Microfluidics for Peptidomics, Proteomics, and Cell Analysis
Abstract
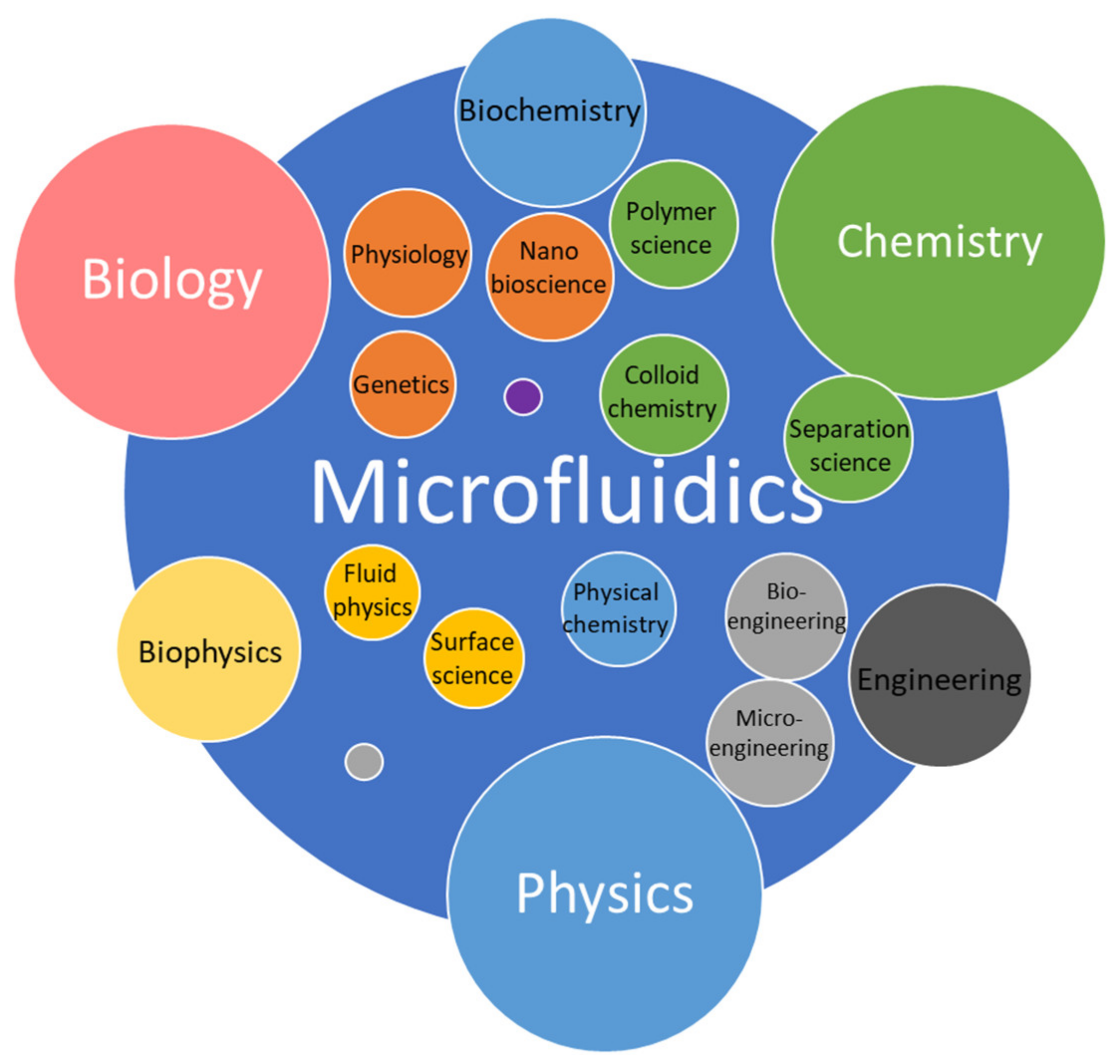
1. Introduction
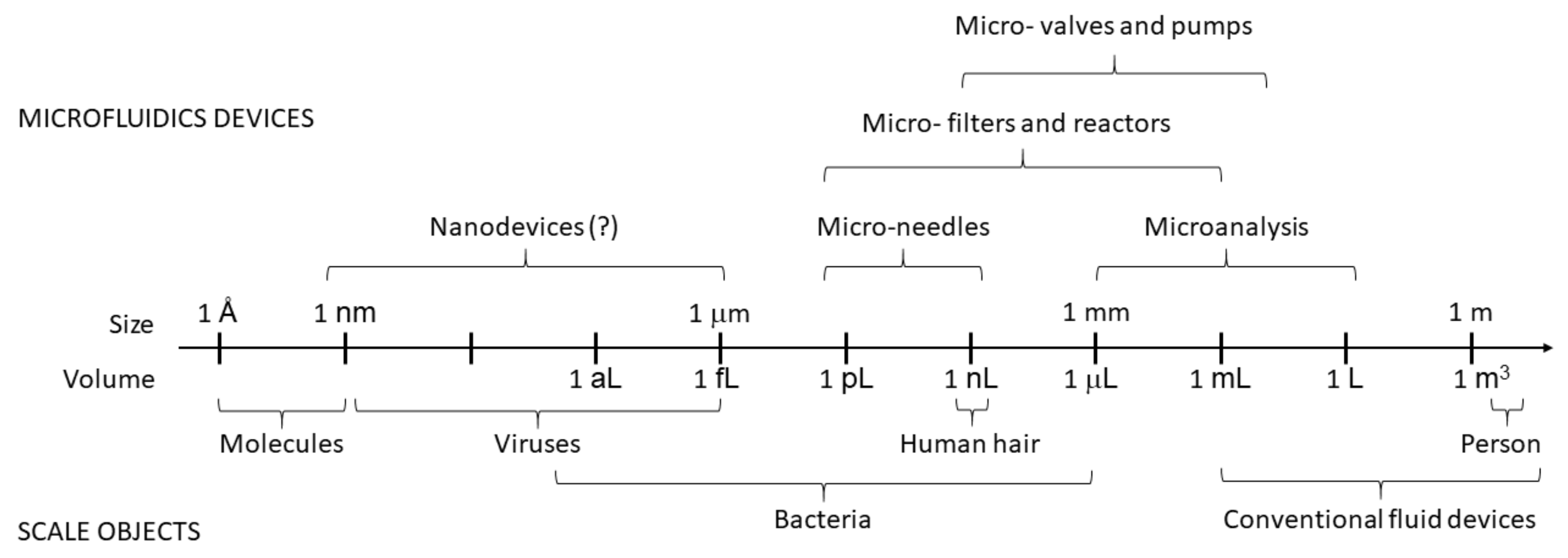
2. Fundamentals of Microfluidics
2.1. Microfluidic Chips
2.1.1. Glass vs. Plastic Chips
2.1.2. Fabrication Process of the Chips
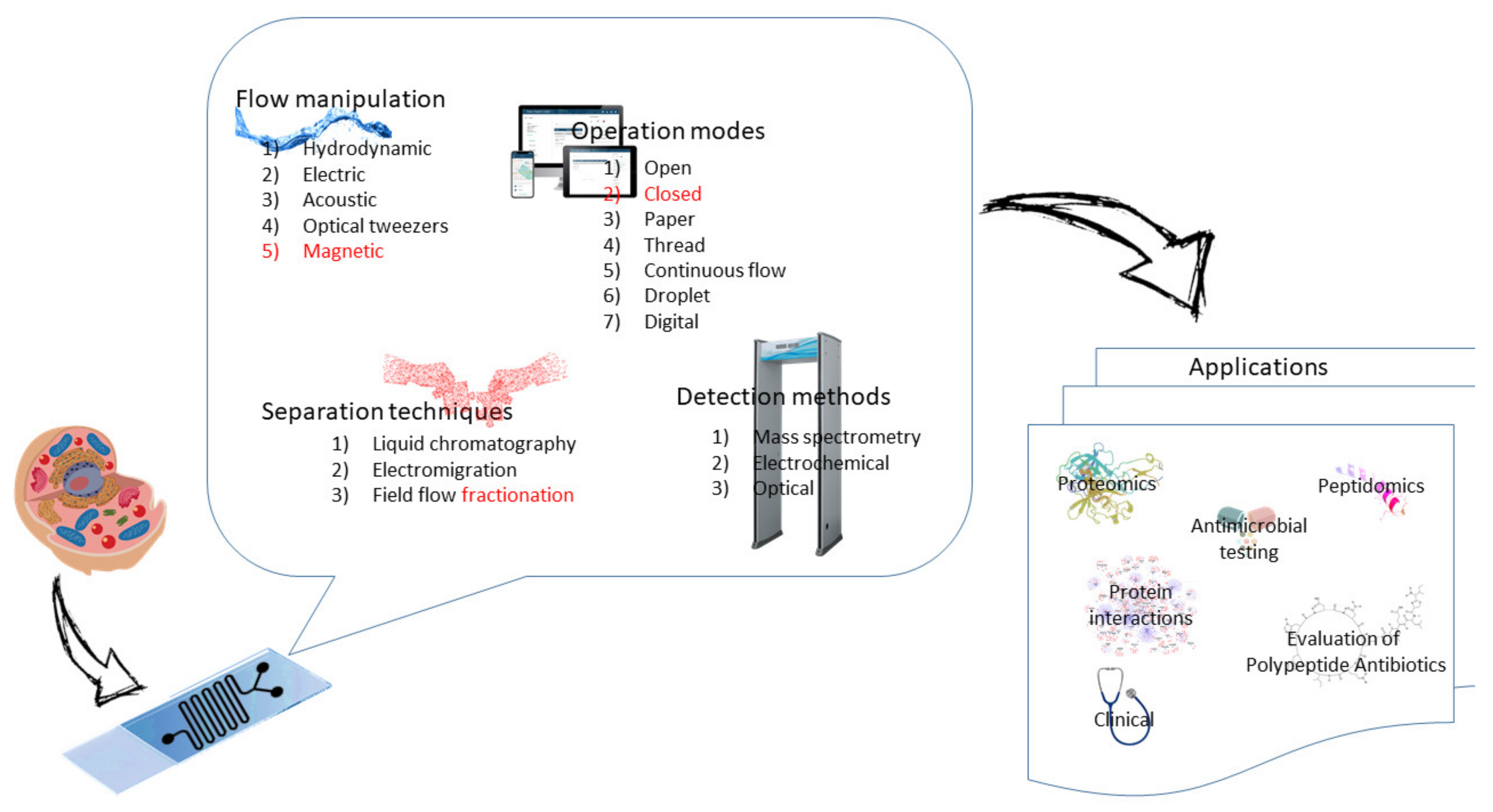
2.2. Flow Manipulation
2.2.1. Hydrodynamic Methods
2.2.2. Electric Methods
2.2.3. Acoustic Method
2.2.4. Optical Methods
2.2.5. Magnetic Methods
2.3. Basic Modes of Microfluidics
2.4. Separation Techniques Implemented in Microchips
2.4.1. Liquid Chromatographic Methods
2.4.2. Electromigration Methods
2.4.3. Field-Flow Fractionation
2.5. Detection Schemes
3. Applications of Microfluidics
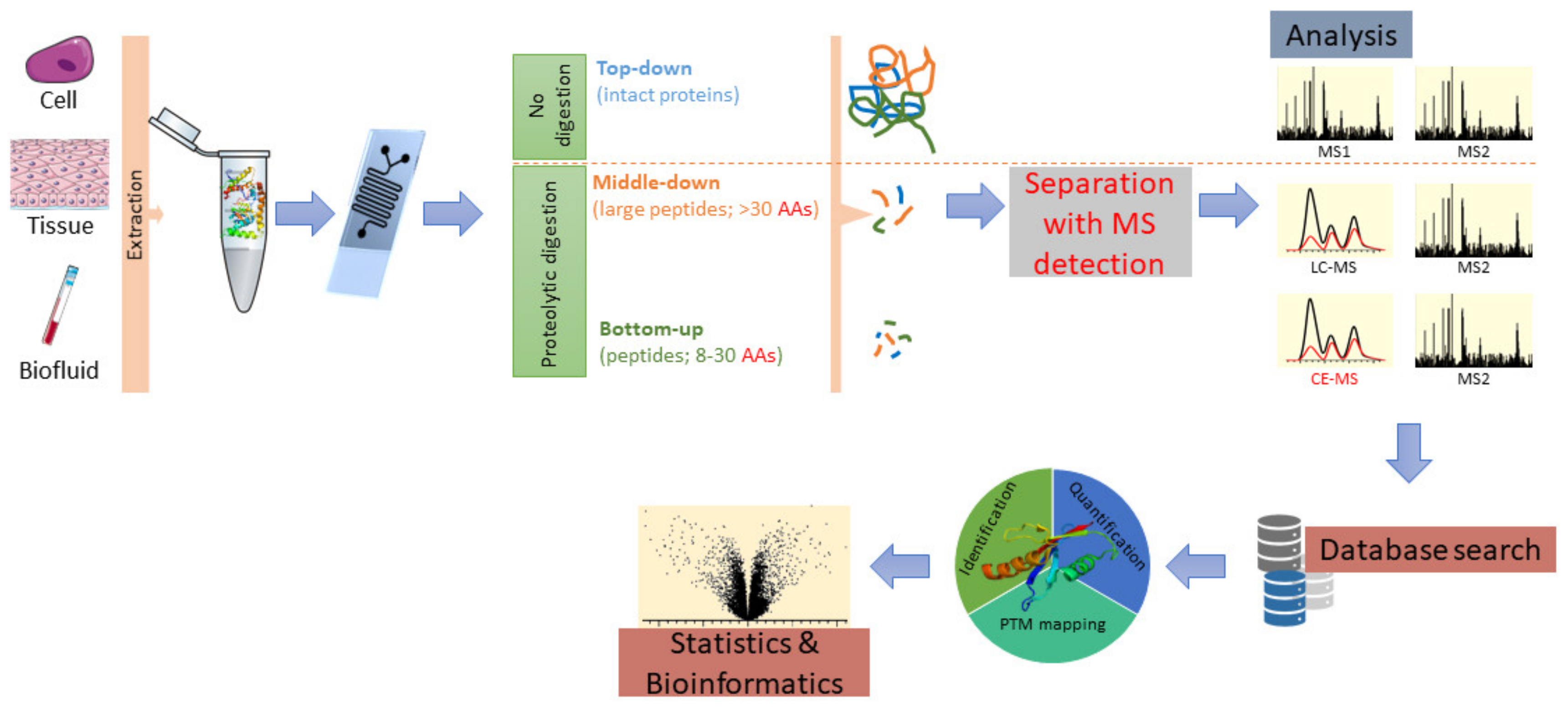
3.1. Analysis of Peptides and Proteins; Peptidomics and Proteomics
3.1.1. Introduction
3.1.2. Evaluation of Polypeptide Antibiotics
3.1.3. Antimicrobial Susceptibility Testing
3.1.4. Study of Protein–Protein Interactions
3.1.5. Proteome Analysis
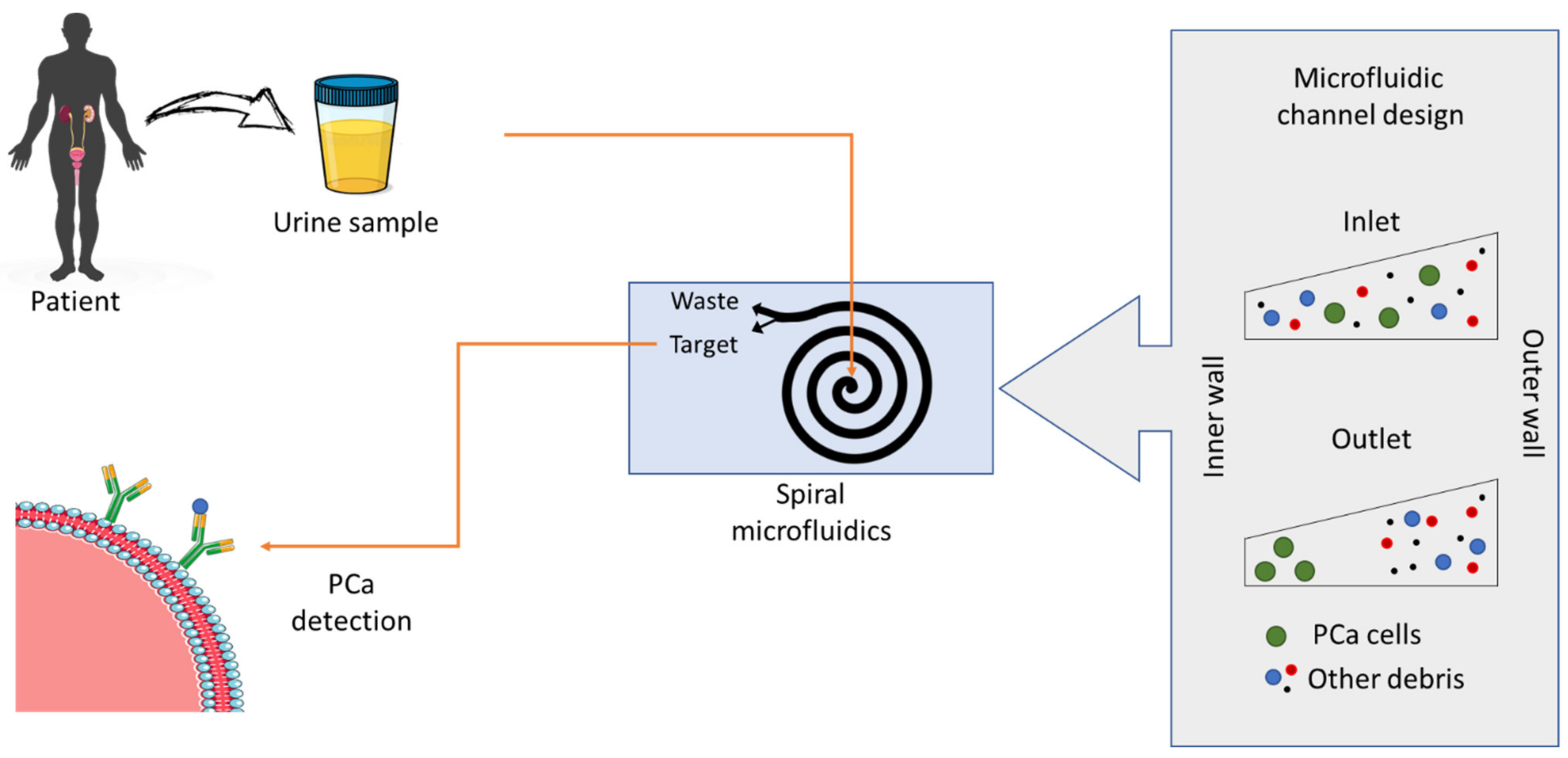
3.1.6. Clinical Applications
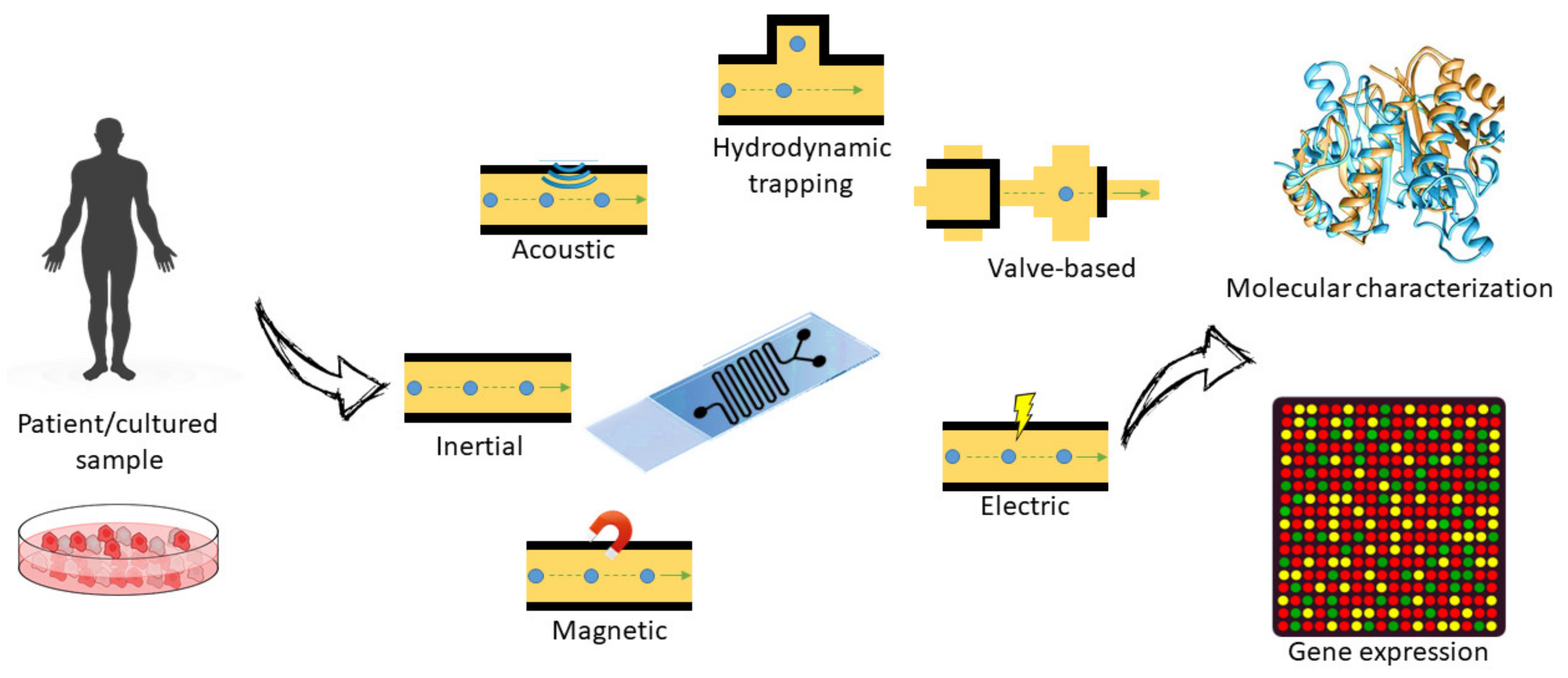
3.2. Separation and Analysis of Cells
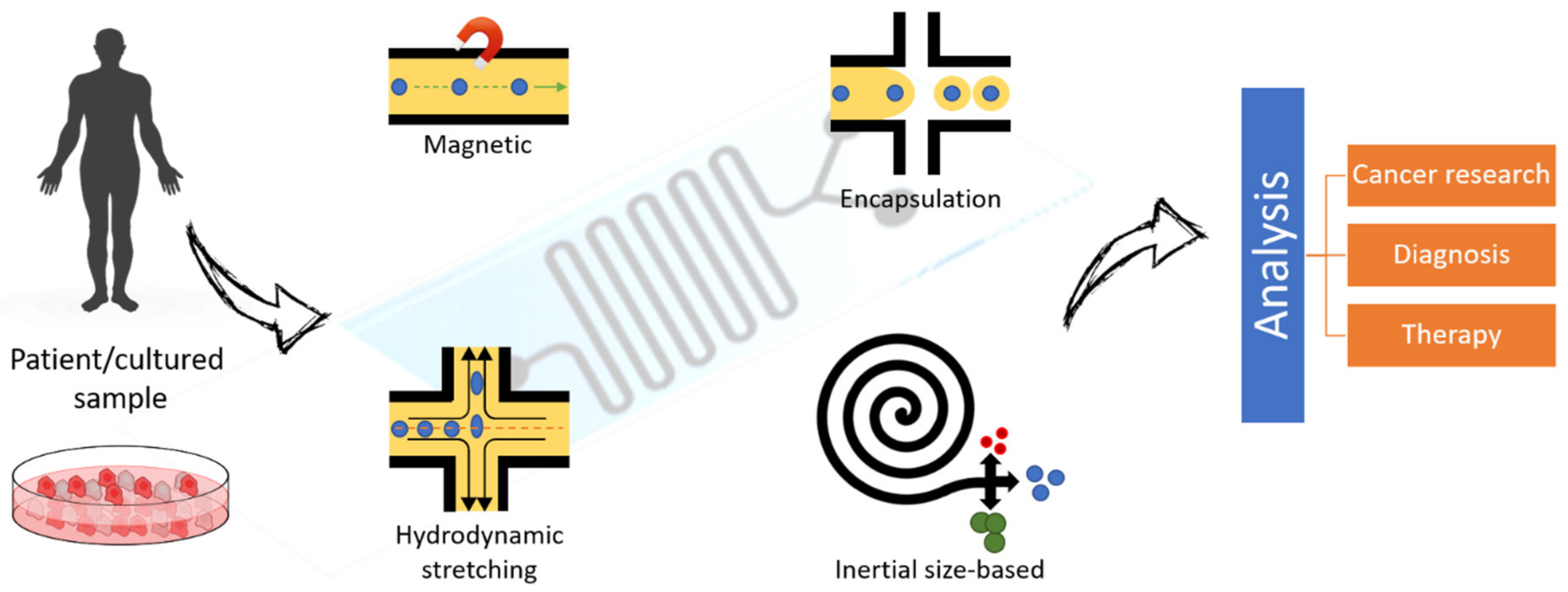
3.2.1. Cell Sorting and Single-Cell Analysis
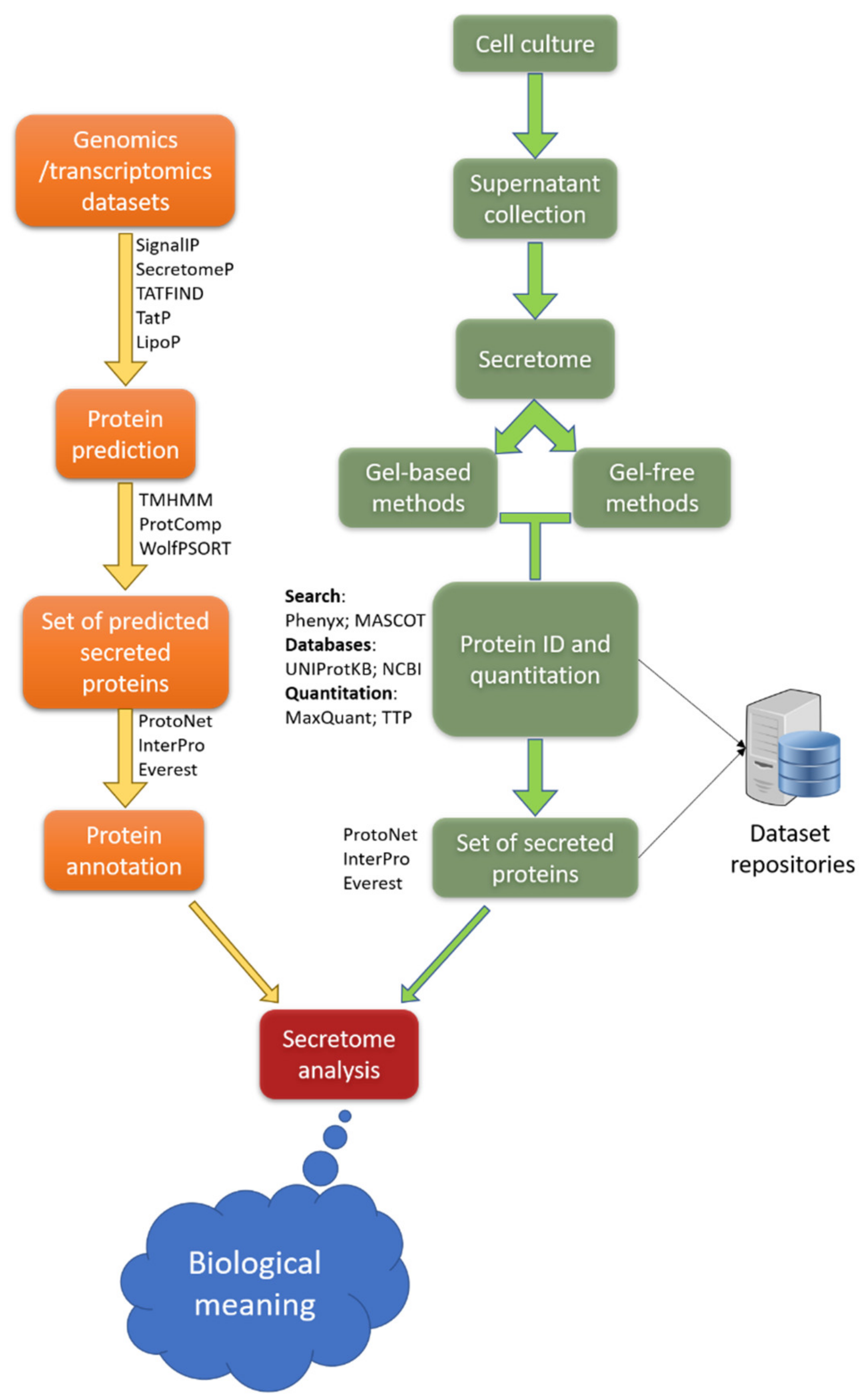
3.2.2. Secretome Analysis and Single-Cell Omics Analysis
3.2.3. Investigation of Stimulus-Driven Cell Behavior
3.2.4. Investigation of Biomolecular Coronas Nanoparticles
4. The Future of Microfluidics
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Lee, W.; Tseng, P.; Di Carlo, D. Microfluidic Cell Sorting and Separation Technology. In Microtechnology for Cell Manipulation and Sorting; Springer: Berlin/Heidelberg, Germany, 2017; pp. 1–14. [Google Scholar]
- Gao, D.; Song, C.; Lin, J.-M. Microfluidics-Mass Spectrometry Combination Systems for Single-Cell Analysis. In Microfluidics for Single-Cell Analysis; Springer: Berlin/Heidelberg, Germany, 2019; pp. 163–195. [Google Scholar]
- Liu, Y.; Yang, Q.; Cao, L.; Xu, F. Analysis of Leukocyte Behaviors on Microfluidic Chips. Adv. Healthc. Mater. 2019, 8, 1801406. [Google Scholar] [CrossRef]
- Chiu, D.T.; deMello, A.J.; Di Carlo, D.; Doyle, P.S.; Hansen, C.; Maceiczyk, R.M.; Wootton, R.C. Small but perfectly formed? Successes, challenges, and opportunities for microfluidics in the chemical and biological sciences. Chem 2017, 2, 201–223. [Google Scholar] [CrossRef]
- Conde, J.P.; Madaboosi, N.; Soares, R.R.; Fernandes, J.T.S.; Novo, P.; Moulas, G.; Chu, V. Lab-on-chip systems for integrated bioanalyses. Essays Biochem. 2016, 60, 121–131. [Google Scholar]
- Narayanamurthy, V.; Jeroish, Z.; Bhuvaneshwari, K.; Bayat, P.; Premkumar, R.; Samsuri, F.; Yusoff, M.M. Advances in passively driven microfluidics and lab-on-chip devices: A comprehensive literature review and patent analysis. RSC Adv. 2020, 10, 11652–11680. [Google Scholar] [CrossRef]
- Manz, A.; Graber, N.; Widmer, H.Á. Miniaturized total chemical analysis systems: A novel concept for chemical sensing. Sens. Actuators B Chem. 1990, 1, 244–248. [Google Scholar] [CrossRef]
- Bragheri, F.; Vázquez, R.M.; Osellame, R. Microfluidics. In Three-Dimensional Microfabrication Using Two-Photon Polymerization; Elsevier: Amsterdam, The Netherlands, 2020; pp. 493–526. [Google Scholar]
- Bai, Y.; Gao, M.; Wen, L.; He, C.; Chen, Y.; Liu, C.; Fu, X.; Huang, S. Applications of Microfluidics in Quantitative Biology. Biotechnol. J. 2018, 13, e1700170. [Google Scholar] [CrossRef] [PubMed]
- Loo, J.A.; Udseth, H.R.; Smith, R.D. Peptide and protein analysis by electrospray ionization-mass spectrometry and capillary electrophoresis-mass spectrometry. Anal. Biochem. 1989, 179, 404–412. [Google Scholar] [CrossRef]
- Kammeijer, G.S.; Kohler, I.; Jansen, B.C.; Hensbergen, P.J.; Mayboroda, O.A.; Falck, D.; Wuhrer, M. Dopant enriched nitrogen gas combined with sheathless capillary electrophoresis–electrospray ionization-mass spectrometry for improved sensitivity and repeatability in glycopeptide analysis. Anal. Chem. 2016, 88, 5849–5856. [Google Scholar] [CrossRef] [PubMed]
- Barry, R.; Ivanov, D. Microfluidics in biotechnology. J. Nanobiotechnol. 2004, 2, 2. [Google Scholar] [CrossRef]
- Soloviev, M.; Barry, R.; Scrivener, E.; Terrett, J. Combinatorial peptidomics: A generic approach for protein expression profiling. J. Nanobiotechnol. 2003, 1, 4. [Google Scholar] [CrossRef][Green Version]
- Caicedo, H.H.; Brady, S.T. Microfluidics: The challenge is to bridge the gap instead of looking for a ‘killer app’. Trends Biotechnol. 2016, 34, 1–3. [Google Scholar] [CrossRef]
- Fu, Y.Q.; Luo, J.; Nguyen, N.-T.; Walton, A.; Flewitt, A.J.; Zu, X.-T.; Li, Y.; McHale, G.; Matthews, A.; Iborra, E. Advances in piezoelectric thin films for acoustic biosensors, acoustofluidics and lab-on-chip applications. Prog. Mater. Sci. 2017, 89, 31–91. [Google Scholar] [CrossRef]
- Dekker, S.; Buesink, W.; Blom, M.; Alessio, M.; Verplanck, N.; Hihoud, M.; Dehan, C.; César, W.; Le Nel, A.; van den Berg, A. Standardized and modular microfluidic platform for fast Lab on Chip system development. Sens. Actuators B Chem. 2018, 272, 468–478. [Google Scholar] [CrossRef]
- Sollier, E.; Kochersperger, M.L.; Englert, R.F.; Che, J.; Huang, K.-W.; Boyce-Jacino, M.; Neddersen, A.; Passernig, A.; Richardson, B.; Choi, I. Microfluidic Chips and Cartridges and Systems Utilizing Microfluidic Chips and Cartridges. International Patent Application No. PCT/US2017/027959, 17 April 2017. [Google Scholar]
- Wu, J.; He, Z.; Chen, Q.; Lin, J.-M. Biochemical analysis on microfluidic chips. Trac Trends Anal. Chem. 2016, 80, 213–231. [Google Scholar] [CrossRef]
- Tran, D.Q. Microfluidic Studies on Flow Manipulation to Assist Metastasis Research. Ph.D. Thesis, Nanyang Technological University, Singapore, 2017. [Google Scholar]
- Mao, X.; Huang, T.J. Microfluidic diagnostics for the developing world. Lab Chip 2012, 12, 1412–1416. [Google Scholar] [CrossRef]
- Piccin, E.; Ferraro, D.; Sartori, P.; Chiarello, E.; Pierno, M.; Mistura, G. Generation of water-in-oil and oil-in-water microdroplets in polyester-toner microfluidic devices. Sens. Actuators B Chem. 2014, 196, 525–531. [Google Scholar] [CrossRef]
