Abstract
Nanomaterials are increasingly being used in new products and devices with a great impact on different fields from sensoristics to biomedicine. Biosynthesis of nanomaterials by microorganisms is recently attracting interest as a new, exciting approach towards the development of ‘greener’ nanomanufacturing compared to traditional chemical and physical approaches. This review provides an insight about microbial biosynthesis of nanomaterials by bacteria, yeast, molds, and microalgae for the manufacturing of sensoristic devices and therapeutic/diagnostic applications. The last ten-year literature was selected, focusing on scientific works where aspects like biosynthesis features, characterization, and applications have been described. The knowledge, challenges, and potentiality of microbial-mediated biosynthesis was also described. Bacteria and microalgae are the main microorganism used for nanobiosynthesis, principally for biomedical applications. Some bacteria and microalgae have showed the ability to synthetize unique nanostructures: bacterial nanocellulose, exopolysaccharides, bacterial nanowires, and biomineralized nanoscale materials (magnetosomes, frustules, and coccoliths). Yeasts and molds are characterized by extracellular synthesis, advantageous for possible reuse of cell cultures and reduced purification processes of nanomaterials. The intrinsic variability of the microbiological systems requires a greater protocols standardization to obtain nanomaterials with increasingly uniform and reproducible chemical-physical characteristics. A deeper knowledge about biosynthetic pathways and the opportunities from genetic engineering are stimulating the research towards a breakthrough development of microbial-based nanosynthesis for the future scaling-up and possible industrial exploitation of these promising ‘nanofactories’.
1. Introduction
During the period of 2016–2022 the global nanomaterials market is expected to grow with a compound annual growth rate of about 20% or more [1]. One of the major challenges for the global advancement of nanomaterials market is the environmental sustainability of nanomanufacturing processes. Indeed, traditional top-down or bottom-up chemical and physical nanomanufacturing approaches have a greater energy-intensity compared to manufacturing processes of bulk materials. Further, they are often characterized by low process yields (using acidic/basic chemicals and organic solvents), generation of greenhouse gases, and they require specific facilities, operative conditions (e.g., from moderate to high vacuum), and high purity levels of starting materials [2,3,4]. The principles of green chemistry (“the invention, design and application of chemical products and processes to reduce or to eliminate the use and generation of hazardous substances”) combined with white biotechnology (“biotechnology that uses living cells—yeasts, molds, bacteria, plants, and enzymes to synthesize products at industrial scale”) can really contribute to the development of more sustainable industrial processes [5], also for nanomanufacturing. The microbial-mediated biosynthesis of nanomaterials is a promising biotechnological-based nanomanufacturing process that represents a ‘green’ alternative approach to physical and chemical strategies of nanosynthesis [6,7]. The microbial-mediated biosynthesis of metallic (also as alloys), non-metallic, or metal oxides nanoparticles have been reported for many microbial strains of bacteria, yeast, molds, and microalgae [8] (Figure 1).
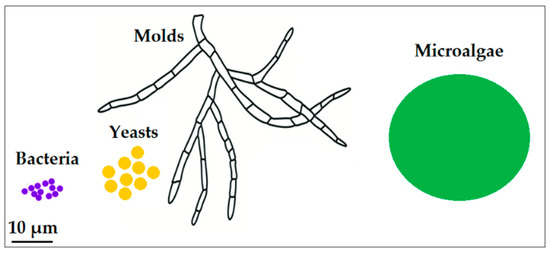
Figure 1.
Schematic comparing average sizes of the microorganisms described in this review.
In addition, some microorganisms have shown the capability to biosynthesize unique nanostructured materials, i.e., biomineralized nanostructures like silicified frustules [9], calcified coccoliths [10], magnetosomes [11], and organic nanomaterials like bacterial nanocellulose [12] exopolysaccharide nanoparticles [13] and bacterial nanowires [14]. The microbial-mediated biosynthesis of nanomaterials has been extensively explored showing different advantages and features including the following: (i) synthetized nanomaterials have defined chemical composition, size and morphology, (ii) biosynthesis is performed at mild physico-chemical conditions, (iii) easily handling and cultivation of microbial cells and possibility of cell culture scale-up, (iv) possibility of in vivo tuning nanomaterial characteristics by changing key parameters of cell culture operational set up or through genetically engineering tools [15]. In order to enable a broad applicability of microbial-mediated biosynthesis of nanomaterials as a real alternative to ‘traditional’ synthetic approaches to nanomanufacturing, many hurdles still need to be overcome: a reduction of polidispersity of nanoparticles, a more complete characterization of biocapping layer agents, effectiveness of removal procedures of biocapping layer and nanomaterials purifications, standardization of microbial cell culture protocols for reproducibility of nanosynthesis processess, as well as production costs and yields. Overeaching the challenge for the development of reliable eco-friendly nanotechnologies for nanomaterial synthesis is of utmost importance for future exploitations of broad-impact nanostructured-based technologies and applications, like innovative optical and electrochemical (bio) sensoristic devices [16] and therapeutic and diagnostic applications of nanostructured materials e.g., for drug delivery, in vivo/in vitro imaging and development of antimicrobial and antitumoral drugs [17,18]. In the first part of this review, we reported an overview of scientific literature (mainly from the last ten years) about in vivo microbial biosynthesis of nanomaterials that have been used for (bio) sensoristic and biomedical purposes. We focused on works that have covered several key aspects of nanomaterials: (i) type of biosynthesis (in some cases post-biosynthesis functionalization), (ii) biosynthetic pathways (presumptive or demonstrated), (iii) characterization, (iv) applications. In the second part, main acquired knowledge, challenges, and potentiality of microbial-mediated biosynthesis has been described.
