Designed Functional Dispersion for Insulin Protection from Pepsin Degradation and Skeletal Muscle Cell Proliferation: In Silico and In Vitro Study
Abstract
1. Introduction
2. Materials and Methods
2.1. Molecular Docking Simulation of Insulin Binding to PEGylated SWCNT
2.2. Materials
2.3. Preparation of Functional Dispersions
2.4. Enzymatic Degradation Study
2.5. Qualitative Analysis of Insulin Using MALDI-TOF
2.6. Quantitative Analysis of Free Insulin in Functional Dispersion Using UV Spectroscopy
2.7. Cell Culture
2.8. Cell Proliferation Assay
2.9. Insulin Quantification in Cell Supernatant Using the UV-Spectroscopy Technique
2.10. Absolute Glucose Quantification in Cell Supernatant Using 1H-NMR Spectroscopy
2.11. Statistical Analysis
3. Results and Discussion
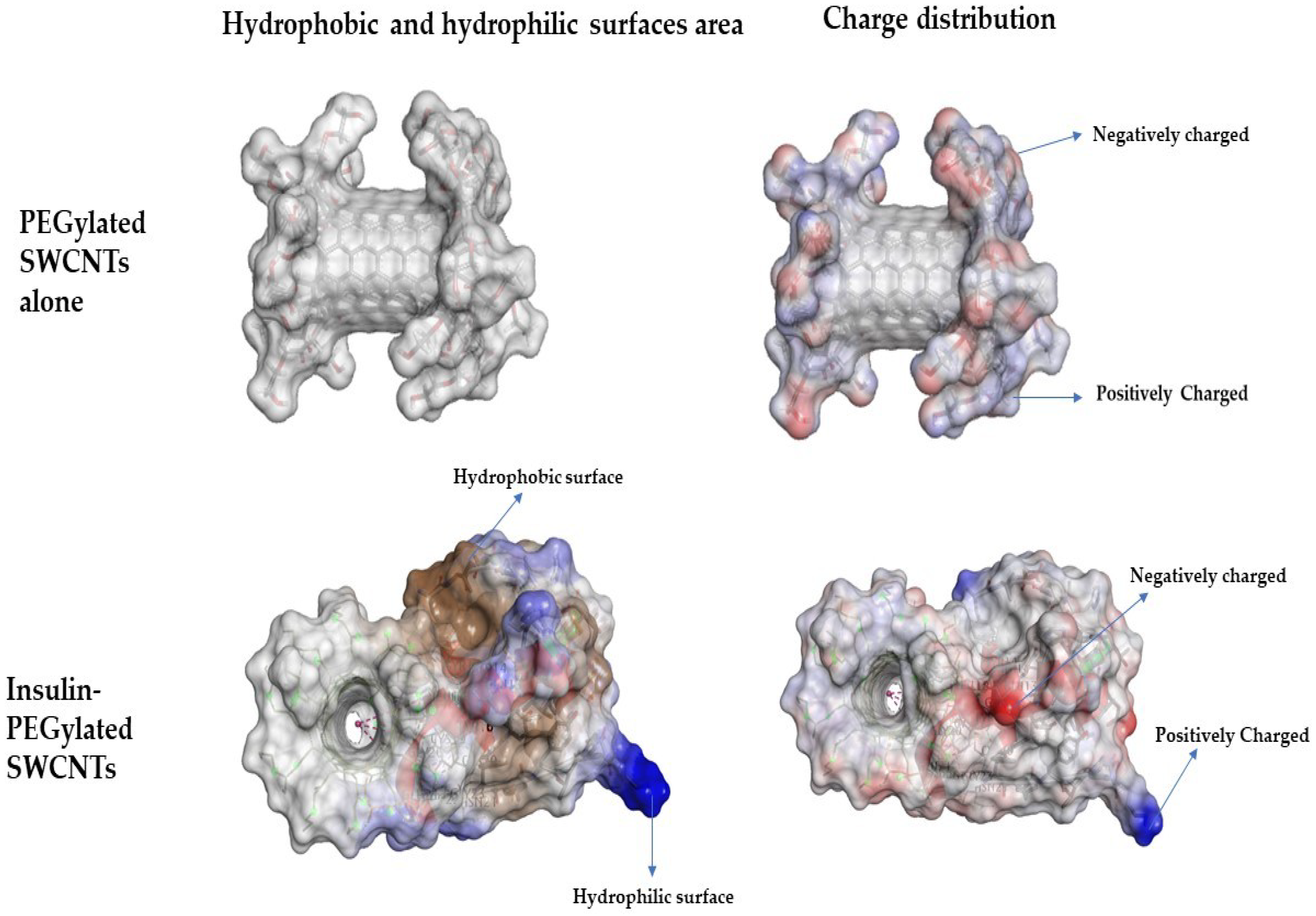
3.1. Non-Covalent Binding of Insulin to PEGylated SWCNT—Molecular Docking Simulations Study
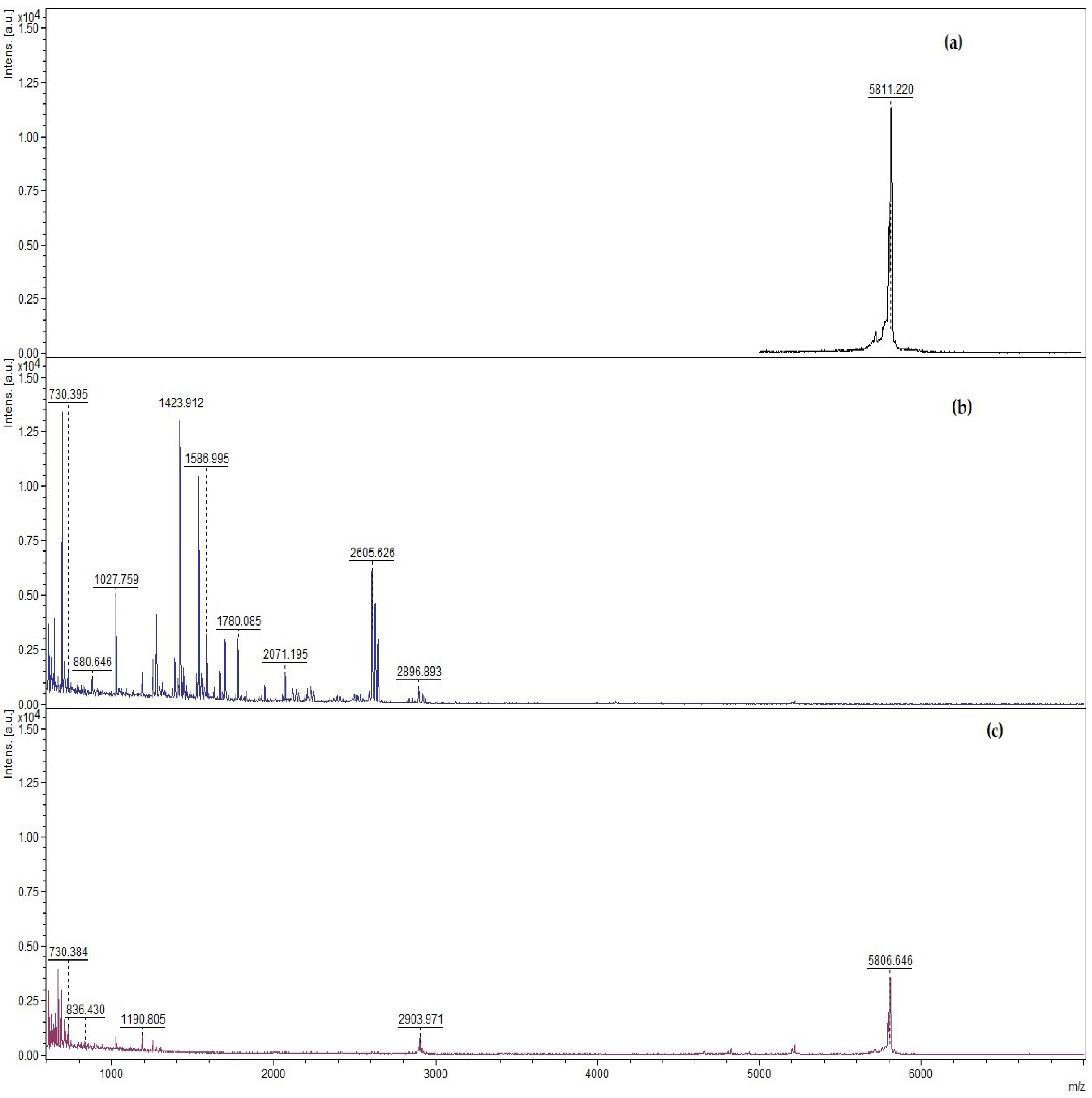
3.2. Qualitative and Quantitative Determination of Insulin Protected from Pepsin Digestion Using Enzymatic Degradation Assay
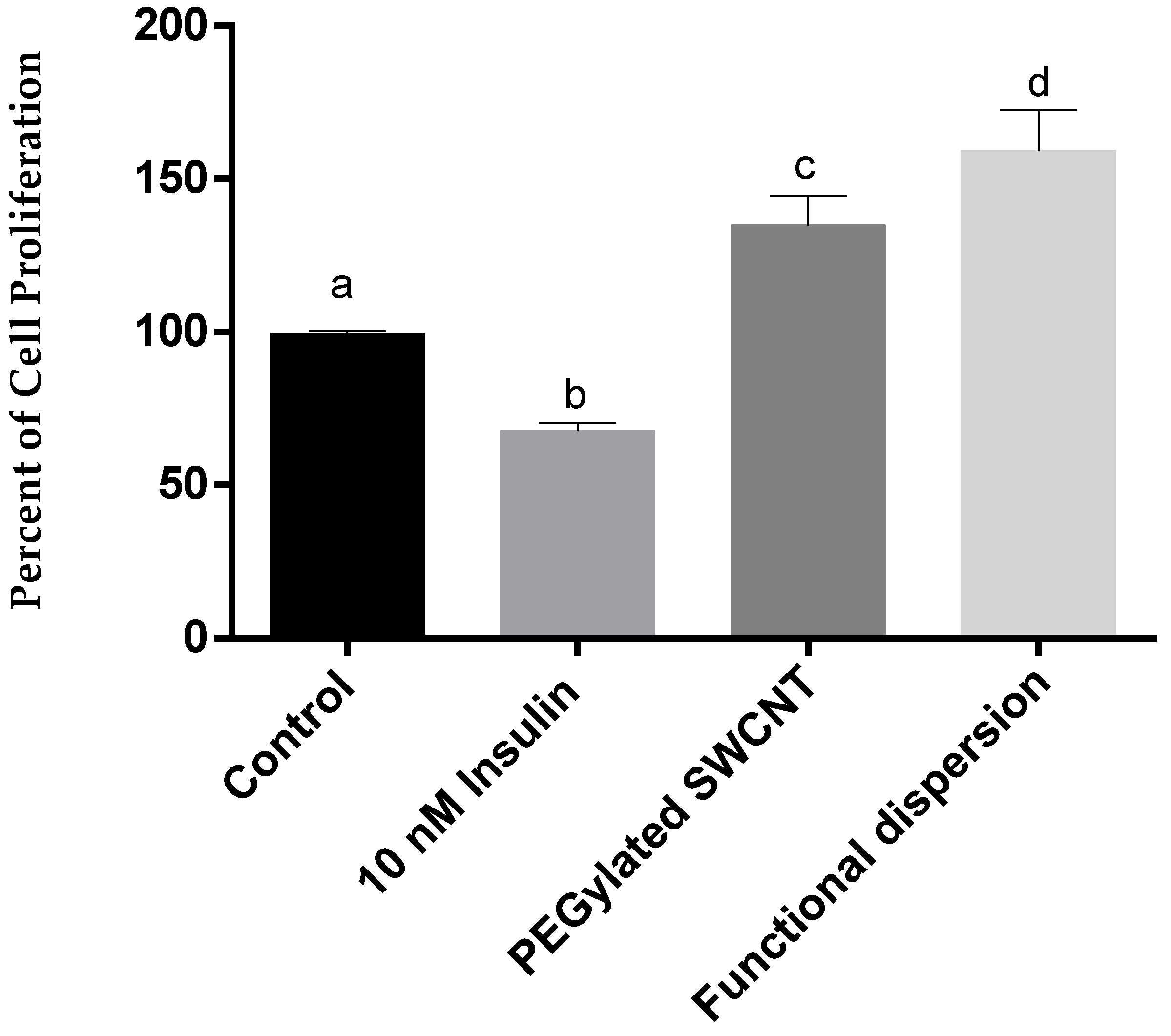
3.3. Cell Proliferation Study
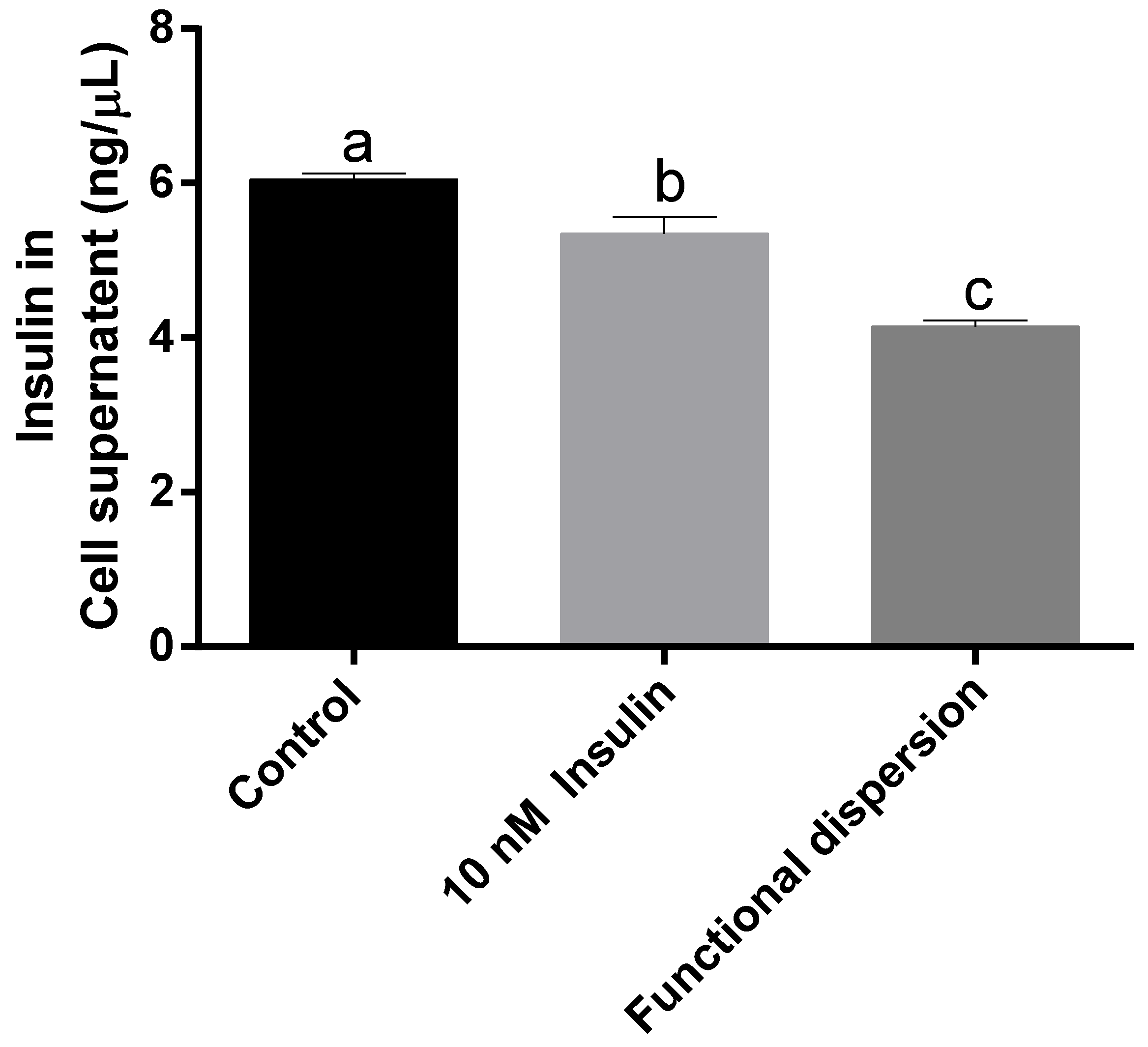
3.4. Insulin Levels in Cell Supernatant
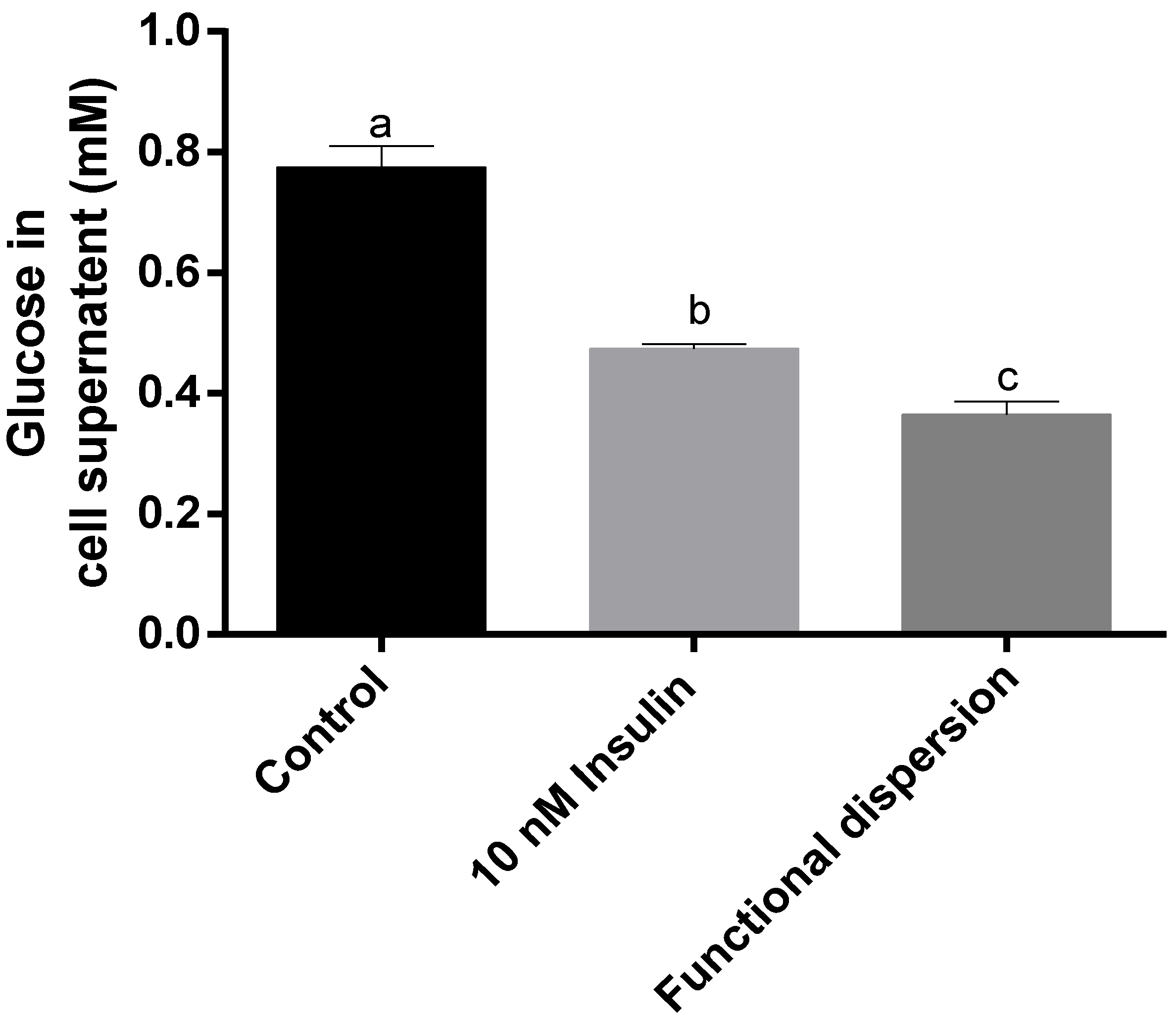
3.5. Glucose Levels in Cell Supernatant
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Boisseau, P.; Loubaton, B. Nanomedicine, nanotechnology in medicine. Comptes Rendus Phys. 2011, 12, 620–636. [Google Scholar] [CrossRef]
- Bawarski, W.E.; Chidlowsky, E.; Bharali, D.J.; Mousa, S.A. Emerging nanopharmaceuticals. Nanomed. Nanotechnol. Biol. Med. 2008. [Google Scholar] [CrossRef] [PubMed]
- Nikalje, A.P. Nanotechnology and its applications in medicine. Med. Chem. 2015. [Google Scholar] [CrossRef]
- Weissig, V.; Pettinger, T.K.; Murdock, N. Nanopharmaceuticals (part 1): Products on the market. Int. J. Nanomed. 2014, 9, 4357–4373. [Google Scholar] [CrossRef] [PubMed]
