Genetic and Epigenetic Etiology Underlying Autism Spectrum Disorder
Abstract
1. Introduction
2. Environmental and Prenatal Factors that Cause ASD
2.1. Viral Infection
2.2. Parental Age
2.3. Zinc Deficiency
3. Genetic Epidemiology
3.1. Chromosome Loci that Affect ASD
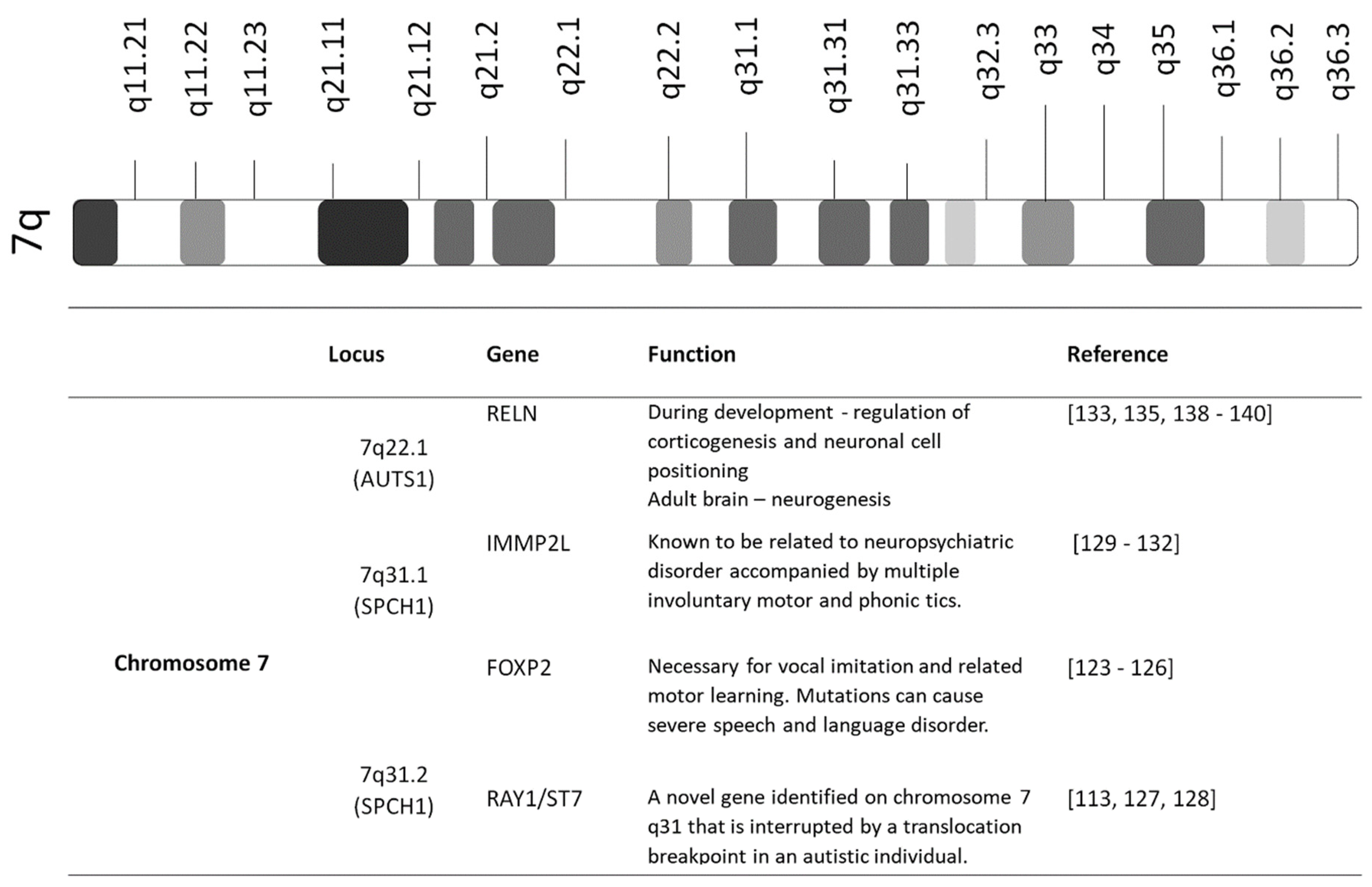
3.2. Candidate Genes on Chromosome 7
3.2.1. FOXP2
3.2.2. RAY1/ST7
3.2.3. IMMP2L
3.2.4. RELN
3.3. Neurodevelopmental Disorders and ASD by CNV
4. Epigenetic Dysregulation Underlying ASD
4.1. DNA Methylation
4.1.1. MeCP2
4.1.2. OXTR
4.1.3. SHANK3
4.2. Histone Modification and Chromatin Remodeling
4.3. MicroRNAs
5. Social Interaction Genes Associated with ASD
6. Conclusion and Future Perspectives
Author Contributions
Funding
Conflicts of Interest
References
- Quaak, I.; Brouns, M.R.; Van de Bor, M. The dynamics of autism spectrum disorders: How neurotoxic compounds and neurotransmitters interact. Int. J. Environ. Res. Public Health 2013, 10, 3384–3408. [Google Scholar] [CrossRef]
- Kanner, L.J. Autistic disturbances of affective contact. Nerv. Child 1943, 2, 217–250. [Google Scholar]
- Fombonne, E. The epidemiology of autism: A review. Psychol. Med. 1999, 29, 769–786. [Google Scholar] [CrossRef]
- Kim, Y.S.; Leventhal, B.L.; Koh, Y.-J.; Fombonne, E.; Laska, E.; Lim, E.-C.; Cheon, K.-A.; Kim, S.-J.; Kim, Y.-K.; Lee, H.; et al. Prevalence of Autism Spectrum Disorders in a Total Population Sample. Am. J. Psychiatry 2011, 168, 904–912. [Google Scholar] [CrossRef]
- Saemundsen, E.; Magnússon, P.; Georgsdóttir, I.; Egilsson, E.; Rafnsson, V. Prevalence of autism spectrum disorders in an Icelandic birth cohort. BMJ Open 2013, 3, e002748. [Google Scholar] [CrossRef]
- Wan, Y.; Hu, Q.; Li, T.; Jiang, L.; Du, Y.; Feng, L.; Wong, J.C.-M.; Li, C.-B. Prevalence of autism spectrum disorders among children in China: A systematic review. Shanghai Arch. Psychiatry 2013, 25, 70–80. [Google Scholar]
- Russell, G.; Rodgers, L.R.; Ukoumunne, O.C.; Ford, T. Prevalence of Parent-Reported ASD and ADHD in the UK: Findings from the Millennium Cohort Study. J. Autism Dev. Disord. 2013, 44, 31–40. [Google Scholar] [CrossRef]
- Zeldovich, L. The Evolution of ‘Autism’As a Diagnosis, Explained; Spectrum: New York, NY, USA, 2018. [Google Scholar]
- Baio, J.; Wiggins, L.; Christensen, D.L.; Maenner, M.J.; Daniels, J.; Warren, Z.; Kurzius-Spencer, M.; Zahorodny, W.; Robinson, C.; Rosenberg, C.R.; et al. Prevalence of Autism Spectrum Disorder Among Children Aged 8 Years—Autism and Developmental Disabilities Monitoring Network, 11 Sites, United States, 2014. Mmwr. Surveill. Summ. 2018, 67, 1–23. [Google Scholar] [CrossRef]
- Canitano, R. Epilepsy in autism spectrum disorders. Eur. Child Adolesc. Psychiatry 2006, 16, 61–66. [Google Scholar] [CrossRef]
- Yasuhara, A. Correlation between EEG abnormalities and symptoms of autism spectrum disorder (ASD). Brain Dev. 2010, 32, 791–798. [Google Scholar] [CrossRef]
- Valicenti-McDermott, M.; McVicar, K.A.; Rapin, I.; Wershil, B.K.; Cohen, H.; Shinnar, S. Frequency of Gastrointestinal Symptoms in Children with Autistic Spectrum Disorders and Association with Family History of Autoimmune Disease. J. Dev. Behav. Pediatr. 2006, 27, S128–S136. [Google Scholar] [CrossRef]
- Richdale, A.L.; Schreck, K.A. Sleep problems in autism spectrum disorders: Prevalence, nature, & possible biopsychosocial aetiologies. Sleep Med. Rev. 2009, 13, 403–411. [Google Scholar]
- White, S.; Oswald, D.; Ollendick, T.H.; Scahill, L. Anxiety in children and adolescents with autism spectrum disorders. Clin. Psychol. Rev. 2009, 29, 216–229. [Google Scholar] [CrossRef]
- De Rubeis, S.; He, X.; Goldberg, A.P.; Poultney, C.S.; Samocha, K.; Cicek, A.E.; Kou, Y.; Liu, L.; Fromer, M.; Walker, S. Synaptic, transcriptional and chromatin genes disrupted in autism. Nature 2014, 515, 209–215. [Google Scholar] [CrossRef]
- Ng, M.; de Montigny, J.G.; Ofner, M.; Docé, M.T. Health promotion and Chronic Disease Prevention in Canada: Research, Practice. Environmental Factors Associated with Autism Spectrum Disorder: A Scoping Review for the Years 2003–2013. HPCDP 2017, 37, 1. [Google Scholar]
- Bailey, A.; Le Couteur, A.; Gottesman, I.; Bolton, P.; Simonoff, E.; Yuzda, E.; Rutter, M. Autism as a strongly genetic disorder: Evidence from a British twin study. Psychol. Med. 1995, 25, 63–77. [Google Scholar] [CrossRef]
- Ciernia, A.V.; LaSalle, J. The landscape of DNA methylation amid a perfect storm of autism aetiologies. Nat. Rev. Neurosci. 2016, 17, 411–423. [Google Scholar] [CrossRef]
- Hultman, C.M.; Sparen, P.; Cnattingius, S. Perinatal risk factors for infantile autism. Epidemiology 2002, 13, 417–423. [Google Scholar] [CrossRef]
- Clemens, L.E.; Siiteri, P.K.; Stites, D.P. Mechanism of immunosuppression of progesterone on maternal lymphocyte activation during pregnancy. J. Immunol. 1979, 122, 1978–1985. [Google Scholar]
- Deykin, E.Y.; MacMahon, B. Viral exposure and autism. Am. J. Epidemiol. 1979, 109, 628–638. [Google Scholar] [CrossRef]
- Atladottir, H.O.; Thorsen, P.; Ostergaard, L.; Schendel, D.E.; Lemcke, S.; Abdallah, M.; Parner, E.T. Maternal infection requiring hospitalization during pregnancy and autism spectrum disorders. J. Autism. Dev. Disord. 2010, 40, 1423–1430. [Google Scholar] [CrossRef]
- Zerbo, O.; Qian, Y.; Yoshida, C.; Grether, J.K.; Van de Water, J.; Croen, L.A. Maternal Infection During Pregnancy and Autism Spectrum Disorders. J. Autism. Dev. Disord. 2015, 45, 4015–4025. [Google Scholar] [CrossRef]
- Karimi, P.; Kamali, E.; Mousavi, S.M.; Karahmadi, M. Environmental factors influencing the risk of autism. J. Res. Med. Sci. 2017, 22, 27. [Google Scholar] [CrossRef]
- Chess, S. Follow-up report on autism in congenital rubella. J. Autism. Child Schizophr. 1977, 7, 69–81. [Google Scholar] [CrossRef]
- Madsen, K.M.; Hviid, A.; Vestergaard, M.; Schendel, D.; Wohlfahrt, J.; Thorsen, P.; Olsen, J.; Melbye, M. A population-based study of measles, mumps, and rubella vaccination and autism. N. Engl. J. Med. 2002, 347, 1477–1482. [Google Scholar] [CrossRef]
- Libbey, J.E.; Sweeten, T.L.; McMahon, W.M.; Fujinami, R.S. Autistic disorder and viral infections. J. Neurovirol. 2005, 11, 1–10. [Google Scholar] [CrossRef]
- Diav-Citrin, O.; Ornoy, A. Adverse environment and prevention of early pregnancy disorders. Early Pregnancy 2000, 4, 5–18. [Google Scholar]
- Mahic, M.; Mjaaland, S.; Bovelstad, H.M.; Gunnes, N.; Susser, E.; Bresnahan, M.; Oyen, A.S.; Levin, B.; Che, X.; Hirtz, D.; et al. Maternal Immunoreactivity to Herpes Simplex Virus 2 and Risk of Autism Spectrum Disorder in Male Offspring. mSphere 2017, 2. [Google Scholar] [CrossRef]
- Gillberg, C.; Mary, C. The Biology of the Autistic Syndromes; Cambridge University Press: Cambridge, UK, 2000. [Google Scholar]
- Yamashita, Y.; Fujimoto, C.; Nakajima, E.; Isagai, T.; Matsuishi, T. Possible association between congenital cytomegalovirus infection and autistic disorder. J. Autism. Dev. Disord. 2003, 33, 455–459. [Google Scholar] [CrossRef]
- Sweeten, T.L.; Posey, D.J.; McDougle, C.J. Brief report: Autistic disorder in three children with cytomegalovirus infection. J. Autism. Dev. Disord. 2004, 34, 583–586. [Google Scholar] [CrossRef]
- Kawatani, M.; Nakai, A.; Okuno, T.; Kobata, R.; Moriuchi, M.; Moriuchi, H.; Tsukahara, H.; Mayumi, M. Detection of cytomegalovirus in preserved umbilical cord from a boy with autistic disorder. Pediatr. Int. 2010, 52, 304–307. [Google Scholar] [CrossRef] [PubMed]
- Malm, G.; Engman, M.L. Congenital cytomegalovirus infections. Semin. Fetal Neonatal Med. 2007, 12, 154–159. [Google Scholar] [CrossRef]
- Mouridsen, S.E.; Rich, B.; Isager, T.; Nedergaard, N.J. Autoimmune diseases in parents of children with infantile autism: A case-control study. Dev. Med. Child Neurol. 2007, 49, 429–432. [Google Scholar] [CrossRef] [PubMed]
- Keil, A.; Daniels, J.L.; Forssen, U.; Hultman, C.; Cnattingius, S.; Soderberg, K.C.; Feychting, M.; Sparen, P. Parental autoimmune diseases associated with autism spectrum disorders in offspring. Epidemiology 2010, 21, 805–808. [Google Scholar] [CrossRef]
