Vaccines and Therapies in Development for SARS-CoV-2 Infections
Abstract
1. Introduction to SARS-CoV-2
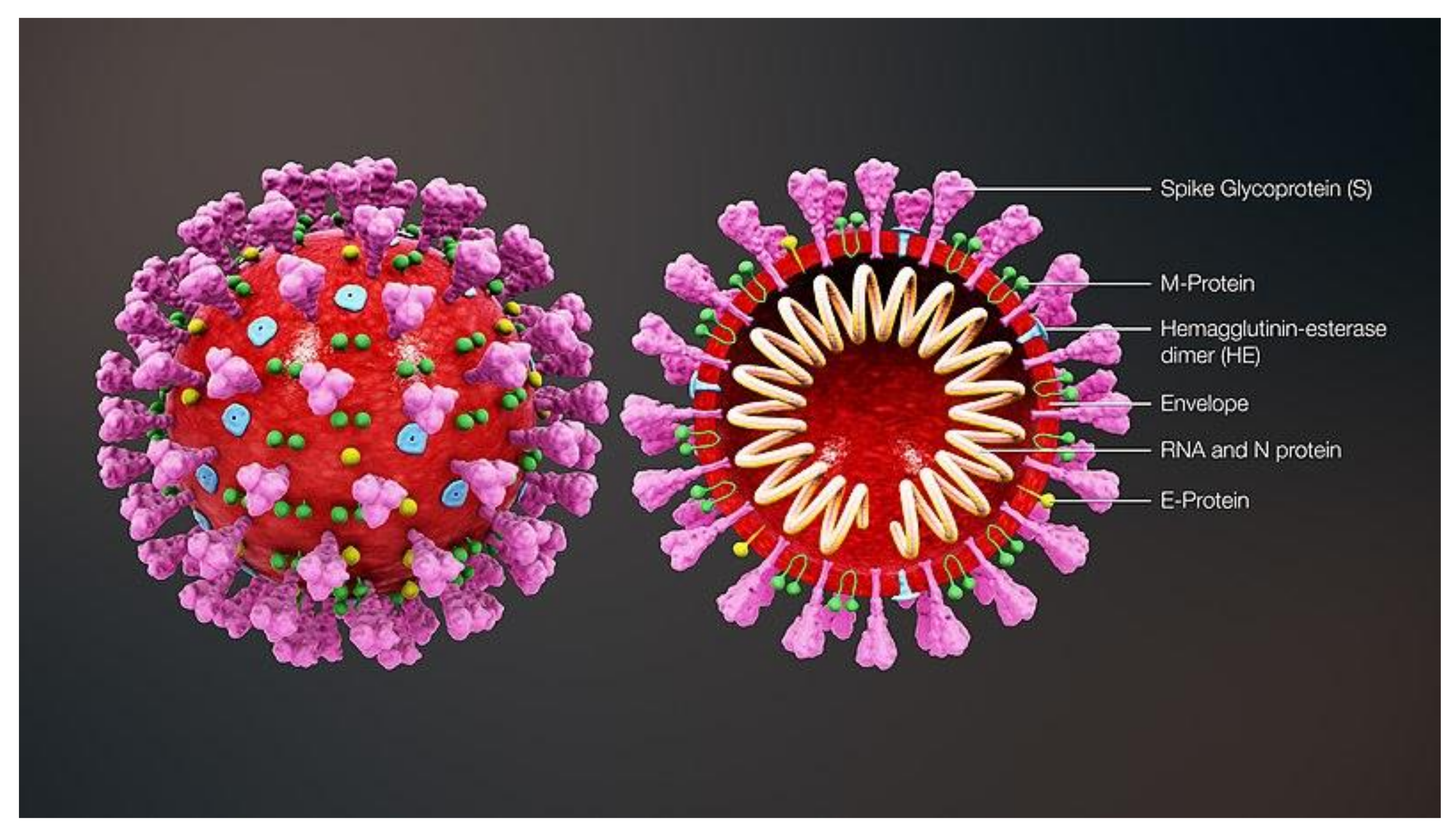
2. Structure of SARS-CoV-2 Protein Targets and Comparison to SARS-Classic
3. Viral Factors for Drug Development
3.1. Therapeutics Targeting Spike Protein
3.2. Therapeutics Targeting NSPs
3.3. RNA Dependent RNA Polymerase
3.4. Molecules Targeting Helicase
3.5. Host Factors for Drug Development
4. Therapies in Development
4.1. Remdesivir, Galidesivir, and Favipiravir
4.2. Experimental Therapy: Plasma Therapy
4.3. Experimental Therapy: Hydroxychloroquine/Chloroquine
4.4. Indomethacin
4.5. Ivermectin
5. Vaccines in Development: mRNA-1273 and ChAdOx1 nCoV-19
6. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Tu, Y.; Chien, C.; Yarmishyn, A.A.; Lin, Y.; Luo, Y.; Lin, Y.; Lai, W.; Yang, D.; Chou, S.; Yang, Y.; et al. A Review of SARS-CoV-2 and the Ongoing Clinical Trials. Int. J. Mol. Sci. 2020, 21. [Google Scholar] [CrossRef]
- Coronavirus Update (Live): 2,536,673 Cases and 175,759 Deaths from COVID-19 Virus Pandemic—Worldometer. Available online: https://www.worldometers.info/coronavirus/ (accessed on 21 April 2020).
- SARS Frequently Asked Questions CDC. Available online: https://www.cdc.gov/sars/about/faq.html (accessed on 21 April 2020).
- 2009 H1N1 Pandemic. Available online: https://www.cdc.gov/flu/pandemic-resources/2009-h1n1-pandemic.html (accessed on 21 April 2020).
- Valencia, D.N. Brief Review on COVID-19: The 2020 Pandemic Caused by SARS-CoV-2. Cureus Journal of Medical Science 2020, 12. [Google Scholar] [CrossRef]
- Coronavirus Human Coronavirus Types CDC. Available online: https://www.cdc.gov/coronavirus/types.html (accessed on 21 April 2020).
