Patterns of Genetic Diversity and Implications for In Situ Conservation of Wild Celery (Apium graveolens L. ssp. graveolens)
Abstract
:1. Introduction
2. Results
3. Discussion
4. Material and Methods
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- AGMRC (Agricultural Marketing Resource Center). Available online: http://www.agmrc.org/ (accessed on 16 February 2017).
- EU (European Union); EuroStat. Your Key to European Statistics. Available online: http://ec.europa.eu/eurostat/web/main/home (accessed on 16 February 2017).
- Schultze-Motel, J. Rudolf Mansfelds Verzeichnis Landwirtschaftlich und Gärtnerisch Kultivierter Pflanzenarten, 2nd ed.; Akademie Verlag: Berlin, Germany, 1986; pp. 999–1001. ISBN 3-05-500-164-8. [Google Scholar]
- Solberg, S.Ø. Celery. In Leaf Medicinal Herbs: Botany, Chemistry, Postharvest Technology and Uses; Ambrose, D.C.P., Manickavasagan, A., Naik, R., Eds.; CAP International: Wallingford, UK, 2016; pp. 74–84. ISBN 9781780645599. [Google Scholar]
- Barbour, J.D. Vegetable Crops. Search for Arthropod Resistance in Genetic Resources. In Global Plant Genetic Resources for Insect-Resistant Crops, 1st ed.; Clement, S.L., Quisenberry, S.S., Eds.; CRC Press: Boca Raton, FL, USA, 1998; pp. 171–189. ISBN 0-84932695-8. [Google Scholar]
- Melchinger, A.E.; Lübberstedt, T. Abschlussbericht zum GFP-Forschungsvorhaben ghg 1/98 (97 HS 044). Sortendifferenzierung und Verwandtschaftsuntersuchungen bei Feldsalat (Valerianella locusta L.), Radies (Raphanus sativus L.) und Knollensellerie (Apium graveolens L.) mit Hilfe PCR-Gestützter Genetischer Marker; Final report of the GFP research project ghg 1/98 (97 HS 044). Variety differentiation and investigation of the relationships in corn salad (Valerianella locusta L.), radish (Raphanus sativus L.) and celeriac (Apium graveolens L.) using PCR-based genetic marker; Universität Hohenheim, Institut für Pflanzenzüchtung, Saatgutforschung und Populationsgenetik, Lehrstuhl Angewandte Genetik und Pflanzenzüchtung: Stuttgart, Germany, 2003. [Google Scholar]
- Torricelli, R.; Tiranti, B.; Spataro, G.; Castellini, G.; Albertini, E.; Falcinelli, M.; Negri, V. Differentiation and structure of an Italian landrace of celery (Apium graveolens L.): Inferences for on farm conservation. Genet. Resour. Crop Evol. 2013, 60, 995–1006. [Google Scholar] [CrossRef]
- Wang, S.; Yang, W.; Shen, H. Genetic diversity in Apium graveolens and related species revealed by SRAP and SSR markers. Sci. Hortic. 2011, 129, 1–8. [Google Scholar] [CrossRef]
- Acquadro, A.; Magurno, F.; Portis, E.; Lanteri, S. dbEST-derived microsatellite markers in celery (Apium graveolens L. var. dulce). Mol. Ecol. Notes 2006, 6, 1080–1082. [Google Scholar] [CrossRef]
- Fu, N.; Wang, Q.; Shen, H.-L. De Novo Assembly, Gene Annotation and Marker Development Using Illumina Paired-End Transcriptome Sequences in Celery (Apium graveolens L.). PLoS ONE 2013, 8, e57686. [Google Scholar] [CrossRef] [PubMed]
- Fu, N.; Wang, P.-Y.; Liu, X.-D.; Shen, H.-L. Use of EST-SSR Markers for Evaluating Genetic Diversity and Fingerprinting Celery (Apium graveolens L.) Cultivars. Molecules 2014, 19, 1939–1955. [Google Scholar] [CrossRef] [PubMed]
- EURISCO. The European Search Catalogue for Plant Genetic Resources. Available online: http://eurisco.ecpgr.org (accessed on 8 May 2018).