- Silvestrini, S.; Ferraro, D.; Tóth, T.; Pierno, M.; Carofiglio, T.; Mistura, G.; Maggini, M. Tailoring the wetting properties of thiolene microfluidic materials. Lab Chip 2012, 12, 4041–4043. [Google Scholar] [CrossRef] [PubMed]
- Sollier, K.; Mandon, C.A.; Heyries, K.A.; Blum, L.J.; Marquette, C.A. “Print-n-Shrink” technology for the rapid production of microfluidic chips and protein microarrays. Lab Chip 2009, 9, 3489–3494. [Google Scholar] [CrossRef] [PubMed]
- Taylor, D.; Dyer, D.; Lew, V.; Khine, M. Shrink film patterning by craft cutter: Complete plastic chips with high resolution/high-aspect ratio channel. Lab Chip 2010, 10, 2472–2475. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Zhao, S.; Pan, T. Lab-on-a-print: From a single polymer film to three-dimensional integrated microfluidics. Lab Chip 2009, 9, 1133–1137. [Google Scholar] [CrossRef] [PubMed]
- Yuen, P.K.; Goral, V.N. Low-cost rapid prototyping of flexible microfluidic devices using a desktop digital craft cutter. Lab Chip 2010, 10, 384–387. [Google Scholar] [CrossRef] [PubMed]
- Halldorsson, S.; Lucumi, E.; Gómez-Sjöberg, R.; Fleming, R.M. Advantages and challenges of microfluidic cell culture in polydimethylsiloxane devices. Biosens. Bioelectron. 2015, 63, 218–231. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, H.-T.; Thach, H.; Roy, E.; Huynh, K.; Perrault, C.M.-T. Low-Cost, Accessible Fabrication Methods for Microfluidics Research in Low-Resource Settings. Micromachines 2018, 9, 461. [Google Scholar] [CrossRef] [PubMed]
- Tsao, C.-W. Polymer microfluidics: Simple, low-cost fabrication process bridging academic lab research to commercialized production. Micromachines 2016, 7, 225. [Google Scholar] [CrossRef]
- Wang, J.D.; Douville, N.J.; Takayama, S.; El Sayed, M. Quantitative analysis of molecular absorption into PDMS microfluidic channels. Ann. Biomed. Eng. 2012, 40, 1862–1873. [Google Scholar] [CrossRef]
- Thach, H.; Nguyen, H.-T.; Tong, U.; Hoang, T.; Vuong, T.-A.; Perrault, C.M.; Huynh, K. Comparison of Nail Polish Meth (Acrylates)(MA) Gel Photoresist and Vinyl Adhesive Paper for Low-Cost Microfluidics Fabrication. In Proceedings of the 6th International Conference on the Development of Biomedical Engineering in Vietnam (BME6), Ho Chi Minh, Vietnam, 27–29 June 2018; Springer: Berlin/Heidelberg, Germany, 2018; pp. 323–329. [Google Scholar]
- Song, Y.; Cheng, D.; Zhao, L.; Lei, K. Introduction: The Origin, Current Status, and Future of Microfluidics. In Microfluidics: Fundamentals, Devices and Applications; Wiley: Hoboken, NJ, USA, 2017. [Google Scholar]
- Arabghahestani, M.; Poozesh, S.; Akafuah, N.K. Advances in computational fluid mechanics in cellular flow manipulation: A review. Appl. Sci. 2019, 9, 4041. [Google Scholar] [CrossRef]
- Viovy, J.-L.; Malaquin, L.; Begolo, S.; Cherif, A.A.; Descroix, S. Microfluidic System. U.S. Patent 20140342373, 20 November 2014. [Google Scholar]
- Kaminski, T.S.; Scheler, O.; Garstecki, P. Droplet microfluidics for microbiology: Techniques, applications and challenges. Lab Chip 2016, 16, 2168–2187. [Google Scholar] [CrossRef]
- Karimi, A.; Yazdi, S.; Ardekani, A. Hydrodynamic mechanisms of cell and particle trapping in microfluidics. Biomicrofluidics 2013, 7, 021501. [Google Scholar] [CrossRef]
- Di Carlo, D.; Irimia, D.; Tompkins, R.G.; Toner, M. Continuous inertial focusing, ordering, and separation of particles in microchannels. Proc. Natl. Acad. Sci. USA 2007, 104, 18892–18897. [Google Scholar] [CrossRef]
- Evander, M.; Johansson, L.; Lilliehorn, T.; Piskur, J.; Lindvall, M.; Johansson, S.; Almqvist, M.; Laurell, T.; Nilsson, J. Noninvasive acoustic cell trapping in a microfluidic perfusion system for online bioassays. Anal. Chem. 2007, 79, 2984–2991. [Google Scholar] [CrossRef]
- Ozcelik, A.; Rufo, J.; Guo, F.; Gu, Y.; Li, P.; Lata, J.; Huang, T.J. Acoustic tweezers for the life sciences. Nat. Methods 2018, 15, 1021–1028. [Google Scholar] [CrossRef]
- Chu, S.; Bjorkholm, J.; Ashkin, A.; Cable, A. Experimental observation of optically trapped atoms. Phys. Rev. Lett. 1986, 57, 314. [Google Scholar] [CrossRef]
- Lenshof, A.; Laurell, T. Continuous separation of cells and particles in microfluidic systems. Chem. Soc. Rev. 2010, 39, 1203–1217. [Google Scholar] [CrossRef]
- Van Reenen, S.; Vitorino, M.; Meskers, S.; Janssen, R.; Kemerink, M. Photoluminescence quenching in films of conjugated polymers by electrochemical doping. Phys. Rev. B 2014, 89, 205206. [Google Scholar] [CrossRef]
- Puri, I.K.; Ganguly, R. Particle transport in therapeutic magnetic fields. Annu. Rev. Fluid Mech. 2014, 46, 407–440. [Google Scholar] [CrossRef]
- Berthier, J.; Brakke, K.A.; Berthier, E. Open Microfluidics; John Wiley & Sons: Hoboken, NJ, USA, 2016. [Google Scholar]
- Melin, J.; van der Wijngaart, W.; Stemme, G. Behaviour and design considerations for continuous flow closed-open-closed liquid microchannels. Lab Chip 2005, 5, 682–686. [Google Scholar] [CrossRef]
- Li, X.; Ballerini, D.R.; Shen, W. A perspective on paper-based microfluidics: Current status and future trends. Biomicrofluidics 2012, 6, 011301. [Google Scholar] [CrossRef]
- Ballerini, D.R.; Li, X.; Shen, W. Flow control concepts for thread-based microfluidic devices. Biomicrofluidics 2011, 5, 014105. [Google Scholar] [CrossRef]
- Jahn, A.; Reiner, J.E.; Vreeland, W.N.; DeVoe, D.L.; Locascio, L.E.; Gaitan, M. Preparation of nanoparticles by continuous-flow microfluidics. J. Nanopart. Res. 2008, 10, 925–934. [Google Scholar] [CrossRef]
- Seemann, R.; Brinkmann, M.; Pfohl, T.; Herminghaus, S. Droplet based microfluidics. Rep. Prog. Phys. 2011, 75, 016601. [Google Scholar] [CrossRef]
- Zhang, Y.; Nguyen, N.-T. Magnetic digital microfluidics—A review. Lab Chip 2017, 17, 994–1008. [Google Scholar] [CrossRef]
- Samiei, E.; Tabrizian, M.; Hoorfar, M. A review of digital microfluidics as portable platforms for lab-on a-chip applications. Lab Chip 2016, 16, 2376–2396. [Google Scholar] [CrossRef]
- Choi, K.; Ng, A.H.; Fobel, R.; Wheeler, A.R. Digital microfluidics. Annu. Rev. Anal. Chem. 2012, 5, 413–440. [Google Scholar] [CrossRef]
- Jebrail, M.J.; Wheeler, A.R. Let’s get digital: Digitizing chemical biology with microfluidics. Curr. Opin. Chem. Biol. 2010, 14, 574–581. [Google Scholar] [CrossRef] [PubMed]