- Smith, S.E.; Li, J.; Garbett, K.; Mirnics, K.; Patterson, P.H. Maternal immune activation alters fetal brain development through interleukin-6. J. Neurosci. Off. 2007, 27, 10695–10702. [Google Scholar] [CrossRef]
- Short, S.J.; Lubach, G.R.; Karasin, A.I.; Olsen, C.W.; Styner, M.; Knickmeyer, R.C.; Gilmore, J.H.; Coe, C.L. Maternal influenza infection during pregnancy impacts postnatal brain development in the rhesus monkey. Biol. Psychiatry 2010, 67, 965–973. [Google Scholar] [CrossRef]
- Gillberg, C. Maternal age and infantile autism. J. Autism. Dev. Disord. 1980, 10, 293–297. [Google Scholar] [CrossRef]
- Glasson, E.J.; Bower, C.; Petterson, B.; de Klerk, N.; Chaney, G.; Hallmayer, J.F. Perinatal factors and the development of autism: A population study. Arch. Gen. Psychiatry 2004, 61, 618–627. [Google Scholar] [CrossRef]
- Maimburg, R.D.; Vaeth, M. Perinatal risk factors and infantile autism. Acta Psychiatr. Scand. 2006, 114, 257–264. [Google Scholar] [CrossRef]
- Reichenberg, A.; Gross, R.; Weiser, M.; Bresnahan, M.; Silverman, J.; Harlap, S.; Rabinowitz, J.; Shulman, C.; Malaspina, D.; Lubin, G.; et al. Advancing paternal age and autism. Arch. Gen. Psychiatry 2006, 63, 1026–1032. [Google Scholar] [CrossRef]
- Croen, L.A.; Najjar, D.V.; Fireman, B.; Grether, J.K. Maternal and paternal age and risk of autism spectrum disorders. Arch. Pediatr. Adolesc. Med. 2007, 161, 334–340. [Google Scholar] [CrossRef] [PubMed]
- Durkin, M.S.; Maenner, M.J.; Newschaffer, C.J.; Lee, L.C.; Cunniff, C.M.; Daniels, J.L.; Kirby, R.S.; Leavitt, L.; Miller, L.; Zahorodny, W.; et al. Advanced parental age and the risk of autism spectrum disorder. Am. J. Epidemiol. 2008, 168, 1268–1276. [Google Scholar] [CrossRef] [PubMed]
- Reichenberg, A.; Gross, R.; Sandin, S.; Susser, E.S. Advancing paternal and maternal age are both important for autism risk. Am. J. Public Health 2010, 100, 772–773. [Google Scholar] [CrossRef]
- Sandin, S.; Hultman, C.M.; Kolevzon, A.; Gross, R.; MacCabe, J.H.; Reichenberg, A. Advancing maternal age is associated with increasing risk for autism: A review and meta-analysis. J. Am. Acad. Child Adolesc. Psychiatry 2012, 51, 477–486. [Google Scholar] [CrossRef]
- De la Rochebrochard, E.; Thonneau, P. Paternal age and maternal age are risk factors for miscarriage; results of a multicentre European study. Hum. Reprod. 2002, 17, 1649–1656. [Google Scholar] [CrossRef]
- De La Rochebrochard, E.; Thonneau, P. Paternal age > or = 40 years: An important risk factor for infertility. Am. J. Obstet. Gynecol. 2003, 189, 901–905. [Google Scholar] [CrossRef]
- Kleinhaus, K.; Perrin, M.; Friedlander, Y.; Paltiel, O.; Malaspina, D.; Harlap, S. Paternal age and spontaneous abortion. Obstet. Gynecol. 2006, 108, 369–377. [Google Scholar] [CrossRef]
- Jaleel, R.; Khan, A. Paternal factors in spontaneous first trimester miscarriage. Pak. J. Med. Sci. 2013, 29, 748–752. [Google Scholar] [CrossRef]
- Nybo Andersen, A.M.; Hansen, K.D.; Andersen, P.K.; Davey Smith, G. Advanced paternal age and risk of fetal death: A cohort study. Am. J. Epidemiol. 2004, 160, 1214–1222. [Google Scholar] [CrossRef]
- Slama, R.; Bouyer, J.; Windham, G.; Fenster, L.; Werwatz, A.; Swan, S.H. Influence of paternal age on the risk of spontaneous abortion. Am. J. Epidemiol. 2005, 161, 816–823. [Google Scholar] [CrossRef]
- Yip, B.H.; Pawitan, Y.; Czene, K. Parental age and risk of childhood cancers: A population-based cohort study from Sweden. Int. J. Epidemiol. 2006, 35, 1495–1503. [Google Scholar] [CrossRef] [PubMed]
- Urhoj, S.K.; Raaschou-Nielsen, O.; Hansen, A.V.; Mortensen, L.H.; Andersen, P.K.; Nybo Andersen, A.M. Advanced paternal age and childhood cancer in offspring: A nationwide register-based cohort study. Int. J. Cancer 2017, 140, 2461–2472. [Google Scholar] [CrossRef] [PubMed]
- Malaspina, D.; Harlap, S.; Fennig, S.; Heiman, D.; Nahon, D.; Feldman, D.; Susser, E.S. Advancing paternal age and the risk of schizophrenia. Arch. Gen. Psychiatry 2001, 58, 361–367. [Google Scholar] [CrossRef] [PubMed]
- Brown, A.S.; Schaefer, C.A.; Wyatt, R.J.; Begg, M.D.; Goetz, R.; Bresnahan, M.A.; Harkavy-Friedman, J.; Gorman, J.M.; Malaspina, D.; Susser, E.S. Paternal age and risk of schizophrenia in adult offspring. Am. J. Psychiatry 2002, 159, 1528–1533. [Google Scholar] [CrossRef]
- Zammit, S.; Allebeck, P.; Dalman, C.; Lundberg, I.; Hemmingson, T.; Owen, M.J.; Lewis, G. Paternal age and risk for schizophrenia. Br. J. Psychiatry J. Ment. Sci. 2003, 183, 405–408. [Google Scholar] [CrossRef]
- Sipos, A.; Rasmussen, F.; Harrison, G.; Tynelius, P.; Lewis, G.; Leon, D.A.; Gunnell, D. Paternal age and schizophrenia: A population based cohort study. BMJ 2004, 329, 1070. [Google Scholar] [CrossRef]
- Penrose, L.S. Parental age and mutation. Lancet 1955, 269, 312–313. [Google Scholar] [CrossRef]
- Jones, K.L.; Smith, D.W.; Harvey, M.A.; Hall, B.D.; Quan, L. Older paternal age and fresh gene mutation: Data on additional disorders. J. Pediatr. 1975, 86, 84–88. [Google Scholar] [CrossRef]
- Keilin, D.; Mann, T. Carbonic anhydrase. Purification and nature of the enzyme. Biochem. J. 1940, 34, 1163–1176. [Google Scholar] [CrossRef]
- Vallee, B.L.; Falchuk, K.H. The biochemical basis of zinc physiology. Physiol. Rev. 1993, 73, 79–118. [Google Scholar] [CrossRef]
- Vallee, B.L.; Galdes, A. The metallobiochemistry of zinc enzymes. Adv. Enzymol. Relat. Areas Mol. Biol. 1984, 56, 283–430. [Google Scholar] [CrossRef] [PubMed]
- Choi, D.W.; Koh, J.Y. Zinc and brain injury. Annu. Rev. Neurosci. 1998, 21, 347–375. [Google Scholar] [CrossRef] [PubMed]
- Prasad, A.S. Zinc: An overview. Nutrition 1995, 11, 93–99. [Google Scholar] [PubMed]
- Vallee, B.L.; Coleman, J.E.; Auld, D.S. Zinc fingers, zinc clusters, and zinc twists in DNA-binding protein domains. Proc. Natl. Acad. Sci. USA 1991, 88, 999–1003. [Google Scholar] [CrossRef]
- Simmer, K.; Thompson, R.P. Zinc in the fetus and newborn. Acta Paediatr. Scand. Suppl. 1985, 319, 158–163. [Google Scholar] [CrossRef]
- Fabris, N.; Mocchegiani, E. Zinc, human diseases and aging. Aging 1995, 7, 77–93. [Google Scholar] [CrossRef]
- Bhatnagar, S.; Taneja, S. Zinc and cognitive development. Br. J. Nutr. 2001, 85, S139–S145. [Google Scholar] [CrossRef]
- Black, M.M. The evidence linking zinc deficiency with children’s cognitive and motor functioning. J. Nutr. 2003, 133, 1473S–1476S. [Google Scholar] [CrossRef]
- Black, M.M. Micronutrient deficiencies and cognitive functioning. J. Nutr. 2003, 133, 3927S–3931S. [Google Scholar] [CrossRef]
- Clarkson, T.W. Metal toxicity in the central nervous system. Environ. Health Perspect. 1987, 75, 59–64. [Google Scholar] [CrossRef]
- Mantyh, P.W.; Ghilardi, J.R.; Rogers, S.; DeMaster, E.; Allen, C.J.; Stimson, E.R.; Maggio, J.E. Aluminum, iron, and zinc ions promote aggregation of physiological concentrations of beta-amyloid peptide. J. Neurochem. 1993, 61, 1171–1174. [Google Scholar] [CrossRef] [PubMed]
- Gaeta, A.; Hider, R.C. The crucial role of metal ions in neurodegeneration: The basis for a promising therapeutic strategy. Br. J. Pharmacol. 2005, 146, 1041–1059. [Google Scholar] [CrossRef] [PubMed]
- Zatta, P.; Drago, D.; Bolognin, S.; Sensi, S.L. Alzheimer’s disease, metal ions and metal homeostatic therapy. Trends Pharmacol. Sci. 2009, 30, 346–355. [Google Scholar] [CrossRef]
- Karri, V.; Schuhmacher, M.; Kumar, V. Heavy metals (Pb, Cd, As and MeHg) as risk factors for cognitive dysfunction: A general review of metal mixture mechanism in brain. Environ. Toxicol. Pharmacol. 2016, 48, 203–213. [Google Scholar] [CrossRef]
- Cristovao, J.S.; Santos, R.; Gomes, C.M. Metals and Neuronal Metal Binding Proteins Implicated in Alzheimer’s Disease. Oxidative Med. Cell. Longev. 2016, 2016, 9812178. [Google Scholar] [CrossRef]
- Dexter, D.T.; Wells, F.R.; Lees, A.J.; Agid, F.; Agid, Y.; Jenner, P.; Marsden, C.D. Increased nigral iron content and alterations in other metal ions occurring in brain in Parkinson’s disease. J. Neurochem. 1989, 52, 1830–1836. [Google Scholar] [CrossRef]
