Biosensors for Detection of Biochemical Markers Relevant to Osteoarthritis
Abstract
1. Introduction
2. Materials and Methods
2.1. Eligibility Criteria and Search Strategy
2.2. Study Selection and Data Collection
2.3. Quality Assessment
2.4. Data Synthesis and Analysis
3. Results
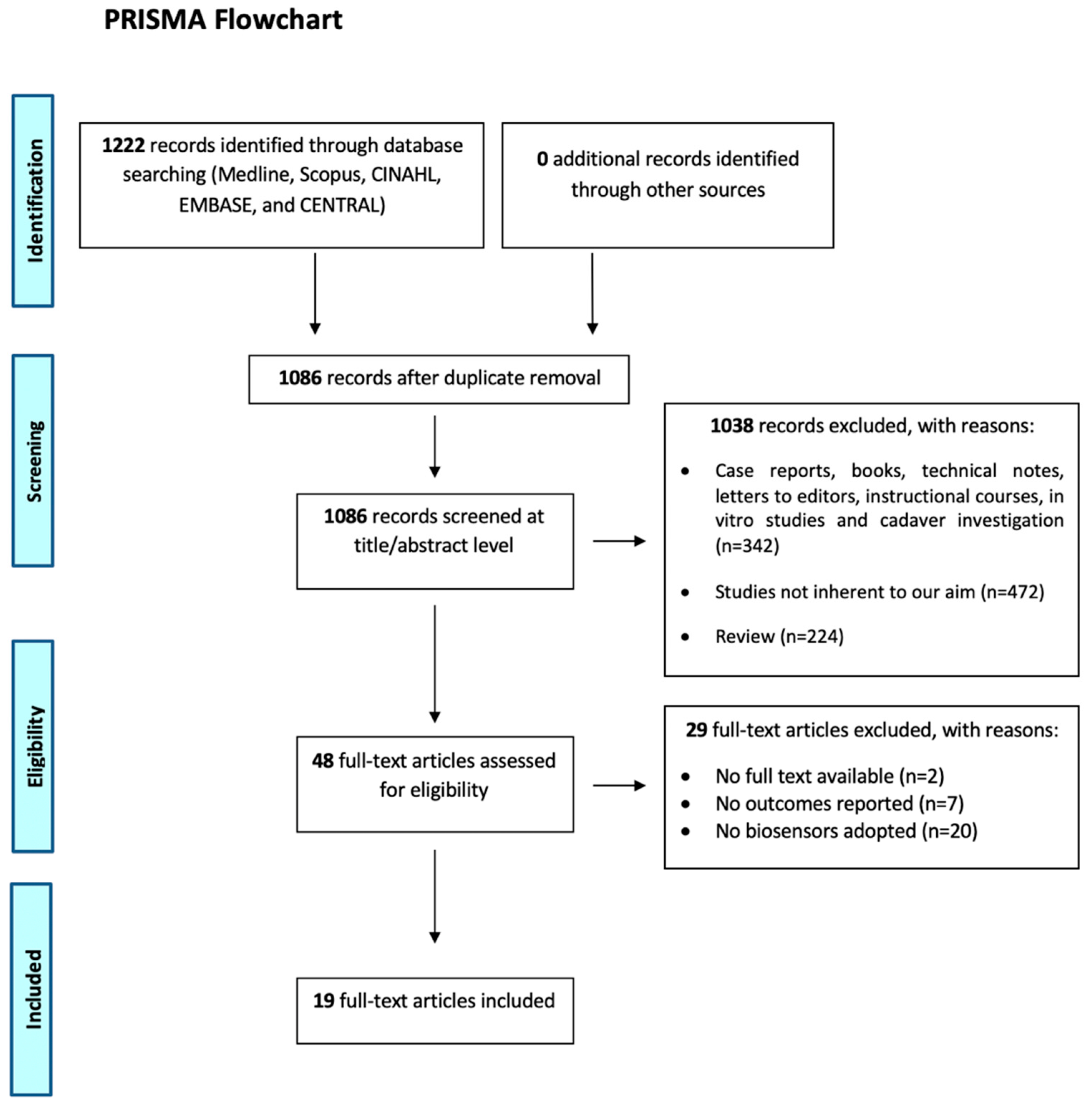
3.1. Study Selection and Patient Characteristics
3.2. Quality Assessment
3.3. Results of Individual Studies
3.3.1. Outcome: Accuracy
3.3.2. Outcome: Rapidity of Diagnosis
3.3.3. Outcome: Costs
3.3.4. Outcome: Ease of Use
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AMPK | AMP-activated protein kinase |
| COMP | Cartilage oligomeric matrix protein |
| CRP | C-reactive protein |
| DdRFP | matriptase sensitive protein biosensor based on dimerization-dependent red fluorescent protein |
| ECM | Articular cartilage extracellular matrix |
| ELISA | Enzyme-Linked Immunosorbent Assays |
| FBG | New fiber Bragg grating |
| FCD | fluid control device |
| FMGC | Fluoro-microbead guiding chip |
| FO-PPR | Fiber optic-particle plasmon resonance biosensor |
| GPI | antibodies against glucose 6-phosphate isomerase |
| IDE | Immunoassay with the specific antibody for uCTX-II |
| MIP | Molecular Imprinted Polymer sensor |
| MMP-1 | Matrix metalloproteinase 1 |
| MMP-3 | Matrix metalloproteinase 3 |
| OA | osteoarthritis |
| QCM | Quartz crystal microbalance |
| SAM | Biosensor based on label-free immuno-sensing with self-assembled monolayer |
| sCTx-I | Serum C-terminal telopeptide of type I collagen |
| sCTX-II | Serum C-terminal telopeptide fragment of type II collagen |
| SPRi | Plasmon resonance biosensor |
| TNF-a | Tumor necrosis factor. |
| uCTX-II | Urinary C-terminal telopeptide fragment of type II collagen |
References
- Blanco, F.J.; Rego-Pérez, I. Mitochondria and mitophagy: Biosensors for cartilage degradation and osteoarthritis. Osteoarthr. Cartil. 2018, 26, 989–991. [Google Scholar] [CrossRef]
- Wang, S.; Su, S.; Yu, C.; Gopinath, S.C.B.; Yang, Z. Immunodetection of urinary C-terminal telopeptide fragment of type II collagen: An osteoarthritis biomarker analysis. Biotechnol. Appl. Biochem. 2020. [Google Scholar] [CrossRef]
- Hosnijeh, F.S.; Siebuhr, A.S.; Uitterlinden, A.G.; Oei, E.H.; Hofman, A.; Karsdal, M.A.; Bierma-Zeinstra, S.M.; Bay-Jensen, A.C.; van Meurs, J.B. Association between biomarkers of tissue inflammation and progression of osteoarthritis: Evidence from the Rotterdam study cohort. Arthritis Res. Ther. 2016, 18, 81. [Google Scholar] [CrossRef]
- Lai, Y.; Yu, X.P.; Zhang, Y.; Tian, Q.; Song, H.; Mucignat, M.T.; Perris, R.; Samuels, J.; Krasnokutsky, S.; Attur, M.; et al. Enhanced COMP catabolism detected in serum of patients with arthritis and animal disease models through a novel capture ELISA. Osteoarthr. Cartil. 2012, 20, 854–862. [Google Scholar] [CrossRef][Green Version]