- Ericson, C.; Holm, J.; Ericson, T.; Hjerten, S. Electroosmosis-and pressure-driven chromatography in chips using continuous beds. Anal. Chem. 2000, 72, 81–87. [Google Scholar] [CrossRef]
- Kecskemeti, A.; Gaspar, A. Particle-based liquid chromatographic separations in microfluidic devices—A review. Anal. Chim. Acta 2018, 1021, 1–19. [Google Scholar] [CrossRef]
- Godinho, J.M.; Reising, A.E.; Tallarek, U.; Jorgenson, J.W. Implementation of high slurry concentration and sonication to pack high-efficiency, meter-long capillary ultrahigh pressure liquid chromatography columns. J. Chromatogr. A 2016, 1462, 165–169. [Google Scholar] [CrossRef]
- Depluverez, S.; Daled, S.; De Waele, S.; Planckaert, S.; Schoovaerts, J.; Deforce, D.; Devreese, B.; Devreese, B.; Ledeganckstraat, K. Microfluidics-based LC-MS MRM approach for the relative quantification of Burkholderia cenocepacia secreted virulence factors. Rapid Commun. Mass Spectrom. 2018. [Google Scholar] [CrossRef]
- Li, D.; Chen, H.; Ren, S.; Zhang, Y.; Yang, Y.; Chang, H. Portable liquid chromatography for point-of-care testing of glycated haemoglobin. Sens. Actuators B Chem. 2020, 305, 127484. [Google Scholar] [CrossRef]
- Piendl, S.K.; Geissler, D.; Weigelt, L.; Belder, D. Multiple Heart-Cutting Two-Dimensional Chip-HPLC Combined with Deep-UV Fluorescence and Mass Spectrometric Detection. Anal. Chem. 2020, 92, 3795–3803. [Google Scholar] [CrossRef]
- Wang, S.; Wang, X.; Wang, L.; Pu, Q.; Du, W.; Guo, G. Plasma-assisted alignment in the fabrication of microchannel-array-based in-tube solid-phase microextraction microchips packed with TiO(2) nanoparticles for phosphopeptide analysis. Anal. Chim. Acta 2018, 1018, 70–77. [Google Scholar] [CrossRef]
- Rodríguez-Ruiz, I.; Babenko, V.; Martínez-Rodríguez, S. Protein separation under a microfluidic regime. Analyst 2018, 143, 606–619. [Google Scholar] [CrossRef]
- Nge, P.N.; Pagaduan, J.V.; Yu, M.; Woolley, A.T. Microfluidic chips with reversed-phase monoliths for solid phase extraction and on-chip labeling. J. Chromatogr. A 2012, 1261, 129–135. [Google Scholar] [CrossRef]
- Dziomba, S.; Araya-Farias, M.; Smadja, C.; Taverna, M.; Carbonnier, B.; Tran, N.T. Solid supports for extraction and preconcentration of proteins and peptides in microfluidic devices: A review. Anal. Chim. Acta 2017, 955, 1–26. [Google Scholar] [CrossRef]
- Giddings, J.C. Harnessing electrical forces for separation. Capillary zone electrophoresis, isoelectric focusing, field-flow fractionation, split-flow thin-cell continuous-separation and other techniques. J. Chromatogr. 1989, 480, 21–33. [Google Scholar] [CrossRef]
- Dolník, V. Wall coating for capillary electrophoresis on microchips. Electrophoresis 2004, 25, 3589–3601. [Google Scholar] [CrossRef]
- Zhang, Z.; Yan, B.; Liao, Y.; Liu, H. Nanoparticle: Is it promising in capillary electrophoresis? Anal. Bioanal. Chem. 2008, 391, 925–927. [Google Scholar] [CrossRef]
- Wei, Q.; Becherer, T.; Angioletti-Uberti, S.; Dzubiella, J.; Wischke, C.; Neffe, A.T.; Lendlein, A.; Ballauff, M.; Haag, R. Protein interactions with polymer coatings and biomaterials. Angew. Chem. Int. Ed. 2014, 53, 8004–8031. [Google Scholar] [CrossRef]
- Ouimet, C.M.; D’amico, C.I.; Kennedy, R.T. Advances in capillary electrophoresis and the implications for drug discovery. Expert Opin. Drug Discov. 2017, 12, 213–224. [Google Scholar] [CrossRef]
- Hajba, L.; Guttman, A. Recent advances in column coatings for capillary electrophoresis of proteins. Trac Trends Anal. Chem. 2017, 90, 38–44. [Google Scholar] [CrossRef]
- Sýkora, D.; Kasicka, V.; Miksík, I.; Rezanka, P.; Záruba, K.; Matejka, P.; Král, V. Application of gold nanoparticles in separation sciences. J. Sep. Sci. 2010, 33, 372–387. [Google Scholar] [CrossRef] [PubMed]
- Tu, Q.; Wang, J.-C.; Zhang, Y.; Liu, R.; Liu, W.; Ren, L.; Shen, S.; Xu, J.; Zhao, L.; Wang, J. Surface modification of poly (dimethylsiloxane) and its applications in microfluidics-based biological analysis. Rev. Anal. Chem. 2012, 31, 177–192. [Google Scholar] [CrossRef]
- Zhou, J.; Khodakov, D.A.; Ellis, A.V.; Voelcker, N.H. Surface modification for PDMS-based microfluidic devices. Electrophoresis 2012, 33, 89–104. [Google Scholar] [CrossRef] [PubMed]
- Breadmore, M.C.; Grochocki, W.; Kalsoom, U.; Alves, M.N.; Phung, S.C.; Rokh, M.T.; Cabot, J.M.; Ghiasvand, A.; Li, F.; Shallan, A.I.; et al. Recent advances in enhancing the sensitivity of electrophoresis and electrochromatography in capillaries and microchips (2016–2018). Electrophoresis 2019, 40, 17–39. [Google Scholar] [CrossRef] [PubMed]
- Šlampová, A.; Malá, Z.; Gebauer, P. Recent progress of sample stacking in capillary electrophoresis (2016–2018). Electrophoresis 2019, 40, 40–54. [Google Scholar] [CrossRef] [PubMed]
- Arvin, N.E.; Dawod, M.; Lamb, D.T.; Anderson, J.P.; Furtaw, M.D.; Kennedy, R.T. Fast Immunoassay for Microfluidic Western Blotting by Direct Deposition of Reagents onto Capture Membrane. Anal. Methods Adv. Methods Appl. 2020, 12, 1606–1616. [Google Scholar] [CrossRef] [PubMed]
- Jin, S.; Furtaw, M.D.; Chen, H.; Lamb, D.T.; Ferguson, S.A.; Arvin, N.E.; Dawod, M.; Kennedy, R.T. Multiplexed Western Blotting Using Microchip Electrophoresis. Anal. Chem. 2016, 88, 6703–6710. [Google Scholar] [CrossRef]
- Štěpánová, S.; Kašička, V. Analysis of proteins and peptides by electromigration methods in microchips. J. Sep. Sci. 2017, 40, 228–250. [Google Scholar] [CrossRef]
- Giddings, J.C. Field-Flow Fractionation. Chem. Eng. News Arch. 1988, 66, 34–45. [Google Scholar] [CrossRef]
- Davanlou, A.; Reddy, V. Numerical Simulation and Optimization of Electric Field-Based Sorting in Particle Laden Flows in Microchannels. In Proceedings of the ASME 2018 5th Joint US-European Fluids Engineering Division Summer Meeting, Montreal, QC, Canada, 15–20 July 2018; p. V002T009A020. [Google Scholar]
- Geissler, D.; Heiland, J.J.; Lotter, C.; Belder, D. Microchip HPLC separations monitored simultaneously by coherent anti-Stokes Raman scattering and fluorescence detection. Microchim. Acta 2017, 184, 315–321. [Google Scholar] [CrossRef]
- Toraño, J.S.; Ramautar, R.; de Jong, G. Advances in capillary electrophoresis for the life sciences. J. Chromatogr. B 2019, 1118, 116–136. [Google Scholar] [CrossRef] [PubMed]
