Metabolic Assessment of Human Induced Pluripotent Stem Cells-Derived Astrocytes and Fetal Primary Astrocytes: Lactate and Glucose Turnover
Abstract
1. Introduction
2. Materials and Methods
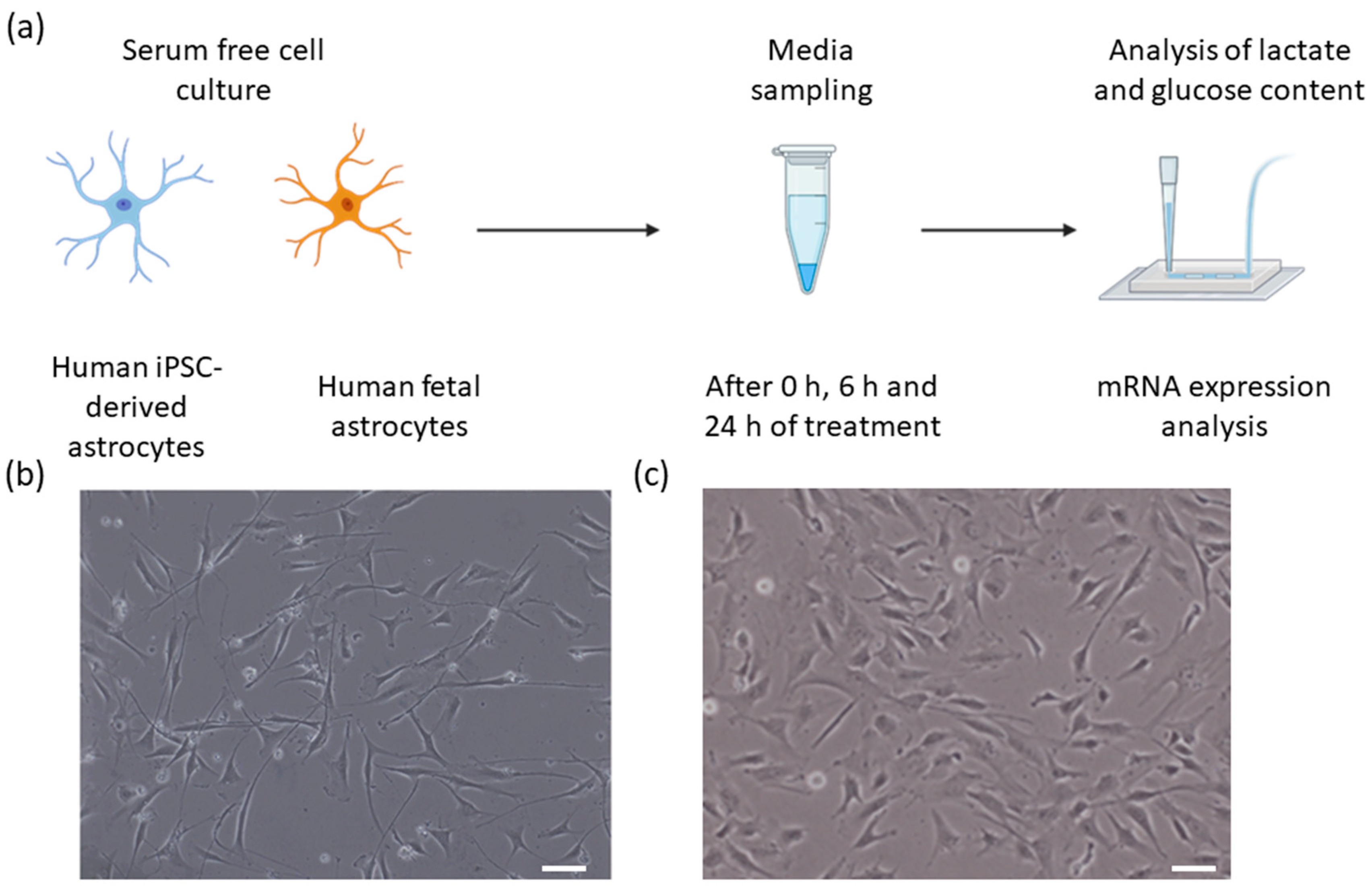
2.1. Cell Models/Source and Cell Culture
2.2. Sample Collection
2.3. mRNA and dsDNA Quantification
2.4. Lactate Assay
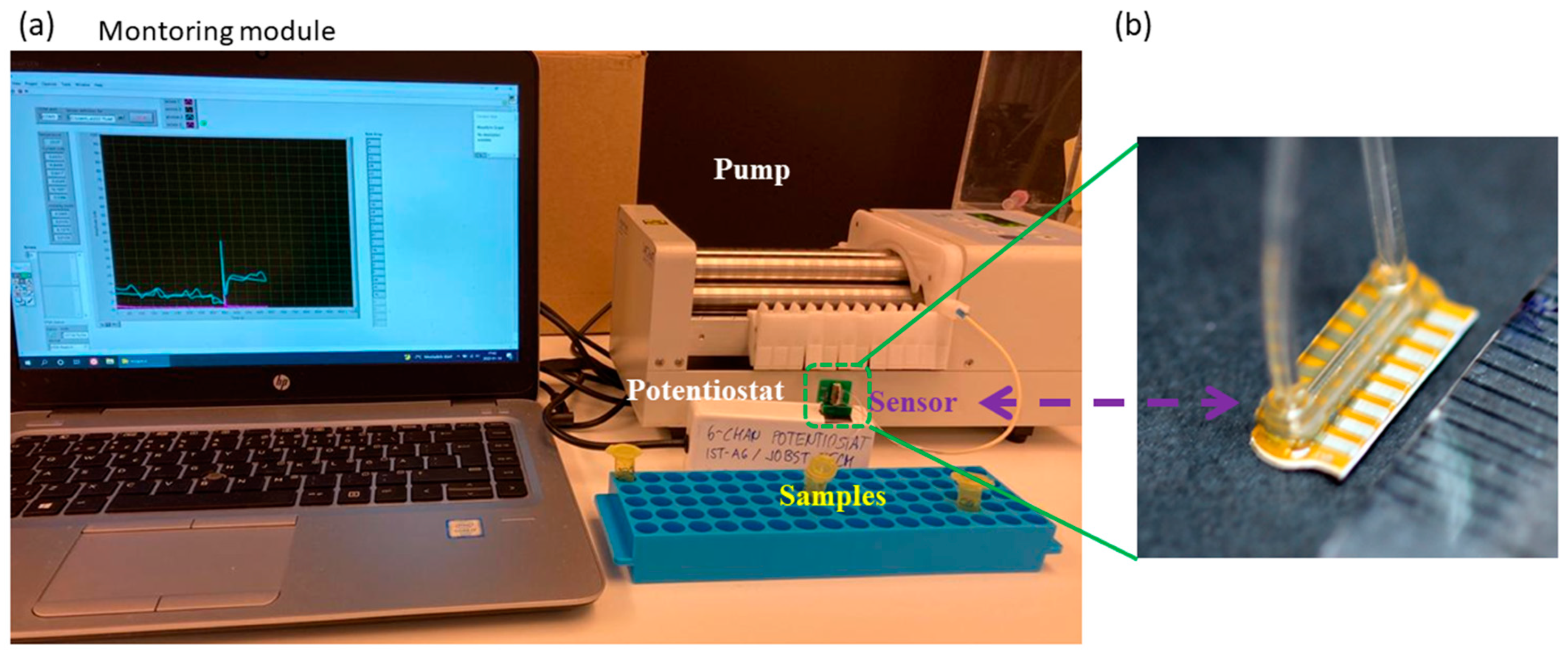
2.5. Biosensor Metabolic Readout
2.6. Statistical Analysis
3. Results and Discussion
3.1. mRNA Analysis
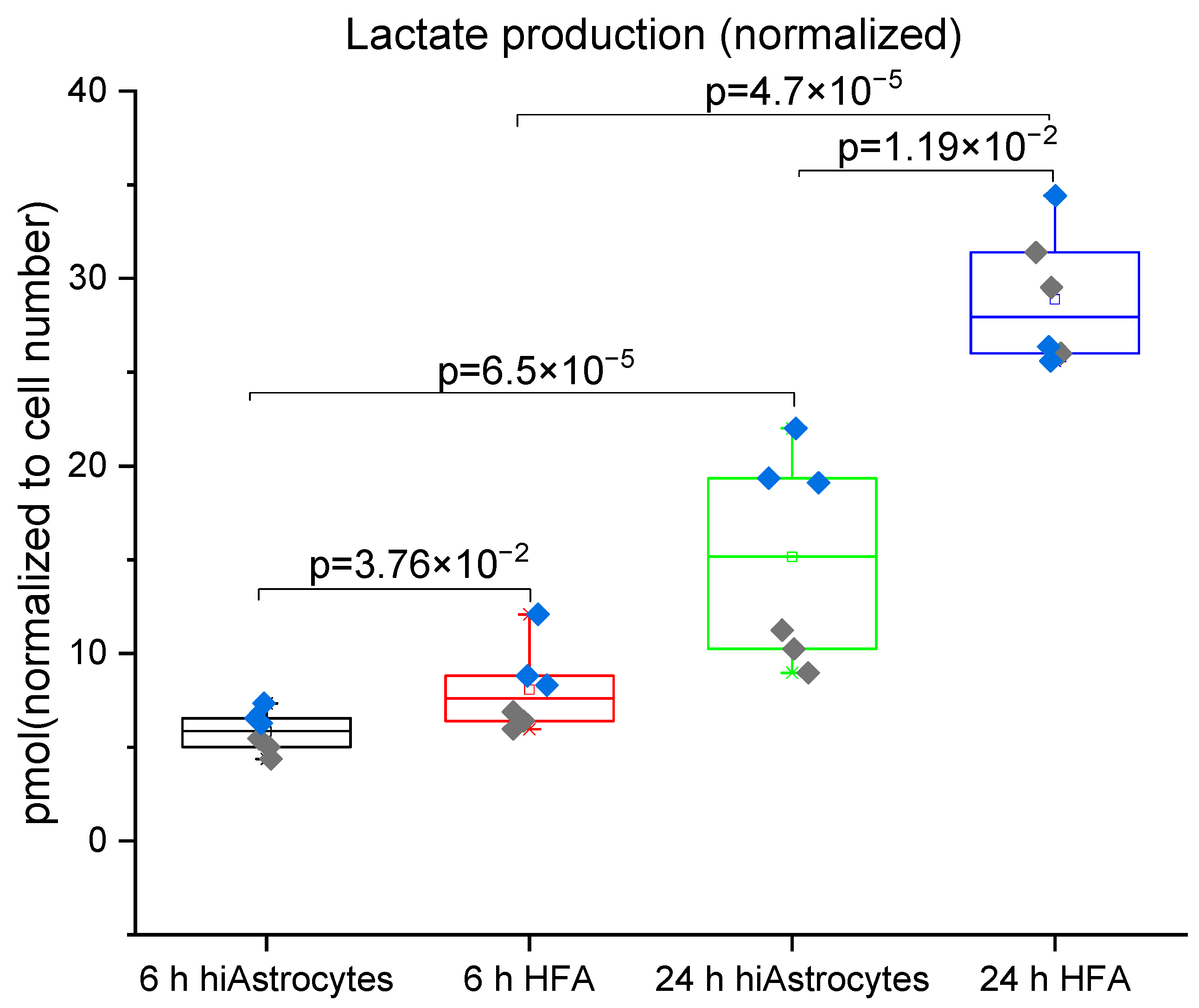
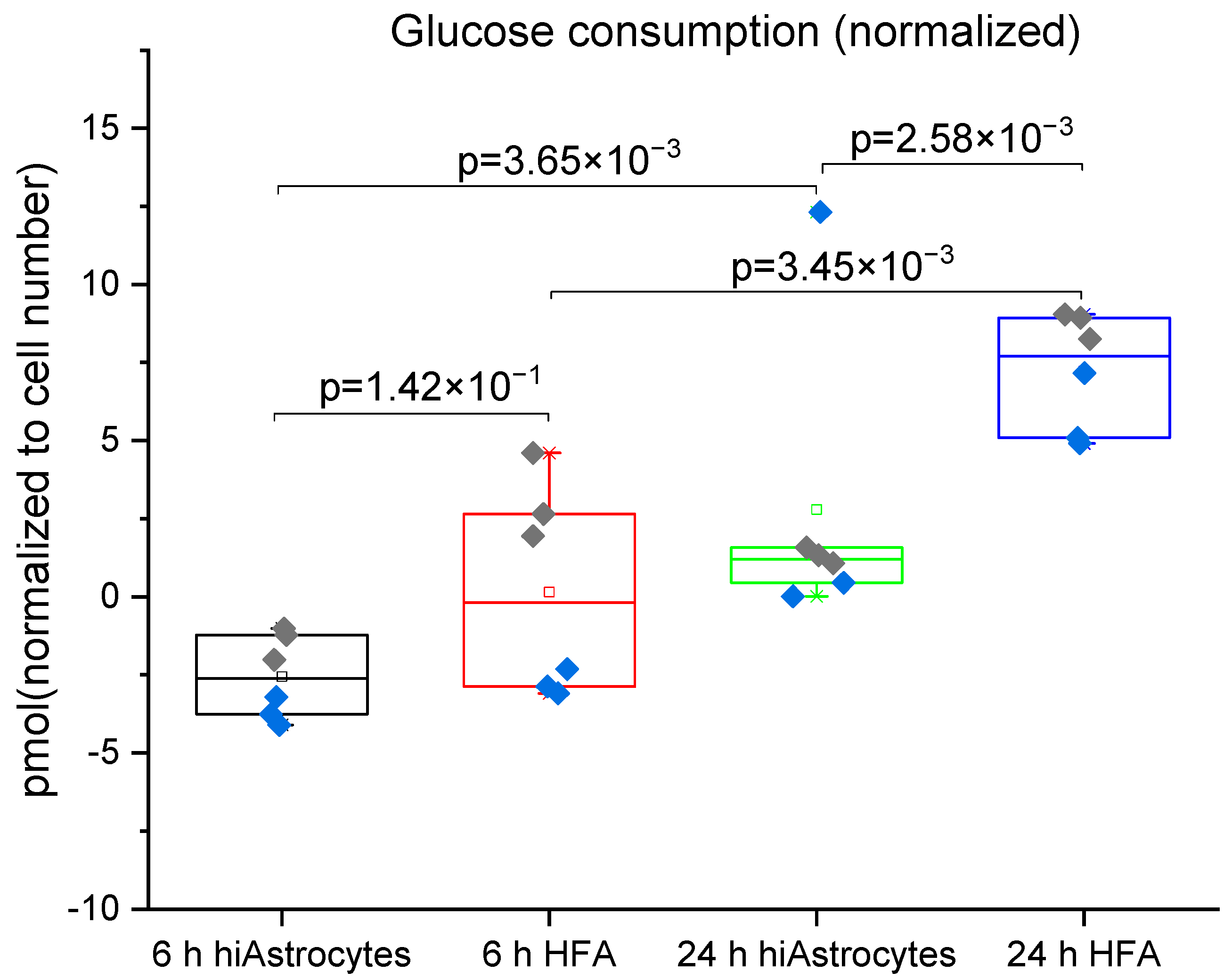
3.2. Glucose Consumption and Lactate Production
| Analyte | HFA (Our Study) | hiAstrocytes (Our Study) | HFA Ref. [4] | Human Hepatocytes Ref. [18] | Human Hepatocytes Ref. [17] |
|---|---|---|---|---|---|
| Glucose consumption rate [pmol/cell/h] | 0.398 ± 0.11 | 0.189 ± 0.05 | NA | 0.259 | 0.144 |
| Lactate production rate [pmol/cell/h] | 1.177 ± 0.15 | 0.510 ± 0.23 | 0.298 | 0.296 | 0.180 |
| Lactate/glucose ratio | 2.95 | 2.69 | NA | 1.14 | 1.25 |
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Rose, J.; Brian, C.; Pappa, A.; Panayiotidis, M.I.; Franco, R. Mitochondrial Metabolism in Astrocytes Regulates Brain Bioenergetics, Neurotransmission and Redox Balance. Front. Neurosci. 