Genotoxicity of Nanomaterials: Advanced In Vitro Models and High Throughput Methods for Human Hazard Assessment—A Review
Abstract
:1. Introduction
2. Methods
3. Results of Literature Search
4. General Mechanisms of Genotoxicity
5. Nanomaterials–Characteristics in Cell Culture Media and Assay Interference
6. Cytotoxicity Testing before Genotoxicity Studies
7. Biomarkers for Genotoxicity
7.1. Mammalian Gene Mutation Test
7.2. Chomosomal Damage
7.3. DNA Damage
8. Carcinogenicity
9. New Endpoints of Genotoxicity
10. High Throughput Methods
10.1. Comet Assay
10.2. In Vitro Micronucleus Assay
10.3. γH2AX Assay
10.4. ToxTracker Assay
11. New Advanced In Vitro Models (3D, Organ-on-a-Chip)
12. Conclusions and Future Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Commission Recommendation of 18 October 2011 on the definiton of a nanomaterial, (2011/696/EU). Off. J. Eur. Union 2011.
- Azqueta, A.; Dusinska, M. The use of the comet assay for the evaluation of the genotoxicity of nanomaterials. Front. Genet. 2015, 6, 239. [Google Scholar] [CrossRef] [Green Version]
- Clift, M.J.D.; Raemy, D.O.; Endes, C.; Ali, Z.; Lehmann, A.D.; Brandenberger, C.; Petri-Fink, A.; Wick, P.; Parak, W.J.; Gehr, P.; et al. Can the Ames test provide an insight into nano-object mutagenicity? Investigating the interaction between nano-objects and bacteria. Nanotoxicology 2013, 7, 1373–1385. [Google Scholar] [CrossRef] [PubMed]
- Doak, S.H.; Manshian, B.; Jenkins, G.J.S.; Singh, N. In vitro genotoxicity testing strategy for nanomaterials and the adaptation of current OECD guidelines. Mutat. Res. 2012, 745, 104–111. [Google Scholar] [CrossRef]
- OECD. Genotoxicity of Manufactured Nanomaterials: Report of the OECD Expert Meeting; Elsevier: Amsterdam, The Netherlands, 2014; Volume 472, pp. 1–37. [Google Scholar]
- Ventura, C.; Sousa-Uva, A.; Lavinha, J.; Silva, M.J. Conventional and novel “omics”-based approaches to the study of carbon nanotubes pulmonary toxicity. Environ. Mol. Mutagen. 2018, 59, 334–362. [Google Scholar] [CrossRef]
- Lan, J.; Gou, N.; Gao, C.; He, M.; Gu, A.Z. Comparative and mechanistic genotoxicity assessment of nanomaterials via a quantitative toxicogenomics approach across multiple species. Environ. Sci. Technol. 2014, 48, 12937–12945. [Google Scholar] [CrossRef] [Green Version]
- Evans, S.J.; Clift, M.J.D.; Singh, N.; De Oliveira Mallia, J.; Burgum, M.; Wills, J.W.; Wilkinson, T.S.; Jenkins, G.J.S.; Doak, S.H. Critical review of the current and future challenges associated with advanced in vitro systems towards the study of nanoparticle (secondary) genotoxicity. Mutagenesis 2017, 32, 233–241. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pfuhler, S.; Downs, T.R.; Allemang, A.J.; Shan, Y.; Crosby, M.E. Weak silica nanomaterial-induced genotoxicity can be explained by indirect DNA damage as shown by the OGG1-modified comet assay and genomic analysis. Mutagenesis 2017, 32, 5–12. [Google Scholar] [CrossRef]
- Pfuhler, S.; van Benthem, J.; Curren, R.; Doak, S.H.; Dusinska, M.; Hayashi, M.; Heflich, R.H.; Kidd, D.; Kirkland, D.; Luan, Y.; et al. Use of in vitro 3D tissue models in genotoxicity testing: Strategic fit, validation status and way forward. Report of the working group from the 7th International Workshop on Genotoxicity Testing (IWGT). Mutat. Res. Toxicol. Environ. Mutagen. 2020, 850–851, 503135. [Google Scholar] [CrossRef]
- Magdolenova, Z.; Drlickova, M.; Henjum, K.; Rundén-Pran, E.; Tulinska, J.; Bilanicova, D.; Pojana, G.; Kazimirova, A.; Barancokova, M.; Kuricova, M.; et al. Coating-dependent induction of cytotoxicity and genotoxicity of iron oxide nanoparticles. Nanotoxicology 2015, 9, 44–56. [Google Scholar] [CrossRef]
- Doak, S.H.; Dusinska, M. NanoGenotoxicology: Present and the future. Mutagenesis 2017, 32, 1–4. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nelson, B.C.; Wright, C.W.; Ibuki, Y.; Moreno-Villanueva, M.; Karlsson, H.L.; Hendriks, G.; Sims, C.M.; Singh, N.; Doak, S.H. Emerging metrology for high-throughput nanomaterial genotoxicology. Mutagenesis 2017, 32, 215–232. [Google Scholar] [CrossRef] [Green Version]
- Elje, E.; Hesler, M.; Rundén-Pran, E.; Mann, P.; Mariussen, E.; Wagner, S.; Dusinska, M.; Kohl, Y. The comet assay applied to HepG2 liver spheroids. Mutat. Res. Toxicol. Environ. Mutagen. 2019, 0–1. [Google Scholar] [CrossRef]
- Collins, A.R.; Annangi, B.; Rubio, L.; Marcos, R.; Dorn, M.; Merker, C.; Estrela-Lopis, I.; Cimpan, M.R.; Ibrahim, M.; Cimpan, E.; et al. High throughput toxicity screening and intracellular detection of nanomaterials. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 2017, 9. [Google Scholar] [CrossRef]
- Moreno-Villanueva, M.; Eltze, T.; Dressler, D.; Bernhardt, J.; Hirsch, C.; Wick, P.; von Scheven, G.; Lex, K.; Bürkle, A. The automated FADU-assay, a potential high-throughput in vitro method for early screening of DNA breakage. ALTEX 2011, 28, 295–303. [Google Scholar] [CrossRef] [Green Version]
- Sramkova, M.; Kozics, K.; Masanova, V.; Uhnakova, I.; Razga, F.; Nemethova, V.; Mazancova, P.; Kapka-Skrzypczak, L.; Kruszewski, M.; Novotova, M.; et al. Kidney nanotoxicity studied in human renal proximal tubule epithelial cell line TH1. Mutat. Res. Toxicol. Environ. Mutagen. 2019. [Google Scholar] [CrossRef]
- Brown, D.M.; Danielsen, P.H.; Derr, R.; Moelijker, N.; Fowler, P.; Stone, V.; Hendriks, G.; Møller, P.; Kermanizadeh, A. The mechanism-based toxicity screening of particles with use in the food and nutrition sector via the ToxTracker reporter system. Toxicol. In Vitro 2019, 61, 104594. [Google Scholar] [CrossRef]
- Hufnagel, M.; Schoch, S.; Wall, J.; Strauch, B.M.; Hartwig, A. Toxicity and Gene Expression Profiling of Copper- and Titanium-Based Nanoparticles Using Air-Liquid Interface Exposure. Chem. Res. Toxicol. 2020, 33, 1237–1249. [Google Scholar] [CrossRef]
- Bajpayee, M.; Kumar, A.; Dhawan, A. The Comet Assay: Assessment of In Vitro and In Vivo DNA Damage. In Genotoxicity Assessment; Humana: New York, NY, USA, 2019; pp. 237–257. [Google Scholar]
- Gao, C.-H.; Mortimer, M.; Zhang, M.; Holden, P.A.; Cai, P.; Wu, S.; Xin, Y.; Wu, Y.; Huang, Q. Impact of metal oxide nanoparticles on in vitro DNA amplification. PeerJ 2019, 7, e7228. [Google Scholar] [CrossRef]
- El Yamani, N.; Collins, A.R.; Rundén-Pran, E.; Fjellsbø, L.M.; Shaposhnikov, S.; Zienolddiny, S.; Dusinska, M. In vitro genotoxicity testing of four reference metal nanomaterials, titanium dioxide, zinc oxide, cerium oxide and silver: Towards reliable hazard assessment. Mutagenesis 2017, 32, 117–126. [Google Scholar] [CrossRef] [Green Version]
- Alhadrami, H.A. Biosensors: Classifications, medical applications, and future prospective. Biotechnol. Appl. Biochem. 2018, 65, 497–508. [Google Scholar] [CrossRef] [PubMed]
- Townsend, T.A.; Parrish, M.C.; Engelward, B.P.; Manjanatha, M.G. The development and validation of EpiComet-Chip, a modified high-throughput comet assay for the assessment of DNA methylation status. Environ. Mol. Mutagen. 2017, 58, 508–521. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wasalathanthri, D.P.; Li, D.; Song, D.; Zheng, Z.; Choudhary, D.; Jansson, I.; Lu, X.; Schenkman, J.B.; Rusling, J.F. Elucidating Organ-Specific Metabolic Toxicity Chemistry from Electrochemiluminescent Enzyme/DNA Arrays and Bioreactor Bead-LC-MS/MS. Chem. Sci. 2015, 6, 2457–2468. [Google Scholar] [CrossRef] [PubMed]
- Akagi, J.; Hall, C.J.; Crosier, K.E.; Cooper, J.M.; Crosier, P.S.; Wlodkowic, D. OpenSource lab-on-a-chip physiometer for accelerated zebrafish embryo biotests. Curr. Protoc. Cytom. 2014, 67, 9.44.1–9.44.16. [Google Scholar] [CrossRef] [PubMed]
- Klepeisz, P.; Sagmeister, S.; Haudek-Prinz, V.; Pichlbauer, M.; Grasl-Kraupp, B.; Gerner, C. Phenobarbital induces alterations in the proteome of hepatocytes and mesenchymal cells of rat livers. PLoS ONE 2013, 8, e76137. [Google Scholar] [CrossRef] [PubMed]