- Wenstrup, D.; Ehmann, W.D.; Markesbery, W.R. Trace element imbalances in isolated subcellular fractions of Alzheimer’s disease brains. Brain Res. 1990, 533, 125–131. [Google Scholar] [CrossRef]
- Constantinidis, J. The hypothesis of zinc deficiency in the pathogenesis of neurofibrillary tangles. Med. Hypotheses 1991, 35, 319–323. [Google Scholar] [CrossRef]
- Jenner, P.; Dexter, D.T.; Sian, J.; Schapira, A.H.; Marsden, C.D. Oxidative stress as a cause of nigral cell death in Parkinson’s disease and incidental Lewy body disease. The Royal Kings and Queens Parkinson’s Disease Research Group. Ann. Neurol. 1992, 32, S82–S87. [Google Scholar] [CrossRef]
- Corrigan, F.M.; Reynolds, G.P.; Ward, N.I. Hippocampal tin, aluminum and zinc in Alzheimer’s disease. Biometals Int. J. Role Met. Ions Biol. Biochem. Med. 1993, 6, 149–154. [Google Scholar] [CrossRef]
- Andrasi, E.; Farkas, E.; Scheibler, H.; Reffy, A.; Bezur, L. Al, Zn, Cu, Mn and Fe levels in brain in Alzheimer’s disease. Arch. Gerontol. Geriatr. 1995, 21, 89–97. [Google Scholar] [CrossRef]
- Deibel, M.A.; Ehmann, W.D.; Markesbery, W.R. Copper, iron, and zinc imbalances in severely degenerated brain regions in Alzheimer’s disease: Possible relation to oxidative stress. J. Neurol. Sci. 1996, 143, 137–142. [Google Scholar] [CrossRef]
- Cuajungco, M.P.; Lees, G.J. Zinc metabolism in the brain: Relevance to human neurodegenerative disorders. Neurobiol. Dis. 1997, 4, 137–169. [Google Scholar] [CrossRef] [PubMed]
- Arora, M.; Reichenberg, A.; Willfors, C.; Austin, C.; Gennings, C.; Berggren, S.; Lichtenstein, P.; Anckarsater, H.; Tammimies, K.; Bolte, S. Fetal and postnatal metal dysregulation in autism. Nat. Commun. 2017, 8, 15493. [Google Scholar] [CrossRef]
- Bush, A.I.; Multhaup, G.; Moir, R.D.; Williamson, T.G.; Small, D.H.; Rumble, B.; Pollwein, P.; Beyreuther, K.; Masters, C.L. A novel zinc(II) binding site modulates the function of the beta A4 amyloid protein precursor of Alzheimer’s disease. J. Biol. Chem. 1993, 268, 16109–16112. [Google Scholar]
- Bush, A.I.; Pettingell, W.H.; Multhaup, G.; d Paradis, M.; Vonsattel, J.P.; Gusella, J.F.; Beyreuther, K.; Masters, C.L.; Tanzi, R.E. Rapid induction of Alzheimer A beta amyloid formation by zinc. Science 1994, 265, 1464–1467. [Google Scholar] [CrossRef] [PubMed]
- Clements, A.; Allsop, D.; Walsh, D.M.; Williams, C.H. Aggregation and metal-binding properties of mutant forms of the amyloid A beta peptide of Alzheimer’s disease. J. Neurochem. 1996, 66, 740–747. [Google Scholar] [CrossRef]
- Esler, W.P.; Stimson, E.R.; Ghilardi, J.R.; Vinters, H.V.; Lee, J.P.; Mantyh, P.W.; Maggio, J.E. In vitro growth of Alzheimer’s disease beta-amyloid plaques displays first-order kinetics. Biochemistry 1996, 35, 749–757. [Google Scholar] [CrossRef]
- Strozyk, D.; Launer, L.J.; Adlard, P.A.; Cherny, R.A.; Tsatsanis, A.; Volitakis, I.; Blennow, K.; Petrovitch, H.; White, L.R.; Bush, A.I. Zinc and copper modulate Alzheimer Abeta levels in human cerebrospinal fluid. Neurobiol. Aging 2009, 30, 1069–1077. [Google Scholar] [CrossRef]
- Naisbitt, S.; Kim, E.; Tu, J.C.; Xiao, B.; Sala, C.; Valtschanoff, J.; Weinberg, R.J.; Worley, P.F.; Sheng, M. Shank, a novel family of postsynaptic density proteins that binds to the NMDA receptor/PSD-95/GKAP complex and cortactin. Neuron 1999, 23, 569–582. [Google Scholar] [CrossRef]
- Boeckers, T.M.; Winter, C.; Smalla, K.H.; Kreutz, M.R.; Bockmann, J.; Seidenbecher, C.; Garner, C.C.; Gundelfinger, E.D. Proline-rich synapse-associated proteins ProSAP1 and ProSAP2 interact with synaptic proteins of the SAPAP/GKAP family. Biochem. Biophys. Res. Commun. 1999, 264, 247–252. [Google Scholar] [CrossRef] [PubMed]
- Grabrucker, S.; Jannetti, L.; Eckert, M.; Gaub, S.; Chhabra, R.; Pfaender, S.; Mangus, K.; Reddy, P.P.; Rankovic, V.; Schmeisser, M.J.; et al. Zinc deficiency dysregulates the synaptic ProSAP/Shank scaffold and might contribute to autism spectrum disorders. Brain 2014, 137, 137–152. [Google Scholar] [CrossRef] [PubMed]
- Durand, C.M.; Betancur, C.; Boeckers, T.M.; Bockmann, J.; Chaste, P.; Fauchereau, F.; Nygren, G.; Rastam, M.; Gillberg, I.C.; Anckarsater, H.; et al. Mutations in the gene encoding the synaptic scaffolding protein SHANK3 are associated with autism spectrum disorders. Nat. Genet. 2007, 39, 25–27. [Google Scholar] [CrossRef] [PubMed]
- Moessner, R.; Marshall, C.R.; Sutcliffe, J.S.; Skaug, J.; Pinto, D.; Vincent, J.; Zwaigenbaum, L.; Fernandez, B.; Roberts, W.; Szatmari, P.; et al. Contribution of SHANK3 mutations to autism spectrum disorder. Am. J. Hum. Genet. 2007, 81, 1289–1297. [Google Scholar] [CrossRef]
- Gauthier, J.; Spiegelman, D.; Piton, A.; Lafreniere, R.G.; Laurent, S.; St-Onge, J.; Lapointe, L.; Hamdan, F.F.; Cossette, P.; Mottron, L. Novel de novo SHANK3 mutation in autistic patients. Am. J. Med. Genet. Part B Neuropsychiatr. Genet. 2009, 150, 421–424. [Google Scholar] [CrossRef]
- Gundelfinger, E.D.; Boeckers, T.M.; Baron, M.K.; Bowie, J.U. A role for zinc in postsynaptic density asSAMbly and plasticity? Trends Biochem. Sci. 2006, 31, 366–373. [Google Scholar] [CrossRef]
- Grabrucker, A.M. A role for synaptic zinc in ProSAP/Shank PSD scaffold malformation in autism spectrum disorders. Dev. Neurobiol. 2014, 74, 136–146. [Google Scholar] [CrossRef]
- Pfaender, S.; Sauer, A.K.; Hagmeyer, S.; Mangus, K.; Linta, L.; Liebau, S.; Bockmann, J.; Huguet, G.; Bourgeron, T.; Boeckers, T.M.; et al. Zinc deficiency and low enterocyte zinc transporter expression in human patients with autism related mutations in SHANK3. Sci. Rep. 2017, 7, 45190. [Google Scholar] [CrossRef]
- Yasuda, H.; Yoshida, K.; Yasuda, Y.; Tsutsui, T. Infantile zinc deficiency: Association with autism spectrum disorders. Sci. Rep. 2011, 1, 129. [Google Scholar] [CrossRef]
- Schoen, M.; Asoglu, H.; Bauer, H.F.; Muller, H.P.; Abaei, A.; Sauer, A.K.; Zhang, R.; Song, T.J.; Bockmann, J.; Kassubek, J.; et al. Shank3 Transgenic and Prenatal Zinc-Deficient Autism Mouse Models Show Convergent and Individual Alterations of Brain Structures in MRI. Front. Neural. Circuits 2019, 13, 6. [Google Scholar] [CrossRef]
- Sweetman, D.U.; O’Donnell, S.M.; Lalor, A.; Grant, T.; Greaney, H. Zinc and vitamin A deficiency in a cohort of children with autism spectrum disorder. Child Care Health Dev. 2019, 45, 380–386. [Google Scholar] [CrossRef]
- Fluegge, K.J. Zinc and Copper Metabolism and Risk of Autism: A reply to Sayehmiri et al. Iran. J. Child Neurol. 2017, 11, 66. [Google Scholar]
- World Health Oganization. The ICD-10 Classification of Mental and Behavioral Disorders: Clinical Descriptions and Diagnostic Guidelines; World Health Oganization: Geneva, Switzerland, 1992. [Google Scholar]
- Association, A.P. Diagnostic Criteria from dsM-iV-tr; American Psychiatric Publishing, Inc.: Washington, DC, USA, 2000. [Google Scholar]
- Amir, R.E.; Van den Veyver, I.B.; Wan, M.; Tran, C.Q.; Francke, U.; Zoghbi, H.Y. Rett syndrome is caused by mutations in X-linked MECP2, encoding methyl-CpG-binding protein 2. Nat. Genet. 1999, 23, 185–188. [Google Scholar] [CrossRef] [PubMed]
- Piven, J.; Palmer, P.; Jacobi, D.; Childress, D.; Arndt, S. Broader autism phenotype: Evidence from a family history study of multiple-incidence autism families. Am. J. Psychiatry 1997, 154, 185–190. [Google Scholar] [PubMed]
- Muhle, R.; Trentacoste, S.V.; Rapin, I. The genetics of autism. Pediatrics 2004, 113, e472–e486. [Google Scholar] [CrossRef]
- Neale, B.M.; Kou, Y.; Liu, L.; Ma’Ayan, A.; Samocha, K.E.; Sabo, A.; Lin, C.-F.; Stevens, C.; Wang, L.-S.; Makarov, V. Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature 2012, 485, 242–245. [Google Scholar] [CrossRef]
- Iossifov, I.; Ronemus, M.; Levy, D.; Wang, Z.; Hakker, I.; Rosenbaum, J.; Yamrom, B.; Lee, Y.-h.; Narzisi, G.; Leotta, A. De novo gene disruptions in children on the autistic spectrum. Neuron 2012, 74, 285–299. [Google Scholar] [CrossRef]