- Battigelli, A.; Ménard-Moyon, C.; Da Ros, T.; Prato, M.; Bianco, A. Endowing carbon nanotubes with biological and biomedical properties by chemical modifications. Adv. Drug Deliv. Rev. 2013, 65, 1899–1920. [Google Scholar] [CrossRef] [PubMed]
- Heister, E.; Brunner, E.W.; Dieckmann, G.R.; Jurewicz, I.; Dalton, A.B. Are carbon nanotubes a natural solution? applications in biology and medicine. ACS Appl. Mater. Interfaces 2013, 5, 1870–1891. [Google Scholar] [CrossRef] [PubMed]
- Tasis, D.; Tagmatarchis, N.; Bianco, A.; Prato, M. Chemistry of carbon nanotubes. Chem. Rev. 2006, 106, 1105–1136. [Google Scholar] [CrossRef] [PubMed]
- Singh, P.; Campidelli, S.; Giordani, S.; Bonifazi, D.; Bianco, A.; Prato, M. Organic functionalisation and characterisation of single-walled carbon nanotubes. Chem. Soc. Rev. 2009, 38, 2214–2230. [Google Scholar] [CrossRef] [PubMed]
- Antonucci, A.; Kupis-Rozmysłowicz, J.; Boghossian, A.A. Noncovalent protein and peptide functionalization of single-walled carbon nanotubes for biodelivery and optical sensing applications. ACS Appl. Mater. Interfaces 2017, 9, 11321–11331. [Google Scholar] [CrossRef] [PubMed]
- Klumpp, C.; Kostarelos, K.; Prato, M.; Bianco, A. Functionalized carbon nanotubes as emerging nanovectors for the delivery of therapeutics. Biochim. Biophys. Acta 2006, 1758, 404–412. [Google Scholar] [CrossRef] [PubMed]
- Balasubramanian, K.; Burghard, M. Biosensors based on carbon nanotubes. Anal. Bioanal. Chem. 2006, 385, 452–468. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Meng, L.; Lu, Q.; Fei, Z.; Dyson, P.J. Targeted delivery and controlled release of doxorubicin to cancer cells using modified single wall carbon nanotubes. Biomaterials 2009. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Cai, W.; He, L.; Nakayama, N.; Chen, K.; Sun, X.; Chen, X.; Dai, H. In vivo biodistribution and highly efficient tumour targeting of carbon nanotubes in mice. Nat. Nanotechnol. 2007. [Google Scholar] [CrossRef] [PubMed]