- Spoor, W.; Simmonds, N.W. Base-broadening: Introgression and incorporation. In Broadening the Genetic Base of Crop Production; Cooper, H.D., Spillane, C., Hodgkin, T., Eds.; CAP International: Wallingford, UK, 2001; pp. 71–79. ISBN 0-85199-411-3. [Google Scholar]
- Breiteneder, H.; Hoffmann-Sommergruber, K.; O’Riordain, G.; Susani, M.; Ahorn, H.; Ebner, C.; Kraft, D.; Scheiner, O. Molecular Characterization of Api g 1, the Major Allergen of Celery (Apium graveolens), and its Immunological and Structural Relationships to a Group of 17kDa Tree Pollen Allergens. Eur. J. Biochem. 1995, 233, 484–489. [Google Scholar] [CrossRef] [PubMed]
- Gadermaier, G.; Egger, M.; Girbl, T.; Erler, A.; Harrer, A.; Vejvar, E.; Liso, M.; Richter, K.; Zuidmeer, L.; Mari, A.; et al. Molecular characterization of Api g 2, a novel allergenic member of the lipid-transfer protein 1 family from celery stalks. Mol. Nutr. Food Res. 2011, 55, 568–577. [Google Scholar] [CrossRef] [PubMed]
- Vejvar, E.; Himly, M.; Briza, P.; Eichhorn, S.; Ebner, C.; Hemmer, W.; Ferreira, F.; Gadermaier, G. Allergenic relevance of nonspecific lipid transfer proteins 2: Identification and characterization of Api g 6 from celery tuber as representative of a novel IgE-binding protein family. Mol. Nutr. Food Res. 2013, 57, 2061–2070. [Google Scholar] [CrossRef] [PubMed]
- Ochoa, O.; Quiros, C.F. Apium wild species: Novel sources for resistance to late blight in celery. Plant Breed. 1989, 102, 317–321. [Google Scholar] [CrossRef]
- Diawara, M.M.; Trumble, J.; Quiros, C.F.; Millar, J.G. Resistance to Spodoptera exigua in Apium prostratum. Entomol. Exp. Appl. 1992, 64, 125–133. [Google Scholar] [CrossRef]
- Trumble, J.T.; Dercks, W.; Quiros, C.F.; Beier, R.C. Host plant resistance and linear furanocoumarin content of Apium accessions. J. Econ. Entomol. 1990, 88, 519–525. [Google Scholar] [CrossRef]
- Trumble, J.T.; Diawara, M.M.; Quiros, C.F. Breeding resistance in Apium graveolens to Liriomyza trifolii: Antibiosis and linear furanocoumarin content. Acta Hortic. 2000, 513, 29–38. [Google Scholar] [CrossRef]
- Constance, L.; Chuang, T.-L.; Bell, C.R. Chromosome numbers in Umbelliferae. V. Am. J. Bot. 1976, 63, 608–625. [Google Scholar] [CrossRef]
- IUCN (International Union for Conservation of Nature and Natural Resources). The IUCN Red List of Threatened Species. Available online: http://www.iucnredlist.org/details/164203/0 (accessed on 8 May 2018).
- Frese, L.; Bönisch, M.; Herden, T.; Zander, M.; Friesen, N. In-situ-Erhaltung von Wildselleriearten. Ein Beispiel für genetische Ressourcen von Wildlebenden Verwandten von Kulturarten (WVK-Arten). Nat. Prot. Landsc. Plan. 2018, 50, 155–163. [Google Scholar]
- Berg, C.; Dengler, J.; Abdank, A.; Isermann, M. Die Pflanzengesellschaften Mecklenburg-Vorpommerns und ihre Gefährdung; Textband; Landesamt für Umwelt, Naturschutz und Geologie Mecklenburg-Vorpommern, Weißdorn-Verlag: Jena, Germany, 2004; ISBN 3-936055-03-3. [Google Scholar]
- Preising, E.; Vahle, H.C.; Brandes, D.; Hofmeister, H.; Tüxen, J.; Weber, H.E. Die Pflanzengesellschaften Niedersachsens—Bestandsentwicklung, Gefährdung und Schutzprobleme Salzpflanzengesellschaften der Meeresküste und des Binnenlandes; Naturschutz und Landschaftspflege in Niedersachsen; Niedersächsische Landesverwaltungsamt—Fachbehörde für Naturschutz: Hannover, Germany, 1990; ISBN 392232150X. [Google Scholar]
- Bettinger, A. Die Binnensalzstellen in Lothringen, im Saarland und in Rheinland-Pfalz. In Binnensalzstellen Mitteleuropas. Internationale Tagung, Bad Frankenhausen, Germany, 8–10 September 2005; Thüringer Ministerium für Landwirtschaft, Naturschutz und Umwelt: Erfurt, Germany, 2007; pp. 143–148. [Google Scholar]
- Böttcher, H. Das EU-Life-Natur-Projekt, Erhaltung und Entwicklung der Binnensalzstellen Nordthüringens (2003–2008). In Binnensalzstellen Mitteleuropas. Internationale Tagung, Bad Frankenhausen, Germany, 8–10. September 2005; Thüringer Ministerium für Landwirtschaft, Naturschutz und Umwelt: Erfurt, Germany, 2007; pp. 54–62. [Google Scholar]
- Herrmann, A. Binnensalzstellen in Brandenburg—Verbreitung und Zustand salzbeeinflusster Lebensräume. In Binnensalzstellen Mitteleuropas. Internationale Tagung, Bad Frankenhausen, Germany, 8–10. September 2005; Thüringer Ministerium für Landwirtschaft, Naturschutz und Umwelt: Erfurt, Germany, 2007; pp. 135–142. [Google Scholar]
- Kell, S. Farmer’s Pride. Conserving Plant Diversity for Future Generations. Available online: http://www.farmerspride.eu/ (accessed on 8 May 2018).