- Wagner, M.; Holzschuh, S.; Traeger, A.; Fahr, A.; Schubert, U.S. Asymmetric flow field-flow fractionation in the field of nanomedicine. Anal. Chem. 2014, 86, 5201–5210. [Google Scholar] [CrossRef] [PubMed]
- Shendruk, T.N.; Tahvildari, R.; Catafard, N.M.; Andrzejewski, L.; Gigault, C.; Todd, A.; Gagne-Dumais, L.; Slater, G.W.; Godin, M. Field-flow fractionation and hydrodynamic chromatography on a microfluidic chip. Anal. Chem. 2013, 85, 5981–5988. [Google Scholar] [CrossRef] [PubMed]
- Condina, M.R.; Dilmetz, B.A.; Bazaz, S.R.; Meneses, J.; Warkiani, M.E.; Hoffmann, P. Rapid separation and identification of beer spoilage bacteria by inertial microfluidics and MALDI-TOF mass spectrometry. Lab Chip 2019, 19, 1961–1970. [Google Scholar] [CrossRef] [PubMed]
- Stolz, A.; Jooß, K. Recent advances in capillary electrophoresis-mass spectrometry: Instrumentation, methodology and applications. Electrophoresis 2019, 40, 79–112. [Google Scholar] [CrossRef]
- Sierra, T.; Crevillen, A.G.; Escarpa, A. Electrochemical detection based on nanomaterials in CE and microfluidic systems. Electrophoresis 2019, 40, 113–123. [Google Scholar] [CrossRef]
- García-Carmona, L.; Martín, A.; Sierra, T.; González, M.C.; Escarpa, A. Electrochemical detectors based on carbon and metallic nanostructures in capillary and microchip electrophoresis. Electrophoresis 2017, 38, 80–94. [Google Scholar] [CrossRef]
- Martín, A.; Kim, J.; Kurniawan, J.F.; Sempionatto, J.R.; Moreto, J.R.; Tang, G.; Campbell, A.S.; Shin, A.; Lee, M.Y.; Liu, X. Epidermal microfluidic electrochemical detection system: Enhanced sweat sampling and metabolite detection. ACS Sens. 2017, 2, 1860–1868. [Google Scholar] [CrossRef]
- Wang, Y.; Xu, H.; Luo, J.; Liu, J.; Wang, L.; Fan, Y.; Yan, S.; Yang, Y.; Cai, X. A novel label-free microfluidic paper-based immunosensor for highly sensitive electrochemical detection of carcinoembryonic antigen. Biosens. Bioelectron. 2016, 83, 319–326. [Google Scholar] [CrossRef] [PubMed]
- Liang, W.; Lin, H.; Chen, J.; Chen, C. Utilization of nanoparticles in microfluidic systems for optical detection. Microsyst. Technol. 2016, 22, 2363–2370. [Google Scholar] [CrossRef]
- Gubatayao, T.C.; Handique, K.; Ganesan, K.; Drummond, D.M. Scanning Real-Time Microfluidic Thermocycler and Methods for Synchronized Thermocycling and Scanning Optical Detection. U.S. Patent US9765389B2, 19 September 2017. [Google Scholar]
- Jeon, S.; Kwon, Y.W.; Park, J.Y.; Hong, S.W. Fluorescent Detection of Bovine Serum Albumin Using Surface Imprinted Films Formed on PDMS Microfluidic Channels. J. Nanosci. Nanotechnol. 2019, 19, 4736–4739. [Google Scholar] [CrossRef]
- Burger, R.; Amato, L.; Boisen, A. Detection methods for centrifugal microfluidic platforms. Biosens. Bioelectron. 2016, 76, 54–67. [Google Scholar] [CrossRef]
- Srinivas, P.R. Introduction to Protein Electrophoresis. In Electrophoretic Separation of Proteins; Springer: Berlin/Heidelberg, Germany, 2019; pp. 23–29. [Google Scholar]
- Westermeier, R. Looking at proteins from two dimensions: A review on five decades of 2D electrophoresis. Arch. Physiol. Biochem. 2014, 120, 168–172. [Google Scholar] [CrossRef]
- Righetti, P.G.; Candiano, G. Recent advances in electrophoretic techniques for the characterization of protein biomolecules: A poker of aces. J. Chromatogr. A 2011, 1218, 8727–8737. [Google Scholar] [CrossRef]
- Fekete, S.; Guillarme, D.; Sandra, P.; Sandra, K. Chromatographic, electrophoretic, and mass spectrometric methods for the analytical characterization of protein biopharmaceuticals. Anal. Chem. 2016, 88, 480–507. [Google Scholar] [CrossRef] [PubMed]
- Kašička, V. Recent developments in capillary and microchip electroseparations of peptides (2017–mid 2019). Electrophoresis 2020, 41, 10–35. [Google Scholar] [CrossRef] [PubMed]
- Štěpánová, S.; Kašička, V. Recent applications of capillary electromigration methods to separation and analysis of proteins. Anal. Chim. Acta 2016, 933, 23–42. [Google Scholar] [CrossRef]
- Štěpánová, S.; Kašička, V. Recent developments and applications of capillary and microchip electrophoresis in proteomics and peptidomics (2015–mid 2018). J. Sep. Sci. 2019, 42, 398–414. [Google Scholar] [CrossRef]
- Catherman, A.D.; Skinner, O.S.; Kelleher, N.L. Top down proteomics: Facts and perspectives. Biochem. Biophys. Res. Commun. 2014, 445, 683–693. [Google Scholar] [CrossRef] [PubMed]
- Puangpila, C.; Mayadunne, E.; El Rassi, Z. Liquid phase based separation systems for depletion, prefractionation, and enrichment of proteins in biological fluids and matrices for in-depth proteomics analysis—An update covering the period 2011–2014. Electrophoresis 2015, 36, 238–252. [Google Scholar] [CrossRef] [PubMed]
- Giordano, B.C.; Burgi, D.S.; Hart, S.J.; Terray, A. On-line sample pre-concentration in microfluidic devices: A review. Anal. Chim. Acta 2012, 718, 11–24. [Google Scholar] [CrossRef]
- Gharari, H.; Farjaminezhad, M.; Marefat, A.; Fakhari, A.R. All-in-one solid-phase microextraction: Development of a selective solid-phase microextraction fiber assembly for the simultaneous and efficient extraction of analytes with different polarities. J. Sep. Sci. 2016, 39, 1709–1716. [Google Scholar] [CrossRef]
- Bang, Y.; Hwang, Y.; Lee, S.; Park, S.; Bae, S. Sol–gel-adsorbent-coated extraction needles to detect volatile compounds in spoiled fish. J. Sep. Sci. 2017, 40, 3839–3847. [Google Scholar] [CrossRef]
- Dawod, M.; Arvin, N.E.; Kennedy, R.T. Recent advances in protein analysis by capillary and microchip electrophoresis. Analyst 2017, 142, 1847–1866. [Google Scholar] [CrossRef]
- Ríos, Á.; Zougagh, M.; Avila, M. Miniaturization through lab-on-a-chip: Utopia or reality for routine laboratories? A review. Anal. Chim. Acta 2012, 740, 1–11. [Google Scholar] [CrossRef]
- Sonker, M.; Parker, E.K.; Nielsen, A.V.; Sahore, V.; Woolley, A.T. Electrokinetically operated microfluidic devices for integrated immunoaffinity monolith extraction and electrophoretic separation of preterm birth biomarkers. Analyst 2018, 143, 224–231. [Google Scholar] [CrossRef]
