple, D.L.; Armes, N.A. DNA Detection Using Recombination Proteins. PLOS Biol. 2006, 4, e204. [Google Scholar] [CrossRef]
- Jauset-Rubio, M.; Svobodová, M.; Mairal, T.; McNeil, C.; Keegan, N.; Saeed, A.; Abbas, M.N.; El-Shahawi, M.S.; Bashammakh, A.S.; Alyoubi, A.O.; et al. Ultrasensitive, Rapid and Inexpensive Detection of DNA Using Paper Based Lateral Flow Assay. Sci. Rep. 2016, 6, 37732. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Liao, L.; Ma, C.; Zhang, X.; Yu, J.; Yi, L.; Liu, X.; Shen, H.; Gao, S.; Lu, Q. Duplex On-Site Detection of Vibrio Cholerae and Vibrio Vulnificus by Recombinase Polymerase Amplification and Three-Segment Lateral Flow Strips. Biosensors 2021, 11, 151. [Google Scholar] [CrossRef] [PubMed]
- Talwar, C.S.; Park, K.-H.; Ahn, W.-C.; Kim, Y.-S.; Kwon, O.S.; Yong, D.; Kang, T.; Woo, E. Detection of Infectious Viruses Using CRISPR-Cas12-Based Assay. Biosensors 2021, 11, 301. [Google Scholar] [CrossRef]
- Trinh, K.T.L.; Lee, N.Y. Fabrication of Wearable PDMS Device for Rapid Detection of Nucleic Acids via Recombinase Polymerase Amplification Operated by Human Body Heat. Biosensors 2022, 12, 72. [Google Scholar] [CrossRef] [PubMed]
- Roumani, F.; Rodrigues, C.; Barros-Velázquez, J.; Garrido-Maestu, A.; Prado, M. Development of a Panfungal Recombinase Polymerase Amplification (RPA) Method Coupled with Lateral Flow Strips for the Detection of Spoilage Fungi. Food Anal. Methods 2022, 1–10. [Google Scholar] [CrossRef]
- Karakkat, B.B.; Hockemeyer, K.; Franchett, M.; Olson, M.; Mullenberg, C.; Koch, P.L. Detection of Root-Infecting Fungi on Cool-Season Turfgrasses Using Loop-Mediated Isothermal Amplification and Recombinase Polymerase Amplification. J. Microbiol. Methods 2018, 151, 90–98. [Google Scholar] [CrossRef]
- Ju, Y.; Lin, Y.; Yang, G.; Wu, H.; Pan, Y. Development of Recombinase Polymerase Amplification Assay for Rapid Detection of Meloidogyne incognita, M. javanica, M. arenaria, and M. enterolobii. Eur. J. Plant Pathol. 2019, 155, 1155–1163. [Google Scholar] [CrossRef]
- Castellanos-Gonzalez, A.; White, A.C.; Melby, P.; Travi, B. Molecular Diagnosis of Protozoan Parasites by Recombinase Polymerase Amplification. Acta Trop. 2018, 182, 4–11. [Google Scholar] [CrossRef]
- Crannell, Z.A.; Rohrman, B.; Richards-Kortum, R. Equipment-Free Incubation of Recombinase Polymerase Amplification Reactions Using Body Heat. PLoS ONE 2014, 9, e112146. [Google Scholar] [CrossRef]
- Kong, M.; Li, Z.; Wu, J.; Hu, J.; Sheng, Y.; Wu, D.; Lin, Y.; Li, M.; Wang, X.; Wang, S. A Wearable Microfluidic Device for Rapid Detection of HIV-1 DNA Using Recombinase Polymerase Amplification. Talanta 2019, 205, 120155. [Google Scholar] [CrossRef]
- Zaghloul, H.; El-Shahat, M. Recombinase Polymerase Amplification as a Promising Tool in Hepatitis C Virus Diagnosis. World J. Hepatol. 2014, 6, 916–922. [Google Scholar] [CrossRef] [PubMed]
- Ahn, H.; Batule, B.S.; Seok, Y.; Kim, M.-G. Single-Step Recombinase Polymerase Amplification Assay Based on a Paper Chip for Simultaneous Detection of Multiple Foodborne Pathogens. Anal. Chem. 2018, 90, 10211–10216. [Google Scholar] [CrossRef] [PubMed]
- Magro, L.; Jacquelin, B.; Escadafal, C.; Garneret, P.; Kwasiborski, A.; Manuguerra, J.-C.; Monti, F.; Sakuntabhai, A.; Vanhomwegen, J.; Lafaye, P.; et al. Paper-Based RNA Detection and Multiplexed Analysis for Ebola Virus Diagnostics. Sci. Rep. 2017, 7, 1347. [Google Scholar] [CrossRef] [PubMed]
- Daher, R.K.; Stewart, G.; Boissinot, M.; Bergeron, M.G. Recombinase Polymerase Amplification for Diagnostic Applications. Clin. Chem. 2016, 62, 947–958. [Google Scholar] [CrossRef] [PubMed]
- Kaminski, M.M.; Abudayyeh, O.O.; Gootenberg, J.S.; Zhang, F.; Collins, J.J. CRISPR-Based Diagnostics. Nat. Biomed. Eng. 2021, 5, 643–656. [Google Scholar] [CrossRef] [PubMed]
- Joung, J.; Ladha, A.; Saito, M.; Kim, N.-G.; Woolley, A.E.; Segel, M.; Barretto, R.P.J.; Ranu, A.; Macrae, R.K.; Faure, G.; et al. Detection of SARS-CoV-2 with SHERLOCK One-Pot Testing. N. Engl. J. Med. 2020, 383, 1492–1494. [Google Scholar] [CrossRef] [PubMed]
- Zou, Y.; Mason, M.G.; Botella, J.R. Evaluation and Improvement of Isothermal Amplification Methods for Point-of-Need Plant Disease Diagnostics. PLoS ONE 2020, 15, e0235216. [Google Scholar] [CrossRef] [PubMed]
- Kim, N.-Y.; Oh, J.; Lee, S.-H.; Kim, H.; Moon, J.S.; Jeong, R.-D. Rapid and Specific Detection of Apple Stem Grooving Virus by Reverse Transcription-Recombinase Polymerase Amplification. Plant Pathol. J. 2018, 34, 575–579. [Google Scholar] [CrossRef]
- Wilson, I.G. Inhibition and Facilitation of Nucleic Acid Amplification. Appl. Environ. Microbiol. 1997, 63, 3741–3751. [Google Scholar] [CrossRef]
- Nagamine, K.; Hase, T.; Notomi, T. Accelerated Reaction by Loop-Mediated Isothermal Amplification Using Loop Primers. Mol. Cell. Probes 2002, 16, 223–229. [Google Scholar] [CrossRef]
- Notomi, T.; Mori, Y.; Tomita, N.; Kanda, H. Loop-Mediated Isothermal Amplification (LAMP): Principle, Features, and Future Prospects. J. Microbiol. 2015, 53, 60–69. [Google Scholar] [CrossRef] [PubMed]