- Sacchetti, C.; Motamedchaboki, K.; Magrini, A.; Palmieri, G.; Mattei, M.; Bernardini, S.; Rosato, N.; Bottini, N.; Bottini, M. Surface polyethylene glycol conformation influences the protein corona of polyethylene glycol-modified single-walled carbon nanotubes: Potential implications on biological performance. ACS Nano 2013. [Google Scholar] [CrossRef] [PubMed]
- El-Sayed, R.; Bhattacharya, K.; Gu, Z.; Yang, Z.; Weber, J.K.; Li, H.; Leifer, K.; Zhao, Y.; Toprak, M.S.; Zhou, R.; et al. Single-walled carbon nanotubes inhibit the cytochrome P450 enzyme, CYP3A4. Sci. Rep. 2016, 6, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Son, K.H.; Hong, J.H.; Lee, J.W. Carbon nanotubes as cancer therapeutic carriers and mediators. Int. J. Nanomed. 2016, 11, 5163–5185. [Google Scholar] [CrossRef] [PubMed]
- Park, Y.K.; Bold, B.; Lee, W.K.; Jeon, M.H.; An, K.H.; Jeong, S.Y.; Shim, Y.K. D-(+)-galactose-conjugated single-walled carbon nanotubes as new chemical probes for electrochemical biosensors for the cancer marker galectin-3. Int. J. Mol. Sci. 2011. [Google Scholar] [CrossRef] [PubMed]
- Fujigaya, T.; Nakashima, N. Non-covalent polymer wrapping of carbon nanotubes and the role of wrapped polymers as functional dispersants. Sci. Technol. Adv. Mater. 2015, 16, 1–21. [Google Scholar] [CrossRef] [PubMed]
- WHO Global Action Plan for the Prevention and Control of Noncommunicable Diseases 2013–2020. Available online: ww.who.int/ncd (accessed on 17 October 2018).
- Classification and Diagnosis of Diabetes. Available online: http://care.diabetesjournals.org/content/40/Supplement_1/S11 (accessed on 17 October 2018).
- McCall, A.L. Insulin therapy and hypoglycemia. Endocrinol. Metab. Clin. N. Am. 2012, 41, 57–87. [Google Scholar] [CrossRef] [PubMed]
