Breeding for Nutritional and Organoleptic Quality in Vegetable Crops: The Case of Tomato and Cauliflower
Abstract
1. Introduction
2. Breeding for Old and New Challenges in Vegetable Crops
3. Botanical Aspects of Tomato and Cauliflower
4. Quality in Tomato and Cauliflower
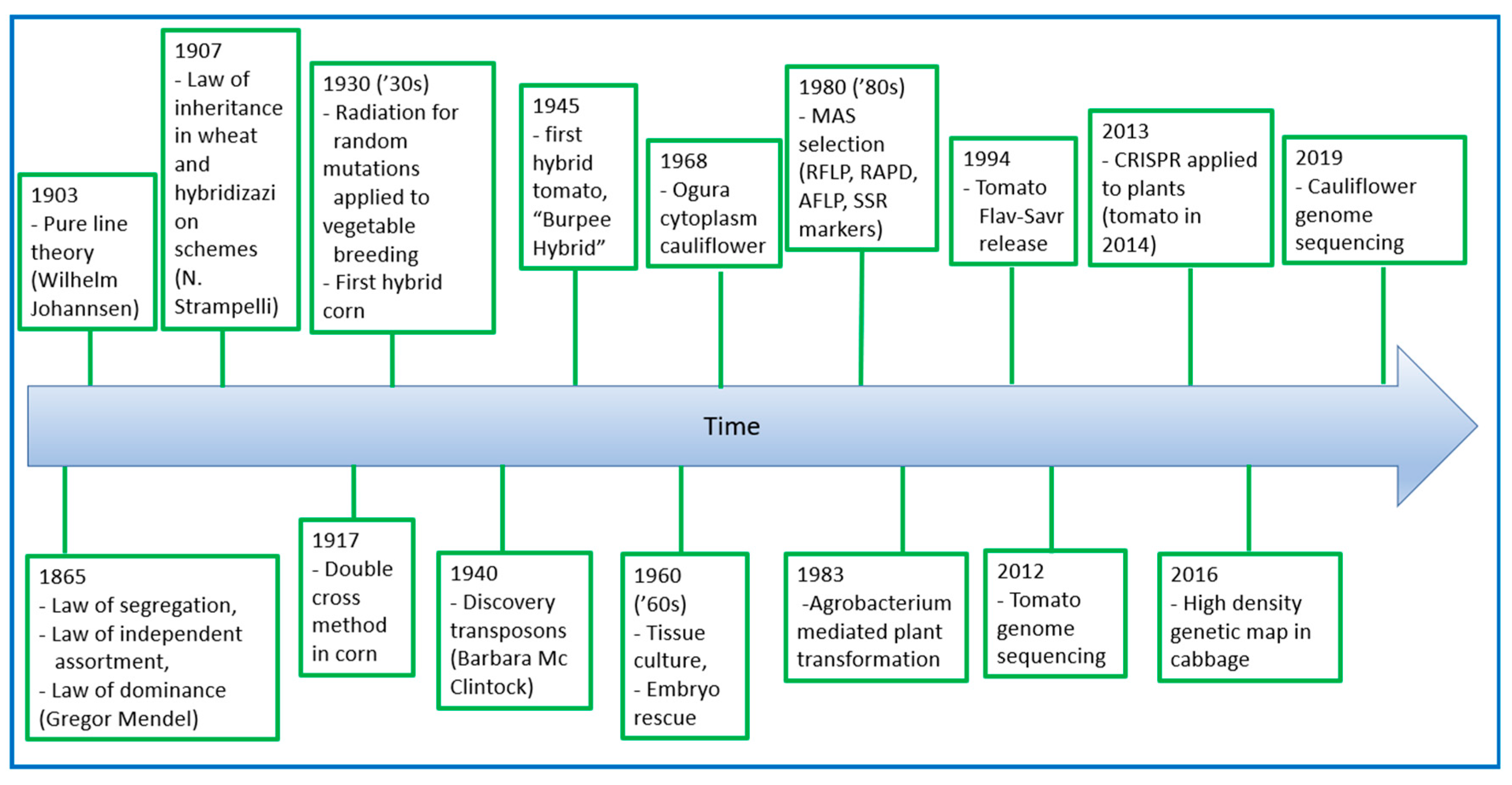
5. Evolution of Breeding for Nutritional and Organoleptic Quality
6. Biotechnological Approaches: From Transgenesis to Genome Editing
7. Conclusions and Perspectives
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Pott, D.M.; Durán-Soria, S.; Osorio, S.; Vallarino, J.G. Combining metabolomic and transcriptomic approaches to assess and improve crop quality traits. CABI Agric. Biosci. 2021, 2, 1–20. [Google Scholar] [CrossRef]
- Crossa, J.; Fritsche-Neto, R.; Montesinos-Lopez, O.A.; Costa-Neto, G.; Dreisigacker, S.; Montesinos-Lopez, A.; Bentley, A.R. The Modern Plant Breeding Triangle: Optimizing the Use of Genomics, Phenomics, and Enviromics Data. Front. Plant Sci. 2021, 12, 651480. [Google Scholar] [CrossRef]
- Aharoni, A.; O’Connell, A.P. Gene expression analysis of strawberry achene and receptacle maturation using DNA microarrays. J. Exp. Bot. 2002, 53, 2073–2087. [Google Scholar] [CrossRef] [PubMed]
- Dey, S.S.; Singh, N.; Bhatia, R.; Parkash, C.; Chandel, C. Genetic combining ability and heterosis for important vitamins and antioxidant pigments in cauliflower (Brassica oleracea var. botrytis L.). Euphytica 2014, 195, 169–181. [Google Scholar] [CrossRef]
- Cardi, T.; D’Agostino, N.; Tripodi, P. Genetic Transformation and Genomic Resources for Next-Generation Precise Genome Engineering in Vegetable Crops. Front. Plant Sci. 2017, 8, 241. [Google Scholar] [CrossRef]
- Jorasch, P. The global need for plant breeding innovation. Transgenic Res. 2019, 28, 81–86. [Google Scholar] [CrossRef] [PubMed]
- Churchill, F.B. William Johannsen and the genotype concept. J. Hist. Biol. 1974, 7, 5–30. [Google Scholar] [CrossRef]
- D’Amato, F. Introduction to Plant Genetic. In Italian Contribution to Plant Genetic and Breeding; Scarascia Mugnozza, G.T., Pagnotta, M.A., Eds.; Tipolitografia Quatrini A&F: Viterbo, Italy, 1998; pp. 21–53. [Google Scholar]
- Strampelli, N. Alla Ricerca e Creazione di Nuove Varietà di Frumenti a Mezzo di Ibridazione; Ti-Pografia Unione Coop. Editrice: Rome, Italy, 1907. [Google Scholar]
- Borojevic, K. Historic role of the wheat variety Akakomugi in Southern and Central European Wheat Breeding Programs. Breed. Sci. 2005, 55, 253–256. [Google Scholar] [CrossRef][Green Version]
- Pray, L.; Zhaurova, K. Barbara McClintock and the discovery of jumping genes (transposons). Nat. Sci. Educ. 2008, 1, 169. [Google Scholar]
- Somssich, M. A short history of plant transformation. PeerJ 2019. [Google Scholar] [CrossRef]
- Swaminathan, M.S. Preface. In Marker-Assisted Plant Breeding, Principles and Practices; Singh, B.D., Singh, A.K., Eds.; Springer: New Delhi, India, 2015; pp. vii–viii. [Google Scholar]
- Grube, R.C.; Radwanski, E.R.; Jahn, M. Comparative Genetics of Disease Resistance within the Solanaceae. Genetics 2000, 155, 873–887. [Google Scholar] [CrossRef]
- Cardi, T.; Stewart, C.N. Progress of targeted genome modification approaches in higher plants. Plant. Cell Rep. 2016, 35, 1401–1416. [Google Scholar] [CrossRef] [PubMed]
- Bortesi, L.; Fischer, R. The CRISPR/Cas9 system for plant genome editing and beyond. Biotechnol. Adv. 2015, 33, 41–52. [Google Scholar] [CrossRef] [PubMed]
- Klee, H.J.; Tieman, D.M. The genetics of fruit flavour preferences. Nat. Rev. Genet. 2018, 19, 347–356. [Google Scholar] [CrossRef]
- Chan, E.K.F.; Rowe, H.C.; Hansen, B.G.; Kliebenstein, D.J. The Complex Genetic Architecture of the Metabolome. PLoS Genet. 2010, 6, e1001198. [Google Scholar] [CrossRef] [PubMed]
- Sauvage, C.; Segura, V.; Bauchet, G.; Stevens, R.; Do, P.T.; Nikoloski, Z.; Fernie, A.R.; Causse, M. Genome-Wide Association in Tomato Reveals 44 Candidate Loci for Fruit Metabolic Traits. Plant Physiol. 2014, 165, 1120–1132. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Wang, W.; Peng, M.; Gong, L.; Gao, Y.; Wan, J.; Wang, S.; Shi, L.; Zhou, B.; Li, Z.; et al. Comparative and parallel genome-wide association studies for metabolic and agronomic traits in cereals. Nat. Commun. 2016, 7, 12767. [Google Scholar] [CrossRef]
- Iqbal, R.K.; Saeed, K.; Khan, A.; Noreen, I.; Bashir, M. Tomato (Lycopersicon esculetum) fruit improvement through breeding. Sch. J. Appl. Sci. Res. 2019, 2, 21–25. [Google Scholar]
- Campanelli, G.; Sestili, S.; Acciarri, N.; Montemurro, F.; Palma, D.; Leteo, F.; Beretta, M. Multi-Parental Advances Generation Inter-Cross Population, to Develop Organic Tomato Genotypes by Participatory Plant Breeding. Agronomy 2019, 9, 119. [Google Scholar] [CrossRef]
