On Facing the SARS-CoV-2 (COVID-19) with Combination of Nanomaterials and Medicine: Possible Strategies and First Challenges
Abstract
1. Introduction
2. New SARS-CoV-2 (COVID-19)
2.1. Society Impact, Diagnosis, Treatment Approaches
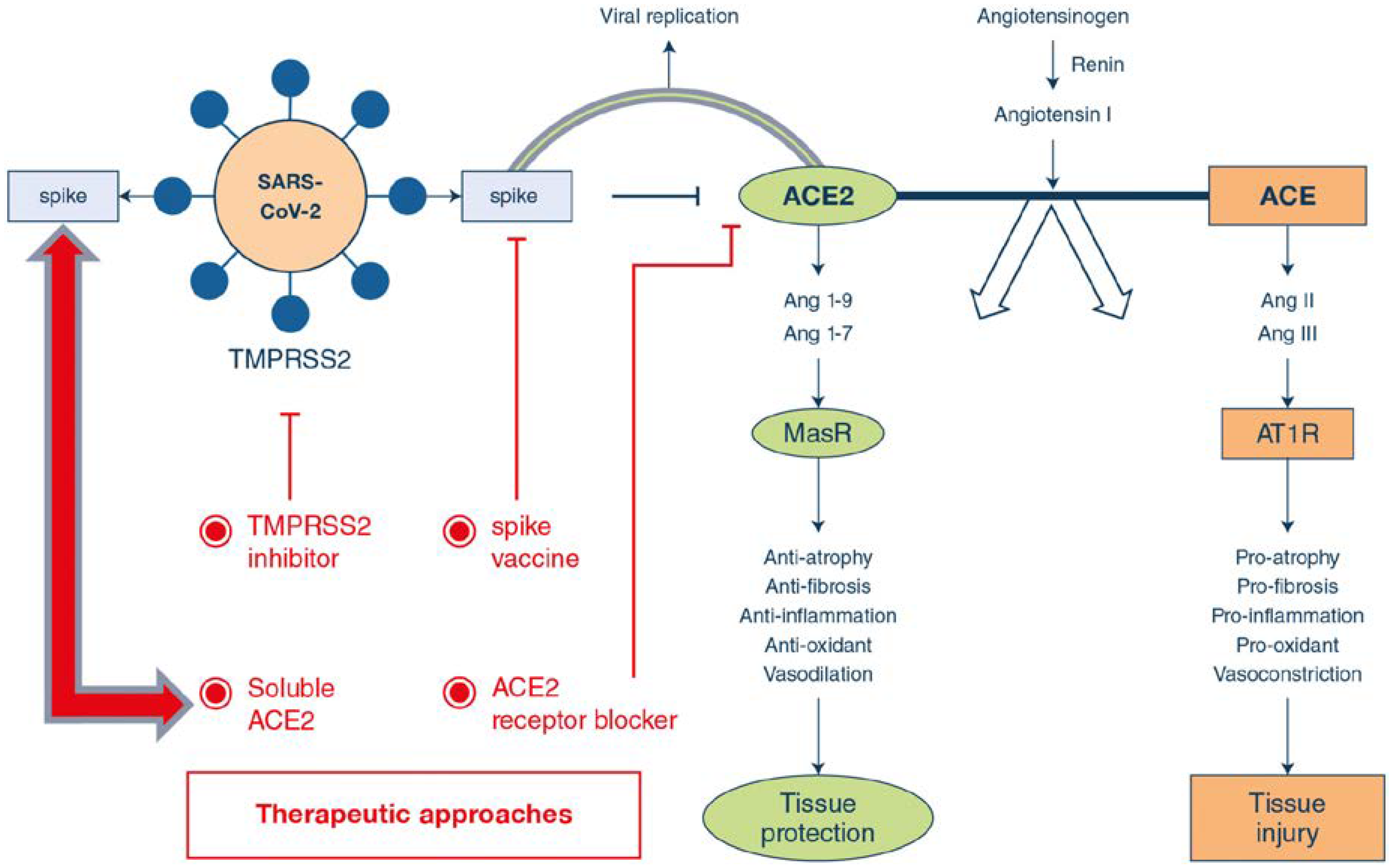
2.2. Pathophysiology and Related Facts
3. Drug Development for SARS-CoV-2
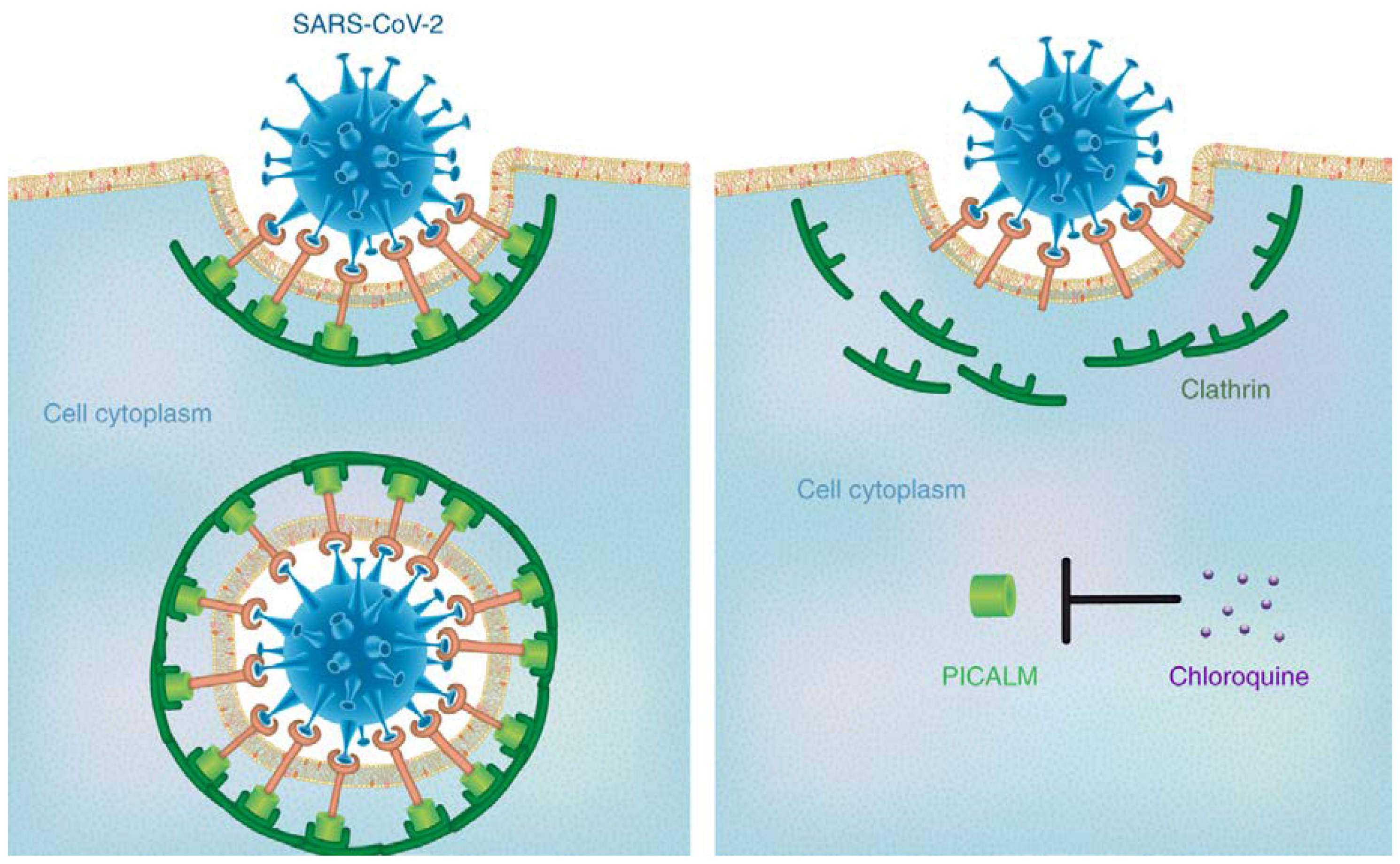
4. Old Drug Recipes for the New Target
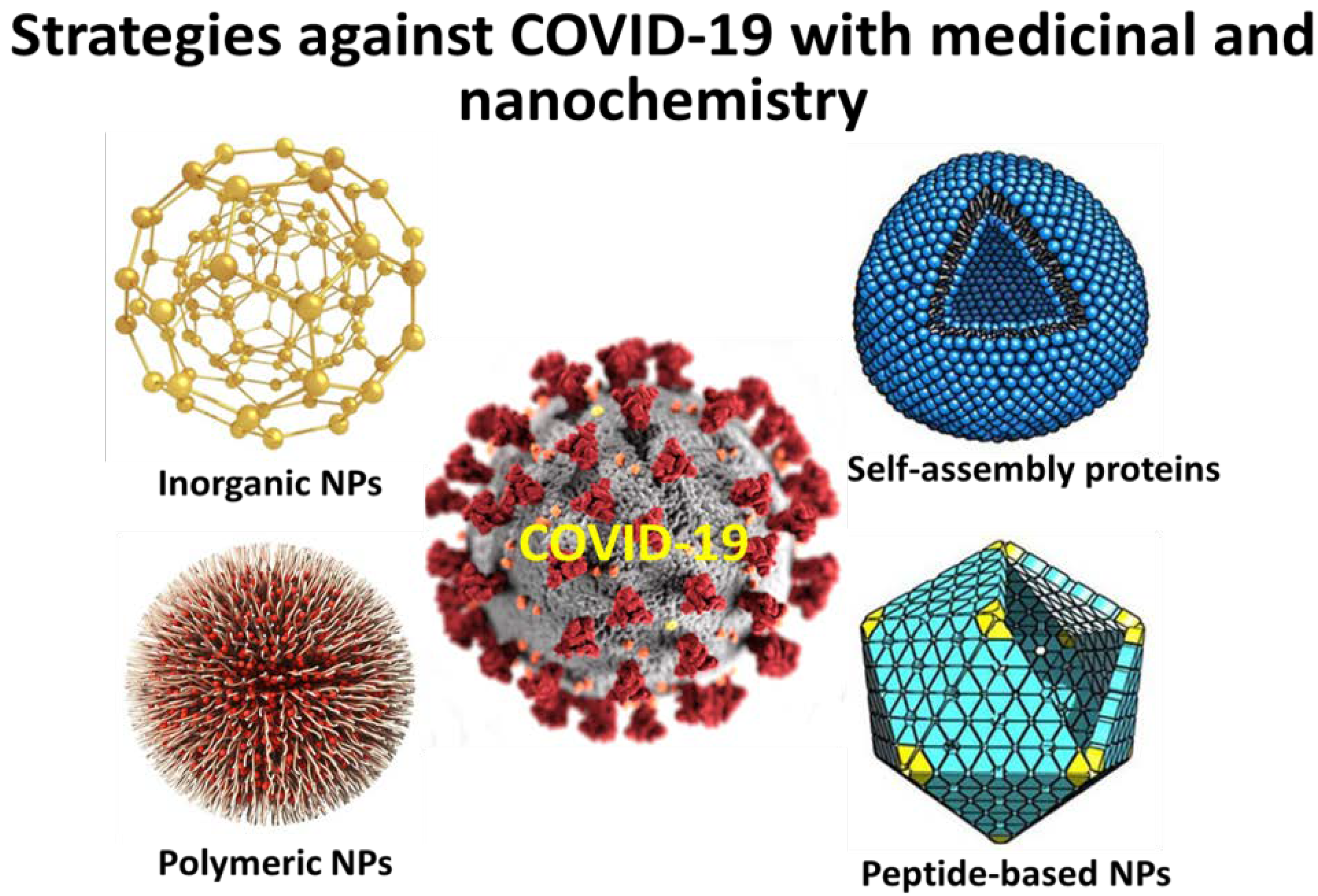
5. Perspectives in Nanoscience against Respiratory Viruses
5.1. Nanoscience to Face Various Viruses
5.2. Specific Examples of Nanoparticles against Viruses
6. Nanosensors
7. Safety and Limitations
8. Unsolved Concerns and Perspectives
9. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Casadevall, A.; Pirofski, L.-A. The convalescent sera option for containing COVID-19. J. Clin. Investig. 2020, 130, 1545–1548. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Hu, B.; Hu, C.; Zhu, F.; Liu, X.; Zhang, J.; Wang, B.; Xiang, H.; Cheng, Z.; Xiong, Y.; et al. Clinical characteristics of 138 hospitalized patients with 2019 novel Coronavirus–infected pneumonia in Wuhan, China. JAMA 2020, 323, 1061–1069. [Google Scholar] [CrossRef] [PubMed]
- Hui, D.S.I.; Azhar, E.I.; Madani, T.A.; Ntoumi, F.; Kock, R.; Dar, O.; Ippolito, G.; McHugh, T.D.; Memish, Z.A.; Drosten, C.; et al. The continuing 2019-nCoV epidemic threat of novel coronaviruses to global health —The latest 2019 novel coronavirus outbreak in Wuhan, China. Int. J. Infect. Dis. 2020, 91, 264–266. [Google Scholar] [CrossRef] [PubMed]
- Paules, C.I.; Marston, H.D.; Fauci, A.S. Coronavirus infections—More than just the common cold. JAMA 2020, 323, 707–708. [Google Scholar] [CrossRef]
- COVID-19 Coronavirus Pandemic. Available online: https://www.worldometers.info/coronavirus/ (accessed on 28 March 2020).
- Cascella, M.; Rajnik, M.; Cuomo, A.; Dulebohn, S.C.; Di Napoli, R. Features, evaluation and treatment Coronavirus (COVID-19). In StatPearls; StatPearls Publishing: Treasure Island, FL, USA, 2020. [Google Scholar]
- Fauci, A.S.; Lane, H.C.; Redfield, R.R. Covid-19—Navigating the uncharted. N. Engl. J. Med. 2020, 382, 1268–1269. [Google Scholar] [CrossRef]