- Hermansen, K.; Davies, M. Does insulin detemir have a role in reducing risk of insulin-associated weight gain? Diabetes Obes. Metab. 2007, 9, 209–217. [Google Scholar] [CrossRef] [PubMed]
- Bowker, S.L.; Majumdar, S.R.; Veugelers, P.; Johnson, J.A. Increased cancer-related mortality for patients with type 2 diabetes who use sulfonylureas or insulin. Diabetes Care 2006. [Google Scholar] [CrossRef]
- Mo, R.; Jiang, T.; Di, J.; Tai, W.; Gu, Z. Emerging micro- and nanotechnology based synthetic approaches for insulin delivery. Chem. Soc. Rev. 2014, 43, 3595–3629. [Google Scholar] [CrossRef] [PubMed]
- Stanford, K.I.; Goodyear, L.J. Exercise and type 2 diabetes: Molecular mechanisms regulating glucose uptake in skeletal muscle. Adv. Physiol. Educ. 2014. [Google Scholar] [CrossRef] [PubMed]
- Mann, C.J.; Ayuso, E.; Anguela, X.M.; Bosch, F. Skeletal muscle metabolism in the pathology and treatment of type 1 diabetes. Curr. Pharm. Des. 2010, 16, 1002–1020. [Google Scholar] [CrossRef] [PubMed]
- Andersen, H.; Gadeberg, P.C.; Brock, B.; Jakobsen, J. Muscular atrophy in diabetic neuropathy: A stereological magnetic resonance imaging study. Diabetologia 1997. [Google Scholar] [CrossRef] [PubMed]
- Kivelä, R.; Silvennoinen, M.; Touvra, A.M.; Lehti, T.M.; Kainulainen, H.; Vihko, V. Effects of experimental type 1 diabetes and exercise training on angiogenic gene expression and capillarization in skeletal muscle. FASEB J. 2006. [Google Scholar] [CrossRef] [PubMed]
- Prior, S.J.; Mckenzie, M.J.; Joseph, L.J.; Ivey, F.; Macko, R.F.; Hafer-Macko, C.E.; Ryan, A.S. Reduced skeletal muscle capillarization and glucose intolerance. Microcirculation 2009. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, J.; Mogensen, M.; Vind, B.F.; Sahlin, K.; Hojlund, K.; Schroder, H.D.; Ortenblad, N. Increased subsarcolemmal lipids in type 2 diabetes: Effect of training on localization of lipids, mitochondria, and glycogen in sedentary human skeletal muscle. AJP Endocrinol. Metab. 2010. [Google Scholar] [CrossRef] [PubMed]