- Gramazio, P.; Pereira-Dias, L.; Vilanova, S.; Prohens, J.; Soler, S.; Esteras, J.; Garmendia, A.; Díez, M.J. Morphoagronomic characterization and whole-genome resequencing of eight highly diverse wild and weedy S. pimpinellifolium and S. lycopersicum var. cerasiforme accessions used for the first interspecific tomato MAGIC population. Hortic. Res. 2020, 7, 1–16. [Google Scholar] [CrossRef]
- Esposito, S.; Cardi, T.; Campanelli, G.; Sestili, S.; Díez, M.J.; Soler, S.; Prohens, J.; Tripodi, P. ddRAD sequencing-based genotyping for population structure analysis in cultivated tomato provides new insights into the genomic diversity of Mediterranean ‘da serbo’ type long shelf-life germplasm. Hortic. Res. 2020, 7, 1–14. [Google Scholar] [CrossRef]
- Martínez-Cuenca, M.-R.; Pereira-Dias, L.; Soler, S.; López-Serrano, L.; Alonso, D.; Calatayud, Á.; Díez, M.J. Adaptation to Water and Salt Stresses of Solanum pimpinellifolium and Solanum lycopersicum var. cerasiforme. Agronomy 2020, 10, 1169. [Google Scholar] [CrossRef]
- Rosa-Martínez, E.; Adalid, A.M.; Alvarado, L.E.; Burguet, R.; García-Martínez, M.D.; Pereira-Dias, L.; Casanova, C.; Soler, E.; Figàs, M.R.; Plazas, M.; et al. Variation for Composition and Quality in a Collection of the Resilient Mediterranean ‘de penjar’ Long Shelf-Life Tomato Under High and Low N Fertilization Levels. Front. Plant Sci. 2021, 12, 633957. [Google Scholar] [CrossRef]
- Gao, Y.; Zhu, N.; Zhu, X.; Wu, M.; Jiang, C.-Z.; Grierson, D.; Luo, Y.; Shen, W.; Zhong, S.; Fu, D.-Q.; et al. Diversity and redundancy of the ripening regulatory networks revealed by the fruit ENCODE and the new CRISPR/Cas9 CNR and NOR mutants. Hortic. Res. 2019, 6, 1–10. [Google Scholar] [CrossRef]
- The Tomato Genome Consortium. The tomato genome sequence provides insights into fleshy fruit evolution. Nature 2012, 485, 635–641. [Google Scholar] [CrossRef]
- Cheema, D.S.; Dhaliwal, M.S. Hybrid Tomato Breeding. J. New Seeds 2005, 6, 1–14. [Google Scholar] [CrossRef]
- Schiavi, M.; Vitelli, G.; Maestrelli, A.; Forni, E.; Giovannessi, L. Breeding of green curded cauliflower (Brassica oleracea L., botrytis group) for fresh market and freezing. Acta Hortic. 1998, 459, 403–410. [Google Scholar] [CrossRef]
- Acciarri, N.; Branca, F.; Sabatini, E.; Argento, S.; Magnifico, V. Miglioramento genetico dei cavolfiori a corimbo bianco e colourato. L’Inf. Agrar. 2004, 25, 33–36. [Google Scholar]
- Schulz, O.E. Cruciferae-Brassicaceae part I, Brassicinae and raphaninae. In Das Pflanzenreich; Wilhelm Engelmann: Leipzig, Germany, 1919; pp. 1–290. [Google Scholar]
- Smith, L.B.; King, G.J. The distribution of BoCAL-a alleles in Brassica oleracea is consistent with a genetic model for curd development and domestication of the cauliflower. Mol. Breed. 2000, 6, 603–613. [Google Scholar] [CrossRef]
- Singh, B.K.; Singh, B.; Singh, P.M. Breeding Cauliflower: A Review. Int. J. Veg. Sci. 2018, 24, 58–84. [Google Scholar] [CrossRef]
- Pelletier, G.; Primard, C.; Vedel, F.; Chétrit, P.; Rémy, R.; Roussell, P.; Renard, M. Intergeneric cytoplasmic hybridization in Cruciferae by protoplast fusion. Mol. Gen. Genet. 1983, 191, 244–250. [Google Scholar] [CrossRef]
- Kucera, V.; Chytilov, V.; Vyvadilov, M.; Klima, M. Hybrid breeding of cauliflower using self-incompatibility and cytoplasmic male sterility. Hortic. Sci. 2006, 33, 148–152. [Google Scholar] [CrossRef]
- Kyriacou, M.; Rouphael, Y. Towards a new definition of quality for fresh fruits and vegetables. Sci. Hortic. 2018, 234, 463–469. [Google Scholar] [CrossRef]
- Dorais, A.; Gosselin, A.; Papadopolus, M. Greenhouse Tomato Fruit Quality. Hortic. Rev. 2001, 26, 238–262. [Google Scholar]
- Barry, C.; Giovannoni, J.J. Ethylene and Fruit Ripening. J. Plant. Growth Regul. 2007, 26, 143–159. [Google Scholar] [CrossRef]
- Conesa, M.À.; Fullana-Pericàs, M.; Granell, A.; Galmés, J. Mediterranean Long Shelf-Life Landraces: An Untapped Genetic Resource for Tomato Improvement. Front. Plant Sci. 2020, 10, 1651. [Google Scholar] [CrossRef]
- Wang, D.; Seymour, G.B. Tomato Flavor: Lost and Found? Mol. Plant 2017, 10, 782–784. [Google Scholar] [CrossRef] [PubMed]
- Rambla, J.L.; Tikunov, Y.M.; Monforte, A.J.; Bovy, A.G.; Granell, A. The expanded tomato fruit volatile landscape. J. Exp. Bot. 2013, 65, 4613–4623. [Google Scholar] [CrossRef]
- Zhao, Z.; Gu, H.; Sheng, X.; Yu, H.; Wang, J.; Huang, L.; Wang, D. Genome-Wide Single-Nucleotide Polymorphisms Discovery and High-Density Genetic Map Construction in Cauliflower Using Specific-Locus Amplified Fragment Sequencing. Front. Plant Sci. 2016, 7, 334. [Google Scholar] [CrossRef]
- Raja, M.M.; Raja, M.; Imran, M.; Rahmsn, H.A. Quality aspects of cauliflower during storage. Int. Food Res. J. 2011, 18, 427–431. [Google Scholar]
- Fernie, A.R.; Tadmor, Y.; Zamir, D. Natural genetic variation for improving crop quality. Curr. Opin. Plant Biol. 2006, 9, 196–202. [Google Scholar] [CrossRef]
- Varalakshmi, B. Heterosis and combining ability for yield and its components in early cauliflower. Indian J. Hortic. 2009, 66, 198–203. [Google Scholar]