2. Microbial-Mediated Biosynthesis of Nanomaterials for Sensoristic and Biomedical Applications
2.1. Bacteria
In the last ten years, bacteria have been used to synthesize inorganic nanomaterials (mainly selenium, gold, and silver nanoparticles) with interesting properties for the development of voltammetric sensoristic devices [19], and third-generation biosensors [20], for possible diagnostic applications [21] like cell imaging and biolabeling [22] and for applications where no surface coat is required, like annealing and thin film formation [23] (Table 1). Bacterial-biosynthesized nanoparticles have mainly shown in vitro antimicrobial activity against some pathogenic bacterial strains [24,25,26,27,28] and properties i.e., antioxidant [29], anti-proliferative, anti-migration [30], anticoagulant [31], and anticancer [26,27,28,29,30,31,32,33]. Biochemical mechanisms which mediate the bacterial biosynthesis of nanoparticles have been proposed or they are currently under investigation. Many of these biochemical mechanisms have been described as part of microbial resistance mechanisms for cellular detoxification which involves changes in solubility of inorganic ions by enzymatic reduction and/or precipitation of soluble toxic to insoluble non-toxic nanostructures. Both extracellular and intracellular biocatalytic synthesis (with and possible excretion) mainly involves oxidoreductase enzymes (e.g., NADH-dependent nitrate reductase, NADPH-dependent sulphite reductase flavoprotein subunit α, and cysteine desulfhydrase) and cellular transporters. Physicochemical processes like biosorption, complexation, nucleation, growth, and stabilization mediated by biomolecules (e.g., proteins and carbohydrates) have also been described. In addition to inorganic nanomaterials, some bacteria genera have shown the ability to biosynthesize very peculiar organic nanostructures. Bacterial nanocellulose is a 3-D network of cellulose nanofibrils produced by aerobic acetic bacteria like those belonging to the genus Gluconacetobacter, the most efficient bacteria for nanocellulose biosynthesis. Compared to the nanocrystalline cellulose and nanofibrillated cellulose, bacterial nanocellulose shows higher purity, crystallinity and mechanical stability [34]. Therefore, bacterial nanocellulose is a nanomaterial which has attracted great attention for use in biomedical applications (e.g., as antimicrobial agent, for drug delivery systems and scaffolds for tissue engineering) and on biosensoristic platforms (as nanocomposite and as support for the immobilization of biological recognition elements) [35,36,37]. Exopolysaccharides are microbial extracellular biopolymers with different roles in adhesion of bacterial biofilms and as protection agents. In a recent work a novel self-assembled and spherical nanosized non-glucan exopolysaccharide has been described for bacteria Lactobacillus plantarum-605. Results have showed its reducing actions for rapid (30 min.) biosynthesis of good monodispersed gold and silver nanoparticles without any pretreatment or modification [38]. Bacterial nanowires are conductive proteinaceous pilus-like nanostructures involved in extracellular electron transport processes of anaerobic dissimilatory metal-reducing bacteria like Geobacter and Shewanella genera [39], aerobic bacteria like Pseudomonas aeruginosa [40] and aerobic photosynthetic cyanobacteria like Microcystis and Synechocystis genera [41]. Metallic-like conductivity (due to aromatic amino acids-richness in PilA proteic fibers) and a redox-based conductivity (mediated by cytochrome OmcS present on fibers surface) have been hypothesized for bacterial nanowires in G. sulfurreducens [39]. Studies on nanowires Shewanella oneidensis MR-1 strain have showed a p-type, tunable electronic behavior with electrical conductivities comparable to moderately doped inorganic semiconductors used in synthetic organic semiconductor-based devices like field-effect transistors [42]. The bacterium S. oneidensis have been also described for biosynthesis of gold and silver nanomaterials [23,24]. Bacterial nanowires are also very promising nanostructures in the bioelectronic field for the development of new biomaterial for microbial fuel cells and electrochemical (bio) sensoristic devices i.e., as direct electron transfer mediator between bacteria biofilm and the solid-state electrode surfaces. Different silicon-based electrodes for rapid biochemical oxygen demand (BOD) determination and water integral toxicity monitoring have been described in recent literature [43,44,45]. Bacterial magnetosomes are organic-coated intracellular nanocrystals of Fe3O4 and/or Fe3S4, biosynthesized by both magnetotactic and non-magnetotactic bacteria. The composition of magnetic inorganic part is species-specific, and the external organic coating layer is derived from bacterial phospholipid bilayer membrane. The putative functions of protein component of the external organic coating layer in the magnetosome biomineralization process have been hypothesized [11]. Bacterial Fe3O4 magnetosomes are stable single-magnetic domains permanently magnetic at ambient temperature, possessing peculiar characteristics of high chemical purity, a narrow size range and consistent crystal morphology [46]. Some recent applications include molecular imaging [47], cancer therapy [48], and the development of a chip-based whole-cell biosensor for toxicity assessment [49].

Table 1.
Nanomaterials synthesized by bacteria.
2.2. Yeasts and Molds
The research focused on biosynthesis of nanomaterials by fungi, like yeasts and molds, have brought to the coinage of the term ‘myconanotechnology’, in order to refer to a newly emerging domain of nanotechnology. Yeasts are unicellular fungi mainly known in nanosynthesis for their ability to produce semiconductor nanoparticles [8]. Biosynthesis of high water-soluble and biocompatible cadmium telluride quantum dots by model organism yeast Saccharomyces cerevisiae have been reported in literature. These cadmium telluride quantum dots have showed interesting characteristics of size-tunable (changing culture time and temperature) emission and photoluminescence quantum yield as good candidate for bio-imaging and bio-labelling applications [50]. S. cerevisiae have been also used for biosynthesis of Au–Ag alloy nanoparticles for electrochemical sensors fabrication [51,52], aimed to the determination of paracetamol in tablet samples and vanillin in vanilla bean and vanilla tea sample, respectively (see Table 2). Possible biosynthesis mechanisms of nanoparticles by S. cerevisiae could involve membrane bound and cytosolic oxidoreductases as well as extracellular 1,3-β-glucan synthase-mediated formation and growth of nanoparticles [50,51,52,53]. Molds are a large group of microscopic filamentous fungi that include many genera like to Penicillum, Aspergillus, and Fusarium. Compared to bacteria, molds possess many distinctive advantages for biosynthesis of nanomaterials: (i) higher metal tolerance, (ii) higher metal binding and uptake capabilities, (iii) easy culturing and fast growing; (iv) higher extracellular nanosynthesis (mediated by extracellular enzyme, reductive proteins, and secondary secreted metabolites). Extracellular biosynthesis of nanomaterials poses advantages in terms of a possible reuse of cell cultures for new biosynthesis (cell lysis not required) and reduced nanoparticle downstream purification processes [54]. Proposed mechanisms behind fungal synthesis of nanoparticles hypothesized a possible involvement of biomolecules secreted in formation and stabilization of nanoparticles [55], secreted reductases [56,57] and possible trapping of metal ions by electrostatic interaction with positively charged groups in enzymes present in cell wall of the mycelia [58]. In the last ten years, several works have described mold-based biosynthesis of nanoparticles (silver, gold, and tellurium) and quantum dots (zinc sulfide, zinc sulfide with gadolinium, and lead sulfide). These nanoparticles have shown both antibacterial activity [54,57,59] and antitumoral activity [55,56,58] beside possible employment in optical detection of heavy metals and arsenic in water [60,61] (Table 2).

Table 2.
Nanomaterials synthesized by yeasts and molds.
2.3. Microalgae
Microalgae are unicellular photosynthetic microorganisms that have attracted significant interest in the field of nanomanufacturing [62] (see Table 3). Microalgae like Tetraselmis kochinensis, Scenedesmus and Desmodesmus have been used for the biosynthesis of noble metal nanoparticles with good antimicrobial activity, useful for applications in biomedical tool designing but also in drug delivery, catalysis and electronics [63,64,65]. For these microalgae, mechanisms described for nanoparticles biosynthesis include phenomena of nucleation, control of dimension, and stabilization of nanoparticle structure, mediated by reducing agents [64], enzymes present in the cell wall cytoplasmic membrane [63], biomolecules like polysaccharides, proteins, polyphenols and phenolic compounds [65]. Mechanisms behind biological mineralization (or biomineralization), i.e., the in vivo inorganic minerals formation, have been extensively studied for possible development of new nanomaterials. Diatoms are unicellular microalgae with very peculiar biomineralized silica cell wall called frustules. Diatom frustules possess a highly periodic and hierarchical 3D-porous micro-nanostructure of different morphology (pennate and centric). Hypotheses about their natural functions include mechanical protection, biological protections, filtration, DNA protection from UV and optimization of light harvesting [66,67,68]. Compared to analogous synthetic mesoporous silica materials, e.g., MCM-4, diatom frustules possess different advantages, including higher biocompatibility, reduced toxicity and easily purification. Diatom frustules also exhibit interesting optical and optoelectronic properties [66,69,70,71,72,73,74]. The abundance of silanol groups (Si-OH) make the diatom frustules surface easily functionalizable (also in vivo), thus allowing to fully exploit the potentiality of structural nanopatterning of diatom frustules [75]. Functionalization with antibodies [72,73,74,75,76,77] and gold nanoparticles [78,79] has been described for different sensitive materials in optical or electrochemical immunosensors. Diatom frustules have showed potential application in drug delivery systems [80,81,82]. Diatomaceous earth (or diatomite) is a large available microfossil material from diatom frustules with extensive commercial use in abrasives or filters. Recently, diatomite—gold nanoparticle nanocomposites have been described for on-chip chromatography and surface-enhanced Raman scattering-based sensoristic devices for the detection of cocaine in biological samples [83] and of histamine in salmon and tuna samples [84]. The “diatom nanotechnology” is a rapidly evolving research field which aims to fully exploit the unique properties of diatom frustules and the great potential of silica biomineralization cellular pathway for the development of new functionalized nanomaterials for emerging applications in sensing, photonic and drug delivery [85]. Another characteristic of diatom frustules is the presence of xanthophyll pigment fucoxanthin. Recent studies have highlighted the active role of fucoxanthin as photo-reducing agent of metal ions to stabilize silver nanoparticles. These silver nanoparticles have showed a significant in vitro antimicrobial activity against Escherichia coli, Bacillus stearothermophilus, and Streptococcus mutans [86] and possible application in optical chemosensing of dissolved ammonia in water samples [87]. Compared to diatoms, microalgae coccolithophores have received less attention. Coccolithophores are calcifying nanoplankton that produces CaCO3 microparticles (coccoliths), in form of arrays of nanoscaled substructures. Interesting optical properties (e.g., light scattering) of coccoliths have been described but also some drawbacks including low electrical conductivity, dissolution at low pH values and scarcity of surface functional groups. Despite these, coccoliths morphologies have showed a great applicative potential for nanodevices fabrications, especially following appropriate in vivo or in vitro modification and functionalizations [10,78]. In very recent work the fabrication of an electrochemical aptamer-based sandwich-type biosensor for the detection of type 2 diabetes biomarker Vaspin has been described [88].

