Assembly and Characterization of HBc Derived Virus-like Particles with Magnetic Core
Abstract
1. Introduction
2. Materials and Methods
2.1. Reagents
2.2. Superparamagnetic Iron Oxide Nanoparticles (SPIONs) Synthesis
2.3. SPIONs Functionalization
2.4. Concentration Measurement via Thermogravimetric Analysis
2.5. HBc Production and Preparation
2.6. SPION-HBc Preparation
2.7. VLPs Antigenicity
2.8. Statistical Analysis
2.9. Characterization Methods
3. Results
3.1. SPIONs Synthesis and Functionalization
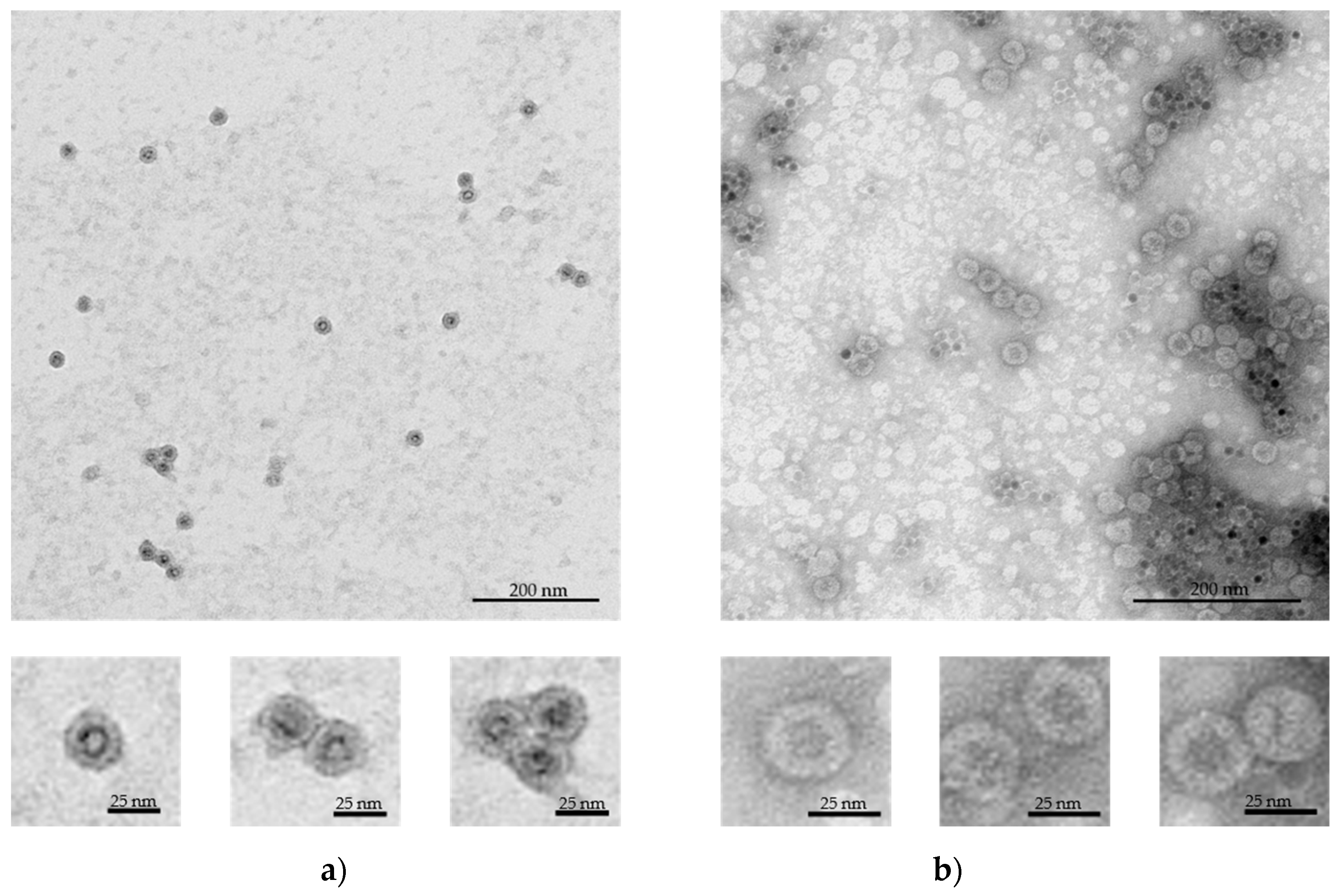
3.2. VLPs-SPION Assembly
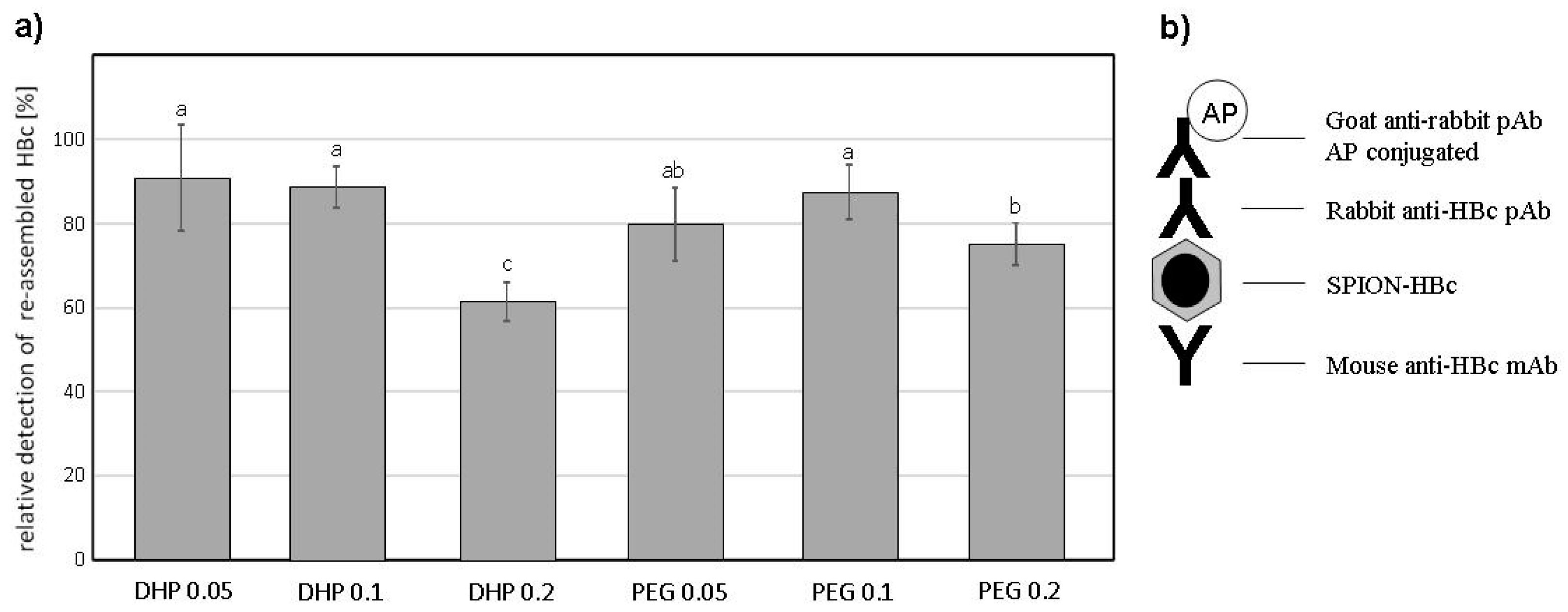
3.3. VLPs ELISA
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- López-Sagaseta, J.; Malito, E.; Rappuoli, R.; Bottomley, M.J. Self-assembling protein nanoparticles in the design of vaccines. Comput. Struct. Biotechnol. J. 2016, 14, 58–68. [Google Scholar] [CrossRef] [PubMed]
- Bachmann, M.F.; Jennings, G.T. Vaccine delivery: a matter of size, geometry, kinetics and molecular patterns. Nat. Rev. Immunol. 2010, 10, 787–796. [Google Scholar] [CrossRef] [PubMed]
- Sominskaya, I.; Skrastina, D.; Dislers, A.; Vasiljev, D.; Mihailova, M.; Ose, V.; Dreilina, D.; Pumpens, P. Construction and immunological evaluation of multivalent hepatitis B virus (HBV) core virus-like particles carrying HBV and HCV epitopes. Clin. Vaccine Immunol. 2010, 17, 1027–1033. [Google Scholar] [CrossRef] [PubMed]
- Galarza, J.M.; Latham, T.; Cupo, A. Virus-Like Particle (VLP) Vaccine Conferred Complete Protection against a Lethal Influenza Virus Challenge. Viral Immunol. 2005, 18, 244–251. [Google Scholar] [CrossRef] [PubMed]
- Jackwood, D.J. Multivalent Virus-Like–Particle Vaccine Protects Against Classic and Variant Infectious Bursal Disease Viruses. Avian Dis. 2013, 57, 41–50. [Google Scholar] [CrossRef] [PubMed]