- Davidson, J.L.; Wang, J.; Maruthamuthu, M.K.; Dextre, A.; Pascual-Garrigos, A.; Mohan, S.; Putikam, S.V.S.; Osman, F.O.I.; McChesney, D.; Seville, J.; et al. A Paper-Based Colorimetric Molecular Test for SARS-CoV-2 in Saliva. Biosens. Bioelectron. X 2021, 9, 100076. [Google Scholar] [CrossRef]
- Mohan, S.; Pascual-Garrigos, A.; Brouwer, H.; Pillai, D.; Koziol, J.; Ault, A.; Schoonmaker, J.; Johnson, T.; Verma, M.S. Loop-Mediated Isothermal Amplification for the Detection of Pasteurella multocida, Mannheimia haemolytica, and Histophilus somni in Bovine Nasal Samples. ACS Agric. Sci. Technol. 2021, 1, 100–108. [Google Scholar] [CrossRef]
- Zhu, Z.; Li, R.; Zhang, H.; Wang, J.; Lu, Y.; Zhang, D.; Yang, L. PAM-Free Loop-Mediated Isothermal Amplification Coupled with CRISPR/Cas12a Cleavage (Cas-PfLAMP) for Rapid Detection of Rice Pathogens. Biosens. Bioelectron. 2022, 204, 114076. [Google Scholar] [CrossRef]
- Francois, P.; Tangomo, M.; Hibbs, J.; Bonetti, E.-J.; Boehme, C.C.; Notomi, T.; Perkins, M.D.; Schrenzel, J. Robustness of a Loop-Mediated Isothermal Amplification Reaction for Diagnostic Applications. FEMS Immunol. Med. Microbiol. 2011, 62, 41–48. [Google Scholar] [CrossRef] [PubMed]
- Edwards, T.; Burke, P.A.; Smalley, H.B.; Gillies, L.; Hobbs, G. Loop-Mediated Isothermal Amplification Test for Detection of Neisseria gonorrhoeae in Urine Samples and Tolerance of the Assay to the Presence of Urea. J. Clin. Microbiol. 2014, 52, 2163–2165. [Google Scholar] [CrossRef] [PubMed]
- Walker, F.M.; Hsieh, K. Advances in Directly Amplifying Nucleic Acids from Complex Samples. Biosensors 2019, 9, 117. [Google Scholar] [CrossRef]
- Kaarj, K.; Akarapipad, P.; Yoon, J.-Y. Simpler, Faster, and Sensitive Zika Virus Assay Using Smartphone Detection of Loop-Mediated Isothermal Amplification on Paper Microfluidic Chips. Sci. Rep. 2018, 8, 12438. [Google Scholar] [CrossRef]
- Torres, C.; Vitalis, E.A.; Baker, B.R.; Gardner, S.N.; Torres, M.W.; Dzenitis, J.M. LAVA: An Open-Source Approach to Designing LAMP (Loop-Mediated Isothermal Amplification) DNA Signatures. BMC Bioinform. 2011, 12, 240. [Google Scholar] [CrossRef]
- Badolo, A.; Okado, K.; Guelbeogo, W.M.; Aonuma, H.; Bando, H.; Fukumoto, S.; Sagnon, N.; Kanuka, H. Development of an Allele-Specific, Loop-Mediated, Isothermal Amplification Method (AS-LAMP) to Detect the L1014F Kdr-w Mutation in Anopheles Gambiae s. l. Malar. J. 2012, 11, 227. [Google Scholar] [CrossRef]
- Carlos, F.F.; Veigas, B.; Matias, A.S.; Doria, G.; Flores, O.; Baptista, P.V. Allele Specific LAMP—Gold Nanoparticle for Characterization of Single Nucleotide Polymorphisms. Biotechnol. Rep. 2017, 16, 21–25. [Google Scholar] [CrossRef]
- Kalofonou, M.; Malpartida-Cardenas, K.; Alexandrou, G.; Rodriguez-Manzano, J.; Yu, L.-S.; Miscourides, N.; Allsopp, R.; Gleason, K.L.T.; Goddard, K.; Fernandez-Garcia, D.; et al. A Novel Hotspot Specific Isothermal Amplification Method for Detection of the Common PIK3CA p.H1047R Breast Cancer Mutation. Sci. Rep. 2020, 10, 4553. [Google Scholar] [CrossRef] [PubMed]
- Baek, Y.H.; Um, J.; Antigua, K.J.C.; Park, J.-H.; Kim, Y.; Oh, S.; Kim, Y.; Choi, W.-S.; Kim, S.G.; Jeong, J.H.; et al. Development of a Reverse Transcription-Loop-Mediated Isothermal Amplification as a Rapid Early-Detection Method for Novel SARS-CoV-2. Emerg. Microbes Infect. 2020, 9, 998–1007. [Google Scholar] [CrossRef] [PubMed]
- Shirato, K. Detecting Amplicons of Loop-Mediated Isothermal Amplification. Microbiol. Immunol. 2019, 63, 407–412. [Google Scholar] [CrossRef] [PubMed]
- Tanner, N.A.; Zhang, Y.; Evans, T.C. Visual Detection of Isothermal Nucleic Acid Amplification Using PH-Sensitive Dyes. BioTechniques 2015, 58, 59–68. [Google Scholar] [CrossRef]
- Seok, Y.; Joung, H.-A.; Byun, J.-Y.; Jeon, H.-S.; Shin, S.J.; Kim, S.; Shin, Y.-B.; Han, H.S.; Kim, M.-G. A Paper-Based Device for Performing Loop-Mediated Isothermal Amplification with Real-Time Simultaneous Detection of Multiple DNA Targets. Theranostics 2017, 7, 2220–2230. [Google Scholar] [CrossRef]
- Aman, R.; Mahas, A.; Mahfouz, M. Nucleic Acid Detection Using CRISPR/Cas Biosensing Technologies. ACS Synth. Biol. 2020, 9, 1226–1233. [Google Scholar] [CrossRef]
- Jiang, F.; Doudna, J.A. CRISPR-Cas9 Structures and Mechanisms. Annu. Rev. Biophys. 2017, 46, 505–529. [Google Scholar] [CrossRef]
- Bonini, A.; Poma, N.; Vivaldi, F.; Kirchhain, A.; Salvo, P.; Bottai, D.; Tavanti, A.; Di Francesco, F. Advances in Biosensing: The CRISPR/Cas System as a New Powerful Tool for the Detection of Nucleic Acids. J. Pharm. Biomed. Anal. 2021, 192, 113645. [Google Scholar] [CrossRef]
- Chen, J.S.; Ma, E.; Harrington, L.B.; Costa, M.D.; Tian, X.; Palefsky, J.M.; Doudna, J.A. CRISPR-Cas12a Target Binding Unleashes Indiscriminate Single-Stranded DNase Activity. Science 2018, 360, 436–439. [Google Scholar] [CrossRef] [PubMed]
- Mustafa, M.I.; Makhawi, A.M. SHERLOCK and DETECTR: CRISPR-Cas Systems as Potential Rapid Diagnostic Tools for Emerging Infectious Diseases. J. Clin. Microbiol. 2021, 59, e00745-20. [Google Scholar] [CrossRef] [PubMed]