- Singh, S.; Kalia, P.; Meena, R.K.; Mangal, M.; Islam, S.; Saha, S.; Tomar, B.S. Genetics and Expression Analysis of Anthocyanin Accumulation in Curd Portion of Sicilian Purple to Facilitate Biofortification of Indian Cauliflower. Front. Plant Sci. 2020, 10, 1766. [Google Scholar] [CrossRef]
- Chiu, L.-W.; Zhou, X.; Burke, S.; Wu, X.; Prior, R.L.; Li, L. The Purple Cauliflower Arises from Activation of a MYB Transcription Factor. Plant Physiol. 2010, 154, 1470–1480. [Google Scholar] [CrossRef] [PubMed]
- Chiu, L.W. Transposon Insertion at the Promoter of a MYB Transcription Factor Results in Ectopic Anthocyanins Accumulation in Purple Cauliflower (Brassica oleracea var. botrytis). Ph.D. Thesis, Cornell University, Ithaca, NY, USA, 2010. [Google Scholar]
- Johnson, E.J.; Hammond, B.; Yeum, K.-J.; Qin, J.; Wang, X.D.; Castaneda, C.; Snodderly, D.; Russell, R.M. Relation among serum and tissue concentrations of lutein and zeaxanthin and macular pigment density. Am. J. Clin. Nutr. 2000, 71, 1555–1562. [Google Scholar] [CrossRef]
- Crisp, P.; Walkey, D.G.A.; Bellman, E.; Roberts, E. A mutation affecting curd colour in cauliflower (Brassica oleracea L. var. Botrytis DC). Euphytica 1975, 24, 173–176. [Google Scholar] [CrossRef]
- Dickson, M.H.; Lee, C.Y.; Blamble, A.E. Orange-curd high carotene cauliflower inbreds, NY 156, NY 163, and NY Hort165. Science 1998, 23, 778–779. [Google Scholar]
- Li, L.; Paolillo, D.J.; Parthasarathy, M.V.; DiMuzio, E.M.; Garvin, D.F. A novel gene mutation that confers abnormal patterns of β-carotene accumulation in cauliflower (Brassica oleracea var. botrytis). Plant J. 2001, 26, 59–67. [Google Scholar] [CrossRef]
- Wang, L.-S.; Stoner, G.D. Anthocyanins and their role in cancer prevention. Cancer Lett. 2008, 269, 281–290. [Google Scholar] [CrossRef]
- Volden, J.; Bengtsson, G.B.; Wicklund, T. Glucosinolates, l-ascorbic acid, total phenols, anthocyanins, antioxidant capacities and colour in cauliflower (Brassica oleracea L. ssp. botrytis); effects of long-term freezer storage. Food Chem. 2009, 112, 967–976. [Google Scholar] [CrossRef]
- Scalzo, R.L.; Genna, A.; Branca, F.; Chedin, M.; Chassaigne, H. Anthocyanin composition of cauliflower (Brassica oleracea L. var. botrytis) and cabbage (B. oleracea L. var. capitata) and its stability in relation to thermal treatments. Food Chem. 2008, 107, 136–144. [Google Scholar] [CrossRef]
- Tommonaro, G.; De Prisco, R.; Abbamondi, G.R.; Marzocco, S.; Saturnino, C.; Poli, A.; Nicolaus, B. Evaluation of Antioxidant Properties, Total Phenolic Content, and Biological Activities of New Tomato Hybrids of Industrial Interest. J. Med. Food 2012, 15, 483–489. [Google Scholar] [CrossRef]
- Blando, F.; Berland, H.; Maiorano, G.; Durante, M.; Mazzucato, A.; Picarella, M.E.; Nicoletti, I.; Gerardi, C.; Mita, G.; Andersen, Ø.M. Nutraceutical Characterization of Anthocyanin-Rich Fruits Produced by “Sun Black” Tomato Line. Front. Nutr. 2019, 6, 133. [Google Scholar] [CrossRef] [PubMed]
- Mazzucato, A.; Willems, D.; Bernini, R.; Picarella, M.E.; Santangelo, E.; Ruiu, F.; Tilesi, F.; Soressi, G.P. Novel phenotypes related to the breeding of purple-fruited tomatoes and effect of peel extracts on human cancer cell proliferation. Plant Physiol. Biochem. 2013, 72, 125–133. [Google Scholar] [CrossRef] [PubMed]
- Dono, G.; Rambla, J.L.; Frusciante, S.; Granell, A.; Diretto, G.; Mazzucato, A. Color Mutations Alter the Biochemical Composition in the San Marzano Tomato Fruit. Metabolites 2020, 10, 110. [Google Scholar] [CrossRef] [PubMed]
- Gapper, N.; Giovannoni, J.J.; Watkins, C. Understanding development and ripening of fruit crops in an ‘omics’ era. Hortic. Res. 2014, 1, 14034. [Google Scholar] [CrossRef]
- Carli, P.; Barone, A.; Fogliano, V.; Frusciante, L.; Ercolano, M.R. Dissection of genetic and environmental factors involved in tomato organoleptic quality. BMC Plant Biol. 2011, 11, 58. [Google Scholar] [CrossRef]
- Ercolano, M.R.; Sacco, A.; Ferriello, F.; D’Alessandro, R.; Tononi, P.; Traini, A.; Barone, A.; Zago, E.; Chiusano, M.L.; Buson, G.; et al. Patchwork sequencing of tomato San Marzano and Vesuviano varieties highlights genome-wide variations. BMC Genom. 2014, 15, 138. [Google Scholar] [CrossRef]
- Tieman, D.; Zhu, G., Jr.; Lin, T.; Nguyen, C.; Bies, D.; Rambla, J.L.; Beltran, K.S.O.; Taylor, M.; Zhang, B.; Ikeda, H. A chemical genetic roadmap to improved tomato flavor. Science 2017, 355, 391–394. [Google Scholar] [CrossRef]
- Tranchida-Lombardo, V.; Cigliano, R.A.; Anzar, I.; Landi, S.; Palombieri, S.; Colantuono, C.; Bostan, H.; Termolino, P.; Aversano, R.; Batelli, G.; et al. Wholegenome resequencing of two Italian tomato landraces reveals sequence variations in genes associated with stress tolerance, fruit quality and long shelf-life traits. DNA Res. 2018, 25, 149–160. [Google Scholar] [CrossRef]