- Mohsen, M.O.; Zha, L.; Cabral-Miranda, G.; Bachmann, M.F. Major findings and recent advances in virus–like particle (VLP)-based vaccines. Semin. Immunol. 2017, 34, 123–132. [Google Scholar] [CrossRef]
- Park, J.-S.; Cho, M.K.; Lee, E.J.; Ahn, K.-Y.; Lee, K.E.; Jung, J.H.; Cho, Y.; Han, S.-S.; Kim, Y.K.; Lee, J. A highly sensitive and selective diagnostic assay based on virus nanoparticles. Nat. Nanotechnol. 2009, 4, 259–264. [Google Scholar] [CrossRef]
- Stephanopoulos, N.; Carrico, Z.M.; Francis, M.B. Nanoscale Integration of Sensitizing Chromophores and Porphyrins with Bacteriophage MS2. Angew. Chem. Int. Ed. 2009, 48, 9498–9502. [Google Scholar] [CrossRef]
- Merzlyak, A.; Indrakanti, S.; Lee, S.-W. Genetically Engineered Nanofiber-Like Viruses For Tissue Regenerating Materials. Nano Lett. 2009, 9, 846–852. [Google Scholar] [CrossRef]
- De la Escosura, A.; Nolte, R.J.M.; Cornelissen, J.J.L.M. Viruses and protein cages as nanocontainers and nanoreactors. J. Mater. Chem. 2009, 19, 2274. [Google Scholar] [CrossRef]
- Uchida, M.; Klem, M.T.; Allen, M.; Suci, P.; Flenniken, M.; Gillitzer, E.; Varpness, Z.; Liepold, L.O.; Young, M.; Douglas, T. Biological Containers: Protein Cages as Multifunctional Nanoplatforms. Adv. Mater. 2007, 19, 1025–1042. [Google Scholar] [CrossRef]
- Liang, T.J. Hepatitis B: The virus and disease. Hepatology 2009, 49, S13–S21. [Google Scholar] [CrossRef]
- Nassal, M. The arginine-rich domain of the hepatitis B virus core protein is required for pregenome encapsidation and productive viral positive-strand DNA synthesis but not for virus assembly. J. Virol. 1992, 66, 4107–4116. [Google Scholar] [PubMed]
- Crowther, R.A.; Kiselev, N.A.; Böttcher, B.; Berriman, J.A.; Borisova, G.P.; Ose, V.; Pumpens, P. Three-dimensional structure of hepatitis B virus core particles determined by electron cryomicroscopy. Cell 1994, 77, 943–950. [Google Scholar] [CrossRef]