- Noach-Hirsh, M.; Nevenzal, H.; Glick, Y.; Chorni, E.; Avrahami, D.; Barbiro-Michaely, E.; Gerber, D.; Tzur, A. Integrated microfluidics for protein modification discovery. Mol. Cell. Proteom. 2015, 14, 2824–2832. [Google Scholar] [CrossRef] [PubMed]
- Lazar, I.M.; Gulakowski, N.S.; Lazar, A.C. Protein and proteome measurements with microfluidic chips. Anal. Chem. 2019, 92, 169–182. [Google Scholar] [CrossRef] [PubMed]
- Fan, B.; Li, X.; Chen, D.; Peng, H.; Wang, J.; Chen, J. Development of microfluidic systems enabling high-throughput single-cell protein characterization. Sensors 2016, 16, 232. [Google Scholar] [CrossRef]
- Santos, R.S.; Figueiredo, C.; Azevedo, N.F.; Braeckmans, K.; De Smedt, S.C. Nanomaterials and molecular transporters to overcome the bacterial envelope barrier: Towards advanced delivery of antibiotics. Adv. Drug Deliv. Rev. 2018, 136, 28–48. [Google Scholar] [CrossRef] [PubMed]
- Kolarević, S.; Milovanović, D.; Avdović, M.; Oalđe, M.; Kostić, J.; Sunjog, K.; Nikolić, B.; Knežević-Vukčević, J.; Vuković-Gačić, B. Optimisation of the microdilution method for detection of minimum inhibitory concentration values in selected bacteria. Bot. Serbica 2016, 40, 29–36. [Google Scholar]
- Al Nahas, K.; Cama, J.; Schaich, M.; Hammond, K.; Deshpande, S.; Dekker, C.; Ryadnov, M.; Keyser, U. A microfluidic platform for the characterisation of membrane active antimicrobials. Lab Chip 2019, 19, 837–844. [Google Scholar] [CrossRef]
- Epand, R.M.; Walker, C.; Epand, R.F.; Magarvey, N.A. Molecular mechanisms of membrane targeting antibiotics. Biochim. Biophys. Acta BBA Biomembr. 2016, 1858, 980–987. [Google Scholar] [CrossRef]
- Estemalik, J.; Demko, C.; Bissada, N.F.; Joshi, N.; Bodner, D.; Shankar, E.; Gupta, S. Simultaneous Detection of Oral Pathogens in Subgingival Plaque and Prostatic Fluid of Men with Periodontal and Prostatic Diseases. J. Periodontol. 2017, 88, 823–829. [Google Scholar] [CrossRef] [PubMed]
- Martínez, M.; Polizzotto, A.; Flores, N.; Semorile, L.; Maffía, P.C. Antibacterial, anti-biofilm and in vivo activities of the antimicrobial peptides P5 and P6. 2. Microb. Pathog. 2020, 139, 103886. [Google Scholar] [CrossRef] [PubMed]
- Deshpande, S.; Caspi, Y.; Meijering, A.E.; Dekker, C. Octanol-assisted liposome assembly on chip. Nat. Commun. 2016, 7, 1–9. [Google Scholar] [CrossRef]
- Yandrapalli, N.; Robinson, T. Ultra-high capacity microfluidic trapping of giant vesicles for high-throughput membrane studies. Lab Chip 2019, 19, 626–633. [Google Scholar] [CrossRef]
- Prestinaci, F.; Pezzotti, P.; Pantosti, A. Antimicrobial resistance: A global multifaceted phenomenon. Pathog. Glob. Health 2015, 109, 309–318. [Google Scholar] [CrossRef] [PubMed]
- Ferri, M.; Ranucci, E.; Romagnoli, P.; Giaccone, V. Antimicrobial resistance: A global emerging threat to public health systems. Crit. Rev. Food Sci. Nutr. 2017, 57, 2857–2876. [Google Scholar] [CrossRef] [PubMed]
- Ayukekbong, J.A.; Ntemgwa, M.; Atabe, A.N. The threat of antimicrobial resistance in developing countries: Causes and control strategies. Antimicrob. Resist. Infect. Control 2017, 6, 47. [Google Scholar] [CrossRef] [PubMed]
- Marston, H.D.; Dixon, D.M.; Knisely, J.M.; Palmore, T.N.; Fauci, A.S. Antimicrobial resistance. JAMA 2016, 316, 1193–1204. [Google Scholar] [CrossRef]
- Lee, W.-B.; Chien, C.-C.; You, H.-L.; Kuo, F.-C.; Lee, M.S.; Lee, G.-B. An integrated microfluidic system for antimicrobial susceptibility testing with antibiotic combination. Lab Chip 2019, 19, 2699–2708. [Google Scholar] [CrossRef] [PubMed]
- Périllaud-Dubois, C.; Pilmis, B.; Diep, J.; de Ponfilly, G.P.; Perreau, S.; d’Epenoux, L.R.; Mizrahi, A.; Couzigou, C.; Vidal, B.; Le Monnier, A. Performance of rapid antimicrobial susceptibility testing by disk diffusion on MHR-SIR agar directly on urine specimens. Eur. J. Clin. Microbiol. Infect. Dis. 2019, 38, 185–189. [Google Scholar] [CrossRef]
- Stern, E.; Flentie, K.; Phelan, N. Assays for Improving Automated Antimicrobial Susceptibility Testing Accuracy. International Patent Application No. PCT/US2020/041547, 14 January 2021. [Google Scholar]
- Liu, Z.; Banaei, N.; Ren, K. Microfluidics for combating antimicrobial resistance. Trends Biotechnol. 2017, 35, 1129–1139. [Google Scholar] [CrossRef] [PubMed]
- Hassan, S.-U.; Zhang, X. Microfluidics as an Emerging Platform for Tackling Antimicrobial Resistance (AMR): A Review. Curr. Anal. Chem. 2020, 16, 41–51. [Google Scholar] [CrossRef]
- Chiang, Y.-L.; Lin, C.-H.; Yen, M.-Y.; Su, Y.-D.; Chen, S.-J.; Chen, H.-f. Innovative antimicrobial susceptibility testing method using surface plasmon resonance. Biosens. Bioelectron. 2009, 24, 1905–1910. [Google Scholar] [CrossRef]
- Needs, S.H.; Donmez, S.I.; Bull, S.P.; McQuaid, C.; Osborn, H.M.I.; Edwards, A.D. Challenges in Microfluidic and Point-of-Care Phenotypic Antimicrobial Resistance Tests. Front. Mech. Eng. 2020, 6. [Google Scholar] [CrossRef]
- He, C.; Yin, L.; Tang, C.; Yin, C. Multifunctional polymeric nanoparticles for oral delivery of TNF-α siRNA to macrophages. Biomaterials 2013, 34, 2843–2854. [Google Scholar] [CrossRef]
- Avesar, J.; Rosenfeld, D.; Truman-Rosentsvit, M.; Ben-Arye, T.; Geffen, Y.; Bercovici, M.; Levenberg, S. Rapid phenotypic antimicrobial susceptibility testing using nanoliter arrays. Proc. Natl. Acad. Sci. USA 2017, 114, E5787–E5795. [Google Scholar] [CrossRef]
- Lee, W.-B.; Fu, C.-Y.; Chang, W.-H.; You, H.-L.; Wang, C.-H.; Lee, M.S.; Lee, G.-B. A microfluidic device for antimicrobial susceptibility testing based on a broth dilution method. Biosens. Bioelectron. 2017, 87, 669–678. [Google Scholar] [CrossRef]
- Sun, H.; Chan, C.-W.; Wang, Y.; Yao, X.; Mu, X.; Lu, X.; Zhou, J.; Cai, Z.; Ren, K. Reliable and reusable whole polypropylene plastic microfluidic devices for a rapid, low-cost antimicrobial susceptibility test. Lab Chip 2019, 19, 2915–2924. [Google Scholar] [CrossRef] [PubMed]
- Boedicker, J.Q.; Li, L.; Kline, T.R.; Ismagilov, R.F. Detecting bacteria and determining their susceptibility to antibiotics by stochastic confinement in nanoliter droplets using plug-based microfluidics. Lab Chip 2008, 8, 1265–1272. [Google Scholar] [CrossRef] [PubMed]