- Wang, S.H.; Shen, C.Y.; Weng, T.C.; Lin, P.H.; Yang, J.J.; Chen, I.F.; Kuo, S.M.; Chang, S.J.; Tu, Y.K.; Kao, Y.H.; et al. Detection of cartilage oligomeric matrix protein using a quartz crystal microbalance. Sensors 2010, 10, 11633–11643. [Google Scholar] [CrossRef]
- Bernotiene, E.; Bagdonas, E.; Kirdaite, G.; Bernotas, P.; Kalvaityte, U.; Uzieliene, I.; Thudium, C.S.; Hannula, H.; Lorite, G.S.; Dvir-Ginzberg, M.; et al. Emerging Technologies and Platforms for the Immunodetection of Multiple Biochemical Markers in Osteoarthritis Research and Therapy. Front. Med. 2020, 7, 572977. [Google Scholar] [CrossRef]
- Creemers, L.B.; Rutgers, M.; Saris, D.B. Evaluation of histological scoring systems for tissue-engineered, repaired and osteoarthritic cartilage. Osteoarthr. Cartil. 2010, 18, P1002. [Google Scholar] [CrossRef][Green Version]
- Song, S.Y.; Han, Y.D.; Hong, S.Y.; Kim, K.; Yang, S.S.; Min, B.H.; Yoon, H.C. Chip-based cartilage oligomeric matrix protein detection in serum and synovial fluid for osteoarthritis diagnosis. Anal. Biochem. 2012, 420, 139–146. [Google Scholar] [CrossRef]
- Mobasheri, A. Osteoarthritis year 2012 in review: Biomarkers. Osteoarthr. Cartil. 2012, 20, 1451–1464. [Google Scholar] [CrossRef][Green Version]
- Park, Y.M.; Kim, S.J.; Lee, K.J.; Yang, S.S.; Min, B.H.; Yoon, H.C. Detection of CTX-II in serum and urine to diagnose osteoarthritis by using a fluoro-microbeads guiding chip. Biosens. Bioelectron. 2015, 67, 192–199. [Google Scholar] [CrossRef]
- Filardo, G.; Kon, E.; Longo, U.G.; Madry, H.; Marchettini, P.; Marmotti, A.; Van Assche, D.; Zanon, G.; Peretti, G.M. Non-surgical treatments for the management of early osteoarthritis. Knee Surg. Sports Traumatol. Arthrosc. 2016, 24, 1775–1785. [Google Scholar] [CrossRef]
- Kim, J.Y.; Lee, M.H.; Jung, K.I.; Na, H.Y.; Cha, H.S.; Ko, E.M.; Kim, T.J. Detection of antibodies against glucose 6-phosphate isomerase in synovial fluid of rheumatoid arthritis using surface plasmon resonance (BIAcore). Exp. Mol. Med. 2003, 35, 310–316. [Google Scholar] [CrossRef]
- Camacho-Encina, M.; Balboa-Barreiro, V.; Rego-Perez, I.; Picchi, F.; VanDuin, J.; Qiu, J.; Fuentes, M.; Oreiro, N.; LaBaer, J.; Ruiz-Romero, C.; et al. Discovery of an autoantibody signature for the early diagnosis of knee osteoarthritis: Data from the Osteoarthritis Initiative. Ann. Rheum. Dis. 2019, 78, 1699–1705. [Google Scholar] [CrossRef]
- Afsarimanesh, N.; Mukhopadhyay, S.C.; Kruger, M. Molecularly Imprinted Polymer-Based Electrochemical Biosensor for Bone Loss Detection. IEEE Trans. Biomed. Eng. 2018, 65, 1264–1271. [Google Scholar] [CrossRef]
- Clark, L.C.; Lyons, C. Electrode systems for continuous monitoring in cardiovascular surgery. Ann. N. Y. Acad. Sci. 1962, 102, 29–45. [Google Scholar] [CrossRef]