- Hardwick, R.N.; Betts, C.J.; Whritenour, J.; Sura, R.; Thamsen, M.; Kaufman, E.H.; Fabre, K. Drug-induced skin toxicity: Gaps in preclinical testing cascade as opportunities for complex in vitro models and assays. Lab Chip 2020, 20, 199–214. [Google Scholar] [CrossRef] [PubMed]
- Israel, J.W.; Chappell, G.A.; Simon, J.M.; Pott, S.; Safi, A.; Lewis, L.; Cotney, P.; Boulos, H.S.; Bodnar, W.; Lieb, J.D.; et al. Tissue- and strain-specific effects of a genotoxic carcinogen 1,3-butadiene on chromatin and transcription. Mamm. Genome 2018, 29, 153–167. [Google Scholar] [CrossRef]
- Zhang, Y.; Xu, S.; Wu, T.; Hu, K.; Chen, S.; Xu, A.; Wu, L. Assessment of Genotoxic Effects by Constructing a 3D Cellular System with Highly Sensitive Mutagenic Human-Hamster Hybrid Cells. Chem. Res. Toxicol. 2018, 31, 594–600. [Google Scholar] [CrossRef]
- Kermanizadeh, A.; Løhr, M.; Roursgaard, M.; Messner, S.; Gunness, P.; Kelm, J.M.; Møller, P.; Stone, V.; Loft, S. Hepatic toxicology following single and multiple exposure of engineered nanomaterials utilising a novel primary human 3D liver microtissue model. Part. Fibre Toxicol. 2014, 11, 1–15. [Google Scholar] [CrossRef] [Green Version]
- Wills, J.W.; Hondow, N.; Thomas, A.D.; Chapman, K.E.; Fish, D.; Maffeis, T.G.; Penny, M.W.; Brown, R.A.; Jenkins, G.J.S.; Brown, A.P.; et al. Genetic toxicity assessment of engineered nanoparticles using a 3D in vitro skin model (EpiDermTM). Part. Fibre Toxicol. 2016, 13, 1–21. [Google Scholar] [CrossRef] [Green Version]
- Haase, A.; Dommershausen, N.; Schulz, M.; Landsiedel, R.; Reichardt, P.; Krause, B.-C.; Tentschert, J.; Luch, A. Genotoxicity testing of different surface-functionalized SiO2, ZrO2 and silver nanomaterials in 3D human bronchial models. Arch. Toxicol. 2017, 91, 3991–4007. [Google Scholar] [CrossRef] [PubMed]
- Elje, E.; Mariussen, E.; Moriones, O.H.; Bastús, N.G.; Puntes, V.; Kohl, Y.; Dusinska, M.; Rundén-Pran, E. Hepato(Geno)toxicity assessment of nanoparticles in a hepg2 liver spheroid model. Nanomaterials 2020, 10, 545. [Google Scholar] [CrossRef] [Green Version]
- Shoeb, M.; Kodali, V.K.; Farris, B.Y.; Bishop, L.M.; Meighan, T.G.; Salmen, R.; Eye, T.; Friend, S.; Schwegler-Berry, D.; Roberts, J.R.; et al. Oxidative Stress, DNA Methylation, and Telomere Length Changes in Peripheral Blood Mononuclear Cells after Pulmonary Exposure to Metal-Rich Welding Nanoparticles. NanoImpact 2017, 5, 61–69. [Google Scholar] [CrossRef] [PubMed]
- Efeoglu, E.; Maher, M.A.; Casey, A.; Byrne, H.J. Toxicological assessment of nanomaterials: The role of in vitro Raman microspectroscopic analysis. Anal. Bioanal. Chem. 2018, 410, 1631–1646. [Google Scholar] [CrossRef] [Green Version]
- Conway, G.E.; Shah, U.-K.; Llewellyn, S.; Cervena, T.; Evans, S.J.; Al Ali, A.S.; Jenkins, G.J.; Clift, M.J.D.; Doak, S.H. Adaptation of the in vitro micronucleus assay for genotoxicity testing using 3D liver models supporting longer-term exposure durations. Mutagenesis 2020. [Google Scholar] [CrossRef] [PubMed]
- Llewellyn, S.V.; Conway, G.E.; Shah, U.-K.; Evans, S.J.; Jenkins, G.J.S.; Clift, M.J.D.; Doak, S.H. Advanced 3D Liver Models for In vitro Genotoxicity Testing Following Long-Term Nanomaterial Exposure. J. Vis. Exp. 2020. [Google Scholar] [CrossRef]
- Kiratipaiboon, C.; Stueckle, T.A.; Ghosh, R.; Rojanasakul, L.W.; Chen, Y.C.; Dinu, C.Z.; Rojanasakul, Y. Acquisition of Cancer Stem Cell-like Properties in Human Small Airway Epithelial Cells after a Long-term Exposure to Carbon Nanomaterials. Environ. Sci. Nano 2019, 6, 2152–2170. [Google Scholar] [CrossRef]
- Kirsch-Volders, M.; Decordier, I.; Elhajouji, A.; Plas, G.; Aardema, M.J.; Fenech, M. In vitro genotoxicity testing using the micronucleus assay in cell lines, human lymphocytes and 3D human skin models. Mutagenesis 2011, 26, 177–184. [Google Scholar] [CrossRef]
- Dusinska, M.; Mariussen, E.; Rundén-Pran, E.; Hudecova, A.M.; Elje, E.; Kazimirova, A.; El Yamani, N.; Dommershausen, N.; Tharmann, J.; Fieblinger, D.; et al. In vitro approaches for assessing the genotoxicity of nanomaterials. In Nanotxicity; Methods in Molecular Biology Book Series; Humana Press: New York, NY, USA, 2019; Volume 1894, pp. 83–122. [Google Scholar]
- Regulation (EC) No 1907/2006 of the European Parliament and of the Councilconcerning the Registration, Evaluation, Authorisation and Restriction of Chemicals (REACH), establishing a European Chemicals Agency, amending Directive 1999/45/EC and repealing Council Regulation (EEC) No 793/93 and Commission Regulation (EC) No 1488/94 as well as Council Directive 76/769/EEC and Commission Directives 91/155/EEC, 93/67/EEC, 93/105/EC and 2000/21/EC. Off. J. Eur. Union 2006.
- Russell, W.; Burch, R. The Principles of Humane Experimental Technique; Methuen: London, UK, 1959; pp. 1–238. [Google Scholar]
- Hackenberg, S.; Zimmermann, F.-Z.; Scherzed, A.; Friehs, G.; Froelich, K.; Ginzkey, C.; Koehler, C.; Burghartz, M.; Hagen, R.; Kleinsasser, N. Repetitive exposure to zinc oxide nanoparticles induces dna damage in human nasal mucosa mini organ cultures. Environ. Mol. Mutagen. 2011, 52, 582–589. [Google Scholar] [CrossRef] [PubMed]
- Kazimirova, A.; El Yamani, N.; Rubio, L.; García-Rodríguez, A.; Barancokova, M.; Marcos, R.; Dusinska, M. Effects of titanium dioxide nanoparticles on the Hprt gene mutations in V79 hamster cells. Nanomaterials 2020, 10, 465. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kisin, E.R.; Murray, A.R.; Sargent, L.; Lowry, D.; Chirila, M.; Siegrist, K.J.; Schwegler-Berry, D.; Leonard, S.; Castranova, V.; Fadeel, B.; et al. Genotoxicity of carbon nanofibers: Are they potentially more or less dangerous than carbon nanotubes or asbestos? Toxicol. Appl. Pharmacol. 2011, 252, 1–10. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bartek, J.; Mistrik, M.; Bartkova, J. Long-distance inflammatory and genotoxic impact of cancer in vivo. Proc. Natl. Acad. Sci. USA 2010, 107, 17861–17862. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fubini, B.; Ghiazza, M.; Fenoglio, I. Physico-chemical features of engineered nanoparticles relevant to their toxicity. Nanotoxicology 2010, 4, 347–363. [Google Scholar] [CrossRef]
- Danielsen, P.H.; Knudsen, K.B.; Štrancar, J.; Umek, P.; Koklič, T.; Garvas, M.; Vanhala, E.; Savukoski, S.; Ding, Y.; Madsen, A.M.; et al. Effects of physicochemical properties of TiO2 nanomaterials for pulmonary inflammation, acute phase response and alveolar proteinosis in intratracheally exposed mice. Toxicol. Appl. Pharmacol. 2020, 386, 114830. [Google Scholar] [CrossRef]
- Ling, C.; An, H.; Li, L.; Wang, J.; Lu, T.; Wang, H.; Hu, Y.; Song, G.; Liu, S. Genotoxicity Evaluation of Titanium Dioxide Nanoparticles In Vitro: A Systematic Review of the Literature and Meta-analysis. Biol. Trace Elem. Res. 2020. [Google Scholar] [CrossRef]
- Magdolenova, Z.; Collins, A.; Kumar, A.; Dhawan, A.; Stone, V.; Dusinska, M. Mechanisms of genotoxicity. A review of in vitro and in vivo studies with engineered nanoparticles. Nanotoxicology 2014, 8, 233–278. [Google Scholar]
- Huk, A.; Izak-Nau, E.; Reidy, B.; Boyles, M.; Duschl, A.; Lynch, I.; Dušinska, M. Is the toxic potential of nanosilver dependent on its size? Part. Fibre Toxicol. 2014, 11, 65. [Google Scholar] [CrossRef] [Green Version]
- Kononenko, V.; Repar, N.; Marušič, N.; Drašler, B.; Romih, T.; Hočevar, S.; Drobne, D. Comparative in vitro genotoxicity study of ZnO nanoparticles, ZnO macroparticles and ZnCl2 to MDCK kidney cells: Size matters. Toxicol. In Vitro 2017, 40, 256–263. [Google Scholar] [CrossRef]
- Guo, X.; Li, Y.; Yan, J.; Ingle, T.; Jones, M.Y.; Mei, N.; Boudreau, M.D.; Cunningham, C.K.; Abbas, M.; Paredes, A.M.; et al. Size- and coating-dependent cytotoxicity and genotoxicity of silver nanoparticles evaluated using in vitro standard assays. Nanotoxicology 2016, 10, 1373–1384. [Google Scholar] [CrossRef]