Table 3.
Nanomaterials synthesized by microalgae.
3. Towards a Large-Scale Applicability: Knowledge, Issues, and Potentiality
The in vivo microbial nanobiosynthesis and possible control and tuning of nanomaterial properties represent a concrete opportunity for future development and promising uses in biosensoristics and biomedical fields. Despite all the advantages, microbial nanotechnology still has very limited uses [89]. Bacteria have showed the ability to synthetize nanomaterials either by extracellular or intracellular mechanisms. These mechanisms generally produce opposite advantages and disvantages in terms of metal nanoparticles dispersity and purification. Extracellularly produced nanoparticles are generally more polydispersed (i.e., with a great variability in size) than intracellularly produced nanoparticles. By contrast, in extracellular nanomaterial productions less downstream extraction/purification steps (e.g., ultrasound treatment and detergent uses) are required. Thus, the extracellular mediated synthesis described for yeast and molds can greatly simplify the purification steps, besides being an advantage for a possible reuse of microorganisms for more biosynthesis cycles. However, the characterization and identification of the enzymes responsible for nanobiosynthesis in molds is still uncomplete. The photoautotrophic metabolism of microalgae and cyanobacteria is based on carbon dioxide (as carbon source), light (as energy source), inorganic nutrients and water. This condition generally reduce the costs of culture media (compared to culture media used for the growth of bacteria, yeasts, and molds) and it can strongly spur the future scaling-up from the laboratory to the industrial scale, also through the design and the development of solar photobioreactors for the fixation (and reduction) of atmospheric carbon dioxide.
3.1. Nanoparticles Dispersion and Capping Layers
One of the main challenges in microbial nanobiosynthesis is the control of dispersity of nanostructure materials, which heavily influence electronic and optical properties, and the isolation and purification of plural form. Dispersity, i.e., the size distribution of the nanoparticle population, is a key property that strongly influences the particle’s behavior in fluids. Improvement and optimization of extraction and purification protocols are required, both for intracellular and extracellular biosynthesis: methods like freeze-thawing, osmotic shock and centrifugation could lead to changes in nanoparticle structures as well as aggregation and precipitation phenomena. Through the adoption of suitable strategies, microbial biosynthesis of nanoparticles could be improved. Selection of appropriate microbial strains (in terms of growth rate and biocatalytic activities), optimization of culturing conditions and uses of genetic engineering tools could help to overcome drawbacks linked to slower producing rate and polidispersity (compared to chemical-based nanomanufacturing) [90]. Microbial biosynthetic nanoparticles are characterized by the presence of a capping layer of biomolecules adsorbed on the surface that act as stabilizing agent and biological active layer of nanoparticles [21]. A deep knowledge of capping characteristics, a clear identification of capping agents (mainly peptides like glutathione, metallothioneins, membrane associated proteins etc.), and possible purification of nanoparticles [23] are fundamental for future in vivo medical applications [15,91].
3.2. Cell Culture Conditions
For future large-scale productions, costs of culture media for microbial growth should be seriously considered to not limit the applications of microbial biosynthetic nanomaterials. One current example is bacterial nanocellulose whose applications are still limited to a few biomedical devices, mainly because of costs of culture medium [92]. In addition, optimization and standardization of microbial cell culture growth protocols and modifications culture conditions are pivotal for control, tune, and to improve characteristics of microbial biosynthesized nanomaterials. The influence of physico-chemical parameters of cell culture operational set up on nanomaterials biosynthesis have been previously highlighted. These factors include (i) microbial cell concentration; (ii) precursor concentration; (iii) pH; and (iv) temperature. The optimum conditions of pH, temperature, and NaCl concentration have been studied to achieve high purity and high synthesis rate of cadmium selenide nanoparticles by bacterium Pseudomonas aeruginosa strain RB. Interestingly, the results of this work have showed that optimum conditions for nanoparticles synthesis did not match with optimum growth conditions for Pseudomonas aeruginosa strain RB [93]. Effects of precursor concentration, temperature, and pH on silver nanoparticle synthesis and particle sizes have been described for the bacterium E. coli strain DH5α [94]. Other recent examples include: (i) the temperature-dependence of size and monodispersity of silver nanoparticles biosynthesis by mold Trichoderma viride [95], (ii) the influence on the type of gold nanostructures synthetized (nanoparticles or nanoplates) and them relative size in yeast Yarrowia lipolytica strain NCIM 3589 by changing the proportion of cell concentration and precursor gold salt concentration, (iii) the effect of temperature on gold nanostructures release from cell wall into the aqueous phase [96] and control of bacterial growth kinetics of the bacterium Morganella psychrotolerans to achieve shape anisotropy of silver nanoparticles [97]. Elsoud et al. (2018) observed an improvement in tellurium nanoparticles production by a 1 kGy of gamma irradiation of mold Aspergillus welwitschiae KY766958 broth culture (compared to the non-irradiated broth culture control). These results have been ascribed to the activation of enzyme (s) involved in biosynthetic pathway [59]. Although the controlling of biomineralization process still remains a challenge, the optimization of frustules morphological properties (e.g., pore sizes and pore density) has been explored by changing operational parameters of experimental setup (e.g., pH, salinity, temperature, nutrient concentration, precursor Si(OH)4 concentration, and light regime) [98]. Interestingly Townley et al. (2007) reported alteration in pore sizes of Coscinodiscus wailesii frustules when exposed to sublethal concentration of nickel [99]. The possibility of in vivo chemical modification of frustules or other biomineralized structures has been recently described. These in vivo chemical modifications lead to the inorganic elements/compounds-doping of biomineralized structures through the addition to the culture medium of given precursors at sublethal concentrations. Several works described the doping of diatom Pinnularia sp. frustules or diatoms Thalassiosira weissflogii frustules with titania (TiO2) [100,101] nanobiosynthesis containing Si-Ge oxides nanocomb in diatom Nitzschia and Pinnularia sp. by adding Ge(OH)4 or GeO2 in the diatom culture medium at photobioreactor scale-productions [102,103,104,105]. Compared to diatoms, possible nanotechnological applications of the calcareous-based shell of marine protozoa foraminifera have not been so widely explored. A recent work described the in vivo preparation of a bionic material through the inclusion of fluorescent magnetite nanoparticles within calcite skeletal structure of the unicellular organism foraminifer Amphistrigina lesson. Such in vitro synthetic approach exploited the natural biomineralization process of growth in the presence of magnetic nanoparticles functionalized with a hydroxylated dextran shell [106].