- Kelley, D.E.; Mandarino, L.J. Fuel selection in human skeletal muscle in insulin resistance: A reexamination. Diabetes 2000. [Google Scholar] [CrossRef]
- Park, S.W.; Goodpaster, B.H.; Strotmeyer, E.S.; De Rekeneire, N.; Harris, T.B.; Schwartz, A.V.; Tylavsky, F.A.; Newman, A.B. Decreased muscle strength and quality in older adults with type 2 diabetes: The health, aging, and body composition study. Diabetes 2006. [Google Scholar] [CrossRef] [PubMed]
- Mastrocola, R.; Reffo, P.; Penna, F.; Elena Tomasinelli, C.; Boccuzzi, G.; Maria Baccino, F.; Aragno, M.; Costelli, P. Muscle wasting in diabetic and in tumor-bearing rats: Role of oxidative stress. Free Radic. Biol. Med. 2008. [Google Scholar] [CrossRef] [PubMed]
- Shapiro, A.M.J.; Pokrywczynska, M.; Ricordi, C. Clinical pancreatic islet transplantation. Nat. Rev. Endocrinol. 2016. [Google Scholar] [CrossRef] [PubMed]
- Assady, S.; Maor, G.; Amit, M.; Itskovitz-Eldor, J.; Skorecki, K.L.; Tzukerman, M. Insulin production by human embryonic stem cells. Diabetes 2001, 50, 1691–1697. [Google Scholar] [CrossRef] [PubMed]
- Palmer, J.P. Immunomodulatory therapy of human type 1 diabetes: Lessons from the mouse. J. Clin. Investig. 2001, 108, 31–33. [Google Scholar] [CrossRef] [PubMed]
- Talchai, C.; Xuan, S.; Kitamura, T.; DePinho, R.A.; Accili, D. Generation of functional insulin-producing cells in the gut by Foxo1 ablation. Nat. Genet. 2012. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Lee, H.H.; Kwon, H.; Shin, S.; Yoon, J.W.; Jun, H.S. Engineered enteroendocrine cells secrete insulin in response to glucose and reverse hyperglycemia in diabetic mice. Mol. Ther. 2007. [Google Scholar] [CrossRef] [PubMed]