- Kellner, M.J.; Koob, J.G.; Gootenberg, J.S.; Abudayyeh, O.O.; Zhang, F. SHERLOCK: Nucleic Acid Detection with CRISPR Nucleases. Nat. Protoc. 2019, 14, 2986–3012. [Google Scholar] [CrossRef]
- Broughton, J.P.; Deng, X.; Yu, G.; Fasching, C.L.; Servellita, V.; Singh, J.; Miao, X.; Streithorst, J.A.; Granados, A.; Sotomayor-Gonzalez, A.; et al. CRISPR–Cas12-Based Detection of SARS-CoV-2. Nat. Biotechnol. 2020, 38, 870–874. [Google Scholar] [CrossRef]
- Van Dongen, J.E.; Berendsen, J.T.W.; Steenbergen, R.D.M.; Wolthuis, R.M.F.; Eijkel, J.C.T.; Segerink, L.I. Point-of-Care CRISPR/Cas Nucleic Acid Detection: Recent Advances, Challenges and Opportunities. Biosens. Bioelectron. 2020, 166, 112445. [Google Scholar] [CrossRef] [PubMed]
- Gootenberg, J.S.; Abudayyeh, O.O.; Lee, J.W.; Essletzbichler, P.; Dy, A.J.; Joung, J.; Verdine, V.; Donghia, N.; Daringer, N.M.; Freije, C.A.; et al. Nucleic Acid Detection with CRISPR-Cas13a/C2c2. Science 2017, 356, 438–442. [Google Scholar] [CrossRef]
- Selvam, K.; Najib, M.A.; Khalid, M.F.; Mohamad, S.; Palaz, F.; Ozsoz, M.; Aziah, I. RT-LAMP CRISPR-Cas12/13-Based SARS-CoV-2 Detection Methods. Diagnostics 2021, 11, 1646. [Google Scholar] [CrossRef]
- Wang, R.; Qian, C.; Pang, Y.; Li, M.; Yang, Y.; Ma, H.; Zhao, M.; Qian, F.; Yu, H.; Liu, Z.; et al. OpvCRISPR: One-Pot Visual RT-LAMP-CRISPR Platform for SARS-Cov-2 Detection. Biosens. Bioelectron. 2021, 172, 112766. [Google Scholar] [CrossRef] [PubMed]
- Gootenberg, J.S.; Abudayyeh, O.O.; Kellner, M.J.; Joung, J.; Collins, J.J.; Zhang, F. Multiplexed and Portable Nucleic Acid Detection Platform with Cas13, Cas12a, and Csm6. Science 2018, 360, 439–444. [Google Scholar] [CrossRef]
- Zhang, J.; Lv, H.; Li, L.; Chen, M.; Gu, D.; Wang, J.; Xu, Y. Recent Improvements in CRISPR-Based Amplification-Free Pathogen Detection. Front. Microbiol. 2021, 12, 2898. [Google Scholar] [CrossRef]
- Liu, T.Y.; Knott, G.J.; Smock, D.C.J.; Desmarais, J.J.; Son, S.; Bhuiya, A.; Jakhanwal, S.; Prywes, N.; Agrawal, S.; Díaz de León Derby, M.; et al. Accelerated RNA Detection Using Tandem CRISPR Nucleases. Nat. Chem. Biol. 2021, 17, 982–988. [Google Scholar] [CrossRef] [PubMed]
- Suea-Ngam, A.; Howes, P.D.; deMello, A.J. An Amplification-Free Ultra-Sensitive Electrochemical CRISPR/Cas Biosensor for Drug-Resistant Bacteria Detection. Chem. Sci. 2021, 12, 12733–12743. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Liu, J.; Zhou, X. A CRISPR-Based and Post-Amplification Coupled SARS-CoV-2 Detection with a Portable Evanescent Wave Biosensor. Biosens. Bioelectron. 2021, 190, 113418. [Google Scholar] [CrossRef] [PubMed]
- Yoo, H.; Jo, H.; Soo Oh, S. Detection and beyond: Challenges and Advances in Aptamer-Based Biosensors. Mater. Adv. 2020, 1, 2663–2687. [Google Scholar] [CrossRef]
- Zhuo, Z.; Yu, Y.; Wang, M.; Li, J.; Zhang, Z.; Liu, J.; Wu, X.; Lu, A.; Zhang, G.; Zhang, B. Recent Advances in SELEX Technology and Aptamer Applications in Biomedicine. Int. J. Mol. Sci. 2017, 18, 2142. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.; Wu, X.; Li, S.; Lu, R.; Li, Y.; Wan, Z.; Qin, J.; Yu, G.; Jin, X.; Zhang, C. Comparative Evaluation of 19 Reverse Transcription Loop-Mediated Isothermal Amplification Assays for Detection of SARS-CoV-2. Sci. Rep. 2021, 11, 2936. [Google Scholar] [CrossRef]
- Fu, J.; Zhang, Y.; Cai, G.; Meng, G.; Shi, S. Rapid and Sensitive RPA-Cas12a-Fluorescence Assay for Point-of-Care Detection of African Swine Fever Virus. PLoS ONE 2021, 16, e0254815. [Google Scholar] [CrossRef]
- Pascual-Garrigos, A.; Maruthamuthu, M.K.; Ault, A.; Davidson, J.L.; Rudakov, G.; Pillai, D.; Koziol, J.; Schoonmaker, J.P.; Johnson, T.; Verma, M.S. On-Farm Colorimetric Detection of Pasteurella multocida, Mannheimia haemolytica, and Histophilus somni in Crude Bovine Nasal Samples. Vet. Res. 2021, 52, 126. [Google Scholar] [CrossRef]
- Wang, J.; Dextre, A.; Pascual-Garrigos, A.; Davidson, J.L.; Maruthamuthu, M.K.; McChesney, D.; Seville, J.; Verma, M.S. Fabrication of a Paper-Based Colorimetric Molecular Test for SARS-CoV-2. MethodsX 2021, 8, 101586. [Google Scholar] [CrossRef]
- An, B.; Zhang, H.; Su, X.; Guo, Y.; Wu, T.; Ge, Y.; Zhu, F.; Cui, L. Rapid and Sensitive Detection of Salmonella spp. Using CRISPR-Cas13a Combined with Recombinase Polymerase Amplification. Front. Microbiol. 2021, 12, 732426. [Google Scholar] [CrossRef]
- Liu, H.; Wang, J.; Zeng, H.; Liu, X.; Jiang, W.; Wang, Y.; Ouyang, W.; Tang, X. RPA-Cas12a-FS: A Frontline Nucleic Acid Rapid Detection System for Food Safety Based on CRISPR-Cas12a Combined with Recombinase Polymerase Amplification. Food Chem. 2021, 334, 127608. [Google Scholar] [CrossRef]
- Jiang, W.; Ren, Y.; Han, X.; Xue, J.; Shan, T.; Chen, Z.; Liu, Y.; Wang, Q. Recombinase Polymerase Amplification-Lateral Flow (RPA-LF) Assay Combined with Immunomagnetic Separation for Rapid Visual Detection of Vibrio Parahaemolyticus in Raw Oysters. Anal. Bioanal. Chem. 2020, 412, 2903–2914. [Google Scholar] [CrossRef] [PubMed]
- Abraham, J.P.; Plourde, B.D.; Cheng, L. Using Heat to Kill SARS-CoV-2. Rev. Med. Virol. 2020, 30, e2115. [Google Scholar] [CrossRef] [PubMed]
