Liquid Biopsy is Instrumental for 3PM Dimensional Solutions in Cancer Management
Abstract
1. Introduction
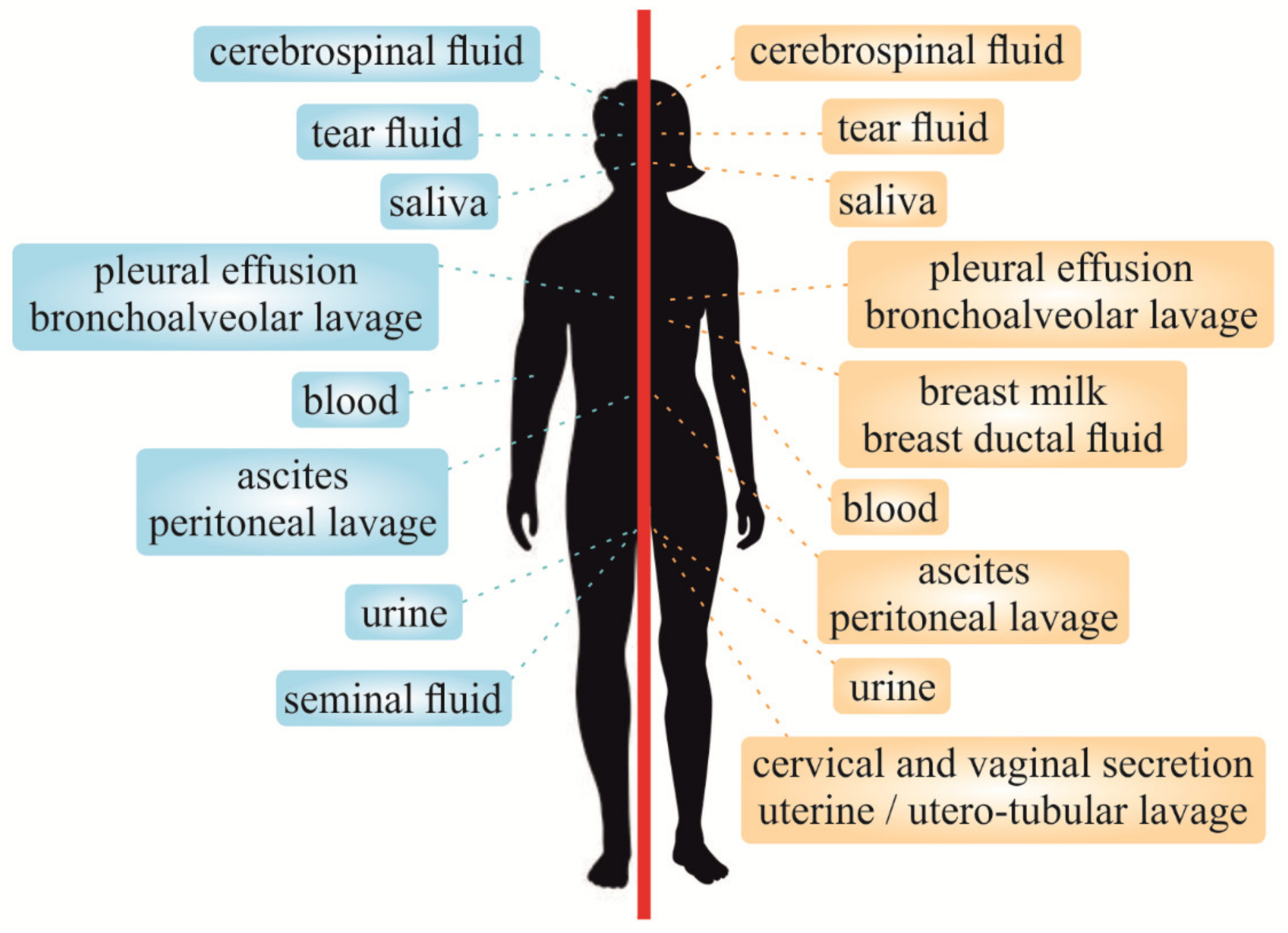
2. Liquid Biopsy Classification
3. Liquid Biopsy Analysis: Advanced Technologies Manifest New Horizons in Cancer Management
3.1. Real-Time PCR
3.2. Digital PCR
3.3. Next-Generation Sequencing
3.4. Proteomics Analyses of Liquid Biopsy
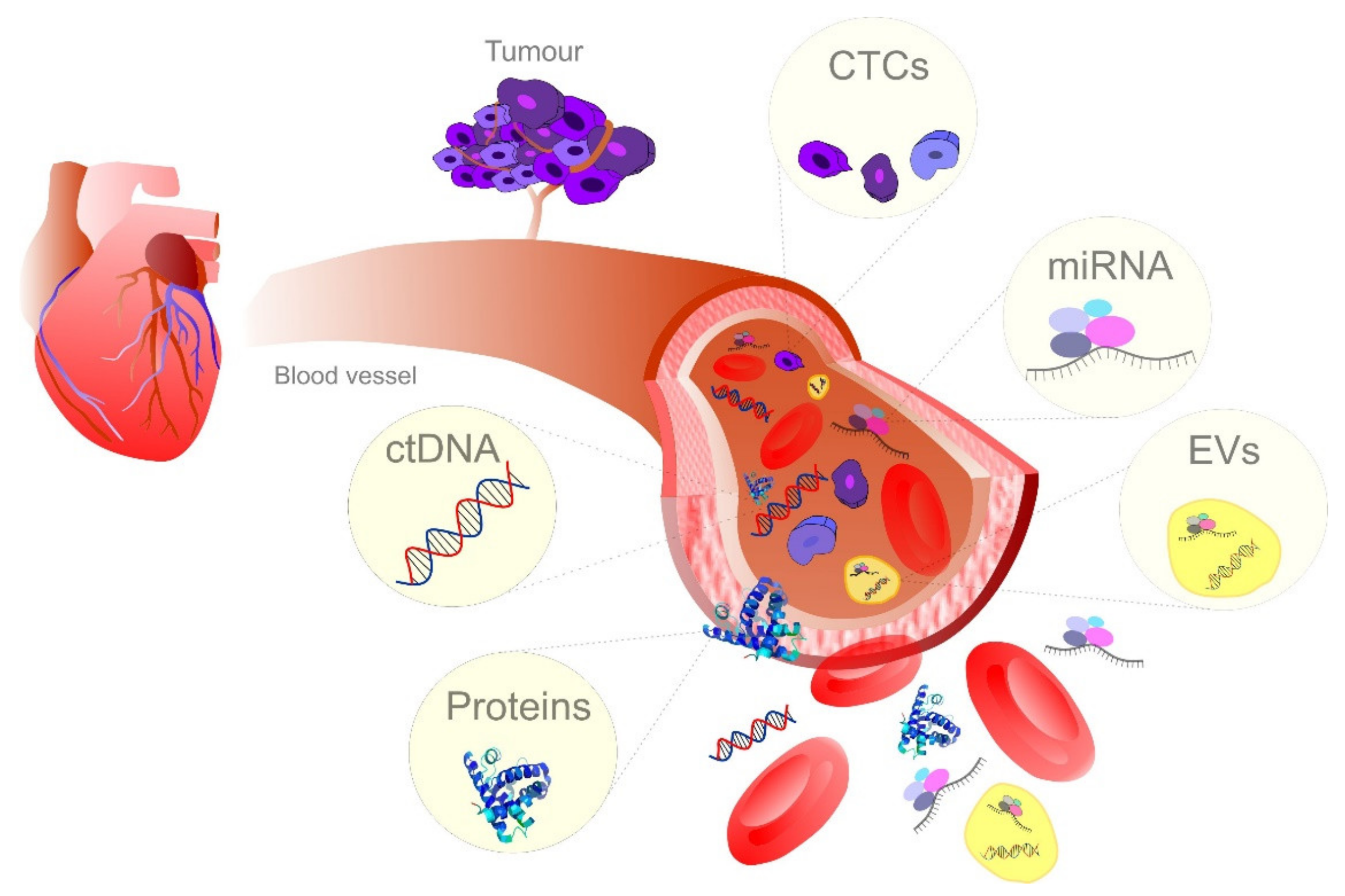
4. Blood Samples as Currently Most Frequently Used Liquid Biopsy in Cancer Diagnostics
4.1. Circulating Tumor DNA
4.2. Circulating Cell-Free miRNAs
4.3. Extracellular Vesicles in Liquid Biopsy
4.4. The Significance of CTCs in Blood-Based Liquid Biopsy for Cancer Management
4.5. Other Biomarkers of Blood-Based Liquid Biopsy or Their Combinations
5. Other Liquid Biopsy Types
5.1. Urine-Based Liquid Biopsy
5.2. Salivary Liquid Biopsy
5.3. Cerebrospinal Fluid-Based Liquid Biopsy
5.4. Liquid Biopsy Based on Other Bio-Fluids
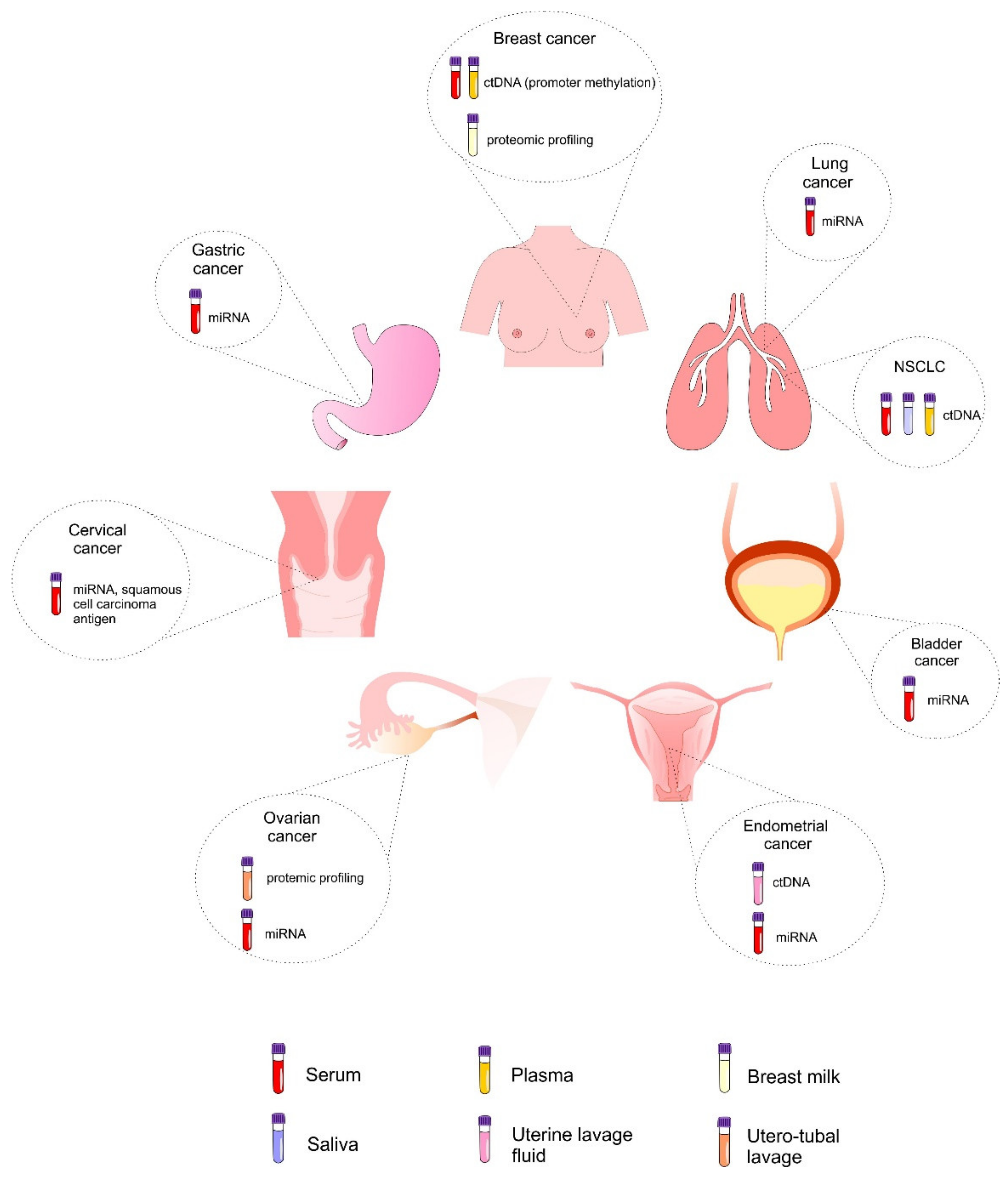
6. Liquid Biopsy as a Tool for Screening and Early Cancer Diagnosis
7. Concluding Remarks and 3PM-Related Expert Recommendations
Author Contributions
Funding
Conflicts of Interest
References
- Cancer. Available online: https://www.who.int/health-topics/cancer#tab=tab_1 (accessed on 15 July 2020).