- Mohan, R.; Mukherjee, A.; Sevgen, S.E.; Sanpitakseree, C.; Lee, J.; Schroeder, C.M.; Kenis, P.J. A multiplexed microfluidic platform for rapid antibiotic susceptibility testing. Biosens. Bioelectron. 2013, 49, 118–125. [Google Scholar] [CrossRef] [PubMed]
- Schudel, B.R.; Choi, C.J.; Cunningham, B.T.; Kenis, P.J. Microfluidic chip for combinatorial mixing and screening of assays. Lab Chip 2009, 9, 1676–1680. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Liu, Z.; Hu, C.; Ren, K. Cell-on-hydrogel platform made of agar and alginate for rapid, low-cost, multidimensional test of antimicrobial susceptibility. Lab Chip 2016, 16, 3130–3138. [Google Scholar] [CrossRef]
- Furlan, C.; Dirks, R.A.; Thomas, P.C.; Jones, R.C.; Wang, J.; Lynch, M.; Marks, H.; Vermeulen, M. Miniaturised interaction proteomics on a microfluidic platform with ultra-low input requirements. Nat. Commun. 2019, 10, 1–8. [Google Scholar] [CrossRef]
- Branon, T.C.; Bosch, J.A.; Sanchez, A.D.; Udeshi, N.D.; Svinkina, T.; Carr, S.A.; Feldman, J.L.; Perrimon, N.; Ting, A.Y. Efficient proximity labeling in living cells and organisms with TurboID. Nat. Biotechnol. 2018, 36, 880–887. [Google Scholar] [CrossRef]
- Chen, J.-X.; Cipriani, P.G.; Mecenas, D.; Polanowska, J.; Piano, F.; Gunsalus, K.C.; Selbach, M. In vivo interaction proteomics in Caenorhabditis elegans embryos provides new insights into P granule dynamics. Mol. Cell. Proteom. 2016, 15, 1642–1657. [Google Scholar] [CrossRef]
- Lam, S.S.; Martell, J.D.; Kamer, K.J.; Deerinck, T.J.; Ellisman, M.H.; Mootha, V.K.; Ting, A.Y. Directed evolution of APEX2 for electron microscopy and proximity labeling. Nat. Methods 2015, 12, 51. [Google Scholar] [CrossRef]
- Malovannaya, A.; Lanz, R.B.; Jung, S.Y.; Bulynko, Y.; Le, N.T.; Chan, D.W.; Ding, C.; Shi, Y.; Yucer, N.; Krenciute, G. Analysis of the human endogenous coregulator complexome. Cell 2011, 145, 787–799. [Google Scholar] [CrossRef]
- Rolland, T.; Taşan, M.; Charloteaux, B.; Pevzner, S.J.; Zhong, Q.; Sahni, N.; Yi, S.; Lemmens, I.; Fontanillo, C.; Mosca, R. A proteome-scale map of the human interactome network. Cell 2014, 159, 1212–1226. [Google Scholar] [CrossRef]
- Roux, K.J.; Kim, D.I.; Raida, M.; Burke, B. A promiscuous biotin ligase fusion protein identifies proximal and interacting proteins in mammalian cells. J. Cell Biol. 2012, 196, 801–810. [Google Scholar] [CrossRef]
- Smits, A.H.; Vermeulen, M. Characterizing protein–protein interactions using mass spectrometry: Challenges and opportunities. Trends Biotechnol. 2016, 34, 825–834. [Google Scholar] [CrossRef] [PubMed]
- Swami, M. A discovery strategy for novel cancer biomarkers. Nat. Rev. Cancer 2010, 10, 597. [Google Scholar] [CrossRef] [PubMed]
- Hunyadi-Gulyás, É.; Medzihradszky, K.F. Factors that contribute to the complexity of protein digests. Drug Discov. Today Targets 2004, 3, 3–10. [Google Scholar] [CrossRef]
- Luk, V.N.; Fiddes, L.K.; Luk, V.M.; Kumacheva, E.; Wheeler, A.R. Digital microfluidic hydrogel microreactors for proteomics. Proteomics 2012, 12, 1310–1318. [Google Scholar] [CrossRef] [PubMed]
- Leipert, J.; Tholey, A. Miniaturized sample preparation on a digital microfluidics device for sensitive bottom-up microproteomics of mammalian cells using magnetic beads and mass spectrometry-compatible surfactants. Lab Chip 2019, 19, 3490–3498. [Google Scholar] [CrossRef] [PubMed]
- Yi, L.; Piehowski, P.D.; Shi, T.; Smith, R.D.; Qian, W.J. Advances in microscale separations towards nanoproteomics applications. J. Chromatogr. A 2017, 1523, 40–48. [Google Scholar] [CrossRef]
- Specht, H.; Slavov, N. Transformative Opportunities for Single-Cell Proteomics. J. Proteome Res. 2018, 17, 2565–2571. [Google Scholar] [CrossRef] [PubMed]
- Kasuga, K.; Katoh, Y.; Nagase, K.; Igarashi, K. Microproteomics with microfluidic-based cell sorting: Application to 1000 and 100 immune cells. Proteomics 2017, 17. [Google Scholar] [CrossRef]
- Liu, W.-W.; Zhu, Y. “Development and application of analytical detection techniques for droplet-based microfluidics”—A Review. Anal. Chim. Acta 2020. [Google Scholar] [CrossRef]
- Pang, L.; Ding, J.; Liu, X.-X.; Fan, S.-K. Digital microfluidics for cell manipulation. TrAC Trends Anal. Chem. 2019. [Google Scholar] [CrossRef]
- Rackus, D.G.; de Campos, R.P.; Chan, C.; Karcz, M.M.; Seale, B.; Narahari, T.; Dixon, C.; Chamberlain, M.D.; Wheeler, A.R. Pre-concentration by liquid intake by paper (P-CLIP): A new technique for large volumes and digital microfluidics. Lab Chip 2017, 17, 2272–2280. [Google Scholar] [CrossRef] [PubMed]
- Rahimi, A.; Mahdavi, H. Zwitterionic-functionalized GO/PVDF nanocomposite membranes with improved anti-fouling properties. J. Water Process Eng. 2019, 32, 100960. [Google Scholar] [CrossRef]
- Pedde, R.D.; Li, H.; Borchers, C.H.; Akbari, M. Microfluidic-mass spectrometry interfaces for translational proteomics. Trends Biotechnol. 2017, 35, 954–970. [Google Scholar] [CrossRef]
- Lin, L.; Lin, J.-M. Microfluidics-Mass Spectrometry for Cell Analysis. In Cell Analysis on Microfluidics; Springer: Berlin/Heidelberg, Germany, 2018; pp. 291–311. [Google Scholar]
- Reddy, P.J.; Gollapalli, K.; Ghantasala, S.; Das, T.; Patel, S.K.; Chanukuppa, V.; Srivastava, S.; Rapole, S. Basics of Mass Spectrometry and Its Applications in Biomarker Discovery. In Biomarker Discovery in the Developing World: Dissecting the Pipeline for Meeting the Challenges; Springer: Berlin/Heidelberg, Germany, 2016; pp. 41–63. [Google Scholar]
- Abouleila, Y.; Onidani, K.; Ali, A.; Shoji, H.; Kawai, T.; Lim, C.T.; Kumar, V.; Okaya, S.; Kato, K.; Hiyama, E. Live single cell mass spectrometry reveals cancer-specific metabolic profiles of circulating tumor cells. Cancer Sci. 2019, 110, 697–706. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.; Araujo, R.; Howard, M.; Magni, R.; Liotta, L.A.; Luchini, A. Affinity enrichment for mass spectrometry: Improving the yield of low abundance biomarkers. Expert Rev. Proteom. 2018, 15, 353–366. [Google Scholar] [CrossRef]
- El-Aneed, A.; Cohen, A.; Banoub, J. Mass Spectrometry, Review of the Basics: Electrospray, MALDI, and Commonly Used Mass Analyzers. Appl. Spectrosc. Rev. 2009, 44, 210–230. [Google Scholar] [CrossRef]
- Wang, H.; Chen, L.; Sun, L. Digital microfluidics: A promising technique for biochemical applications. Front. Mech. Eng. 2017, 12, 510–525. [Google Scholar] [CrossRef]