- Maxted, N.; Avagyan, A.; Frese, L.; Iriondo, J.M.; Magos Brehm, J.; Singer, A.; Kell, S.P. ECPGR Concept for in situ Conservation of Crop Wild Relatives in Europe; European Cooperative Programme for Plant Genetic Resources: Rome, Italy, 2015. [Google Scholar]
- Maxted, N.; Hawkes, J.G.; Ford-Lloyd, B.V.; Williams, J.T. A practical model for in situ genetic conservation. In Plant Genetic Conservation: The in situ Approach; Maxted, N., Ford-Lloyd, B.V., Hawkes, J.G., Eds.; Kluwer Academic Publishers: Dordrecht, The Netherlands, 1997; pp. 339–364. ISBN 0-412-63400-7. [Google Scholar]
- Iriondo, J.M.; Maxted, N.; Kell, S.; Ford-Lloyd, B.V.; Lara-Romero, C.; Labokas, J.; Magos Brehm, J. Quality Standards for Genetic Reserve Conservation of Crop Wild Relatives. In Agrobiodiversity Conserving: Securing the Diversity of Crop Wild Relatives and Landraces; Maxted, N., Dulloo, M.E., Ford-Lloyd, B.V., Frese, L., Iriondo, J.M., Pinheiro de Carvalho, M.Â.A., Eds.; CAB International: Wallingford, UK, 2012; pp. 72–77. [Google Scholar]
- Kell, S.; Maxted, N.; Frese, L.; Iriondo, J.M. In situ conservation of crop wild relatives: A strategy for identifying priority genetic reserves sites. In Agrobiodiversity Conservation: Securing the Diversity of Crop Wild Relatives and Landraces; Maxted, N., Dulloo, M.E., Ford-Lloyd, B.V., Frese, L., Iriondo, J.M., Pinheiro de Carvalho, M.Â.A., Eds.; CAB International: Wallingford, UK, 2012; pp. 7–19. [Google Scholar]
- Manel, S.; Schwartz, M.K.; Luikart, G.; Taberlet, P. Landscape genetics: Combining landscape ecology and population genetics. Trends Ecol. Evol. 2003, 18, 189–197. [Google Scholar] [CrossRef]
- Hodel, R.G.J.; Segovia-Salcedo, M.C.; Landis, J.B.; Crowl, A.A.; Sun, M. The report of my death was an exaggeration: A review for researchers using microsatellites in the 21st century. Appl. Plant Sci. 2016, 4. [Google Scholar] [CrossRef] [PubMed]
- Arus, P.; Orton, T.J. Inheritance patterns and linkage relationships of eight genes of celery. J. Hered. 1984, 75, 11–14. [Google Scholar] [CrossRef]
- Huestis, G.M.; McGrath, J.M.; Quiros, C.F. Development of genetic markers in celery based on restriction fragment length polymorphisms. Theor. Appl. Genet. 1993, 85, 889–896. [Google Scholar] [CrossRef] [PubMed]
- Feng, K.; Hou, X.-L.; Li, M.-Y.; Jiang, Q.; Xu, Z.-S.; Liu, J.-X.; Xiong, A.-S. CeleryDB: A genomic database for celery. Database 2018, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Li, M.Y.; Wang, F.; Jiang, Q.; Ma, J.; Xiong, A.-S. Identification of SSRs and differentially expressed genes in two cultivars of celery (Apium graveolens L.) by deep transcriptome sequencing. Hortic. Res. 2014, 1. [Google Scholar] [CrossRef] [PubMed]
- Gillet, E.M.; Gregorius, H.-R. Measuring differentiation among populations at different levels of genetic integration. BMC Genet. 2008, 9, 60. [Google Scholar] [CrossRef] [PubMed]
- Gillet, E.M. DifferInt: Compositional differentiation among populations at three levels of genetic integration. Mol. Ecol. Resour. 2013, 13, 953–964. [Google Scholar] [CrossRef] [PubMed]
- Meynen, E.; Schmithüsen, J. Handbuch der Naturräumlichen Gliederung Deutschlands/Unter Mitwirkung des Zentralausschusses für Deutsche Landeskunde, 1953–1962; Bundesanstalt für Landeskunde und Raumforschung: Bad Godesberg, Germany, 1962. [Google Scholar]
- Gregorius, H.-R.; Gillet, E.M.; Ziehe, M. Measuring Differences of Trait Distributions between Populations. Biom. J. 2003, 45, 959–973. [Google Scholar] [CrossRef]
- Goudet, J. FSTAT (Version 1.2): A computer program to calculate F-statistics. J. Hered. 1995, 86, 485–486. [Google Scholar] [CrossRef]
- Jombart, T.; Devillard, S.; Balloux, F. Discriminant analysis of principal components: A new method for the analysis of genetically structured populations. BMC Genet. 2010, 11, 94. [Google Scholar] [CrossRef] [PubMed]
- Perrier, X.; Jacquemound-Collet, J.P. DARwin5, Version 5.0.158 Software. Available online: http://darwin.cirad.fr/darwin (accessed on 10 October 2010).