- Hofmarcher, T.; Brådvik, G.; Svedman, C.; Lindgren, P.; Jönsson, B.; Wilking, N. Comparator Report on Cancer in Europe 2019. Processes 2019, 230, 747. [Google Scholar]
- Golubnitschaja, O.; Baban, B.; Boniolo, G.; Wang, W.; Bubnov, R.; Kapalla, M.; Krapfenbauer, K.; Mozaffari, M.S.; Costigliola, V. Medicine in the early twenty-first century: Paradigm and anticipation—EPMA position paper 2016. EPMA J. 2016, 7, 23. [Google Scholar] [CrossRef] [PubMed]
- Grech, G.; Zhan, X.; Yoo, B.C.; Bubnov, R.; Hagan, S.; Danesi, R.; Vittadini, G.; Desiderio, D.M. EPMA position paper in cancer: Current overview and future perspectives. EPMA J. 2015, 6, 9. [Google Scholar] [CrossRef] [PubMed]
- Kucera, R.; Pecen, L.; Topolcan, O.; Dahal, A.R.; Costigliola, V.; Giordano, F.A.; Golubnitschaja, O. Prostate cancer management: Long-term beliefs, epidemic developments in the early twenty-first century and 3PM dimensional solutions. EPMA J. 2020. [Google Scholar] [CrossRef]
- Mandel, P.C.; Huland, H.; Tiebel, A.; Haese, A.; Salomon, G.; Budäus, L.; Tilki, D.; Chun, F.; Heinzer, H.; Graefen, M.; et al. Enumeration and Changes in Circulating Tumor Cells and Their Prognostic Value in Patients Undergoing Cytoreductive Radical Prostatectomy for Oligometastatic Prostate Cancer-Translational Research Results from the Prospective ProMPT trial. Eur. Urol. Focus 2019. [Google Scholar] [CrossRef]
- Gerner, C.; Costigliola, V.; Golubnitschaja, O. Multiomic patterns in body fluids: Technological challenge with a great potential to implement the advances paradigm of 3P medicine. Mass Spectrom. Rev. 2019. [Google Scholar] [CrossRef]
- Golubnitschaja, O.; Polivka, J.; Yeghiazaryan, K.; Berliner, L. Liquid biopsy and multiparametric analysis in management of liver malignancies: New concepts of the patient stratification and prognostic approach. EPMA J. 2018, 9, 271–285. [Google Scholar] [CrossRef]
- Sakuma, Y.; Fujii, K.; Han, J.; Takahashi, R. Recent Advances in Liquid Biopsy Based on Circulating Tumor DNA. J. Clin. Med. 2019, 8, 1957. [Google Scholar] [CrossRef]
- Casas-Arozamena, C.; Díaz, E.; Moiola, C.P.; Alonso-Alconada, L.; Ferreiros, A.; Abalo, A.; López Gil, C.; Oltra, S.S.; de Santiago, J.; Cabrera, S.; et al. Genomic Profiling of Uterine Aspirates and cfDNA as an Integrative Liquid Biopsy Strategy in Endometrial Cancer. J. Clin. Med. 2020, 9, 585. [Google Scholar] [CrossRef]
- Rapado-González, Ó.; Majem, B.; Álvarez-Castro, A.; Díaz-Peña, R.; Abalo, A.; Suárez-Cabrera, L.; Gil-Moreno, A.; Santamaría, A.; López-López, R.; Muinelo-Romay, L.; et al. A Novel Saliva-Based miRNA Signature for Colorectal Cancer Diagnosis. J. Clin. Med. 2019, 8, 2029. [Google Scholar] [CrossRef]
- Eisenstein, M. Could liquid biopsies help deliver better treatment? Nature 2020, 579, S6–S8. [Google Scholar] [CrossRef] [PubMed]
- Miranda-Castro, R.; Palchetti, I.; de-los-Santos-Álvarez, N. The Translational Potential of Electrochemical DNA-Based Liquid Biopsy. Front. Chem. 2020, 8. [Google Scholar] [CrossRef] [PubMed]
- Van Paemel, R.; Vlug, R.; De Preter, K.; Van Roy, N.; Speleman, F.; Willems, L.; Lammens, T.; Laureys, G.; Schleiermacher, G.; Tytgat, G.A.M.; et al. The pitfalls and promise of liquid biopsies for diagnosing and treating solid tumors in children: A review. Eur. J. Pediatr. 2020, 179, 191–202. [Google Scholar] [CrossRef] [PubMed]
- Gu, X.; He, J.; Ji, G. Combined use of circulating tumor cells and salivary mRNA to detect non–small-cell lung cancer. Medicine 2020, 99. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Lázaro, D.; García Hernández, J.L.; García, A.C.; Córdova Martínez, A.; Mielgo-Ayuso, J.; Cruz-Hernández, J.J. Liquid Biopsy as Novel Tool in Precision Medicine: Origins, Properties, Identification and Clinical Perspective of Cancer’s Biomarkers. Diagnostics 2020, 10, 215. [Google Scholar] [CrossRef]
- Khatami, F.; Larijani, B.; Nasiri, S.; Tavangar, S.M. Liquid Biopsy as a Minimally Invasive Source of Thyroid Cancer Genetic and Epigenetic Alterations. Int. J. Mol. Cell. Med. 2019, 8, 19–29. [Google Scholar] [CrossRef]
- Castro-Giner, F.; Gkountela, S.; Donato, C.; Alborelli, I.; Quagliata, L.; Ng, C.K.Y.; Piscuoglio, S.; Aceto, N. Cancer Diagnosis Using a Liquid Biopsy: Challenges and Expectations. Diagnostics 2018, 8, 31. [Google Scholar] [CrossRef]
- Bai, Y.; Zhao, H. Liquid biopsy in tumors: Opportunities and challenges. Ann. Transl. Med. 2018, 6. [Google Scholar] [CrossRef]
- Alix-Panabières, C.; Pantel, K. Circulating Tumor Cells: Liquid Biopsy of Cancer. Clin. Chem. 2013, 59, 110–118. [Google Scholar] [CrossRef]
- Thomas, M.L.; Marcato, P. Epigenetic Modifications as Biomarkers of Tumor Development, Therapy Response, and Recurrence across the Cancer Care Continuum. Cancers 2018, 10, 101. [Google Scholar] [CrossRef]
- Kawaguchi, T.; Komatsu, S.; Ichikawa, D.; Tsujiura, M.; Takeshita, H.; Hirajima, S.; Miyamae, M.; Okajima, W.; Ohashi, T.; Imamura, T.; et al. Circulating MicroRNAs: A Next-Generation Clinical Biomarker for Digestive System Cancers. Int. J. Mol. Sci. 2016, 17, 1459. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Wang, D. Liquid Biopsy in the OMICS Era of Tumor Medicine. Open Access J. Biomed. Eng. Appl. 2018, 1, 115. [Google Scholar] [PubMed]
- Skotland, T.; Ekroos, K.; Kauhanen, D.; Simolin, H.; Seierstad, T.; Berge, V.; Sandvig, K.; Llorente, A. Molecular lipid species in urinary exosomes as potential prostate cancer biomarkers. Eur. J. Cancer 2017, 70, 122–132. [Google Scholar] [CrossRef] [PubMed]
- Ballester, L.Y.; Lu, G.; Zorofchian, S.; Vantaku, V.; Putluri, V.; Yan, Y.; Arevalo, O.; Zhu, P.; Riascos, R.F.; Sreekumar, A.; et al. Analysis of cerebrospinal fluid metabolites in patients with primary or metastatic central nervous system tumors. Acta Neuropathol. Commun. 2018, 6, 85. [Google Scholar] [CrossRef] [PubMed]
- Saarenheimo, J.; Eigeliene, N.; Andersen, H.; Tiirola, M.; Jekunen, A. The Value of Liquid Biopsies for Guiding Therapy Decisions in Non-small Cell Lung Cancer. Front. Oncol. 2019, 9. [Google Scholar] [CrossRef] [PubMed]
- De Luca, F.; Rotunno, G.; Salvianti, F.; Galardi, F.; Pestrin, M.; Gabellini, S.; Simi, L.; Mancini, I.; Vannucchi, A.M.; Pazzagli, M.; et al. Mutational analysis of single circulating tumor cells by next generation sequencing in metastatic breast cancer. Oncotarget 2016, 7, 26107–26119. [Google Scholar] [CrossRef] [PubMed]
- Jain, S.; Lin, S.Y.; Song, W.; Su, Y.-H. Urine-Based Liquid Biopsy for Nonurological Cancers. Genet. Test Mol. Biomark. 2019, 23, 277–283. [Google Scholar] [CrossRef]
- Siravegna, G.; Marsoni, S.; Siena, S.; Bardelli, A. Integrating liquid biopsies into the management of cancer. Nat. Rev. Clin. Oncol. 2017, 14, 531–548. [Google Scholar] [CrossRef]
- Nakano, M.; Ito, M.; Tanaka, R.; Yamaguchi, K.; Ariyama, H.; Mitsugi, K.; Yoshihiro, T.; Ohmura, H.; Tsuruta, N.; Hanamura, F.; et al. PD-1+ TIM-3+ T cells in malignant ascites predict prognosis of gastrointestinal cancer. Cancer Sci. 2018, 109, 2986–2992. [Google Scholar] [CrossRef]
- Mannello, F. New horizon for breast cancer biomarker discoveries: What might the liquid biopsy of nipple aspirate fluid hold? Proteom. Clin. Appl. 2017, 11. [Google Scholar] [CrossRef]
- Halvaei, S.; Daryani, S.; Zahra, E.-S.; Samadi, T.; Jafarbeik-Iravani, N.; Bakhshayesh, T.O.; Majidzadeh, A.K.; Esmaeili, R. Exosomes in Cancer Liquid Biopsy: A Focus on Breast Cancer. Mol. Ther. Nucleic Acids 2018, 10, 131–141. [Google Scholar] [CrossRef] [PubMed]
- Ponti, G.; Maccaferri, M.; Manfredini, M.; Micali, S.; Torricelli, F.; Milandri, R.; Del Prete, C.; Ciarrocchi, A.; Ruini, C.; Benassi, L.; et al. Quick assessment of cell-free DNA in seminal fluid and fragment size for early non-invasive prostate cancer diagnosis. Clin. Chim. Acta 2019, 497, 76–80. [Google Scholar] [CrossRef] [PubMed]
- Prigenzi, K.C.K.; Heinke, T.; Salim, R.C.; Focchi, G.R. de A. Dual p16 and Ki-67 Expression in Liquid-Based Cervical Cytological Samples Compared to Pap Cytology Findings, Biopsies, and HPV Testing in Cervical Cancer Screening: A Diagnostic Accuracy Study. Acta Cytol. 2018, 62, 104–114. [Google Scholar] [CrossRef] [PubMed]
- Barnabas, G.D.; Bahar-Shany, K.; Sapoznik, S.; Helpman, L.; Kadan, Y.; Beiner, M.; Weitzner, O.; Arbib, N.; Korach, J.; Perri, T.; et al. Microvesicle Proteomic Profiling of Uterine Liquid Biopsy for Ovarian Cancer Early Detection. Mol. Cell Proteom. 2019, 18, 865–875. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Li, L.; Douville, C.; Cohen, J.D.; Yen, T.-T.; Kinde, I.; Sundfelt, K.; Kjær, S.K.; Hruban, R.H.; Shih, I.-M.; et al. Evaluation of liquid from the Papanicolaou test and other liquid biopsies for the detection of endometrial and ovarian cancers. Sci. Transl. Med. 2018, 10. [Google Scholar] [CrossRef] [PubMed]
- Domagala-Kulawik, J. New Frontiers for Molecular Pathology. Front. Med. 2019, 6, 284. [Google Scholar] [CrossRef]
- Roman-Canal, B.; Tarragona, J.; Moiola, C.P.; Gatius, S.; Bonnin, S.; Ruiz-Miró, M.; Sierra, J.E.; Rufas, M.; González, E.; Porcel, J.M.; et al. EV-associated miRNAs from peritoneal lavage as potential diagnostic biomarkers in colorectal cancer. J. Transl. Med. 2019, 17, 208. [Google Scholar] [CrossRef]
- García-Romero, N.; Carrión-Navarro, J.; Areal-Hidalgo, P.; Ortiz de Mendivil, A.; Asensi-Puig, A.; Madurga, R.; Núñez-Torres, R.; González-Neira, A.; Belda-Iniesta, C.; González-Rumayor, V.; et al. BRAF V600E Detection in Liquid Biopsies from Pediatric Central Nervous System Tumors. Cancers 2019, 12, 66. [Google Scholar] [CrossRef]
- Augustus, E.; Van Casteren, K.; Sorber, L.; van Dam, P.; Roeyen, G.; Peeters, M.; Vorsters, A.; Wouters, A.; Raskin, J.; Rolfo, C.; et al. The art of obtaining a high yield of cell-free DNA from urine. PLoS ONE 2020, 15. [Google Scholar] [CrossRef]
- Grölz, D.; Hauch, S.; Schlumpberger, M.; Guenther, K.; Voss, T.; Sprenger-Haussels, M.; Oelmüller, U. Liquid Biopsy Preservation Solutions for Standardized Pre-Analytical Workflows—Venous Whole Blood and Plasma. Curr. Pathobiol. Rep. 2018, 6, 275–286. [Google Scholar] [CrossRef]
- Martignano, F. Cell-Free DNA: An Overview of Sample Types and Isolation Procedures. Methods Mol. Biol. 2019, 1909, 13–27. [Google Scholar] [CrossRef] [PubMed]
- Hench, I.B.; Hench, J.; Tolnay, M. Liquid Biopsy in Clinical Management of Breast, Lung, and Colorectal Cancer. Front. Med. 2018, 5. [Google Scholar] [CrossRef] [PubMed]
- Heitzer, E.; Ulz, P.; Geigl, J.B. Circulating Tumor DNA as a Liquid Biopsy for Cancer. Clin. Chem. 2015, 61, 112–123. [Google Scholar] [CrossRef] [PubMed]
- Poulet, G.; Massias, J.; Taly, V. Liquid Biopsy: General Concepts. Acta Cytol. 2019, 63, 449–455. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Zhao, H. Next-generation sequencing in liquid biopsy: Cancer screening and early detection. Hum. Genom. 2019, 13, 34. [Google Scholar] [CrossRef]