- Zlotnick, A.; Palmer, I.; Kaufman, J.D.; Stahl, S.J.; Steven, A.C.; Wingfield, P.T. Separation and crystallization of T = 3 and T = 4 icosahedral complexes of the hepatitis B virus core protein. Acta Crystallogr. Sect. D Biol. Crystallogr. 1999, 55, 717–720. [Google Scholar] [CrossRef]
- Milich, D.R.; McLachlan, A.; Moriarty, A.; Thornton, G.B. Immune response to hepatitis B virus core antigen (HBcAg): localization of T cell recognition sites within HBcAg/HBeAg. J. Immunol. 1987, 139, 1223–1231. [Google Scholar] [PubMed]
- Haracz, S.; Mróz, B.; Rybka, J.D.; Giersig, M.; Mrõz, B.; Rybka, J.D.; Giersig, M. Magnetic behaviour of non-interacting colloidal iron oxide nanoparticles in physiological solutions. Cryst. Res. Technol. 2015, 50, 791–796. [Google Scholar] [CrossRef]
- Krȩcisz, M.; Rybka, J.D.; Strugała, A.J.; Skalski, B.; Figlerowicz, M.; Kozak, M.; Giersig, M. Interactions between magnetic nanoparticles and model lipid bilayers—Fourier transformed infrared spectroscopy (FTIR) studies of the molecular basis of nanotoxicity. J. Appl. Phys. 2016, 120, 124701. [Google Scholar] [CrossRef]
- Haracz, S.; Hilgendorff, M.; Rybka, J.D.; Giersig, M. Effect of surfactant for magnetic properties of iron oxide nanoparticles. Nucl. Instruments Methods Phys. Res. Sect. B Beam Interact. Mater. Atoms 2015, 364, 120–126. [Google Scholar] [CrossRef]
- Mahmoudi, M.; Sant, S.; Wang, B.; Laurent, S.; Sen, T. Superparamagnetic iron oxide nanoparticles (SPIONs): Development, surface modification and applications in chemotherapy. Adv. Drug Deliv. Rev. 2011, 63, 24–46. [Google Scholar] [CrossRef]
- Neuberger, T.; Schöpf, B.; Hofmann, H.; Hofmann, M.; von Rechenberg, B. Superparamagnetic nanoparticles for biomedical applications: Possibilities and limitations of a new drug delivery system. J. Magn. Magn. Mater. 2005, 293, 483–496. [Google Scholar] [CrossRef]