- Jie, M.; Mao, S.; Li, H.; Lin, J.-M. Multi-channel microfluidic chip-mass spectrometry platform for cell analysis. Chin. Chem. Lett. 2017, 28, 1625–1630. [Google Scholar] [CrossRef]
- Gao, D.; Jin, F.; Zhou, M.; Jiang, Y. Recent advances in single cell manipulation and biochemical analysis on microfluidics. Analyst 2019, 144, 766–781. [Google Scholar] [CrossRef] [PubMed]
- Shang, L.; Cheng, Y.; Zhao, Y. Emerging droplet microfluidics. Chem. Rev. 2017, 117, 7964–8040. [Google Scholar] [CrossRef]
- Cui, P.; Wang, S. Application of microfluidic chip technology in pharmaceutical analysis: A review. J. Pharm. Anal. 2019, 9, 238–247. [Google Scholar] [CrossRef] [PubMed]
- Týčová, A.; Ledvina, V.; Klepárník, K. Recent advances in CE-MS coupling: Instrumentation, methodology, and applications. Electrophoresis 2017, 38, 115–134. [Google Scholar] [CrossRef] [PubMed]
- Hughes, A.J.; Lin, R.K.; Peehl, D.M.; Herr, A.E. Microfluidic integration for automated targeted proteomic assays. Proc. Natl. Acad. Sci. USA 2012, 109, 5972–5977. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Sun, Y.; Neoh, K.H.; Chen, A.; Li, W.; Yang, X.; Han, R.P.S. Microfluidic assay of circulating endothelial cells in coronary artery disease patients with angina pectoris. PLoS ONE 2017, 12, e0181249. [Google Scholar] [CrossRef]
- Jones, A.L.; Dhanapala, L.; Baldo, T.A.; Sharafeldin, M. Prostate Cancer Diagnosis in the Clinic Using an 8-Protein Biomarker Panel. Anal. Chem. 2021, 93, 1059–1067. [Google Scholar] [CrossRef]
- Bohr, A.; Colombo, S.; Jensen, H. Future of Microfluidics in Research and in the Market. In Microfluidics for Pharmaceutical Applications; Elsevier: Amsterdam, The Netherlands, 2019; pp. 425–465. [Google Scholar]
- Du, G.; Fang, Q.; den Toonder, J.M. Microfluidics for cell-based high throughput screening platforms—A review. Anal. Chim. Acta 2016, 903, 36–50. [Google Scholar] [CrossRef]
- Wang, Y.; Li, Q.; Shi, H.; Tang, K.; Qiao, L.; Yu, G.; Ding, C.; Yu, S. Microfluidic Raman biochip detection of exosomes: A promising tool for prostate cancer diagnosis. Lab Chip 2020, 20, 4632–4637. [Google Scholar] [CrossRef]
- Kang, J.H.; Park, J.-K. Technical paper on microfluidic devices-cell separation technology. Asia Pac. Biotech News 2005, 9, 1135–1146. [Google Scholar]
- Chiu, Y.-Y.; Huang, C.-K.; Lu, Y.-W. Enhancement of microfluidic particle separation using cross-flow filters with hydrodynamic focusing. Biomicrofluidics 2016, 10, 011906. [Google Scholar] [CrossRef] [PubMed]
- Menon, N.V.; Lim, S.B.; Lim, C.T. Microfluidics for personalized drug screening of cancer. Curr. Opin. Pharmacol. 2019, 48, 155–161. [Google Scholar] [CrossRef] [PubMed]
- Kulasinghe, A.; Wu, H.; Punyadeera, C.; Warkiani, M.E. The use of microfluidic technology for cancer applications and liquid biopsy. Micromachines 2018, 9, 397. [Google Scholar] [CrossRef] [PubMed]
- Pakjesm Pourfard, P. Single Cell Enrichment with High Throughput Microfluidic Devices. Master’s Thesis, University of California Irvine, Irvine, CA, USA, 2017. [Google Scholar]
- Zhu, P.; Wang, L. Passive and active droplet generation with microfluidics: A review. Lab Chip 2017, 17, 34–75. [Google Scholar] [CrossRef] [PubMed]
- Lei, C.; Kobayashi, H.; Wu, Y.; Li, M.; Isozaki, A.; Yasumoto, A.; Mikami, H.; Ito, T.; Nitta, N.; Sugimura, T. High-throughput imaging flow cytometry by optofluidic time-stretch microscopy. Nat. Protoc. 2018, 13, 1603–1631. [Google Scholar] [CrossRef]
- Liu, N.; Petchakup, C.; Tay, H.M.; Li, K.H.H.; Hou, H.W. Spiral Inertial Microfluidics for Cell Separation and Biomedical Applications. In Applications of Microfluidic Systems in Biology and Medicine; Springer: Berlin/Heidelberg, Germany, 2019; pp. 99–150. [Google Scholar]
- Su, W.; Li, H.; Chen, W.; Qin, J. Microfluidic strategies for label-free exosomes isolation and analysis. TrAC Trends Anal. Chem. 2019. [Google Scholar] [CrossRef]
- Chagas, C.L.; Moreira, R.C.; Bressan, L.P.; de Jesus, D.P.; da Silva, J.A.; Coltro, W.K. Instrumental Platforms for Capillary and Microchip Electromigration Separation Techniques. In Capillary Electromigration Separation Methods; Elsevier: Amsterdam, The Netherlands, 2018; pp. 269–292. [Google Scholar]
- Pamme, N. Continuous flow separations in microfluidic devices. Lab Chip 2007, 7, 1644–1659. [Google Scholar] [CrossRef]
- Kielpinski, M.; Walther, O.; Cao, J.; Henkel, T.; Köhler, J.M.; Groß, G.A. Microfluidic Chamber Design for Controlled Droplet Expansion and Coalescence. Micromachines 2020, 11, 394. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Zhang, N.; Chen, J.; Rodgers, V.G.; Brisk, P.; Grover, W.H. Finding the optimal design of a passive microfluidic mixer. Lab Chip 2019, 19, 3618–3627. [Google Scholar] [CrossRef]
- Prakash, J.; Tripathi, D. Electroosmotic flow of Williamson ionic nanoliquids in a tapered microfluidic channel in presence of thermal radiation and peristalsis. J. Mol. Liq. 2018, 256, 352–371. [Google Scholar] [CrossRef]
- Ahmad, I.L.; Ahmad, M.R.; Takeuchi, M.; Nakajima, M.; Hasegawa, Y. Tapered Microfluidic for Continuous Micro-Object Separation Based on Hydrodynamic Principle. IEEE Trans. Biomed. Circuits Syst. 2017, 11, 1413–1421. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, I.L.; Ahmad, M.R. Tapered microchannel for multi-particles passive separation based on hydrodynamic resistance. Indones. J. Electr. Eng. Comput. Sci. 2017, 5, 628–635. [Google Scholar] [CrossRef]
- Voigt, A.; Schreiter, J.; Frank, P.; Pini, C.; Mayr, C.; Richter, A. Method for the Computer-aided Schematic Design and Simulation of Hydrogel-based Microfluidic Systems. IEEE Trans. Comput. Aided Des. Integr. Circuits Syst. 2019. [Google Scholar] [CrossRef]
- Naderi, A.; Bhattacharjee, N.; Folch, A. Digital Manufacturing for Microfluidics. Annu. Rev. Biomed. Eng. 2019, 21, 325–364. [Google Scholar] [CrossRef]
- Sun, J.; Moore, L.; Xue, W.; Kim, J.; Zborowski, M.; Chalmers, J.J. Correlation of simulation/finite element analysis to the separation of intrinsically magnetic spores and red blood cells using a microfluidic magnetic deposition system. Biotechnol. Bioeng. 2018, 115, 1288–1300. [Google Scholar] [CrossRef]