3.3. Biochemistry, Molecular Biology, and Genetics
A deeper knowledge of molecular biology and genetic aspects behind microbial nanobiosynthetic pathways is strongly required. For instance, the characterization of not fully understood biochemical mechanisms and a complete identification of extracellular enzymes secreted by filamentous fungi could lead to an improved control in chemical compositions, shapes, and sizes of nanoparticles [55]. The availability of microorganism genome sequences could considerably increase the range of possibilities in genetic manipulation of microorganism to implement the nanoparticles biosynthesis and the in vivo tuning of nanoparticle characteristics. A recent example has come from the study of Zhang et al. (2017), which showed how CdSe quantum dots biosynthesis can be improved through genetic modification of the ATP metabolism pathway in yeast S. cerevisiae [107]. Biotechnological approaches based on genetic engineering and recombinant technologies could allow the identification of sequences of gene involved in nanoparticle synthesis and a possible heterologous expression (i.e., controlled expression of one or more gene sequences in a host organism) to enhance nanomaterial production efficiency [91]. The bacterium E. coli is a highly efficient model host microorganism that has been exploited as heterologous expression system for phytochelatin synthase and/or metallothionein for the in vivo synthesis of various metal nanoparticles (e.g., CdSeZn, PrGd, CdCs, and FeCo) never synthesized before by chemical methods [108,109] or cadmium selenide quantum dots [110]. Thanks to recent advances in the characterization of biochemical mechanisms involved in bacterial nanocellulose biosynthesis, the future development of new genetically engineered bacterial nanocellulose-producing strains could be achieved. This could eventually lead to a reduction of production costs, an improvement of production yield, and biosynthesis of nanocellulose with new properties, suitable for broader range of technological applications [92,111]. Concerning genetic manipulation of nanostructure-producing microorganisms, Tan et al. (2016) have reported a 2000-fold increase in electrical conductivity and diameter of nanowires filaments produced by model microorganism Geobacter sulfurreducens. In this case, genetic modification has been concerned the modification of aminoacidic composition of the carboxyl end of PilA protein, the structural component of bacterial nanowires [112]. The biomineralization process behind magnetosome biogenesis in bacterial species (Magnetospirillum species are the most studied) is very complex and not completely elucidated. The mechanism of magnetosome formation has been shown to be under tight genetic control and induced by growth conditions. To date, six different models have been proposed to elucidate magnetosome formation, but they are still not completed [11]. Delalat et al. (2015) have reported a genetically engineered diatom Thalassiosira pseudonana (whose genome has been completely sequenced) to incorporate immunoglubulin G-binding domain of protein G (GB1) into the frustules surface. Such antibody-labelled genetically modified diatom enables an in vitro selectively cell targeting and selectively killing of neuroblastoma and B-lymphoma cells [113]. Furthermore, the role of gene Silicanin-1 in the control of biosilica morphology has been recently highlighted in Thalassiosira pseudonana, opening new possibilities for future genetic engineering of frustules architectures [114]. A deeper knowledge of biosilicification process as well as the role of organic components of diatom frustules (i.e., proteins silaffins and long-chain polyamines) in biogenesis and formation of nanopatterns in diatom frustules are still a challenge. To date, biocalcification system of coccolithophores remains unclear, even though a very recent genetic and proteomic study about expression of transcripts and proteins in coccolithophore Emiliania huxleyi will help future identification and more detailed characterization of molecular mechanisms and metabolic pathways underlying calcification in coccolithophores [115].
4. Conclusions
In the light of recent literature herein reported, microbial nanotechnology is a fascinating and booming field for future breakthrough nanomaterial synthesis. Through a ‘green’ and sustainable approach, microbial nanotechnology can really spur innovation in nanomanufacturing with a potential strong impact in several fields, including sensoristics and biomedicine.
Author Contributions
Conceptualization, G.G.; writing—original draft preparation, G.G., D.Z., R.D.; writing—review and editing, G.G., D.Z., R.D.; visualization, G.G., D.Z., R.D.; supervision, R.D. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Inshakova, E.; Inshakov, O. World market for nanomaterials: Structure and trends. In Proceedings of the MATEC Web of Conferences, Sevastopol, Russia, 11–15 September 2017; EDP Sciences: Les Ulis, France, 2017; Volume 129, p. 02013. [Google Scholar] [CrossRef]
- Şengül, H.; Theis, T.L.; Ghosh, S. Toward sustainable nanoproducts: An overview of nanomanufacturing methods. J. Ind. Ecol. 2008, 12, 329–359. [Google Scholar] [CrossRef]
- Fang, F.Z.; Zhang, X.D.; Gao, W.; Guo, Y.B.; Byrne, G.; Hansen, H.N. Nanomanufacturing—Perspective and applications. CIRP Ann. 2017, 66, 683–705. [Google Scholar] [CrossRef]
- Yuan, C.; Zhang, T. Environmental Implications of Nano-manufacturing. In Green Manufacturing; Dornfeld, D., Ed.; Springer: Boston, MA, USA, 2013. [Google Scholar] [CrossRef]
- Ribeiro, B.D.; Coelho, M.A.Z.; de Castro, A.M. Principles of Green Chemistry and White Biotechnology. In White Biotechnology for Sustainable Chemistry; Coelho, M.A.Z., Ribeiro, B.D., Eds.; The Royal Society of Chemistry: London, UK, 2016; pp. 1–8. [Google Scholar] [CrossRef]
- Li, X.; Xu, H.; Chen, Z.S.; Chen, G. Biosynthesis of nanoparticles by microorganisms and their applications. J. Nanomater. 2011, 2011. [Google Scholar] [CrossRef]
- Khan, T.; Abbas, S.; Fariq, A.; Yasmin, A. Microbes: Nature’s cell factories of nanoparticles synthesis. In Exploring the Realms of Nature for Nanosynthesis; Springer: Cham, Switzerland, 2018; pp. 25–50. [Google Scholar] [CrossRef]