- Gao, L.; Gonda, I.; Sun, H.; Ma, Q.; Bao, K.; Tieman, D.M.; Chang, E.; Fish, T.L.; Stromberg, K.A.; Sacks, G.L.; et al. The tomato pan-genome uncovers new genes and a rare allele regulating fruit flavor. Nat. Genet. 2019, 51, 1044–1051. [Google Scholar] [CrossRef] [PubMed]
- Wu, F.; Tanksley, S.D. Chromosomal evolution in the plant family Solanaceae. BMC Genom. 2010, 11, 182. [Google Scholar] [CrossRef]
- Zhu, G.; Wang, S.; Huang, Z.; Zhang, S.; Liao, Q.; Zhang, C.; Lin, T.; Qin, M.; Peng, M.; Yang, C.; et al. Rewiring of the Fruit Metabolome in Tomato Breeding. Cell 2018, 172, 249–261.e12. [Google Scholar] [CrossRef] [PubMed]
- Baxter, C.J.; Carrari, F.; Bauke, A.; Overy, S.; Hill, S.A.; Quick, P.W.; Fernie, A.R.; Sweetlove, L.J. Fruit carbohydrate me-tabolism in an introgression line of tomato with increased fruit soluble solids. Plant Cell Physiol. 2005, 46, 425–437. [Google Scholar] [CrossRef] [PubMed]
- Colanero, S.; Perata, P.; Gonzali, S. The atroviolacea Gene Encodes an R3-MYB Protein Repressing Anthocyanin Synthesis in Tomato Plants. Front. Plant Sci. 2018, 9, 830. [Google Scholar] [CrossRef]
- Tanksley, S.D. The Genetic, Developmental, and Molecular Bases of Fruit Size and Shape Variation in Tomato. Plant Cell 2004, 16, S181–S189. [Google Scholar] [CrossRef]
- Adhikari, P.; McNellie, J.; Panthee, D.R. Detection of Quantitative Trait Loci (QTL) Associated with the Fruit Morphology of Tomato. Genes 2020, 11, 1117. [Google Scholar] [CrossRef]
- Casas-Diaz, A.V.; Hewitt, J.D.; Lapushner, D. Effects of parthenocarpy on fruit quality in tomato. J. Am. Soc. Hortic. Sci. 1987, 112, 634–637. [Google Scholar]
- Mazzucato, A.; Taddei, A.; Soressi, G. The parthenocarpic fruit (pat) mutant of tomato (Lycopersicon esculentum Mill.) sets seedless fruits and has aberrant anther and ovule development. Development 1998, 125, 107–114. [Google Scholar] [CrossRef]
- Acciarri, N.; Ferrari, V.; Vitelli, G.; Ficcadenti, N.; Pandolfini, T.; Spena, A.; Rotino, G.L. Effetto della partenocarpia in ibridi di pomodoro geneticamente modificati. L’Inf. Agrar. 2000, 4, 117–122. [Google Scholar]
- Beraldi, D.; Picarella, M.E.; Soressi, G.P.; Mazzucato, A. Fine mapping of the parthenocarpic fruit (pat) mutation in tomato. Theor. Appl. Genet. 2003, 108, 209–216. [Google Scholar] [CrossRef] [PubMed]
- Rotino, G.L.; Acciarri, N.; Sabatini, E.; Mennella, G.; Scalzo, R.L.; Maestrelli, A.; Molesini, B.; Pandolfini, T.; Scalzo, J.; Mezzetti, B.; et al. Open field trial of genetically modified parthenocarpic tomato: Seedlessness and fruit quality. BMC Biotechnol. 2005, 5, 32. [Google Scholar] [CrossRef]
- Ewitzel, K.; Eneugart, S.; Eruppel, S.; Schreiner, M.; Ewiesner, M.; Ebaldermann, S. Recent progress in the use of ‘omics technologies in brassicaceous vegetables. Front. Plant Sci. 2015, 6, 244. [Google Scholar] [CrossRef]
- Francisco, M.; Soengas, P.; Velasco, P.; Bhadauria, V.; Cartea, M.; Rodríguez, V.M. Omics Approach to Identify Factors Involved in Brassica Disease Resistance. Curr. Issues Mol. Biol. 2015, 19, 31–42. [Google Scholar]
- Park, S.-Y.; Lim, S.-H.; Ha, S.-H.; Yeo, Y.; Park, W.T.; Kwon, D.Y.; Park, S.U.; Kim, J.K. Metabolite Profiling Approach Reveals the Interface of Primary and Secondary Metabolism in Colored Cauliflowers (Brassica oleracea L. ssp. botrytis). J. Agric. Food Chem. 2013, 61, 6999–7007. [Google Scholar] [CrossRef]
- Ding, Y.; Jian, Y. An Orange Cauliflower Hybrid ‘Jinyu 60’ with High Β-Carotene. Acta Hortic. 2010, 856, 261–264. [Google Scholar] [CrossRef]
- Wang, Y.; Li, X.; Xu, J.; Xu, Y.; Liu, L. Cultivar identification and genetic diversity analysis of cauliflower with molecular markers. Acta Hortic. 2011, 918, 315–321. [Google Scholar] [CrossRef]
- Lv, H.; Wang, Q.; Zhang, Y.; Yang, L.; Fang, Z.; Wang, X.; Liu, Y.; Zhuang, M.; Lin, Y.; Yu, H.; et al. Linkage map construction using InDel and SSR markers and QTL analysis of heading traits in Brassica oleracea var. capitata L. Mol. Breed. 2014, 34, 87–98. [Google Scholar] [CrossRef]
- Ogura, H. Studies on the new male sterility in Japanese radish with special reference to the utilisation of this sterility towards the pratical raising of hybrid seeds. Mem. Fac. Agric. Kagoshima Univ. 1968, 6, 39–78. [Google Scholar]
- Dey, S.S.; Bhatia, R.; Parkash, C.; Sharma, S.; Dabral, M.; Mishra, V.; Bhardwaj, I.; Sharma, K.; Sharma, V.K.; Kumar, R. Alteration in important quality traits and antioxidant activities in Brassica oleracea with Oguracybrid cytoplasm. Plant Breed. 2017, 136, 400–409. [Google Scholar] [CrossRef]
- Acciarri, N.; Cardi, T. Brassica. In Italian Contribution to Plant Genetic and Breeding; Scarascia Mugnozza, G.T., Pagnotta, M.A., Eds.; Tipolitografia Quatrini A&F: Viterbo, Italy, 1998; pp. 481–486. [Google Scholar]
- Sun, B.; Zheng, A.; Jiang, M.; Xue, S.; Yuan, Q.; Jiang, L.; Chen, Q.; Li, M.; Wang, Y.; Zhang, Y.; et al. CRISPR/Cas9-mediated mutagenesis of homologous genes in Chinese kale. Sci. Rep. 2018, 8, 16786. [Google Scholar] [CrossRef]