- Proquin, H.; Rodríguez-Ibarra, C.; Moonen, C.G.J.; Urrutia Ortega, I.M.; Briedé, J.J.; de Kok, T.M.; van Loveren, H.; Chirino, Y.I. Titanium dioxide food additive (E171) induces ROS formation and genotoxicity: Contribution of micro and nano-sized fractions. Mutagenesis 2017, 32, 139–149. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xia, Q.; Li, H.; Liu, Y.; Zhang, S.; Feng, Q.; Xiao, K. The effect of particle size on the genotoxicity of gold nanoparticles. J. Biomed. Mater. Res. A 2017, 105, 710–719. [Google Scholar] [CrossRef] [PubMed]
- Raghunathan, V.K.; Devey, M.; Hawkins, S.; Hails, L.; Davis, S.A.; Mann, S.; Chang, I.T.; Ingham, E.; Malhas, A.; Vaux, D.J.; et al. Influence of particle size and reactive oxygen species on cobalt chrome nanoparticle-mediated genotoxicity. Biomaterials 2013, 34, 3559–3570. [Google Scholar] [CrossRef] [PubMed]
- Hesler, M.; Aengenheister, L.; Ellinger, B.; Drexel, R.; Straskraba, S.; Jost, C.; Wagner, S.; Meier, F.; von Briesen, H.; Büchel, C.; et al. Multi-endpoint toxicological assessment of polystyrene nano- and microparticles in different biological models in vitro. Toxicol. In Vitro 2019, 61, 104610. [Google Scholar] [CrossRef] [PubMed]
- Zhou, F.; Liao, F.; Chen, L.; Liu, Y.; Wang, W.; Feng, S. The size-dependent genotoxicity and oxidative stress of silica nanoparticles on endothelial cells. Environ. Sci. Pollut. Res. Int. 2019, 26, 1911–1920. [Google Scholar] [CrossRef] [PubMed]
- Butler, K.S.; Peeler, D.J.; Casey, B.J.; Dair, B.J.; Elespuru, R.K. Silver nanoparticles: Correlating nanoparticle size and cellular uptake with genotoxicity. Mutagenesis 2015, 30, 577–591. [Google Scholar] [CrossRef]
- Rodriguez-Garraus, A.; Azqueta, A.; Vettorazzi, A.; López de Cerain, A. Genotoxicity of Silver Nanoparticles. Nanomaterials 2020, 10, 251. [Google Scholar] [CrossRef] [Green Version]
- Kawanishi, M.; Yoneda, R.; Totsuka, Y.; Yagi, T. Genotoxicity of micro- and nano-particles of kaolin in human primary dermal keratinocytes and fibroblasts. Genes Environ. 2020, 42, 16. [Google Scholar] [CrossRef]
- Naz, S.; Gul, A.; Zia, M. Toxicity of copper oxide nanoparticles: A review study. IET Nanobiotechnology 2020, 14, 1–13. [Google Scholar] [CrossRef]
- Lebedová, J.; Hedberg, Y.S.; Odnevall Wallinder, I.; Karlsson, H.L. Size-dependent genotoxicity of silver, gold and platinum nanoparticles studied using the mini-gel comet assay and micronucleus scoring with flow cytometry. Mutagenesis 2018, 33, 77–85. [Google Scholar] [CrossRef] [Green Version]
- Huk, A.; Izak-Nau, E.; el Yamani, N.; Uggerud, H.; Vadset, M.; Zasonska, B.; Duschl, A.; Dusinska, M. Impact of nanosilver on various DNA lesions and HPRT gene mutations—effects of charge and surface coating. Part. Fibre Toxicol. 2015, 12, 25. [Google Scholar] [CrossRef] [Green Version]
- Gábelová, A.; El Yamani, N.; Alonso, T.I.; Buliaková, B.; Srančíková, A.; Bábelová, A.; Pran, E.R.; Fjellsbø, L.M.; Elje, E.; Yazdani, M.; et al. Fibrous shape underlies the mutagenic and carcinogenic potential of nanosilver while surface chemistry affects the biosafety of iron oxide nanoparticles. Mutagenesis 2017, 32, 193–202. [Google Scholar] [CrossRef] [PubMed]
- Vales, G.; Suhonen, S.; Siivola, K.M.; Savolainen, K.M.; Catalán, J.; Norppa, H. Genotoxicity and Cytotoxicity of Gold Nanoparticles In Vitro: Role of Surface Functionalization and Particle Size. Nanomaterials 2020, 10, 271. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Falck, G.; Lindberg, H.; Suhonen, S.; Vippola, M.; Vanhala, E.; Catalán, J.; Savolainen, K.; Norppa, H. Genotoxic effects of nanosized and fine TiO2. Hum. Exp. Toxicol. 2009, 28, 339–352. [Google Scholar] [CrossRef]
- Vales, G.; Rubio, L.; Marcos, R. Long-term exposures to low doses of titanium dioxide nanoparticles induce cell transformation, but not genotoxic damage in BEAS-2B cells. Nanotoxicology 2015, 9, 568–578. [Google Scholar] [CrossRef] [PubMed]
- Osman, I.F.; Baumgartner, A.; Cemeli, E.; Fletcher, J.N.; Anderson, D. Genotoxicity and cytotoxicity of zinc oxide and titanium dioxide in HEp-2 cells. Nanomedicine 2010, 5, 1193–1203. [Google Scholar] [CrossRef] [Green Version]
- Charles, S.; Jomini, S.; Fessard, V.; Bigorgne-Vizade, E.; Rousselle, C.; Michel, C. Assessment of the in vitro genotoxicity of TiO2nanoparticles in a regulatory context. Nanotoxicology 2018, 12, 357–374. [Google Scholar] [CrossRef]
- Ghosh, S.; Ghosh, I.; Chakrabarti, M.; Mukherjee, A. Genotoxicity and biocompatibility of superparamagnetic iron oxide nanoparticles: Influence of surface modification on biodistribution, retention, DNA damage and oxidative stress. Food Chem. Toxicol. 2020, 136. [Google Scholar] [CrossRef]
- Pöttler, M.; Staicu, A.; Zaloga, J.; Unterweger, H.; Weigel, B.; Schreiber, E.; Hofmann, S.; Wiest, I.; Jeschke, U.; Alexiou, C.; et al. Genotoxicity of superparamagnetic iron oxide nanoparticles in granulosa cells. Int. J. Mol. Sci. 2015, 16, 26280–26290. [Google Scholar] [CrossRef]
- Zhang, W.; Jiang, P.; Chen, W.; Zheng, B.; Mao, Z.; Antipov, A.; Correia, M.; Larsen, E.H.; Gao, C. Genotoxicity of copper oxide nanoparticles with different surface chemistry on rat bone marrow mesenchymal stem cells. J. Nanosci. Nanotechnol. 2016, 16, 5489–5497. [Google Scholar] [CrossRef]
- Laban, B.; Ralević, U.; Petrović, S.; Leskovac, A.; Vasić-Anićijević, D.; Marković, M.; Vasić, V. Green synthesis and characterization of nontoxic L-methionine capped silver and gold nanoparticles. J. Inorg. Biochem. 2020, 204. [Google Scholar] [CrossRef] [PubMed]
- Platel, A.; Carpentier, R.; Becart, E.; Mordacq, G.; Betbeder, D.; Nesslany, F. Influence of the surface charge of PLGA nanoparticles on their in vitro genotoxicity, cytotoxicity, ROS production and endocytosis. J. Appl. Toxicol. 2016, 36, 434–444. [Google Scholar] [CrossRef] [PubMed]
- Niu, M.; Zhong, H.; Shao, H.; Hong, D.; Ma, T.; Xu, K.; Chen, X.; Han, J.; Sun, J. Shape-dependent genotoxicity of mesoporous silica nanoparticles and cellular mechanisms. J. Nanosci. Nanotechnol. 2016, 16, 2313–2318. [Google Scholar] [CrossRef]
- Gea, M.; Bonetta, S.; Iannarelli, L.; Giovannozzi, A.M.; Maurino, V.; Bonetta, S.; Hodoroaba, V.D.; Armato, C.; Rossi, A.M.; Schilirò, T. Shape-engineered titanium dioxide nanoparticles (TiO2–NPs): Cytotoxicity and genotoxicity in bronchial epithelial cells. Food Chem. Toxicol. 2019, 127, 89–100. [Google Scholar] [CrossRef] [PubMed]
- Cowie, H.; Magdolenova, Z.; Saunders, M.; Drlickova, M.; Correia Carreira, S.; Halamoda Kenzaoi, B.; Gombau, L.; Guadagnini, R.; Lorenzo, Y.; Walker, L.; et al. Suitability of human and mammalian cells of different origin for the assessment of genotoxicity of metal and polymeric engineered nanoparticles. Nanotoxicology 2015, 9, 57–65. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.; Liu, C.; Yang, D.; Zhang, H.; Xi, Z. Comparative study of cytotoxicity, oxidative stress and genotoxicity induced by four typical nanomaterials: The role of particle size, shape and composition. J. Appl. Toxicol. 2009, 29, 69–78. [Google Scholar] [CrossRef] [PubMed]
- Kisin, E.R.; Murray, A.R.; Keane, M.J.; Shi, X.C.; Schwegler-Berry, D.; Gorelik, O.; Arepalli, S.; Castranova, V.; Wallace, W.E.; Kagan, V.E.; et al. Single-walled carbon nanotubes: Geno- and cytotoxic effects in lung fibroblast V79 cells. J. Toxicol. Environ. Heal. Part A Curr. Issues 2007, 70, 2071–2079. [Google Scholar] [CrossRef]
- Catalán, J.; Siivola, K.M.; Nymark, P.; Lindberg, H.; Suhonen, S.; Järventaus, H.; Koivisto, A.J.; Moreno, C.; Vanhala, E.; Wolff, H.; et al. In vitro and in vivo genotoxic effects of straight versus tangled multi-walled carbon nanotubes. Nanotoxicology 2016, 10, 794–806. [Google Scholar] [CrossRef]
- Kim, J.S.; Lee, K.; Lee, Y.H.; Cho, H.S.; Kim, K.H.; Choi, K.H.; Lee, S.H.; Song, K.S.; Kang, C.S.; Yu, I.J. Aspect ratio has no effect on genotoxicity of multi-wall carbon nanotubes. Arch. Toxicol. 2011, 85, 775–786. [Google Scholar] [CrossRef]
- Xu, L.; Zhao, J.; Wang, Z. Genotoxic response and damage recovery of macrophages to graphene quantum dots. Sci. Total Environ. 2019, 664, 536–545. [Google Scholar] [CrossRef]