- El-Tholoth, M.; Bau, H.H.; Song, J. A Single and Two-Stage, Closed-Tube, Molecular Test for the 2019 Novel Coronavirus (COVID-19) at Home, Clinic, and Points of Entry. ChemRxiv 2020. [Google Scholar] [CrossRef]
- Katzmeier, F.; Aufinger, L.; Dupin, A.; Quintero, J.; Lenz, M.; Bauer, L.; Klumpe, S.; Sherpa, D.; Dürr, B.; Honemann, M.; et al. A Low-Cost Fluorescence Reader for In Vitro Transcription and Nucleic Acid Detection with Cas13a. PLoS ONE 2019, 14, e0220091. [Google Scholar] [CrossRef] [PubMed]
- García-Bernalt Diego, J.; Fernández-Soto, P.; Crego-Vicente, B.; Alonso-Castrillejo, S.; Febrer-Sendra, B.; Gómez-Sánchez, A.; Vicente, B.; López-Abán, J.; Muro, A. Progress in Loop-Mediated Isothermal Amplification Assay for Detection of Schistosoma mansoni DNA: Towards a Ready-to-Use Test. Sci. Rep. 2019, 9, 14744. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Gallagher, R.; Bishop, J.; Kline, E.; Buser, J.; Lafleur, L.; Shah, K.; Lutz, B.; Yager, P. Long-Term Dry Storage of Enzyme-Based Reagents for Isothermal Nucleic Acid Amplification in a Porous Matrix for Use in Point-of-Care Diagnostic Devices. Analyst 2020, 145, 6875–6886. [Google Scholar] [CrossRef]
- Silveira, C.M.; Monteiro, T.; Almeida, M.G. Biosensing with Paper-Based Miniaturized Printed Electrodes—A Modern Trend. Biosensors 2016, 6, 51. [Google Scholar] [CrossRef]
- Liu, C.C. Electrochemical Based Biosensors. Biosensors 2012, 2, 269–272. [Google Scholar] [CrossRef]
- Farshchi, F.; Saadati, A.; Fathi, N.; Hasanzadeh, M.; Samiei, M. Flexible Paper-Based Label-Free Electrochemical Biosensor for the Monitoring of MiRNA-21 Using Core–Shell Ag@Au/GQD Nano-Ink: A New Platform for the Accurate and Rapid Analysis by Low Cost Lab-on-Paper Technology. Anal. Methods 2021, 13, 1286–1294. [Google Scholar] [CrossRef]
- Moccia, M.; Caratelli, V.; Cinti, S.; Pede, B.; Avitabile, C.; Saviano, M.; Imbriani, A.L.; Moscone, D.; Arduini, F. Paper-Based Electrochemical Peptide Nucleic Acid (PNA) Biosensor for Detection of MiRNA-492: A Pancreatic Ductal Adenocarcinoma Biomarker. Biosens. Bioelectron. 2020, 165, 112371. [Google Scholar] [CrossRef]
- Nielsen, P.E.; Haaima, G. Peptide Nucleic Acid (PNA). A DNA Mimic with a Pseudopeptide Backbone. Chem. Soc. Rev. 1997, 26, 73–78. [Google Scholar] [CrossRef]
- Ananthanawat, C.; Vilaivan, T.; Hoven, V.P.; Su, X. Comparison of DNA, Aminoethylglycyl PNA and Pyrrolidinyl PNA as Probes for Detection of DNA Hybridization Using Surface Plasmon Resonance Technique. Biosens. Bioelectron. 2010, 25, 1064–1069. [Google Scholar] [CrossRef] [PubMed]
- Teengam, P.; Siangproh, W.; Tuantranont, A.; Henry, C.S.; Vilaivan, T.; Chailapakul, O. Electrochemical Paper-Based Peptide Nucleic Acid Biosensor for Detecting Human Papillomavirus. Anal. Chim. Acta 2017, 952, 32–40. [Google Scholar] [CrossRef] [PubMed]
- Fu, P.; Xing, S.; Xu, M.; Zhao, Y.; Zhao, C. Peptide Nucleic Acid-Based Electrochemical Biosensor for Simultaneous Detection of Multiple MicroRNAs from Cancer Cells with Catalytic Hairpin Assembly Amplification. Sens. Actuators B Chem. 2020, 305, 127545. [Google Scholar] [CrossRef]
- Vilaivan, T. Pyrrolidinyl PNA with α/β-Dipeptide Backbone: From Development to Applications. Acc. Chem. Res. 2015, 48, 1645–1656. [Google Scholar] [CrossRef]
- Srisomwat, C.; Yakoh, A.; Chuaypen, N.; Tangkijvanich, P.; Vilaivan, T.; Chailapakul, O. Amplification-Free DNA Sensor for the One-Step Detection of the Hepatitis B Virus Using an Automated Paper-Based Lateral Flow Electrochemical Device. Anal. Chem. 2021, 93, 2879–2887. [Google Scholar] [CrossRef]
- Horny, M.-C.; Lazerges, M.; Siaugue, J.-M.; Pallandre, A.; Rose, D.; Bedioui, F.; Deslouis, C.; Haghiri-Gosnet, A.-M.; Gamby, J. Electrochemical DNA Biosensors Based on Long-Range Electron Transfer: Investigating the Efficiency of a Fluidic Channel Microelectrode Compared to an Ultramicroelectrode in a Two-Electrode Setup. Lab Chip 2016, 16, 4373–4381. [Google Scholar] [CrossRef]
- Bhatt, G.; Bhattacharya, S. Biosensors on Chip: A Critical Review from an Aspect of Micro/Nanoscales. J. Micromanuf. 2019, 2, 198–219. [Google Scholar] [CrossRef]
- Khaliliazar, S.; Toldrà, A.; Chondrogiannis, G.; Hamedi, M.M. Electroanalytical Paper-Based Nucleic Acid Amplification Biosensors with Integrated Thread Electrodes. Anal. Chem. 2021, 93, 14187–14195. [Google Scholar] [CrossRef]
- Teengam, P.; Siangproh, W.; Tuantranont, A.; Vilaivan, T.; Chailapakul, O.; Henry, C.S. Electrochemical Impedance-Based DNA Sensor Using Pyrrolidinyl Peptide Nucleic Acids for Tuberculosis Detection. Anal. Chim. Acta 2018, 1044, 102–109. [Google Scholar] [CrossRef]
- Santhanam, M.; Algov, I.; Alfonta, L. DNA/RNA Electrochemical Biosensing Devices a Future Replacement of PCR Methods for a Fast Epidemic Containment. Sensors 2020, 20, 4648. [Google Scholar] [CrossRef] [PubMed]
- Kudr, J.; Michalek, P.; Ilieva, L.; Adam, V.; Zitka, O. COVID-19: A Challenge for Electrochemical Biosensors. Trends Anal. Chem. 2021, 136, 116192. [Google Scholar] [CrossRef] [PubMed]
- Bruch, R.; Baaske, J.; Chatelle, C.; Meirich, M.; Madlener, S.; Weber, W.; Dincer, C.; Urban, G.A. CRISPR/Cas13a-Powered Electrochemical Microfluidic Biosensor for Nucleic Acid Amplification-Free MiRNA Diagnostics. Adv. Mater. 2019, 31, 1905311. [Google Scholar] [CrossRef] [PubMed]