- de Wit, E.; van Doremalen, N.; Falzarano, D.; Munster, V.J. SARS and MERS: Recent insights into emerging coronaviruses. Nat. Rev. Microbiol. 2016, 14, 523–534. [Google Scholar] [CrossRef]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef]
- Timmer, J. COVID-19: The Biology of an Effective Therapy. Available online: https://arstechnica.com/science/2020/03/covid-19-the-biology-of-an-effective-therapy/ (accessed on 26 March 2020).
- Perlman, S.; Netland, J. Coronaviruses post-SARS: Update on replication and pathogenesis. Nat. Rev. Microbiol. 2009, 7, 439–450. [Google Scholar] [CrossRef]
- Zhou, P.; Yang, X.-L.; Wang, X.-G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.-R.; Zhu, Y.; Li, B.; Huang, C.-L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef]
- Li, W.; Shi, Z.; Yu, M.; Ren, W.; Smith, C.; Epstein, J.H.; Wang, H.; Crameri, G.; Hu, Z.; Zhang, H.; et al. Bats are natural reservoirs of SARS-like Coronaviruses. Science 2005, 310, 676–679. [Google Scholar] [CrossRef]
- Yang, L.; Wu, Z.; Ren, X.; Yang, F.; He, G.; Zhang, J.; Dong, J.; Sun, L.; Zhu, Y.; Du, J.; et al. Novel SARS-like Betacoronaviruses in Bats, China, 2011. Emerg. Infect. Dis. 2013, 19, 989–991. [Google Scholar] [CrossRef] [PubMed]
- Hu, B.; Zeng, L.-P.; Yang, X.-L.; Ge, X.-Y.; Zhang, W.; Li, B.; Xie, J.-Z.; Shen, X.-R.; Zhang, Y.-Z.; Wang, N.; et al. Discovery of a rich gene pool of bat SARS-related coronaviruses provides new insights into the origin of SARS coronavirus. PLoS Pathog. 2017, 13, e1006698. [Google Scholar] [CrossRef] [PubMed]
- Min, W.F.; Huizhi, G.; Jessica, Y.W.; Jingyi, X.; Eunice, Y.C.S.; Sukhyun, R.; Benjamin, J.C. Nonpharmaceutical measures for pandemic influenza in nonhealthcare settings—Social distancing measures. Emerg. Infect. Dis. J. 2020, 26, 976–984. [Google Scholar]
- Kampf, G.; Todt, D.; Pfaender, S.; Steinmann, E. Persistence of coronaviruses on inanimate surfaces and their inactivation with biocidal agents. J. Hosp. Infect. 2020, 104, 246–251. [Google Scholar] [CrossRef]
- Li, Q.; Guan, X.; Wu, P.; Wang, X.; Zhou, L.; Tong, Y.; Ren, R.; Leung, K.S.M.; Lau, E.H.Y.; Wong, J.Y.; et al. Early Transmission Dynamics in Wuhan, China, of Novel Coronavirus–Infected Pneumonia. N. Engl. J. Med. 2020, 382, 1199–1207. [Google Scholar] [CrossRef]
- CDC, 2019 Novel Coronavirus, Wuhan, China. Available online: https://www.cdc.gov/coronavirus/2019-nCoV/summary.html (accessed on 1 February 2020).