- Elazezy, M.; Joosse, S.A. Techniques of using circulating tumor DNA as a liquid biopsy component in cancer management. Comput. Struct. Biotechnol. J. 2018, 16, 370–378. [Google Scholar] [CrossRef]
- DeVos, T.; Tetzner, R.; Model, F.; Weiss, G.; Schuster, M.; Distler, J.; Steiger, K.V.; Grützmann, R.; Pilarsky, C.; Habermann, J.K.; et al. Circulating methylated SEPT9 DNA in plasma is a biomarker for colorectal cancer. Clin. Chem. 2009, 55, 1337–1346. [Google Scholar] [CrossRef]
- Al-Sheikh, Y.A.; Ghneim, H.K.; Softa, K.I.; Al-Jobran, A.A.; Al-Obeed, O.; Mohamed, M.A.V.; Abdulla, M.; Aboul-Soud, M.A.M. Expression profiling of selected microRNA signatures in plasma and tissues of Saudi colorectal cancer patients by qPCR. Oncol. Lett. 2016, 11, 1406–1412. [Google Scholar] [CrossRef]
- Gorur, A.; Balci Fidanci, S.; Dogruer Unal, N.; Ayaz, L.; Akbayir, S.; Yildirim Yaroglu, H.; Dirlik, M.; Serin, M.S.; Tamer, L. Determination of plasma microRNA for early detection of gastric cancer. Mol. Biol. Rep. 2013, 40, 2091–2096. [Google Scholar] [CrossRef]
- Akers, J.C.; Ramakrishnan, V.; Kim, R.; Skog, J.; Nakano, I.; Pingle, S.; Kalinina, J.; Hua, W.; Kesari, S.; Mao, Y.; et al. MiR-21 in the extracellular vesicles (EVs) of cerebrospinal fluid (CSF): A platform for glioblastoma biomarker development. PLoS ONE 2013, 8, e78115. [Google Scholar] [CrossRef]
- Drusco, A.; Bottoni, A.; Laganà, A.; Acunzo, M.; Fassan, M.; Cascione, L.; Antenucci, A.; Kumchala, P.; Vicentini, C.; Gardinam, M.P.; et al. A differentially expressed set of microRNAs in cerebro-spinal fluid (CSF) can diagnose CNS malignancies. Oncotarget 2015, 6, 20829–20839. [Google Scholar] [CrossRef] [PubMed]
- Mengual, L.; Burset, M.; Ribal, M.J.; Ars, E.; Marín-Aguilera, M.; Fernández, M.; Ingelmo-Torres, M.; Villavicencio, H.; Alcaraz, A. Gene expression signature in urine for diagnosing and assessing aggressiveness of bladder urothelial carcinoma. Clin. Cancer Res. 2010, 16, 2624–2633. [Google Scholar] [CrossRef] [PubMed]
- Yun, S.J.; Jeong, P.; Kim, W.-T.; Kim, T.H.; Lee, Y.-S.; Song, P.H.; Choi, Y.-H.; Kim, I.Y.; Moon, S.-K.; Kim, W.-J. Cell-free microRNAs in urine as diagnostic and prognostic biomarkers of bladder cancer. Int. J. Oncol. 2012, 41, 1871–1878. [Google Scholar] [CrossRef] [PubMed]
- Kim, W.T.; Jeong, P.; Yan, C.; Kim, Y.H.; Lee, I.-S.; Kang, H.-W.; Kim, Y.-J.; Lee, S.-C.; Kim, S.J.; Kim, Y.T.; et al. UBE2C cell-free RNA in urine can discriminate between bladder cancer and hematuria. Oncotarget 2016, 7, 58193–58202. [Google Scholar] [CrossRef] [PubMed]
- Park, N.J.; Zhou, H.; Elashoff, D.; Henson, B.S.; Kastratovic, D.A.; Abemayor, E.; Wong, D.T. Salivary microRNA: Discovery, characterization, and clinical utility for oral cancer detection. Clin. Cancer Res. 2009, 15, 5473–5477. [Google Scholar] [CrossRef] [PubMed]
- Xie, Z.; Chen, G.; Zhang, X.; Li, D.; Huang, J.; Yang, C.; Zhang, P.; Qin, Y.; Duan, Y.; Gong, B.; et al. Salivary MicroRNAs as Promising Biomarkers for Detection of Esophageal Cancer. PLoS ONE 2013, 8. [Google Scholar] [CrossRef] [PubMed]
- Xie, Z.-J.; Chen, G.; Zhang, X.-C.; Li, D.-F.; Huang, J.; Li, Z.-J. Saliva supernatant miR-21: A novel potential biomarker for esophageal cancer detection. Asian Pac. J. Cancer Prev. 2012, 13, 6145–6149. [Google Scholar] [CrossRef]
- Bao, Q.; Xu, Y.; Ding, M.; Chen, P. Identification of differentially expressed miRNAs in differentiating benign from malignant pleural effusion. Hereditas 2020, 157. [Google Scholar] [CrossRef]
- Quan, P.-L.; Sauzade, M.; Brouzes, E. dPCR: A Technology Review. Sensors 2018, 18, 1271. [Google Scholar] [CrossRef]
- Van Ginkel, J.H.; Huibers, M.M.H.; van Es, R.J.J.; de Bree, R.; Willems, S.M. Droplet digital PCR for detection and quantification of circulating tumor DNA in plasma of head and neck cancer patients. BMC Cancer 2017, 17, 428. [Google Scholar] [CrossRef]
- Burjanivova, T.; Malicherova, B.; Grendar, M.; Minarikova, E.; Dusenka, R.; Vanova, B.; Bobrovska, M.; Pecova, T.; Homola, I.; Lasabova, Z.; et al. Detection of BRAFV600E Mutation in Melanoma Patients by Digital PCR of Circulating DNA. Genet. Test Mol. Biomark. 2019, 23, 241–245. [Google Scholar] [CrossRef] [PubMed]
- Guibert, N.; Pradines, A.; Farella, M.; Casanova, A.; Gouin, S.; Keller, L.; Favre, G.; Mazieres, J. Monitoring KRAS mutations in circulating DNA and tumor cells using digital droplet PCR during treatment of KRAS-mutated lung adenocarcinoma. Lung Cancer 2016, 100, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Hiemcke-Jiwa, L.S.; Minnema, M.C.; Radersma-van Loon, J.H.; Jiwa, N.M.; de Boer, M.; Leguit, R.J.; de Weger, R.A.; Huibers, M.M.H. The use of droplet digital PCR in liquid biopsies: A highly sensitive technique for MYD88 p.(L265P) detection in cerebrospinal fluid. Hematol. Oncol. 2018, 36, 429–435. [Google Scholar] [CrossRef] [PubMed]
- Russo, I.J.; Ju, Y.; Gordon, N.S.; Zeegers, M.P.; Cheng, K.K.; James, N.D.; Bryan, R.T.; Ward, D.G. Toward Personalised Liquid Biopsies for Urothelial Carcinoma: Characterisation of ddPCR and Urinary cfDNA for the Detection of the TERT 228 G>A/T Mutation. Bladder Cancer 2018, 4, 41–48. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Zhao, J.; Cui, L.; Liu, Y. Urinary circulating DNA detection for dynamic tracking of EGFR mutations for NSCLC patients treated with EGFR-TKIs. Clin. Transl. Oncol. 2017, 19, 332–340. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.W.; Balaj, L.; Liau, L.M.; Samuels, M.L.; Kotsopoulos, S.K.; Maguire, C.A.; LoGuidice, L.; Soto, H.; Garrett, M.; Zhu, L.D.; et al. BEAMing and Droplet Digital PCR Analysis of Mutant IDH1 mRNA in Glioma Patient Serum and Cerebrospinal Fluid Extracellular Vesicles. Mol. Ther. Nucleic Acids 2013, 2. [Google Scholar] [CrossRef]
- García-Foncillas, J.; Alba, E.; Aranda, E.; Díaz-Rubio, E.; López-López, R.; Tabernero, J.; Vivancos, A. Incorporating BEAMing technology as a liquid biopsy into clinical practice for the management of colorectal cancer patients: An expert taskforce review. Ann. Oncol. 2017, 28, 2943–2949. [Google Scholar] [CrossRef]
- Vessies, D.C.L.; Greuter, M.J.E.; van Rooijen, K.L.; Linders, T.C.; Lanfermeijer, M.; Ramkisoensing, K.L.; Meijer, G.A.; Koopman, M.; Coupé, V.M.H.; Vink, G.R.; et al. Performance of four platforms for KRAS mutation detection in plasma cell-free DNA: ddPCR, Idylla, COBAS z480 and BEAMing. Sci. Rep. 2020, 10. [Google Scholar] [CrossRef]
- Li, X.; Liu, Y.; Shi, W.; Xu, H.; Hu, H.; Dong, Z.; Zhu, G.; Sun, Y.; Liu, B.; Gao, H.; et al. Droplet digital PCR improved the EGFR mutation diagnosis with pleural fluid samples in non-small-cell lung cancer patients. Clin. Chim. Acta 2017, 471, 177–184. [Google Scholar] [CrossRef]
- Yang, Y.; Xie, B.; Yan, J. Application of Next-generation Sequencing Technology in Forensic Science. Genomics Proteom. Bioinform. 2014, 12, 190–197. [Google Scholar] [CrossRef]
- Kamps, R.; Brandão, R.D.; Bosch, B.J.v.d.; Paulussen, A.D.C.; Xanthoulea, S.; Blok, M.J.; Romano, A. Next-Generation Sequencing in Oncology: Genetic Diagnosis, Risk Prediction and Cancer Classification. Int. J. Mol. Sci. 2017, 18, 308. [Google Scholar] [CrossRef] [PubMed]
- Abou Daya, S.; Mahfouz, R. Circulating tumor DNA, liquid biopsy, and next generation sequencing: A comprehensive technical and clinical applications review. Meta Gene 2018, 17, 192–201. [Google Scholar] [CrossRef]
- Li, B.T.; Janku, F.; Jung, B.; Hou, C.; Madwani, K.; Alden, R.; Razavi, P.; Reis-Filho, J.S.; Shen, R.; Isbell, J.M.; et al. Ultra-deep next-generation sequencing of plasma cell-free DNA in patients with advanced lung cancers: Results from the Actionable Genome Consortium. Ann. Oncol. 2019, 30, 597–603. [Google Scholar] [CrossRef] [PubMed]
- Ge, M.; Zhan, Q.; Zhang, Z.; Ji, X.; Zhou, X.; Huang, R.; Liang, X. Different next-generation sequencing pipelines based detection of tumor DNA in cerebrospinal fluid of lung adenocarcinoma cancer patients with leptomeningeal metastases. BMC Cancer 2019, 19, 143. [Google Scholar] [CrossRef] [PubMed]
- Zheng, M.-M.; Li, Y.-S.; Jiang, B.-Y.; Tu, H.-Y.; Tang, W.-F.; Yang, J.-J.; Zhang, X.-C.; Ye, J.-Y.; Yan, H.-H.; Su, J.; et al. Clinical Utility of Cerebrospinal Fluid Cell-Free DNA as Liquid Biopsy for Leptomeningeal Metastases in ALK-Rearranged NSCLC. J. Thorac Oncol. 2019, 14, 924–932. [Google Scholar] [CrossRef]
- Tong, L.; Ding, N.; Tong, X.; Li, J.; Zhang, Y.; Wang, X.; Xu, X.; Ye, M.; Li, C.; Wu, X.; et al. Tumor-derived DNA from pleural effusion supernatant as a promising alternative to tumor tissue in genomic profiling of advanced lung cancer. Theranostics 2019, 9, 5532–5541. [Google Scholar] [CrossRef]
- Wu, J.; Hu, S.; Zhang, L.; Xin, J.; Sun, C.; Wang, L.; Ding, K.; Wang, B. Tumor circulome in the liquid biopsies for cancer diagnosis and prognosis. Theranostics 2020, 10, 4544–4556. [Google Scholar] [CrossRef]
- Bratulic, S.; Gatto, F.; Nielsen, J. The Translational Status of Cancer Liquid Biopsies. Regen. Eng. Transl. Med. 2019. [Google Scholar] [CrossRef]
- Long, S.; Qin, Q.; Wang, Y.; Yang, Y.; Wang, Y.; Deng, A.; Qiao, L.; Liu, B. Nanoporous silica coupled MALDI-TOF MS detection of Bence-Jones proteins in human urine for diagnosis of multiple myeloma. Talanta 2019, 200, 288–292. [Google Scholar] [CrossRef]
- Park, H.-G.; Jang, K.-S.; Park, H.-M.; Song, W.-S.; Jeong, Y.-Y.; Ahn, D.-H.; Kim, S.-M.; Yang, Y.-H.; Kim, Y.-G. MALDI-TOF MS-based total serum protein fingerprinting for liver cancer diagnosis. Analyst 2019, 144, 2231–2238. [Google Scholar] [CrossRef]
- Zhao, J.; Fan, Y.-X.; Yang, Y.; Liu, D.-L.; Wu, K.; Wen, F.-B.; Zhang, C.-Y.; Zhu, D.-Y.; Zhao, S. Identification of potential plasma biomarkers for esophageal squamous cell carcinoma by a proteomic method. Int. J. Clin. Exp. Pathol. 2015, 8, 1535–1544. [Google Scholar] [PubMed]
- Ueda, M.; Iguchi, T.; Masuda, T.; Nakahara, Y.; Hirata, H.; Uchi, R.; Niida, A.; Momose, K.; Sakimura, S.; Chiba, K.; et al. Somatic mutations in plasma cell-free DNA are diagnostic markers for esophageal squamous cell carcinoma recurrence. Oncotarget 2016, 7, 62280–62291. [Google Scholar] [CrossRef] [PubMed]
- Marrugo-Ramírez, J.; Mir, M.; Samitier, J. Blood-Based Cancer Biomarkers in Liquid Biopsy: A Promising Non-Invasive Alternative to Tissue Biopsy. Int. J. Mol. Sci. 2018, 19, 2877. [Google Scholar] [CrossRef]
- Crowley, E.; Di Nicolantonio, F.; Loupakis, F.; Bardelli, A. Liquid biopsy: Monitoring cancer-genetics in the blood. Nat. Rev. Clin. Oncol. 2013, 10, 472–484. [Google Scholar] [CrossRef] [PubMed]
- Soliman, S.E.-S.; Alhanafy, A.M.; Habib, M.S.E.; Hagag, M.; Ibrahem, R.A.L. Serum circulating cell free DNA as potential diagnostic and prognostic biomarker in non small cell lung cancer. Biochem. Biophys. Rep. 2018, 15, 45–51. [Google Scholar] [CrossRef]