- Choopara, I.; Suea-Ngam, A.; Teethaisong, Y.; Howes, P.D.; Schmelcher, M.; Leelahavanichkul, A.; Thunyaharn, S.; Wongsawaeng, D.; deMello, A.J.; Dean, D.; et al. Fluorometric Paper-Based, Loop-Mediated Isothermal Amplification Devices for Quantitative Point-of-Care Detection of Methicillin-Resistant Staphylococcus Aureus (MRSA). ACS Sens. 2021, 6, 742–751. [Google Scholar] [CrossRef] [PubMed]
- Kaur, N.; Michael, J.S.; Toley, B.J. A Modular Paper-and-Plastic Device for Tuberculosis Nucleic Acid Amplification Testing in Limited-Resource Settings. Sci. Rep. 2019, 9, 15367. [Google Scholar] [CrossRef]
- Xu, G.; Nolder, D.; Reboud, J.; Oguike, M.C.; van Schalkwyk, D.A.; Sutherland, C.J.; Cooper, J.M. Paper-Origami-Based Multiplexed Malaria Diagnostics from Whole Blood. Angew. Chem. Int. Ed. Engl. 2016, 55, 15250–15253. [Google Scholar] [CrossRef]
- Deng, X.; Wang, C.; Gao, Y.; Li, J.; Wen, W.; Zhang, X.; Wang, S. Applying Strand Displacement Amplification to Quantum Dots-Based Fluorescent Lateral Flow Assay Strips for HIV-DNA Detection. Biosens. Bioelectron. 2018, 105, 211–217. [Google Scholar] [CrossRef]
- Xu, Y.; Liu, Y.; Wu, Y.; Xia, X.; Liao, Y.; Li, Q. Fluorescent Probe-Based Lateral Flow Assay for Multiplex Nucleic Acid Detection. Anal. Chem. 2014, 86, 5611–5614. [Google Scholar] [CrossRef]
- Magiati, M.; Sevastou, A.; Kalogianni, D.P. A Fluorometric Lateral Flow Assay for Visual Detection of Nucleic Acids Using a Digital Camera Readout. Microchim. Acta 2018, 185, 314. [Google Scholar] [CrossRef]
- Farrar, J.S.; Wittwer, C.T. Chapter 6—High-Resolution Melting Curve Analysis for Molecular Diagnostics. In Molecular Diagnostics, 3rd ed.; Patrinos, G.P., Ed.; Academic Press: Cambridge, MA, USA, 2017; pp. 79–102. ISBN 978-0-12-802971-8. [Google Scholar]
- Dragan, A.I.; Pavlovic, R.; McGivney, J.B.; Casas-Finet, J.R.; Bishop, E.S.; Strouse, R.J.; Schenerman, M.A.; Geddes, C.D. SYBR Green I: Fluorescence Properties and Interaction with DNA. J. Fluoresc. 2012, 22, 1189–1199. [Google Scholar] [CrossRef]
- Safavieh, M.; Kanakasabapathy, M.K.; Tarlan, F.; Ahmed, M.U.; Zourob, M.; Asghar, W.; Shafiee, H. Emerging Loop-Mediated Isothermal Amplification-Based Microchip and Microdevice Technologies for Nucleic Acid Detection. ACS Biomater. Sci. Eng. 2016, 2, 278–294. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Huang, S.; Liu, N.; Dong, D.; Yang, Z.; Tang, Y.; Ma, W.; He, X.; Ao, D.; Xu, Y.; et al. Establishment of an Accurate and Fast Detection Method Using Molecular Beacons in Loop-Mediated Isothermal Amplification Assay. Sci. Rep. 2017, 7, 40125. [Google Scholar] [CrossRef] [PubMed]
- Scott, A.T.; Layne, T.R.; O’Connell, K.C.; Tanner, N.A.; Landers, J.P. Comparative Evaluation and Quantitative Analysis of Loop-Mediated Isothermal Amplification Indicators. Anal. Chem. 2020, 92, 13343–13353. [Google Scholar] [CrossRef] [PubMed]
- He, S.; Jang, H.; Zhao, C.; Xu, K.; Wang, J.; Pang, B.; Si, X.; Jin, M.; Song, X.; Li, J. Rapid Visualized Isothermal Nucleic Acid Testing of Vibrio Parahaemolyticus by Polymerase Spiral Reaction. Anal. Bioanal. Chem. 2020, 412, 93–101. [Google Scholar] [CrossRef]
- Shah, K.G.; Yager, P. Wavelengths and Lifetimes of Paper Autofluorescence: A Simple Substrate Screening Process to Enhance the Sensitivity of Fluorescence-Based Assays in Paper. Anal. Chem. 2017, 89, 12023–12029. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Jiang, H.; Li, X.; Lv, X. Recent Advances in Sensitivity Enhancement for Lateral Flow Assay. Microchim. Acta 2021, 188, 379. [Google Scholar] [CrossRef] [PubMed]
- Sapountzi, E.A.; Tragoulias, S.S.; Kalogianni, D.P.; Ioannou, P.C.; Christopoulos, T.K. Lateral Flow Devices for Nucleic Acid Analysis Exploiting Quantum Dots as Reporters. Anal. Chim. Acta 2015, 864, 48–54. [Google Scholar] [CrossRef]
- Wu, J.; Lu, Y.; Ren, N.; Jia, M.; Wang, R.; Zhang, J. DNAzyme-Functionalized R-Phycoerythrin as a Cost-Effective and Environment-Friendly Fluorescent Biosensor for Aqueous Pb2+ Detection. Sensors 2019, 19, 2732. [Google Scholar] [CrossRef]
- Vicente, F.A.; Cardoso, I.S.; Martins, M.; Gonçalves, C.V.M.; Dias, A.C.R.V.; Domingues, P.; Coutinho, J.A.P.; Ventura, S.P.M. R-Phycoerythrin Extraction and Purification from Fresh Gracilaria sp. Using Thermo-Responsive Systems. Green Chem. 2019, 21, 3816–3826. [Google Scholar] [CrossRef]
- Karawajew, L.; Behrsing, O.; Kaiser, G.; Micheel, B. Production and ELISA Application of Bispecific Monoclonal Antibodies against Fluorescein Isothiocyanate (FITC) and Horseradish Peroxidase (HRP). J. Immunol. Methods 1988, 111, 95–99. [Google Scholar] [CrossRef]
- Lee, D.; Chu, C.-H.; Sarioglu, A.F. Point-of-Care Toolkit for Multiplex Molecular Diagnosis of SARS-CoV-2 and Influenza A and B Viruses. ACS Sens. 2021, 6, 3204–3213. [Google Scholar] [CrossRef]
- Trinh, T.N.D.; Lee, N.Y. Spinning and Fully Integrated Microdevice for Rapid Screening of Vancomycin-Resistant Enterococcus. ACS Sens. 2021, 6, 2902–2910. [Google Scholar] [CrossRef]