- Meselson, M. Droplets and Aerosols in the Transmission of SARS-CoV-2. N. Engl. J. Med. 2020, 382, 2063. [Google Scholar] [CrossRef] [PubMed]
- Van Doremalen, N.; Bushmaker, T.; Morris, D.H.; Holbrook, M.G.; Gamble, A.; Williamson, B.N.; Tamin, A.; Harcourt, J.L.; Thornburg, N.J.; Gerber, S.I.; et al. Aerosol and Surface Stability of SARS-CoV-2 as Compared with SARS-CoV-1. N. Engl. J. Med. 2020, 382, 1564–1567. [Google Scholar] [CrossRef] [PubMed]
- Guo, Z.; Wang, Z.; Zhang, S.; Li, X.; Li, L.; Li, C.; Cui, Y.; Fu, R.; Dong, Y.; Chi, X.; et al. Early Release—Aerosol and Surface Distribution of Severe Acute Respiratory Syndrome Coronavirus 2 in Hospital Wards, Wuhan, China. Emerg. Infect. Dis. 2020, 26. [Google Scholar] [CrossRef]
- Zhang, S.; Diao, M.; Yu, W.; Pei, L.; Lin, Z.; Chen, D. Estimation of the reproductive number of novel coronavirus (COVID-19) and the probable outbreak size on the Diamond Princess cruise ship: A data-driven analysis. Int. J. Infect. Dis. 2020, 93, 201–204. [Google Scholar] [CrossRef]
- Sanche, S.; Lin, Y.T.; Xu, C.; Romero-Severson, E.; Hengartner, N.; Ke, R. Early Release—High Contagiousness and Rapid Spread of Severe Acute Respiratory Syndrome Coronavirus 2. Emerg. Infect. Dis. 2020, 26. [Google Scholar] [CrossRef]
- Coronavirus Disease 2019 (COVID-19)? In Symptoms. Available online: https://www.cdc.gov/coronavirus/2019-ncov/symptoms-testing/symptoms.html (accessed on 21 April 2020).
- Wang, D.; Hu, B.; Hu, C.; Zhu, F.; Liu, X.; Zhang, J.; Wang, B.; Xiang, H.; Cheng, Z.; Xiong, Y.; et al. Clinical Characteristics of 138 Hospitalized Patients With 2019 Novel Coronavirus–Infected Pneumonia in Wuhan, China. JAMA 2020, 323, 1061–1069. [Google Scholar] [CrossRef]
- Wu, P.; Duan, F.; Luo, C.; Liu, Q.; Qu, X.; Liang, L.; Wu, K. Characteristics of Ocular Findings of Patients With Coronavirus Disease 2019 (COVID-19) in Hubei Province, China. JAMA Ophthalmol. 2020, 138, 575–578. [Google Scholar] [CrossRef]
- Wu, C.; Liu, Y.; Yang, Y.; Zhang, P.; Zhong, W.; Wang, Y.; Wang, Q.; Xu, Y.; Li, M.; Li, X.; et al. Analysis of therapeutic targets for SARS-CoV-2 and discovery of potential drugs by computational methods. Acta Pharm. Sin. B 2020, 10, 766–788. [Google Scholar] [CrossRef]
- Xu, H.; Zhong, L.; Deng, J.; Peng, J.; Dan, H.; Zeng, X.; Li, T.; Chen, G. High expression of ACE2 receptor of 2019-nCoV on the epithelial cells of oral mucosa. Int. J. Oral Sci. 2020, 12, 8. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Krüger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 52, 731–733. [Google Scholar] [CrossRef] [PubMed]
- Zhou, P.; Yang, X.; Wang, X.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.; Zhu, Y.; Li, B.; Huang, C.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef] [PubMed]
- Simmons, G.; Zmora, P.; Gierer, S.; Heurich, A.; Pöhlmann, S. Proteolytic activation of the SARS-coronavirus spike protein: Cutting enzymes at the cutting edge of antiviral research. Antivir. Res. 2013, 100, 605–614. [Google Scholar] [CrossRef] [PubMed]
- Ou, X.; Liu, Y.; Lei, X.; Li, P.; Mi, D.; Ren, L.; Guo, L.; Guo, R.; Chen, T.; Hu, J.; et al. Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nature Communications 2020, 11, 1620. [Google Scholar] [CrossRef]
- Yan, R.; Zhang, Y.; Li, Y.; Xia, L.; Guo, Y.; Zhou, Q. Structural basis for the recognition of SARS-CoV-2 by full-length human ACE2. Science 2020, 367, 1444–1448. [Google Scholar] [CrossRef] [PubMed]
- Shang, J.; Ye, G.; Shi, K.; Wan, Y.; Luo, C.; Aihara, H.; Geng, Q.; Auerbach, A.; Li, F. Structural basis of receptor recognition by SARS-CoV-2. Nature 2020, 581, 221–224. [Google Scholar] [CrossRef]
- Ortega, J.T.; Serrano, M.L.; Pujol, F.H.; Rangel, H.R. Role of changes in SARS-CoV-2 spike protein in the interaction with the human ACE2 receptor: An in silico analysis. EXCLI J. 2020, 19, 410–417. [Google Scholar] [CrossRef]
- Wang, Q.; Zhang, Y.; Wu, L.; Niu, S.; Song, C.; Zhang, Z.; Lu, G.; Qiao, C.; Hu, Y.; Yuen, K.; et al. Structural and Functional Basis of SARS-CoV-2 Entry by Using Human ACE2. Cell 2020, 181, 894–904. [Google Scholar] [CrossRef]
- Zhang, L.; Lin, D.; Sun, X.; Curth, U.; Drosten, C.; Sauehering, L.; Becker, S.; Rox, K.; Hilgenfeld, R. Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved α-ketoamide inhibitors. Science 2020, 369, 409–412. [Google Scholar] [CrossRef]
- Chen, Y.W.; Yiu, C.B.; Wong, K. Prediction of the SARS-CoV-2 (2019-nCoV) 3C-like protease (3CL pro) structure: Virtual screening reveals velpatasvir, ledipasvir, and other drug repurposing candidates. F1000Res 2020, 9, 129. [Google Scholar] [CrossRef] [PubMed]
- Xia, S.; Liu, M.; Wang, C.; Xu, W.; Lan, Q.; Feng, S.; Qi, F.; Bao, L.; Du, L.; Liu, S.; et al. Inhibition of SARS-CoV-2 (previously 2019-nCoV) infection by a highly potent pan-coronavirus fusion inhibitor targeting its spike protein that harbors a high capacity to mediate membrane fusion. Cell Res. 2020, 30, 343–355. [Google Scholar] [CrossRef] [PubMed]
- Jin, Z.; Du, X.; Yang, H. Structure of Mpro from SARS-CoV-2 and discovery of its inhibitors. Nature 2020, 582, 289–293. [Google Scholar] [CrossRef]