- Iossifov, I.; O’roak, B.J.; Sanders, S.J.; Ronemus, M.; Krumm, N.; Levy, D.; Stessman, H.A.; Witherspoon, K.T.; Vives, L.; Patterson, K.E. The contribution of de novo coding mutations to autism spectrum disorder. Nature 2014, 515, 216–221. [Google Scholar] [CrossRef]
- Vincent, J.B.; Herbrick, J.A.; Gurling, H.M.; Bolton, P.F.; Roberts, W.; Scherer, S.W. Identification of a novel gene on chromosome 7q31 that is interrupted by a translocation breakpoint in an autistic individual. Am. J. Hum. Genet. 2000, 67, 510–514. [Google Scholar] [CrossRef]
- Lopreiato, J.O.; Wulfsberg, E.A. A complex chromosome rearrangement in a boy with autism. J. Dev. Behav. Pediatrics JDBP 1992, 13, 281–283. [Google Scholar] [CrossRef]
- Ashley-Koch, A.; Wolpert, C.M.; Menold, M.M.; Zaeem, L.; Basu, S.; Donnelly, S.L.; Ravan, S.A.; Powell, C.M.; Qumsiyeh, M.B.; Aylsworth, A. Genetic studies of autistic disorder and chromosome 7. Genomics 1999, 61, 227–236. [Google Scholar] [CrossRef] [PubMed]
- Warburton, P.; Baird, G.; Chen, W.; Morris, K.; Jacobs, B.W.; Hodgson, S.; Docherty, Z. Support for linkage of autism and specific language impairment to 7q3 from two chromosome rearrangements involving band 7q31. Am. J. Med Genet. 2000, 96, 228–234. [Google Scholar] [CrossRef]
- A full genome screen for autism with evidence for linkage to a region on chromosome 7q. International Molecular Genetic Study of Autism Consortium. Hum. Mol. Genet. 1998, 7, 571–578. [CrossRef]
- Hurst, J.A.; Baraitser, M.; Auger, E.; Graham, F.; Norell, S. An extended family with a dominantly inherited speech disorder. Dev. Med. Child Neurol. 1990, 32, 352–355. [Google Scholar] [CrossRef]
- Hutcheson, H.B.; Bradford, Y.; Folstein, S.E.; Gardiner, M.B.; Santangelo, S.L.; Sutcliffe, J.S.; Haines, J.L. Defining the autism minimum candidate gene region on chromosome 7. Am. J. Med Genet. Part B 2003, 117B, 90–96. [Google Scholar] [CrossRef] [PubMed]
- Shao, Y.; Wolpert, C.M.; Raiford, K.L.; Menold, M.M.; Donnelly, S.L.; Ravan, S.A.; Bass, M.P.; McClain, C.; von Wendt, L.; Vance, J.M.; et al. Genomic screen and follow-up analysis for autistic disorder. Am. J. Med. Genet. 2002, 114, 99–105. [Google Scholar] [CrossRef] [PubMed]
- Philippe, A.; Martinez, M.; Guilloud-Bataille, M.; Gillberg, C.; Råstam, M.; Sponheim, E.; Coleman, M.; Zappella, M.; Aschauer, H.; Van Maldergem, L. Genome-wide scan for autism susceptibility genes. Hum. Mol. Genet. 1999, 8, 805–812. [Google Scholar] [CrossRef]
- Barrett, S.; Beck, J.C.; Bernier, R.; Bisson, E.; Braun, T.A.; Casavant, T.L.; Childress, D.; Folstein, S.E.; Garcia, M.; Gardiner, M.B. An autosomal genomic screen for autism. Collaborative linkage study of autism. Am. J. Med Genet. 1999, 88, 609–615. [Google Scholar] [PubMed]
- Lai, C.S.; Fisher, S.E.; Hurst, J.A.; Vargha-Khadem, F.; Monaco, A.P. A forkhead-domain gene is mutated in a severe speech and language disorder. Nature 2001, 413, 519–523. [Google Scholar] [CrossRef]
- Lai, C.S.; Gerrelli, D.; Monaco, A.P.; Fisher, S.E.; Copp, A.J. FOXP2 expression during brain development coincides with adult sites of pathology in a severe speech and language disorder. Brain 2003, 126, 2455–2462. [Google Scholar] [CrossRef]
- Shu, W.; Yang, H.; Zhang, L.; Lu, M.M.; Morrisey, E.E. Characterization of a new subfamily of winged-helix/forkhead (Fox) genes that are expressed in the lung and act as transcriptional repressors. J. Biol. Chem. 2001, 276, 27488–27497. [Google Scholar] [CrossRef] [PubMed]
- Fisher, S.E.; Vargha-Khadem, F.; Watkins, K.E.; Monaco, A.P.; Pembrey, M.E. Localisation of a gene implicated in a severe speech and language disorder. Nat. Genet. 1998, 18, 168–170. [Google Scholar] [CrossRef]
- Newbury, D.F.; Bonora, E.; Lamb, J.A.; Fisher, S.E.; Lai, C.S.; Baird, G.; Jannoun, L.; Slonims, V.; Stott, C.M.; Merricks, M.J.; et al. FOXP2 is not a major susceptibility gene for autism or specific language impairment. Am. J. Hum. Genet. 2002, 70, 1318–1327. [Google Scholar] [CrossRef] [PubMed]
- Tolosa, A.; Sanjuán, J.; Dagnall, A.M.; Moltó, M.D.; Herrero, N.; de Frutos, R.J. FOXP2 gene and language impairment in schizophrenia: Association and epigenetic studies. BMC Med. Genet. 2010, 11, 114. [Google Scholar] [CrossRef] [PubMed]
- Toma, C.; Hervas, A.; Torrico, B.; Balmaña, N.; Salgado, M.; Maristany, M.; Vilella, E.; Martínez-Leal, R.; Planelles, M.I.; Cusco, I.; et al. Analysis of two language-related genes in autism. Psychiatr. Genet. 2013, 23, 82–85. [Google Scholar] [CrossRef] [PubMed]
- Liyanage, V.R.B. Role of RNA methylation and non-coding RNAs in pathobiology of autism spectrum disorders. Biomed. Sci. 2016, 2, 24. [Google Scholar]
- Vincent, J.B.; Petek, E.; Thevarkunnel, S.; Kolozsvari, D.; Cheung, J.; Patel, M.; Scherer, S.W. The RAY1/ST7 tumor-suppressor locus on chromosome 7q31 represents a complex multi-transcript system. Genomics 2002, 80, 283–294. [Google Scholar] [CrossRef]
- Maestrini, E.; Pagnamenta, A.T.; Lamb, J.A.; Bacchelli, E.; Sykes, N.H.; Sousa, I.; Toma, C.; Barnby, G.; Butler, H.; Winchester, L.; et al. High-density SNP association study and copy number variation analysis of the AUTS1 and AUTS5 loci implicate the IMMP2L-DOCK4 gene region in autism susceptibility. Mol. Psychiatry 2010, 15, 954–968. [Google Scholar] [CrossRef]
- Robertson, M.M. Annotation: Gilles de la Tourette syndrome—An update. J. Child Psychol. Psychiatry Allied Discip. 1994, 35, 597–611. [Google Scholar] [CrossRef]
- Kreilaus, F.; Chesworth, R.; Eapen, V.; Clarke, R.; Karl, T. First behavioural assessment of a novel Immp2l knockdown mouse model with relevance for Gilles de la Tourette syndrome and Autism spectrum disorder. Behav. Brain Res. 2019, 374, 112057. [Google Scholar] [CrossRef]
- Petek, E.; Schwarzbraun, T.; Noor, A.; Patel, M.; Nakabayashi, K.; Choufani, S.; Windpassinger, C.; Stamenkovic, M.; Robertson, M.M.; Aschauer, H.N.; et al. Molecular and genomic studies of IMMP2L and mutation screening in autism and Tourette syndrome. Mol. Genet. Genom. 2007, 277, 71–81. [Google Scholar] [CrossRef] [PubMed]
- Liang, S.; Wang, X.L.; Zou, M.Y.; Wang, H.; Zhou, X.; Sun, C.H.; Xia, W.; Wu, L.J.; Fujisawa, T.X.; Tomoda, A. Family-based association study of ZNF533, DOCK4 and IMMP2L gene polymorphisms linked to autism in a northeastern Chinese Han population. J. Zhejiang Univ. Sci. B 2014, 15, 264–271. [Google Scholar] [CrossRef] [PubMed]
- Ogawa, M.; Miyata, T.; Nakajimat, K.; Yagyu, K.; Seike, M.; Ikenaka, K.; Yamamoto, H.; Mikoshibat, K. The reeler gene-associated antigen on Cajal-Retzius neurons is a crucial molecule for laminar organization of cortical neurons. Neuron 1995, 14, 899–912. [Google Scholar] [CrossRef]
- D’Arcangelo, G.; Miao, G.G.; Chen, S.-C.; Scares, H.D.; Morgan, J.I.; Curran, T. A protein related to extracellular matrix proteins deleted in the mouse mutant reeler. Nature 1995, 374, 719–723. [Google Scholar] [CrossRef]
- Yip, J.W.; Yip, Y.P.L.; Nakajima, K.; Capriotti, C. Reelin controls position of autonomic neurons in the spinal cord. Proc. Natl. Acad. Sci. USA 2000, 97, 8612–8616. [Google Scholar] [CrossRef]
- Dulabon, L.; Olson, E.C.; Taglienti, M.G.; Eisenhuth, S.; McGrath, B.; Walsh, C.A.; Kreidberg, J.A.; Anton, E. Reelin binds α3β1 integrin and inhibits neuronal migration. Neuron 2000, 27, 33–44. [Google Scholar] [CrossRef]
- Persico, A.; D’agruma, L.; Maiorano, N.; Totaro, A.; Militerni, R.; Bravaccio, C.; Wassink, T.; Schneider, C.; Melmed, R.; Trillo, S. Reelin gene alleles and haplotypes as a factor predisposing to autistic disorder. Mol. Psychiatry 2001, 6, 150–159. [Google Scholar] [CrossRef]
- Zhang, H.; Liu, X.; Zhang, C.; Mundo, E.; Macciardi, F.; Grayson, D.; Guidotti, A.; Holden, J. Reelin gene alleles and susceptibility to autism spectrum disorders. Mol. Psychiatry 2002, 7, 1012–1017. [Google Scholar] [CrossRef][Green Version]
- Skaar, D.; Shao, Y.; Haines, J.; Stenger, J.; Jaworski, J.; Martin, E.R.; DeLong, G.; Moore, J.; McCauley, J.L.; Sutcliffe, J. Analysis of the RELN gene as a genetic risk factor for autism. Mol. Psychiatry 2005, 10, 563–571. [Google Scholar] [CrossRef]