- Grieshaber, D.; MacKenzie, R.; Vörös, J.; Reimhult, E. Electrochemical Biosensors—Sensor Principles and Architectures. Sensors 2008, 8, 1400–1458. [Google Scholar] [CrossRef]
- Rodovalho, V.R.; Araujo, G.R.; Vaz, E.R.; Ueira-Vieira, C.; Goulart, L.R.; Madurro, J.M.; Brito-Madurro, A.G. Peptide-based electrochemical biosensor for juvenile idiopathic arthritis detection. Biosens. Bioelectron. 2018, 100, 577–582. [Google Scholar] [CrossRef]
- Thévenot, D.R.; Toth, K.; Durst, R.A.; Wilson, G.S. Electrochemical biosensors: Recommended definitions and classification. Biosens. Bioelectron. 2001, 16, 121–131. [Google Scholar] [CrossRef]
- Juska, V.B.; Pemble, M.E. A Critical Review of Electrochemical Glucose Sensing: Evolution of Biosensor Platforms Based on Advanced Nanosystems. Sensors 2020, 20, 6013. [Google Scholar] [CrossRef]
- D’Orazio, P. Biosensors in clinical chemistry. Clin. Chim. Acta 2003, 334, 41–69. [Google Scholar] [CrossRef]
- Kim, J.; Campbell, A.S.; de Ávila, B.E.; Wang, J. Wearable biosensors for healthcare monitoring. Nat. Biotechnol. 2019, 37, 389–406. [Google Scholar] [CrossRef]
- Duk Han, Y.; Jin Chun, H.; Yoon, H.C. The transformation of common office supplies into a low-cost optical biosensing platform. Biosens. Bioelectron. 2014, 59, 259–268. [Google Scholar] [CrossRef]
- Bwambok, D.K.; Siraj, N.; Macchi, S.; Larm, N.E.; Baker, G.A.; Pérez, R.L.; Ayala, C.E.; Walgama, C.; Pollard, D.; Rodriguez, J.D.; et al. QCM Sensor Arrays, Electroanalytical Techniques and NIR Spectroscopy Coupled to Multivariate Analysis for Quality Assessment of Food Products, Raw Materials, Ingredients and Foodborne Pathogen Detection: Challenges and Breakthroughs. Sensors 2020, 20, 6982. [Google Scholar] [CrossRef]
- Mitchell, A.C.; Alford, S.C.; Hunter, S.A.; Kannan, D.; Sperberg, R.A.P.; Chang, C.H.; Cochran, J.R. Development of a Protease Biosensor Based on a Dimerization-Dependent Red Fluorescent Protein. ACS Chem. Biol. 2018, 13, 66–72. [Google Scholar] [CrossRef]
- Parthasarathy, P.; Vivekanandan, S. Investigation on uric acid biosensor model for enzyme layer thickness for the application of arthritis disease diagnosis. Health Inf. Sci. Syst. 2018, 6, 5. [Google Scholar] [CrossRef]
- Couto, R.A.; Lima, J.L.; Quinaz, M.B. Recent developments, characteristics and potential applications of screen-printed electrodes in pharmaceutical and biological analysis. Talanta 2016, 146, 801–814. [Google Scholar] [CrossRef]
- Ahn, K.Y.; Kwon, K.; Huh, J.; Kim, G.T.; Lee, E.B.; Park, D.; Lee, J. A sensitive diagnostic assay of rheumatoid arthritis using three-dimensional ZnO nanorod structure. Biosens. Bioelectron. 2011, 28, 378–385. [Google Scholar] [CrossRef]