- Esposito Abate, R.; Pasquale, R.; Sacco, A.; Piccirillo, M.C.; Morabito, A.; Bidoli, P.; Finocchiaro, G.; Chiari, R.; Foltran, L.; Buosi, R.; et al. Liquid Biopsy Testing Can Improve Selection of Advanced Non-Small-Cell Lung Cancer Patients to Rechallenge with Gefitinib. Cancers 2019, 11, 1431. [Google Scholar] [CrossRef]
- Cicchillitti, L.; Corrado, G.; De Angeli, M.; Mancini, E.; Baiocco, E.; Patrizi, L.; Zampa, A.; Merola, R.; Martayan, A.; Conti, L.; et al. Circulating cell-free DNA content as blood based biomarker in endometrial cancer. Oncotarget 2017, 8, 115230–115243. [Google Scholar] [CrossRef]
- Nunes, S.P.; Diniz, F.; Moreira-Barbosa, C.; Constâncio, V.; Silva, A.V.; Oliveira, J.; Soares, M.; Paulino, S.; Cunha, A.L.; Rodrigues, J.; et al. Subtyping Lung Cancer Using DNA Methylation in Liquid Biopsies. J. Clin. Med. 2019, 8, 1500. [Google Scholar] [CrossRef]
- Delmonico, L.; Silva Magalhães Costa, M.A.; Gomes, R.J.; De Oliveira Vieira, P.; Da Silva, A.B.P.; Fournier, M.V.; Scherrer, L.R.; De Azevedo, C.M.; Ornellas, M.H.F.; Alves, G. Methylation profiling in promoter sequences of ATM and CDKN2A (p14ARF/p16INK4a ) genes in blood and cfDNA from women with impalpable breast lesions. Oncol. Lett. 2020, 19, 3003–3010. [Google Scholar] [CrossRef]
- Zhao, A.; Guo, L.; Xu, J.; Zheng, L.; Guo, Z.; Ling, Z.; Wang, L.; Mao, W. Identification and validation of circulating exosomes-based liquid biopsy for esophageal cancer. Cancer Med. 2019, 8, 3566–3574. [Google Scholar] [CrossRef]
- Yang, S.; Che, S.P.Y.; Kurywchak, P.; Tavormina, J.L.; Gansmo, L.B.; Correa de Sampaio, P.; Tachezy, M.; Bockhorn, M.; Gebauer, F.; Haltom, A.R.; et al. Detection of mutant KRAS and TP53 DNA in circulating exosomes from healthy individuals and patients with pancreatic cancer. Cancer Biol. Ther. 2017, 18, 158–165. [Google Scholar] [CrossRef] [PubMed]
- De Miguel Pérez, D.; Rodriguez Martínez, A.; Ortigosa Palomo, A.; Delgado Ureña, M.; Garcia Puche, J.L.; Robles Remacho, A.; Exposito Hernandez, J.; Lorente Acosta, J.A.; Ortega Sánchez, F.G.; Serrano, M.J. Extracellular vesicle-miRNAs as liquid biopsy biomarkers for disease identification and prognosis in metastatic colorectal cancer patients. Sci. Rep. 2020, 10. [Google Scholar] [CrossRef] [PubMed]
- Link, T.; Passek, S.; Wimberger, P.; Frank, K.; Vassileva, Y.D.; Kramer, M.; Kuhlmann, J.D. Serum calretinin as an independent predictor for platinum resistance and prognosis in ovarian cancer. Int. J. Cancer 2020, 146, 2608–2618. [Google Scholar] [CrossRef]
- Yoon, J.H.; Park, Y.G.; Nam, S.W.; Park, W.S. The diagnostic value of serum gastrokine 1 (GKN1) protein in gastric cancer. Cancer Med. 2019, 8, 5507–5514. [Google Scholar] [CrossRef] [PubMed]
- Reddy, D.; Khade, B.; Pandya, R.; Gupta, S. A novel method for isolation of histones from serum and its implications in therapeutics and prognosis of solid tumours. Clin. Epigenet. 2017, 9. [Google Scholar] [CrossRef] [PubMed]
- Gezer, U.; Yörüker, E.E.; Keskin, M.; Kulle, C.B.; Dharuman, Y.; Holdenrieder, S. Histone Methylation Marks on Circulating Nucleosomes as Novel Blood-Based Biomarker in Colorectal Cancer. Int. J. Mol. Sci. 2015, 16, 29654–29662. [Google Scholar] [CrossRef]
- Du, S.; Zhao, Y.; Lv, C.; Wei, M.; Gao, Z.; Meng, X. Applying Serum Proteins and MicroRNA as Novel Biomarkers for Early-Stage Cervical Cancer Detection. Sci. Rep. 2020, 10. [Google Scholar] [CrossRef]
- Bellassai, N.; Spoto, G. Biosensors for liquid biopsy: Circulating nucleic acids to diagnose and treat cancer. Anal. Bioanal. Chem. 2016, 408, 7255–7264. [Google Scholar] [CrossRef]
- Muinelo-Romay, L.; Casas-Arozamena, C.; Abal, M. Liquid Biopsy in Endometrial Cancer: New Opportunities for Personalized Oncology. Int. J. Mol. Sci. 2018, 19, 2311. [Google Scholar] [CrossRef]
- Francis, G.; Stein, S. Circulating Cell-Free Tumour DNA in the Management of Cancer. Int. J. Mol. Sci. 2015, 16, 14122–14142. [Google Scholar] [CrossRef]
- Wu, X.; Li, J.; Gassa, A.; Buchner, D.; Alakus, H.; Dong, Q.; Ren, N.; Liu, M.; Odenthal, M.; Stippel, D.; et al. Circulating tumor DNA as an emerging liquid biopsy biomarker for early diagnosis and therapeutic monitoring in hepatocellular carcinoma. Int. J. Biol. Sci. 2020, 16, 1551–1562. [Google Scholar] [CrossRef] [PubMed]
- Pittella-Silva, F.; Chin, Y.M.; Chan, H.T.; Nagayama, S.; Miyauchi, E.; Low, S.-K.; Nakamura, Y. Plasma or Serum: Which Is Preferable for Mutation Detection in Liquid Biopsy? Clin. Chem. 2020. [Google Scholar] [CrossRef] [PubMed]
- Xu, R.-H.; Wei, W.; Krawczyk, M.; Wang, W.; Luo, H.; Flagg, K.; Yi, S.; Shi, W.; Quan, Q.; Li, K.; et al. Circulating tumour DNA methylation markers for diagnosis and prognosis of hepatocellular carcinoma. Nat. Mat. 2017, 16, 1155–1161. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Zhang, Z.-H.; Wang, S.; Lang, J.-H. Circulating Cell-Free DNA or Circulating Tumor DNA in the Management of Ovarian and Endometrial Cancer. Onco Targets Ther. 2019, 12, 11517–11530. [Google Scholar] [CrossRef] [PubMed]
- Kodahl, A.R.; Ehmsen, S.; Pallisgaard, N.; Jylling, A.M.B.; Jensen, J.D.; Lænkholm, A.; Knoop, A.S.; Ditzel, H.J. Correlation between circulating cell--free PIK3CA tumor DNA levels and treatment response in patients with PIK3CA--mutated metastatic breast cancer. Mol. Oncol. 2018, 12, 925–935. [Google Scholar] [CrossRef]
- Cheung, T.H.; Yim, S.F.; Yu, M.Y.; Worley, M.J.; Fiascone, S.J.; Chiu, R.W.K.; Lo, K.W.K.; Siu, N.S.S.; Wong, M.C.S.; Yeung, A.C.M.; et al. Liquid biopsy of HPV DNA in cervical cancer. J. Clin. Virol. 2019, 114, 32–36. [Google Scholar] [CrossRef]
- Krug, A.K.; Enderle, D.; Karlovich, C.; Priewasser, T.; Bentink, S.; Spiel, A.; Brinkmann, K.; Emenegger, J.; Grimm, D.G.; Castellanos-Rizaldos, E.; et al. Improved EGFR mutation detection using combined exosomal RNA and circulating tumor DNA in NSCLC patient plasma. Ann. Oncol. 2018, 29, 700–706. [Google Scholar] [CrossRef]
- Wang, J.; Han, X.; Sun, Y. DNA methylation signatures in circulating cell-free DNA as biomarkers for the early detection of cancer. Sci. China Life Sci. 2017, 60, 356–362. [Google Scholar] [CrossRef]
- Constâncio, V.; Barros-Silva, D.; Jerónimo, C.; Henrique, R. Known epigenetic biomarkers for prostate cancer detection and management: Exploring the potential of blood-based liquid biopsies. Expert Rev. Mol. Diagn. 2019, 19, 367–375. [Google Scholar] [CrossRef]
- Locke, W.J.; Guanzon, D.; Ma, C.; Liew, Y.J.; Duesing, K.R.; Fung, K.Y.C.; Ross, J.P. DNA Methylation Cancer Biomarkers: Translation to the Clinic. Front. Genet. 2019, 10. [Google Scholar] [CrossRef]
- Mijnes, J.; Tiedemann, J.; Eschenbruch, J.; Gasthaus, J.; Bringezu, S.; Bauerschlag, D.; Maass, N.; Arnold, N.; Weimer, J.; Anzeneder, T.; et al. SNiPER: A novel hypermethylation biomarker panel for liquid biopsy based early breast cancer detection. Oncotarget 2019, 10, 6494–6508. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.C.; Oxnard, G.R.; Klein, E.A.; Swanton, C.; Seiden, M.V.; Liu, M.C.; Oxnard, G.R.; Klein, E.A.; Smith, D.; Richards, D.; et al. Sensitive and specific multi-cancer detection and localization using methylation signatures in cell-free DNA. Ann. Oncol. 2020, 31, 745–759. [Google Scholar] [CrossRef]
- Lu, Y.-T.; Delijani, K.; Mecum, A.; Goldkorn, A. Current status of liquid biopsies for the detection and management of prostate cancer. Cancer Manag. Res. 2019, 11, 5271–5291. [Google Scholar] [CrossRef] [PubMed]
- Glinge, C.; Clauss, S.; Boddum, K.; Jabbari, R.; Jabbari, J.; Risgaard, B.; Tomsits, P.; Hildebrand, B.; Kääb, S.; Wakili, R.; et al. Stability of Circulating Blood-Based MicroRNAs—Pre-Analytic Methodological Considerations. PLoS ONE 2017, 12. [Google Scholar] [CrossRef] [PubMed]
- Fortunato, O.; Gasparini, P.; Boeri, M.; Sozzi, G. Exo-miRNAs as a New Tool for Liquid Biopsy in Lung Cancer. Cancers 2019, 11, 888. [Google Scholar] [CrossRef]
- Ono, A.; Fujimoto, A.; Yamamoto, Y.; Akamatsu, S.; Hiraga, N.; Imamura, M.; Kawaoka, T.; Tsuge, M.; Abe, H.; Hayes, C.N.; et al. Circulating Tumor DNA Analysis for Liver Cancers and Its Usefulness as a Liquid Biopsy. Cell. Mol. Gastroenterol. Hepatol. 2015, 1, 516–534. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Yang, D.; Zhang, H.; Wei, X.; Ma, T.; Cheng, Z.; Hong, Q.; Hu, J.; Zhuo, H.; Song, Y.; et al. Early Detection of Lung Cancer in Serum by a Panel of MicroRNA Biomarkers. Clin. Lung Cancer 2015, 16, 313–319.e1. [Google Scholar] [CrossRef] [PubMed]
- Urabe, F.; Matsuzaki, J.; Yamamoto, Y.; Kimura, T.; Hara, T.; Ichikawa, M.; Takizawa, S.; Aoki, Y.; Niida, S.; Sakamoto, H.; et al. Large-scale Circulating microRNA Profiling for the Liquid Biopsy of Prostate Cancer. Clin. Cancer Res. 2019, 25, 3016–3025. [Google Scholar] [CrossRef]
- Usuba, W.; Urabe, F.; Yamamoto, Y.; Matsuzaki, J.; Sasaki, H.; Ichikawa, M.; Takizawa, S.; Aoki, Y.; Niida, S.; Kato, K.; et al. Circulating miRNA panels for specific and early detection in bladder cancer. Cancer Sci. 2019, 110, 408–419. [Google Scholar] [CrossRef]
- Ritter, A.; Hirschfeld, M.; Berner, K.; Jaeger, M.; Grundner-Culemann, F.; Schlosser, P.; Asberger, J.; Weiss, D.; Noethling, C.; Mayer, S.; et al. Discovery of potential serum and urine-based microRNA as minimally-invasive biomarkers for breast and gynecological cancer. Cancer Biomark. 2020, 27, 225–242. [Google Scholar] [CrossRef]
- Lucchetti, D.; Fattorossi, A.; Sgambato, A. Extracellular Vesicles in Oncology: Progress and Pitfalls in the Methods of Isolation and Analysis. Biotechnol. J. 2019, 14, e1700716. [Google Scholar] [CrossRef] [PubMed]
- Carnino, J.M.; Lee, H.; Jin, Y. Isolation and characterization of extracellular vesicles from Broncho-alveolar lavage fluid: A review and comparison of different methods. Respir. Res. 2019, 20. [Google Scholar] [CrossRef] [PubMed]
- Furi, I.; Momen-Heravi, F.; Szabo, G. Extracellular vesicle isolation: Present and future. Ann. Transl. Med. 2017, 5. [Google Scholar] [CrossRef]
- Nakai, W.; Yoshida, T.; Diez, D.; Miyatake, Y.; Nishibu, T.; Imawaka, N.; Naruse, K.; Sadamura, Y.; Hanayama, R. A novel affinity-based method for the isolation of highly purified extracellular vesicles. Sci. Rep. 2016, 6, 33935. [Google Scholar] [CrossRef] [PubMed]
- Smith, J.T.; Wunsch, B.H.; Dogra, N.; Ahsen, M.E.; Lee, K.; Yadav, K.K.; Weil, R.; Pereira, M.A.; Patel, J.V.; Duch, E.A.; et al. Integrated nanoscale deterministic lateral displacement arrays for separation of extracellular vesicles from clinically-relevant volumes of biological samples. Lab Chip 2018, 18, 3913–3925. [Google Scholar] [CrossRef] [PubMed]
- Amrollahi, P.; Rodrigues, M.; Lyon, C.J.; Goel, A.; Han, H.; Hu, T.Y. Ultra-Sensitive Automated Profiling of EpCAM Expression on Tumor-Derived Extracellular Vesicles. Front. Genet. 2019, 10. [Google Scholar] [CrossRef]