- Hulkoti, N.I.; Taranath, T.C. Biosynthesis of nanoparticles using microbes—A review. Colloids Surf. B Biointerfaces 2014, 121, 474–483. [Google Scholar] [CrossRef] [PubMed]
- Kröger, N.; Poulsen, N. Diatoms—From cell wall biogenesis to nanotechnology. Annu. Rev. Genet. 2008, 42, 83–107. [Google Scholar] [CrossRef] [PubMed]
- Skeffington, A.W.; Scheffel, A. Exploiting algal mineralization for nanotechnology: Bringing coccoliths to the fore. Curr. Opin. Biotechnol. 2018, 49, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Yan, L.; Da, H.; Zhang, S.; Lopez, V.M.; Wang, W. Bacterial magnetosome and its potential application. Microbiol. Res. 2017, 203, 19–28. [Google Scholar] [CrossRef]
- Gama, M.; Gatenholm, P.; Klemm, D. (Eds.) Bacterial Nanocellulose: A sophisticated Multifunctional Material; CRC Press: Boca Raton, FL, USA, 2012. [Google Scholar]
- Nwodo, U.; Green, E.; Okoh, A. Bacterial exopolysaccharides: Functionality and prospects. Int. J. Mol. Sci. 2012, 13, 14002–14015. [Google Scholar] [CrossRef]
- Malvankar, N.S.; Lovley, D.R. Microbial nanowires: A new paradigm for biological electron transfer and bioelectronics. ChemSusChem 2012, 5, 1039–1046. [Google Scholar] [CrossRef]
- Prasad, R.; Pandey, R.; Barman, I. Engineering tailored nanoparticles with microbes: Quo vadis? Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 2016, 8, 316–330. [Google Scholar] [CrossRef]
- Dragone, R.; Grasso, G.; Muccini, M.; Toffanin, S. Portable bio/chemosensoristic devices: Innovative systems for environmental health and food safety diagnostics. Front. Public Health 2017, 5, 80. [Google Scholar] [CrossRef] [PubMed]
- Petros, R.A.; DeSimone, J.M. Strategies in the design of nanoparticles for therapeutic applications. Nature Rev. Drug Discov. 2010, 9, 615. [Google Scholar] [CrossRef] [PubMed]
- Kiessling, F.; Mertens, M.E.; Grimm, J.; Lammers, T. Nanoparticles for imaging: Top or flop? Radiology 2014, 273, 10–28. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.; Yang, L.; Zhang, B.; Liu, J. Extracellular biosynthesis and transformation of selenium nanoparticles and application in H2O2 biosensor. Colloids Surf. B Biointerfaces 2010, 80, 94–102. [Google Scholar] [CrossRef] [PubMed]
- Du, L.; Jiang, H.; Liu, X.; Wang, E. Biosynthesis of gold nanoparticles assisted by Escherichia coli DH5α and its application on direct electrochemistry of hemoglobin. Electrochem. Commun. 2007, 9, 1165–1170. [Google Scholar] [CrossRef]
- Ramya, S.; Shanmugasundaram, T.; Balagurunathan, R. Biomedical potential of actinobacterially synthesized selenium nanoparticles with special reference to anti-biofilm, anti-oxidant, wound healing, cytotoxic and anti-viral activities. J. Trace Elem. Med. Biol. 2015, 32, 30–39. [Google Scholar] [CrossRef]
- Bao, H.; Lu, Z.; Cui, X.; Qiao, Y.; Guo, J.; Anderson, J.M.; Li, C.M. Extracellular microbial synthesis of biocompatible CdTe quantum dots. Acta Biomater. 2010, 6, 3534–3541. [Google Scholar] [CrossRef]
- Suresh, A.K.; Pelletier, D.A.; Wang, W.; Broich, M.L.; Moon, J.W.; Gu, B.; Allison, D.P.; Joy, D.C.; Phelps, T.J.; Doktycz, M.J. Biofabrication of discrete spherical gold nanoparticles using the metal-reducing bacterium Shewanella oneidensis. Acta Biomater. 2011, 7, 2148–2152. [Google Scholar] [CrossRef]
- Suresh, A.K.; Pelletier, D.A.; Wang, W.; Moon, J.W.; Gu, B.; Mortensen, N.P.; David, P.; Allison, D.C.; Joy, C.; Phelps, T.J.; et al. Silver nanocrystallites: Biofabrication using Shewanella oneidensis, and an evaluation of their comparative toxicity on gram-negative and gram-positive bacteria. Environ. Sci. Technol. 2010, 44, 5210–5215. [Google Scholar] [CrossRef]
- Manivasagan, P.; Alam, M.S.; Kang, K.H.; Kwak, M.; Kim, S.K. Extracellular synthesis of gold bionanoparticles by Nocardiopsis sp. and evaluation of its antimicrobial, antioxidant and cytotoxic activities. Bioprocess Biosyst. Eng. 2015, 38, 1167–1177. [Google Scholar] [CrossRef]
- Roychoudhury, P.; Gopal, P.K.; Paul, S.; Pal, R. Cyanobacteria assisted biosynthesis of silver nanoparticles—A potential antileukemic agent. J. Appl. Phycol. 2016, 28, 3387–3394. [Google Scholar] [CrossRef]
- Al-Dhabi, N.; Mohammed Ghilan, A.K.; Arasu, M. Characterization of silver nanomaterials derived from marine Streptomyces sp. al-dhabi-87 and its in vitro application against multidrug resistant and extended-spectrum beta-lactamase clinical pathogens. Nanomaterials 2018, 8, 279. [Google Scholar] [CrossRef] [PubMed]
- Shao, W.; Liu, H.; Liu, X.; Sun, H.; Wang, S.; Zhang, R. pH-responsive release behavior and anti-bacterial activity of bacterial cellulose-silver nanocomposites. Int. J. Biol. Macromol. 2015, 76, 209–217. [Google Scholar] [CrossRef] [PubMed]
- Torres, S.K.; Campos, V.L.; León, C.G.; Rodríguez-Llamazares, S.M.; Rojas, S.M.; Gonzalez, M.; Mondaca, M.A. Biosynthesis of selenium nanoparticles by Pantoea agglomerans and their antioxidant activity. J. Nanopart. Res. 2012, 14, 1236. [Google Scholar] [CrossRef]
- Kalishwaralal, K.; Banumathi, E.; Pandian, S.R.K.; Deepak, V.; Muniyandi, J.; Eom, S.H.; Gurunathan, S. Silver nanoparticles inhibit VEGF induced cell proliferation and migration in bovine retinal endothelial cells. Colloids Surf. B Biointerfaces 2009, 73, 51–57. [Google Scholar] [CrossRef]
- Kalishwaralal, K.; Deepak, V.; Pandian, S.R.K.; Kottaisamy, M.; BarathManiKanth, S.; Kartikeyan, B.; Gurunathan, S. Biosynthesis of silver and gold nanoparticles using Brevibacterium casei. Colloids Surf. B Biointerfaces 2010, 77, 257–262. [Google Scholar] [CrossRef]
- Ahmad, M.S.; Yasser, M.M.; Sholkamy, E.N.; Ali, A.M.; Mehanni, M.M. Anticancer activity of biostabilized selenium nanorods synthesized by Streptomyces bikiniensis strain Ess_amA-1. Int. J. Nanomed. 2015, 10, 3389. [Google Scholar] [CrossRef]
- Gurunathan, S.; Han, J.W.; Eppakayala, V.; Kim, J.H. Green synthesis of graphene and its cytotoxic effects in human breast cancer cells. Int. J. Nanomed. 2013, 8, 1015. [Google Scholar] [CrossRef]
- Golmohammadi, H.; Morales-Narvaez, E.; Naghdi, T.; Merkoci, A. Nanocellulose in sensing and biosensing. Chem. Mater. 2017, 29, 5426–5446. [Google Scholar] [CrossRef]
- Saxena, I.M.; Dandekar, T.; Brown, R.M. Mechanisms in cellulose biosynthesis. Mol. Biol. 2004. Available online: http://www.esf.edu/outreach/pd/2000/cellulose/saxena.pdf (accessed on 10 November 2019).