- Harcourt, B.H.; Jukneliene, D.; Kanjanahaluethai, A.; Bechill, J.; Severson, K.M.; Smith, C.M.; Rota, P.A.; Baker, S.C. Identification of severe acute respiratory syndrome coronavirus replicase products and characterization of papain-like protease activity. J. Virol. 2004, 78, 13600–13612. [Google Scholar] [CrossRef] [PubMed]
- Chu, C.K.; Gadthula, S.; Chen, X.; Choo, H.; Olgen, S.; Barnard, D.L.; Sidwell, R.W. Antiviral activity of nucleoside analogues against SARS-coronavirus (SARS-coV). Antivir. Chem. Chemother. 2006, 17, 285–289. [Google Scholar] [CrossRef] [PubMed]
- Narayanan, N.; Nair, D.T. Vitamin B12 May Inhibit RNA-Dependent-RNA Polymerase Activity of nsp12 from the SARS-CoV-2 Virus. 2020. [Google Scholar] [CrossRef]
- Ruan, Z.; Liu, C.; Guo, Y.; He, Z.; Huang, X.; Jia, X.; Yang, T. Potential Inhibitors Targeting RNA-Dependent RNA Polymerase Activity (NSP12) of SARS-CoV-2. 2020. [Google Scholar] [CrossRef]
- Ivanov, K.A.; Ziebuhr, J. Human coronavirus 229E nonstructural protein 13: Characterization of duplex-unwinding, nucleoside triphosphatase, and RNA 5′-triphosphatase activities. J. Virol. 2004, 78, 7833–7838. [Google Scholar] [CrossRef]
- Shum, K.T.; Tanner, J.A. Differential inhibitory activities and stabilisation of DNA aptamers against the SARS coronavirus helicase. ChemBioChem 2008, 9, 3037–3045. [Google Scholar] [CrossRef]
- Jang, K.J.; Lee, N.; Yeo, W.; Jeong, Y.; Kim, D. Isolation of inhibitory RNA aptamers against severe acute respiratory syndrome (SARS) coronavirus NTPase/Helicase. Biochem. Biophys. Res. Commun. 2008, 366, 738–744. [Google Scholar] [CrossRef]
- Yu, M.; Lee, J.; Lee, J.M.; Kim, Y.; Chin, Y.; Jee, J.; Keum, Y.; Jeong, Y. Identification of myricetin and scutellarein as novel chemical inhibitors of the SARS coronavirus helicase, nsP13. Bioorg. Med. Chemistry Lett. 2012, 22, 4049–4054. [Google Scholar] [CrossRef] [PubMed]
- Bukreyeva, N.; Mantlo, E.K.; Sattler, R.A.; Huang, C.; Paessler, S.; Zeldis, J. The impdh inhibitor merimepodib suppresses sars cov 2 replication in vitro. 2020. [Google Scholar] [CrossRef]
- Madu, I.G.; Chu, V.C.; Lee, H.; Regan, A.D.; Bauman, B.E.; Whittaker, G.R. Heparan sulfate is a selective attachment factor for the avian coronavirus infectious bronchitis virus Beaudette. Avian Dis. 2007, 51, 45–51. [Google Scholar] [CrossRef]
- Watanabe, R.; Sawicki, S.G.; Taguchi, F. Heparan sulfate is a binding molecule but not a receptor for CEACAM1-independent infection of murine coronavirus. Virology 2007, 366, 16–22. [Google Scholar] [CrossRef]
- Milewska, A.; Nowak, P.; Owczarek, K.; Szczepanski, A.; Zarebski, M.; Hoang, A.; Berniak, K.; Wojarski, J.; Zeglen, S.; Baster, Z.; et al. Entry of human coronavirus NL63 into the cell. J. Virol 2018, 92, e01933-17. [Google Scholar] [CrossRef]
- Naskalska, A.; Dabrowska, A.; Szczepanski, A.; Milewska, A.; Jasik, K.P.; Pyrc, K. Membrane Protein of Human Coronavirus NL63 Is Responsible for Interaction with the Adhesion Receptor. J. Virol. 2019, 93, e00355-19. [Google Scholar] [CrossRef] [PubMed]
- Lang, J.; Yang, N.; Deng, J.; Liu, K.; Yang, P.; Zhang, G.; Jiang, C. Inhibition of SARS pseudovirus cell entry by lactoferrin binding to heparan sulfate proteoglycans. PLoS ONE 2011, 6, e23710. [Google Scholar] [CrossRef]
- Zhou, Y.; Jiang, X.; Tong, T.; Fang, L.; Wu, Y.; Liang, J.; Xiao, S. High antiviral activity of mercaptoethane sulfonate functionalized Te/BSA nanostars against arterivirus and coronavirus. RSC Adv. 2020, 10, 14161–14169. [Google Scholar] [CrossRef]
- Tiwari, V.; Liu, J.; Valyi-Nagy, T.; Shukla, D. Anti-heparan sulfate peptides that block herpes simplex virus infection in vivo. J. Biol. Chem. 2011, 286, 25406–25415. [Google Scholar] [CrossRef]
- Jaishankar, D.; Yakoub, A.M.; Bogdanov, A.; Valyi-Nagy, T.; Shukla, D. Characterization of a proteolytically stable D-peptide that suppresses herpes simplex virus 1 infection: Implications for the development of entry-based antiviral therapy. J. Virol. 2015, 89, 1932–1938, DOI 101128/JVI02979. [Google Scholar] [CrossRef]
- Jaishankar, D.; Buhrman, J.S.; Valyi-Nagy, T.; Gemeinhart, R.A.; Shukla, D. Extended Release of an Anti–Heparan Sulfate Peptide From a Contact Lens Suppresses Corneal Herpes Simplex Virus-1 Infection. Investigative ophthalmology & visual science 2016, 57, 169–180. [Google Scholar] [CrossRef]
- Hadigal, S.R.; Agelidis, A.M.; Karasneh, G.A.; Antoine, T.E.; Yakoub, A.M.; Ramani, V.C.; Djalilian, A.R.; Sanderson, R.D.; Shukla, D. Heparanase is a host enzyme required for herpes simplex virus-1 release from cells. Nat. Commun. 2015, 6, 6985. [Google Scholar] [CrossRef] [PubMed]
- Monteil, V.; Kwon, H.; Prado, P.; Hangelkruys, A.; Wimmer, R.; Stahl, M.; Leopoldi, A.; Garreta, E.; Penninger, J. Inhibition of SARS-CoV-2 infections in engineered human tissues using clinical-grade soluble human ACE2. Cell Press 2020, 181, 905–913. [Google Scholar]
- Adedeji, A.O.; Severson, W.; Jonsson, C.; Singh, K.; Weiss, S.R.; Sarafianos, S.G. Novel inhibitors of severe acute respiratory syndrome coronavirus entry that act by three distinct mechanisms. J. Virol. 2013, 87, 8017–8028. [Google Scholar] [CrossRef]
- Zhang, H.; Penninger, J.M.; Li, Y.; Zhong, N.; Slutsky, A.S. Angiotensin-converting enzyme 2 (ACE2) as a SARS-CoV-2 receptor: Molecular mechanisms and potential therapeutic target. Intensive Care Med. 2020, 46, 586–590. [Google Scholar] [CrossRef]