- Bolger, A.; Scossa, F.; Bolger, M.E.; Lanz, C.; Maumus, F.; Tohge, T.; Quesneville, H.; Alseekh, S.; Sørensen, I.; Lichtenstein, G.; et al. The genome of the stress-tolerant wild tomato species Solanum pennellii. Nat. Genet. 2014, 46, 1034–1038. [Google Scholar] [CrossRef]
- Wang, X.; Gao, L.; Jiao, C.; Stravoravdis, S.; Hosmani, P.S.; Saha, S.; Zhang, J.; Mainiero, S.; Strickler, S.R.; Catala, C.; et al. Genome of Solanum pimpinellifolium provides insights into structural variants during tomato breeding. Nat. Commun. 2020, 11, 1–11. [Google Scholar] [CrossRef]
- Razali, R.; Bougouffa, S.; Morton, M.J.L.; Lightfoot, D.J.; Alam, I.; Essack, M.; Arold, S.T.; Kamau, A.A.; Schmöckel, S.M.; Pailles, Y.; et al. The Genome Sequence of the Wild Tomato Solanum pimpinellifolium Provides Insights into Salinity Tolerance. Front. Plant Sci. 2018, 9, 1402. [Google Scholar] [CrossRef]
- Sun, D.; Wang, C.; Zhang, X.; Zhang, W.; Jiang, H.; Yao, X.; Liu, L.; Wen, Z.; Niu, G.; Shan, X. Draft genome sequence of cauliflower (Brassica oleracea L. var. botrytis) provides new insights into the C genome in Brassica species. Hortic. Res. 2019, 6, 1–11. [Google Scholar] [CrossRef]
- Wang, X.; Wang, H.; Wang, J.; Sun, R.; Wu, J.; Liu, S.; Bai, Y.; Mun, J.-H.; Bancroft, I.; Cheng, F.; et al. The genome of the mesopolyploid crop species Brassica rapa. Nat. Genet. 2011, 43, 1035–1039. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Liu, D.; Wang, X.; Ji, C.; Cheng, F.; Liu, B.; Hu, Z.; Chen, S.; Pental, D.; Ju, Y.; et al. The genome sequence of allopolyploid Brassica juncea and analysis of differential homoeolog gene expression influencing selection. Nat. Genet. 2016, 48, 1225–1232. [Google Scholar] [CrossRef]
- Bayer, P.E.; Hurgobin, B.; Golicz, A.A.; Chan, C.K.; Yuan, Y.; Lee, H.; Renton, M.; Meng, J.; Li, R.; Long, Y.; et al. Assembly and comparison of two closely related Brassica napus genomes. Plant Biotechnol. J. 2017, 15, 1602–1610. [Google Scholar] [CrossRef] [PubMed]
- Gosal, S.S.; Wani, S. Plant Genetic Transformation and Transgenic Crops: Methods and Applications. In Biotechnologies of Crop Improvement; Springer: Cham, Switzerland, 2018; Volume 2, pp. 1–23. [Google Scholar]
- Di Matteo, A.; Manuela, M.; Sacco, A.; Frusciante, L.; Barone, A. Genetic Transformation in Tomato: Novel Tools to Improve Fruit Quality and Pharmaceutical Production. In Genetic Transformation; InTech: Wheeling, IL, USA, 2011. [Google Scholar]
- Krieger, E.K.; Allen, E.; Gilbertson, L.A.; Roberts, J.K.; Hiatt, W.; Sanders, R.A. The Flavr Savr Tomato, an Early Example of RNAi Technology. HortScience 2008, 43, 962–964. [Google Scholar] [CrossRef]
- Gerszberg, A.; Hnatuszko-Konka, K.; Kowalczyk, T.; Kononowicz, A.K. Tomato (Solanum lycopersicum L.) in the service of biotechnology. Plant. Cell Tissue Organ. Cult. 2015, 120, 881–902. [Google Scholar] [CrossRef]
- Barg, R.; Shabtai, S.; Salts, Y. Transgenic Tomato (Lycopersicom esculentum). In Biotechnology in Agriculture and Forestry 47, Transgenic Crop; Bajaj, A., Ed.; Springer: New York, NY, USA, 2012; Volume 2, pp. 219–233. [Google Scholar]
- Rosati, C.; Aquilani, R.; Dharmapuri, S.; Pallara, P.; Marusic, C.; Tavazza, R.; Bouvier, F.; Camara, B.; Giuliano, G. Metabolic engineering of beta-carotene and lycopene content in tomato fruit. Plant J. 2000, 24, 413–420. [Google Scholar] [CrossRef]
- Long, M.; Millar, D.J.; Kimura, Y.; Donovan, G.; Rees, J.; Fraser, P.D.; Bramley, P.M.; Bolwell, G.P. Metabolite profiling of carotenoid and phenolic pathways in mutant and transgenic lines of tomato: Identification of a high antioxidant fruit line. Phytochemistry 2006, 67, 1750–1757. [Google Scholar] [CrossRef]
- Giuliano, G. Plant carotenoids: Genomics meets multi-gene engineering. Curr. Opin. Plant Biol. 2014, 19, 111–117. [Google Scholar] [CrossRef]
- Wurbs, D.; Ruf, S.; Bock, R. Contained metabolic engineering in tomatoes by expression of carotenoid biosynthesis genes from the plastid genome. Plant J. 2006, 49, 276–288. [Google Scholar] [CrossRef]
- Apel, W.; Bock, R. Enhancement of Carotenoid Biosynthesis in Transplastomic Tomatoes by Induced Lycopene-to-Provitamin A Conversion. Plant Physiol. 2009, 151, 59–66. [Google Scholar] [CrossRef] [PubMed]
- Orzaez, D.; Medina, A.; Torre, S.; Fernández-Moreno, J.P.; Rambla, J.L.; Fernández-Del-Carmen, A.; Butelli, E.; Martin, C.; Granell, A. A Visual Reporter System for Virus-Induced Gene Silencing in Tomato Fruit Based on Anthocyanin Accumulation. Plant Physiol. 2009, 150, 1122–1134. [Google Scholar] [CrossRef]
- Butelli, E.; Titta, L.; Giorgio, M.; Mock, H.-P.; Matros, A.; Peterek, S.; Schijlen, E.G.W.M.; Hall, R.D.; Bovy, A.G.; Luo, J.; et al. Enrichment of tomato fruit with health-promoting anthocyanins by expression of select transcription factors. Nat. Biotechnol. 2008, 26, 1301–1308. [Google Scholar] [CrossRef] [PubMed]