- Fievet, V.; Touzet, P.; Arnaud, J.-F.; Cuguen, J. Spatial analysis of nuclear and cytoplasmic DNA diversity in wild sea beet (Beta vulgaris ssp. maritima) populations: Do marine currents shape the genetic structure? Mol. Ecol. 2007, 16, 1847–1864. [Google Scholar] [PubMed]
- Petit, R.J.; El Mousadik, A.; Pons, O. Identifying populations for conservation on the basis of genetic markers. Conserv. Biol. 1998, 12, 844–855. [Google Scholar] [CrossRef]
- Zander, M.; Albrecht Daniel Thaer-Institut für Agrar- und Gartenbauwissenschaften, Urbane Ökophysiologie der Pflanzen, Humboldt-Universität zu Berlin, Berlin, Germany. Personal communication, 2018.
- Piorr, H.-P. Entwicklung der pflanzenbaulichen Flächennutzung in den Agrarlandschaften Deutschlands. In Biologische Vielfalt mit der Land- und Forstwirtschaft? Welling, P., Ed.; Reihe A: Angewandte Wissenschaft; Schriftenreihe des Bundesministeriums für Verbraucherschutz, Ernährung und Landwirtschaft: Bonn, Germany, 2002; pp. 127–135. [Google Scholar]
- Klotz, S.; Kühn, I.; Durka, W. BIOLFLOR—Eine Datenbank zu biologisch-ökologischen Merkmalen der Gefäßpflanzen in Deutschland. In Schriftenreihe für Vegetationskunde 38; Bundesamt für Naturschutz: Bad Godesberg, Germany, 2002. [Google Scholar]
- BMVEL (Bundesministerium für Verbraucherschutz, Ernährung und Landwirtschaft). Tiergenetische Ressourcen; Nationales Fachprogramm: Bonn, Germany, 2004. [Google Scholar]
- Wehling, P.; Scholz, M.; Ruge-Wehling, B.; Hackauf, B.; Frese, L. Anpassung landwirtschaftlicher Kulturarten an den Klimawandel—Optionen aus Sicht der Züchtungsforschung. J. Cultiv. Plants 2017, 69, 47–50. [Google Scholar] [CrossRef]
- Bilz, M.; Kell, S.; Maxted, N.; Lansdown, R.V. European Red List of Vascular Plants; Office of the European Union: Luxembourg, 2011. [Google Scholar]
- EU (European Union). Halbzeitbewertung der EU-Biodiversitätsstrategie. Natura 2000, 2016, 39. [Google Scholar]
- Parmesan, C.; Hanley, M.E. Plants and climate change: Complexities and surprises. Ann. Bot. 2015, 116, 849–864. [Google Scholar] [CrossRef] [PubMed]
- Araujo, M.B.; Cabeza, M.; Thuiller, W.; Hannah, L.; Williams, P.H. Would climate change drive species out of reserves? An assessment of existing reserve-selection methods. Glob. Chang. Biol. 2004, 10, 1618–1626. [Google Scholar] [CrossRef]
- Castaneda-Alvarez, N.P.; Khoury, C.K.; Achicanoy, H.A.; Bernau, V.; Dempewolf, H.; Eastwood, R.J.; Guarino, L.; Harker, R.H.; Jarvis, A.; Maxted, N.; et al. Global conservation priorities for crop wild relatives. Nat. Plants 2016, 2. [Google Scholar] [CrossRef] [PubMed]
- CBD (Convention on Biological Diversity). Convention on Biological Diversity; Convention on Biological Diversity: Montreal, QC, Canada, 1992. [Google Scholar]
- FAO (Food and Agriculture Organization). International Treaty on Plant Genetic Resources for Food and Agriculture. Available online: http://www.planttreaty.org/ (accessed on 16 February 2017).
- ECPGR (European Cooperative Programme for Plant Genetic Resources). The ECPGR Concept for in situ Conservation of Crop Wild Relatives in Europe. Available online: http://www.ecpgr.cgiar.org/working-groups/wild-species-conservation/ (accessed on 20 June 2017).
- CWRnl. Crop Wild Relatives (CWR) in the Netherlands. Available online: https://www.cwrnl.nl/nl/CWRnl/CWRspergewas/Apium-graveolens-L.htm (accessed on 8 May 2018).
- Aguirre-Gutiérrez, J.; van Treuren, R.; Hoekstra, R.; van Hintum, T.J.L. Crop wild relatives range shifts and conservation in Europe under climate change. Divers. Distrib. 2017, 23. [Google Scholar] [CrossRef]
- Hoekstra, R.; Centre for Genetic Resources, Droevendaalsesteeg 1, Building 107, 6708 PB Wageningen, The Netherlands. Personal communication, 2017.