- Fatemi, S.H.; Snow, A.V.; Stary, J.M.; Araghi-Niknam, M.; Reutiman, T.J.; Lee, S.; Brooks, A.I.; Pearce, D.A. Reelin signaling is impaired in autism. Biol. Psychiatry 2005, 57, 777–787. [Google Scholar] [CrossRef]
- Lammert, D.B.; Howell, B.W. RELN mutations in autism spectrum disorder. Front. Cell. Neurosci. 2016, 10, 84. [Google Scholar] [CrossRef] [PubMed]
- Seripa, D.; Matera, M.G.; Franceschi, M.; Daniele, A.; Bizzarro, A.; Rinaldi, M.; Panza, F.; Fazio, V.M.; Gravina, C.; D’Onofrio, G.; et al. The RELN locus in Alzheimer’s disease. J. Alzheimer Dis. JAD 2008, 14, 335–344. [Google Scholar] [CrossRef] [PubMed]
- Kramer, P.L.; Xu, H.; Woltjer, R.L.; Westaway, S.K.; Clark, D.; Erten-Lyons, D.; Kaye, J.A.; Welsh-Bohmer, K.A.; Troncoso, J.C.; Markesbery, W.R.; et al. Alzheimer disease pathology in cognitively healthy elderly: A genome-wide study. Neurobiol. Aging 2011, 32, 2113–2122. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Chen, P.L.; McGrath, J.; Wolyniec, P.; Fallin, D.; Nestadt, G.; Liang, K.Y.; Pulver, A.; Valle, D.; Avramopoulos, D. Replication of an association of a common variant in the Reelin gene (RELN) with schizophrenia in Ashkenazi Jewish women. Psychiatr. Genet. 2010, 20, 184–186. [Google Scholar] [CrossRef] [PubMed]
- Ovadia, G.; Shifman, S. The genetic variation of RELN expression in schizophrenia and bipolar disorder. PLoS ONE 2011, 6, e19955. [Google Scholar] [CrossRef]
- Hong, S.E.; Shugart, Y.Y.; Huang, D.T.; Shahwan, S.A.; Grant, P.E.; Hourihane, J.O.; Martin, N.D.; Walsh, C.A. Autosomal recessive lissencephaly with cerebellar hypoplasia is associated with human RELN mutations. Nat. Genet. 2000, 26, 93–96. [Google Scholar] [CrossRef]
- Chang, B.S.; Duzcan, F.; Kim, S.; Cinbis, M.; Aggarwal, A.; Apse, K.A.; Ozdel, O.; Atmaca, M.; Zencir, S.; Bagci, H.; et al. The role of RELN in lissencephaly and neuropsychiatric disease. Am. J. Med Genet. Part B 2007, 144B, 58–63. [Google Scholar] [CrossRef]
- Fatemi, S.H.; Stary, J.M.; Egan, E.A. Reduced blood levels of reelin as a vulnerability factor in pathophysiology of autistic disorder. Cell. Mol. Neurobiol. 2002, 22, 139–152. [Google Scholar] [CrossRef]
- Wang, Z.; Hong, Y.; Zou, L.; Zhong, R.; Zhu, B.; Shen, N.; Chen, W.; Lou, J.; Ke, J.; Zhang, T.; et al. Reelin gene variants and risk of autism spectrum disorders: An integrated meta-analysis. Am. J. Med Genet. Part B 2014, 165B, 192–200. [Google Scholar] [CrossRef]
- Abrahams, B.S.; Arking, D.E.; Campbell, D.B.; Mefford, H.C.; Morrow, E.M.; Weiss, L.A.; Menashe, I.; Wadkins, T.; Banerjee-Basu, S.; Packer, A. SFARI Gene 2.0: A community-driven knowledgebase for the autism spectrum disorders (ASDs). Mol. Autism 2013, 4, 36. [Google Scholar] [CrossRef]
- Prontera, P.; Serino, D.; Caldini, B.; Scarponi, L.; Merla, G.; Testa, G.; Muti, M.; Napolioni, V.; Mazzotta, G.; Piccirilli, M.; et al. Brief Report: Functional MRI of a Patient with 7q11.23 Duplication Syndrome and Autism Spectrum Disorder. J. Autism Dev. Disord. 2014, 44, 2608–2613. [Google Scholar] [CrossRef] [PubMed]
- Lammert, D.; Middleton, F.A.; Pan, J.Q.; Olson, E.; Howell, B. The de novo autism spectrum disorder RELN R2290C mutation reduces Reelin secretion and increases protein disulfide isomerase expression. J. Neurochem. 2017, 142, 89–102. [Google Scholar] [CrossRef] [PubMed]
- Lintas, C.; Sacco, R.; Persico, A.M. Differential methylation at the RELN gene promoter in temporal cortex from autistic and typically developing post-puberal subjects. J. Neurodev. Disord. 2016, 8, 18. [Google Scholar] [CrossRef] [PubMed]
- Marshall, C.R.; Noor, A.; Vincent, J.B.; Lionel, A.C.; Feuk, L.; Skaug, J.; Shago, M.; Moessner, R.; Pinto, D.; Ren, Y.; et al. Structural variation of chromosomes in autism spectrum disorder. Am. J. Hum. Genet. 2008, 82, 477–488. [Google Scholar] [CrossRef]
- Guo, H.; Peng, Y.; Hu, Z.; Li, Y.; Xun, G.; Ou, J.; Sun, L.; Xiong, Z.; Liu, Y.; Wang, T.; et al. Genome-wide copy number variation analysis in a Chinese autism spectrum disorder cohort. Sci. Rep. 2017, 7, 44155. [Google Scholar] [CrossRef]
- Willemsen, M.H.; Fernandez, B.A.; Bacino, C.A.; Gerkes, E.; de Brouwer, A.P.; Pfundt, R.; Sikkema-Raddatz, B.; Scherer, S.W.; Marshall, C.R.; Potocki, L.; et al. Identification of ANKRD11 and ZNF778 as candidate genes for autism and variable cognitive impairment in the novel 16q24.3 microdeletion syndrome. Eur. J. Hum. Genet. EJHG 2010, 18, 429–435. [Google Scholar] [CrossRef]
- Kumar, R.A.; KaraMohamed, S.; Sudi, J.; Conrad, D.F.; Brune, C.; Badner, J.A.; Gilliam, T.C.; Nowak, N.J.; Cook, E.H., Jr.; Dobyns, W.B.; et al. Recurrent 16p11.2 microdeletions in autism. Hum. Mol. Genet. 2008, 17, 628–638. [Google Scholar] [CrossRef]
- Kim, H.-G.; Kishikawa, S.; Higgins, A.W.; Seong, I.-S.; Donovan, D.J.; Shen, Y.; Lally, E.; Weiss, L.A.; Najm, J.; Kutsche, K. Disruption of neurexin 1 associated with autism spectrum disorder. Am. J. Hum. Genet. 2008, 82, 199–207. [Google Scholar] [CrossRef]
- He, H.; Tan, C.K.; Downey, K.M.; So, A.G. A tumor necrosis factor alpha- and interleukin 6-inducible protein that interacts with the small subunit of DNA polymerase delta and proliferating cell nuclear antigen. Proc. Natl. Acad. Sci. USA 2001, 98, 11979–11984. [Google Scholar] [CrossRef] [PubMed]
- Golzio, C.; Willer, J.; Talkowski, M.E.; Oh, E.C.; Taniguchi, Y.; Jacquemont, S.; Reymond, A.; Sun, M.; Sawa, A.; Gusella, J.F.; et al. KCTD13 is a major driver of mirrored neuroanatomical phenotypes of the 16p11.2 copy number variant. Nature 2012, 485, 363–367. [Google Scholar] [CrossRef]
- Escamilla, C.O.; Filonova, I.; Walker, A.K.; Xuan, Z.X.; Holehonnur, R.; Espinosa, F.; Liu, S.; Thyme, S.B.; Lopez-Garcia, I.A.; Mendoza, D.B.; et al. Kctd13 deletion reduces synaptic transmission via increased RhoA. Nature 2017, 551, 227–231. [Google Scholar] [CrossRef] [PubMed]
- Leblond, C.S.; Cliquet, F.; Carton, C.; Huguet, G.; Mathieu, A.; Kergrohen, T.; Buratti, J.; Lemiere, N.; Cuisset, L.; Bienvenu, T.; et al. Both rare and common genetic variants contribute to autism in the Faroe Islands. NPJ Genom. Med. 2019, 4, 1. [Google Scholar] [CrossRef] [PubMed]
- Yan, Y.; Eipper, B.A.; Mains, R.E. Kalirin-9 and Kalirin-12 Play Essential Roles in Dendritic Outgrowth and Branching. Cereb. Cortex 2015, 25, 3487–3501. [Google Scholar] [CrossRef] [PubMed]
- Qu, B.X.; Gong, Y.; Sinclair, D.; Fu, M.; Perl, D.; Diaz-Arrastia, R. cPLA2alpha knockout mice exhibit abnormalities in the architecture and synapses of cortical neurons. Brain Res. 2013, 1497, 101–105. [Google Scholar] [CrossRef]
- Wang, K.; Zhang, H.; Ma, D.; Bucan, M.; Glessner, J.T.; Abrahams, B.S.; Salyakina, D.; Imielinski, M.; Bradfield, J.P.; Sleiman, P.M.; et al. Common genetic variants on 5p14.1 associate with autism spectrum disorders. Nature 2009, 459, 528–533. [Google Scholar] [CrossRef]
- Mirzaa, G.M.; Chong, J.X.; Piton, A.; Popp, B.; Foss, K.; Guo, H.; Harripaul, R.; Xia, K.; Scheck, J.; Aldinger, K.A.; et al. De novo and inherited variants in ZNF292 underlie a neurodevelopmental disorder with features of autism spectrum disorder. Genet. Med. Off. 2019. [Google Scholar] [CrossRef]
- Irimia, M.; Weatheritt, R.J.; Ellis, J.D.; Parikshak, N.N.; Gonatopoulos-Pournatzis, T.; Babor, M.; Quesnel-Vallieres, M.; Tapial, J.; Raj, B.; O’Hanlon, D.; et al. A highly conserved program of neuronal microexons is misregulated in autistic brains. Cell 2014, 159, 1511–1523. [Google Scholar] [CrossRef]
- Sebat, J.; Lakshmi, B.; Malhotra, D.; Troge, J.; Lese-Martin, C.; Walsh, T.; Yamrom, B.; Yoon, S.; Krasnitz, A.; Kendall, J.; et al. Strong association of de novo copy number mutations with autism. Science 2007, 316, 445–449. [Google Scholar] [CrossRef]
- Alvarez-Baron, E.; Michel, K.; Mittelstaedt, T.; Opitz, T.; Schmitz, F.; Beck, H.; Dietrich, D.; Becker, A.J.; Schoch, S. RIM3gamma and RIM4gamma are key regulators of neuronal arborization. J. Neurosci. Off. 2013, 33, 824–839. [Google Scholar] [CrossRef]