- Bassolino, L.; Zhang, Y.; Schoonbeek, H.-J.; Kiferle, C.; Perata, P.; Martin, C. Accumulation of anthocyanins in tomato skin extends shelf life. New Phytol. 2013, 200, 650–655. [Google Scholar] [CrossRef]
- Giovinazzo, G.; Ingrosso, I.; Taurino, M.; Santino, A. Metabolic Engineering for Functional Foods: Tomato Fruits and Stilbenes. In Natural Products; Springer: Cham, Switzerland, 2013; pp. 1581–1597. [Google Scholar]
- Giovinazzo, G.; D’Amico, L.; Paradiso, A.; Bollini, R.; Sparvoli, F.; De Gara, L. Antioxidant metabolite profiles in tomato fruit constitutively expressing the grapevine stilbene synthase gene. Plant Biotechnol. J. 2004, 3, 57–69. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Wang, Y.; Chen, S.; Tian, H.; Fu, D.; Zhu, B.; Luo, Y.; Zhu, H. Lycopene Is Enriched in Tomato Fruit by CRISPR/Cas9-Mediated Multiplex Genome Editing. Front. Plant Sci. 2018, 9, 559. [Google Scholar] [CrossRef]
- D’Ambrosio, C.; Stigliani, A.L.; Giorio, G. CRISPR/Cas9 editing of carotenoid genes in tomato. Transgenic Res. 2018, 27, 367–378. [Google Scholar] [CrossRef]
- Deng, L.; Wang, H.; Sun, C.; Li, Q.; Jiang, H.; Du, M.; Li, C.-B.; Li, C. Efficient generation of pink-fruited tomatoes using CRISPR/Cas9 system. J. Genet. Genom. 2018, 45, 51–54. [Google Scholar] [CrossRef] [PubMed]
- Čermák, T.; Baltes, N.J.; Čegan, R.; Zhang, Y.; Voytas, D.F. High-frequency, precise modification of the tomato genome. Genome Biol. 2015, 16, 1–15. [Google Scholar] [CrossRef]
- Nonaka, S.; Arai, C.; Takayama, M.; Matsukura, C.; Ezura, H. Efficient increase of ɣ-aminobutyric acid (GABA) content in tomato fruits by targeted mutagenesis. Sci. Rep. 2017, 7, 1–14. [Google Scholar] [CrossRef]
- Li, R.; Li, R.; Li, X.; Fu, D.; Zhu, B.; Tian, H.; Luo, Y.; Zhu, H. Multiplexed CRISPR/Cas9-mediated metabolic engineering of γ-aminobutyric acid levels in Solanum lycopersicum. Plant. Biotechnol. J. 2018, 16, 415–427. [Google Scholar] [CrossRef] [PubMed]
- Gago, C.; Drosou, V.; Paschalidis, K.; Guerreiro, A.; Miguel, G.; Antunes, D.; Hilioti, Z. Targeted gene disruption coupled with metabolic screen approach to uncover the LEAFY COTYLEDON1-LIKE4 (L1L4) function in tomato fruit metabolism. Plant Cell Rep. 2017, 36, 1065–1082. [Google Scholar] [CrossRef] [PubMed]
- Ito, Y.; Nishizawa-Yokoi, A.; Endo, M.; Mikami, M.; Toki, S. CRISPR/Cas9-mediated mutagenesis of the RIN locus that regulates tomato fruit ripening. Biochem. Biophys. Res. Commun. 2015, 467, 76–82. [Google Scholar] [CrossRef]
- Ito, Y.; Nishizawa-Yokoi, A.; Endo, M.; Mikami, M.; Shima, Y.; Nakamura, N.; Kotake-Nara, E.; Kawasaki, S.; Toki, S. Re-evaluation of the rin mutation and the role of RIN in the induction of tomato ripening. Nat. Plants 2017, 3, 866–874. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Zhu, B.; Pirrello, J.; Xu, C.; Zhang, B.; Bouzayen, M.; Chen, K.; Grierson, D. Roles of RIN and ethylene in tomato fruit ripening and ripening-associated traits. New Phytol. 2020, 226, 460–475. [Google Scholar] [CrossRef]
- Gao, Y.; Wei, W.; Zhao, X.; Tan, X.; Fan, Z.; Zhang, Y.; Jing, Y.; Meng, L.; Zhu, B.; Zhu, H.; et al. A NAC transcription factor, NOR-like1, is a new positive regulator of tomato fruit ripening. Hortic. Res. 2018, 5, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Tavano, E.C.D.R.; Lammers, M.; Martinelli, A.P.; Angenent, G.C.; De Maagd, R.A. Re-evaluation of transcription factor function in tomato fruit development and ripening with CRISPR/Cas9-mutagenesis. Sci. Rep. 2019, 9, 1–10. [Google Scholar] [CrossRef]
- Lang, Z.; Wang, Y.; Tang, K.; Tang, D.; Datsenka, T.; Cheng, J.; Zhang, Y.; Handa, A.K.; Zhu, J.-K. Critical roles of DNA demethylation in the activation of ripening-induced genes and inhibition of ripening-repressed genes in tomato fruit. Proc. Natl. Acad. Sci. USA 2017, 114, E4511–E4519. [Google Scholar] [CrossRef]
- Yang, Y.; Zhu, G.; Li, R.; Yan, S.; Fu, D.; Zhu, B.; Tian, H.; Luo, Y.; Zhu, H. The RNA Editing Factor SlORRM4 Is Required for Normal Fruit Ripening in Tomato. Plant Physiol. 2017, 175, 1690–1702. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Fu, D.; Zhu, B.; Luo, Y.; Zhu, H. CRISPR/Cas9-mediated mutagenesis of lncRNA1459 alters tomato fruit ripening. Plant J. 2018, 94, 513–524. [Google Scholar] [CrossRef] [PubMed]
- Uluisik, S.; Chapman, N.H.; Smith, R.; Poole, M.; Adams, G.; Gillis, R.B.; Besong, T.M.D.; Sheldon, J.; Stiegelmeyer, S.; Perez-Fons, L.; et al. Genetic improvement of tomato by targeted control of fruit softening. Nat. Biotechnol. 2016, 34, 950–952. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Samsulrizal, N.H.; Yan, C.; Allcock, N.S.; Craigon, J.; Blanco-Ulate, B.; Ortega-Salazar, I.; Marcus, S.E.; Bagheri, H.M.; Perez-Fons, L.; et al. Characterisation of CRISPR mutants targeting genes modulating pectin degradation in ripening tomato. Plant Physiol. 2018, 179, 544–557. [Google Scholar] [CrossRef]