- Kleinschmit, J.R.G.; Kownatzki, D.; Gregorius, H.-R. Adaptational characteristics of autochthonous populations—Consequences for provenance delineation. For. Ecol. Manag. 2004, 197, 213–224. [Google Scholar] [CrossRef]
- GBIF (Global Biological Information Facility). Available online: https://www.gbif.org/species/search (accessed on 8 August 2018).
- Nachtigall, N.; Bülow, L.; Schubert, J.; Frese, L. Development of SSR Markers for the Genus Patellifolia (Chenopodiaceae). Appl. Plant Sci. 2016, 4, 8. [Google Scholar] [CrossRef] [PubMed]
- Enders, M. Entwicklung und Anwendung Molekularer und Informatorischer Werkzeuge zum Genetischen Monitoring bei Wildrüben; Diplomarbeit im Fach Bioinformatik, Martin-Luther-Universität Halle-Wittenberg, Naturwissenschaftliche Fakultät III, Institut für Informatik: Halle, Germany, 2010. [Google Scholar]
- Nachtigall, M.; Frese, L.; Bönisch, M. Microsatellite Marker Data of Apium graveolens L. ssp. graveolens; Open Agrar Repositorium: Quedlinburg, Germany, 2018. [Google Scholar]
- R Core Team. R: A Language and Environment for Statistical Computing, version 3.5.1; R Foundation for Statistical Computing: Vienna, Austria, 2013; Available online: https://www.R-project.org (accessed on 8 August 2018).
- Jombart, T. Adegenet: A R package for the multivariate analysis of genetic markers. Bioinformatics 2008, 24, 1403–1405. [Google Scholar] [CrossRef] [PubMed]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar] [PubMed]
- Gillet, E.M. DifferInt Compositional Differentiation among Populations at Three Levels of Genetic Integration. User Manual. Available online: http://www.uni-goettingen.de/forstgenetik/differint (accessed on 17 November 2016).
- Slatkin, M. Linkage disequilibrium—Understanding the evolutionary past and mapping the medical future. Nat. Rev. Genet. 2008, 9, 477–485. [Google Scholar] [CrossRef] [PubMed]
 = Weser-Aller-Flachland,
= Weser-Aller-Flachland,  = Nordmecklenburgisches Flachland,
= Nordmecklenburgisches Flachland,  = Schleswig-Holsteinisches Hügelland,
= Schleswig-Holsteinisches Hügelland,  = Osthessisches Bergland,
= Osthessisches Bergland,  = Mecklenburgisch-Vorpommersches Küstengebiet,
= Mecklenburgisch-Vorpommersches Küstengebiet,  = Mecklenburgische Seenplatte,
= Mecklenburgische Seenplatte,  = Mittelbrandenburgische Platten und Niederungen,
= Mittelbrandenburgische Platten und Niederungen,  = Gäuplatten im Neckar- und Tauberland,
= Gäuplatten im Neckar- und Tauberland,  = Weser-Leinebergland,
= Weser-Leinebergland,  = Mitteldeutsches Schwarzerdegebiet,
= Mitteldeutsches Schwarzerdegebiet,  = Elbtalniederung,
= Elbtalniederung,  = Nördliches Harzvorland,
= Nördliches Harzvorland,  = Westfälische Tieflandsbucht,
= Westfälische Tieflandsbucht,  = Thüringer Becken und Randplatten.
= Thüringer Becken und Randplatten.
 = Weser-Aller-Flachland,
= Weser-Aller-Flachland,  = Nordmecklenburgisches Flachland,
= Nordmecklenburgisches Flachland,  = Schleswig-Holsteinisches Hügelland,
= Schleswig-Holsteinisches Hügelland,  = Osthessisches Bergland,
= Osthessisches Bergland,  = Mecklenburgisch-Vorpommersches Küstengebiet,
= Mecklenburgisch-Vorpommersches Küstengebiet,  = Mecklenburgische Seenplatte,
= Mecklenburgische Seenplatte,  = Mittelbrandenburgische Platten und Niederungen,
= Mittelbrandenburgische Platten und Niederungen,  = Gäuplatten im Neckar- und Tauberland,
= Gäuplatten im Neckar- und Tauberland,  = Weser-Leinebergland,
= Weser-Leinebergland,  = Mitteldeutsches Schwarzerdegebiet,
= Mitteldeutsches Schwarzerdegebiet,  = Elbtalniederung,
= Elbtalniederung,  = Nördliches Harzvorland,
= Nördliches Harzvorland,  = Westfälische Tieflandsbucht,
= Westfälische Tieflandsbucht,  = Thüringer Becken und Randplatten.