- Li, F.; Li, W.; Farzan, M.; Harrison, S.C. Structure of SARS coronavirus spike receptor-binding domain complexed with receptor. Science 2005, 309, 1864–1868. [Google Scholar] [CrossRef]
- Chappell, M.; Leibbrandt, A.; Liu, Y.; Gao, H.; Rao, S.; Guo, F.; Zheng, D.; Qin, C.; Liu, D.; Deng, W.; et al. A crucial role of angiotensin converting enzyme 2 (ACE2) in SARS coronavirus-induced lung injury. Nat. Med. 2005, 11, 875–879. [Google Scholar] [CrossRef]
- Imai, Y.; Kuba, K.; Rao, S.; Huan, Y.; Guo, F.; Guan, B.; Yang, P.; Sarao, R.; Wada, T.; Leong-Poi, H.; et al. Angiotensin-converting enzyme 2 protects from severe acute lung failure. Nature 2005, 436, 112–116. [Google Scholar] [CrossRef]
- Amanat, F.; Krammer, F. SARS-CoV-2 Vaccines: Status Report. Immunity 2020, 52, 583–589. [Google Scholar] [CrossRef]
- Agostini, M.L.; Andres, E.L.; Sims, A.C.; Graham, R.L.; Sheahan, T.P.; Lu, X.; Smith, E.C.; Case, J.B.; Feng, J.Y.; Jordan, R.; et al. Coronavirus Susceptibility to the Antiviral Remdesivir (GS-5734) Is Mediated by the Viral Polymerase and the Proofreading Exoribonuclease. mBio 2018, 9. [Google Scholar] [CrossRef]
- Elfiky, A.A. Ribavirin, Remdesivir, Sofosbuvir, Galidesivir, and Tenofovir against SARS-CoV-2 RNA dependent RNA polymerase (RdRp): A molecular docking study. Life Sci. 2020, 253. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, A.K.; Brindisi, M.; Shahabi, D.; Chapman, M.E.; Mesecar, A.D. Drug Development and Medicinal Chemistry Efforts Toward SARS-Coronavirus and Covid-19 Therapeutics. ChemMedChem 2020, 15, 907–932. [Google Scholar] [CrossRef] [PubMed]
- Grein, J.; Ohmagari, N.; Shin, D.; Diaz, G.; Asperges, E.; Castagna, A.; Feldt, T.; Green, G.; Green, M.L.; Lescure, F.; et al. Compassionate Use of Remdesivir for Patients with Severe Covid-19. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef]
- Kakodkar, P.; Kaka, N.; Baig, M.N. A Comprehensive Literature Review on the Clinical Presentation, and Management of the Pandemic Coronavirus Disease 2019 (COVID-19). Cureus 2020, 12. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, D.; Du, G.; Du, R.; Zhao, J.; Jin, Y.; Fu, S.; Gao, L.; Cheng, Z.; Lu, Q.; et al. Remdesivir in adults with severe COVID-19: A randomised, double-blind, placebo-controlled, multicentre trial. Lancet 2020, 395, 1569–1578. [Google Scholar] [CrossRef]
- Peer-Reviewed Data Shows Remdesivir for COVID-19 Improves Time to Recovery. Available online: https://www.nih.gov/news-events/news-releases/peer-reviewed-data-shows-remdesivir-covid-19-improves-time-recovery (accessed on 22 May 2020).
- Krause, K.L.; Furneaux, R.; Benjes, P.; Brimble, M.; Davidson, T.; Denny, W.; Harris, L.; Hinkley, S.; Tyler, P.; Ussher, J.E.; et al. The post-lockdown period should be used to acquire effective therapies for future resurgence in SARS-Cov-2 infections. N. Z. Med. J. 2020, 133, 107–111. [Google Scholar]
- Li, G.; Clercq, E.D. Therapeutic options for the 2019 novel coronavirus (2019-nCoV). Nat. Rev. Drug Discov. 2020, 19, 149–150. [Google Scholar] [CrossRef]
- Elfiky, A.A. Anti-HCV, nucleotide inhibitors, repurposing against COVID-19. Life Sci. 2020, 248, 117477. [Google Scholar] [CrossRef]
- Furuta, Y.; Komeno, T.; Nakamura, T. Favipiravir (T-705), a broad spectrum inhibitor of viral RNA polymerase. Proc. Jpn. Acad. Ser. B 2017, 93, 449–463. [Google Scholar] [CrossRef]
- Baranovich, T.; Wong, S.-S.; Armstrong, J.; Marjuki, H.; Webby, R.J.; Webster, R.G.; Govorkova, E.A. T-705 (Favipiravir) Induces Lethal Mutagenesis in Influenza A H1N1 Viruses In Vitro. J. Virol. 2013, 87, 3741–3751. [Google Scholar] [CrossRef]
- Jin, Z.; Smith, L.K.; Rajwanshi, V.K.; Kim, B.; Deval, J. The Ambiguous Base-Pairing and High Substrate Efficiency of T-705 (Favipiravir) Ribofuranosyl 5′-Triphosphate towards Influenza A Virus Polymerase. PLoS ONE 2013, 8, e68347. [Google Scholar] [CrossRef] [PubMed]
- Abd El-Aziz, T.M.; Stockand, J.D. Recent progress and challenges in drug development against COVID-19 coronavirus (SARS-CoV-2)—An update on the status. Infect. Genet. Evol. 2020, 83, 104327. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Wang, Y.M.; Xu, J.Y.; Cao, B. Potential antiviral therapeutics for 2019 Novel Coronavirus. Zhonghua Jie He He Hu Xi Za Zhi 2020, 43, 170–172. [Google Scholar] [CrossRef] [PubMed]
- Treatment with CytoDyn′s Leronlimab Indicates Significant Trend Toward Immunological Restoration in Severely Ill COVID-19 Patients. Available online: https://www.cytodyn.com/newsroom/press-releases/detail/405/treatment-with-cytodyns-leronlimab-indicates-significant (accessed on 27 April 2020).
- Brilacidin. Available online: http://www.ipharminc.com/brilacidin-1 (accessed on 27 April 2020).
- Roivant Sciences. Roivant Announces Development of Anti-GM-CSF Monoclonal Antibody to Prevent and Treat Acute Respiratory Distress Syndrome (ARDS) in Patients with COVID-19. Available online: https://roivant.com/roivant-announces-development-of-anti-gm-csf-monoclonal-antibody-to-prevent-and-treat-acute-respiratory-distress-syndrome-ards-in-patients-with-covid-19.html, (accessed on 9 June 2020).