- Tang, W.-J. Targeting insulin-degrading enzyme to treat type 2 diabetes mellitus. Trends Endocrinol. Metab. 2015. [Google Scholar] [CrossRef] [PubMed]
- Berman, H.M. The protein data bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Andrusier, N.; Nussinov, R.; Wolfson, H.J. FireDock: Fast interaction refinement in molecular docking. Proteins Struct. Funct. Genet. 2007. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, N.; Sato, T.; Kawasaki, K.; Murosaki, S.; Yamamoto, Y. A nonradioisotope, enzymatic assay for 2-deoxyglucose uptake in L6 skeletal muscle cells cultured in a 96-well microplate. Anal. Biochem. 2006. [Google Scholar] [CrossRef] [PubMed]
- Skehan, P.; Storeng, R.; Scudiero, D.; Monks, A.; McMahon, J.; Vistica, D.; Warren, J.T.; Bokesch, H.; Kenney, S.; Boyd, M.R. New colorimetric cytotoxicity assay for anti- cancer-drug screening. J. Natl. Cancer Inst. 1990, 13, 1107–1112. [Google Scholar] [CrossRef]
- Royatvand, S.; Fallah Hoseini, H.; Ezzatpanah, H.; Sekehchi, M. Determination of Insulin Concentration in Camel Milk Using Ultra Violet –Visible Absorption Spectroscopy. J. Food Biosci. Technol. Sci. Res. Branch 2013, 3, 53–60. [Google Scholar]
- Langley, J.N.; Edkins, J.S. Pepsinogen and pepsin. J. Physiol. 1886. [Google Scholar] [CrossRef]
- Bisker, G.; Bakh, N.A.; Lee, M.A.; Ahn, J.; Park, M.; O’Connell, E.B.; Iverson, N.M.; Strano, M.S. Insulin detection using a corona phase molecular recognition site on single-walled carbon nanotubes. ACS Sens. 2018, 3, 367–377. [Google Scholar] [CrossRef] [PubMed]
- Matsuura, K.; Saito, T.; Okazaki, T.; Ohshima, S.; Yumura, M.; Iijima, S. Selectivity of water-soluble proteins in single-walled carbon nanotube dispersions. Chem. Phys. Lett. 2006, 429, 497–502. [Google Scholar] [CrossRef]