= Thüringer Becken und Randplatten.
| LabID | Population ID | Location | Ecogeographic Unit | No. of Analyzed Individuals | Arguments For or Against the Nomination as Candidate MAWPs |
|---|---|---|---|---|---|
| AgBW | BW-UB-20150728-0934 | Baden-Württemberg: Ubstadt-Weiher (samples from ex situ cultivation in Botanical Garden of KIT and Tübingen) | Gäuplatten im Neckar- und Tauberland | 30 | The only representative of the ecogeographic unit, most southern occurrence in Germany, site management exists already |
| AgSEHL | SH-SEHL-20150824-0900, -1000 | Schleswig-Holstein: Sehlendorf | Schleswig-Holsteinisches Hügelland | 30 | Adjustment of management would be required |
| AgEICH | SH-EICH-20150824-1000, -1010, -1100, -1110, -1120, -1200 | Schleswig-Holstein: Eichholzniederung | Schleswig-Holsteinisches Hügelland | 30 | Large site and population size, site management exists already |
| AgKREM | SH-KREM-20150902-1315, -1500, -1510, -1520 | Schleswig-Holstein: Neustadt | Schleswig-Holsteinisches Hügelland | 30 | Within the same ecogeographic unit AgEICH matches genetic reserves quality criteria better than AgKREM |
| AgFEH | SH-FEH-20150902-1000, -1010, -1020 | Schleswig-Holstein: Fehmarn | Schleswig-Holsteinisches Hügelland | 30 | Within the same ecogeographic unit AgEICH matches genetic reserves quality criteria better than AgFEH |
| AgOEH | SH-OEH-20150826-1200, 1300, -1320, -1330, -1340, -1350, -1400 | Schleswig-Holstein: Schleimündung | Schleswig-Holsteinisches Hügelland | 30 | Within the same ecogeographic unit AgEICH matches genetic reserves quality criteria better than AgOEH |
| AgBBG | Bbg-JÜ-20150727-1739 | Brandenburg: Gröben | Mittelbrandenburgische Platten und Niederungen | 30 | The only representative of the ecogeographic unit, site management exists already |
| AgFRI | ST-FRIED-20150621-1437 | Sachsen-Anhalt: Friedeburg | Mitteldeutsches Schwarzerdegebiet | 30 | Within the same ecogeographic unit AgHEC matches genetic reserves quality criteria better than AgFRI |
| AgWA | ST-WALBE-20150720-1304 | Sachsen-Anhalt: Walbeck | Weser-Aller-Flachland | 30 | The only representative of the natural region, highest deviation in genetic composition from the complement |
| AgHEC | ST-HECBE-20150709-0957, ST-HECSA-20150709-0854 | Sachsen-Anhalt: Hecklingen | Mitteldeutsches Schwarzerdegebiet | 30 | Represents the genetic composition of the complement best, site management exists already |
| AgZIE | ST-ZIELI-20150709-1223 | Sachsen-Anhalt: Zielitz | Elbtalniederung | 30 | The only representative of the ecogeographic unit |
| AgSUE | ST-SÜLLW-20150714-1054, ST-SÜLNO-20150714-1003 | Sachen-Anhalt: Sülldorf | Mitteldeutsches Schwarzerdegebiet | 30 | Large site and population size, site management exists already |
| AgROS | ST-ROSSL-20150702-1029 | Thüringen/Sachsen-Anhalt: Roßleben | Thüringer Becken und Randplatten | 30 | The only representative of the ecogeographic unit |