- Yu, Q.-H.; Wang, B.; Li, N.; Tang, Y.; Yang, S.; Yang, T.; Xu, J.; Guo, C.; Yan, P.; Wang, Q.; et al. CRISPR/Cas9-induced Targeted Mutagenesis and Gene Replacement to Generate Long-shelf Life Tomato Lines. Sci. Rep. 2017, 7, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Zsögön, A.; Čermák, T.; Naves, E.R.; Notini, M.M.; Edel, K.H.; Weinl, S.; Freschi, L.; Voytas, D.; Kudla, J.; Peres, L.E.P. De novo domestication of wild tomato using genome editing. Nat. Biotechnol. 2018, 36, 1211–1216. [Google Scholar] [CrossRef]
- Gao, Y.; Liu, J.; Chen, Y.; Tang, H.; Wang, Y.; He, Y.; Ou, Y.; Sun, X.; Wang, S.; Yao, Y. Tomato SlAN11 regulates flavonoid biosynthesis and seed dormancy by interaction with bHLH proteins but not with MYB proteins. Hortic. Res. 2018. [Google Scholar] [CrossRef] [PubMed]
- Zuluaga, D.L.; Gonzali, S.; Loreti, E.; Pucciariello, C.; Degl’Innocenti, E.; Guidi, L.; Alpi, A.; Perata, P. Arabidopsis thaliana MYB75/PAP1 transcription factor induces anthocyanin production in transgenic tomato plants. Funct. Plant Biol. 2008, 35, 606–618. [Google Scholar] [CrossRef] [PubMed]
- Xu, W.; Dubos, C.; Lepiniec, L. Transcriptional control of flavonoid biosynthesis by MYB–bHLH–WDR complexes. Trends Plant Sci. 2015, 20, 176–185. [Google Scholar] [CrossRef] [PubMed]
- Naves, E.R.; Silva, L.D.; Sulpice, R.; Araújo, W.; Nunes-Nesi, A.; Peres, L.E.; Zsögön, A. Capsaicinoids: Pungency beyond Capsicum. Trends Plant Sci. 2019, 24, 109–120. [Google Scholar] [CrossRef] [PubMed]
- Martín-Pizarro, C.; Posé, D. Genome Editing as a Tool for Fruit Ripening Manipulation. Front. Plant Sci. 2018, 9, 1415. [Google Scholar] [CrossRef]
- Nugent, G.D.; Coyne, S.; Nguyen, T.T.; Kavanagh, T.A.; Dix, P.J. Nuclear and plastid transformation of Brassica oleracea var. botrytis (cauliflower) using PEG-mediated uptake of DNA into protoplasts. Plant Sci. 2006, 170, 135–142. [Google Scholar] [CrossRef]
- Nguyen, K.; Jordi, W.; Van Dun, K.; Schepers, F.; Davelaar, E.; Stoopen, G.; Dix, P.J.; Kane, E.J. Delayed Senescence in Cauliflower Transformed with an Autoregulated Isopentenyl Transferase Gene. Int. J. Plant Sci. 2008, 169, 339–347. [Google Scholar] [CrossRef]
- Kowalczyk, T.; Gerszberg, A.; Durańska, P.; Biłas, R.; Hnatuszko-Konka, K. High efficiency transformation of Brassica oleracea var. botrytis plants by Rhizobium rhizogenes. AMB Express 2018, 8, 1–11. [Google Scholar] [CrossRef]
- Zhou, X.; Van Eck, J.; Li, L. Use of the cauliflower or gene for improving crop nutritional quality. Biotechnol. Annu. Rev. 2008, 14, 171–190. [Google Scholar] [CrossRef] [PubMed]
- Lawrenson, T.; Shorinola, O.; Stacey, N.; Li, C.; Østergaard, L.; Patron, N.; Uauy, C.; Harwood, W. Induction of targeted, heritable mutations in barley and Brassica oleracea using RNA-guided Cas9 nuclease. Genome Biol. 2015, 16, 1–13. [Google Scholar] [CrossRef]
- Ma, C.; Liu, M.; Li, Q.; Si, J.; Ren, X.; Song, H. Efficient BoPDS Gene Editing in Cabbage by the CRISPR/Cas9 System. Hortic. Plant J. 2019, 5, 164–169. [Google Scholar] [CrossRef]
- Ma, C.; Zhu, C.; Zheng, M.; Liu, M.; Zhang, D.; Liu, B.; Li, Q.; Si, J.; Ren, X.; Song, H. CRISPR/Cas9-mediated multiple gene editing in Brassica oleracea var. capitata using the endogenous tRNA-processing system. Hortic. Res. 2019, 6, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Murovec, J.; Guček, K.; Bohanec, B.; Avbelj, M.; Jerala, R. DNA-Free Genome Editing of Brassica oleracea and B. rapa Protoplasts Using CRISPR-Cas9 Ribonucleoprotein Complexes. Front. Plant Sci. 2018, 9, 1594. [Google Scholar] [CrossRef] [PubMed]
- Schouten, H.J.; Tikunov, Y.; Verkerke, W.; Finkers, R.; Bovy, A.; Bai, Y.; Visser, R.G. Breeding Has Increased the Diversity of Cultivated Tomato in The Netherlands. Front. Plant Sci. 2019, 10, 1606. [Google Scholar] [CrossRef]
- Luby, J.J.; Shaw, D.V. Plant Breeders’ Perspectives on Improving Yield and Quality Traits in Horticultural Food Crops. HortScience 2009, 44, 20–22. [Google Scholar] [CrossRef]
- Van Vu, T.; Das, S.; Tran, M.T.; Hong, J.C.; Kim, J.-Y. Precision Genome Engineering for the Breeding of Tomatoes: Recent Progress and Future Perspectives. Front. Genome Ed. 2020, 2, 2. [Google Scholar] [CrossRef]
- Dias, J.S.; Ortiz, R. Transgenic Vegetable Breeding for Nutritional Quality and Health Benefits. Food Nutr. Sci. 2012, 3, 1209–1219. [Google Scholar] [CrossRef]
- Kato-Nitta, N.; Maeda, T.; Inagaki, Y.; Tachikawa, M. Expert and public perceptions of gene-edited crops: Attitude changes in relation to scientific knowledge. Palgrave Commun. 2019, 5, 1–14. [Google Scholar] [CrossRef]
- Jorasch, P. Potential, Challenges, and Threats for the Application of New Breeding Techniques by the Private Plant Breeding Sector in the EU. Front. Plant Sci. 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- Jorasch, P. Will the EU stay out of step with science and the rest of the world on plant breeding innovation? Plant Cell Rep. 2019, 39, 163–167. [Google Scholar] [CrossRef] [PubMed]