- WHO. Novel Coronavirus–China. 2020. Available online: https://www.who.int/csr/don/12-january-2020-novel-coronavirus-china/en/ (accessed on 1 February 2020).
- Liu, T.; Hu, J.; Kang, M.; Lin, L.; Zhong, H.; Xiao, J.; He, G.; Song, T.; Huang, Q.; Rong, Z.; et al. Transmission dynamics of 2019 novel coronavirus (2019-nCoV). bioRxiv 2020, in press. [Google Scholar] [CrossRef]
- Zhu, Y.; Chen, L.; Ji, H.; Xi, M.; Fang, Y.; Li, Y. The Risk and Prevention of Novel Coronavirus Pneumonia Infections Among Inpatients in Psychiatric Hospitals. Neurosci. Bull. 2020, 36, 299–302. [Google Scholar] [CrossRef]
- Adhikari, S.P.; Meng, S.; Wu, Y.-J.; Mao, Y.-P.; Ye, R.-X.; Wang, Q.-Z.; Sun, C.; Sylvia, S.; Rozelle, S.; Raat, H.; et al. Epidemiology, causes, clinical manifestation and diagnosis, prevention and control of coronavirus disease (COVID-19) during the early outbreak period: A scoping review. Infect. Dis. Poverty 2020, 9, 29. [Google Scholar] [CrossRef]
- WHO. International Clinical Trials Registry Platform. Available online: https://apps.who.int/trialsearch/default.aspx (accessed on 23 February 2020).
- Xu, M.; Zhang, Y. Investigation on the psychological status of the first batch of clinical first-line support nurses to fight against pneumonia caused by novel coronavirus. Chin. Nurs. Res. 2020, 34, 1–3. [Google Scholar]
- Li, H.; Wang, Y.; Xu, J.; Cao, B. Potential antiviral therapeutics for 2019 Novel Coronavirus. Zhonghua Jie He He Hu Xi Za Zhi = Zhonghua Jiehe He Huxi Zazhi = Chinese J. Tuberc. Respir. Dis. 2020, 43, 2. [Google Scholar]
- Holshue, M.L.; DeBolt, C.; Lindquist, S.; Lofy, K.H.; Wiesman, J.; Bruce, H.; Spitters, C.; Ericson, K.; Wilkerson, S.; Tural, A.; et al. First Case of 2019 Novel Coronavirus in the United States. N. Engl. J. Med. 2020, 382, 929–936. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.-H.; Cheng, Y. Increasing Host Cellular Receptor—Angiotensin-Converting Enzyme 2 (ACE2) Expression by Coronavirus may Facilitate 2019-nCoV Infection. bioRxiv 2020, in press. [Google Scholar] [CrossRef]
- Li, R.; Qiao, S.; Zhang, G. Analysis of angiotensin-converting enzyme 2 (ACE2) from different species sheds some light on cross-species receptor usage of a novel coronavirus 2019-nCoV. J. Infect. 2020, 80, 469–496. [Google Scholar] [CrossRef] [PubMed]
- Zeng, Q.; Langereis, M.A.; van Vliet, A.L.W.; Huizinga, E.G.; de Groot, R.J. Structure of coronavirus hemagglutinin-esterase offers insight into corona and influenza virus evolution. Proc. Natl. Acad. Sci. USA 2008, 105, 9065–9069. [Google Scholar] [CrossRef] [PubMed]
- Bakkers, M.J.G.; Lang, Y.; Feitsma, L.J.; Hulswit, R.J.G.; de Poot, S.A.H.; van Vliet, A.L.W.; Margine, I.; de Groot-Mijnes, J.D.F.; van Kuppeveld, F.J.M.; Langereis, M.A.; et al. Betacoronavirus Adaptation to Humans Involved Progressive Loss of Hemagglutinin-Esterase Lectin Activity. Cell Host Microbe 2017, 21, 356–366. [Google Scholar] [CrossRef] [PubMed]
- Simmons, G.; Bertram, S.; Glowacka, I.; Steffen, I.; Chaipan, C.; Agudelo, J.; Lu, K.; Rennekamp, A.J.; Hofmann, H.; Bates, P.; et al. Different host cell proteases activate the SARS-coronavirus spike-protein for cell–cell and virus–cell fusion. Virology 2011, 413, 265–274. [Google Scholar] [CrossRef]
- Wang, M.; Cao, R.; Zhang, L.; Yang, X.; Liu, J.; Xu, M.; Shi, Z.; Hu, Z.; Zhong, W.; Xiao, G. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell Res. 2020, 30, 269–271. [Google Scholar] [CrossRef]
- Impellizzeri, D.; Bruschetta, G.; Esposito, E.; Cuzzocrea, S. Emerging drugs for acute lung injury. Expert Opin. Emerg. Drugs 2015, 20, 75–89. [Google Scholar] [CrossRef]
- Sweeney, R.M.; Griffiths, M.; McAuley, D. Treatment of Acute Lung Injury: Current and Emerging Pharmacological Therapies. Semin. Respir. Crit. Care Med. 2013, 34, 487–498. [Google Scholar] [CrossRef]
- Kuba, K.; Imai, Y.; Rao, S.; Gao, H.; Guo, F.; Guan, B.; Huan, Y.; Yang, P.; Zhang, Y.; Deng, W.; et al. A crucial role of angiotensin converting enzyme 2 (ACE2) in SARS coronavirus–induced lung injury. Nat. Med. 2005, 11, 875–879. [Google Scholar] [CrossRef]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Krüger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.-H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280. [Google Scholar] [CrossRef]
- Zhao, Y.; Zhao, Z.; Wang, Y.; Zhou, Y.; Ma, Y.; Zuo, W. Single-cell RNA expression profiling of ACE2, the putative receptor of Wuhan 2019-nCoV. bioRxiv 2020, in press. [Google Scholar] [CrossRef]
- Zou, Z.; Yan, Y.; Shu, Y.; Gao, R.; Sun, Y.; Li, X.; Ju, X.; Liang, Z.; Liu, Q.; Zhao, Y.; et al. Angiotensin-converting enzyme 2 protects from lethal avian influenza A H5N1 infections. Nat. Commun. 2014, 5, 3594. [Google Scholar] [CrossRef] [PubMed]
- Fang, L.; Karakiulakis, G.; Roth, M. Are patients with hypertension and diabetes mellitus at increased risk for COVID-19 infection? Lancet Respir. Med. 2020, 8, e21. [Google Scholar] [CrossRef]
- Li, X.; Geng, M.; Peng, Y.; Meng, L.; Lu, S. Molecular immune pathogenesis and diagnosis of COVID-19. J. Pharm. Anal. 2020, in press. [Google Scholar] [CrossRef]
- Ramos-Casals, M.; Brito-Zerón, P.; López-Guillermo, A.; Khamashta, M.A.; Bosch, X. Adult haemophagocytic syndrome. Lancet 2014, 383, 1503–1516. [Google Scholar] [CrossRef]
- Seguin, A.; Galicier, L.; Boutboul, D.; Lemiale, V.; Azoulay, E. Pulmonary involvement in patients with hemophagocytic Lymphohistiocytosis. Chest 2016, 149, 1294–1301. [Google Scholar] [CrossRef]
- Wan, S.; Yi, Q.; Fan, S.; Lv, J.; Zhang, X.; Guo, L.; Lang, C.; Xiao, Q.; Xiao, K.; Yi, Z.; et al. Characteristics of lymphocyte subsets and cytokines in peripheral blood of 123 hospitalized patients with 2019 novel coronavirus pneumonia (NCP). medRxiv 2020, in press. [Google Scholar] [CrossRef]
- Mehta, P.; McAuley, D.F.; Brown, M.; Sanchez, E.; Tattersall, R.S.; Manson, J.J. COVID-19: Consider cytokine storm syndromes and immunosuppression. Lancet 2020, 395, 1033–1034. [Google Scholar] [CrossRef]
- Rut, W.; Groborz, K.; Zhang, L.; Sun, X.; Zmudzinski, M.; Hilgenfeld, R.; Drag, M. Substrate specificity profiling of SARS-CoV-2 MPRO; protease provides basis for anti-COVID-19 drug design. bioRxiv 2020, in press. [Google Scholar] [CrossRef]