| AgBEN | ST-BENWÜ-20150702-0828 | Sachsen-Anhalt: Bennstedt | Mitteldeutsches Schwarzerdegebiet | 30 | Within the same ecogeographic unit AgHEC matches genetic reserves quality criteria better than AgBEN |
| AgWU | MV-WU-20150824-1100 | Mecklenburg-Vorpommern: Halbinsel Wustrow | Mecklenburgisch-Vorpommersches Küstengebiet | 30 | Large site and population, best representative of the ecogeographic unit |
| AgDA | MV-DA-20150901-0900 | Mecklenburg-Vorpommern: Dabitz | Mecklenburgisch-Vorpommersches Küstengebiet | 23 | Critically small population size. Within the same ecogeographic unit AgDZ matches genetic reserves quality criteria better than AgDA |
| AgDZ | MV-DSZ-20150903-1430 | Mecklenburg-Vorpommern: Dassow | Mecklenburgische Seenplatte | 30 | Best representative of the ecogeographic unit |
| AgUW | MV-UW-20150902-1000 | Mecklenburg-Vorpommern: Rostock | Mecklenburgisch-Vorpommersches Küstengebiet | 30 | Within the same ecogeographic unit AgDZ matches genetic reserves quality criteria better than AgUW |
| AgHEU | MV-HEU-20150921-1600 | Mecklenburg-Vorpommern: Bodden bei Rügen | Mecklenburgisch-Vorpommersches Küstengebiet | 30 | Access to population is complicated. Within the same ecogeographic unit AgDZ matches genetic reserves quality criteria better than AgHEU |
| AgG | MV-G-20150831-1200 | Mecklenburg-Vorpommern: Gristow | Nordostmecklenburgisches Flachland | 30 | The only representative of the ecogeographic unit |
| AgHID | MV-HiABs-20150912-1400 | Mecklenburg-Vorpommern: Hiddensee | Mecklenburgisch-Vorpommersches Küstengebiet | 30 | Within the same ecogeographic unit AgDZ matches genetic reserves quality criteria better than AgHID |
| AgSL | MV-Sü-20150902-1540 | Mecklenburg-Vorpommern: Sülten | Mecklenburgische Seenplatte | 20 | Critically small population size. Within the same ecogeographic unit AgDZ matches genetic reserves quality criteria better than AgSL |
| AgSK | NRW-SK-20180818-1530 | Nordrhein-Westfalen: Salzkotten | Westfälische Tieflandsbucht | 30 | The only representative of the ecogeographic unit, site management exists already |
| AgGOE | NI-GÖ-20150825-1000 | Niedersachsen: Nörten-Hardenberg | Weser-Leinebergland | 30 | Within the same ecogeographic unit AgHI matches genetic reserves quality criteria better than AgGOE |
| AgHI | NI-HI-20150824-1600 | Niedersachsen: Hildesheim | Weser-Leinebergland | 30 | Best representative of the ecogeographic unit |
| AgJX | NI-JX-20150724-1100 | Niedersachsen: Jerxheim | Nördliches Harzvorland | 30 | The only representative of the ecogeographic unit |
| AgHES | HE-BS-20150925-1020 | Hessen: Bad Salzhausen | Osthessisches Bergland | 27 | The only representative of the ecogeographic unit, site management exists already |
| Locus | Number of Individuals | Number of Alleles | Polymorphic Information Content (PIC) | Observed Heterozygosity (Ho) | Expected Heterozygosity (He) |
|---|---|---|---|---|---|
| ECMS01 (1) | 790 | 7 | 0.733 | 0.291 | 0.769 |
| ECMS11 | 790 | 9 | 0.774 | 0.523 | 0.801 |