- Farris, W.; Mansourian, S.; Chang, Y.; Lindsley, L.; Eckman, E.A.; Frosch, M.P.; Eckman, C.B.; Tanzi, R.E.; Selkoe, D.J.; Guenette, S. Insulin-degrading enzyme regulates the levels of insulin, amyloid beta-protein, and the beta-amyloid precursor protein intracellular domain in vivo. Proc. Natl. Acad. Sci. USA 2003. [Google Scholar] [CrossRef] [PubMed]
- Di Guglielmo, G.M.; Drake, P.G.; Baass, P.C.; Authier, F.; Posner, B.I.; Bergeron, J.J. Insulin receptor internalization and signalling. Mol. Cell. Biochem. 1998. [Google Scholar] [CrossRef]
- Ge, C.; Du, J.; Zhao, L.; Wang, L.; Liu, Y.; Li, D.; Yang, Y.; Zhou, R.; Zhao, Y.; Chai, Z.; et al. Binding of blood proteins to carbon nanotubes reduces cytotoxicity. Proc. Natl. Acad. Sci. USA 2011, 108, 16968–16973. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.H.; Kwon, C.H.; Lee, C.; An, J.; Phuong, T.T.T.; Park, S.H.; Lima, M.D.; Baughman, R.H.; Kang, T.M.; Kim, S.J. Bio-inspired hybrid carbon nanotube muscles. Sci. Rep. 2016. [Google Scholar] [CrossRef] [PubMed]
- Joddar, B.; Garcia, E.; Casas, A.; Stewart, C.M. Development of functionalized multi-walled carbon-nanotube-based alginate hydrogels for enabling biomimetic technologies. Sci. Rep. 2016. [Google Scholar] [CrossRef] [PubMed]
- Deligianni, D.D. Multiwalled carbon nanotubes enhance human bone marrow mesenchymal stem cells’ spreading but delay their proliferation in the direction of differentiation acceleration. Cell Adhes. Migr. 2014, 8. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Han, J.; Li, H.; Liu, M.; Zeng, B. The establishment of insulin resistance model in FL83B and L6 cell. AIP Conf. Proc. 2017, 1890, 030006. [Google Scholar]
- Guo, X.; Li, H.; Xu, H.; Woo, S.; Dong, H.; Lu, F.; Lange, A.J.; Wu, C. Glycolysis in the control of blood glucose homeostasis. Acta Pharm. Sin. B 2012. [Google Scholar] [CrossRef]
- Harada, S.; Smith, R.M.; Jarett, L. Mechanisms of nuclear translocation of insulin. Cell Biochem. Biophys. 1999, 31, 307–319. [Google Scholar] [CrossRef] [PubMed]