- Ton, A.-T.; Gentile, F.; Hsing, M.; Ban, F.; Cherkasov, A. Rapid Identification of potential inhibitors of SARS-CoV-2 main protease by deep docking of 1.3 billion compounds. Mol. Inform. 2020. [Google Scholar] [CrossRef] [PubMed]
- Bzówka, M.; Mitusińska, K.; Raczyńska, A.; Samol, A.; Tuszyński, J.; Góra, A. Molecular dynamics simulations indicate the COVID-19 Mpro is not a viable target for small-molecule Inhibitors design. bioRxiv 2020, in press. [Google Scholar] [CrossRef]
- Park, T.; Lee, S.-Y.; Kim, S.; Kim, M.J.; Kim, H.G.; Jun, S.; Kim, S.I.; Kim, B.T.; Park, E.C.; Park, D. Spike protein binding prediction with neutralizing antibodies of SARS-CoV-2. bioRxiv 2020, in press. [Google Scholar] [CrossRef]
- ul Qamar, M.T.; Rehman, A.; Ashfaq, U.A.; Awan, M.Q.; Fatima, I.; Shahid, F.; Chen, L.-L. Designing of a next generation multiepitope based vaccine (MEV) against SARS-COV-2: Immunoinformatics and in silico approaches. bioRxiv 2020, in press. [Google Scholar] [CrossRef]
- Ko, M.; Chang, S.Y.; Byun, S.Y.; Choi, I.; d’Orengiani, A.-L.P.H.D.A.; Shum, D.; Min, J.-Y.; Windisch, M.P. Screening of FDA-approved drugs using a MERS-CoV clinical isolate from South Korea identifies potential therapeutic options for COVID-19. bioRxiv 2020, in press. [Google Scholar] [CrossRef]
- Zhang, D.-H.; Wu, K.-L.; Zhang, X.; Deng, S.-Q.; Peng, B. In silico screening of Chinese herbal medicines with the potential to directly inhibit 2019 novel coronavirus. J. Integr. Med. 2020, 18, 152–158. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.; Wang, X.; Yang, Y.; Zhang, X.; Mu, K.; Shi, Y.; Peng, C.; Xu, Z.; Zhu, W. D3Similarity: A ligand-based approach for predicting drug targets and for virtual screening of active compounds against COVID-19. ChemRxiv 2020, in press. [Google Scholar] [CrossRef]
- Zhang, H.; Penninger, J.M.; Li, Y.; Zhong, N.; Slutsky, A.S. Angiotensin-converting enzyme 2 (ACE2) as a SARS-CoV-2 receptor: Molecular mechanisms and potential therapeutic target. Intensive Care Med. 2020, 46, 586–590. [Google Scholar] [CrossRef]
- Strittmatter, S.M. Overcoming drug development bottlenecks with repurposing: Old drugs learn new tricks. Nat. Med. 2014, 20, 590–591. [Google Scholar] [CrossRef]
- Guo, D. Old weapon for new enemy: Drug repurposing for treatment of newly emerging viral diseases. Virol. Sin. 2020, 1–3. [Google Scholar] [CrossRef]
- Mohammed, A.; Tam, D.N.H.; Vu, T.L.H.; Tieu, T.M.; Elfaituri, M.K.; Trinh, N.N.; Sagheir, E.A.; Tran, L.; Loc, T.T.H.; Low, S.K.; et al. Morinda morindoides: A systematic review of its therapeutic activities. S. Afr. J. Bot. 2020, 131, 93–103. [Google Scholar] [CrossRef]
- Varisli, L.; Cen, O.; Vlahopoulos, S. Dissecting pharmacological effects of chloroquine in cancer treatment: Interference with inflammatory signaling pathways. Immunology 2020, 159, 257–278. [Google Scholar] [CrossRef] [PubMed]
- Schrezenmeier, E.; Dörner, T. Mechanisms of action of hydroxychloroquine and chloroquine: Implications for rheumatology. Nat. Rev. Rheumatol. 2020, 16, 155–166. [Google Scholar] [CrossRef] [PubMed]
- Devaux, C.A.; Rolain, J.-M.; Colson, P.; Raoult, D. New insights on the antiviral effects of chloroquine against coronavirus: What to expect for COVID-19? Int. J. Antimicrob. Agents 2020, in press. [Google Scholar] [CrossRef]
- Savarino, A.; Boelaert, J.R.; Cassone, A.; Majori, G.; Cauda, R. Effects of chloroquine on viral infections: An old drug against today’s diseases. Lancet Infect. Dis. 2003, 3, 722–727. [Google Scholar] [CrossRef]
- Hu, T.Y.; Frieman, M.; Wolfram, J. Insights from nanomedicine into chloroquine efficacy against COVID-19. Nat. Nanotechnol. 2020, 15, 247–249. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Tian, Z.; Yang, X. Breakthrough: Chloroquine phosphate has shown apparent efficacy in treatment of COVID-19 associated pneumonia in clinical studies. Biosci. Trends 2020, 14, 72–73. [Google Scholar] [CrossRef] [PubMed]
- Zhou, D.; Dai, S.-M.; Tong, Q. COVID-19: A recommendation to examine the effect of hydroxychloroquine in preventing infection and progression. J. Antimicrob. Chemother. 2020, in press. [Google Scholar] [CrossRef]
- Gautret, P.; Lagier, J.-C.; Parola, P.; Hoang, V.T.; Meddeb, L.; Mailhe, M.; Doudier, B.; Courjon, J.; Giordanengo, V.; Vieira, V.E.; et al. Hydroxychloroquine and azithromycin as a treatment of COVID-19: Results of an open-label non-randomized clinical trial. Int. J. Antimicrob. Agents 2020, in press. [Google Scholar] [CrossRef]
- Xu, J.; Shi, P.-Y.; Li, H.; Zhou, J. Broad spectrum antiviral agent Niclosamide and its therapeutic potential. ACS Infect. Dis. 2020, in press. [Google Scholar] [CrossRef]
- Fan, H.-H.; Wang, L.-Q.; Liu, W.-L.; An, X.-P.; Liu, Z.-D.; He, X.-Q.; Song, L.-H.; Tong, Y.-G. Repurposing of clinically approved drugs for treatment of coronavirus disease 2019 in a 2019-novel coronavirus (2019-nCoV) related coronavirus model. Chin. Med. J. 2020, in press. [Google Scholar] [CrossRef] [PubMed]
- Cao, B.; Wang, Y.; Wen, D.; Liu, W.; Wang, J.; Fan, G.; Ruan, L.; Song, B.; Cai, Y.; Wei, M.; et al. A trial of Lopinavir–Ritonavir in adults hospitalized with severe Covid-19. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef] [PubMed]
- Russell, C.D.; Millar, J.E.; Baillie, J.K. Clinical evidence does not support corticosteroid treatment for 2019-nCoV lung injury. Lancet 2020, 395, 473–475. [Google Scholar] [CrossRef]
- Favalli, E.G.; Ingegnoli, F.; De Lucia, O.; Cincinelli, G.; Cimaz, R.; Caporali, R. COVID-19 infection and rheumatoid arthritis: Faraway, so close! Autoimmun. Rev. 2020, 19, 102523. [Google Scholar] [CrossRef]
- Liu, X.; Li, Z.; Liu, S.; Chen, Z.; Zhao, Z.; Huang, Y.-y.; Zhang, Q.; Wang, J.; Shi, Y.; Xu, Y.; et al. Therapeutic effects of dipyridamole on COVID-19 patients with coagulation dysfunction. medRxiv 2020, in press. [Google Scholar] [CrossRef]
- Sang, P.; Tian, S.; Meng, Z.; Yang, L. Insight derived from molecular docking and molecular dynamics simulations into the binding interactions between HIV-1 protease inhibitors and SARS-CoV-2 3CLPRO. ChemRxiv 2020, in press. [Google Scholar] [CrossRef]