- Khan, W.S.; Longo, U.G. The Role of Surgery in Osteoarthritis, Rheumatoid Arthritis and Ankylosing Spondylitis. Open Orthop. J. 2015, 9, 460–462. [Google Scholar] [CrossRef]
- Liberati, A.; Altman, D.G.; Tetzlaff, J.; Mulrow, C.; Gotzsche, P.C.; Ioannidis, J.P.; Clarke, M.; Devereaux, P.J.; Kleijnen, J.; Moher, D. The PRISMA statement for reporting systematic reviews and meta-analyses of studies that evaluate health care interventions: Explanation and elaboration. PLoS Med. 2009, 6, e1000100. [Google Scholar] [CrossRef]
- Slim, K.; Nini, E.; Forestier, D.; Kwiatkowski, F.; Panis, Y.; Chipponi, J. Methodological index for non-randomized studies (minors): Development and validation of a new instrument. ANZ J. Surg. 2003, 73, 712–716. [Google Scholar] [CrossRef]
- Hsu, W.T.; Hsieh, W.H.; Cheng, S.F.; Jen, C.P.; Wu, C.C.; Li, C.H.; Lee, C.Y.; Li, W.Y.; Chau, L.K.; Chiang, C.Y.; et al. Integration of fiber optic-particle plasmon resonance biosensor with microfluidic chip. Anal. Chim. Acta 2011, 697, 75–82. [Google Scholar] [CrossRef]
- Huang, Y.C.; Chiang, C.Y.; Li, C.H.; Chang, T.C.; Chiang, C.S.; Chau, L.K.; Huang, K.W.; Wu, C.W.; Wang, S.C.; Lyu, S.R. Quantification of tumor necrosis factor-α and matrix metalloproteinases-3 in synovial fluid by a fiber-optic particle plasmon resonance sensor. Analyst 2013, 138, 4599–4606. [Google Scholar] [CrossRef]
- Hartmann, B.; Marchi, G.; Alberton, P.; Farkas, Z.; Aszodi, A.; Roths, J.; Clausen-Schaumann, H. Early Detection of Cartilage Degeneration: A Comparison of Histology, Fiber Bragg Grating-Based Micro-Indentation, and Atomic Force Microscopy-Based Nano-Indentation. Int. J. Mol. Sci. 2020, 21, 7384. [Google Scholar] [CrossRef]
- Ahmad, N.; Colak, B.; Zhang, D.W.; Gibbs, M.J.; Watkinson, M.; Becer, C.R.; Gautrot, J.E.; Krause, S. Peptide Cross-Linked Poly (Ethylene Glycol) Hydrogel Films as Biosensor Coatings for the Detection of Collagenase. Sensors 2019, 19, 1677. [Google Scholar] [CrossRef]
- Chiang, C.Y.; Hsieh, M.L.; Huang, K.W.; Chau, L.K.; Chang, C.M.; Lyu, S.R. Fiber-optic particle plasmon resonance sensor for detection of interleukin-1β in synovial fluids. Biosens. Bioelectron. 2010, 26, 1036–1042. [Google Scholar] [CrossRef]
- Vance, S.A.; Sandros, M.G. Zeptomole detection of C-reactive protein in serum by a nanoparticle amplified surface plasmon resonance imaging aptasensor. Sci. Rep. 2014, 4, 5129. [Google Scholar] [CrossRef]
- Yun, Y.H.; Bhattacharya, A.; Watts, N.B.; Schulz, M.J. A label-free electronic biosensor for detection of bone turnover markers. Sensors 2009, 9, 7957–7969. [Google Scholar] [CrossRef]
- Park, Y.M.; Han, Y.D.; Chun, H.J.; Yoon, H.C. Ambient light-based optical biosensing platform with smartphone-embedded illumination sensor. Biosens. Bioelectron. 2017, 93, 205–211. [Google Scholar] [CrossRef]