- Mab Explores TJM2 In Treating Severe COVID-19 Disease. Available online: https://www.contractpharma.com/contents/view_breaking-news/2020-03-13/i-mab-explores-tjm2-in-treating-severe-covid-19-disease/ (accessed on 27 April 2020).
- Airway Therapeutics Announces Filing with NIH to Evaluate AT-100 as a Therapy for Novel Coronavirus. Available online: https://www.airwaytherapeutics.com/airway-therapeutics-announces-filing-with-nih-to-evaluate-at-100-as-a-therapy-for-novel-coronavirus/ (accessed on 8 May 2020).
- Tiziana Life Sci PLC - TZLS-501 / COVID-19. Available online: https://ir.tizianalifesciences.com/news-releases/news-release-details/tiziana-life-sci-plc-tzls-501-covid-19/ (accessed on 9 June 2020).
- OyaGen, Inc. Announces a Compound in Development with Broad Antiviral Activity Against Coronaviruses, including SARS-CoV-2. Available online: https://www.biospace.com/article/oyagen-inc-announces-a-compound-in-development-with-broad-antiviral-activity-against-coronaviruses-including-sars-cov-2/ (accessed on 27 April 2020).
- BeyondSpring Files for Patent Protection on BPI-002 for the Treatment of Viral Infections Including COVID-19. Available online: https://www.biospace.com/article/beyondspring-files-for-patent-protection-on-bpi-002-for-the-treatment-of-viral-infections-including-covid-19/ (accessed on 27 April 2020).
- NP-120 (Ifenprodil). Available online: https://algernonpharmaceuticals.com/ipf-np-120/ (accessed on 9 June 2020).
- Pilot Clinical Trial in China to Test UBC Researcher’s Findings as a Targeted Therapy for COVID-19. Available online: https://www.med.ubc.ca/news/pilot-clinical-trial-in-china-to-test-ubc-researchers-findings-as-a-targeted-therapy-for-covid-19/ (accessed on 27 April 2020).
- Roche Ups Actemra Production Anticipating Increased Demand. Available online: https://www.biopharma-reporter.com/Article/2020/03/25/Roche-ups-Actemra-production-for-coronavirus-trials (accessed on 27 April 2020).
- Regeneron and Sanofi Begin Global Kevzara® (sarilumab) Clinical Trial Program in Patients with Severe COVID-19. Available online: https://pipelinereview.com/index.php/2020031674032/Antibodies/Regeneron-and-Sanofi-Begin-Global-Kevzara-sarilumab-Clinical-Trial-Program-in-Patients-with-Severe-COVID-19.html (accessed on 27 April 2020).
- COVID-19. Available online: https://www.synairgen.com/covid-19/ (accessed on 27 April 2020).
- AmnioBoost For Covid-19 Clincal Study Recruitment. Available online: http://www.latticebiologics.com/amnioboost-for-covid-19-clincal-study-recruitment/ (accessed on 9 June 2020).
- Hydroxychloroquine or Chloroquine for COVID-19: Drug Safety Communication—FDA Cautions Against Use Outside of the Hospital Setting or a Clinical Trial Due to Risk of Heart Rhythm Problems. Available online: https://www.fda.gov/safety/medical-product-safety-information/hydroxychloroquine-or-chloroquine-covid-19-drug-safety-communication-fda-cautions-against-use (accessed on 27 April 2020).
- Enanta Pharmaceuticals Announces Efforts to Discover a Treatment for the Novel Coronavirus Disease COVID-19. Available online: https://www.enanta.com/investors/news-releases/press-release/2020/Enanta-Pharmaceuticals-Announces-Efforts-to-Discover-a-Treatment-for-the-Novel-Coronavirus-Disease-COVID-19/default.aspx (accessed on 27 April 2020).
- Emergent BioSolutions Signs Agreement to be U.S. Manufacturing Partner for Johnson & Johnson’s Lead Vaccine Candidate for COVID-19. Available online: https://investors.emergentbiosolutions.com/news-releases/news-release-details/emergent-biosolutions-signs-agreement-be-us-manufacturing (accessed on 27 April 2020).
- Embattled Cel-Sci to tackle COVID-19 with immunotherapy platform tech LEAPS. Available online: https://www.fiercebiotech.com/research/embattled-cel-sci-to-tackle-covid-19-immunotherapy-platform-tech-leaps (accessed on 27 April 2020).
- Takeda Initiates Development of a Plasma-Derived Therapy for COVID-19. Available online: https://www.takeda.com/newsroom/newsreleases/2020/takeda-initiates-development-of-a-plasma-derived-therapy-for-covid-19/ (accessed on 27 April 2020).
- Coronavirus vaccine from BioNTech, Pfizer Set to Enter Testing in Germany. Available online: https://www.biopharmadive.com/news/coronavirus-pfizer-biontech-vaccine-study-start-germany/576522/ (accessed on 27 April 2020).
- Mateon Announces the Filing of an IND with US FDA to Evaluate its Antisense Drug Candidate OT-101 in COVID-19 patients. Available online: http://investor.mateon.com/news-releases/news-release-details/mateon-announces-filing-ind-us-fda-evaluate-its-antisense-drug (accessed on 27 April 2020).
- SwRI’s Virtual Screening Software Rhodium Increases Processing Power for Coronavirus Research. Available online: https://www.swri.org/press-release/virtual-screening-software-rhodium-coronavirus-research (accessed on 27 April 2020).
- Coronavirus Drug Development Update from NanoViricides, Inc. Available online: https://www.marketwatch.com/press-release/coronavirus-drug-development-update-from-nanoviricides-inc-2020-03-16-62641516 (accessed on 27 April 2020).
- Vir Biotechnology Identifies Two Antibodies That Bind to the Spike Protein of 2019-nCoV, Newly Named as SARS-CoV-2. Available online: https://investors.vir.bio/news-releases/news-release-details/vir-biotechnology-identifies-two-antibodies-bind-spike-protein (accessed on 27 April 2020).
- Chu, C.M.; Cheng, V.C.C.; Hung, I.F.N.; Wong, M.M.L.; Chan, K.H.; Chan, K.S.; Kao, R.Y.T.; Poon, L.L.M.; Wong, C.L.P.; Guan, Y.; et al. Role of lopinavir/ritonavir in the treatment of SARS: Initial virological and clinical findings. Thorax 2004, 59, 252–256. [Google Scholar] [CrossRef]
- Lopinavir; A Potent Drug against Coronavirus Infection: Insight from Molecular Docking Study. Available online: http://archcid.com/en/articles/13823.html (accessed on 27 April 2020).