- Teja, A.S.; Koh, P.-Y. Synthesis, properties, and applications of magnetic iron oxide nanoparticles. Prog. Cryst. Growth Charact. Mater. 2009, 55, 22–45. [Google Scholar] [CrossRef]
- Malyutin, A.G.; Easterday, R.; Lozovyy, Y.; Spilotros, A.; Cheng, H.; Sanchez-Felix, O.R.; Stein, B.D.; Morgan, D.G.; Svergun, D.I.; Dragnea, B.; et al. Viruslike nanoparticles with maghemite cores allow for enhanced mri contrast agents. Chem. Mater. 2015, 27, 327–335. [Google Scholar] [CrossRef]
- Huang, X.; Bronstein, L.M.; Retrum, J.; Dufort, C.; Tsvetkova, I.; Aniagyei, S.; Stein, B.; Stucky, G.; McKenna, B.; Remmes, N.; et al. Self-assembled virus-like particles with magnetic cores. Nano Lett. 2007, 7, 2407–2416. [Google Scholar] [CrossRef] [PubMed]
- Okuda, M.; Eloi, J.-C.; Jones, S.E.W.; Verwegen, M.; Cornelissen, J.J.L.M.; Schwarzacher, W. Pt, Co–Pt and Fe–Pt alloy nanoclusters encapsulated in virus capsids. Nanotechnology 2016, 27, 095605. [Google Scholar] [CrossRef]
- Gubin, S.P.; Koksharov, Y.A.; Khomutov, G.B.; Yurkov, G.Y. Magnetic nanoparticles: preparation, structure and properties. Russ. Chem. Rev. 2005, 74, 489–520. [Google Scholar] [CrossRef]
- Fatima, H.; Kim, K.S. Magnetic nanoparticles for bioseparation. Korean J. Chem. Eng. 2017, 34, 589–599. [Google Scholar] [CrossRef]
- Shen, L.; Zhou, J.; Wang, Y.; Kang, N.; Ke, X.; Bi, S.; Ren, L. Efficient Encapsulation of Fe3O4 Nanoparticles into Genetically Engineered Hepatitis B Core Virus-Like Particles Through a Specific Interaction for Potential Bioapplications. Small 2015, 11, 1190–1196. [Google Scholar] [CrossRef]
- Roose, K.; De Baets, S.; Schepens, B.; Saelens, X. Hepatitis B core-based virus-like particles to present heterologous epitopes. Expert Rev. Vaccines 2013, 12, 183–198. [Google Scholar] [CrossRef]