- Chen, L.Y.; Wang, Y.; Terkeltaub, R.; Liu-Bryan, R. Activation of AMPK-SIRT3 signaling is chondroprotective by preserving mitochondrial DNA integrity and function. Osteoarthr. Cartil. 2018, 26, 1539–1550. [Google Scholar] [CrossRef]
- Vina, E.R.; Kwoh, C.K. Epidemiology of osteoarthritis: Literature update. Curr. Opin. Rheumatol. 2018, 30, 160–167. [Google Scholar] [CrossRef]
- Bergmann, J.H.; Chandaria, V.; McGregor, A. Wearable and implantable sensors: The patient’s perspective. Sensors 2012, 12, 16695–16709. [Google Scholar] [CrossRef]
- Liu, Y.; Peng, H.; Meng, Z.; Wei, M. Correlation of IL-17 Level in Synovia and Severity of Knee Osteoarthritis. Med. Sci. Monit. 2015, 21, 1732–1736. [Google Scholar] [CrossRef]
- Sharif, M.; Saxne, T.; Shepstone, L.; Kirwan, J.R.; Elson, C.J.; Heinegård, D.; Dieppe, P.A. Relationship between serum cartilage oligomeric matrix protein levels and disease progression in osteoarthritis of the knee joint. Br. J. Rheumatol. 1995, 34, 306–310. [Google Scholar] [CrossRef]
| Author and Year | Type of Study and Level of Evidence | Sample Test | Biosensor | Biochemical Markers | Characteristics | Advantages |
|---|---|---|---|---|---|---|
| Afsarimanesh 2017 [14] | Case-control study, Level III | Human serum | MIP sensor | sCTx-I | between 0.1 and 2.5 ng/mL | The proposed biosensorexhibited good selectivity and quick rebinding capacity towards target molecules. |
| Ahmad 2019 [34] | Cross-sectional study, Level III | Synovial fluid | Quartz crystal microbalance biosensor. | MMP-1 | Between 2 to 2000 nM | Reaction time advantage |
| Chen 2018 [39] | Cross-sectional study, Level III | DNA is extracted with the D-Neasy Blood & Tissue kit | Metabolic biosensor AMPK | Mitochondrial DNA | AMPK activation limits oxidative stress and improves mitochondrial DNA integrity and function in OA chondrocytes. | |
| Chiang 2010 [35] | Cross-sectional study, Level III | Synovial fluid | FOPPR | Interleukin-1B | 0.050–10 ng/mL | High sensitivity |
| Duk Han 2014 [22] | Case-control, Level III | uCTX-II controls | Ultraviolet-visible spectroscopy | CTX-II | Detection range: 1.3–10 ng/mL | This biosensor has high sensitivity, facile fabrication, and the high obtainability and cost-effectiveness of the components used to make it |
| Hartmann 2020 [33] | Cross-sectional, Level III | Bovine articular cartilage | FBG-based optoelectronic micro-indenter | ECM | 5, 50, 100 and 500 μg/mL | High sensitivity |
| Hsu 2011 [31] | Case-control study, Level III | Synovial fluid | FO-PPR | MMP-3 | A low-cost and portable biosensor | |
| Huang 2013 [32] | Cross-sectional study, Level III | Synovial fluid | FO-PPR | TNF and MMP-3 | TNF-a: 8.2 pg/mL; MMP-3: 8.2 pg/mL | Reaction time advantage, simple usage, high sensitivity, high selectivity |