- Johnson & Johnson. Johnson & Johnson Launches Multi-Pronged Response to Coronavirus Global Public Health Threat. Available online: https://www.jnj.com/johnson-johnson-launches-multi-pronged-response-to-coronavirus-global-public-health-threat (accessed on 27 April 2020).
- The Convalescent Sera Option for Containing COVID-19. Available online: https://www.jci.org/articles/view/138003/pdf (accessed on 25 April 2020).
- Bloch, E. Deployment of Convalescent Plasma for the Prevention and Treatment of COVID-19. Available online: https://www.jci.org/articles/view/138745/pdf (accessed on 25 April 2020).
- Keith, P.; Day, M.; Perkins, L.; Moyer, L.; Hewitt, K.; Wells, A. A novel treatment approach to the novel coronavirus: An argument for the use of therapeutic plasma exchange for fulminant COVID-19. Crit. Care 2020, 24, 128. [Google Scholar] [CrossRef]
- Shen, C.; Wang, Z.; Zhao, F.; Yang, Y.; Li, J.; Yuan, J.; Wang, F.; Li, D.; Yang, M.; Xing, L.; et al. Treatment of 5 Critically Ill Patients With COVID-19 With Convalescent Plasma. JAMA 2020, 323, 1582–1589. [Google Scholar] [CrossRef]
- Schrezenmeier, E.; Dörner, T. Mechanisms of action of hydroxychloroquine and chloroquine: Implications for rheumatology. Nat. Rev. Rheumatol. 2020, 16, 155–166. [Google Scholar] [CrossRef]
- Lentini, G.; Cavalluzzi, M.M.; Habtemariam, S. COVID-19, Chloroquine Repurposing, and Cardiac Safety Concern: Chirality Might Help. Molecules 2020, 25. [Google Scholar] [CrossRef]
- Mehra, M.; Desai, S.; Ruschitzka, F.; Patel, A. Hydroxychloroquine or chloroquine with or without a macrolide for treatment of COVID-19: a multinational registry analysis. Lancet 2020. [Google Scholar] [CrossRef]
- Raaben, M.; Einerhand, A.W.C.; Taminiau, L.J.A.; van Houdt, M.; Bouma, J.; Raatgeep, R.H.; Büller, H.A.; de Haan, C.A.M.; Rossen, J.W.A. Cyclooxygenase activity is important for efficient replication of mouse hepatitis virus at an early stage of infection. Virol. J. 2007, 4, 55. [Google Scholar] [CrossRef]
- Vane, J.R.; Botting, R.M. Mechanism of action of nonsteroidal anti-inflammatory drugs. Am. J. Med. 1998, 104, 2S–22S. [Google Scholar] [CrossRef]
- Amici, C.; Di Caro, A.; Ciucci, A.; Chiappa, L.; Castilletti, C.; Martella, V.; Decaro, N.; Buonavoglia, C.; Capobianchi, M.R.; Santoro, M.G. Indomethacin has a potent antiviral activity against SARS coronavirus. Antivir. Ther. 2006, 11, 1021–1030. [Google Scholar] [PubMed]
- Caly, L.; Druce, J.D.; Catton, M.G.; Jans, D.A.; Wagstaff, K.M. The FDA-approved drug ivermectin inhibits the replication of SARS-CoV-2 in vitro. Antivir. Res. 2020, 178, 104787. [Google Scholar] [CrossRef]
- Wagstaff, K.M.; Sivakumaran, H.; Heaton, S.M.; Harrich, D.; Jans, D.A. Ivermectin is a specific inhibitor of importin α/β-mediated nuclear import able to inhibit replication of HIV-1 and dengue virus. Biochem. J. 2012, 443, 851–856. [Google Scholar] [CrossRef] [PubMed]
- Lundberg, L.; Pinkham, C.; Baer, A.; Amaya, M.; Narayanan, A.; Wagstaff, K.M.; Jans, D.A.; Kehn-Hall, K. Nuclear import and export inhibitors alter capsid protein distribution in mammalian cells and reduce Venezuelan Equine Encephalitis Virus replication. Antivir. Res. 2013, 100, 662–672. [Google Scholar] [CrossRef]
- Prompetchara, E.; Ketloy, C.; Palaga, T. Immune responses in COVID-19 and potential vaccines: Lessons learned from SARS and MERS epidemic. Asian Pac. J. Allergy Immunol. 2020, 38, 1–9. [Google Scholar] [CrossRef]
- Moderna Announces Positive Interim Phase 1 Data for its mRNA Vaccine (mRNA-1273) Against Novel Coronavirus. Available online: https://investors.modernatx.com/news-releases/news-release-details/moderna-announces-positive-interim-phase-1-data-its-mrna-vaccine (accessed on 9 June 2020).
- Shi, Y.; Wang, N.; Zou, Q.M. Progress and challenge of vaccine development against 2019 novel coronavirus (2019-nCoV). Zhonghua Yu Fang Yi Xue Za Zhi 2020, 54, E029. [Google Scholar] [CrossRef]
- Pardi, N.; Hogan, M.J.; Porter, F.W.; Weissman, D. mRNA vaccines—A new era in vaccinology. Nat. Rev. Drug Discov. 2018, 17, 261–279. [Google Scholar] [CrossRef]
- Fusogenix Drug Delivery System | Entos Pharmaceuticals. Available online: https://www.entospharma.com/fusogenix (accessed on 27 April 2020).
- Altimmune AdCOVID™ | Altimmune. 2020. Available online: https://altimmune.com/adcovid/ (accessed on 27 April 2020).
- Medicago Develops Plant-Based Coronavirus Vaccine Candidate. Available online: https://www.biospace.com/article/medicago-successfully-produces-a-viable-vaccine-candidate-for-covid-19/ (accessed on 27 April 2020).
- INO-4800 DNA Coronavirus Vaccine. Available online: https://www.precisionvaccinations.com/vaccines/ino-4800-dna-coronavirus-vaccine (accessed on 27 April 2020).
- MIGAL’s Coronavirus Vaccine Project. Available online: https://www.migal.org.il/en/node/7010 (accessed on 27 April 2020).
- TNX-1800 (Coronavirus Vaccine). Available online: https://www.tonixpharma.com/pipeline/tnx-1800-coronavirus-vaccine (accessed on 27 April 2020).
- CEPI announces COVID-19 vaccine development partnership with Clover Biopharmaceuticals’ Australian Subsidiary. Available online: http://www.cloverbiopharma.com/index.php?m=content&c=index&a=show&catid=11&id=44 (accessed on 27 April 2020).