- Burgum, M.J.; Clift, M.J.D.; Evans, S.J.; Hondow, N.; Miller, M.; Lopez, S.B.; Williams, A.; Tarat, A.; Jenkins, G.J.; Doak, S.H. In Vitro Primary-Indirect Genotoxicity in Bronchial Epithelial Cells Promoted by Industrially Relevant Few-Layer Graphene. Small 2020, e2002551. [Google Scholar] [CrossRef] [PubMed]
- Syama, S.; Mohanan, P. V Safety and biocompatibility of graphene: A new generation nanomaterial for biomedical application. Int. J. Biol. Macromol. 2016, 86, 546–555. [Google Scholar] [CrossRef] [PubMed]
- Domenech, J.; Hernández, A.; Demir, E.; Marcos, R.; Cortés, C. Interactions of graphene oxide and graphene nanoplatelets with the in vitro Caco-2/HT29 model of intestinal barrier. Sci. Rep. 2020, 10, 2793. [Google Scholar] [CrossRef] [PubMed]
- Samadian, H.; Salami, M.S.; Jaymand, M.; Azarnezhad, A.; Najafi, M.; Barabadi, H.; Ahmadi, A. Genotoxicity assessment of carbon-based nanomaterials; Have their unique physicochemical properties made them double-edged swords? Mutat. Res. Mutat. Res. 2020, 783, 108296. [Google Scholar] [CrossRef] [PubMed]
- Panek, A.; Błażewicz, M.; Frączek-Szczypta, A.; Adamczyk, J.; Wiltowska-Zuber, J.; Paluszkiewicz, C. Applications of Comet Assay for the Evaluation of Genotoxicity and DNA Repair Efficiency in Nanomaterials Research. Acta Phys. Pol. A 2018, 133, 280–282. [Google Scholar] [CrossRef]
- Guo, F.; Ma, N.; Horibe, Y.; Kawanishi, S.; Murata, M.; Hiraku, Y. Nitrative DNA damage induced by multi-walled carbon nanotube via endocytosis in human lung epithelial cells. Toxicol. Appl. Pharmacol. 2012, 260, 183–192. [Google Scholar] [CrossRef] [PubMed]
- Lindberg, H.K.; Falck, G.C.M.; Suhonen, S.; Vippola, M.; Vanhala, E.; Catalán, J.; Savolainen, K.; Norppa, H. Genotoxicity of nanomaterials: DNA damage and micronuclei induced by carbon nanotubes and graphite nanofibres in human bronchial epithelial cells in vitro. Toxicol. Lett. 2009, 186, 166–173. [Google Scholar] [CrossRef]
- Wang, J.; Wang, J.; Liu, Y.; Nie, Y.; Si, B.; Wang, T.; Waqas, A.; Zhao, G.; Wang, M.; Xu, A. Aging-independent and size-dependent genotoxic response induced by titanium dioxide nanoparticles in mammalian cells. J. Environ. Sci. 2019, 85, 94–106. [Google Scholar] [CrossRef]
- Foltête, A.S.; Masfaraud, J.F.; Bigorgne, E.; Nahmani, J.; Chaurand, P.; Botta, C.; Labille, J.; Rose, J.; Férard, J.F.; Cotelle, S. Environmental impact of sunscreen nanomaterials: Ecotoxicity and genotoxicity of altered TiO2 nanocomposites on Vicia faba. Environ. Pollut. 2011, 159, 2515–2522. [Google Scholar] [CrossRef]
- Pereira, R.; Rocha-Santos, T.A.P.; Antunes, F.E.; Rasteiro, M.G.; Ribeiro, R.; Gonçalves, F.; Soares, A.M.V.M.; Lopes, I. Screening evaluation of the ecotoxicity and genotoxicity of soils contaminated with organic and inorganic nanoparticles: The role of ageing. J. Hazard. Mater. 2011, 194, 345–354. [Google Scholar] [CrossRef]
- Senapati, V.A.; Kansara, K.; Shanker, R.; Dhawan, A.; Kumar, A. Monitoring characteristics and genotoxic effects of engineered nanoparticle-protein corona. Mutagenesis 2017, 32, 479–490. [Google Scholar] [CrossRef] [PubMed]
- Dubey, P.; Matai, I.; Kumar, S.U.; Sachdev, A.; Bhushan, B.; Gopinath, P. Perturbation of cellular mechanistic system by silver nanoparticle toxicity: Cytotoxic, genotoxic and epigenetic potentials. Adv. Colloid Interface Sci. 2015, 221, 4–21. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Xu, M.; Wang, R.; Yin, Y.; Lynch, I.; Liu, S. The Crucial Role of Environmental Coronas in Determining the Biological Effects of Engineered Nanomaterials. Small 2020, 16. [Google Scholar] [CrossRef] [PubMed]
- Rasmussen, K.; Rauscher, H.; Mech, A.; Riego Sintes, J.; Gilliland, D.; González, M.; Kearns, P.; Moss, K.; Visser, M.; Groenewold, M.; et al. Physico-chemical properties of manufactured nanomaterials—Characterisation and relevant methods. An outlook based on the OECD Testing Programme. Regul. Toxicol. Pharmacol. 2018, 92, 8–28. [Google Scholar] [CrossRef]
- Hansen, U.; Thünemann, A.F. Characterization of Silver Nanoparticles in Cell Culture Medium Containing Fetal Bovine Serum. Langmuir 2015, 31, 6842–6852. [Google Scholar] [CrossRef]
- Precupas, A.; Gheorghe, D.; Botea-Petcu, A.; Leonties, A.R.; Sandu, R.; Popa, V.T.; Mariussen, E.; Naouale, E.Y.; Rundén-Pran, E.; Dumit, V.; et al. Thermodynamic Parameters at Bio-Nano Interface and Nanomaterial Toxicity: A Case Study on BSA Interaction with ZnO, SiO2, and TiO2. Chem. Res. Toxicol. 2020, 33, 2054–2071. [Google Scholar] [CrossRef]
- Savage, D.T.; Hilt, J.Z.; Dziubla, T.D. In Vitro Methods for Assessing Nanoparticle Toxicity. In Nanotoxicity; Humana Press: New York, NY, USA, 2019; pp. 1–29. [Google Scholar]
- DeLoid, G.M.; Cohen, J.M.; Pyrgiotakis, G.; Demokritou, P. An integrated dispersion preparation, characterization and in vitro dosimetry methodology for engineered nanomaterials. Nat Protoc. 2017, 12, 335–371. [Google Scholar] [CrossRef]
- Pulido-Reyes, G.; Leganes, F.; Fernández-Piñas, F.; Rosal, R. Bio-nano interface and environment: A critical review. Environ. Toxicol. Chem. 2017, 36, 3181–3193. [Google Scholar] [CrossRef]
- Moore, T.L.; Rodriguez-Lorenzo, L.; Hirsch, V.; Balog, S.; Urban, D.; Jud, C.; Rothen-Rutishauser, B.; Lattuada, M.; Petri-Fink, A. Nanoparticle colloidal stability in cell culture media and impact on cellular interactions. Chem. Soc. Rev. 2015, 44, 6287–6305. [Google Scholar] [CrossRef] [Green Version]
- Clogston, J.D.; Patri, A.K. Zeta Potential Measurement. In Characterization of Nanoparticles Intended for Drug Delivery; Methods in Molecular Biology; Humana Press: New York, NY, USA, 2011; Volume 697, pp. 63–70. [Google Scholar] [CrossRef]
- Bednar, A.J.; Poda, A.R.; Mitrano, D.M.; Kennedy, A.J.; Gray, E.P.; Ranville, J.F.; Hayes, C.A.; Crocker, F.H.; Steevens, J.A. Comparison of on-line detectors for field flow fractionation analysis of nanomaterials. Talanta 2013, 104, 140–148. [Google Scholar] [CrossRef]
- Almutary, A.; Sanderson, B.J.S. The MTT and Crystal Violet Assays: Potential Confounders in Nanoparticle Toxicity Testing. Int. J. Toxicol. 2016, 35, 454–462. [Google Scholar] [CrossRef] [PubMed]
- Oostingh, G.J.; Casals, E.; Italiani, P.; Colognato, R.; Stritzinger, R.; Ponti, J.; Pfaller, T.; Kohl, Y.; Ooms, D.; Favilli, F.; et al. Problems and challenges in the development and validation of human cell-based assays to determine nanoparticle-induced immunomodulatory effects. Part. Fibre Toxicol. 2011, 8, 8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Majtnerová, P.; Roušar, T. An overview of apoptosis assays detecting DNA fragmentation. Mol. Biol. Rep. 2018, 45, 1469–1478. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Koppen, G.; Azqueta, A.; Pourrut, B.; Brunborg, G.; Collins, A.R.; Langie, S.A.S. The next three decades of the comet assay: A report of the 11th International Comet Assay Workshop. Mutagenesis 2017, 32, 397–408. [Google Scholar] [CrossRef]
- Reisinger, K.; Blatz, V.; Brinkmann, J.; Downs, T.R.; Fischer, A.; Henkler, F.; Hoffmann, S.; Krul, C.; Liebsch, M.; Luch, A.; et al. Validation of the 3D Skin Comet assay using full thickness skin models: Transferability and reproducibility. Mutat. Res. Genet. Toxicol. Environ. Mutagen. 2018, 827, 27–41. [Google Scholar] [CrossRef]
- Riss, T.; Niles, A.; Moravec, R.; Karassina, N.; Vidugiriene, J. Cytotoxicity Assays: In Vitro Methods to Measure Dead Cells. In Assay Guidance Manual; Markossian, S., Sittampalam, G.S., Grossman, A., Brimacombe, K., Arkin, M., Auld., D., Austin, C.P., Baell, J., Caaveiro, J.M.M., Chung, T.D.Y., et al., Eds.; The Assay Guidance Manual, Eli Lilly & Company and the National Center for Advancing Translational Sciences: Bethesda, MD, USA, 2004. [Google Scholar]
- ISO 10993-5:2009—Biological Evaluation of Medical Devices—Part 5: Tests for In Vitro Cytotoxicity. Available online: https://www.iso.org/standard/36406.html (accessed on 27 August 2020).