- Belyi, V.A.; Muthukumar, M. Electrostatic origin of the genome packing in viruses. Proc. Natl. Acad. Sci. USA 2006, 103, 17174–17178. [Google Scholar] [CrossRef]
- Mieloch, A.A.; Kręcisz, M.; Rybka, J.D.; Strugała, A.; Krupiński, M.; Urbanowicz, A.; Kozak, M.; Skalski, B.; Figlerowicz, M.; Giersig, M. The influence of ligand charge and length on the assembly of Brome mosaic virus derived virus-like particles with magnetic core. AIP Adv. 2018, 8, 035005. [Google Scholar] [CrossRef]
- Giersig, M.; Hilgendorff, M. Magnetic nanoparticle superstructures. Eur. J. Inorg. Chem. 2005, 2005, 3571–3583. [Google Scholar] [CrossRef]
- Peyret, H.; Lomonossoff, G.P. The pEAQ vector series: the easy and quick way to produce recombinant proteins in plants. Plant Mol. Biol. 2013, 83, 51–58. [Google Scholar] [CrossRef] [PubMed]
- Pyrski, M.; Rugowska, A.; Wierzbiński, K.R.; Kasprzyk, A.; Bogusiewicz, M.; Bociąg, P.; Samardakiewicz, S.; Czyż, M.; Kurpisz, M.; Pniewski, T. HBcAg produced in transgenic tobacco triggers Th1 and Th2 response when intramuscularly delivered. Vaccine 2017, 35, 5714–5721. [Google Scholar] [CrossRef] [PubMed]
- Hagan, M.F. A theory for viral capsid assembly around electrostatic cores. J. Chem. Phys. 2009, 130, 114902. [Google Scholar] [CrossRef] [PubMed]
- Šiber, A.; Zandi, R.; Podgornik, R. Thermodynamics of nanospheres encapsulated in virus capsids. Phys. Rev. E Stat. Nonlinear Soft Matter Phys. 2010, 81, 051919. [Google Scholar] [CrossRef]
- Wu, W.; Chen, Z.; Cheng, N.; Watts, N.R.; Stahl, S.J.; Farci, P.; Purcell, R.H.; Wingfield, P.T.; Steven, A.C. Specificity of an anti-capsid antibody associated with Hepatitis B Virus-related acute liver failure. J. Struct. Biol. 2013, 181, 53–60. [Google Scholar] [CrossRef]