- Thai, D.A.; Lee, N.Y. A Paper-Based Colorimetric Chemosensor for Rapid and Highly Sensitive Detection of Sulfide for Environmental Monitoring. Anal. Methods 2021, 13, 1332–1339. [Google Scholar] [CrossRef] [PubMed]
- Trinh, T.N.D.; Thai, D.A.; Lee, N.Y. Pop-up Paper-Based and Fully Integrated Microdevice for Point-of-Care Testing of Vancomycin-Resistant Enterococcus. Sens. Actuators B Chem. 2021, 345, 130362. [Google Scholar] [CrossRef]
- Ding, X.; Li, Z.; Avery, L.; Ballesteros, E.; Makol, R.; Liu, C. PH-EVD: A PH-Paper-Based Extraction and Visual Detection System for Instrument-Free SARS-CoV-2 Diagnostics. Adv. NanoBiomed Res. 2021, 2, 2100101. [Google Scholar] [CrossRef] [PubMed]
- Roy, S.; Mohd-Naim, N.F.; Safavieh, M.; Ahmed, M.U. Colorimetric Nucleic Acid Detection on Paper Microchip Using Loop Mediated Isothermal Amplification and Crystal Violet Dye. ACS Sens. 2017, 2, 1713–1720. [Google Scholar] [CrossRef]
- Trieu, P.T.; Lee, N.Y. Paper-Based All-in-One Origami Microdevice for Nucleic Acid Amplification Testing for Rapid Colorimetric Identification of Live Cells for Point-of-Care Testing. Anal. Chem. 2019, 91, 11013–11022. [Google Scholar] [CrossRef]
- Diep Trinh, T.N.; Yoon Lee, N. A Foldable Isothermal Amplification Microdevice for Fuchsin-Based Colorimetric Detection of Multiple Foodborne Pathogens. Lab Chip 2019, 19, 1397–1405. [Google Scholar] [CrossRef]
- Trinh, K.T.L.; Trinh, T.N.D.; Lee, N.Y. Fully Integrated and Slidable Paper-Embedded Plastic Microdevice for Point-of-Care Testing of Multiple Foodborne Pathogens. Biosens. Bioelectron. 2019, 135, 120–128. [Google Scholar] [CrossRef]
- Tsai, T.-T.; Huang, C.-Y.; Chen, C.-A.; Shen, S.-W.; Wang, M.-C.; Cheng, C.-M.; Chen, C.-F. Diagnosis of Tuberculosis Using Colorimetric Gold Nanoparticles on a Paper-Based Analytical Device. ACS Sens. 2017, 2, 1345–1354. [Google Scholar] [CrossRef]
- Naorungroj, S.; Teengam, P.; Vilaivan, T.; Chailapakul, O. Paper-Based DNA Sensor Enabling Colorimetric Assay Integrated with Smartphone for Human Papillomavirus Detection. New J. Chem. 2021, 45, 6960–6967. [Google Scholar] [CrossRef]
- Gao, Y.; Deng, X.; Wen, W.; Zhang, X.; Wang, S. Ultrasensitive Paper Based Nucleic Acid Detection Realized by Three-Dimensional DNA-AuNPs Network Amplification. Biosens. Bioelectron. 2017, 92, 529–535. [Google Scholar] [CrossRef]
- Teengam, P.; Siangproh, W.; Tuantranont, A.; Vilaivan, T.; Chailapakul, O.; Henry, C.S. Multiplex Paper-Based Colorimetric DNA Sensor Using Pyrrolidinyl Peptide Nucleic Acid-Induced AgNPs Aggregation for Detecting MERS-CoV, MTB, and HPV Oligonucleotides. Anal. Chem. 2017, 89, 5428–5435. [Google Scholar] [CrossRef]
- Kalish, B.; Zhang, J.; Edema, H.; Luong, J.; Roper, J.; Beaudette, C.; Echodu, R.; Tsutsui, H. Distance and Microsphere Aggregation-Based DNA Detection in a Paper-Based Microfluidic Device. SLAS Technol. Transl. Life Sci. Innov. 2020, 25, 58–66. [Google Scholar] [CrossRef]
- Kalish, B.; Luong, J.; Roper, J.; Beaudette, C.; Tsutsui, H. Distance-Based Quantitative DNA Detection in a Paper-Based Microfluidic Device. In Proceedings of the 2017 IEEE 12th International Conference on Nano/Micro Engineered and Molecular Systems (NEMS), Los Angeles, CA, USA, 9–12 April 2017; pp. 337–341. [Google Scholar]
- Hongwarittorrn, I.; Chaichanawongsaroj, N.; Laiwattanapaisal, W. Semi-Quantitative Visual Detection of Loop Mediated Isothermal Amplification (LAMP)-Generated DNA by Distance-Based Measurement on a Paper Device. Talanta 2017, 175, 135–142. [Google Scholar] [CrossRef] [PubMed]
- Gadkar, V.J.; Goldfarb, D.M.; Gantt, S.; Tilley, P.A.G. Real-Time Detection and Monitoring of Loop Mediated Amplification (LAMP) Reaction Using Self-Quenching and De-Quenching Fluorogenic Probes. Sci. Rep. 2018, 8, 5548. [Google Scholar] [CrossRef] [PubMed]
- Miyamoto, S.; Sano, S.; Takahashi, K.; Jikihara, T. Method for Colorimetric Detection of Double-Stranded Nucleic Acid Using Leuco Triphenylmethane Dyes. Anal. Biochem. 2015, 473, 28–33. [Google Scholar] [CrossRef] [PubMed]
- Kurozimi, T.; Kurihara, K.; Hachimoto, Y. Interaction of Methyl Green with Deoxyribonucleic Acid as Observed by a New Method of Using Hydrogen Peroxide. J. Biochem. 1963, 53, 135–142. [Google Scholar] [CrossRef] [PubMed]
- Elghanian, R.; Storhoff, J.J.; Mucic, R.C.; Letsinger, R.L.; Mirkin, C.A. Selective Colorimetric Detection of Polynucleotides Based on the Distance-Dependent Optical Properties of Gold Nanoparticles. Science 1997, 277, 1078–1081. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, W.-Y.; Lin, C.-H.; Lin, T.-C.; Lin, C.-H.; Chang, H.-F.; Tsai, C.-H.; Wu, H.-T.; Lin, C.-S. Development and Efficacy of Lateral Flow Point-of-Care Testing Devices for Rapid and Mass COVID-19 Diagnosis by the Detections of SARS-CoV-2 Antigen and Anti-SARS-CoV-2 Antibodies. Diagnostics 2021, 11, 1760. [Google Scholar] [CrossRef]
- Goto, M.; Honda, E.; Ogura, A.; Nomoto, A.; Hanaki, K.-I. Colorimetric Detection of Loop-Mediated Isothermal Amplification Reaction by Using Hydroxy Naphthol Blue. Available online: https://www.future-science.com/doi/abs/10.2144/000113072 (accessed on 3 January 2022).