- OECD. Test No. 476: In vitro Mammalian Cell Gene Mutation Test; OECD Publishing for the Testing Chemicals Section: Paris, France, 1997. [Google Scholar]
- OECD. Test No. 490: In Vitro Mammalian Cell Gene Mutation Tests Using the Thymidine Kinase Gene; OECD: Paris, France, 2015. [Google Scholar]
- Rubio, L.; El Yamani, N.; Kazimirova, A.; Dusinska, M.; Marcos, R. Multi-walled carbon nanotubes (NM401) induce ROS-mediated HPRT mutations in Chinese hamster lung fibroblasts. Environ. Res. 2016, 146, 185–190. [Google Scholar] [CrossRef]
- Cheng, T.F.; Patton, G.W.; Muldoon-Jacobs, K. Can the L5178Y Tk+/- mouse lymphoma assay detect epigenetic silencing? Food Chem. Toxicol. 2013, 59, 187–190. [Google Scholar] [CrossRef]
- Shibai-Ogata, A.; Kakinuma, C.; Hioki, T.; Kasahara, T. Evaluation of high-throughput screening for in vitro micronucleus test using fluorescence-based cell imaging. Mutagenesis 2011, 26, 709–719. [Google Scholar] [CrossRef] [Green Version]
- Kirsch-Volders, M.; Plas, G.; Elhajouji, A.; Lukamowicz, M.; Gonzalez, L.; Vande Loock, K.; Decordier, I. The in vitro MN assay in 2011: Origin and fate, biological significance, protocols, high throughput methodologies and toxicological relevance. Arch. Toxicol. 2011, 85, 873–899. [Google Scholar] [CrossRef]
- OECD. Test No. 487: In Vitro Mammalian Cell Micronucleus Test; OECD Guidelines for the Testing of Chemicals, Section 4; OECD: Paris, France, 2016; ISBN 9789264264861. [Google Scholar]
- García-Rodríguez, A.; Kazantseva, L.; Vila, L.; Rubio, L.; Velázquez, A.; Ramírez, M.J.; Marcos, R.; Hernández, A. Micronuclei detection by flow cytometry as a high-throughput approach for the genotoxicity testing of nanomaterials. Nanomaterials 2019, 9, 1677. [Google Scholar] [CrossRef] [Green Version]
- Di Bucchianico, S.; Cappellini, F.; Le Bihanic, F.; Zhang, Y.; Dreij, K.; Karlsson, H.L. Genotoxicity of TiO2 nanoparticles assessed by mini-gel comet assay and micronucleus scoring with flow cytometry. Mutagenesis 2017, 32, 127–137. [Google Scholar] [CrossRef] [Green Version]
- Watson, C.; Ge, J.; Cohen, J.; Pyrgiotakis, G.; Engelward, B.P.; Demokritou, P. High-throughput screening platform for engineered nanoparticle-mediated genotoxicity using cometchip technology. ACS Nano 2014, 8, 2118–2133. [Google Scholar] [CrossRef] [Green Version]
- Collins, A.; El Yamani, N.; Dusinska, M. Sensitive detection of DNA oxidation damage induced by nanomaterials. Free Radic. Biol. Med. 2017, 107, 69–76. [Google Scholar] [CrossRef]
- Collins, A.R. The use of bacterial repair endonucleases in the comet assay. In Drug Safety Evaluation; Methods in Molecular Biology; Gautier, J.C., Ed.; Humana Press: New York, NY, USA, 2010; Volume 1641. [Google Scholar] [CrossRef]
- Mah, L.J.; El-Osta, A.; Karagiannis, T.C. γh2AX: A sensitive molecular marker of DNA damage and repair. Leukemia 2010, 24, 679–686. [Google Scholar] [CrossRef] [Green Version]
- Rothkamm, K.; Barnard, S.; Moquet, J.; Ellender, M.; Rana, Z.; Burdak-Rothkamm, S. DNA damage foci: Meaning and significance. Environ. Mol. Mutagen. 2015, 56, 491–504. [Google Scholar] [CrossRef]
- Wan, R.; Mo, Y.; Tong, R.; Gao, M.; Zhang, Q. Determination of phosphorylated histone H2AX in nanoparticle-induced genotoxic studies. In Nanotoxicity; Methods in Molecular Biology; Humana Press: New York, NY, USA, 2019; Volume 1894, pp. 145–159. [Google Scholar] [CrossRef]
- Elespuru, R.; Pfuhler, S.; Aardema, M.J.; Chen, T.; Doak, S.H.; Doherty, A.; Farabaugh, C.S.; Kenny, J.; Manjanatha, M.; Mahadevan, B.; et al. Genotoxicity assessment of nanomaterials: Recommendations on best practices, assays, and methods. Toxicol. Sci. 2018, 164, 391–416. [Google Scholar] [CrossRef]
- Bin-Jumah, M.N.; Al-Abdan, M.; Al-Basher, G.; Alarifi, S. Molecular Mechanism of Cytotoxicity, Genotoxicity, and Anticancer Potential of Green Gold Nanoparticles on Human Liver Normal and Cancerous Cells. Dose Response 2020, 18, 1559325820912154. [Google Scholar] [CrossRef] [Green Version]
- El-Hussein, A.; Hamblin, M.R. ROS generation and DNA damage with photo-inactivation mediated by silver nanoparticles in lung cancer cell line. IET Nanobiotechnol. 2017, 11, 173–178. [Google Scholar] [CrossRef] [Green Version]
- Audebert, M.; Dolo, L.; Perdu, E.; Cravedi, J.-P.; Zalko, D. Use of the γH2AX assay for assessing the genotoxicity of bisphenol A and bisphenol F in human cell lines. Arch. Toxicol. 2011, 85, 1463–1473. [Google Scholar] [CrossRef]
- Bonner, W.M.; Redon, C.E.; Dickey, J.S.; Nakamura, A.J.; Sedelnikova, O.A.; Solier, S.; Pommier, Y. γH2AX and cancer. Nat. Rev. Cancer 2008, 8, 957–967. [Google Scholar] [CrossRef]
- Zhu, L.; Chang, D.W.; Dai, L.; Hong, Y. DNA damage induced by multiwalled carbon nanotubes in mouse embryonic stem cells. Nano Lett. 2007, 7, 3592–3597. [Google Scholar] [CrossRef] [PubMed]
- Cveticanin, J.; Joksic, G.; Leskovac, A.; Petrovic, S.; Sobot, A.V.; Neskovic, O. Using carbon nanotubes to induce micronuclei and double strand breaks of the DNA in human cells. Nanotechnology 2010, 21, 015102. [Google Scholar] [CrossRef] [PubMed]
- Valdiglesias, V.; Costa, C.; Kiliç, G.; Costa, S.; Pásaro, E.; Laffon, B.; Teixeira, J.P. Neuronal cytotoxicity and genotoxicity induced by zinc oxide nanoparticles. Environ. Int. 2013, 55, 92–100. [Google Scholar] [CrossRef]
- Chattopadhyay, N.; Cai, Z.; Kwon, Y.L.; Lechtman, E.; Pignol, J.P.; Reilly, R.M. Molecularly targeted gold nanoparticles enhance the radiation response of breast cancer cells and tumor xenografts to X-radiation. Breast Cancer Res. Treat. 2013, 137, 81–91. [Google Scholar] [CrossRef]
- Msiska, Z.; Pacurari, M.; Mishra, A.; Leonard, S.S.; Castranova, V.; Vallyathan, V. DNA Double-Strand Breaks by Asbestos, Silica, and Titanium Dioxide. Am. J. Respir. Cell Mol. Biol. 2010, 43, 210–219. [Google Scholar] [CrossRef]
- Paget, V.; Dekali, S.; Kortulewski, T.; Grall, R.; Gamez, C.; Blazy, K.; Aguerre-Chariol, O.; Chevillard, S.; Braun, A.; Rat, P.; et al. Specific uptake and genotoxicity induced by polystyrene nanobeads with distinct surface chemistry on human lung epithelial cells and macrophages. PLoS ONE 2015, 10. [Google Scholar] [CrossRef] [Green Version]
- Hernández, L.G.; van Steeg, H.; Luijten, M.; van Benthem, J. Mechanisms of non-genotoxic carcinogens and importance of a weight of evidence approach. Mutat. Res. Rev. Mutat. Res. 2009, 682, 94–109. [Google Scholar] [CrossRef]
- Trouiller, B.; Reliene, R.; Westbrook, A.; Solaimani, P.; Schiestl, R.H. Titanium dioxide nanoparticles induce DNA damage and genetic instability in vivo in mice. Cancer Res. 2009, 69, 8784–8789. [Google Scholar] [CrossRef] [Green Version]
- Toyooka, T.; Amano, T.; Ibuki, Y. Titanium dioxide particles phosphorylate histone H2AX independent of ROS production. Mutat. Res. Genet. Toxicol. Environ. Mutagen. 2012, 742, 84–91. [Google Scholar] [CrossRef]
- Kerckaert, G.A.; LeBoeuf, R.A.; Isfort, R.J. Use of the Syrian hamster embryo cell transformation assay for determining the carcinogenic potential of heavy metal compounds. Fundam. Appl. Toxicol. 1996, 34, 67–72. [Google Scholar] [CrossRef]
- LeBoeuf, R.A.; Kerckaert, G.A.; Aardema, M.J.; Gibson, D.P.; Brauninger, R.; Isfort, R.J. The pH 6.7 Syrian hamster embryo cell transformation assay for assessing the carcinogenic potential of chemicals. Mutat. Res. 1996, 356, 85–127. [Google Scholar] [CrossRef]