- Vaxart Announces Positive Pre-Clinical Data for its Oral COVID-19 Vaccine Program. Available online: https://investors.vaxart.com/news-releases/news-release-details/vaxart-announces-positive-pre-clinical-data-its-oral-covid-19 (accessed on 27 April 2020).
- Applied DNA Sciences Applied DNA Sciences Subsidiary, LineaRx, and Takis Biotech Collaborate for Development of a Linear DNA Vaccine Candidate Against Wuhan Coronavirus 2019-nCoV. 2020. Available online: https://www.biospace.com/article/releases/applied-dna-sciences-subsidiary-linearx-and-takis-biotech-collaborate-for-development-of-a-linear-dna-vaccine-candidate-against-wuhan-coronavirus-2019-ncov/ (accessed on 27 April 2020).
- Modjarrad, K. MERS-CoV vaccine candidates in development: The current landscape. Vaccine 2016, 34, 2982–2987. [Google Scholar] [CrossRef]
- INO-4700 MERS-CoV Vaccine. Available online: https://www.precisionvaccinations.com/vaccines/ino-4700-mers-cov-vaccine-0 (accessed on 27 April 2020).
- Predictive Oncology Launches New AI Platform for Vaccine and Drug Development Targeting Coronaviruses and Acute Respiratory Syndromes (COVID-19, MERS, and SARS). Available online: https://apnews.com/Globe%20Newswire/46fd8fb75ea454dca6a07b38fa20c7d6 (accessed on 27 April 2020).
- Integral Molecular Integral Molecular Accelerates Coronavirus Vaccine Research. 2020. Available online: https://www.integralmolecular.com/integral-molecular-accelerates-coronavirus-vaccine-research/ (accessed on 27 April 2020).
- AJ Vaccines To Develop Vaccine for COVID-19. Available online: https://www.biospace.com/article/aj-vaccines-to-develop-vaccine-for-covid-19-/ (accessed on 27 April 2020).
- Heat Biologics Announces Research Collaboration with University of Miami to Develop Vaccine Designed to Protect Against COVID-19 Coronavirus. Available online: https://www.heatbio.com/news-media/news-releases/detail/649/heat-biologics-announces-research-collaboration-with (accessed on 27 April 2020).
- Hong Kong researchers develop coronavirus vaccine. Available online: https://www.bioworld.com/articles/432835-hong-kong-researchers-develop-coronavirus-vaccine?v=preview (accessed on 27 April 2020).
- Generex Signs Contract with EpiVax to Develop Ii-Key Peptide Vaccines to Address the Coronavirus Pandemic. Available online: https://www.biospace.com/article/generex-signs-contract-with-epivax-to-develop-ii-key-peptide-vaccines-to-address-the-coronavirus-pandemic/ (accessed on 27 April 2020).
- Tulane University launches new coronavirus research program to develop a vaccine and advanced diagnostics. Available online: https://news.tulane.edu/pr/tulane-university-launches-new-coronavirus-research-program-develop-vaccine-and-advanced (accessed on 27 April 2020).
- Inc, P. ImmunoPrecise Launches Coronavirus Vaccine and Therapeutic Antibody Program. 2020. Available online: https://www.pharmiweb.com/press-release/2020-02-20/immunoprecise-launches-coronavirus-vaccine-and-therapeutic-antibody-program (accessed on 27 April 2020).
- Codagenix and Serum Institute of India Initiate Co-Development of a Scalable, Live-Attenuated Vaccine Against the 2019 Novel Coronavirus, COVID-19. Available online: https://www.biospace.com/article/codagenix-and-serum-institute-of-india-initiate-co-development-of-a-scalable-live-attenuated-vaccine-against-the-2019-novel-coronavirus-covid-19/ (accessed on 27 April 2020).
- Zydus Cadila launches a fast tracked programme to develop vaccine for the novel coronavirus, 2019-nCoV (COVID-19). Available online: https://pipelinereview.com (accessed on 27 April 2020).
- Investigational ChAdOx1 nCoV-19 Vaccine Protects Monkeys Against COVID-19 Pneumonia. Available online: https://www.nih.gov/news-events/news-releases/investigational-chadox1-ncov-19-vaccine-protects-monkeys-against-covid-19-pneumonia (accessed on 22 May 2020).
- Munster, V.J.; Wells, D.; Lambe, T.; Wright, D.; Fischer, R.J.; Bushmaker, T.; Saturday, G.; van Doremalen, N.; Gilbert, S.C.; de Wit, E.; et al. Protective efficacy of a novel simian adenovirus vaccine against lethal MERS-CoV challenge in a transgenic human DPP4 mouse model. NPJ Vaccines 2017, 2. [Google Scholar] [CrossRef]
- Alharbi, N.K.; Padron-Regalado, E.; Thompson, C.P.; Kupke, A.; Wells, D.; Sloan, M.A.; Grehan, K.; Temperton, N.; Lambe, T.; Warimwe, G.; et al. ChAdOx1 and MVA based vaccine candidates against MERS-CoV elicit neutralising antibodies and cellular immune responses in mice. Vaccine 2017, 35, 3780–3788. [Google Scholar] [CrossRef] [PubMed]
- Folegatti, P.M.; Bittaye, M.; Flaxman, A.; Lopez, F.R.; Bellamy, D.; Kupke, A.; Mair, C.; Makinson, R.; Sheridan, J.; Rohde, C.; et al. Safety and immunogenicity of a candidate Middle East respiratory syndrome coronavirus viral-vectored vaccine: A dose-escalation, open-label, non-randomised, uncontrolled, phase 1 trial. Lancet Infect. Dis. 2020. [Google Scholar] [CrossRef]
- Mousavizadeh, L.; Ghasemi, S. Genotype and phenotype of COVID-19: Their roles in pathogenesis. J. Microbiol. Immunol. Infect. 2020. [Google Scholar] [CrossRef] [PubMed]


| Sr. No. | Name of Viral Protein | Function |
|---|---|---|
| Structural Proteins | ||
| 1 | Nucleocapsid (N) protein | Houses the viral genome. |
| 2 | Envelope (E) protein | Encases nucleocapsid and makes complete viral envelope along with M protein. |
| 3 | Membrane (M) glycoprotein | M and E protein makes complete viral envelope. |