- Lewis, J.M.; Vyas, A.D.; Qiu, Y.; Messer, K.S.; White, R.; Heller, M.J. Integrated Analysis of Exosomal Protein Biomarkers on Alternating Current Electrokinetic Chips Enables Rapid Detection of Pancreatic Cancer in Patient Blood. ACS Nano 2018, 12, 3311–3320. [Google Scholar] [CrossRef]
- Heinzelman, P.; Powers, D.N.; Wohlschlegel, J.A.; John, V. Shotgun Proteomic Profiling of Bloodborne Nanoscale Extracellular Vesicles. Methods Mol. Biol. 2019, 1897, 403–416. [Google Scholar] [CrossRef]
- Tadimety, A.; Closson, A.; Li, C.; Song, Y.; Shen, T.; Zhang, J.X.J. Advances in Liquid Biopsy On-Chip for Cancer Management: Technologies, Biomarkers, and Clinical Analysis. Crit. Rev. Clin. Lab. Sci. 2018, 55, 140–162. [Google Scholar] [CrossRef]
- Sharma, S.; Zhuang, R.; Long, M.; Pavlovic, M.; Kang, Y.; Ilyas, A.; Asghar, W. Circulating Tumor Cell Isolation, Culture, and Downstream Molecular Analysis. Biotechnol. Adv. 2018, 36, 1063–1078. [Google Scholar] [CrossRef]
- Lousada-Fernandez, F.; Rapado-Gonzalez, O.; Lopez-Cedrun, J.-L.; Lopez-Lopez, R.; Muinelo-Romay, L.; Suarez-Cunqueiro, M.M. Liquid Biopsy in Oral Cancer. Int. J. Mol. Sci. 2018, 19, 1704. [Google Scholar] [CrossRef] [PubMed]
- Fabisiewicz, A.; Grzybowska, E. CTC clusters in cancer progression and metastasis. Med. Oncol. 2017, 34, 12. [Google Scholar] [CrossRef] [PubMed]
- Cabel, L.; Proudhon, C.; Gortais, H.; Loirat, D.; Coussy, F.; Pierga, J.-Y.; Bidard, F.-C. Circulating tumor cells: Clinical validity and utility. Int. J. Clin. Oncol. 2017, 22, 421–430. [Google Scholar] [CrossRef] [PubMed]
- Neumann, M.H.D.; Bender, S.; Krahn, T.; Schlange, T. ctDNA and CTCs in Liquid Biopsy—Current Status and Where We Need to Progress. Comput. Struct. Biotechnol. J. 2018, 16, 190–195. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.; Xia, B.-R.; Jin, W.-L.; Lou, G. Circulating tumor cells in precision oncology: Clinical applications in liquid biopsy and 3D organoid model. Cancer Cell Int. 2019, 19, 341. [Google Scholar] [CrossRef] [PubMed]
- Lorente, D.; Olmos, D.; Mateo, J.; Dolling, D.; Bianchini, D.; Seed, G.; Flohr, P.; Crespo, M.; Figueiredo, I.; Miranda, S.; et al. Circulating tumour cell increase as a biomarker of disease progression in metastatic castration-resistant prostate cancer patients with low baseline CTC counts. Ann. Oncol. 2018, 29, 1554–1560. [Google Scholar] [CrossRef]
- Ito, M.; Horimoto, Y.; Tokuda, E.; Murakami, F.; Uomori, T.; Himuro, T.; Nakai, K.; Orihata, G.; Iijima, K.; Saito, M. Impact of circulating tumour cells on survival of eribulin-treated patients with metastatic breast cancer. Med. Oncol. 2019, 36, 89. [Google Scholar] [CrossRef]
- Messaritakis, I.; Politaki, E.; Koinis, F.; Stoltidis, D.; Apostolaki, S.; Plataki, M.; Dermitzaki, E.-K.; Georgoulias, V.; Kotsakis, A. Dynamic changes of phenotypically different circulating tumor cells sub-populations in patients with recurrent/refractory small cell lung cancer treated with pazopanib. Sci. Rep. 2018, 8, 2238. [Google Scholar] [CrossRef]
- Zhang, X.; Li, H.; Yu, X.; Li, S.; Lei, Z.; Li, C.; Zhang, Q.; Han, Q.; Li, Y.; Zhang, K.; et al. Analysis of Circulating Tumor Cells in Ovarian Cancer and Their Clinical Value as a Biomarker. Cell. Physiol. Biochem. 2018, 48, 1983–1994. [Google Scholar] [CrossRef]
- Scharpenseel, H.; Hanssen, A.; Loges, S.; Mohme, M.; Bernreuther, C.; Peine, S.; Lamszus, K.; Goy, Y.; Petersen, C.; Westphal, M.; et al. EGFR and HER3 expression in circulating tumor cells and tumor tissue from non-small cell lung cancer patients. Sci. Rep. 2019, 9. [Google Scholar] [CrossRef]
- Guadagni, S.; Clementi, M.; Masedu, F.; Fiorentini, G.; Sarti, D.; Deraco, M.; Kusamura, S.; Papasotiriou, I.; Apostolou, P.; Aigner, K.R.; et al. A Pilot Study of the Predictive Potential of Chemosensitivity and Gene Expression Assays Using Circulating Tumour Cells from Patients with Recurrent Ovarian Cancer. Int. J. Mol. Sci. 2020, 21, 4813. [Google Scholar] [CrossRef] [PubMed]
- Benini, S.; Gamberi, G.; Cocchi, S.; Garbetta, J.; Alberti, L.; Righi, A.; Gambarotti, M.; Picci, P.; Ferrari, S. Detection of circulating tumor cells in liquid biopsy from Ewing sarcoma patients. Cancer Manag. Res. 2018, 10, 49–60. [Google Scholar] [CrossRef] [PubMed]
- Tsutsuyama, M.; Nakanishi, H.; Yoshimura, M.; Oshiro, T.; Kinoshita, T.; Komori, K.; Shimizu, Y.; Ichinosawa, Y.; Kinuta, S.; Wajima, K.; et al. Detection of circulating tumor cells in drainage venous blood from colorectal cancer patients using a new filtration and cytology-based automated platform. PLoS ONE 2019, 14, e0212221. [Google Scholar] [CrossRef] [PubMed]
- Buderath, P.; Schwich, E.; Jensen, C.; Horn, P.A.; Kimmig, R.; Kasimir-Bauer, S.; Rebmann, V. Soluble Programmed Death Receptor Ligands sPD-L1 and sPD-L2 as Liquid Biopsy Markers for Prognosis and Platinum Response in Epithelial Ovarian Cancer. Front. Oncol. 2019, 9, 1015. [Google Scholar] [CrossRef]
- Marcuello, M.; Duran-Sanchon, S.; Moreno, L.; Lozano, J.J.; Bujanda, L.; Castells, A.; Gironella, M. Analysis of A 6-Mirna Signature in Serum from Colorectal Cancer Screening Participants as Non-Invasive Biomarkers for Advanced Adenoma and Colorectal Cancer Detection. Cancers 2019, 11, 1542. [Google Scholar] [CrossRef]
- Mellby, L.D.; Nyberg, A.P.; Johansen, J.S.; Wingren, C.; Nordestgaard, B.G.; Bojesen, S.E.; Mitchell, B.L.; Sheppard, B.C.; Sears, R.C.; Borrebaeck, C.A.K. Serum Biomarker Signature-Based Liquid Biopsy for Diagnosis of Early-Stage Pancreatic Cancer. J. Clin. Oncol. 2018, 36, 2887–2894. [Google Scholar] [CrossRef]
- Rodrigo, A.; Ojeda, J.L.; Vega, S.; Sanchez-Gracia, O.; Lanas, A.; Isla, D.; Velazquez-Campoy, A.; Abian, O. Thermal Liquid Biopsy (TLB): A Predictive Score Derived from Serum Thermograms as a Clinical Tool for Screening Lung Cancer Patients. Cancers 2019, 11, 1012. [Google Scholar] [CrossRef]
- Villatoro, S.; Mayo-de-Las-Casas, C.; Jordana-Ariza, N.; Viteri-Ramírez, S.; Garzón-Ibañez, M.; Moya-Horno, I.; García-Peláez, B.; González-Cao, M.; Malapelle, U.; Balada-Bel, A.; et al. Prospective detection of mutations in cerebrospinal fluid, pleural effusion, and ascites of advanced cancer patients to guide treatment decisions. Mol. Oncol. 2019, 13, 2633–2645. [Google Scholar] [CrossRef]
- Lebrecht, A.; Boehm, D.; Schmidt, M.; Koelbl, H.; Schwirz, R.L.; Grus, F.H. Diagnosis of breast cancer by tear proteomic pattern. Cancer Genom. Proteom. 2009, 6, 177–182. [Google Scholar]
- Aslebagh, R.; Channaveerappa, D.; Arcaro, K.F.; Darie, C.C. Proteomics analysis of human breast milk to assess breast cancer risk. Electrophoresis 2018, 39, 653–665. [Google Scholar] [CrossRef]
- Ponti, G.; Maccaferri, M.; Mandrioli, M.; Manfredini, M.; Micali, S.; Cotugno, M.; Bianchi, G.; Ozben, T.; Pellacani, G.; Del Prete, C.; et al. Seminal Cell-Free DNA Assessment as a Novel Prostate Cancer Biomarker. Pathol. Oncol. Res. 2018, 24, 941–945. [Google Scholar] [CrossRef] [PubMed]
- Hur, J.Y.; Lee, J.S.; Kim, I.A.; Kim, H.J.; Kim, W.S.; Lee, K.Y. Extracellular vesicle-based EGFR genotyping in bronchoalveolar lavage fluid from treatment-naive non-small cell lung cancer patients. Transl. Lung Cancer Res. 2019, 8, 1051–1060. [Google Scholar] [CrossRef]
- Do Canto, L.M.; Marian, C.; Willey, S.; Sidawy, M.; da Cunha, P.A.; Rone, J.D.; Li, X.; Gusev, Y.; Haddad, B.R. MicroRNA analysis of breast ductal fluid in breast cancer patients. Int. J. Oncol. 2016, 48, 2071–2078. [Google Scholar] [CrossRef]
- Flammer Syndrome: From Phenotype to Associated Pathologies, Prediction, Prevention and Personalisation; Advances in Predictive, Preventive and Personalised Medicine; Golubnitschaja, O., Ed.; Springer International Publishing: Cham, Switzerland, 2019; Volume 11, ISBN 978-3-030-13549-2. [Google Scholar]
- Satyal, U.; Srivastava, A.; Abbosh, P.H. Urine Biopsy—Liquid Gold for Molecular Detection and Surveillance of Bladder Cancer. Front. Oncol. 2019, 9. [Google Scholar] [CrossRef] [PubMed]
- Lu, T.; Li, J. Clinical applications of urinary cell-free DNA in cancer: Current insights and promising future. Am. J. Cancer Res. 2017, 7, 2318–2332. [Google Scholar] [PubMed]
- Christensen, E.; Birkenkamp-Demtröder, K.; Nordentoft, I.; Høyer, S.; van der Keur, K.; van Kessel, K.; Zwarthoff, E.; Agerbæk, M.; Ørntoft, T.F.; Jensen, J.B.; et al. Liquid Biopsy Analysis of FGFR3 and PIK3CA Hotspot Mutations for Disease Surveillance in Bladder Cancer. Eur. Urol. 2017, 71, 961–969. [Google Scholar] [CrossRef]
- Nekrasov, K.A.; Vikarchuk, M.V.; Rudenko, E.E.; Ivanitskiy, I.V.; Grygorenko, V.M.; Danylets, R.O.; Kondratov, A.G.; Stoliar, L.A.; Sharopov, B.R.; Kashuba, V.I. 6-gene promoter methylation assay is potentially applicable for prostate cancer clinical staging based on urine collection following prostatic massage. Oncol. Lett. 2019, 18, 6917–6925. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.; Han, L.; Yuan, J.; Sun, Y. Circulating tumor cell free DNA from plasma and urine in the clinical management of colorectal cancer. Cancer Biomark. 2020, 27, 29–37. [Google Scholar] [CrossRef]
- Wu, Z.; Yang, Z.; Li, C.-S.; Zhao, W.; Liang, Z.-X.; Dai, Y.; Zeng, J.; Zhu, Q.; Miao, K.-L.; Cui, D.-H.; et al. Non-invasive detection of EGFR and TP53 mutations through the combination of plasma, urine and sputum in advanced non-small cell lung cancer. Oncol. Lett. 2019, 18, 3581–3590. [Google Scholar] [CrossRef]
- Mall, C.; Rocke, D.M.; Durbin-Johnson, B.; Weiss, R.H. Stability of miRNA in human urine supports its biomarker potential. Biomark. Med. 2013, 7. [Google Scholar] [CrossRef]
- Li, G.; Zhao, A.; Péoch, M.; Cottier, M.; Mottet, N. Detection of urinary cell-free miR-210 as a potential tool of liquid biopsy for clear cell renal cell carcinoma. Urol. Oncol. 2017, 35, 294–299. [Google Scholar] [CrossRef]
- Pardini, B.; Cordero, F.; Naccarati, A.; Viberti, C.; Birolo, G.; Oderda, M.; Di Gaetano, C.; Arigoni, M.; Martina, F.; Calogero, R.A.; et al. microRNA profiles in urine by next-generation sequencing can stratify bladder cancer subtypes. Oncotarget 2018, 9, 20658–20669. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, A.; Moxley, K.; Ruskin, R.; Dhanasekaran, D.N.; Zhao, Y.D.; Ramesh, R. A Non-invasive Liquid Biopsy Screening of Urine-Derived Exosomes for miRNAs as Biomarkers in Endometrial Cancer Patients. AAPS J. 2018, 20, 82. [Google Scholar] [CrossRef] [PubMed]
- Kaczor-Urbanowicz, K.E.; Wei, F.; Rao, S.L.; Kim, J.; Shin, H.; Cheng, J.; Tu, M.; Wong, D.T.W.; Kim, Y. Clinical validity of saliva and novel technology for cancer detection. Biochim. Biophys. Acta Rev. Cancer 2019, 1872, 49–59. [Google Scholar] [CrossRef] [PubMed]