- Mauthe, R.J.; Gibson, D.P.; Bunch, R.T.; Custer, L.; Syrian Hamster Embryo Assay Working Group. Response to Alternative models for carcinogenicity testing: Weight of evidence across models. Sam Cohen, Toxicologic Pathology (2001) 29(suppl.): 183–190. Toxicol. Pathol. 2002, 30, 292–293. [Google Scholar] [CrossRef] [PubMed]
- Creton, S.; Aardema, M.J.; Carmichael, P.L.; Harvey, J.S.; Martin, F.L.; Newbold, R.F.; O’Donovan, M.R.; Pant, K.; Poth, A.; Sakai, A.; et al. Cell transformation assays for prediction of carcinogenic potential: State of the science and future research needs. Mutagenesis 2012, 27, 93–101. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fontana, C.; Kirsch, A.; Seidel, C.; Marpeaux, L.; Darne, C.; Gaté, L.; Remy, A.; Guichard, Y. In vitro cell transformation induced by synthetic amorphous silica nanoparticles. Mutat. Res. 2017, 823, 22–27. [Google Scholar] [CrossRef]
- Cui, Y.; Liu, H.; Ze, Y.; Zengli, Z.; Hu, Y.; Cheng, Z.; Cheng, J.; Hu, R.; Gao, G.; Wang, L.; et al. Gene expression in liver injury caused by long-term exposure to titanium dioxide nanoparticles in mice. Toxicol. Sci. 2012, 128. [Google Scholar] [CrossRef] [Green Version]
- Dawson, M.A.; Kouzarides, T. Cancer epigenetics: From mechanism to therapy. Cell 2012, 150, 12–27. [Google Scholar] [CrossRef] [Green Version]
- Gao, G.; Ze, Y.; Li, B.; Zhao, X.; Zhang, T.; Sheng, L.; Hu, R.; Gui, S.; Sang, X.; Sun, Q.; et al. Ovarian dysfunction and gene-expressed characteristics of female mice caused by long-term exposure to titanium dioxide nanoparticles. J. Hazard. Mater. 2012, 243, 19–27. [Google Scholar] [CrossRef]
- Gojova, A.; Lee, J.-T.; Jung, H.S.; Guo, B.; Barakat, A.I.; Kennedy, I.M. Effect of cerium oxide nanoparticles on inflammation in vascular endothelial cells. Inhal. Toxicol. 2009, 21 (Suppl. 1), 123–130. [Google Scholar] [CrossRef] [Green Version]
- Moos, P.J.; Olszewski, K.; Honeggar, M.; Cassidy, P.; Leachman, S.; Woessner, D.; Cutler, N.S.; Veranth, J.M. Responses of human cells to ZnO nanoparticles: A gene transcription study. Metallomics 2011, 3, 1199–1211. [Google Scholar] [CrossRef] [Green Version]
- Park, E.-J.; Choi, J.; Park, Y.-K.; Park, K. Oxidative stress induced by cerium oxide nanoparticles in cultured BEAS-2B cells. Toxicology 2008, 245, 90–100. [Google Scholar] [CrossRef]
- Smolkova, B.; El Yamani, N.; Collins, A.R.; Gutleb, A.C.; Dusinska, M. Nanoparticles in food. Epigenetic changes induced by nanomaterials and possible impact on health. Food Chem. Toxicol. 2015, 77, 64–73. [Google Scholar] [CrossRef]
- Rossner, P.; Vrbova, K.; Rossnerova, A.; Zavodna, T.; Milcova, A.; Klema, J.; Vecera, Z.; Mikuska, P.; Coufalik, P.; Capka, L.; et al. Gene Expression and Epigenetic Changes in Mice Following Inhalation of Copper(II) Oxide Nanoparticles. Nanomaterials 2020, 10, 550. [Google Scholar] [CrossRef] [Green Version]
- Smolkova, B.; Dusinska, M.; Gabelova, A. Nanomedicine and epigenome. Possible health risks. Food Chem. Toxicol. 2017, 109, 780–796. [Google Scholar] [CrossRef]
- Cavalli, G.; Heard, E. Advances in epigenetics link genetics to the environment and disease. Nature 2019, 571, 489–499. [Google Scholar] [CrossRef] [Green Version]
- Smolkova, B.; Dusinska, M.; Gabelova, A. Epigenetic Effects of Nanomaterials. Encycl. Environ. Heal. 2019, 678–685. [Google Scholar] [CrossRef]
- Stoccoro, A.; Karlsson, H.L.; Coppedè, F.; Migliore, L. Epigenetic effects of nano-sized materials. Toxicology 2013, 313, 3–14. [Google Scholar] [CrossRef]
- Lamas, A.; Franco, C.M.; Regal, P.; Miranda, J.M.; Vázquez, B.; Cepeda, A. High-Throughput Platforms in Real-Time PCR and Applications. In Polymerase Chain Reaction for Biomedical Applications; InTech: London, UK, 2016; Chapter 2; ISBN 978-953-51-2796-3. [Google Scholar] [CrossRef] [Green Version]
- Dusinska, M.; Tulinska, J.; El Yamani, N.; Kuricova, M.; Liskova, A.; Rollerova, E.; Rundén-Pran, E.; Smolkova, B. Immunotoxicity, genotoxicity and epigenetic toxicity of nanomaterials: New strategies for toxicity testing? Food Chem. Toxicol. 2017, 109, 797–811. [Google Scholar] [CrossRef]
- Nel, A.; Xia, T.; Meng, H.; Wang, X.; Lin, S.; Ji, Z.; Zhang, H. Nanomaterial toxicity testing in the 21st century: Use of a predictive toxicological approach and high-throughput screening. Acc. Chem. Res. 2013, 46, 607–621. [Google Scholar] [CrossRef]
- Seo, J.-E.; Tryndyak, V.; Wu, Q.; Dreval, K.; Pogribny, I.; Bryant, M.; Zhou, T.; Robison, T.W.; Mei, N.; Guo, X. Quantitative comparison of in vitro genotoxicity between metabolically competent HepaRG cells and HepG2 cells using the high-throughput high-content CometChip assay. Arch. Toxicol. 2019, 93, 1433–1448. [Google Scholar] [CrossRef]
- Mack, M.; Schweinlin, K.; Mirsberger, N.; Zubel, T.; Bürkle, A. Automated screening for oxidative or methylation-induced DNA damage in human cells. ALTEX 2020. [Google Scholar] [CrossRef]
- Baltazar, M.T.; Cable, S.; Carmichael, P.L.; Cubberley, R.; Cull, T.; Delagrange, M.; Dent, M.P.; Hatherell, S.; Houghton, J.; Kukic, P.; et al. A Next-Generation Risk Assessment Case Study for Coumarin in Cosmetic Products. Toxicol. Sci. 2020, 176, 236–252. [Google Scholar] [CrossRef]
- Johann, S.; Goßen, M.; Behnisch, P.A.; Hollert, H.; Seiler, T.-B. Combining Different In Vitro Bioassays to Evaluate Genotoxicity of Water-Accommodated Fractions from Petroleum Products. Toxics 2020, 8, 45. [Google Scholar] [CrossRef]
- Seo, J.-E.; Wu, Q.; Bryant, M.; Ren, L.; Shi, Q.; Robison, T.W.; Mei, N.; Manjanatha, M.G.; Guo, X. Performance of high-throughput CometChip assay using primary human hepatocytes: A comparison of DNA damage responses with in vitro human hepatoma cell lines. Arch. Toxicol. 2020, 94, 2207–2224. [Google Scholar] [CrossRef]
- Takeiri, A.; Matsuzaki, K.; Motoyama, S.; Yano, M.; Harada, A.; Katoh, C.; Tanaka, K.; Mishima, M. High-content imaging analyses of γH2AX-foci and micronuclei in TK6 cells elucidated genotoxicity of chemicals and their clastogenic/aneugenic mode of action. Genes Environ. Off. J. Jpn. Environ. Mutagen Soc. 2019, 41, 4. [Google Scholar] [CrossRef] [Green Version]
- Damoiseaux, R.; George, S.; Li, M.; Pokhrel, S.; Ji, Z.; France, B.; Xia, T.; Suarez, E.; Rallo, R.; Mädler, L.; et al. No time to lose—high throughput screening to assess nanomaterial safety. Nanoscale 2011, 3, 1345. [Google Scholar] [CrossRef]
- George, S.; Xia, T.; Rallo, R.; Zhao, Y.; Ji, Z.; Lin, S.; Wang, X.; Zhang, H.; France, B.; Schoenfeld, D.; et al. Use of a high-throughput screening approach coupled with in vivo zebrafish embryo screening to develop hazard ranking for engineered nanomaterials. ACS Nano 2011, 5, 1805–1817. [Google Scholar] [CrossRef] [Green Version]
- Moller, P.; Moller, L.; Godschalk, R.W.L.; Jones, G.D.D. Assessment and reduction of comet assay variation in relation to DNA damage: Studies from the European Comet Assay Validation Group. Mutagenesis 2010, 25, 109–111. [Google Scholar] [CrossRef] [Green Version]
- Wilde, S.; Dambowsky, M.; Hempt, C.; Sutter, A.; Queisser, N. Classification of in vitro genotoxicants using a novel multiplexed biomarker assay compared to the flow cytometric micronucleus test. Environ. Mol. Mutagen. 2017, 58, 662–677. [Google Scholar] [CrossRef] [Green Version]