2020, 14, 536682. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.-Y.; Yang, Y.; Ju, W.-N.; Wang, X.; Zhang, H. Emerging Roles of Astrocytes in Neuro-Vascular Unit and the Tripartite Synapse With Emphasis on Reactive Gliosis in the Context of Alzheimer’s Disease. Front. Cell. Neurosci. 2018, 12, 193. [Google Scholar] [CrossRef] [PubMed]
- Boison, D.; Steinhäuser, C. Epilepsy and astrocyte energy metabolism. Glia 2018, 66, 1235–1243. [Google Scholar] [CrossRef] [PubMed]
- Herland, A.; Maoz, B.M.; FitzGerald, E.A.; Grevesse, T.; Vidoudez, C.; Sheehy, S.P.; Budnik, N.; Dauth, S.; Mannix, R.; Budnik, B.; et al. Proteomic and Metabolomic Characterization of Human Neurovascular Unit Cells in Response to Methamphetamine. Adv. Biosyst. 2020, 4, 1900230. [Google Scholar] [CrossRef] [PubMed]
- Chung, W.-S.; Allen, N.J.; Eroglu, C. Astrocytes control synapse formation, function, and elimination. Cold Spring Harb. Perspect. Biol. 2015, 7, a020370. [Google Scholar] [CrossRef]
- Deitmer, J.W.; Theparambil, S.M.; Ruminot, I.; Noor, S.I.; Becker, H.M. Energy Dynamics in the Brain: Contributions of Astrocytes to Metabolism and pH Homeostasis. Front. Neurosci. 2019, 13, 1301. [Google Scholar] [CrossRef]