- Seok, Y.; Batule, B.S.; Kim, M.-G. Lab-on-Paper for All-in-One Molecular Diagnostics (LAMDA) of Zika, Dengue, and Chikungunya Virus from Human Serum. Biosens. Bioelectron. 2020, 165, 112400. [Google Scholar] [CrossRef] [PubMed]
- Tomita, N.; Mori, Y.; Kanda, H.; Notomi, T. Loop-Mediated Isothermal Amplification (LAMP) of Gene Sequences and Simple Visual Detection of Products. Nat. Protoc. 2008, 3, 877–882. [Google Scholar] [CrossRef] [PubMed]
- Alafeef, M.; Dighe, K.; Moitra, P.; Pan, D. Rapid, Ultrasensitive, and Quantitative Detection of SARS-CoV-2 Using Antisense Oligonucleotides Directed Electrochemical Biosensor Chip. ACS Nano 2020, 14, 17028–17045. [Google Scholar] [CrossRef] [PubMed]
- Quyen, T.L.; Ngo, T.A.; Bang, D.D.; Madsen, M.; Wolff, A. Classification of Multiple DNA Dyes Based on Inhibition Effects on Real-Time Loop-Mediated Isothermal Amplification (LAMP): Prospect for Point of Care Setting. Front. Microbiol. 2019, 10, 2234. [Google Scholar] [CrossRef] [PubMed]
- Martinez, A.W.; Phillips, S.T.; Butte, M.J.; Whitesides, G.M. Patterned Paper as a Platform for Inexpensive, Low Volume, Portable Bioassays. Angew. Chem. Int. Ed. Engl. 2007, 46, 1318–1320. [Google Scholar] [CrossRef] [PubMed]
- Hunt, J.P.; Zhao, E.L.; Free, T.J.; Soltani, M.; Warr, C.A.; Benedict, A.B.; Takahashi, M.K.; Griffitts, J.S.; Pitt, W.G.; Bundy, B.C. Towards Detection of SARS-CoV-2 RNA in Human Saliva: A Paper-Based Cell-Free Toehold Switch Biosensor with a Visual Bioluminescent Output. New Biotechnol. 2022, 66, 53–60. [Google Scholar] [CrossRef]
- Myhrvold, C.; Freije, C.A.; Gootenberg, J.S.; Abudayyeh, O.O.; Metsky, H.C.; Durbin, A.F.; Kellner, M.J.; Tan, A.L.; Paul, L.M.; Parham, L.A.; et al. Field-Deployable Viral Diagnostics Using CRISPR-Cas13. Science 2018, 360, 444–448. [Google Scholar] [CrossRef]
- Batule, B.S.; Seok, Y.; Kim, M.-G. Paper-Based Nucleic Acid Testing System for Simple and Early Diagnosis of Mosquito-Borne RNA Viruses from Human Serum. Biosens. Bioelectron. 2020, 151, 111998. [Google Scholar] [CrossRef]
- Reboud, J.; Xu, G.; Garrett, A.; Adriko, M.; Yang, Z.; Tukahebwa, E.M.; Rowell, C.; Cooper, J.M. Paper-Based Microfluidics for DNA Diagnostics of Malaria in Low Resource Underserved Rural Communities. Proc. Natl. Acad. Sci. USA 2019, 116, 4834–4842. [Google Scholar] [CrossRef]
- Lee, R.A.; Puig, H.D.; Nguyen, P.Q.; Angenent-Mari, N.M.; Donghia, N.M.; McGee, J.P.; Dvorin, J.D.; Klapperich, C.M.; Pollock, N.R.; Collins, J.J. Ultrasensitive CRISPR-Based Diagnostic for Field-Applicable Detection of Plasmodium Species in Symptomatic and Asymptomatic Malaria. Proc. Natl. Acad. Sci. USA 2020, 117, 25722–25731. [Google Scholar] [CrossRef]
- Teengam, P.; Nisab, N.; Chuaypen, N.; Tangkijvanich, P.; Vilaivan, T.; Chailapakul, O. Fluorescent Paper-Based DNA Sensor Using Pyrrolidinyl Peptide Nucleic Acids for Hepatitis C Virus Detection. Biosens. Bioelectron. 2021, 189, 113381. [Google Scholar] [CrossRef] [PubMed]
- Anfossi, L.; Di Nardo, F.; Cavalera, S.; Giovannoli, C.; Baggiani, C. Multiplex Lateral Flow Immunoassay: An Overview of Strategies towards High-Throughput Point-of-Need Testing. Biosensors 2018, 9, 2. [Google Scholar] [CrossRef] [PubMed]
- Phillips, E.A.; Moehling, T.J.; Ejendal, K.F.K.; Hoilett, O.S.; Byers, K.M.; Basing, L.A.; Jankowski, L.A.; Bennett, J.B.; Lin, L.-K.; Stanciu, L.A.; et al. Microfluidic Rapid and Autonomous Analytical Device (MicroRAAD) to Detect HIV from Whole Blood Samples. Lab Chip 2019, 19, 3375–3386. [Google Scholar] [CrossRef] [PubMed]
- Martinez, A.W.; Phillips, S.T.; Whitesides, G.M.; Carrilho, E. Diagnostics for the Developing World: Microfluidic Paper-Based Analytical Devices. Anal. Chem. 2010, 82, 3–10. [Google Scholar] [CrossRef] [PubMed]
- Nishat, S.; Jafry, A.T.; Martinez, A.W.; Awan, F.R. Paper-Based Microfluidics: Simplified Fabrication and Assay Methods. Sens. Actuators B Chem. 2021, 336, 129681. [Google Scholar] [CrossRef]
- Koczula, K.M.; Gallotta, A. Lateral Flow Assays. Essays Biochem. 2016, 60, 111–120. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Shi, W.; Jiang, L.; Qin, J.; Lin, B. Rapid Prototyping of Paper-Based Microfluidics with Wax for Low-Cost, Portable Bioassay. Electrophoresis 2009, 30, 1497–1500. [Google Scholar] [CrossRef]
- Colozza, N.; Caratelli, V.; Moscone, D.; Arduini, F. Origami Paper-Based Electrochemical (Bio)Sensors: State of the Art and Perspective. Biosensors 2021, 11, 328. [Google Scholar] [CrossRef]
- Noviana, E.; Ozer, T.; Carrell, C.S.; Link, J.S.; McMahon, C.; Jang, I.; Henry, C.S. Microfluidic Paper-Based Analytical Devices: From Design to Applications. Chem. Rev. 2021, 121, 11835–11885. [Google Scholar] [CrossRef]
- Carrilho, E.; Martinez, A.W.; Whitesides, G.M. Understanding Wax Printing: A Simple Micropatterning Process for Paper-Based Microfluidics. Anal. Chem. 2009, 81, 7091–7095. [Google Scholar] [CrossRef]
- Lu, Y.; Shi, W.; Qin, J.; Lin, B. Fabrication and Characterization of Paper-Based Microfluidics Prepared in Nitrocellulose Membrane by Wax Printing. Anal. Chem. 2010, 82, 329–335. [Google Scholar] [CrossRef] [PubMed]
- Apilux, A.; Ukita, Y.; Chikae, M.; Chailapakul, O.; Takamura, Y. Development of Automated Paper-Based Devices for Sequential Multistep Sandwich Enzyme-Linked Immunosorbent Assays Using Inkjet Printing. Lab Chip 2013, 13, 126–135. [Google Scholar] [CrossRef] [PubMed]
- Yamada, K.; Henares, T.G.; Suzuki, K.; Citterio, D. Distance-Based Tear Lactoferrin Assay on Microfluidic Paper Device Using Interfacial Interactions on Surface-Modified Cellulose. ACS Appl. Mater. Interfaces 2015, 7, 24864–24875. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, R.; Gopalakrishnan, S.; Savitha, R.; Renganathan, T.; Pushpavanam, S. Fabrication of Laser Printed Microfluidic Paper-Based Analytical Devices (LP-ΜPADs) for Point-of-Care Applications. Sci. Rep. 2019, 9, 7896. [Google Scholar] [CrossRef]