- Ponson, L.; Gomot, M.; Blanc, R.; Barthelemy, C.; Roux, S.; Munnich, A.; Romana, S.; Aguillon-Hernandez, N.; Malan, V.; Bonnet-Brilhault, F. 22q13 deletion syndrome: Communication disorder or autism? Evidence from a specific clinical and neurophysiological phenotype. Transl. Psychiatry 2018, 8, 1–8. [Google Scholar] [CrossRef]
- Zhou, Y.; Sharma, J.; Ke, Q.; Landman, R.; Yuan, J.; Chen, H.; Hayden, D.S.; Fisher, J.W.; Jiang, M.; Menegas, W. Atypical behaviour and connectivity in SHANK3-mutant macaques. Nature 2019, 570, 326–331. [Google Scholar] [CrossRef] [PubMed]
- Jamain, S.; Quach, H.; Betancur, C.; Rastam, M.; Colineaux, C.; Gillberg, I.C.; Soderstrom, H.; Giros, B.; Leboyer, M.; Gillberg, C.; et al. Mutations of the X-linked genes encoding neuroligins NLGN3 and NLGN4 are associated with autism. Nat. Genet. 2003, 34, 27–29. [Google Scholar] [CrossRef] [PubMed]
- Darnell, J.C.; Van Driesche, S.J.; Zhang, C.; Hung, K.Y.; Mele, A.; Fraser, C.E.; Stone, E.F.; Chen, C.; Fak, J.J.; Chi, S.W.; et al. FMRP stalls ribosomal translocation on mRNAs linked to synaptic function and autism. Cell 2011, 146, 247–261. [Google Scholar] [CrossRef] [PubMed]
- Mirakian, R.; Ewan, P.W.; Durham, S.R.; Youlten, L.J.; Dugue, P.; Friedmann, P.S.; English, J.S.; Huber, P.A.; Nasser, S.M.; BSACI. BSACI guidelines for the management of drug allergy. Clin. Exp. Allergy 2009, 39, 43–61. [Google Scholar] [CrossRef] [PubMed]
- Arking, D.E.; Cutler, D.J.; Brune, C.W.; Teslovich, T.M.; West, K.; Ikeda, M.; Rea, A.; Guy, M.; Lin, S.; Cook, E.H., Jr. A common genetic variant in the neurexin superfamily member CNTNAP2 increases familial risk of autism. Am. J. Hum. Genet. 2008, 82, 160–164. [Google Scholar] [CrossRef] [PubMed]
- Morrow, E.M.; Yoo, S.-Y.; Flavell, S.W.; Kim, T.-K.; Lin, Y.; Hill, R.S.; Mukaddes, N.M.; Balkhy, S.; Gascon, G.; Hashmi, A. Identifying autism loci and genes by tracing recent shared ancestry. Science 2008, 321, 218–223. [Google Scholar] [CrossRef]
- Yuan, H.; Meng, Z.; Zhang, L.; Luo, X.; Liu, L.; Chen, M.; Li, X.; Zhao, W.; Liang, L. A rare de novo interstitial duplication of 15q15.3q21.2 in a boy with severe short stature, hypogonadism, global developmental delay and intellectual disability. Mol. Cytogenet. 2016, 9, 2. [Google Scholar] [CrossRef]
- Bassell, G.J.; Warren, S.T. Fragile X syndrome: Loss of local mRNA regulation alters synaptic development and function. Neuron 2008, 60, 201–214. [Google Scholar] [CrossRef]
- Engwerda, A.; Frentz, B.; den Ouden, A.L.; Flapper, B.C.T.; Swertz, M.A.; Gerkes, E.H.; Plantinga, M.; Dijkhuizen, T.; van Ravenswaaij-Arts, C.M.A. The phenotypic spectrum of proximal 6q deletions based on a large cohort derived from social media and literature reports. Eur. J. Hum. Genet. EJHG 2018, 26, 1478–1489. [Google Scholar] [CrossRef]
- Samaco, R.C.; Hogart, A.; LaSalle, J.M. Epigenetic overlap in autism-spectrum neurodevelopmental disorders: MECP2 deficiency causes reduced expression of UBE3A and GABRB3. Hum. Mol. Genet. 2004, 14, 483–492. [Google Scholar] [CrossRef]
- Steffenburg, S.; Gillberg, C.L.; Steffenburg, U.; Kyllerman, M. Autism in Angelman syndrome: A population-based study. Pediatr. Neurol. 1996, 14, 131–136. [Google Scholar] [CrossRef]
- Yashiro, K.; Riday, T.T.; Condon, K.H.; Roberts, A.C.; Bernardo, D.R.; Prakash, R.; Weinberg, R.J.; Ehlers, M.D.; Philpot, B.D. Ube3a is required for experience-dependent maturation of the neocortex. Nat. Neurosci. 2009, 12, 777–783. [Google Scholar] [CrossRef]
- Smith, S.E.; Zhou, Y.D.; Zhang, G.; Jin, Z.; Stoppel, D.C.; Anderson, M.P. Increased gene dosage of Ube3a results in autism traits and decreased glutamate synaptic transmission in mice. Sci. Transl. Med. 2011, 3. [Google Scholar] [CrossRef] [PubMed]
- Adalsteinsson, B.T.; Ferguson-Smith, A.C. Epigenetic control of the genome—Lessons from genomic imprinting. Genes 2014, 5, 635–655. [Google Scholar] [CrossRef]
- Schiele, M.; Domschke, K. Epigenetics at the crossroads between genes, environment and resilience in anxiety disorders. Genes Brain Behav. 2018, 17, e12423. [Google Scholar] [CrossRef]
- Ben-David, E.; Shifman, S. Combined analysis of exome sequencing points toward a major role for transcription regulation during brain development in autism. Mol. Psychiatry 2013, 18, 1054. [Google Scholar] [CrossRef]
- Autism Spectrum Disorders Working Group of The Psychiatric Genomics Consortium. Meta-analysis of GWAS of over 16,000 individuals with autism spectrum disorder highlights a novel locus at 10q24. 32 and a significant overlap with schizophrenia. Mol. Autism 2017, 8, 1–17.
- Waye, M.M.; Cheng, H.Y. Genetics and epigenetics of autism: A Review. Psychiatry Clin. Neurosci. 2018, 72, 228–244. [Google Scholar] [CrossRef]
- Kubota, T.; Mochizuki, K. Epigenetic effect of environmental factors on autism spectrum disorders. Int. J. Environ. Res. Public Health 2016, 13, 504. [Google Scholar] [CrossRef]
- LoParo, D.; Waldman, I. The oxytocin receptor gene (OXTR) is associated with autism spectrum disorder: A meta-analysis. Mol. Psychiatry 2015, 20, 640–646. [Google Scholar] [CrossRef]
- Qin, L.; Ma, K.; Wang, Z.-J.; Hu, Z.; Matas, E.; Wei, J.; Yan, Z. Social deficits in Shank3-deficient mouse models of autism are rescued by histone deacetylase (HDAC) inhibition. Nat. Neurosci. 2018, 21, 564. [Google Scholar] [CrossRef] [PubMed]
- Uchino, S.; Waga, C. SHANK3 as an autism spectrum disorder-associated gene. Brain Dev. 2013, 35, 106–110. [Google Scholar] [CrossRef] [PubMed]
- Goncalves, T.F.; Goncalves, A.P.; Fintelman Rodrigues, N.; dos Santos, J.M.; Pimentel, M.M.; Santos-Reboucas, C.B. KDM5C mutational screening among males with intellectual disability suggestive of X-Linked inheritance and review of the literature. Eur. J. Med. Genet. 2014, 57, 138–144. [Google Scholar] [CrossRef] [PubMed]
- Duffney, L.J.; Valdez, P.; Tremblay, M.W.; Cao, X.; Montgomery, S.; McConkie-Rosell, A.; Jiang, Y.h. Epigenetics and autism spectrum disorder: A report of an autism case with mutation in H1 linker histone HIST1H1E and literature review. Am. J. Med Genet. Part B 2018, 177, 426–433. [Google Scholar] [CrossRef] [PubMed]
- Cotney, J.; Muhle, R.A.; Sanders, S.J.; Liu, L.; Willsey, A.J.; Niu, W.; Liu, W.; Klei, L.; Lei, J.; Yin, J. The autism-associated chromatin modifier CHD8 regulates other autism risk genes during human neurodevelopment. Nat. Commun. 2015, 6, 6404. [Google Scholar] [CrossRef]
- Hamdan, F.F.; Srour, M.; Capo-Chichi, J.-M.; Daoud, H.; Nassif, C.; Patry, L.; Massicotte, C.; Ambalavanan, A.; Spiegelman, D.; Diallo, O. De novo mutations in moderate or severe intellectual disability. PLoS Genet. 2014, 10, e1004772. [Google Scholar] [CrossRef] [PubMed]
- Helsmoortel, C.; Vulto-van Silfhout, A.T.; Coe, B.P.; Vandeweyer, G.; Rooms, L.; Van Den Ende, J.; Schuurs-Hoeijmakers, J.H.; Marcelis, C.L.; Willemsen, M.H.; Vissers, L.E. A SWI/SNF-related autism syndrome caused by de novo mutations in ADNP. Nat. Genet. 2014, 46, 380. [Google Scholar] [CrossRef]
- Bernstein, E.; Kim, S.Y.; Carmell, M.A.; Murchison, E.P.; Alcorn, H.; Li, M.Z.; Mills, A.A.; Elledge, S.J.; Anderson, K.V.; Hannon, G.J. Dicer is essential for mouse development. Nat. Genet. 2003, 35, 215–217. [Google Scholar] [CrossRef]
- Hébert, S.S.; De Strooper, B. miRNAs in neurodegeneration. Science 2007, 317, 1179–1180. [Google Scholar] [CrossRef]
- Davis, T.H.; Cuellar, T.L.; Koch, S.M.; Barker, A.J.; Harfe, B.D.; McManus, M.T.; Ullian, E.M. Conditional loss of Dicer disrupts cellular and tissue morphogenesis in the cortex and hippocampus. J. Neurosci. 2008, 28, 4322–4330. [Google Scholar] [CrossRef]
- Kawase-Koga, Y.; Otaegi, G.; Sun, T. Different timings of Dicer deletion affect neurogenesis and gliogenesis in the developing mouse central nervous system. Dev. Dyn. Off. 2009, 238, 2800–2812. [Google Scholar] [CrossRef] [PubMed]
- LaSalle, J.M. Epigenomic strategies at the interface of genetic and environmental risk factors for autism. J. Hum. Genet. 2013, 58, 396–401. [Google Scholar] [CrossRef]
- James, S.J.; Cutler, P.; Melnyk, S.; Jernigan, S.; Janak, L.; Gaylor, D.W.; Neubrander, J.A. Metabolic biomarkers of increased oxidative stress and impaired methylation capacity in children with autism. Am. J. Clin. Nutr. 2004, 80, 1611–1617. [Google Scholar] [CrossRef] [PubMed]