- Pourreza, N.; Golmohammadi, H.; Naghdi, T.; Yousefi, H. Green in-Situ Synthesized Silver Nanoparticles Embedded in Bacterial Cellulose Nanopaper as a Bionanocomposite Plasmonic Sensor. Biosens. Bioelectron. 2015, 74, 353–359. [Google Scholar] [CrossRef] [PubMed]
- Morales-Narváez, E.; Golmohammadi, H.; Naghdi, T.; Yousefi, H.; Kostiv, U.; Horák, D.; Pourreza, N.; Merkoçi, A. Nanopaper as an Optical Sensing Platform. ACS Nano 2015, 9, 7296–7305. [Google Scholar] [CrossRef]
- Li, C.; Zhou, L.; Yang, H.; Lv, R.; Tian, P.; Li, X.; Zhang, Y.; Chen, Z.; Lin, F. Self-assembled exopolysaccharide nanoparticles for bioremediation and green synthesis of noble metal nanoparticles. ACS Appl. Mater. Interfaces 2017, 9, 22808–22818. [Google Scholar] [CrossRef] [PubMed]
- Simonte, F.; Sturm, G.; Gescher, J.; Sturm-Richter, K. Extracellular electron transfer and biosensors. In Bioelectrosynthesis; Springer: Cham, Switzerland, 2017; pp. 15–38. [Google Scholar] [CrossRef]
- Liu, X.; Wang, S.; Xu, A.; Zhang, L.; Liu, H.; Ma, L.Z. Biological synthesis of high-conductive pili in aerobic bacterium Pseudomonas aeruginosa. Appl. Microbiol. Biotechnol. 2019, 103, 1535–1544. [Google Scholar] [CrossRef] [PubMed]
- Sure, S.; Torriero, A.A.; Gaur, A.; Li, L.H.; Chen, Y.; Tripathi, C.; Adholeya, A.; Ackland, M.L.; Kochar, M. Inquisition of Microcystis aeruginosa and Synechocystis nanowires: Characterization and modelling. Antonie Van Leeuwenhoek 2015, 108, 1213–1225. [Google Scholar] [CrossRef] [PubMed]
- Leung, K.M.; Wanger, G.; El-Naggar, M.Y.; Gorby, Y.; Southam, G.; Lau, W.M.; Yang, J. Shewanella oneidensis MR-1 bacterial nanowires exhibit p-type, tunable electronic behavior. Nano Lett. 2013, 13, 2407–2411. [Google Scholar] [CrossRef]
- Davila, D.; Esquivel, J.P.; Sabate, N.; Mas, J. Silicon-based microfabricated microbial fuel cell toxicity sensor. Biosens. Bioelectron. 2011, 26, 2426–2430. [Google Scholar] [CrossRef]
- Wang, X.; Gao, N.; Zhou, Q. Concentration responses of toxicity sensor with Shewanella oneidensis MR-1 growing in bioelectrochemical systems. Biosens. Bioelectron. 2013, 43, 264–267. [Google Scholar] [CrossRef]
- Webster, D.P.; TerAvest, M.A.; Doud, D.F.; Chakravorty, A.; Holmes, E.C.; Radens, C.M.; Sureka, S.; Gralnick, J.A.; Angenent, L.T. An arsenic-specific biosensor with genetically engineered Shewanella oneidensis in a bioelectrochemical system. Biosens. Bioelectron. 2014, 62, 320–324. [Google Scholar] [CrossRef]
- Vargas, G.; Cypriano, J.; Correa, T.; Leão, P.; Bazylinski, D.A.; Abreu, F. Applications of magnetotactic bacteria, magnetosomes and magnetosome crystals in biotechnology and nanotechnology: Mini-review. Molecules 2018, 23, 2438. [Google Scholar] [CrossRef]
- Boucher, M.; Geffroy, F.; Prévéral, S.; Bellanger, L.; Selingue, E.; Adryanczyk-Perrier, G.; Péan, M.; Lefèvre, C.T.; Pignol, D.; Ginet, N.; et al. Genetically tailored magnetosomes used as MRI probe for molecular imaging of brain tumor. Biomaterials 2017, 121, 167–178. [Google Scholar] [CrossRef] [PubMed]
- Alphandery, E.; Faure, S.; Seksek, O.; Guyot, F.; Chebbi, I. Chains of magnetosomes extracted from AMB-1 magnetotactic bacteria for application in alternative magnetic field cancer therapy. ACS Nano 2011, 5, 6279–6296. [Google Scholar] [CrossRef] [PubMed]
- Roda, A.; Cevenini, L.; Borg, S.; Michelini, E.; Calabretta, M.M.; Schüler, D. Bioengineered bioluminescent magnetotactic bacteria as a powerful tool for chip-based whole-cell biosensors. Lab Chip 2013, 13, 4881–4889. [Google Scholar] [CrossRef][Green Version]
- Luo, Q.Y.; Lin, Y.; Li, Y.; Xiong, L.H.; Cui, R.; Xie, Z.X.; Pang, D.W. Nanomechanical analysis of yeast cells in CdSe quantum dot biosynthesis. Small 2014, 10, 699–704. [Google Scholar] [CrossRef] [PubMed]
- Wei, R. Biosynthesis of Au–Ag alloy nanoparticles for sensitive electrochemical determination of paracetamol. Int. J. Electrochem. Sci. 2017, 12, 9131–9140. [Google Scholar] [CrossRef]
- Zheng, D.; Hu, C.; Gan, T.; Dang, X.; Hu, S. Preparation and application of a novel vanillin sensor based on biosynthesis of Au–Ag alloy nanoparticles. Sens. Actuators B Chem. 2010, 148, 247–252. [Google Scholar] [CrossRef]
- Korbekandi, H.; Mohseni, S.; Mardani Jouneghani, R.; Pourhossein, M.; Iravani, S. Biosynthesis of silver nanoparticles using Saccharomyces cerevisiae. Artif. Cells Nanomed. Biotechnol. 2016, 44, 235–239. [Google Scholar] [CrossRef]
- Dhillon, G.S.; Brar, S.K.; Kaur, S.; Verma, M. Green approach for nanoparticle biosynthesis by fungi: Current trends and applications. Crit. Rev. Biotechnol. 2012, 32, 49–73. [Google Scholar] [CrossRef]
- Syed, A.; Saraswati, S.; Kundu, G.C.; Ahmad, A. Biological synthesis of silver nanoparticles using the fungus Humicola sp. and evaluation of their cytoxicity using normal and cancer cell lines. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2013, 114, 144–147. [Google Scholar] [CrossRef]
- Husseiny, S.M.; Salah, T.A.; Anter, H.A. Biosynthesis of size-controlled silver nanoparticles by Fusarium oxysporum, their antibacterial and antitumor activities. Beni Suef Univ. J. Basic Appl. Sci. 2015, 4, 225–231. [Google Scholar] [CrossRef]
- Saravanakumar, K.; Chelliah, R.; MubarakAli, D.; Jeevithan, E.; Oh, D.H.; Kathiresan, K.; Wang, M.H. Fungal enzyme-mediated synthesis of chitosan nanoparticles and its biocompatibility, antioxidant and bactericidal properties. Int. J. Biol. Macromol. 2018, 118, 1542–1549. [Google Scholar] [CrossRef] [PubMed]
- Mishra, A.; Tripathy, S.K.; Wahab, R.; Jeong, S.H.; Hwang, I.; Yang, Y.B.; Kim, Y.S.; Shin, H.S.; Yun, S.I. Microbial synthesis of gold nanoparticles using the fungus Penicillium brevicompactum and their cytotoxic effects against mouse mayo blast cancer C 2 C 12 cells. Appl. Microbiol. Biotechnol. 2011, 92, 617–630. [Google Scholar] [CrossRef] [PubMed]
- Elsoud, M.M.A.; Al-Hagar, O.E.; Abdelkhalek, E.S.; Sidkey, N.M. Synthesis and investigations on tellurium myconanoparticles. Biotechnol. Rep. 2018, 18, e00247. [Google Scholar] [CrossRef] [PubMed]
- Uddandarao, P.; Balakrishnan, R.M.; Ashok, A.; Swarup, S.; Sinha, P. Bioinspired ZnS: Gd Nanoparticles Synthesized from an Endophytic Fungi Aspergillus flavus for Fluorescence-Based Metal Detection. Biomimetics 2019, 4, 11. [Google Scholar] [CrossRef]