- Magistretti, P.J.; Allaman, I. A Cellular Perspective on Brain Energy Metabolism and Functional Imaging. Neuron 2015, 86, 883–901. [Google Scholar] [CrossRef] [PubMed]
- Delsing, L.; Herland, A.; Falk, A.; Hicks, R.; Synnergren, J.; Zetterberg, H. Models of the blood-brain barrier using iPSC-derived cells. Mol. Cell. Neurosci. 2020, 107, 103533. [Google Scholar] [CrossRef]
- Lundin, A.; Delsing, L.; Clausen, M.; Ricchiuto, P.; Sanchez, J.; Sabirsh, A.; Ding, M.; Synnergren, J.; Zetterberg, H.; Brolén, G.; et al. Human iPS-Derived Astroglia from a Stable Neural Precursor State Show Improved Functionality Compared with Conventional Astrocytic Models. Stem Cell Rep. 2018, 10, 1030–1045. [Google Scholar] [CrossRef]
- Voulgaris, D.; Nikolakopoulou, P.; Herland, A. Generation of Human iPSC-Derived Astrocytes with a mature star-shaped phenotype for CNS modeling. Stem Cell Rev. Rep. 2022, 18, 2494–2512. [Google Scholar] [CrossRef] [PubMed]
- Azmi, A.; Azman, A.A.; Ibrahim, S.; Yunus, M.A. Techniques in Advancing the Capabilities of Various Nitrate Detection Methods: A Review. Int. J. Smart Sens. Intell. Syst. 2017, 10, 1–39. [Google Scholar] [CrossRef]