- Sanjib, B.; Krishna, K.; Franklin, K. Modulating the Glucose Transport by Engineering Gold Nanoparticles. J. Nanomed. Biother. Discov. 2016. [Google Scholar] [CrossRef]

| Free Insulin | Functional Dispersion (Free Insulin) | |
|---|---|---|
| Before digestion (ng/μL) | 0.90 ± 0.13 | 0.13 ± 0.004 |
| After digestion using pepsin 0.4 mg/mL for 30 min (ng/μL) | 0 | 0.09 ± 0.004 |
| % insulin protected | 0 | 71.2 ± 1.27 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chittepu, V.C.S.R.; Kalhotra, P.; Gallardo-Velázquez, T.; Robles-de la Torre, R.R.; Osorio-Revilla, G. Designed Functional Dispersion for Insulin Protection from Pepsin Degradation and Skeletal Muscle Cell Proliferation: In Silico and In Vitro Study. Nanomaterials 2018, 8, 852. https://doi.org/10.3390/nano8100852
Chittepu VCSR, Kalhotra P, Gallardo-Velázquez T, Robles-de la Torre RR, Osorio-Revilla G. Designed Functional Dispersion for Insulin Protection from Pepsin Degradation and Skeletal Muscle Cell Proliferation: In Silico and In Vitro Study. Nanomaterials. 2018; 8(10):852. https://doi.org/10.3390/nano8100852
Chicago/Turabian StyleChittepu, Veera C. S. R., Poonam Kalhotra, Tzayhri Gallardo-Velázquez, Raúl René Robles-de la Torre, and Guillermo Osorio-Revilla. 2018. "Designed Functional Dispersion for Insulin Protection from Pepsin Degradation and Skeletal Muscle Cell Proliferation: In Silico and In Vitro Study" Nanomaterials 8, no. 10: 852. https://doi.org/10.3390/nano8100852
APA StyleChittepu, V. C. S. R., Kalhotra, P., Gallardo-Velázquez, T., Robles-de la Torre, R. R., & Osorio-Revilla, G. (2018). Designed Functional Dispersion for Insulin Protection from Pepsin Degradation and Skeletal Muscle Cell Proliferation: In Silico and In Vitro Study. Nanomaterials, 8(10), 852. https://doi.org/10.3390/nano8100852