| Kim 2003 [12] | Case-control study, Level III | Synovial fluid | SPRi | GPI fused with or without NusA | Increased solubility in recombinant protein production | |
| Lai 2012 [4] | Cross-sectional study, Level III | Human serum | monoclonal antibodies against COMP fragments | COMP | Between 10 to 100 ng/mL | A significant increase in the COMP fragments was noted in the serum of OA patients assayed by this new sensor |
| Mitchell 2018 [24] | Cross-sectional study, Level III | Epithelial cells | DdRFP; | Protease matriptase | Between 0 to 750 nM | Low cost of production, high dynamic range, robust activity under physiological and non-physiological conditions, and ideal spectroscopic properties |
| Park 2015 [10] | Case-control study, Level III | Human serum and urine | FMGC; FCD | uCTX-II; SCTX-II; | uCTX-II: 200–1400 ng/mmol; sCTX-II: 0.1–2.0 ng/mL | Effectively and quantitatively assessed urinary and sCTX-II simultaneously |
| Park 2016 [38] | Cross-sectional study, Level III | uCTX-II epitope-controls | Hand-held optical biosensing system utilizing a smartphone-embedded illumination sensor that is integrated with immuno-blotting assay method | uCTX-II | LOD: 0.2 ng/mL | Simple to operate, thus allowing its use by untrained and non-medical profession personnel; an immediate and accurate analysis without the use of professional equipment and special software under various ambient light conditions |
| Parthasarathy 2018 [25] | Cross-sectional study, Level III | Not reported | Amperometric biosensor | Uricase enzyme layer thickness | Diagnosis can be made by seeing the change Uricase enzyme layer thickness | |
| Song 2011 [8] | Cross-sectional study, Level III | Human blood and synovial fluid | FMGC | COMP | Between 4 and 128 ng/mL | Ease and accuracy of biomarker quantification over a clinically important concentration range. Reaction time advantage |
| Vance 2014 [36] | Cross-sectional study, Level III | Human serum | Ultrasensitive SPRi biosensors | CRP | 5 fg/mL | Ultra-sensitiveSPRi biosensors offer fast turnaround time and a stronger support structure for the capture probe |
| Wang 2020 [2] | Case-control study, Level III | Urine | IDE | uCTX-II | Between 10 and 100 pM | uCTX-II has been found to be a rapidly potential biomarker for OA. |
| Wang 2010 [5] | Cross-sectional study, Level III | Urine | QCM | COMP | Range 1–200 ng/mL | A highly sensitive, user-friendly and cost-effective analytical method for early-stage diagnosis |
| Yun 2009 [37] | Case-control study, Level III | Urine | SAM | CTX-II | Between 3 μg/mL to 50 ng/mL | Reaction time advantages |