- Pontius, K.; Semenova, D.; Silina, Y.E.; Gernaey, K.V.; Junicke, H. Automated Electrochemical Glucose Biosensor Platform as an Efficient Tool Toward On-Line Fermentation Monitoring: Novel Application Approaches and Insights. Front. Bioeng. Biotechnol. 2020, 8, 436. [Google Scholar] [CrossRef] [PubMed]
- Schaller-Ammann, R.; Kreß, S.; Feiel, J.; Schwagerle, G.; Priedl, J.; Birngruber, T.; Kasper, C.; Egger, D. Advanced Online Monitoring of In Vitro Human 3D Full-Thickness Skin Equivalents. Harmaceutics 2022, 14, 1436. [Google Scholar] [CrossRef] [PubMed]
- Sniegowski, T.; Korac, K.; Bhutia, Y.; Ganapathy, V. SLC6A14 and SLC38A5 Drive the Glutaminolysis and Serine–Glycine–One-Carbon Pathways in Cancer. Pharmaceuticals 2021, 14, 216. [Google Scholar] [CrossRef]
- Jin, L.; Alesi, G.N.; Kang, S. Glutaminolysis as a target for cancer therapy. Oncogene 2016, 35, 3619–3625. [Google Scholar] [CrossRef]
- McKenna, M.C. Substrate Competition Studies Demonstrate Oxidative Metabolism of Glucose, Glutamate, Glutamine, Lactate and 3-Hydroxybutyrate in Cortical Astrocytes from Rat Brain. Neurochem. Res. 2012, 37, 2613–2626. [Google Scholar] [CrossRef]