| ECMS13 | 790 | 8 | 0.193 | 0.066 | 0.203 |
| ECMS02 | 790 | 8 | 0.434 | 0.128 | 0.502 |
| ECMS23 | 790 | 8 | 0.384 | 0.168 | 0.442 |
| ECMS39 | 790 | 13 | 0.803 | 0.167 | 0.826 |
| ECMS06 | 790 | 18 | 0.774 | 0.205 | 0.799 |
| ECMS09 | 790 | 8 | 0.457 | 0.101 | 0.556 |
| Fn09 (2) | 790 | 4 | 0.109 | 0.057 | 0.112 |
| Fn100 | 790 | 6 | 0.042 | 0.005 | 0.042 |
| Fn62 | 790 | 8 | 0.163 | 0.010 | 0.169 |
| QC28 (3) | 790 | 6 | 0.734 | 0.115 | 0.772 |
| QC43 | 790 | 18 | 0.641 | 0.075 | 0.667 |
| QC53 | 790 | 5 | 0.286 | 0.044 | 0.310 |
| QC75 | 790 | 7 | 0.432 | 0.963 | 0.538 |
| QC86 | 790 | 5 | 0.364 | 0.099 | 0.399 |
| LabID | AgBBG | AgBEN | AgBW | AgDA | AgDZ | AgEICH | AgFEH |
| FIS | 0.043 | −0.145 | 0.151 | 0.43 | 0.436 | 0.377 | −0.016 |
| LabID | AgFRI | AgG | AgGOE | AgHEC | AgHES | AgHEU | AgHI |
| FIS | 0.558 | −0.167 | 0.363 | 0.231 | 0.441 | −0.006 | 0.099 |
| LabID | AgHID | AgJX | AgKREM | AgOEH | AgROS | AgSEHL | AgSK |
| FIS | 0.39 | 0.174 | 0.338 | 0.444 | 0.265 | 0.508 | 0.054 |
| LabID | AgSL | AgSUE | AgUW | AgWA | AgWU | AgZIE | |
| FIS | 0.137 | 0.353 | 0.348 | −0.08 | 0.49 | 0.14 |
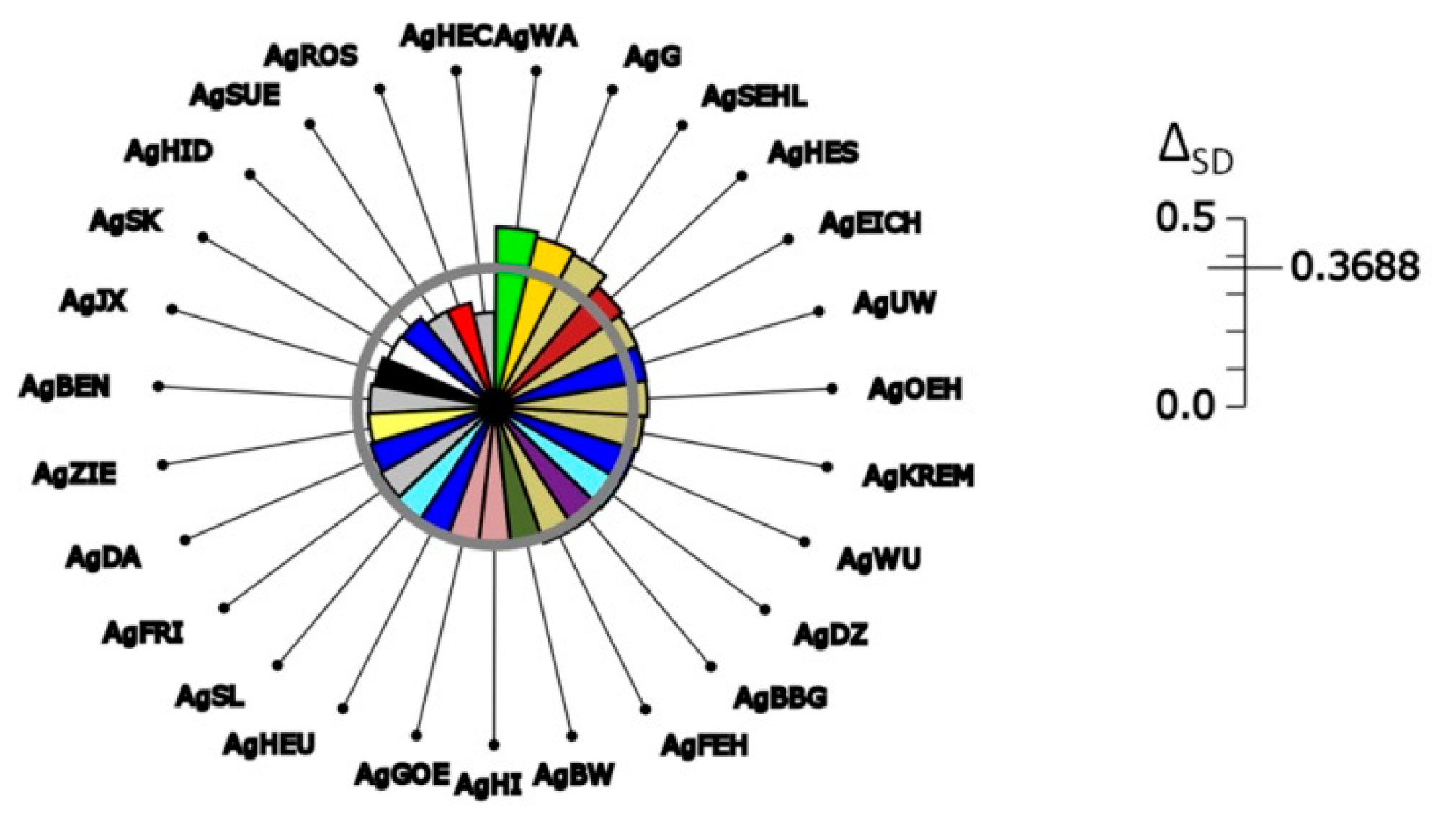
| Permutations | Observed ΔSD Values | Min | 0.95 Confidence Interval | Max | p Value |
|---|---|---|---|---|---|
| Alleles over individuals within occurrences | |||||
| Gene pool level | 0.3688 | not affected | |||
| Mean single-locus genotypes | 0.3771 | 0.3782 | 0.3790; 0.3810 | 0.3821 | 1.0000 **<- |
| Multi-locus genotypes | 0.4001 | 0.4060 | 0.4066; 0.4080 | 0.4085 | 1.0000 **<- |
| All individual genotypes among occurrences | |||||
| Gene pool level | 0.3688 | 0.0960 | 0.1001; 0.1102 | 0.1163 | 0.0000 **-> |
| Mean single-locus genotypes | 0.3771 | 0.1092 | 0.1129; 0.1227 | 0.1286 | 0.0000 **-> |
| Multi-locus genotypes | 0.4001 | 0.2252 | 0.2284; 0.2359 | 0.2415 | 0.0000 **-> |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Frese, L.; Bönisch, M.; Nachtigall, M.; Schirmak, U. Patterns of Genetic Diversity and Implications for In Situ Conservation of Wild Celery (Apium graveolens L. ssp. graveolens). Agriculture 2018, 8, 129. https://doi.org/10.3390/agriculture8090129
Frese L, Bönisch M, Nachtigall M, Schirmak U. Patterns of Genetic Diversity and Implications for In Situ Conservation of Wild Celery (Apium graveolens L. ssp. graveolens). Agriculture. 2018; 8(9):129. https://doi.org/10.3390/agriculture8090129
Chicago/Turabian StyleFrese, Lothar, Maria Bönisch, Marion Nachtigall, and Uta Schirmak. 2018. "Patterns of Genetic Diversity and Implications for In Situ Conservation of Wild Celery (Apium graveolens L. ssp. graveolens)" Agriculture 8, no. 9: 129. https://doi.org/10.3390/agriculture8090129
APA StyleFrese, L., Bönisch, M., Nachtigall, M., & Schirmak, U. (2018). Patterns of Genetic Diversity and Implications for In Situ Conservation of Wild Celery (Apium graveolens L. ssp. graveolens). Agriculture, 8(9), 129. https://doi.org/10.3390/agriculture8090129