- Shaposhnikov, S.; Azqueta, A.; Henriksson, S.; Meier, S.; Gaivão, I.; Huskisson, N.H.; Smart, A.; Brunborg, G.; Nilsson, M.; Collins, A.R. Twelve-gel slide format optimised for comet assay and fluorescent in situ hybridisation. Toxicol. Lett. 2010, 195, 31–34. [Google Scholar] [CrossRef]
- Gutzkow, K.B.; Langleite, T.M.; Meier, S.; Graupner, A.; Collins, A.R.; Brunborg, G. High-throughput comet assay using 96 minigels. Mutagenesis 2013, 28, 333–340. [Google Scholar] [CrossRef] [Green Version]
- Azqueta, A.; Gutzkow, K.B.; Priestley, C.C.; Meier, S.; Walker, J.S.; Brunborg, G.; Collins, A.R. A comparative performance test of standard, medium- and high-throughput comet assays. Toxicol. In Vitro 2013, 27, 768–773. [Google Scholar] [CrossRef] [PubMed]
- Stang, A.; Witte, I. Performance of the comet assay in a high-throughput version. Mutat. Res. Genet. Toxicol. Environ. Mutagen. 2009, 675, 5–10. [Google Scholar] [CrossRef] [PubMed]
- Huk, A.; Collins, A.R.; El Yamani, N.; Porredon, C.; Azqueta, A.; De Lapuente, J.; Dusinska, M. Critical factors to be considered when testing nanomaterials for genotoxicity with the comet assay. Mutagenesis 2015, 30, 85–88. [Google Scholar] [CrossRef]
- García-Rodríguez, A.; Rubio, L.; Vila, L.; Xamena, N.; Velázquez, A.; Marcos, R.; Hernández, A. The comet assay as a tool to detect the genotoxic potential of nanomaterials. Nanomaterials 2019, 9, 1385. [Google Scholar] [CrossRef] [Green Version]
- Rubio, L.; Annangi, B.; Vila, L.; Hernández, A.; Marcos, R. Antioxidant and anti-genotoxic properties of cerium oxide nanoparticles in a pulmonary-like cell system. Arch. Toxicol. 2016, 90, 269–278. [Google Scholar] [CrossRef]
- Fenech, M.; Kirsch-Volders, M.; Rossnerova, A.; Sram, R.; Romm, H.; Bolognesi, C.; Ramakumar, A.; Soussaline, F.; Schunck, C.; Elhajouji, A.; et al. HUMN project initiative and review of validation, quality control and prospects for further development of automated micronucleus assays using image cytometry systems. Int. J. Hyg. Environ. Health 2013, 216, 541–552. [Google Scholar] [CrossRef]
- Roemer, E.; Zenzen, V.; Conroy, L.L.; Luedemann, K.; Dempsey, R.; Schunck, C.; Sticken, E.T. Automation of the in vitro micronucleus and chromosome aberration assay for the assessment of the genotoxicity of the particulate and gas-vapor phase of cigarette smoke. Toxicol. Mech. Methods 2015, 25, 320–333. [Google Scholar] [CrossRef]
- Decordier, I.; Papine, A.; Plas, G.; Roesems, S.; Vande Loock, K.; Moreno-Palomo, J.; Cemeli, E.; Anderson, D.; Fucic, A.; Marcos, R.; et al. Automated image analysis of cytokinesis-blocked micronuclei: An adapted protocol and a validated scoring procedure for biomonitoring. Mutagenesis 2009, 24, 85–93. [Google Scholar] [CrossRef]
- Nüsse, M.; Kramer, J. Flow cytometric analysis of micronuclei found in cells after irradiation. Cytometry 1984, 5, 20–25. [Google Scholar] [CrossRef]
- Avlasevich, S.; Bryce, S.; De Boeck, M.; Elhajouji, A.; Van Goethem, F.; Lynch, A.; Nicolette, J.; Shi, J.; Dertinger, S. Flow cytometric analysis of micronuclei in mammalian cell cultures: Past, present and future. Mutagenesis 2011, 26, 147–152. [Google Scholar] [CrossRef] [Green Version]
- Bryce, S.M.; Avlasevich, S.L.; Bemis, J.C.; Phonethepswath, S.; Dertinger, S.D. Miniaturized flow cytometric in vitro micronucleus assay represents an efficient tool for comprehensively characterizing genotoxicity dose-response relationships. Mutat. Res. Genet. Toxicol. Environ. Mutagen. 2010, 703, 191–199. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bryce, S.M.; Avlasevich, S.L.; Bemis, J.C.; Tate, M.; Walmsley, R.M.; Saad, F.; Van Dijck, K.; De Boeck, M.; Van Goethem, F.; Lukamowicz-Rajska, M.; et al. Flow cytometric 96-well microplate-based in vitro micronucleus assay with human TK6 cells: Protocol optimization and transferability assessment. Environ. Mol. Mutagen. 2013, 54, 180–194. [Google Scholar] [CrossRef] [PubMed]
- Bryce, S.M.; Bemis, J.C.; Avlasevich, S.L.; Dertinger, S.D. In vitro micronucleus assay scored by flow cytometry provides a comprehensive evaluation of cytogenetic damage and cytotoxicity. Mutat. Res. 2007, 630, 78–91. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cai, Z.; Vallis, K.A.; Reilly, R.M. Computational analysis of the number, area and density of gamma-H2AX foci in breast cancer cells exposed to (111)In-DTPA-hEGF or gamma-rays using Image-J software. Int. J. Radiat. Biol. 2009, 85, 262–271. [Google Scholar] [CrossRef] [PubMed]
- Hou, Y.-N.; Lavaf, A.; Huang, D.; Peters, S.; Huq, R.; Friedrich, V.; Rosenstein, B.S.; Kao, J. Development of an Automated γ-H2AX Immunocytochemistry Assay. Radiat. Res. 2009, 171, 360–367. [Google Scholar] [CrossRef] [PubMed]
- Roch-Lefèvre, S.; Mandina, T.; Voisin, P.; Gaëtan, G.; Mesa, J.E.G.; Valente, M.; Bonnesoeur, P.; García, O.; Voisin, P.; Roy, L. Quantification of γ-H2AX Foci in Human Lymphocytes: A Method for Biological Dosimetry after Ionizing Radiation Exposure. Radiat. Res. 2010, 174, 185–194. [Google Scholar] [CrossRef]
- Bhogal, N.; Jalali, F.; Bristow, R.G. Microscopic imaging of DNA repair foci in irradiated normal tissues. Int. J. Radiat. Biol. 2009, 85, 732–746. [Google Scholar] [CrossRef]
- Ivashkevich, A.N.; Martin, O.A.; Smith, A.J.; Redon, C.E.; Bonner, W.M.; Martin, R.F.; Lobachevsky, P.N. γH2AX foci as a measure of DNA damage: A computational approach to automatic analysis. Mutat. Res. 2011, 711, 49–60. [Google Scholar] [CrossRef] [Green Version]
- Harris, G.; Palosaari, T.; Magdolenova, Z.; Mennecozzi, M.; Gineste, J.M.; Saavedra, L.; Milcamps, A.; Huk, A.; Collins, A.R.; Dusinska, M.; et al. Iron oxide nanoparticle toxicity testing using high-throughput analysis and high-content imaging. Nanotoxicology 2015, 9 (Suppl. 1), 87–94. [Google Scholar] [CrossRef]
- Karlsson, H.L.; Gliga, A.R.; Calléja, F.M.G.R.; Gonçalves, C.S.A.G.; Wallinder, I.O.; Vrieling, H.; Fadeel, B.; Hendriks, G. Mechanism-based genotoxicity screening of metal oxide nanoparticles using the ToxTracker panel of reporter cell lines. Part. Fibre Toxicol. 2014, 11, 41. [Google Scholar] [CrossRef] [Green Version]
- Alépée, N. State-of-the-art of 3D cultures (organs-on-a-chip) in safety testing and pathophysiology. ALTEX 2014, 441–477. [Google Scholar] [CrossRef]
- Chen, L.; Li, N.; Liu, Y.; Faquet, B.; Alépée, N.; Ding, C.; Eilstein, J.; Zhong, L.; Peng, Z.; Ma, J.; et al. A new 3D model for genotoxicity assessment: EpiSkinTM Micronucleus Assay. Mutagenesis 2020, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Tahara, H.; Sadamoto, K.; Yamagiwa, Y.; Nemoto, S.; Kurata, M. Investigation of comet assays under conditions mimicking ocular instillation administration in a three-dimensional reconstructed human corneal epithelial model. Cutan. Ocul. Toxicol. 2019, 38, 375–383. [Google Scholar] [CrossRef] [PubMed]
- Mandon, M.; Huet, S.; Dubreil, E.; Fessard, V.; Le Hégarat, L. Three-dimensional HepaRG spheroids as a liver model to study human genotoxicity in vitro with the single cell gel electrophoresis assay. Sci. Rep. 2019, 9, 10548. [Google Scholar] [CrossRef]
- Štampar, M.; Tomc, J.; Filipič, M.; Žegura, B. Development of in vitro 3D cell model from hepatocellular carcinoma (HepG2) cell line and its application for genotoxicity testing. Arch. Toxicol. 2019, 93, 3321–3333. [Google Scholar] [CrossRef]