- Aro, K.; Wei, F.; Wong, D.T.; Tu, M. Saliva Liquid Biopsy for Point-of-Care Applications. Front. Public Health 2017, 5. [Google Scholar] [CrossRef]
- Fröhlich, H.; Patjoshi, S.; Yeghiazaryan, K.; Kehrer, C.; Kuhn, W.; Golubnitschaja, O. Premenopausal breast cancer: Potential clinical utility of a multi-omics based machine learning approach for patient stratification. EPMA J. 2018, 9, 175–186. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Han, B. Liquid Biopsy Promotes Non-Small Cell Lung Cancer Precision Therapy. Technol. Cancer Res. Treat. 2018, 17. [Google Scholar] [CrossRef]
- Shang, X.; Zi, H.; Li, Y.; Gao, Y.; Ge, C.; Sun, Z.; Zhang, Y. Combined use of salivary biomarkers and carcinoembryonic antigen for lung cancer detection in a Chinese population. Medicine 2019, 98. [Google Scholar] [CrossRef]
- Chiabotto, G.; Gai, C.; Deregibus, M.C.; Camussi, G. Salivary Extracellular Vesicle-Associated exRNA as Cancer Biomarker. Cancers 2019, 11, 891. [Google Scholar] [CrossRef]
- Cristaldi, M.; Mauceri, R.; Di Fede, O.; Giuliana, G.; Campisi, G.; Panzarella, V. Salivary Biomarkers for Oral Squamous Cell Carcinoma Diagnosis and Follow-Up: Current Status and Perspectives. Front. Physiol. 2019, 10. [Google Scholar] [CrossRef]
- Machida, T.; Tomofuji, T.; Maruyama, T.; Yoneda, T.; Ekuni, D.; Azuma, T.; Miyai, H.; Mizuno, H.; Kato, H.; Tsutsumi, K.; et al. miR--1246 and miR--4644 in salivary exosome as potential biomarkers for pancreatobiliary tract cancer. Oncol. Rep. 2016, 36, 2375–2381. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Huo, C.; Qiao, Z.; Shang, Z.; Uzzaman, A.; Liu, S.; Jiang, X.; Fan, L.-Y.; Ji, L.; Guan, X.; et al. Comparative Proteomic Analysis of Exosomes and Microvesicles in Human Saliva for Lung Cancer. J. Proteome Res. 2018, 17, 1101–1107. [Google Scholar] [CrossRef]
- Nonaka, T.; Wong, D.T.W. Saliva-Exosomics in Cancer: Molecular Characterization of Cancer-Derived Exosomes in Saliva. Enzymes 2017, 42, 125–151. [Google Scholar] [CrossRef]
- Wang, Z.; Li, F.; Rufo, J.; Chen, C.; Yang, S.; Li, L.; Zhang, J.; Cheng, J.; Kim, Y.; Wu, M.; et al. Acoustofluidic Salivary Exosome Isolation: A Liquid Biopsy Compatible Approach for Human Papillomavirus-Associated Oropharyngeal Cancer Detection. J. Mol. Diagn. 2020, 22, 50–59. [Google Scholar] [CrossRef] [PubMed]
- Tu, M.; Cheng, J.; Chen, Y.-L.; Jea, W.-C.; Chen, W.-L.; Chen, C.-J.; Ho, C.-L.; Huang, W.-L.; Lin, C.-C.; Su, W.-C.; et al. Electric Field-Induced Release and Measurement (EFIRM): Characterization and Technical Validation of a Novel Liquid Biopsy Platform in Plasma and Saliva. J. Mol. Diagn. 2020. [Google Scholar] [CrossRef]
- Mattox, A.K.; Yan, H.; Bettegowda, C. The potential of cerebrospinal fluid-based liquid biopsy approaches in CNS tumors. Neuro-oncology 2019, 21, 1509–1518. [Google Scholar] [CrossRef]
- McEwen, A.E.; Leary, S.E.S.; Lockwood, C.M. Beyond the Blood: CSF-Derived cfDNA for Diagnosis and Characterization of CNS Tumors. Front. Cell Dev. Biol. 2020, 8. [Google Scholar] [CrossRef]
- Ma, C.; Huang, C.; Tang, D.; Ye, X.; Li, Z.; Liu, R.; Mu, N.; Li, J.; Jiang, R.; Zhang, J. Afatinib for Advanced Non-small Cell Lung Cancer in a Case With an Uncommon Epidermal Growth Factor Receptor Mutation (G719A) Identified in the Cerebrospinal Fluid. Front. Oncol. 2019, 9, 628. [Google Scholar] [CrossRef]
- Siravegna, G.; Geuna, E.; Mussolin, B.; Crisafulli, G.; Bartolini, A.; Galizia, D.; Casorzo, L.; Sarotto, I.; Scaltriti, M.; Sapino, A.; et al. Genotyping tumour DNA in cerebrospinal fluid and plasma of a HER2-positive breast cancer patient with brain metastases. ESMO Open 2017, 2. [Google Scholar] [CrossRef]
- Ma, C.; Yang, X.; Xing, W.; Yu, H.; Si, T.; Guo, Z. Detection of circulating tumor DNA from non--small cell lung cancer brain metastasis in cerebrospinal fluid samples. Thorac Cancer 2020, 11, 588–593. [Google Scholar] [CrossRef]
- Yan, W.; Xu, T.; Zhu, H.; Yu, J. Clinical Applications of Cerebrospinal Fluid Circulating Tumor DNA as a Liquid Biopsy for Central Nervous System Tumors. Onco Targets Ther. 2020, 13, 719–731. [Google Scholar] [CrossRef] [PubMed]
- Von Baumgarten, L.; Kumbrink, J.; Jung, A.; Reischer, A.; Flach, M.; Liebmann, S.; Metzeler, K.H.; Holch, J.W.; Niyazi, M.; Thon, N.; et al. Therapeutic management of neuro-oncologic patients—Potential relevance of CSF liquid biopsy. Theranostics 2020, 10, 856–866. [Google Scholar] [CrossRef] [PubMed]
- Jung, M.; Pützer, S.; Gevensleben, H.; Meller, S.; Kristiansen, G.; Dietrich, D. Diagnostic and prognostic value of SHOX2 and SEPT9 DNA methylation and cytology in benign, paramalignant, and malignant ascites. Clin. Epigenet. 2016, 8. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.S.; Hur, J.Y.; Kim, I.A.; Kim, H.J.; Choi, C.M.; Lee, J.C.; Kim, W.S.; Lee, K.Y. Liquid biopsy using the supernatant of a pleural effusion for EGFR genotyping in pulmonary adenocarcinoma patients: A comparison between cell-free DNA and extracellular vesicle-derived DNA. BMC Cancer 2018, 18, 1236. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Wu, X.; Tang, M.; Nie, X.; Li, L. Detection of EGFR gene mutation status from pleural effusions and other body fluid specimens in patients with lung adenocarcinoma. Thorac Cancer 2019, 10, 2218–2224. [Google Scholar] [CrossRef] [PubMed]
- Villalba, M.; Exposito, F.; Pajares, M.J.; Sainz, C.; Redrado, M.; Remirez, A.; Wistuba, I.; Behrens, C.; Jantus-Lewintre, E.; Camps, C.; et al. TMPRSS4: A Novel Tumor Prognostic Indicator for the Stratification of Stage IA Tumors and a Liquid Biopsy Biomarker for NSCLC Patients. J. Clin. Med. 2019, 8, 2134. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Liu, Y.; Zhang, W.; Lin, L.; Zhang, J.; Xiong, Y.; Nie, L.; Liu, X.; Li, H.; Wang, W. A rapid liquid biopsy of lung cancer by separation and detection of exfoliated tumor cells from bronchoalveolar lavage fluid with a dual-layer “PERFECT” filter system. Theranostics 2020, 10, 6517–6529. [Google Scholar] [CrossRef] [PubMed]
- Roman-Canal, B.; Moiola, C.P.; Gatius, S.; Bonnin, S.; Ruiz-Miró, M.; González, E.; González-Tallada, X.; Llordella, I.; Hernández, I.; Porcel, J.M.; et al. EV-Associated miRNAs from Peritoneal Lavage are a Source of Biomarkers in Endometrial Cancer. Cancers 2019, 11, 839. [Google Scholar] [CrossRef]
- Mayo-de-Las-Casas, C.; Velasco, A.; Sanchez, D.; Martínez-Bueno, A.; Garzón-Ibáñez, M.; Gatius, S.; Ruiz-Miró, M.; Gonzalez-Tallada, X.; Llordella, I.; Tresserra, F.; et al. Detection of somatic mutations in peritoneal lavages and plasma of endometrial cancer patients: A proof-of-concept study. Int. J. Cancer 2020, 147, 277–284. [Google Scholar] [CrossRef]
- Martignetti, J.A.; Pandya, D.; Nagarsheth, N.; Chen, Y.; Camacho, O.; Tomita, S.; Brodman, M.; Ascher-Walsh, C.; Kolev, V.; Cohen, S.; et al. Detection of endometrial precancer by a targeted gynecologic cancer liquid biopsy. Cold Spring Harb. Mol. Case Stud. 2018, 4. [Google Scholar] [CrossRef]
- He, S.-M.; Xing, F.; Sui, H.; Wu, Y.; Wang, Y.; Wang, D.; Chen, G.; Kong, Z.; Zhou, S.-F. Determination of CA-125 levels in the serum, cervical and vaginal secretions, and endometrium in Chinese women with precancerous disease or endometrial cancer. Med. Sci. Monit. 2011, 17, CR618–CR625. [Google Scholar] [CrossRef] [PubMed]
- Böhm, D.; Keller, K.; Pieter, J.; Boehm, N.; Wolters, D.; Siggelkow, W.; Lebrecht, A.; Schmidt, M.; Kölbl, H.; Pfeiffer, N.; et al. Comparison of tear protein levels in breast cancer patients and healthy controls using a de novo proteomic approach. Oncol. Rep. 2012, 28, 429–438. [Google Scholar] [CrossRef] [PubMed]
- Inubushi, S.; Kawaguchi, H.; Mizumoto, S.; Kunihisa, T.; Baba, M.; Kitayama, Y.; Takeuchi, T.; Hoffman, R.M.; Sasaki, R. Oncogenic miRNAs Identified in Tear Exosomes From Metastatic Breast Cancer Patients. Anticancer Res. 2020, 40, 3091–3096. [Google Scholar] [CrossRef] [PubMed]
- Moelans, C.B.; Patuleia, S.I.S.; van Gils, C.H.; van der Wall, E.; van Diest, P.J. Application of Nipple Aspirate Fluid miRNA Profiles for Early Breast Cancer Detection and Management. Int. J. Mol. Sci. 2019, 20, 5814. [Google Scholar] [CrossRef] [PubMed]
- Tanos, R.; Thierry, A.R. Clinical relevance of liquid biopsy for cancer screening. Transl. Cancer Res. 2018, 7, S105–S129. [Google Scholar] [CrossRef]
- Qian, S.; Golubnitschaja, O.; Zhan, X. Chronic inflammation: Key player and biomarker-set to predict and prevent cancer development and progression based on individualized patient profiles. EPMA J. 2019, 10, 365–381. [Google Scholar] [CrossRef]
- Stastny, I.; Dankova, Z.; Kajo, K.; Kubatka, P.; Golubnitschaja, O.; Zubor, P. Aberrantly methylated cfDNA in body fluids as a promising diagnostic tool for early detection of breast cancer. Clin. Breast Cancer 2020. [Google Scholar] [CrossRef]
- Lu, M.; Zhan, X. The crucial role of multiomic approach in cancer research and clinically relevant outcomes. EPMA J. 2018, 9, 77–102. [Google Scholar] [CrossRef]
- Li, N.; Zhan, X. Signaling pathway network alterations in human ovarian cancers identified with quantitative mitochondrial proteomics. EPMA J. 2019, 10, 153–172. [Google Scholar] [CrossRef]
- Golubnitschaja, O.; Flammer, J. Individualised patient profile: Clinical utility of Flammer syndrome phenotype and general lessons for predictive, preventive and personalised medicine. EPMA J. 2018, 9, 15–20. [Google Scholar] [CrossRef]
- Janssens, J.P.; Schuster, K.; Voss, A. Preventive, predictive, and personalized medicine for effective and affordable cancer care. EPMA J. 2018, 9, 113–123. [Google Scholar] [CrossRef] [PubMed]
- Polivka, J.; Altun, I.; Golubnitschaja, O. Pregnancy-associated breast cancer: The risky status quo and new concepts of predictive medicine. EPMA J. 2018, 9, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Koklesova, L.; Liskova, A.; Samec, M.; Qaradakhi, T.; Zulli, A.; Smejkal, K.; Kajo, K.; Jakubikova, J.; Behzadi, P.; Pec, M.; et al. Genoprotective activities of plant natural substances in cancer and chemopreventive strategies in the context of 3P medicine. EPMA J. 2020, 11, 261–287. [Google Scholar] [CrossRef] [PubMed]
- Uramova, S.; Kubatka, P.; Dankova, Z.; Kapinova, A.; Zolakova, B.; Samec, M.; Zubor, P.; Zulli, A.; Valentova, V.; Kwon, T.K.; et al. Plant natural modulators in breast cancer prevention: Status quo and future perspectives reinforced by predictive, preventive, and personalized medical approach. EPMA J. 2018, 9, 403–419. [Google Scholar] [CrossRef] [PubMed]
| Technology | Cancer Type | Liquid Biopsy Type | Biomarker | Reference |
|---|---|---|---|---|
| RT-PCR | CRC | Plasma | ↑ Methylation of SEPT9 | [48] |
| CRC | Plasma | ↓ miR-145, -195 ↑ miR-29, 92 | [49] | |
| Gastric cancer | Plasma | ↓ miR-195-5p | [50] | |
| Glioblastoma | CSF | ↑ miR-21 | [51] | |
| CNS malignancies | CSF | Differently expressed miR-771, -451, -223, -935, -125b | [52] | |