- Bavli, D.; Prill, S.; Ezra, E.; Levy, G.; Cohen, M.; Vinken, M.; Vanfleteren, J.; Jaeger, M.; Nahmias, Y. Real-time monitoring of metabolic function in liver-on-chip microdevices tracks the dynamics of mitochondrial dysfunction. Proc. Natl. Acad. Sci. USA 2016, 113, E2231–E2240. [Google Scholar] [CrossRef] [PubMed]
- Dong, J.; Mandenius, C.-F.; Lübberstedt, M.; Urbaniak, T.; Nüssler, A.K.N.; Knobeloch, D.; Gerlach, J.C.; Zeilinger, K. Evaluation and optimization of hepatocyte culture media factors by design of experiments (DoE) methodology. Cytotechnology 2008, 57, 251–261. [Google Scholar] [CrossRef][Green Version]


| Metabolic Enzymes | |
|---|---|
| BDH2 | 3-Hydroxybuturate dehydrogenase 2, initiates ketolysis |
| ABAT | 4-Aminobutyrate aminotransferase, catabolizes GABA into succinic semialdehyde. |
| HMGCL | 3-Hydroxy-3-methylglutaryl-CoA lyase, catalyzes the formation of acetoacetate from HMG-CoA |
| OXCT1 | 3-Oxoacid CoA-Transferase 1, catalyzes CoA from succinyl-CoA to acetoacetate |
| Transporters | |
| SLC6A1 | Solute carrier family 6 member 1, GABA transporter, GABA reuptake in presynaptic neurons and astrocytes |