- Shah, U.-K.; Mallia, J.d.O.; Singh, N.; Chapman, K.E.; Doak, S.H.; Jenkins, G.J.S. A three-dimensional in vitro HepG2 cells liver spheroid model for genotoxicity studies. Mutat. Res. Toxicol. Environ. Mutagen. 2018, 825, 51–58. [Google Scholar] [CrossRef] [Green Version]
- van Duinen, V.; Trietsch, S.J.; Joore, J.; Vulto, P.; Hankemeier, T. Microfluidic 3D cell culture: From tools to tissue models. Curr. Opin. Biotechnol. 2015, 35, 118–126. [Google Scholar] [CrossRef] [Green Version]
- Bhatia, S.N.; Ingber, D.E. Microfluidic organs-on-chips. Nat. Biotechnol. 2014, 32, 760–772. [Google Scholar] [CrossRef]
- Huh, D.; Leslie, D.C.; Matthews, B.D.; Fraser, J.P.; Jurek, S.; Hamilton, G.A.; Thorneloe, K.S.; McAlexander, M.A.; Ingber, D.E. A human disease model of drug toxicity-induced pulmonary edema in a lung-on-a-chip microdevice. Sci. Transl. Med. 2012, 4. [Google Scholar] [CrossRef] [Green Version]
- Bricks, T.; Paullier, P.; Legendre, A.; Fleury, M.-J.; Zeller, P.; Merlier, F.; Anton, P.M.; Leclerc, E. Development of a new microfluidic platform integrating co-cultures of intestinal and liver cell lines. Toxicol. In Vitro 2014, 28, 885–895. [Google Scholar] [CrossRef]
- Gordon, S.; Daneshian, M.; Bouwstra, J.; Caloni, F.; Constant, S.; Davies, D.E.; Dandekar, G.; Guzman, C.A.; Fabian, E.; Haltner, E.; et al. Non-animal models of epithelial barriers (skin, intestine and lung) in research, industrial applications and regulatory toxicology. ALTEX 2015, 32, 327–378. [Google Scholar] [CrossRef] [PubMed]
- Maschmeyer, I.; Lorenz, A.K.; Schimek, K.; Hasenberg, T.; Ramme, A.P.; Hübner, J.; Lindner, M.; Drewell, C.; Bauer, S.; Thomas, A.; et al. A four-organ-chip for interconnected long-term co-culture of human intestine, liver, skin and kidney equivalents. Lab Chip 2015, 15, 2688–2699. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Maschmeyer, I.; Hasenberg, T.; Jaenicke, A.; Lindner, M.; Lorenz, A.K.; Zech, J.; Garbe, L.A.; Sonntag, F.; Hayden, P.; Ayehunie, S.; et al. Chip-based human liver-intestine and liver-skin co-cultures—A first step toward systemic repeated dose substance testing in vitro. Eur. J. Pharm. Biopharm. 2015, 95, 77–87. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Materne, E.-M.; Maschmeyer, I.; Lorenz, A.K.; Horland, R.; Schimek, K.M.S.; Busek, M.; Sonntag, F.; Lauster, R.; Marx, U. The multi-organ chip—A microfluidic platform for long-term multi-tissue coculture. J. Vis. Exp. 2015, e52526. [Google Scholar] [CrossRef] [Green Version]
- Kim, H.J.; Ingber, D.E. Gut-on-a-Chip microenvironment induces human intestinal cells to undergo villus differentiation. Integr. Biol. 2013, 5, 1130–1140. [Google Scholar] [CrossRef] [Green Version]
- Kim, H.J.; Huh, D.; Hamilton, G.; Ingber, D.E. Human gut-on-a-chip inhabited by microbial flora that experiences intestinal peristalsis-like motions and flow. Lab Chip 2012, 12, 2165–2174. [Google Scholar] [CrossRef]
- Yin, F.; Zhu, Y.; Zhang, M.; Yu, H.; Chen, W.; Qin, J. A 3D human placenta-on-a-chip model to probe nanoparticle exposure at the placental barrier. Toxicol. In Vitro 2019, 54, 105–113. [Google Scholar] [CrossRef]
- Zhang, M.; Xu, C.; Jiang, L.; Qin, J. A 3D human lung-on-a-chip model for nanotoxicity testing. Toxicol. Res. 2018, 7, 1048–1060. [Google Scholar] [CrossRef] [Green Version]
- Dong, H.; Sun, H. A Microchip for Integrated Single-Cell Gene Expression Profiling and Genotoxicity Detection. Sensors 2016, 16, 1489. [Google Scholar] [CrossRef] [Green Version]
- Dong, H.; Sun, H.; Zheng, J. A microchip for integrated single-cell genotoxicity assay. Talanta 2016, 161, 804–811. [Google Scholar] [CrossRef]
- Vecchio, G.; Fenech, M.; Pompa, P.P.; Voelcker, N.H. Lab-on-a-chip-based high-throughput screening of the genotoxicity of engineered nanomaterials. Small 2014, 10, 2721–2734. [Google Scholar] [CrossRef] [PubMed]
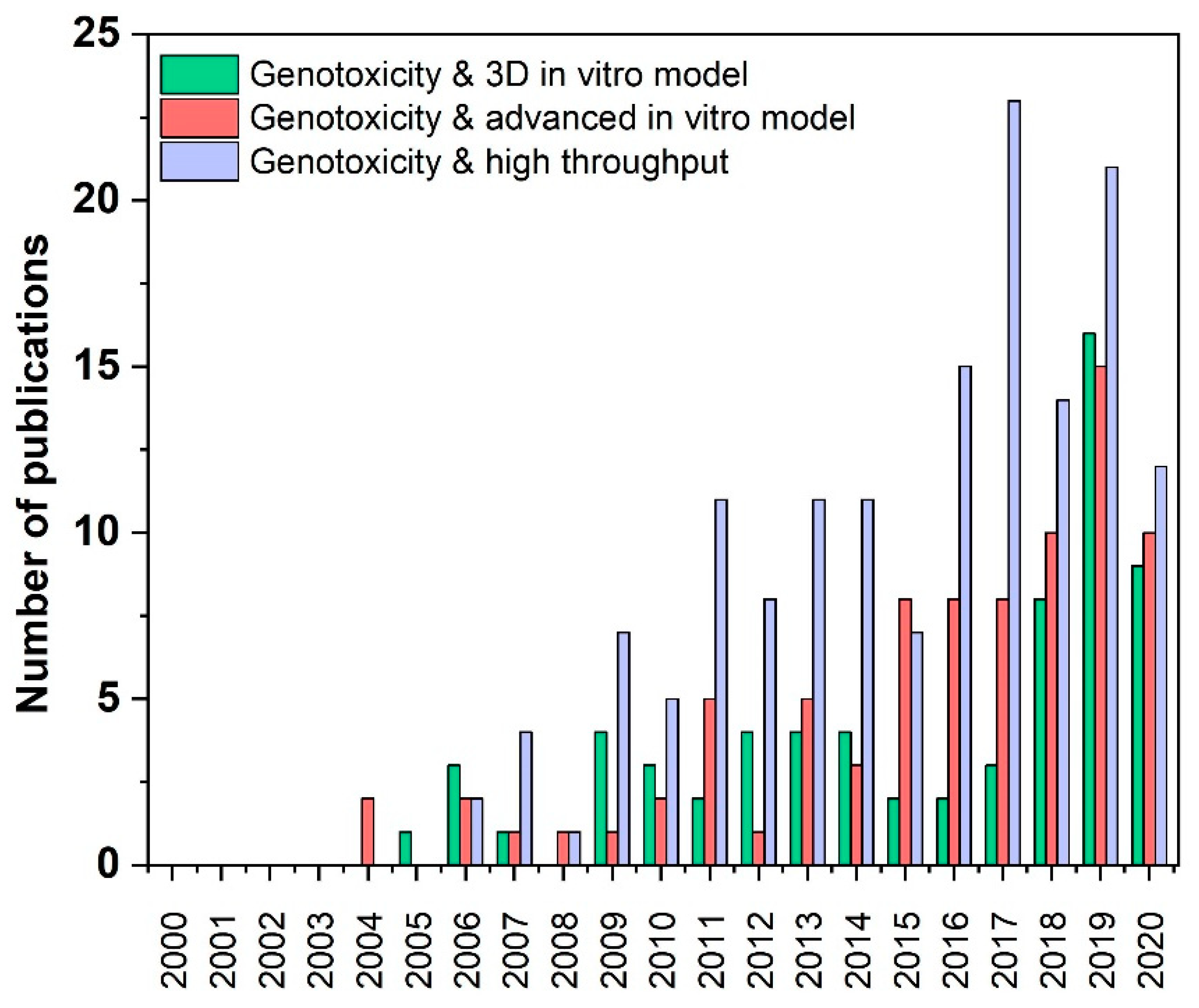
| Keywords of Literature Search | Number of Publications Per Time Period | ||
|---|---|---|---|
| 2000–2020 | 2010–2020 | 2019–2020 | |
| genotoxicity and 3D models in vitro | 59 | 47 | 23 |
| genotoxicity and advanced in vitro model | 73 | 63 | 23 |
| genotoxicity and & high throughput | 134 | 120 | 32 |
| genotoxicity and organ on chip | 7 | 7 * | 2 |
| genotoxicity and 3D models & nanoparticles | 7 | 7 ** | 1 |
| genotoxicity and 3D models & nanomaterials | 11 | 11 *** | 5 |
| genotoxicity and high throughput & nanoparticles | 6 | 6 *** | 4 |
| genotoxicity and high throughput & nanomaterials | 8 | 8 *** | 4 |
| Human Cell Origin | Cell Model | Culture Dimension | Endpoints | Test Methods | NM Tested | Year | Ref |
|---|---|---|---|---|---|---|---|
| Nasal mucosa | mini organ culture | 3D | cytotoxicity, genotoxicity | comet assay, caspase–3 ELISA, ROS assay, TEM | ZnO | 2011 | [44] |
| lung | EpiAirway™ | 3D | cytotoxicity, genotoxicity | comet assay, LDH assay, ATP assay | Ag, SiO2, ZrO2 | 2017 | [33] |
| lung | small airway epithelium | 2D | cytotoxicity, genotoxicity | apoptosis assay, CFA, p53 downregulation, IFL, γH2AX assay, spheroid formation assay | SWCNT, MWCNT, UFCB, ASB | 2019 | [39] |
| skin | EpiDerm™, TK6 | 2D & 3D | cytotoxicity, genotoxicity | MNA, TEM, RPD analysis | SiO2 | 2016 | [32] |
| liver | HepG2 | 2D & 3D | cytotoxicity, genotoxicity | comet assay, LD staining, alamarBlue™ assay | Ag, ZnO, TiO2 | 2020 | [34] |
| liver | HepG2 | 3D | cytotoxicity, genotoxicity, liver functionality | MNA, cytokine secretion | Ag, TiO2 | 2020 | [38] |
| liver | HepG2, HepaRG | 3D | cytotoxicity, genotoxicity, liver functionality | MNA, TB assay, albumin level, urea level | ZnO | 2020 | [37] |
| liver | micro tissue | 3D | cytotoxicity, genotoxicity, liver functionality | comet assay, LD staining, adenylate kinase assay, cytokine secretion, comet assay, albumin ELISA, CYP3A4 activity, lipid peroxidation assay | Ag, ZnO, MWCNT, TiO2 | 2014 | [31] |
| liver, blood, breast | HepG2, TK6, MCF7 | 2D | genotoxicity | comet assay, Epi-comet assay | - | 2017 | [24] |
| liver lung kidney, intestine | n. sp. | 2D | genotoxicity | ECL fluidic chip LC-MS/MS | - | 2015 | [25] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kohl, Y.; Rundén-Pran, E.; Mariussen, E.; Hesler, M.; El Yamani, N.; Longhin, E.M.; Dusinska, M. Genotoxicity of Nanomaterials: Advanced In Vitro Models and High Throughput Methods for Human Hazard Assessment—A Review. Nanomaterials 2020, 10, 1911. https://doi.org/10.3390/nano10101911
Kohl Y, Rundén-Pran E, Mariussen E, Hesler M, El Yamani N, Longhin EM, Dusinska M. Genotoxicity of Nanomaterials: Advanced In Vitro Models and High Throughput Methods for Human Hazard Assessment—A Review. Nanomaterials. 2020; 10(10):1911. https://doi.org/10.3390/nano10101911
Chicago/Turabian StyleKohl, Yvonne, Elise Rundén-Pran, Espen Mariussen, Michelle Hesler, Naouale El Yamani, Eleonora Marta Longhin, and Maria Dusinska. 2020. "Genotoxicity of Nanomaterials: Advanced In Vitro Models and High Throughput Methods for Human Hazard Assessment—A Review" Nanomaterials 10, no. 10: 1911. https://doi.org/10.3390/nano10101911
APA StyleKohl, Y., Rundén-Pran, E., Mariussen, E., Hesler, M., El Yamani, N., Longhin, E. M., & Dusinska, M. (2020). Genotoxicity of Nanomaterials: Advanced In Vitro Models and High Throughput Methods for Human Hazard Assessment—A Review. Nanomaterials, 10(10), 1911. https://doi.org/10.3390/nano10101911