| UCC | Urine | gene expression signatures for UCC | [53] | |
| Bladder cancer | Urine | ↓ miR-145, -200 | [54] | |
| Bladder cancer | Urine | ↑ UBE2C | [55] | |
| OSCC | Saliva | ↓ miR-125, -200a | [56] | |
| EC | Saliva | ↑ miR-10b, -21, -144 and -451 | [57] | |
| EC | Saliva | ↑ miR-21 | [58] | |
| Lung adenocarcinoma | PE | ↑ miR-195-5p, -182-5p, and -34a-5p | ||
| dPCR | HNSCC | Plasma | Detection of TP53 mutation | [61] |
| Melanoma | Plasma | Detection of BRAFV600E mutation | [62] | |
| Metastatic adenocarcinoma | Plasma | Detection of KRAS mutation | [63] | |
| Central nervous system lymphomas | CSF | Detection of MYD88 p.(L265P) | [64] | |
| Glioma | CSF | Analysis of mutant IDH1 mRNA | [67] | |
| Bladder cancer | Urine | Identification of TERT mutation | [65] | |
| NSCLC | Urine | Identification of EGFR mutation | [66] | |
| NSCLC | PE | Identification of EGFR mutation | [70] | |
| NGS | NSCLC | Plasma | Detection of oncogenic drivers and resistance mechanisms | [74] |
| EC | Plasma | Detection of somatic mutations associated with recurrence of disease | [83] | |
| LAC with LM | CSF | Detection of tumor associated mutations | [75] | |
| NSCLC with LM | CSF | Detection of targetable changes | [76] | |
| LC | PE | Identification of alterations in tumor genomics | [77] | |
| MS | Multiple myeloma | Urine | Identification of urine biomarker | [80] |
| Liver cancer | Serum | Analysis of total serum protein fingerprints | [81] | |
| ESCC | Plasma | Identification of plasma biomarkers | [82] |
| Biomarker | Cancer Type | Study Characteristic (Number of Patients) | Study Results | Reference |
|---|---|---|---|---|
| Circulating tumor DNA | ||||
| ctDNA (serum) | NSCLC | NSCLC patients (n = 60), COPD patients (n = 40) and healthy controls (n = 40) | Discrimination of normal individuals from COPD and NSCLC. ctDNA high level → short survival of NSCLC patients. | [86] |
| ctDNA (serum) | Advanced NSCLC | Patients rechallenged with gefitinib at progression after second-line chemotherapy (n = 61) | An identification of EGFR-mutant patients: those not carrying p.T790M variant with no other alternative treatment might benefit from TKI rechallenge. | [87] |
| Formalin-fixed, paraffin-embedded, metastatic tissue and corresponding ctDNA (serum) | Advanced breast cancer | Metastatic breast cancer patients (n = 66) | PIK3CA mutation tumor tissue and detectable PIK3CA mutations in serum ctDNA in 83% of cases. Correlation between changes in PIK3CA ctDNA level and treatment response. | [106] |
| HPV-specific E7 and L1 genes (ctDNA, plasma) | Cervical cancer | Cervical cancer patients (n = 138) | HPV E7 and L1 sequences detected in plasma ctDNA (61.6% of patients). High viral load: increased risk of recurrence and death at 5 years (univariate analysis). | [107] |
| Combined exosomal RNA and ctDNA (plasma) | NSCLC | Mutant EGFR NSCLC patients (n = 84) | Increased sensitivity for EGFR mutation detection, especially in patients with intrathoracic metastatic disease (low levels of ctDNA). | [108] |
| Methylation level in circulating cell-free DNA (serum, plasma) | Breast cancer | Serum test cohort (n = 103), serum validation cohort (n = 368), and plasma cohort (n = 125) | Serum test cohort: panel of SPAG6 and ITIH5 → 63% sensitivity of DCIS and 51% sensitivity for early invasive tumor detection at 80% specificity. The serum validation cohort: 50% sensitivity for DCIS detection (NKX2-6 and ITIH5). | [112] |
| Methylation level in circulating cell-free DNA (plasma) | Lung cancer | Tissue samples (n = 152), plasma samples (n = 129), and benign lesions of lung (n = 28) | Plasma samples: higher methylation of HOXA9 and RASSF1A in SCLC than in NSCLC. | [89] |
| Cell-free DNA sequencing informative methylation patterns (plasma) | More than 50 cancer types | Participants with (n = 8584) and without (n = 6670) cancer | Detection of more than 50 cancer types across stages. | [113] |
| Circulating miRNAs in liquid biopsy | ||||
| miRNA (serum) | Lung cancer | Preliminary marker selection (early-stage lung cancer n = 24 and healthy control n =24) and a validation phase (early-stage lung cancer n = 94, stage II to IV n = 48, and healthy control n = 111) | Potential of combination of miR-125a-5p, miR-25, and miR-126 in early detection of lung cancer. | [118] |
| miRNA (serum) | Prostate cancer | Prostate cancer patients (n = 809), negative prostate biopsies (n = 241), patients with other cancer types (n = 500), and healthy controls (n = 41) | Potential of combination of miR-17-3p and miR-1185-2-3p as a marker of prostate cancer diagnosis. | [119] |
| miRNA (serum) | Bladder cancer | Bladder cancer patients (n = 392), non-cancer samples (n = 100), and other cancer types (n = 480) | Set of 7 miRNAs (miR-6087, miR-1185-1-3p, miR-3960, miR-6724-5p, miR-1343-5p, miR-6831-5p and miR-4695-5p): discrimination of bladder cancer from non-cancer and other types of cancer. | [120] |
| miRNA (serum) | Breast, endometrial and ovarian cancer | Breast cancer (n = 31), endometrial cancer (n = 13), and ovarian cancer (n = 15) patients | miR-518b, -4719 and -6757-3p deregulated in breast cancer. miR-484/-23a diagnostic biomarker for endometrial and ovarian cancer. | [121] |
| Extracellular vesicles in liquid biopsy | ||||
| Circulating exosomes (serum) | ESCC | ESCC patients (n = 200) | Upregulated level of circulating exosomes in ESCC patients. | [91] |
| Circulating exosomal DNA (serum) | PDAC | PDAC patients (n = 48) and healthy subjects (n = 114) | Highlighting the role of circulating exosomal DNA in rapid identification of cancer driving mutations. | [92] |
| EV-miRNAs (serum) | mCRC | mCRC (n = 44) and healthy controls (n = 17) | Baseline miRNA-21 and -92a outperformed carcinoembryonic antigen levels in mCRC patients when compared to healthy controls. Higher levels of miRNA-92a and 222 in patients who died. | [93] |
| Circulating tumor cells | ||||
| CTCs | Prostate cancer patients | Prostate cancer patients with CTCs <5 (n = 511) | Increasing CTCs associated with worse overall survival of patients treated with chemotherapy of endocrine therapy. | [137] |
| Metastatic breast cancer patients | Metastatic breast cancer patients receiving eribulin treatment (n = 21) | Determination of mesenchymal and epithelial CTCs for the prediction of survival. | ||
| SCLC | SCLC patients before pazopanib initiation (n = 56 patients), after one-cycle (n = 35), and on disease progression (n = 45) | Analysis of CTCs as biomarkers of treatment efficacy (pazopanib eliminates CTC subpopulations). | [139] | |
| EOC | EOC patients (n = 109) | Detection of CTCs and their pattern of gene expression could predict the likelihood of chemotherapy resistance and evaluate the prognosis of ovarian cancer patients. | [140] | |
| NSCLC | Advanced-stage NSCLC patients (n = 45) | Identification of CTCs through EGFR/HER3 expression →novel liquid biopsy approach. | [141] | |
| EOC | EOC patients (n = 10) | The feasibility and potential usefulness of chemosensitivity assay using liquid biopsy-derived CTCs in the prediction of response to therapy. | [142] | |
| Ewing sarcoma | Ewing sarcoma patients (n = 18) and healthy volunteers (n = 9) | Identification of CTCs by immunoseparation with CD99 antibody and magnetic microbeads → prognostic and predictive potential. | [143] | |
| CRC (draining venous blood) | CRC patients (n = 26) and healthy volunteers (n = 14) | New filtration and cytology-based automated platform for detection of CTCs → prognostic and predictive potential. | [144] | |
| Other biomarkers of blood-based liquid biopsy or their combinations | ||||
| Soluble PD-L1 and PD-L2 (serum) | EOC | EOC patients (n = 83) and healthy controls (n = 29) | Soluble PD-L1 increased and PD-L2 decreased in EOC. Enhanced soluble PD-L1: residual tumor burden and reduced 5 year overall survival and progression-free survival. Reduced soluble PD-L2: platinum-resistance. | [145] |
| sCRT | Ovarian cancer | Ovarian cancer patients (n = 134) and healthy controls (n = 116) | Increased sCRT in ovarian cancer patients. sCRT predictor of poor prognosis and platinum resistance. | [94] |
| sGKN1 | Gastric cancer | Advanced gastric cancer patients (n = 360), early gastric cancer patients (n = 140), and healthy controls (n = 200) | Increased sGKN1 in healthy subjects when compared with gastric cancer patients. Decreased sGKN1 in advanced gastric cancer when compared with early gastric cancer. | [95] |
| Serum proteins and miRNAs | Cervical cancer | Early-stage cervical cancer patients (n = 140) and healthy controls (n = 140). Independent cohort study (early-stage cervical cancer patients n = 60 and healthy controls n = 60) | Combination of SCC Ag degree and miRNA-29a, miRNA-25, and miRNA-486-5p levels as a marker of early-stage cervical cancer detection. | [98] |
| miRNA and fecal hemoglobin concentration (serum) | Colorectal carcinoma | CRC patients (n = 59), advanced adenomas (n = 74) and control subjects (n = 80) | Potential of a combination of 6 miRNAs (miR-15b-5p, miR-29a-3p, miR-335-5p, miR-18a-5p, miR-19a-3p and miR-19b-3p) and fecal hemoglobin concentration in the detection of advanced colorectal cancer in average risk individuals. | [146] |
| Histone modifications (serum) | Hepatocellular carcinoma | Hepatocellular carcinoma patients’ blood samples (n = 24) and healthy volunteers (n = 6) | Serum purified histones: comparable pattern of modifications like acetylation (H4K16Ac), methylation (H4K20Me3, H3K27Me3, H3K9Me3) and phosphorylation (γ-H2AX and H3S10P) to paired cancer tissues. | [96] |
| Histone modifications (plasma) | CRC | CRC patients (n = 63) and control subjects (n = 40) | Lower H3K27me3 and H4K20me3 in CRC patients when compared to healthy control. | [97] |
| Biomarker | Cancer Type | Study Characteristics | Study Results | Reference |
|---|---|---|---|---|
| Tumor DNA (urine supernatant) | NMIBC | NMIBC patients (n = 216) and patients with bladder cancer undergoing radical cystectomy (n = 27) | An association between high levels of tumor DNA and later disease progression in NMIBC. | [158] |
| 6-gene (APC2, CDH1, FOXP1, LRRC3B, WNT7A and ZIC4) promoter methylation (urine cell-free DNA) | Prostate cancer | Prostate cancer patients (n = 31) and control subjects (n = 33) | NGM increased monotonically from 0.27 in control subjects to 4.6 and 4.25 in patients with highly developed and T2/T3 stage metastatic prostate cancer, respectively. | [159] |
| ctDNA (plasma, urine) | mCRC | mCRC patients (n = 150) | Utilization of both plasma and urine cell-free DNA to address disease progression in CRC patients. | [160] |
| EGFR and TP53 mutations (plasma, urine, sputum) | NSCLC | NSCLC patients (n = 50) | Increase in the detection of EGFR or TP53 mutation with higher sensitivity by a combination of plasma, sputum and urine. | [161] |