- Shamloo, A.; Boodaghi, M. Design and simulation of a microfluidic device for acoustic cell separation. Ultrasonics 2018, 84, 234–243. [Google Scholar] [CrossRef]
- Hu, Q.; Luni, C.; Elvassore, N. Microfluidics for secretome analysis under enhanced endogenous signaling. Biochem. Biophys. Res. Commun. 2018, 497, 480–484. [Google Scholar] [CrossRef]
- Liu, F.; KC, P.; Ni, L.; Zhang, G.; Zhe, J. A microfluidic competitive immuno-aggregation assay for high sensitivity cell secretome detection. Organogenesis 2018, 14, 67–81. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Wang, J.; Wu, L.; Guo, J.; Song, Y.; Tian, T.; Wang, W.; Zhu, Z.; Yang, C. Microfluidic Single-Cell Omics Analysis. Small 2020, 16, 1903905. [Google Scholar] [CrossRef] [PubMed]
- Caen, O.; Lu, H.; Nizard, P.; Taly, V. Microfluidics as a strategic player to decipher single-cell omics? Trends Biotechnol. 2017, 35, 713–727. [Google Scholar] [CrossRef] [PubMed]
- Deng, Y.; Finck, A.; Fan, R. Single-cell omics analyses enabled by microchip technologies. Annu. Rev. Biomed. Eng. 2019, 21, 365–393. [Google Scholar] [CrossRef] [PubMed]
- Lazar, I.M.; Deng, J.; Stremler, M.A.; Ahuja, S. Microfluidic reactors for advancing the MS analysis of fast biological responses. Microsyst. Nanoeng. 2019, 5, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Bahrami, S.; Drabløs, F. Gene regulation in the immediate-early response process. Adv. Biol. Regul. 2016, 62, 37–49. [Google Scholar] [CrossRef] [PubMed]
- Fowler, T.; Sen, R.; Roy, A.L. Regulation of primary response genes. Mol. Cell 2011, 44, 348–360. [Google Scholar] [CrossRef] [PubMed]
- Dungan, J.; Mathews, J.; Levin, M.; Koomson, V. Microfluidic platform to study intercellular connectivity through on-chip electrical impedance measurement. In Proceedings of the 2017 IEEE 60th International Midwest Symposium on Circuits and Systems (MWSCAS), Medford, MA, USA, 6–9 August 2017; pp. 56–59. [Google Scholar]
- Weiss, A.C.; Kempe, K.; Förster, S.; Caruso, F. Microfluidic Examination of the “Hard” Biomolecular Corona Formed on Engineered Particles in Different Biological Milieu. Biomacromolecules 2018, 19, 2580–2594. [Google Scholar] [CrossRef]
- Digiacomo, L.; Palchetti, S.; Giulimondi, F.; Pozzi, D.; Chiozzi, R.Z.; Capriotti, A.L.; Laganà, A.; Caracciolo, G. The biomolecular corona of gold nanoparticles in a controlled microfluidic environment. Lab Chip 2019, 19, 2557–2567. [Google Scholar] [CrossRef] [PubMed]
- Palchetti, S.; Colapicchioni, V.; Digiacomo, L.; Caracciolo, G.; Pozzi, D.; Capriotti, A.L.; La Barbera, G.; Laganà, A. The protein corona of circulating PEGylated liposomes. Biochim. Biophys. Acta BBA Biomembr. 2016, 1858, 189–196. [Google Scholar] [CrossRef] [PubMed]
- Palchetti, S.; Pozzi, D.; Capriotti, A.L.; La Barbera, G.; Chiozzi, R.Z.; Digiacomo, L.; Peruzzi, G.; Caracciolo, G.; Laganà, A. Influence of dynamic flow environment on nanoparticle-protein corona: From protein patterns to uptake in cancer cells. Colloids Surf. B Biointerfaces 2017, 153, 263–271. [Google Scholar] [CrossRef]
- Pozzi, D.; Caracciolo, G.; Digiacomo, L.; Colapicchioni, V.; Palchetti, S.; Capriotti, A.; Cavaliere, C.; Chiozzi, R.Z.; Puglisi, A.; Laganà, A. The biomolecular corona of nanoparticles in circulating biological media. Nanoscale 2015, 7, 13958–13966. [Google Scholar] [CrossRef] [PubMed]
- García-Álvarez, R.; Hadjidemetriou, M.; Sánchez-Iglesias, A.; Liz-Marzán, L.M.; Kostarelos, K. In vivo formation of protein corona on gold nanoparticles. The effect of their size and shape. Nanoscale 2018, 10, 1256–1264. [Google Scholar] [CrossRef]
- Tian, W.-C.; Finehout, E. Current and Future Trends in Microfluidics within Biotechnology Research. In Microfluidics for Biological Applications; Springer: Berlin/Heidelberg, Germany, 2008; pp. 385–411. [Google Scholar]
- Scheler, O.; Postek, W.; Garstecki, P. Recent developments of microfluidics as a tool for biotechnology and microbiology. Curr. Opin. Biotechnol. 2019, 55, 60–67. [Google Scholar] [CrossRef]
- Ahmed, I.; Iqbal, H.M.; Akram, Z. Microfluidics engineering: Recent trends, valorization, and applications. Arab. J. Sci. Eng. 2018, 43, 23–32. [Google Scholar] [CrossRef]
- Wu, H.; Zhu, J.; Huang, Y.; Wu, D.; Sun, J. Microfluidic-based single-cell study: Current status and future perspective. Molecules 2018, 23, 2347. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Wright, M.A.; Saar, K.L.; Challa, P.; Morgunov, A.S.; Peter, Q.A.E.; Devenish, S.; Dobson, C.M.; Knowles, T.P.J. Machine learning aided top-down proteomics on a microfluidic platform. bioRxiv 2020. [Google Scholar] [CrossRef]
- Riordon, J.; Sovilj, D.; Sanner, S.; Sinton, D.; Young, E.W.K. Deep Learning with Microfluidics for Biotechnology. Trends Biotechnol. 2019, 37, 310–324. [Google Scholar] [CrossRef]
- Sarkar, S.; Kang, W.; Jiang, S.; Li, K.; Ray, S.; Luther, E.; Ivanov, A.R.; Fu, Y.; Konry, T. Machine learning-aided quantification of antibody-based cancer immunotherapy by natural killer cells in microfluidic droplets. Lab Chip 2020, 20, 2317–2327. [Google Scholar] [CrossRef] [PubMed]
- Tsur, E.E. Computer-Aided Design of Microfluidic Circuits. Annu. Rev. Biomed. Eng. 2020, 22, 285–307. [Google Scholar] [CrossRef]
- Lamanna, J.; Scott, E.Y.; Edwards, H.S.; Chamberlain, M.D.; Dryden, M.D.M. Digital microfluidic isolation of single cells for -Omics. Nat. Commun. 2020, 11, 5632. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vitorino, R.; Guedes, S.; da Costa, J.P.; Kašička, V. Microfluidics for Peptidomics, Proteomics, and Cell Analysis. Nanomaterials 2021, 11, 1118. https://doi.org/10.3390/nano11051118
Vitorino R, Guedes S, da Costa JP, Kašička V. Microfluidics for Peptidomics, Proteomics, and Cell Analysis. Nanomaterials. 2021; 11(5):1118. https://doi.org/10.3390/nano11051118
Chicago/Turabian StyleVitorino, Rui, Sofia Guedes, João Pinto da Costa, and Václav Kašička. 2021. "Microfluidics for Peptidomics, Proteomics, and Cell Analysis" Nanomaterials 11, no. 5: 1118. https://doi.org/10.3390/nano11051118
APA StyleVitorino, R., Guedes, S., da Costa, J. P., & Kašička, V. (2021). Microfluidics for Peptidomics, Proteomics, and Cell Analysis. Nanomaterials, 11(5), 1118. https://doi.org/10.3390/nano11051118