- Kent, S.P.; Ryan, K.H.; Siegel, A.L. Steric hindrance as a factor in the reaction of labeled antibody with cell surface antigenic determinants. J. Histochem. Cytochem. 1978, 26, 618–621. [Google Scholar] [CrossRef]
- De Michele, C.; De Los Rios, P.; Foffi, G.; Piazza, F. Simulation and Theory of Antibody Binding to Crowded Antigen—Covered Surfaces. PLoS Comput. Biol. 2016, 12, e1004752. [Google Scholar] [CrossRef]
- Hagan, M.F. Controlling viral capsid assembly with templating. Phys. Rev. E Stat. Nonlinear Soft Matter Phys. 2008, 77, 051904. [Google Scholar] [CrossRef]
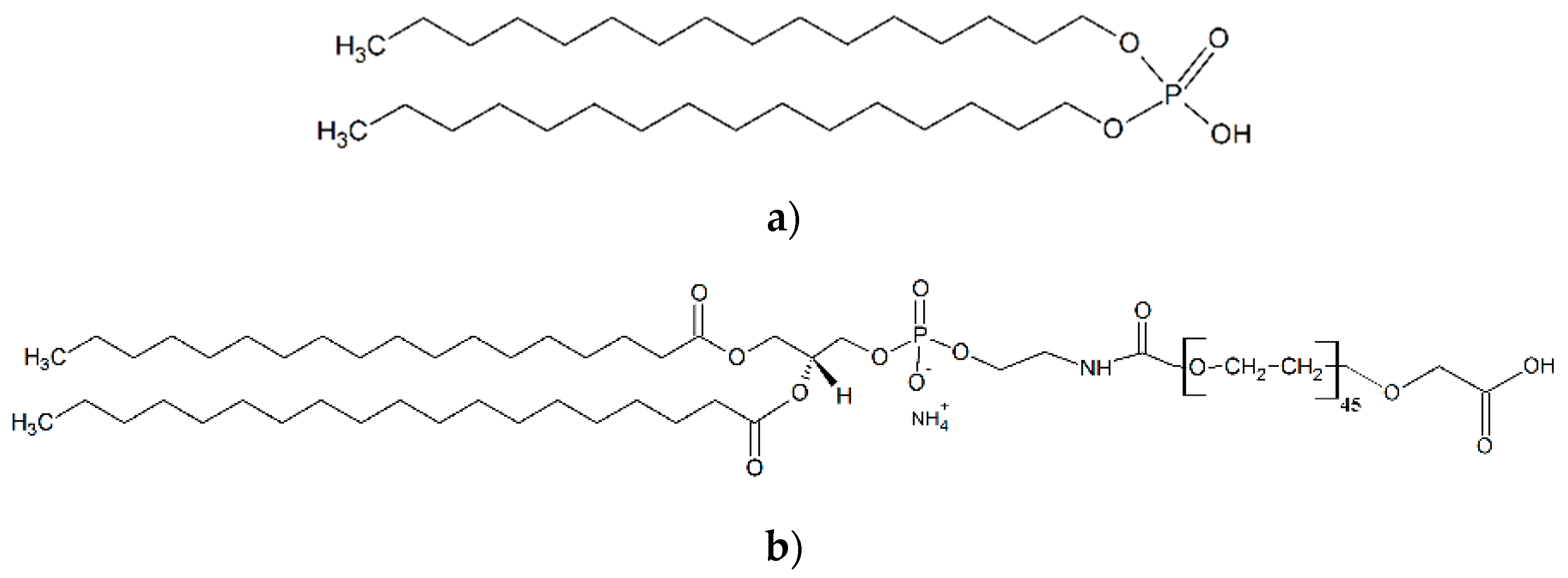

| SPION-DHP | SPION-PEG | |
|---|---|---|
| ζ-potential | −44.0 ± 3.4 mV | −37.3 ± 2.9 mV |
| Hydrodynamic diameter | 53.75 ± 1.93 nm | 29.69 ± 1.57 nm |
| SPION-DHP-HBc | SPION-PEG-HBc | |
|---|---|---|
| Diameter | 28.4 ± 1.2 nm | 29.9 ± 1.5 nm |
| Number of measured VLPs | 41 | 19 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rybka, J.D.; Mieloch, A.A.; Plis, A.; Pyrski, M.; Pniewski, T.; Giersig, M. Assembly and Characterization of HBc Derived Virus-like Particles with Magnetic Core. Nanomaterials 2019, 9, 155. https://doi.org/10.3390/nano9020155
Rybka JD, Mieloch AA, Plis A, Pyrski M, Pniewski T, Giersig M. Assembly and Characterization of HBc Derived Virus-like Particles with Magnetic Core. Nanomaterials. 2019; 9(2):155. https://doi.org/10.3390/nano9020155
Chicago/Turabian StyleRybka, Jakub Dalibor, Adam Aron Mieloch, Alicja Plis, Marcin Pyrski, Tomasz Pniewski, and Michael Giersig. 2019. "Assembly and Characterization of HBc Derived Virus-like Particles with Magnetic Core" Nanomaterials 9, no. 2: 155. https://doi.org/10.3390/nano9020155
APA StyleRybka, J. D., Mieloch, A. A., Plis, A., Pyrski, M., Pniewski, T., & Giersig, M. (2019). Assembly and Characterization of HBc Derived Virus-like Particles with Magnetic Core. Nanomaterials, 9(2), 155. https://doi.org/10.3390/nano9020155