- Dolinoy, D.C.; Weidman, J.R.; Jirtle, R.L. Epigenetic gene regulation: Linking early developmental environment to adult disease. Reprod. Toxicol. 2007, 23, 297–307. [Google Scholar] [CrossRef] [PubMed]
- Jirtle, R.L.; Skinner, M.K. Environmental epigenomics and disease susceptibility. Nat. Rev. Genet. 2007, 8, 253–262. [Google Scholar] [CrossRef]
- Rusiecki, J.A.; Baccarelli, A.; Bollati, V.; Tarantini, L.; Moore, L.E.; Bonefeld-Jorgensen, E.C. Global DNA hypomethylation is associated with high serum-persistent organic pollutants in Greenlandic Inuit. Environ. Health Perspect. 2008, 116, 1547–1552. [Google Scholar] [CrossRef]
- Jang, H.S.; Shin, W.J.; Lee, J.E.; Do, J.T. CpG and Non-CpG Methylation in Epigenetic Gene Regulation and Brain Function. Genes 2017, 8, 148. [Google Scholar] [CrossRef]
- Wong, C.; Meaburn, E.L.; Ronald, A.; Price, T.; Jeffries, A.R.; Schalkwyk, L.; Plomin, R.; Mill, J. Methylomic analysis of monozygotic twins discordant for autism spectrum disorder and related behavioural traits. Mol. Psychiatry 2014, 19, 495. [Google Scholar] [CrossRef]
- Tahiliani, M.; Koh, K.P.; Shen, Y.; Pastor, W.A.; Bandukwala, H.; Brudno, Y.; Agarwal, S.; Iyer, L.M.; Liu, D.R.; Aravind, L. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 2009, 324, 930–935. [Google Scholar] [CrossRef]
- Shcheglovitov, A.; Shcheglovitova, O.; Yazawa, M.; Portmann, T.; Shu, R.; Sebastiano, V.; Krawisz, A.; Froehlich, W.; Bernstein, J.A.; Hallmayer, J.F. SHANK3 and IGF1 restore synaptic deficits in neurons from 22q13 deletion syndrome patients. Nature 2013, 503, 267–271. [Google Scholar] [CrossRef]
- Yi, F.; Danko, T.; Botelho, S.C.; Patzke, C.; Pak, C.; Wernig, M.; Südhof, T.C. Autism-associated SHANK3 haploinsufficiency causes Ih channelopathy in human neurons. Science 2016, 352, aaf2669. [Google Scholar] [CrossRef] [PubMed]
- Katayama, Y.; Nishiyama, M.; Shoji, H.; Ohkawa, Y.; Kawamura, A.; Sato, T.; Suyama, M.; Takumi, T.; Miyakawa, T.; Nakayama, K.I. CHD8 haploinsufficiency results in autistic-like phenotypes in mice. Nature 2016, 537, 675–679. [Google Scholar] [CrossRef] [PubMed]
- Celen, C.; Chuang, J.-C.; Luo, X.; Nijem, N.; Walker, A.K.; Chen, F.; Zhang, S.; Chung, A.S.; Nguyen, L.H.; Nassour, I. Arid1b haploinsufficient mice reveal neuropsychiatric phenotypes and reversible causes of growth impairment. Elife 2017, 6, e25730. [Google Scholar] [CrossRef]
- Jung, E.-M.; Moffat, J.J.; Liu, J.; Dravid, S.M.; Gurumurthy, C.B.; Kim, W.-Y. Arid1b haploinsufficiency disrupts cortical interneuron development and mouse behavior. Nat. Neurosci. 2017, 20, 1694–1707. [Google Scholar] [CrossRef]
- Shibutani, M.; Horii, T.; Shoji, H.; Morita, S.; Kimura, M.; Terawaki, N.; Miyakawa, T.; Hatada, I. Arid1b haploinsufficiency causes abnormal brain gene expression and autism-related behaviors in mice. Int. J. Mol. Sci. 2017, 18, 1872. [Google Scholar] [CrossRef]
- Jung, H.; Park, H.; Choi, Y.; Kang, H.; Lee, E.; Kweon, H.; Roh, J.D.; Ellegood, J.; Choi, W.; Kang, J. Sexually dimorphic behavior, neuronal activity, and gene expression in Chd8-mutant mice. Nat. Neurosci. 2018, 21, 1218–1228. [Google Scholar] [CrossRef]
- Suetterlin, P.; Hurley, S.; Mohan, C.; Riegman, K.L.; Pagani, M.; Caruso, A.; Ellegood, J.; Galbusera, A.; Crespo-Enriquez, I.; Michetti, C. Altered neocortical gene expression, brain overgrowth and functional over-connectivity in Chd8 haploinsufficient mice. Cereb. Cortex 2018, 28, 2192–2206. [Google Scholar] [CrossRef]
- Schaefer, A.; Sampath, S.C.; Intrator, A.; Min, A.; Gertler, T.S.; Surmeier, D.J.; Tarakhovsky, A.; Greengard, P. Control of cognition and adaptive behavior by the GLP/G9a epigenetic suppressor complex. Neuron 2009, 64, 678–691. [Google Scholar] [CrossRef]
- Balemans, M.C.; Huibers, M.M.; Eikelenboom, N.W.; Kuipers, A.J.; van Summeren, R.C.; Pijpers, M.M.; Tachibana, M.; Shinkai, Y.; van Bokhoven, H.; Van der Zee, C.E. Reduced exploration, increased anxiety, and altered social behavior: Autistic-like features of euchromatin histone methyltransferase 1 heterozygous knockout mice. Behav. Brain Res. 2010, 208, 47–55. [Google Scholar] [CrossRef]
- Shu, W.; Cho, J.Y.; Jiang, Y.; Zhang, M.; Weisz, D.; Elder, G.A.; Schmeidler, J.; De Gasperi, R.; Sosa, M.A.G.; Rabidou, D. Altered ultrasonic vocalization in mice with a disruption in the Foxp2 gene. Proc. Natl. Acad. Sci. USA 2005, 102, 9643–9648. [Google Scholar] [CrossRef]
- Araujo, D.J.; Anderson, A.G.; Berto, S.; Runnels, W.; Harper, M.; Ammanuel, S.; Rieger, M.A.; Huang, H.-C.; Rajkovich, K.; Loerwald, K.W. FoxP1 orchestration of ASD-relevant signaling pathways in the striatum. Genes Dev. 2015, 29, 2081–2096. [Google Scholar] [CrossRef] [PubMed]
- Bacon, C.; Schneider, M.; Le Magueresse, C.; Froehlich, H.; Sticht, C.; Gluch, C.; Monyer, H.; Rappold, G. Brain-specific Foxp1 deletion impairs neuronal development and causes autistic-like behaviour. Mol. Psychiatry 2015, 20, 632–639. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.-C.; Kuo, H.-Y.; Bornschein, U.; Takahashi, H.; Chen, S.-Y.; Lu, K.-M.; Yang, H.-Y.; Chen, G.-M.; Lin, J.-R.; Lee, Y.-H. Foxp2 controls synaptic wiring of corticostriatal circuits and vocal communication by opposing Mef2c. Nat. Neurosci. 2016, 19, 1513–1522. [Google Scholar] [CrossRef] [PubMed]
- Araujo, D.J.; Toriumi, K.; Escamilla, C.O.; Kulkarni, A.; Anderson, A.G.; Harper, M.; Usui, N.; Ellegood, J.; Lerch, J.P.; Birnbaum, S.G. Foxp1 in forebrain pyramidal neurons controls gene expression required for spatial learning and synaptic plasticity. J. Neurosci. 2017, 37, 10917–10931. [Google Scholar] [CrossRef] [PubMed]
- Usui, N.; Araujo, D.J.; Kulkarni, A.; Ellegood, J.; Harper, M.; Toriumi, K.; Lerch, J.P.; Konopka, G. Foxp1 regulation of neonatal vocalizations via cortical development. Genes Dev. 2017, 31, 2039–2055. [Google Scholar] [CrossRef]
- Medvedeva, V.P.; Rieger, M.A.; Vieth, B.; Mombereau, C.; Ziegenhain, C.; Ghosh, T.; Cressant, A.; Enard, W.; Granon, S.; Dougherty, J.D. Altered social behavior in mice carrying a cortical Foxp2 deletion. Hum. Mol. Genet. 2019, 28, 701–717. [Google Scholar] [CrossRef]
- French, C.A.; Veloz, M.F.V.; Zhou, K.; Peter, S.; Fisher, S.E.; Costa, R.M.; De Zeeuw, C.I. Differential effects of Foxp2 disruption in distinct motor circuits. Mol. Psychiatry 2019, 24, 447–462. [Google Scholar] [CrossRef]
- Sessa, A.; Fagnocchi, L.; Mastrototaro, G.; Massimino, L.; Zaghi, M.; Indrigo, M.; Cattaneo, S.; Martini, D.; Gabellini, C.; Pucci, C.J.N. SETD5 regulates chromatin methylation state and preserves global transcriptional fidelity during brain development and neuronal wiring. Neuron 2019, 104, 271–289. [Google Scholar] [CrossRef]
- Deliu, E.; Arecco, N.; Morandell, J.; Dotter, C.P.; Contreras, X.; Girardot, C.; Käsper, E.-L.; Kozlova, A.; Kishi, K.; Chiaradia, I. Haploinsufficiency of the intellectual disability gene SETD5 disturbs developmental gene expression and cognition. Nat. Neurosci. 2018, 21, 1717–1727. [Google Scholar] [CrossRef]
- Osipovich, A.; Gangula, R.; Vianna, P.G.; Magnuson, M. Setd5 is essential for mammalian development and the co-transcriptional regulation of histone acetylation. Development 2016, 143, 4595–4607. [Google Scholar] [CrossRef]
- Zhubi, A.; Chen, Y.; Dong, E.; Cook, E.; Guidotti, A.; Grayson, D. Increased binding of MeCP2 to the GAD1 and RELN promoters may be mediated by an enrichment of 5-hmC in autism spectrum disorder (ASD) cerebellum. Transl. Psychiatry 2014, 4, e349. [Google Scholar] [CrossRef] [PubMed]
- Peters, S.U.; Hundley, R.J.; Wilson, A.K.; Warren, Z.; Vehorn, A.; Carvalho, C.M.; Lupski, J.R.; Ramocki, M.B. The Behavioral Phenotype in MECP 2 Duplication Syndrome: A Comparison With Idiopathic Autism. Autism Res. 2013, 6, 42–50. [Google Scholar] [CrossRef] [PubMed]
- Nagarajan, R.; Hogart, A.; Gwye, Y.; Martin, M.R.; LaSalle, J.M. Reduced MeCP2 expression is frequent in autism frontal cortex and correlates with aberrant MECP2 promoter methylation. Epigenetics 2006, 1, 172–182. [Google Scholar] [CrossRef]