- Chen, Y.W.; Yiu, C.-P.B.; Wong, K.-Y. Prediction of the SARS-CoV-2 (2019-nCoV) 3C-like protease (3CL pro) structure: Virtual screening reveals velpatasvir, ledipasvir, and other drug repurposing candidates. F1000Research 2020, 9, 129. [Google Scholar] [CrossRef]
- Shaghaghi, N. Molecular docking study of novel COVID-19 protease with low risk terpenoides compounds of plants. ChemRxiv 2020. [Google Scholar] [CrossRef]
- Elfiky, A.A. Anti-HCV, nucleotide inhibitors, repurposing against COVID-19. Life Sci. 2020, 248, 117477. [Google Scholar] [CrossRef]
- Baron, S.A.; Devaux, C.; Colson, P.; Raoult, D.; Rolain, J.-M. Teicoplanin: An alternative drug for the treatment of coronavirus COVID-19? Int. J. Antimicrob. Agents 2020, 55, 105944. [Google Scholar] [CrossRef]
- Al-Halifa, S.; Gauthier, L.; Arpin, D.; Bourgault, S.; Archambault, D. Nanoparticle-based vaccines against respiratory viruses. Front. Immunol. 2019, 10, 22. [Google Scholar] [CrossRef] [PubMed]
- Nikazar, S.; Sivasankarapillai, V.S.; Rahdar, A.; Gasmi, S.; Anumol, P.S.; Shanavas, M.S. Revisiting the cytotoxicity of quantum dots: An in-depth overview. Biophys. Rev. 2020, in press. [Google Scholar] [CrossRef] [PubMed]
- Jung, J.H.; Park, B.H.; Oh, S.J.; Choi, G.; Seo, T.S. Integration of reverse transcriptase loop-mediated isothermal amplification with an immunochromatographic strip on a centrifugal microdevice for influenza A virus identification. Lab Chip 2015, 15, 718–725. [Google Scholar] [CrossRef] [PubMed]
- Barik, S. New treatments for influenza. BMC Med. 2012, 10, 104. [Google Scholar] [CrossRef]
- Shafagati, N.; Patanarut, A.; Luchini, A.; Lundberg, L.; Bailey, C.; Petricoin, E., III; Liotta, L.; Narayanan, A.; Lepene, B.; Kehn Hall, K. The use of Nanotrap particles for biodefense and emerging infectious disease diagnostics. Pathog. Dis. 2014, 71, 164–176. [Google Scholar] [CrossRef]
- Hendricks, G.L.; Weirich, K.L.; Viswanathan, K.; Li, J.; Shriver, Z.H.; Ashour, J.; Ploegh, H.L.; Kurt-Jones, E.A.; Fygenson, D.K.; Finberg, R.W.; et al. Sialylneolacto-N-tetraose c (LSTc)-bearing liposomal decoys capture influenza A virus. J. Biol. Chem. 2013, 288, 8061–8073. [Google Scholar] [CrossRef]
- Wagner, R.; Matrosovich, M.; Klenk, H.-D. Functional balance between haemagglutinin and neuraminidase in influenza virus infections. Rev. Med. Virol. 2002, 12, 159–166. [Google Scholar] [CrossRef]
- Moscona, A. Global Transmission of Oseltamivir-Resistant Influenza. N. Engl. J. Med. 2009, 360, 953–956. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Lin, Z.; Zhao, M.; Xu, T.; Wang, C.; Hua, L.; Wang, H.; Xia, H.; Zhu, B. Silver Nanoparticle Based Codelivery of Oseltamivir to Inhibit the Activity of the H1N1 Influenza Virus through ROS-Mediated Signaling Pathways. ACS Appl. Mater. Interfaces 2016, 8, 24385–24393. [Google Scholar] [CrossRef]
- Levina, A.S.; Repkova, M.N.; Mazurkova, N.A.; Zarytova, V.F. Nanoparticle-Mediated Nonviral DNA Delivery for Effective Inhibition of Influenza a Viruses in Cells. IEEE Trans. Nanotechnol. 2016, 15, 248–254. [Google Scholar] [CrossRef]
- Saleh, T.; Bolhassani, A.; Shojaosadati, S.A.; Aghasadeghi, M.R. MPG-based nanoparticle: An efficient delivery system for enhancing the potency of DNA vaccine expressing HPV16E7. Vaccine 2015, 33, 3164–3170. [Google Scholar] [CrossRef] [PubMed]
- Chapoy Villanueva, H.; Martinez-Carlin, I.; Lopez-Berestein, G.; Chavez-Reyes, A. Therapeutic silencing of HPV 16 E7 by systemic administration of siRNA-neutral DOPC nanoliposome in a murine cervical cancer model with obesity. J. BU ON 2016, 20, 1471–1479. [Google Scholar]
- Burnett, J.C.; Rossi, J.J.; Tiemann, K. Current progress of siRNA/shRNA therapeutics in clinical trials. Biotechnol. J. 2011, 6, 1130–1146. [Google Scholar] [CrossRef] [PubMed]
- Zamora, M.R.; Budev, M.; Rolfe, M.; Gottlieb, J.; Humar, A.; DeVincenzo, J.; Vaishnaw, A.; Cehelsky, J.; Albert, G.; Nochur, S.; et al. RNA Interference Therapy in Lung Transplant Patients Infected with Respiratory Syncytial Virus. Am. J. Respir. Crit. Care Med. 2011, 183, 531–538. [Google Scholar] [CrossRef] [PubMed]
- Gaikwad, S.; Ingle, A.; Gade, A.; Rai, M.; Falanga, A.; Incoronato, N.; Russo, L.; Galdiero, S.; Galdiero, M. Antiviral activity of mycosynthesized silver nanoparticles against herpes simplex virus and human parainfluenza virus type 3. Int. J. Nanomedicine 2013, 8, 4303–4314. [Google Scholar] [PubMed]
- Lee, Y.-T.; Ko, E.-J.; Hwang, H.S.; Lee, J.S.; Kim, K.-H.; Kwon, Y.-M.; Kang, S.-M. Respiratory syncytial virus-like nanoparticle vaccination induces long-term protection without pulmonary disease by modulating cytokines and T-cells partially through alveolar macrophages. Int. J. Nanomedicine 2015, 10, 4491–4505. [Google Scholar]
- Schmitt, V.; Kesch, C.; Jackson, J.K.; Bidnur, S.; Beraldi, E.; Yago, V.; Bowden, M.; Gleave, M.E. Design and characterization of injectable poly(lactic-co-glycolic acid) pastes for sustained and local drug release. Pharm. Res. 2020, 37, 36. [Google Scholar] [CrossRef]
- Carter, D.C.; Wright, B.; Gray, W.; Rose, J.P.; Wilson, E. A unique protein self-assembling nanoparticle with significant advantages in vaccine development and production. J. Nanomater. 2020, 2020, 4297937. [Google Scholar] [CrossRef]
- Pillai, A.M.; Sivasankarapillai, V.S.; Rahdar, A.; Joseph, J.; Sadeghfar, F.; Anuf, A.R.; Rajesh, K.; Kyzas, G.Z. Green synthesis and characterization of zinc oxide nanoparticles with antibacterial and antifungal activity. J. Mol. Struct. 2020, 1211, 128107. [Google Scholar] [CrossRef]
- Du, Q.; Wang, S.; Wei, D.; Sirois, S.; Chou, K.-C. Molecular modeling and chemical modification for finding peptide inhibitor against severe acute respiratory syndrome coronavirus main proteinase. Anal. Biochem. 2005, 337, 262–270. [Google Scholar] [CrossRef]
- McReynolds, S.; Jiang, S.; Guo, Y.; Celigoy, J.; Schar, C.; Rong, L.; Caffrey, M. Characterization of the Prefusion and Transition States of Severe Acute Respiratory Syndrome Coronavirus S2-HR2. Biochemistry 2008, 47, 6802–6808. [Google Scholar] [CrossRef] [PubMed]