| Author | Clearly Stated Aim | Inclusion of Consecutive Patients | Prospective Data Collection | Endpoints Appropriate to Study Aim | Unbiased Assessment of Study Endpoint | Follow-Up Period Appropriate to Study Aim | <5% Lost to Follow-Up | Prospective Calculation of Study Size | Adequate Control Group | Contemporary Groups | Baseline Equivalence of Groups | Adequate Statistical Analyses | Total Score (…/24) |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Afsarimanesh, 2017 | 2 | NA | 0 | 2 | 2 | 0 | 0 | 0 | 2 | 2 | 0 | 0 | 10 |
| Ahmad, 2019 | 2 | 2 | 0 | 2 | 1 | 0 | 0 | NA | 2 | 2 | 0 | 0 | 11 |
| Chen, 2018 | 2 | 0 | 0 | 2 | 2 | 0 | 0 | 0 | 2 | 2 | 0 | 2 | 12 |
| Chiang, 2010 | 2 | 2 | 0 | 2 | 2 | 0 | 0 | 0 | 2 | 2 | 0 | 2 | 14 |
| Duk Han, 2014 | 2 | 2 | 0 | 2 | 2 | 0 | NA | 0 | 2 | 2 | 0 | 0 | 12 |
| Hartmann, 2020 | 2 | 2 | 0 | 2 | 2 | 0 | 0 | 0 | 2 | 2 | 0 | 2 | 14 |
| Hsu, 2011 | 2 | 2 | NA | 2 | 2 | 0 | 0 | 0 | 2 | 2 | 0 | 0 | 12 |
| Huang, 2013 | 2 | 2 | 0 | 2 | 0 | 0 | 0 | 0 | 2 | 2 | 0 | 2 | 12 |
| Kim, 2003 | 2 | 0 | 2 | 2 | 2 | 0 | 0 | 0 | 2 | 2 | 0 | 2 | 14 |
| Lai, 2012 | 2 | 2 | 0 | 2 | 2 | 0 | 0 | 2 | 2 | 2 | NA | 2 | 16 |
| Mitchel, 2018 | 2 | 0 | 0 | 2 | 2 | NA | NA | NA | 2 | 2 | 0 | 2 | 12 |
| Park, 2015 | 2 | 2 | 0 | 2 | 0 | 0 | 0 | 0 | 2 | 2 | 0 | 0 | 10 |
| Park, 2016 | 2 | 2 | 0 | 2 | 0 | 0 | 0 | 0 | 2 | 2 | 2 | 2 | 14 |
| Parthasarathy, 2018 | 2 | 0 | 0 | 2 | 2 | 0 | 0 | 0 | 2 | 2 | 0 | 0 | 10 |
| Song, 2011 | 2 | 0 | 0 | 2 | 2 | 0 | 0 | 0 | 2 | 2 | 0 | 2 | 12 |
| Vance, 2014 | 2 | NA | 0 | 2 | 2 | 0 | 0 | NA | NA | 2 | NA | 0 | 8 |
| Wang, 2010 | 2 | 2 | 0 | 2 | 0 | NA | NA | 0 | 2 | 2 | 0 | 2 | 12 |
| Wang, 2020 | 2 | 2 | 0 | 2 | 2 | 0 | 0 | 0 | 2 | 2 | 0 | 0 | 12 |
| Yun, 2009 | 2 | 0 | 0 | 2 | 0 | NA | NA | 0 | 2 | 2 | 0 | 2 | 10 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Longo, U.G.; Candela, V.; Berton, A.; De Salvatore, S.; Fioravanti, S.; Giannone, L.; Marchetti, A.; De Marinis, M.G.; Denaro, V. Biosensors for Detection of Biochemical Markers Relevant to Osteoarthritis. Biosensors 2021, 11, 31. https://doi.org/10.3390/bios11020031
Longo UG, Candela V, Berton A, De Salvatore S, Fioravanti S, Giannone L, Marchetti A, De Marinis MG, Denaro V. Biosensors for Detection of Biochemical Markers Relevant to Osteoarthritis. Biosensors. 2021; 11(2):31. https://doi.org/10.3390/bios11020031
Chicago/Turabian StyleLongo, Umile Giuseppe, Vincenzo Candela, Alessandra Berton, Sergio De Salvatore, Sara Fioravanti, Lucia Giannone, Anna Marchetti, Maria Grazia De Marinis, and Vincenzo Denaro. 2021. "Biosensors for Detection of Biochemical Markers Relevant to Osteoarthritis" Biosensors 11, no. 2: 31. https://doi.org/10.3390/bios11020031
APA StyleLongo, U. G., Candela, V., Berton, A., De Salvatore, S., Fioravanti, S., Giannone, L., Marchetti, A., De Marinis, M. G., & Denaro, V. (2021). Biosensors for Detection of Biochemical Markers Relevant to Osteoarthritis. Biosensors, 11(2), 31. https://doi.org/10.3390/bios11020031