- Priyanka, U.; Akshay Gowda, K.M.; Elisha, M.G.; Nitish, N. Biologically synthesized PbS nanoparticles for the detection of arsenic in water. Int. Biodeterior. Biodegrad. 2017, 119, 78–86. [Google Scholar] [CrossRef]
- Dahoumane, S.A.; Mechouet, M.; Alvarez, F.J.; Agathos, S.N.; Jeffryes, C. Microalgae: An outstanding tool in nanotechnology. Bionatura 2016, 1, 196–201. [Google Scholar] [CrossRef]
- Senapati, S.; Syed, A.; Moeez, S.; Kumar, A.; Ahmad, A. Intracellular synthesis of gold nanoparticles using alga Tetraselmis kochinensis. Mater. Lett. 2012, 79, 116–118. [Google Scholar] [CrossRef]
- Jena, J.; Pradhan, N.; Nayak, R.R.; Dash, B.P.; Sukla, L.B.; Panda, P.K.; Mishra, B.K. Microalga Scenedesmus sp.: A potential low-cost green machine for silver nanoparticle synthesis. J. Microbiol. Biotechnol. 2014, 24, 522–533. [Google Scholar] [CrossRef]
- Öztürk, B.Y. Intracellular and extracellular green synthesis of silver nanoparticles using Desmodesmus sp.: Their Antibacterial and antifungal effects. Caryol. Int. J. Cytol. Cytosystem. Cytogenet. 2019, 72, 29–43. [Google Scholar] [CrossRef]
- Fuhrmann, T.; Landwehr, S.; El Rharbi-Kucki, M.; Sumper, M. Diatoms as living photonic crystals. Appl. Phys. B 2004, 78, 257–260. [Google Scholar] [CrossRef]
- Losic, D.; Mitchell, J.G.; Voelcker, N.H. Diatomaceous lessons in nanotechnology and advanced materials. Adv. Mater. 2009, 21, 2947–2958. [Google Scholar] [CrossRef]
- Aguirre, L.E.; Ouyang, L.; Elfwing, A.; Hedblom, M.; Wulff, A.; Inganäs, O. Diatom frustules protect DNA from ultraviolet light. Sci. Rep. 2018, 8, 5138. [Google Scholar] [CrossRef] [PubMed]
- Mazumder, N.; Gogoi, A.; Kalita, R.D.; Ahmed, G.A.; Buragohain, A.K.; Choudhury, A. Luminescence studies of fresh water diatom frustules. Indian J. Phys. 2010, 84, 665–669. [Google Scholar] [CrossRef]
- LeDuff, P.; Roesijadi, G.; Rorrer, G.L. Micro-photoluminescence of single living diatom cells. Luminescence 2016, 31, 1379–1383. [Google Scholar] [CrossRef]
- Ragni, R.; Cicco, S.R.; Vona, D.; Farinola, G.M. Multiple routes to smart nanostructured materials from diatom microalgae: A chemical perspective. Adv. Mater. 2018, 30, 1704289. [Google Scholar] [CrossRef]
- De Stefano, L.; Rotiroti, L.; De Stefano, M.; Lamberti, A.; Lettieri, S.; Setaro, A.; Maddalena, P. Marine diatoms as optical biosensors. Biosens. Bioelectron. 2009, 24, 1580–1584. [Google Scholar] [CrossRef]
- Gale, D.K.; Gutu, T.; Jiao, J.; Chang, C.H.; Rorrer, G.L. Photoluminescence detection of biomolecules by antibody-functionalized diatom biosilica. Adv. Funct. Mater. 2009, 19, 926–933. [Google Scholar] [CrossRef]
- Lin, K.C.; Kunduru, V.; Bothara, M.; Rege, K.; Prasad, S.; Ramakrishna, B.L. Biogenic nanoporous silica-based sensor for enhanced electrochemical detection of cardiovascular biomarkers proteins. Biosens. Bioelectron. 2010, 25, 2336–2342. [Google Scholar] [CrossRef]
- Li, A.; Cai, J.; Pan, J.; Wang, Y.; Yue, Y.; Zhang, D. Multi-layer hierarchical array fabricated with diatom frustules for highly sensitive bio-detection applications. J. Micromech. Microeng. 2014, 24, 025014. [Google Scholar] [CrossRef]
- Kamińska, A.; Sprynskyy, M.; Winkler, K.; Szymborski, T. Ultrasensitive SERS immunoassay based on diatom biosilica for detection of interleukins in blood plasma. Anal. Bioanal. Chem. 2017, 409, 6337–6347. [Google Scholar] [CrossRef]
- Zhen, L.; Ford, N.; Gale, D.K.; Roesijadi, G.; Rorrer, G.L. Photoluminescence detection of 2, 4, 6-trinitrotoluene (TNT) binding on diatom frustule biosilica functionalized with an anti-TNT monoclonal antibody fragment. Biosens. Bioelectron. 2016, 79, 742–748. [Google Scholar] [CrossRef] [PubMed]
- Pannico, M.; Rea, I.; Chandrasekaran, S.; Musto, P.; Voelcker, N.H.; De Stefano, L. Electroless gold-modified diatoms as surface-enhanced Raman scattering supports. Nanoscale Res. Lett. 2016, 11, 315. [Google Scholar] [CrossRef] [PubMed]
- Onesto, V.; Villani, M.; Coluccio, M.L.; Majewska, R.; Alabastri, A.; Battista, E.; Schirato, A.; Calestani, D.; Coppedé, N.; Cesarelli, M.; et al. Silica diatom shells tailored with Au nanoparticles enable sensitive analysis of molecules for biological, safety and environment applications. Nanoscale Res. Lett. 2018, 13, 94. [Google Scholar] [CrossRef] [PubMed]
- Gnanamoorthy, P.; Anandhan, S.; Prabu, V.A. Natural nanoporous silica frustules from marine diatom as a biocarrier for drug delivery. J. Porous Mater. 2014, 21, 789–796. [Google Scholar] [CrossRef]
- Lo Presti, M.; Ragni, R.; Vona, D.; Leone, G. In vivo doped biosilica from living Thalassiosira weissflogii diatoms with a triethoxysilyl functionalized red emitting fluorophore. Biomater. Soft Mater. 2018, 3, 1509–1517. [Google Scholar] [CrossRef]
- Terracciano, M.; De Stefano, L.; Rea, I. Diatoms Green Nanotechnology for Biosilica-Based Drug Delivery Systems. Pharmaceutics 2018, 10, 242. [Google Scholar] [CrossRef] [PubMed]
- Kong, X.; Chong, X.; Squire, K.; Wang, A.X. Microfluidic diatomite analytical devices for illicit drug sensing with ppb-Level sensitivity. Sens. Actuators B Chem. 2018, 259, 587–595. [Google Scholar] [CrossRef]
- Kong, X.; Yu, Q.; Li, E.; Wang, R.; Liu, Q.; Wang, A. Diatomite photonic crystals for facile on-chip chromatography and sensing of harmful ingredients from food. Materials 2018, 11, 539. [Google Scholar] [CrossRef]
- Panwar, V.; Dutta, T. Diatom Biogenic Silica as a Felicitous Platform for Biochemical Engineering: Expanding Frontiers. ACS Appl. Bio Mater. 2019, 2, 2295–2316. [Google Scholar] [CrossRef]
- Jena, J.; Pradhan, N.; Dash, B.P.; Panda, P.K.; Mishra, B.K. Pigment mediated biogenic synthesis of silver nanoparticles using diatom Amphora sp. and its antimicrobial activity. J. Saudi Chem. Soc. 2015, 19, 661–666. [Google Scholar] [CrossRef]
- Chetia, L.; Kalita, D.; Ahmed, G.A. Synthesis of Ag nanoparticles using diatom cells for ammonia sensing. Sens. Bio Sens. Res. 2017, 16, 55–61. [Google Scholar] [CrossRef]
- Kim, S.H.; Nam, O.; Jin, E.; Gu, M.B. A new coccolith modified electrode-based biosensor using a cognate pair of aptamers with sandwich-type binding. Biosens. Bioelectron. 2019, 123, 160–166. [Google Scholar] [CrossRef] [PubMed]
- Luo, C.H.; Shanmugam, V.; Yeh, C.S. Nanoparticle biosynthesis using unicellular and subcellular supports. NPG Asia Mater. 2015, 7, e209. [Google Scholar] [CrossRef]
- Narayanan, K.B.; Sakthivel, N. Biological synthesis of metal nanoparticles by microbes. Adv. Colloid Interface Sci. 2010, 156, 1–13. [Google Scholar] [CrossRef]