| SLC16A1 | Solute carrier family 16 member 1, Monocarboxylate transporter, suggested to transport lactate in astrocytes |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Matthiesen, I.; Nasiri, R.; Tamashiro Orrego, A.; Winkler, T.E.; Herland, A. Metabolic Assessment of Human Induced Pluripotent Stem Cells-Derived Astrocytes and Fetal Primary Astrocytes: Lactate and Glucose Turnover. Biosensors 2022, 12, 839. https://doi.org/10.3390/bios12100839
Matthiesen I, Nasiri R, Tamashiro Orrego A, Winkler TE, Herland A. Metabolic Assessment of Human Induced Pluripotent Stem Cells-Derived Astrocytes and Fetal Primary Astrocytes: Lactate and Glucose Turnover. Biosensors. 2022; 12(10):839. https://doi.org/10.3390/bios12100839
Chicago/Turabian StyleMatthiesen, Isabelle, Rohollah Nasiri, Alessandra Tamashiro Orrego, Thomas E. Winkler, and Anna Herland. 2022. "Metabolic Assessment of Human Induced Pluripotent Stem Cells-Derived Astrocytes and Fetal Primary Astrocytes: Lactate and Glucose Turnover" Biosensors 12, no. 10: 839. https://doi.org/10.3390/bios12100839
APA StyleMatthiesen, I., Nasiri, R., Tamashiro Orrego, A., Winkler, T. E., & Herland, A. (2022). Metabolic Assessment of Human Induced Pluripotent Stem Cells-Derived Astrocytes and Fetal Primary Astrocytes: Lactate and Glucose Turnover. Biosensors, 12(10), 839. https://doi.org/10.3390/bios12100839