- Ng, J.S.; Hashimoto, M. Fabrication of Paper Microfluidic Devices Using a Toner Laser Printer. RSC Adv. 2020, 10, 29797–29807. [Google Scholar] [CrossRef]
- Martinez-Duarte, R.; Madou, M.J. SU-8 photolithography and its impact on microfluidics. In Microfluidics and Nanofluidics Handbook; CRC Press: Boca Raton, FL, USA, 2011. [Google Scholar]
- Li, Z.; Bai, Y.; You, M.; Hu, J.; Yao, C.; Cao, L.; Xu, F. Fully Integrated Microfluidic Devices for Qualitative, Quantitative and Digital Nucleic Acids Testing at Point of Care. Biosens. Bioelectron. 2021, 177, 112952. [Google Scholar] [CrossRef] [PubMed]
- Ranjbaran, M.; Verma, M.S. Microfluidics at the Interface of Bacteria and Fresh Produce. Trends Food Sci. Technol. 2022, 128, 102–117. [Google Scholar] [CrossRef]
- Choi, J.R.; Hu, J.; Tang, R.; Gong, Y.; Feng, S.; Ren, H.; Wen, T.; Li, X.; Abas, W.A.B.W.; Pingguan-Murphy, B.; et al. An Integrated Paper-Based Sample-to-Answer Biosensor for Nucleic Acid Testing at the Point of Care. Lab Chip 2016, 16, 611–621. [Google Scholar] [CrossRef]
- Shin, J.H.; Hong, J.; Go, H.; Park, J.; Kong, M.; Ryu, S.; Kim, K.-P.; Roh, E.; Park, J.-K. Multiplexed Detection of Foodborne Pathogens from Contaminated Lettuces Using a Handheld Multistep Lateral Flow Assay Device. J. Agric. Food Chem. 2018, 66, 290–297. [Google Scholar] [CrossRef]
- Gétaz, M.; Bühlmann, A.; Schneeberger, P.H.H.; van Malderghem, C.; Duffy, B.; Maes, M.; Pothier, J.F.; Cottyn, B. A Diagnostic Tool for Improved Detection of Xanthomonas Fragariae Using a Rapid and Highly Specific LAMP Assay Designed with Comparative Genomics. Plant Pathol. 2017, 66, 1094–1102. [Google Scholar] [CrossRef]
| Fluorometric Reporters | Type of Reporter | Target Nucleic Acid | Excitation Wavelength | LoD (Copies per Reaction) | References |
|---|---|---|---|---|---|
| SYBR Green I | Fluorescent dye | DNA | Maximum excitation 497 nm Secondary excitation peaks at 290 and 380 nm. | 1–10 | [145,146] |
| Hydroxynaphthol blue (HNB) | Fluorescent dye | DNA | 530 nm | 4.1 × 102 | [97] |
| Green calcein | Fluorescent dye | DNA | 365 nm | 25 | [147] |
| Quantum dots | Nanoparticles | DNA | 375 nm | 1.12 × 106 | [148] |
| Fluorescent probes | Oligonucleotide probe | Plasmid DNA | 493 nm/580 nm | 10–100 | [149] |
| R-phycoerythrin and Fluorescein isothiocyanate (FITC) | Organic fluorophore | ssDNA | 496 nm and 495 nm, respectively | 115 | [150] |
| Detected Pathogen | Detection Method | Device Format | Reporting Method | Assay Time | Sample Processing | Sample Used to Determine LoD | LoD (Copies per Reaction) | Reference |
|---|---|---|---|---|---|---|---|---|
| SARS-CoV-2 | Nucleic acid hybridization | µPADs | Electrochemical | RNA extraction time + 5 min | RNA extraction | Extracted viral RNA | 2310 | [187] |
| SARS-CoV-2 | RT-LAMP; Cas12b | LFAs | Colorimetric (GNPs) | 45 min | RNA extraction | In vitro-transcribed RNA in water | 20 | [104] |
| SARS-CoV-2 | RT-LAMP; CRISPR–Cas12 | LFAs | Colorimetric (GNPs) | 90 min | RNA extraction, magnetic beads RNA concentration | Synthetic genomic standards spiked into saliva or nasopharyngeal (NP) swab | 100 | [77] |
| SARS-CoV-2 | RT-LAMP | µPADs | Colorimetric (Phenol red) | 60 min | Saliva dilution | Heat inactivated virus in saliva | 250 | [83] |
| SARS-CoV-2 | Toehold switch | µPADs | Bioluminescent (bioluminescent reporter protein) | RNA extraction + 15 min | RNA extraction | Synthetic RNA targets in saliva | 5.8 × 109 | [190] |
| Zika virus, Dengue virus | RPA; CRISPR–Cas13 | LFAs | Colorimetric (GNPs) | 120 min | RNA extraction | cDNA in water and urine | 20 | [191] |
| Zika virus, Dengue virus | RT-LAMP | µPADs | Fluorometric (Hydroxynaphthol blue) | 60 min | RNA extraction | RNA in 1X PBS and 100% human serum | 1 and 10 | [192] |
| Zika virus, Dengue virus, Chikungunya virus | RT-LAMP | µPADs &LFAs | Fluorometric (Hydroxynaphthol blue) | 60 min | RNA extraction | RNA in human serum | 5 | [185] |
| Plasmodium species | RT-LAMP | µPADs &LFAs | Colorimetric (Streptavidin labeled red particle) | 50 min | DNA extraction, magnetic beads DNA concentration | Genomic DNA in water | 32 | [193] |
| Plasmodium species | RT-RPA; CRISPR–Cas12 | LFAs | Colorimetric (GNPs) | 70 min | DNA extraction | Genomic DNA in blood or serum | 375 | [194] |
| Hepatitis B Virus | Pyrrolidinyl peptide nucleic acid (acpcPNA) | eLFAs | Electrochemical | DNA extraction + 7 min | DNA extraction | DNA in water | 4.3 × 106 | [137] |
| Hepatitis C Virus | acpcPNA | µPADs | Fluorometric (QuantiFluor®) | DNA extraction + 15 min | DNA extraction | Synthetic DNA oligonucleotide in water | 3.0 × 106 | [195] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, J.; Davidson, J.L.; Kaur, S.; Dextre, A.A.; Ranjbaran, M.; Kamel, M.S.; Athalye, S.M.; Verma, M.S. Paper-Based Biosensors for the Detection of Nucleic Acids from Pathogens. Biosensors 2022, 12, 1094. https://doi.org/10.3390/bios12121094
Wang J, Davidson JL, Kaur S, Dextre AA, Ranjbaran M, Kamel MS, Athalye SM, Verma MS. Paper-Based Biosensors for the Detection of Nucleic Acids from Pathogens. Biosensors. 2022; 12(12):1094. https://doi.org/10.3390/bios12121094
Chicago/Turabian StyleWang, Jiangshan, Josiah Levi Davidson, Simerdeep Kaur, Andres A. Dextre, Mohsen Ranjbaran, Mohamed S. Kamel, Shreya Milind Athalye, and Mohit S. Verma. 2022. "Paper-Based Biosensors for the Detection of Nucleic Acids from Pathogens" Biosensors 12, no. 12: 1094. https://doi.org/10.3390/bios12121094
APA StyleWang, J., Davidson, J. L., Kaur, S., Dextre, A. A., Ranjbaran, M., Kamel, M. S., Athalye, S. M., & Verma, M. S. (2022). Paper-Based Biosensors for the Detection of Nucleic Acids from Pathogens. Biosensors, 12(12), 1094. https://doi.org/10.3390/bios12121094