- Pimentel, T.A.P.F.; Yan, Z.; Jeffers, S.A.; Holmes, K.V.; Hodges, R.S.; Burkhard, P. Peptide Nanoparticles as Novel Immunogens: Design and Analysis of a Prototypic Severe Acute Respiratory Syndrome Vaccine. Chem. Biol. Drug Des. 2009, 73, 53–61. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Král, P. Computational Design of ACE2-Based Peptide Inhibitors of SARS-CoV-2. ACS Nano 2020, in press. [Google Scholar] [CrossRef] [PubMed]
- Mansoor, F.; Earley, B.; Cassidy, J.P.; Markey, B.; Doherty, S.; Welsh, M.D. Comparing the immune response to a novel intranasal nanoparticle PLGA vaccine and a commercial BPI3V vaccine in dairy calves. BMC Vet. Res. 2015, 11, 220. [Google Scholar] [CrossRef]
- Dhakal, S.; Hiremath, J.; Bondra, K.; Lakshmanappa, Y.S.; Shyu, D.-L.; Ouyang, K.; Kang, K.-i.; Binjawadagi, B.; Goodman, J.; Tabynov, K.; et al. Biodegradable nanoparticle delivery of inactivated swine influenza virus vaccine provides heterologous cell-mediated immune response in pigs. J. Control. Release 2017, 247, 194–205. [Google Scholar] [CrossRef]
- Okamoto, S.; Matsuura, M.; Akagi, T.; Akashi, M.; Tanimoto, T.; Ishikawa, T.; Takahashi, M.; Yamanishi, K.; Mori, Y. Poly(γ-glutamic acid) nano-particles combined with mucosal influenza virus hemagglutinin vaccine protects against influenza virus infection in mice. Vaccine 2009, 27, 5896–5905. [Google Scholar] [CrossRef]
- Liu, Q.; Zheng, X.; Zhang, C.; Shao, X.; Zhang, X.; Zhang, Q.; Jiang, X. Conjugating influenza a (H1N1) antigen to n-trimethylaminoethylmethacrylate chitosan nanoparticles improves the immunogenicity of the antigen after nasal administration. J. Med. Virol. 2015, 87, 1807–1815. [Google Scholar] [CrossRef]
- Sawaengsak, C.; Mori, Y.; Yamanishi, K.; Mitrevej, A.; Sinchaipanid, N. Chitosan nanoparticle encapsulated hemagglutinin-split influenza virus mucosal vaccine. AAPS PharmSciTech 2014, 15, 317–325. [Google Scholar] [CrossRef]
- Dhakal, S.; Renu, S.; Ghimire, S.; Shaan Lakshmanappa, Y.; Hogshead, B.T.; Feliciano-Ruiz, N.; Lu, F.; HogenEsch, H.; Krakowka, S.; Lee, C.W.; et al. Mucosal immunity and protective efficacy of intranasal inactivated influenza vaccine is improved by chitosan nanoparticle delivery in pigs. Front. Immunol. 2018, 9, 934. [Google Scholar] [CrossRef]
- Dabaghian, M.; Latifi, A.M.; Tebianian, M.; NajmiNejad, H.; Ebrahimi, S.M. Nasal vaccination with r4M2e.HSP70c antigen encapsulated into N-trimethyl chitosan (TMC) nanoparticulate systems: Preparation and immunogenicity in a mouse model. Vaccine 2018, 36, 2886–2895. [Google Scholar] [CrossRef]
- Lynn, G.M.; Laga, R.; Darrah, P.A.; Ishizuka, A.S.; Balaci, A.J.; Dulcey, A.E.; Pechar, M.; Pola, R.; Gerner, M.Y.; Yamamoto, A.; et al. In vivo characterization of the physicochemical properties of polymer-linked TLR agonists that enhance vaccine immunogenicity. Nat. Biotechnol. 2015, 33, 1201–1210. [Google Scholar] [CrossRef] [PubMed]
- Francica, J.R.; Lynn, G.M.; Laga, R.; Joyce, M.G.; Ruckwardt, T.J.; Morabito, K.M.; Chen, M.; Chaudhuri, R.; Zhang, B.; Sastry, M.; et al. Thermoresponsive Polymer Nanoparticles Co-deliver RSV F Trimers with a TLR-7/8 Adjuvant. Bioconjugate Chem. 2016, 27, 2372–2385. [Google Scholar] [CrossRef]
- Ulery, B.D.; Phanse, Y.; Sinha, A.; Wannemuehler, M.J.; Narasimhan, B.; Bellaire, B.H. Polymer chemistry influences monocytic uptake of polyanhydride nanospheres. Pharm. Res. 2009, 26, 683–690. [Google Scholar] [CrossRef] [PubMed]
- McGill, J.L.; Kelly, S.M.; Kumar, P.; Speckhart, S.; Haughney, S.L.; Henningson, J.; Narasimhan, B.; Sacco, R.E. Efficacy of mucosal polyanhydride nanovaccine against respiratory syncytial virus infection in the neonatal calf. Sci. Rep. 2018, 8, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Roux, X.; Dubuquoy, C.; Durand, G.; Tran-Tolla, T.-L.; Castagné, N.; Bernard, J.; Petit-Camurdan, A.; Eléouët, J.-F.; Riffault, S. Sub-nucleocapsid nanoparticles: A nasal vaccine against respiratory syncytial virus. PLoS ONE 2008, 3, e1766. [Google Scholar] [CrossRef] [PubMed]
- Hervé, P.-L.; Deloizy, C.; Descamps, D.; Rameix-Welti, M.-A.; Fix, J.; McLellan, J.S.; Eléouët, J.-F.; Riffault, S. RSV N-nanorings fused to palivizumab-targeted neutralizing epitope as a nanoparticle RSV vaccine. Nanomedicine Nanotechnol. Biol. Med. 2017, 13, 411–420. [Google Scholar] [CrossRef]
- Hervé, P.-L.; Raliou, M.; Bourdieu, C.; Dubuquoy, C.; Petit-Camurdan, A.; Bertho, N.; Eléouët, J.-F.; Chevalier, C.; Riffault, S. A novel subnucleocapsid nanoplatform for mucosal vaccination against influenza virus that targets the ectodomain of matrix protein 2. J. Virol. 2014, 88, 325–338. [Google Scholar] [CrossRef]
- Qi, M.; Zhang, X.-E.; Sun, X.; Zhang, X.; Yao, Y.; Liu, S.; Chen, Z.; Li, W.; Zhang, Z.; Chen, J.; et al. Intranasal nanovaccine confers homo- and hetero-subtypic influenza protection. Small 2018, 14, 1703207. [Google Scholar] [CrossRef]
- Si, Y.; Wen, Y.; Kelly, S.H.; Chong, A.S.; Collier, J.H. Intranasal delivery of adjuvant-free peptide nanofibers elicits resident CD8+ T cell responses. J. Control. Release 2018, 282, 120–130. [Google Scholar] [CrossRef]
- Hsueh, P.-R.; Ko, W.-C.; Wu, J.-J.; Lu, J.-J.; Wang, F.-D.; Wu, H.-Y.; Wu, T.-L.; Teng, L.-J. Consensus Statement on the Adherence to Clinical and Laboratory Standards Institute (CLSI) Antimicrobial Susceptibility Testing Guidelines (CLSI-2010 and CLSI-2010-update) for Enterobacteriaceae in Clinical Microbiology Laboratories in Taiwan. J. Microbiol. Immunol. Infect. 2010, 43, 452–455. [Google Scholar] [CrossRef]
- Quan, F.-S.; Huang, C.; Compans, R.W.; Kang, S.-M. Virus-like particle vaccine induces protective immunity against homologous and heterologous strains of influenza virus. J. Virol. 2007, 81, 3514–3524. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.-T.; Ko, E.-J.; Lee, Y.; Kim, K.-H.; Kim, M.-C.; Lee, Y.-N.; Kang, S.-M. Intranasal vaccination with M2e5x virus-like particles induces humoral and cellular immune responses conferring cross-protection against heterosubtypic influenza viruses. PLoS ONE 2018, 13, e0190868. [Google Scholar] [CrossRef] [PubMed]
- Cai, M.; Wang, C.; Li, Y.; Gu, H.; Sun, S.; Duan, Y.; Lai, C.; Wang, K.; Yang, X.; Xing, L.; et al. Virus-like particle vaccine by intranasal vaccination elicits protective immunity against respiratory syncytial viral infection in mice. Acta Biochim. Biophys. Sin. 2017, 49, 74–82. [Google Scholar] [CrossRef] [PubMed]