| 4 | Spike (S) glycoprotein | On the surface of envelope, characteristic spike proteins. Attachment of virus with receptor at receptor binding domain (RBD). Cellular serine protease (TMPRSS2/Cathepsin B) cleaves S protein this facilitates viral entry. |
| Non-structural proteins (Nsp) | ||
| 5 | Mpro | Also called as 3-chymotrypsin-like protease (3Clpro) It cleaves viral polyproteins pp1a and pp1ab at 11 sites to yield Nsp4 to Nsp16. |
| 6 | PLpro | It cleaves viral polyprotein at N terminus to yield Nsp1 to Nsp3 |
| 7 | Nsp12 | RNA dependent RNA polymerase Replication of virus |
| 8 | Nsp13 | Helicase activity facilitates RNA dependent RNA polymerase (RdRp) during viral RNA replication |
| 9 | Nsp1, Nsp3c, and open reading frame (ORF) 7a | Inhibiting host immune response |
| Drugs in Trial | ||
|---|---|---|
| Name: | Company: | Reference: |
| leronlimab | CytoDyn | [70] |
| Brilacidin | Innovation Pharmaceuticals | [71] |
| Gimsilumab | Roivant Sciences | [72] |
| TJM2 | I-Mab Biopharma | [73] |
| AT-100 | Airway Therapeutics | [74] |
| TZLS-501 | Tiziana Life Sciences | [75] |
| OYA1 | OyaGen | [76] |
| BPI-002 | BeyondSpring | [77] |
| NP-120 (Ifenprodil) | Algernon Pharmaceuticals | [78] |
| APN01 | University of British Columbia and APEIRON Biologics | [79] |
| Remdesivir (GS-5734) | Gilead Sciences | (ClinicalTrials.gov: NCT04257656, NCT04252664). |
| Actemra | Roche | [80] |
| Galidesivir | Biocryst Pharma | (ClinicalTrials.gov: NCT03891420) |
| REGN3048-3051 and Kevzara | Regeneron | [81] |
| SNG001 | Synairgen Research | [82] |
| AmnioBoost | Lattice Biologics | [83] |
| Chloroquine | US FDA | [84] |
| Favilavir | Toyama Chemical | (ClinicalTrials.gov: NCT04336904) |
| Experimental Drugs | ||
| Name: | Company: | Reference: |
| Existing antiviral compound library | Enanta Pharmaceuticals | [85] |
| Plasma-derived product candidates | Emergent BioSolutions | [86] |
| Ligand Epitope Antigen Presentation System (LEAPS) peptide immunotherapy | CEL-SCI | [87] |
| Hyperimmune globulin (H-IG) therapy | Takeda Pharmaceutical Company | [88] |
| Novel compounds for Therapy | Pfizer | [89] |
| Artificial intelligence (AI) platforms for drug discovery | Mateon Therapeutics | [90] |
| Rhodium virtual screening | Southwest Research Institute | [91] |
| Nanoviricide® technology | NanoViricides | [92] |
| Monoclonal antibodies | Vir Biotechnology | [93] |
| HIV drugs for coronavirus treatment | Abbvie | [94] |
| LOPIMUNE, HIV Drug | Cipla | [95] |
| PREZCOBIX® HIV medication (darunavir/cobicistat) | Janssen Pharmaceutical Companies | [96] |
| Vaccines in Development | |||
|---|---|---|---|
| Name: | Company: | Platform | Reference: |
| Fusogenix DNA vaccine | Entos Pharmaceuticals | DNA vaccine | [114] |
| ChAdOx1 nCoV-19 | University of Oxford | Adenovirus Vector | (ClinicalTrials.gov: NCT04324606) |
| AdCOVID | Altimmune | Intranasal Nasovax | [115] |
| Coronavirus vaccine | Medicago | Virus Like Particles | [116] |
| INO-4800 | Inovio Pharmaceuticals and Beijing Advaccine Biotechnology | DNA vaccine | [117] |
| mRNA-1273 vaccine | Moderna and Vaccine Research Center | mRNA vaccine | (ClinicalTrials.gov: NCT0428346). |
| Avian Coronavirus Infectious Bronchitis Virus (IBV) vaccine | MIGAL Research Institute | Protein expression vector | [118] |
| TNX-1800 | Tonix Pharmaceuticals | live modified horsepox vaccine | [119] |
| Recombinant subunit vaccine | Clover Biopharmaceuticals | Protein based Trimer vaccine | [120] |
| Coronavirus vaccine | Vaxart Inc | Oral recombinant Protein vaccine | [121] |
| Linear DNA Vaccine | Applied DNA Sciences and Takis Biotech | DNA vaccine | [122] |
| NVX-CoV2373 | Novavax | Protein based | [123] |
| INO-4700 | Inovio Pharma | DNA vaccine | [124] |
| Other Vaccines | |||
| Name: | Company: | Reference: | |
| AI Platform for the discovery and development of vaccines | Predictive Oncology | N/A | [125] |
| Shotgun Mutagenesis Epitope Mapping Platform | Integral Molecular | N/A | [126] |
| Develop antigens that mimic the native structures of the virus | AJ Vaccines | N/A | [127] |
| gp96 vaccine platform | Heat Biologics | N/A | [128] |
| B-cell and T-cell epitopes for Vaccine development | Hong Kong University of Science and Technology | N/A | [129] |
| COVID-19 vaccine | Vaccine by Generex | N/A | [130] |
| potential coronavirus vaccine | Vaccine by Tulane University | N/A | [131] |
| Cell Select™ and DeepDisplay™ discovery platforms | Coronavirus vaccine by ImmunoPrecise Antibodies | N/A | [132] |
| COVID-19 vaccine | Serum Institute of India | N/A | [133] |
| DNA vaccine/ recombinant measles virus vector -based vaccine | Zydus Cadila | N/A | [134] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, D.; Koganti, R.; Lambe, U.P.; Yadavalli, T.; Nandi, S.S.; Shukla, D. Vaccines and Therapies in Development for SARS-CoV-2 Infections. J. Clin. Med. 2020, 9, 1885. https://doi.org/10.3390/jcm9061885
Wu D, Koganti R, Lambe UP, Yadavalli T, Nandi SS, Shukla D. Vaccines and Therapies in Development for SARS-CoV-2 Infections. Journal of Clinical Medicine. 2020; 9(6):1885. https://doi.org/10.3390/jcm9061885
Chicago/Turabian StyleWu, David, Raghuram Koganti, Upendra P. Lambe, Tejabhiram Yadavalli, Shyam S. Nandi, and Deepak Shukla. 2020. "Vaccines and Therapies in Development for SARS-CoV-2 Infections" Journal of Clinical Medicine 9, no. 6: 1885. https://doi.org/10.3390/jcm9061885
APA StyleWu, D., Koganti, R., Lambe, U. P., Yadavalli, T., Nandi, S. S., & Shukla, D. (2020). Vaccines and Therapies in Development for SARS-CoV-2 Infections. Journal of Clinical Medicine, 9(6), 1885. https://doi.org/10.3390/jcm9061885