- Voeikova, T.A.; Zhuravliova, O.A.; Bulushova, N.V.; Veiko, V.P.; Ismagulova, T.T.; Lupanova, T.N.; Shaitan, K.V.; Debabov, V.G. The “Protein Corona” of Silver-Sulfide Nanoparticles Obtained Using Gram-Negative and-Positive Bacteria. Mol. Genet. Microbiol. Virol. 2017, 32, 204–211. [Google Scholar] [CrossRef]
- Jacek, P.; Dourado, F.; Gama, M.; Bielecki, S. Molecular aspects of bacterial nanocellulose biosynthesis. Microb. Biotechnol. 2019, 12, 633–649. [Google Scholar] [CrossRef]
- Ayano, H.; Kuroda, M.; Soda, S.; Ike, M. Effects of culture conditions of Pseudomonas aeruginosa strain RB on the synthesis of CdSe nanoparticles. J. Biosci. Bioeng. 2015, 119, 440–445. [Google Scholar] [CrossRef]
- Gurunathan, S.; Kalishwaralal, K.; Vaidyanathan, R.; Venkataraman, D.; Pandian, S.R.K.; Muniyandi, J.; Hariharana, N.; Eom, S.H. Biosynthesis, purification and characterization of silver nanoparticles using Escherichia coli. Colloids Surf. B Biointerfaces 2009, 74, 328–335. [Google Scholar] [CrossRef]
- Fayaz, A.M.; Balaji, K.; Kalaichelvan, P.T.; Venkatesan, R. Fungal based synthesis of silver nanoparticles—An effect of temperature on the size of particles. Colloids Surf. B Biointerfaces 2009, 74, 123–126. [Google Scholar] [CrossRef]
- Pimprikar, P.S.; Joshi, S.S.; Kumar, A.R.; Zinjarde, S.S.; Kulkarni, S.K. Influence of biomass and gold salt concentration on nanoparticle synthesis by the tropical marine yeast Yarrowia lipolytica NCIM 3589. Colloids Surf. B Biointerfaces 2009, 74, 309–316. [Google Scholar] [CrossRef]
- Ramanathan, R.; O’Mullane, A.P.; Parikh, R.Y.; Smooker, P.M.; Bhargava, S.K.; Bansal, V. Bacterial kinetics-controlled shape-directed biosynthesis of silver nanoplates using Morganella psychrotolerans. Langmuir 2010, 27, 714–719. [Google Scholar] [CrossRef] [PubMed]
- Su, Y.; Lundholm, N.; Ellegaard, M. Effects of abiotic factors on the nanostructure of diatom frustules—Ranges and variability. Appl. Microbiol. Biotechnol. 2018, 102, 5889–5899. [Google Scholar] [CrossRef] [PubMed]
- Townley, H.E.; Woon, K.L.; Payne, F.P.; White-Cooper, H.; Parker, A.R. Modification of the physical and optical properties of the frustule of the diatom Coscinodiscus wailesii by nickel sulfate. Nanotechnology 2007, 18, 295101. [Google Scholar] [CrossRef]
- Jeffryes, C.; Gutu, T.; Jiao, J.; Rorrer, G.L. Metabolic insertion of nanostructured TiO2 into the patterned biosilica of the diatom Pinnularia sp. by a two-stage bioreactor cultivation process. ACS Nano 2008, 2, 2103–2112. [Google Scholar] [CrossRef]
- Lang, Y.; Del Monte, F.; Rodriguez, B.J.; Dockery, P.; Finn, D.P.; Pandit, A. Integration of TiO2 into the diatom Thalassiosira weissflogii during frustule synthesis. Sci. Rep. 2013, 3, 3205. [Google Scholar] [CrossRef]
- Rorrer, G.L.; Chang, C.H.; Liu, S.H.; Jeffryes, C.; Jiao, J.; Hedberg, J.A. Biosynthesis of silicon–germanium oxide nanocomposites by the marine diatom Nitzschia frustulum. J. Nanosci. Nanotechnol. 2005, 5, 41–49. [Google Scholar] [CrossRef]
- Qin, T.; Gutu, T.; Jiao, J.; Chang, C.H.; Rorrer, G.L. Biological fabrication of photoluminescent nanocomb structures by metabolic incorporation of germanium into the biosilica of the diatom Nitzschia frustulum. ACS Nano 2008, 2, 1296–1304. [Google Scholar] [CrossRef]
- Jeffryes, C.; Gutu, T.; Jiao, J.; Rorrer, G.L. Two-stage photobioreactor process for the metabolic insertion of nanostructured germanium into the silica microstructure of the diatom Pinnularia sp. Mater. Sci. Eng. C 2008, 28, 10–118. [Google Scholar] [CrossRef]
- Jeffryes, C.; Solanki, R.; Rangineni, Y.; Wang, W.; Chang, C.H.; Rorrer, G.L. Electroluminescence and photoluminescence from nanostructured diatom frustules containing metabolically inserted germanium. Adv. Mater. 2008, 20, 2633–2637. [Google Scholar] [CrossRef]
- Magnabosco, G.; Hauzer, H.; Fermani, S.; Calvaresi, M.; Corticelli, F.; Christian, M.; Albonetti, C.; Morandi, V.; Ere, J.; Falini, G. Bionics synthesis of magnetic calcite skeletal structure through living foraminifera. Mater. Horiz. 2019, 6, 1862–1867. [Google Scholar] [CrossRef]
- Zhang, R.; Shao, M.; Han, X.; Wang, C.; Li, Y.; Hu, B.; Pang, D.; Xie, Z. ATP synthesis in the energy metabolism pathway: A new perspective for manipulating CdSe quantum dots biosynthesized in Saccharomyces cerevisiae. Int. J. Nanomed. 2017, 12, 3865. [Google Scholar] [CrossRef] [PubMed]
- Park, T.J.; Lee, S.Y.; Heo, N.S.; Seo, T.S. In vivo synthesis of diverse metal nanoparticles by recombinant Escherichia coli. Angew. Chem. Int. Ed. 2010, 49, 7019–7024. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.; Park, T.J.; Lee, D.C.; Lee, S.Y. Recombinant Escherichia coli as a biofactory for various single-and multi-element nanomaterials. Proc. Natl. Acad. Sci. USA 2018, 115, 5944–5949. [Google Scholar] [CrossRef] [PubMed]
- Mi, C.; Wang, Y.; Zhang, J.; Huang, H.; Xu, L.; Wang, S.; Fang, X.; Fang, J.; Mao, C.; Xu, S. Biosynthesis and characterization of CdS quantum dots in genetically engineered Escherichia coli. J. Biotechnol. 2011, 153, 125–132. [Google Scholar] [CrossRef]
- Taweecheep, P.; Naloka, K.; Matsutani, M.; Yakushi, T.; Matsushita, K.; Theeragool, G. Superfine bacterial nanocellulose produced by reverse mutations in the bcsC gene during adaptive breeding of Komagataeibacter oboediens. Carbohyd. Polym. 2019, 115243. [Google Scholar] [CrossRef]
- Tan, Y.; Adhikari, R.Y.; Malvankar, N.S.; Pi, S.; Ward, J.E.; Woodard, T.L.; Nevin, K.P.; Xia, Q.; Tuominen, M.T.; Lovley, D.R. Synthetic biological protein nanowires with high conductivity. Small 2016, 12, 4481–4485. [Google Scholar] [CrossRef]
- Delalat, B.; Sheppard, V.C.; Ghaemi, S.R.; Rao, S.; Prestidge, C.A.; McPhee, G.; Rogers, M.-L.; Donoghue, J.F.; Pillay, V.; Johns, T.G.; et al. Targeted drug delivery using genetically engineered diatom biosilica. Nat. Commun. 2015, 6, 8791. [Google Scholar] [CrossRef]
- Görlich, S.; Pawolski, D.; Zlotnikov, I.; Kröger, N. Control of biosilica morphology and mechanical performance by the conserved diatom gene Silicanin-1. Commun. Biol. 2019, 2, 245. [Google Scholar] [CrossRef]
- Nam, O.; Park, J.M.; Lee, H.; Jin, E. De novo transcriptome profile of coccolithophorid alga Emiliania huxleyi CCMP371 at different calcium concentrations with proteome analysis. PLoS ONE 2019, 14, e0221938. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).