- Tai, W.; Roberts, L.; Seryshev, A.; Gubatan, J.M.; Bland, C.S.; Zabriskie, R.; Kulkarni, S.; Soong, L.; Mbawuike, I.; Gilbert, B.; et al. Multistrain influenza protection induced by a nanoparticulate mucosal immunotherapeutic. Mucosal Immunol. 2011, 4, 197–207. [Google Scholar] [CrossRef]
- Metcalfe, S.M. Mesenchymal stem cells and management of COVID-19 pneumonia. Med. Drug Discov. 2020, 5, 100019. [Google Scholar] [CrossRef] [PubMed]
- Vishnu Sankar, S.; Jobin, J.; Muhammad Salman, S.; Akash, M.; Sahab, U.; Bijo, M. Silicon quantum dots: Promising theranostic probes for the future. Curr. Drug Targets 2019, 20, 1255–1263. [Google Scholar]
- Zhu, X.; Wang, X.; Han, L.; Chen, T.; Wang, L.; Li, H.; Li, S.; He, L.; Fu, X.; Chen, S.; et al. Reverse transcription loop-mediated isothermal amplification combined with nanoparticles-based biosensor for diagnosis of COVID-19. medRxiv 2020, in press. [Google Scholar] [CrossRef]
- Zhao, Z.; Cui, H.; Song, W.; Ru, X.; Zhou, W.; Yu, X. A simple magnetic nanoparticles-based viral RNA extraction method for efficient detection of SARS-CoV-2. bioRxiv 2020, in press. [Google Scholar] [CrossRef]
- Bai, C.; Zhang, H.; Zeng, L.; Zhao, X.; Ma, L. Inductive magnetic nanoparticle sensor based on microfluidic chip oil detection technology. Micromachines 2020, 11, 183. [Google Scholar] [CrossRef]
- Wang, M.; Fu, A.; Hu, B.; Tong, Y.; Liu, R.; Gu, J.; Liu, J.; Jiang, W.; Shen, G.; Zhao, W.; et al. Nanopore target sequencing for accurate and comprehensive detection of SARS-CoV-2 and other respiratory viruses. medRxiv 2020, in press. [Google Scholar] [CrossRef]
- Yu, L.; Tong, Y.; Shen, G.; Fu, A.; Lai, Y.; Zhou, X.; Yuan, Y.; Wang, Y.; Pan, Y.; Yu, Z.; et al. Immunodepletion with Hypoxemia: A potential high risk subtype of Coronavirus disease 2019. medRxiv 2020, in press. [Google Scholar] [CrossRef]
- Rapid Nano-Gold Tests Can Ease Pressure on Centralised Testing for COVID-19. Nano Magazine—Latest Nanotechnology News 2020. Available online: https://nano-magazine.com/news/2020/3/26/rapid-nano-gold-tests-can-ease-pressure-on-centralised-testing-for-covid-19 (accessed on 28 March 2020).
- Xia, T.; Kovochich, M.; Liong, M.; Zink, J.I.; Nel, A.E. Cationic Polystyrene Nanosphere Toxicity Depends on Cell-Specific Endocytic and Mitochondrial Injury Pathways. ACS Nano 2008, 2, 85–96. [Google Scholar] [CrossRef] [PubMed]
- Park, E.-J.; Yi, J.; Chung, K.-H.; Ryu, D.-Y.; Choi, J.; Park, K. Oxidative stress and apoptosis induced by titanium dioxide nanoparticles in cultured BEAS-2B cells. Toxicol. Lett. 2008, 180, 222–229. [Google Scholar] [CrossRef] [PubMed]
- Nel, A.; Xia, T.; Mädler, L.; Li, N. Toxic Potential of Materials at the Nanolevel. Science 2006, 311, 622. [Google Scholar] [CrossRef] [PubMed]
- Sun, Q.; Tan, D.; Ze, Y.; Sang, X.; Liu, X.; Gui, S.; Cheng, Z.; Cheng, J.; Hu, R.; Gao, G.; et al. Pulmotoxicological effects caused by long-term titanium dioxide nanoparticles exposure in mice. J. Hazard. Mater. 2012, 235, 47–53. [Google Scholar] [CrossRef] [PubMed]
- Ge, C.; Du, J.; Zhao, L.; Wang, L.; Liu, Y.; Li, D.; Yang, Y.; Zhou, R.; Zhao, Y.; Chai, Z.; et al. Binding of blood proteins to carbon nanotubes reduces cytotoxicity. Proc. Natl. Acad. Sci. USA 2011, 108, 16968. [Google Scholar] [CrossRef]
- Duch, M.C.; Budinger, G.R.S.; Liang, Y.T.; Soberanes, S.; Urich, D.; Chiarella, S.E.; Campochiaro, L.A.; Gonzalez, A.; Chandel, N.S.; Hersam, M.C.; et al. Minimizing Oxidation and Stable Nanoscale Dispersion Improves the Biocompatibility of Graphene in the Lung. Nano Lett. 2011, 11, 5201–5207. [Google Scholar] [CrossRef]
- Lee, H.-M.; Shin, D.-M.; Song, H.-M.; Yuk, J.-M.; Lee, Z.-W.; Lee, S.-H.; Hwang, S.M.; Kim, J.-M.; Lee, C.-S.; Jo, E.-K. Nanoparticles up-regulate tumor necrosis factor-α and CXCL8 via reactive oxygen species and mitogen-activated protein kinase activation. Toxicol. Appl. Pharmacol. 2009, 238, 160–169. [Google Scholar] [CrossRef]
- Li, R.; Ning, Z.; Majumdar, R.; Cui, J.; Takabe, W.; Jen, N.; Sioutas, C.; Hsiai, T. Ultrafine particles from diesel vehicle emissions at different driving cycles induce differential vascular pro-inflammatory responses: Implication of chemical components and NF-kappaB signaling. Part. Fibre Toxicol. 2010, 7, 6. [Google Scholar] [CrossRef]
- Hubbs, A.F.; Mercer, R.R.; Benkovic, S.A.; Harkema, J.; Sriram, K.; Schwegler-Berry, D.; Goravanahally, M.P.; Nurkiewicz, T.R.; Castranova, V.; Sargent, L.M. Nanotoxicology—A Pathologist’s Perspective. Toxicol. Pathol. 2010, 39, 301–324. [Google Scholar] [CrossRef]
- Li, S.; He, P.; Dong, J.; Guo, Z.; Dai, L. DNA-Directed Self-Assembling of Carbon Nanotubes. J. Am. Chem. Soc. 2005, 127, 14–15. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.S.; Sung, J.H.; Song, K.S.; Lee, J.H.; Kim, S.M.; Lee, G.H.; Ahn, K.H.; Lee, J.S.; Shin, J.H.; Park, J.D.; et al. Persistent DNA Damage Measured by Comet Assay of Sprague Dawley Rat Lung Cells after Five Days of Inhalation Exposure and 1 Month Post-Exposure to Dispersed Multi-Wall Carbon Nanotubes (MWCNTs) Generated by New MWCNT Aerosol Generation System. Toxicol. Sci. 2012, 128, 439–448. [Google Scholar] [CrossRef] [PubMed]
- Cho, W.-S.; Duffin, R.; Poland, C.A.; Duschl, A.; Oostingh, G.J.; MacNee, W.; Bradley, M.; Megson, I.L.; Donaldson, K. Differential pro-inflammatory effects of metal oxide nanoparticles and their soluble ions in vitro and in vivo; zinc and copper nanoparticles, but not their ions, recruit eosinophils to the lungs. Nanotoxicology 2012, 6, 22–35. [Google Scholar] [CrossRef] [PubMed]
- Shvedova, A.A.; Kisin, E.; Murray, A.R.; Johnson, V.J.; Gorelik, O.; Arepalli, S.; Hubbs, A.F.; Mercer, R.R.; Keohavong, P.; Sussman, N.; et al. Inhalation vs. aspiration of single-walled carbon nanotubes in C57BL/6 mice: Inflammation, fibrosis, oxidative stress, and mutagenesis. Am. J. Physiol. Lung Cell Mol. Physiol. 2008, 295, L552–L565. [Google Scholar] [CrossRef]
- National Science and Technology Council. Committee on Technology, Subcommittee on Nanoscale Science E, and Technology. NNI Environ. Health Saf. Res. Strategy, 2011.