| Lipids in urinary exosomes | Prostate cancer | Prostate cancer patients (n = 15) and healthy controls (n = 13) | Different levels of lipid species in the two groups. | [24] |
| miRNA | ccRCC | ccRCC patients (n = 75) and control subjects (n = 45) | Higher urinary cell-free miRNA-210 in ccRCC vs. control. Decreased urinary cell-free miRNA-210 in ccRCC patients a week after surgery. | [163] |
| miRNA (urine-derived exosomes) | Endometrial cancer | Endometrial cancer patients (n = 22) and symptomatic controls (n = 5) | The potential utilization of differential miRNA in exosomes as biomarker in diagnosis of endometrial cancer (hsa-miR-200c-3p as a candidate). | [165] |
| miRNAs | Bladder cancer | Identification of miRNA fingerprints: bladder cancer patients (n = 66) and control subjects (n = 48). Altered miRNAs validation: bladder cancer patients (n = 112) and control subjects (n = 65) | AUC (miR-30a-5p, let-7c-5p, miR-486-5p) altered in all bladder cancer subtypes → increased accuracy in the discrimination of cases and controls. | [164] |
| Biomarker | Cancer Type | Study Characteristics | Study Results | Reference |
|---|---|---|---|---|
| mRNA (saliva) and blood CTCs | NSCLC | Discovery phase: NSCLC patients (n = 140) and healthy controls (n = 140). Validation phase: NSCLC patients (n = 60) and healthy controls (n = 60). | Panel of CTC level in blood and mRNA markers in saliva (CCNI, EGFR, FGF19, FRS2, GREB1): discrimination of NSCLC from healthy controls. | [15] |
| mRNA (saliva) and CEA (blood) | NSCLC | Discovery phase: NSCLC patients (n = 30) and healthy controls (n = 30). Prediction performance evaluation: NSCLC patients (n = 15) and healthy controls (n = 25). | Panel measuring CEA in blood and GREB1 and FRS2 levels in saliva could be used for the detection of NSCLC. | [170] |
| miRNAs (saliva) | CRC | Discovery phase (healthy controls n = 10 and CRC patients n = 14) and validation phase (healthy controls n = 37, CRC patients n = 51, and adenoma n = 19) | Panel of saliva-based miRNAs (miR-186-5p, miR-29a-3p, miR-29c-3p, miR-766-3p, and miR-491-5p) higher in CRC vs. control → detection of CRC. | [11] |
| miRNAs (salivary exosomes) | Pancreatobiliary tract cancer | Pancreatobiliary tract cancer (n = 12) and healthy controls (n = 13) | Relative expression ratios of miR-1246 and miR-4644 significantly higher in cancer group vs. control. The potential of miR-1246 and miR-4644 in salivary exosomes as candidate biomarkers. | [173] |
| Proteins (salivary exosomes) | Lung cancer | Lung cancer patients and normal subjects | The potential use of informative proteins in salivary EVs for detection of lung cancer. | [174] |
| Salivary exosomes | Isolation of salivary exosomes by the acoustofluidic (the fusion of acoustics and microfluidics) platform → potential in the detection of HPV-OPC. | [176] | ||
| ctDNA containing EGFR mutations (saliva, plasma) | Electric field-induced release and measurement → novel platform detecting ctDNA containing EGFR mutations directly from plasma and saliva in early- and late-stage NSCLC. | [177] | ||
| Biomarker | Cancer Type | Study Characteristics | Study Results | Reference |
|---|---|---|---|---|
| CSF metabolites | Primary or metastatic central nervous system tumors | Patients without a history of cancer (n = 8) and with a variety of CNS tumor types (n = 23) (i.e., glioma IDH mutant, glioma IDH wildtype, metastatic lung cancer and metastatic breast cancer) | Differences in the abundance of 43 metabolites between CSF from control patients and the CSF of patients with primary or metastatic CNS tumors. Alterations in metabolic pathways (e.g., glycine, choline and methionine degradation, diphthamide biosynthesis and glycolysis pathways, among others) between IDH-mutant and IDH-wildtype gliomas. IDH-mutant gliomas: higher levels of D-2-hydroxyglutarate in CSF in comparison to patients with other tumor types or controls. | [25] |
| ctDNA (CSF, plasma) | HER2-positive breast cancer with brain metastases | CSF-derived ctDNA analysis: TP53, PIK3CA mutations and ERBB2 and cMYC amplification. Post-treatment ctDNA analysis: decreased marker levels in plasma (consistent with extra-CNS disease control) and increased CSF (poor treatment benefit in the CNS). | [181] | |
| ctDNA (blood, CSF) | NSCLC with brain metastasis | NSCLC patients with brain metastasis (n = 21) | Specific genetic mutation patterns in driver genes: EGFR mutations: 57.1% (in CSF ctDNA) and 23.8% (in peripheral blood ctDNA and in CTCs). EGFR mutations found in CSF of 81.8% patients with leptomeningeal metastases and 30% patients with brain parenchymal metastases. The status of EGFR and TP53 mutations was consistent between CSF ctDNA and brain lesion tissue in all five patients. | [182] |
| Biomarker | Cancer Type | Study Characteristics | Study Results | Reference |
|---|---|---|---|---|
| Ascites-based liquid biopsy | ||||
| Malignant ascites | Gastrointestinal cancer | Patients diagnosed with malignant ascites of gastrointestinal cancer (n = 27) | Large amount of CD4+ and CD8+ T cells: exhausted phenotype → significant clinical relevance as prognostic and therapeutic target. | [30] |
| Pleural effusion | ||||
| EGFR (pleural effusion, ascites, pericardial effusion and cerebrospinal fluid) | Lung adenocarcinoma patients | Lung adenocarcinoma patients (n = 20) | Higher detection rate sensitivity of tumor-specific EGFR mutations in biofluid-supernatant-free DNA in comparison with biofluid sediment tumor cells and plasma-free DNA samples. | [187] |
| Bronchoalveolar lavage | ||||
| EGFR (bronchoalveolar lavage fluid, EVs) | NSCLC | NSCLC patients (n = 137) | EGFR genotyping by bronchoalveolar lavage fluid obtained from tumor site: high accuracy of diagnosis. | [153] |
| TMPRSS4 methylation (bronchoalveolar lavage and plasma) | NSCLC | Bronchoalveolar lavage: lung cancer patients (n = 79) and healthy controls (n = 26). Plasma: lung cancer patients (n = 89) and healthy controls (n = 25). | Monitoring of surgically resected NSCLC patients. TMPRSS4 methylation status differentiates NSCLC and tumor-free subjects. | [188] |
| Peritoneal lavage | ||||
| EV-isolated miRNAs (peritoneal lavage, ascites) | CRC | CRC patients (n = 25) and control patients (n = 25) | Source of potential biomarkers for CRC diagnosis (miRNA-199b-5p, miRNA-150-5p, miRNA-29c-5p, miRNA-218-5p, miRNA-99a-3p, miRNA-383-5p, miRNA-199a-3p, miRNA-193a-5p, miRNA-10b-5p and miRNA-181c-5p). | [38] |
| EV-isolated miRNAs (peritoneal lavage, ascites) | Endometrial cancer | Endometrial cancer patients (n = 25) and control patients (n = 25) | Deregulated miRNAs in endometrial cancer (n = 114) miRNAs. miRNA-383-5p, miRNA-10b-5p, miRNA-34c-3p, miRNA-449b-5p, miRNA-34c-5p, miRNA-200b-3p, miRNA-2110, and miRNA-34b-3p highlighted as promising biomarkers. | [190] |
| KRAS and PIK3CA mutational analysis (peritoneal lavage, blood) | Endometrial cancer | Endometrial cancer patients (n = 50) | Approved feasibility of mutational analysis. Further studies needed to determine its use in the identification of patients with worse prognosis. | [191] |
| Uterine/utero-tubular lavage | ||||
| PTEN mutations (uterine lavage fluid) | Endometrial cancer | 67-year-old asymptomatic female | Identification of two oncogenic PTEN mutations nearly one year before the occurrence of symptoms | [192] |
| Genomic profiling (uterine aspirate, blood) | Endometrial cancer | Endometrial cancer patients (n = 60) | Potential applicability of combined use of different liquid biopsy for personalized cancer management. | [10] |
| Microvesicle proteomic profiling (utero-tubal lavage) | Ovarian cancer | High-grade ovarian cancer patients (n = 49) and controls (n = 127) | 9-protein classifier with 70% sensitivity and 76.2% specificity (identified all stage I lesions). | [35] |
| Cervical and vaginal secretions | ||||
| CA-125 (cervical and vaginal secretions, serum) | Endometrial cancer | Patients with polyps, hyperplasia or endometrial cancer (n = 97) and healthy subjects (n = 51) | Increased CA-125 in patients with complex hyperplasia and endometrial cancer. | [193] |
| Tear fluid | ||||
| Proteomic pattern | Breast cancer | Primary invasive breast carcinoma patients (n = 25) and healthy controls (n = 25) | Identified diagnostic protein biomarker to differentiate cancer patients from controls. | [150,194] |
| Breast cancer patients (n = 50) and healthy controls (n = 50) | ||||
| Tear exosomes (miRNAs) | Breast cancer | Metastatic breast cancer patients (n = 5) and healthy controls (n = 8) | Higher quantity of exosome markers in tears than in serum. Highly expressed miRNA-21 and miRNA-200c in tear exosomes of metastatic breast cancer patients in comparison with controls. | [195] |
| Breast milk | ||||
| Proteins | Breast cancer | Ten milk samples from eight females | Identification of protein for early detection and accurate assessment of breast cancer. | [151] |
| Breast ductal fluid | ||||
| miRNAs | Unilateral breast cancer | Unilateral breast cancer patients (n = 22) | Feasibility of analyzing miRNAs. Discrimination of tumor histological subtypes. Discrimination of cancer and normal breast samples. | [154] |
| Seminal fluid | ||||
| ctDNA (seminal plasma) | Prostate cancer | Prostate cancer patients (n = 6) and healthy controls (n = 3) | Different concentrations and fragment size of seminal plasma ctDNA in prostate cancer patients, benign prostate hyperplasia and healthy controls. | [33,152] |
| Prostate cancer patients (n = 30), benign prostate hyperplasia patients (n = 33), and healthy controls (n = 21) | ||||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liskova, A.; Samec, M.; Koklesova, L.; Giordano, F.A.; Kubatka, P.; Golubnitschaja, O. Liquid Biopsy is Instrumental for 3PM Dimensional Solutions in Cancer Management. J. Clin. Med. 2020, 9, 2749. https://doi.org/10.3390/jcm9092749
Liskova A, Samec M, Koklesova L, Giordano FA, Kubatka P, Golubnitschaja O. Liquid Biopsy is Instrumental for 3PM Dimensional Solutions in Cancer Management. Journal of Clinical Medicine. 2020; 9(9):2749. https://doi.org/10.3390/jcm9092749
Chicago/Turabian StyleLiskova, Alena, Marek Samec, Lenka Koklesova, Frank A. Giordano, Peter Kubatka, and Olga Golubnitschaja. 2020. "Liquid Biopsy is Instrumental for 3PM Dimensional Solutions in Cancer Management" Journal of Clinical Medicine 9, no. 9: 2749. https://doi.org/10.3390/jcm9092749
APA StyleLiskova, A., Samec, M., Koklesova, L., Giordano, F. A., Kubatka, P., & Golubnitschaja, O. (2020). Liquid Biopsy is Instrumental for 3PM Dimensional Solutions in Cancer Management. Journal of Clinical Medicine, 9(9), 2749. https://doi.org/10.3390/jcm9092749