- Lassoued, R.; Phillips, P.W.; Macall, D.M.; Hesseln, H.; Smyth, S.J. Expert opinions on the regulation of plant genome editing. Plant Biotechnol. J. 2021, 19, 1104–1109. [Google Scholar] [CrossRef]
- Anders, S.; Cowling, W.; Pareek, A.; Gupta, K.J.; Singla-Pareek, S.L.; Foyer, C.H. Gaining Acceptance of Novel Plant Breeding Technologies. Trends Plant Sci. 2021, 26, 575–587. [Google Scholar] [CrossRef]
- Miladinovic, D.; Antunes, D.; Yildirim, K.; Bakhsh, A.; Cvejić, S.; Kondić-Špika, A.; Jeromela, A.M.; Opsahl-Sorteberg, H.-G.; Zambounis, A.; Hilioti, Z. Targeted plant improvement through genome editing: From laboratory to field. Plant Cell Rep. 2021, 40, 935–951. [Google Scholar] [CrossRef] [PubMed]


| Species | Accession/Cultivar/Landrace | Sequencing Technology | Genome Size | References |
|---|---|---|---|---|
| Solanum lycopersicum | Cultivar ‘Heinz 1706’ | Sanger and ‘next generation’ technologies’ (454/Roche GS FLX, SOLiD™ sequencing, Illumina sequencing) | 900 Mb | [28] |
| S. lycopericum | Varieties San Marzano and Vesuviano | Illumina HiSeq | 900 Mb | [63] |
| S. lycopersicum S. pimpinellifolium S. cheesmaniae, S. galapagense | Pan-genome of 725 accessions belonging to wild species, early domesticates, large-fruited heirlooms, modern elite cultivars and hybrids | Illumina NextSeq | Total of 306 Gb of contigs | [27] |
| S. pennellii | LA0716 | Illumina | 942 Mb | [88] |
| S. pimpinellifolium | LA2093 | Illumina | 923 Mb | [89] |
| S. pimpinellifolium | LA0480 | Illumina HiSeq2000 | 811 Mb | [90] |
| B. oleracea var. botrytis | Inbred line C-8 | PacBio and Illumina HiSeq X Ten | 584.60 Mb | [91] |
| B. rapa ssp. pekinensis | Line Chiifu-401-42 | Illumina GA II | 288 Mb | [92] |
| B. juncea var. tumida | Variety T84−66 | Illumina | 955 Mb | [93] |
| B. nigra | Double haploid line YZ12151 | Illumina | 591 Mb | [93] |
| B. napus | Cultivar Tapidor | Illumina | 635 Mb | [94] |
| Species | Traits | Metabolic Pathways | Target Genes | Genome Editing System | Transformation Technology | References |
|---|---|---|---|---|---|---|
| Solanum lycopersicum | Fruit colour | Carotenoids | SGR1, LCY-E, LCY-B1, LCY-B2, Blc | CRISPR/Cas9 | Agrobacterium tumefaciens-mediated | [110] |
| Carotenoids | Psy1 | CRISPR/Cas9 | A. tumefaciens-mediated | [111] | ||
| Flavonoids | MYB12 | CRISPR/Cas9 | A. tumefaciens-mediated | [112] | ||
| Anthocyanins | ANT1 | CRISPR/Cas9 | A. tumefaciens-mediated | [113] | ||
| GABA content in fruits | aminobutiric acid | GAD2, GAD3 | CRISPR/Cas9 | A. tumefaciens-mediated | [114] | |
| aminobutiric acid | GABA-TP1, GABA-TP2, GABA-TP3, CAT9, SSADH | CRISPR/Cas9 | A. tumefaciens-mediated | [115] | ||
| Metabolite content | SSC, fiber, fructose, ascorbic acid, total phenol, carotene, oxalic acid | L1L4 | ZFN | A. tumefaciens-mediated | [116] | |
| Ripening | Ripening regulatory system | RIN | CRISPR/Cas9 | A. tumefaciens | [117,118,119] | |
| Ripening regulatory system | NOR-like1 | CRISPR/Cas9 | A. tumefaciens-mediated | [120] | ||
| Ripening regulatory system | SBP-CNR, NAC-NOR | CRISPR/Cas9 | A. tumefaciens-mediated | [27] | ||
| Ripening regulatory system | AP2a, MAC-NOR, FUL1/TDR4, FUL2/MBP7 | CRISPR/Cas9 | A. tumefaciens-mediated | [121] | ||
| Ripening regulatory system | DML2 | CRISPR/Cas9 | A. tumefaciens-mediated | [122] | ||
| Ripening regulatory system | ORRM4 | CRISPR/Cas9 | A. tumefaciens-mediated | [123] | ||
| Ripening regulatory system | lncRNA1459 | CRISPR/Cas9 | A. tumefaciens-mediated | [124] | ||
| Fruit softening | PL | CRISPR/Cas9 | A. tumefaciens-mediated | [125] | ||
| Fruit softening | PL, PG2a, TBG4 | CRISPR/Cas9 | A. tumefaciens-mediated | [126] | ||
| Shelf life | Alc | CRISPR/Cas9 | A. tumefaciens-mediated | [127] | ||
| S. pimpinellifolium | Metabolite content | Lycopene | CycB | CRISPR/Cas9 | A. tumefaciens-mediated | [128] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Natalini, A.; Acciarri, N.; Cardi, T. Breeding for Nutritional and Organoleptic Quality in Vegetable Crops: The Case of Tomato and Cauliflower. Agriculture 2021, 11, 606. https://doi.org/10.3390/agriculture11070606
Natalini A, Acciarri N, Cardi T. Breeding for Nutritional and Organoleptic Quality in Vegetable Crops: The Case of Tomato and Cauliflower. Agriculture. 2021; 11(7):606. https://doi.org/10.3390/agriculture11070606
Chicago/Turabian StyleNatalini, Alessandro, Nazzareno Acciarri, and Teodoro Cardi. 2021. "Breeding for Nutritional and Organoleptic Quality in Vegetable Crops: The Case of Tomato and Cauliflower" Agriculture 11, no. 7: 606. https://doi.org/10.3390/agriculture11070606
APA StyleNatalini, A., Acciarri, N., & Cardi, T. (2021). Breeding for Nutritional and Organoleptic Quality in Vegetable Crops: The Case of Tomato and Cauliflower. Agriculture, 11(7), 606. https://doi.org/10.3390/agriculture11070606