- Casanova, L.M.; Jeon, S.; Rutala, W.A.; Weber, D.J.; Sobsey, M.D. Effects of air temperature and relative humidity on coronavirus survival on surfaces. Appl. Environ. Microbiol. 2010, 76, 2712–2717. [Google Scholar] [CrossRef]
- Chen, N.; Zhou, M.; Dong, X.; Qu, J.; Gong, F.; Han, Y.; Qiu, Y.; Wang, J.; Liu, Y.; Wei, Y.; et al. Epidemiological and clinical characteristics of 99 cases of 2019 novel coronavirus pneumonia in Wuhan, China: A descriptive study. Lancet 2020, 395, 507–513. [Google Scholar] [CrossRef]
- Global Times China, Chinese Scientists Have Developed a New Weapon to Combat the Coronavirus. Available online: https://twitter.com/globaltimesnews/status/1244244793393217536 (accessed on 29 March 2020).
| Drug | Mechanism of Action | |
|---|---|---|
| Chloroquine | 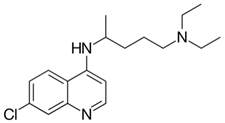 | Anti-viral effect via increase of endosomal pH required for virus/cell fusion inhibition of toll-like receptor |
| Hydroxychloroquine | 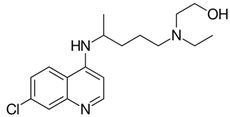 | Activity interference with terminal glycosylation of the cellular receptor ACE 2 |
| IL-6 inhibitors | Treatment of cytokine storm manifestations during ARDS | |
| Baricitinib | 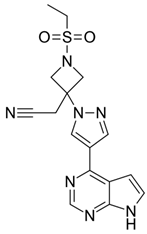 | Interference with viral penetration into the cell by blocking NAK-mediated endocytosis Treatment of cytokine storm manifestations during ARDS |
| TNF-inhibitors | Interference with viral penetration into the cell | |
| Material | Size (nm) | Virus | Antigen/Epitope | Adjuvant | References |
|---|---|---|---|---|---|
| Polymeric nanoparticles | |||||
| PLGA a | 225.4 | Bovine parainfluenza 3 virus (BPI3V) | BPI3V proteins | - | [101] |
| Swine influenza virus (H1N2) | |||||
| 200–300 | Inactivated virus H1N2 virus | - | [102] | ||
| γ- PGA b | 100–200 | Influenza (H1N1) | Hemagglutinin | - | [103] |
| Chitosan | 140 | Influenza (H1N1) | H1N1 antigen | - | [104] |
| 300–350 | Influenza (H1N1) | HA-Split | - | [105] | |
| 571.7 | Swine influenza virus (H1N2) | Killed Swine influenza | - | [106] | |
| 200–250 | Influenza (H1N1) | Antigen M2e | Heat shock protein 70 °C | [107] | |
| HPMA/NIPAM c | 12–25 | Respiratory syncytial virus (RSV) | F protein | TLR-7/8 agonist | [108,109] |
| Polyanhydride | 200–800 | Respiratory syncytial virus (RSV) | F and G glycoproteins | - | [110,111] |
| Self-assembly proteins and peptide-based nanoparticles | |||||
| N nucleocapside protein of RSV | 15 | Respiratory syncytial virus (RSV) | RSV phosphoprotein | R192G | [112] |
| 15 | Respiratory syncytial virus (RSV) | Fsll | Montanide ™ Gel01 | [113] | |
| 15 | Influenza (H1N1) | Antigen M2e | Montanide ™ Gel01 | [114] | |
| Ferritin | 12.5 | Influenza (H1N1) | Antigen M2e | - | [115] |
| Q11 peptide d | - | Influenza (H1N1) | Acid polymerase | - | [116] |
| Inorganic nanoparticles | |||||
| Gold | 12 | Influenza | Antigen M2e | CpG | [117] |
| Other | |||||
| Virus-like particles (VLP) | 80–120 | Influenza (H1N1) | Hemagglutinin | - | [118] |
| 80–120 | Influenza (H1N1, H3N2, H5N1) | Antigen M2e | - | [119] | |
| 80–120 | Respiratory syncytial virus (RSV) | F protein et G glycoprotein of RSV and M1 protein of influenza | - | [120] | |
| ISCOM e | 40 | Influenza (H1N1) | Hemagglutinin | ISCOMATRIC | [119] |
| DLPC liposomes f | 30–100 | Influenza (H1N1) | M2, HA, NP | MPL and trehalose 6,6′ dimycolate | [121] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sivasankarapillai, V.S.; Pillai, A.M.; Rahdar, A.; Sobha, A.P.; Das, S.S.; Mitropoulos, A.C.; Mokarrar, M.H.; Kyzas, G.Z. On Facing the SARS-CoV-2 (COVID-19) with Combination of Nanomaterials and Medicine: Possible Strategies and First Challenges. Nanomaterials 2020, 10, 852. https://doi.org/10.3390/nano10050852
Sivasankarapillai VS, Pillai AM, Rahdar A, Sobha AP, Das SS, Mitropoulos AC, Mokarrar MH, Kyzas GZ. On Facing the SARS-CoV-2 (COVID-19) with Combination of Nanomaterials and Medicine: Possible Strategies and First Challenges. Nanomaterials. 2020; 10(5):852. https://doi.org/10.3390/nano10050852
Chicago/Turabian StyleSivasankarapillai, Vishnu Sankar, Akhilash M. Pillai, Abbas Rahdar, Anumol P. Sobha, Sabya Sachi Das, Athanasios C. Mitropoulos, Mahboobeh Heidari Mokarrar, and George Z. Kyzas. 2020. "On Facing the SARS-CoV-2 (COVID-19) with Combination of Nanomaterials and Medicine: Possible Strategies and First Challenges" Nanomaterials 10, no. 5: 852. https://doi.org/10.3390/nano10050852
APA StyleSivasankarapillai, V. S., Pillai, A. M., Rahdar, A., Sobha, A. P., Das, S. S., Mitropoulos, A. C., Mokarrar, M. H., & Kyzas, G. Z. (2020). On Facing the SARS-CoV-2 (COVID-19) with Combination of Nanomaterials and Medicine: Possible Strategies and First Challenges. Nanomaterials, 10(5), 852. https://doi.org/10.3390/nano10050852