- Lu, Z.; Liu, Z.; Mao, W.; Wang, X.; Zheng, X.; Chen, S.; Cao, B.; Huang, S.; Zhang, X.; Zhou, T.; et al. Locus-specific DNA methylation of Mecp2 promoter leads to autism-like phenotypes in mice. Cell Death Dis. 2020, 11, 85–111. [Google Scholar] [CrossRef]
- Kuwano, Y.; Kamio, Y.; Kawai, T.; Katsuura, S.; Inada, N.; Takaki, A.; Rokutan, K. Autism-associated gene expression in peripheral leucocytes commonly observed between subjects with autism and healthy women having autistic children. PLoS ONE 2011, 6, e24723. [Google Scholar] [CrossRef]
- Gonzales, M.L.; Adams, S.; Dunaway, K.W.; LaSalle, J.M. Phosphorylation of distinct sites in MeCP2 modifies cofactor associations and the dynamics of transcriptional regulation. Mol. Cell. Biol. 2012, 32, 2894–2903. [Google Scholar] [CrossRef]
- Yip, J.; Soghomonian, J.-J.; Blatt, G.J. Decreased GAD67 mRNA levels in cerebellar Purkinje cells in autism: Pathophysiological implications. Acta Neuropathol. 2007, 113, 559–568. [Google Scholar] [CrossRef]
- Yip, J.; Soghomonian, J.J.; Blatt, G.J. Increased GAD67 mRNA expression in cerebellar interneurons in autism: Implications for Purkinje cell dysfunction. J. Neurosci. Res. 2008, 86, 525–530. [Google Scholar] [CrossRef]
- Bosch, C.; Muhaisen, A.; Pujadas, L.; Soriano, E.; Martinez, A. Reelin Exerts Structural, Biochemical and Transcriptional Regulation Over Presynaptic and Postsynaptic Elements in the Adult Hippocampus. Front. Cell. Neurosci. 2016, 10, 138. [Google Scholar] [CrossRef]
- Gregory, S.G.; Connelly, J.J.; Towers, A.J.; Johnson, J.; Biscocho, D.; Markunas, C.A.; Lintas, C.; Abramson, R.K.; Wright, H.H.; Ellis, P. Genomic and epigenetic evidence for oxytocin receptor deficiency in autism. BMC Med. 2009, 7, 62. [Google Scholar] [CrossRef]
- Behnia, F.; Parets, S.E.; Kechichian, T.; Yin, H.; Dutta, E.H.; Saade, G.R.; Smith, A.K.; Menon, R. Fetal DNA methylation of autism spectrum disorders candidate genes: Association with spontaneous preterm birth. Am. J. Obstet. Gynecol. 2015, 212, e531–e533. [Google Scholar]
- Andari, E.; Nishitani, S.; Kaundinya, G.; Caceres, G.A.; Morrier, M.J.; Ousley, O.; Smith, A.K.; Cubells, J.F.; Young, L.J. Epigenetic modification of the oxytocin receptor gene: implications for autism symptom severity and brain functional connectivity. Neuropsychopharmacol. 2020, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Wang, X.; Li, X.L.; Towers, A.; Cao, X.; Wang, P.; Bowman, R.; Yang, H.; Goldstein, J.; Li, Y.J.; et al. Epigenetic dysregulation of SHANK3 in brain tissues from individuals with autism spectrum disorders. Hum. Mol. Genet. 2014, 23, 1563–1578. [Google Scholar] [CrossRef] [PubMed]
- Shilatifard, A. Molecular implementation and physiological roles for histone H3 lysine 4 (H3K4) methylation. Curr. Opin. Cell Biol. 2008, 20, 341–348. [Google Scholar] [CrossRef]
- Gupta, S.; Kim, S.Y.; Artis, S.; Molfese, D.L.; Schumacher, A.; Sweatt, J.D.; Paylor, R.E.; Lubin, F.D. Histone methylation regulates memory formation. J. Neurosci. Off. 2010, 30, 3589–3599. [Google Scholar] [CrossRef]
- Shulha, H.P.; Cheung, I.; Whittle, C.; Wang, J.; Virgil, D.; Lin, C.L.; Guo, Y.; Lessard, A.; Akbarian, S.; Weng, Z. Epigenetic signatures of autism: Trimethylated H3K4 landscapes in prefrontal neurons. Arch. Gen. Psychiatry 2012, 69, 314–324. [Google Scholar] [CrossRef]
- Sanders, S.J.; He, X.; Willsey, A.J.; Ercan-Sencicek, A.G.; Samocha, K.E.; Cicek, A.E.; Murtha, M.T.; Bal, V.H.; Bishop, S.L.; Dong, S. Insights into autism spectrum disorder genomic architecture and biology from 71 risk loci. Neuron 2015, 87, 1215–1233. [Google Scholar] [CrossRef]
- Bernier, R.; Golzio, C.; Xiong, B.; Stessman, H.A.; Coe, B.P.; Penn, O.; Witherspoon, K.; Gerdts, J.; Baker, C.; Vulto-van Silfhout, A.T. Disruptive CHD8 mutations define a subtype of autism early in development. Cell 2014, 158, 263–276. [Google Scholar] [CrossRef]
- Ronan, J.L.; Wu, W.; Crabtree, G.R. From neural development to cognition: Unexpected roles for chromatin. Nat. Rev. Genet. 2013, 14, 347–359. [Google Scholar] [CrossRef]
- Fregeac, J.; Colleaux, L.; Nguyen, L.S. The emerging roles of MicroRNAs in autism spectrum disorders. Neurosci. Biobehav. Rev. 2016, 71, 729–738. [Google Scholar] [CrossRef]
- Mor, M.; Nardone, S.; Sams, D.S.; Elliott, E. Hypomethylation of miR-142 promoter and upregulation of microRNAs that target the oxytocin receptor gene in the autism prefrontal cortex. Mol. Autism 2015, 6, 46. [Google Scholar] [CrossRef]
- Abu-Elneel, K.; Liu, T.; Gazzaniga, F.S.; Nishimura, Y.; Wall, D.P.; Geschwind, D.H.; Lao, K.; Kosik, K.S. Heterogeneous dysregulation of microRNAs across the autism spectrum. Neurogenetics 2008, 9, 153–161. [Google Scholar] [CrossRef] [PubMed]
- Seno, M.M.G.; Hu, P.; Gwadry, F.G.; Pinto, D.; Marshall, C.R.; Casallo, G.; Scherer, S.W. Gene and miRNA expression profiles in autism spectrum disorders. Brain Res. 2011, 1380, 85–97. [Google Scholar] [CrossRef] [PubMed]
- Popov, N.T.; Stoyanova, V.K.; Madzhirova, N.P.; Vachev, T.I. Epigenetic aspects in schizophrenia etiology and pathogenesis. Folia Med. 2012, 54, 12–16. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zhang, Y.; Wang, Z.; Gemeinhart, R.A. Progress in microRNA delivery. J. Control. Release 2013, 172, 962–974. [Google Scholar] [CrossRef] [PubMed]
- Johnson, Z.V.; Young, L.J. Oxytocin and vasopressin neural networks: Implications for social behavioral diversity and translational neuroscience. Neurosci. Biobehav. Rev. 2017, 76, 87–98. [Google Scholar] [CrossRef]
- Freeman, S.M.; Palumbo, M.C.; Lawrence, R.H.; Smith, A.L.; Goodman, M.M.; Bales, K.L. Effect of age and autism spectrum disorder on oxytocin receptor density in the human basal forebrain and midbrain. Transl. Psychiatry 2018, 8, 257. [Google Scholar] [CrossRef]
- Andari, E.; Duhamel, J.R.; Zalla, T.; Herbrecht, E.; Leboyer, M.; Sirigu, A. Promoting social behavior with oxytocin in high-functioning autism spectrum disorders. Proc. Natl. Acad. Sci. USA 2010, 107, 4389–4394. [Google Scholar] [CrossRef]
- Donaldson, Z.R.; Young, L.J. Oxytocin, vasopressin, and the neurogenetics of sociality. Science 2008, 322, 900–904. [Google Scholar] [CrossRef]
- Procyshyn, T.L.; Hurd, P.L.; Crespi, B.J. Association testing of vasopressin receptor 1a microsatellite polymorphisms in non-clinical autism spectrum phenotypes. Autism Res. 2017, 10, 750–756. [Google Scholar] [CrossRef]
- Yirmiya, N.; Rosenberg, C.; Levi, S.; Salomon, S.; Shulman, C.; Nemanov, L.; Dina, C.; Ebstein, R.P. Association between the arginine vasopressin 1a receptor (AVPR1a) gene and autism in a family-based study: Mediation by socialization skills. Mol. Psychiatry 2006, 11, 488–494. [Google Scholar] [CrossRef] [PubMed]
- Umbricht, D.; Del Valle Rubido, M.; Hollander, E.; McCracken, J.T.; Shic, F.; Scahill, L.; Noeldeke, J.; Boak, L.; Khwaja, O.; Squassante, L.; et al. A Single Dose, Randomized, Controlled Proof-Of-Mechanism Study of a Novel Vasopressin 1a Receptor Antagonist (RG7713) in High-Functioning Adults with Autism Spectrum Disorder. Neuropsychopharmacol. Off. 2017, 42, 1914–1923. [Google Scholar] [CrossRef] [PubMed]
- Zarrei, M.; Burton, C.L.; Engchuan, W.; Young, E.J.; Higginbotham, E.J.; MacDonald, J.R.; Trost, B.; Chan, A.J.S.; Walker, S.; Lamoureux, S.; et al. A large data resource of genomic copy number variation across neurodevelopmental disorders. NPJ Genom. Med. 2019, 4, 26. [Google Scholar] [CrossRef]
- Hong, Y.J.; Do, J.T. Neural Lineage Differentiation From Pluripotent Stem Cells to Mimic Human Brain Tissues. Front. Bioeng. Biotechnol. 2019, 7, 400. [Google Scholar] [CrossRef]
- Russo, F.B.; Brito, A.; de Freitas, A.M.; Castanha, A.; de Freitas, B.C.; Beltrao-Braga, P.C.B. The use of iPSC technology for modeling Autism Spectrum Disorders. Neurobiol. Dis. 2019, 130, 104483. [Google Scholar] [CrossRef]
- Mariani, J.; Coppola, G.; Zhang, P.; Abyzov, A.; Provini, L.; Tomasini, L.; Amenduni, M.; Szekely, A.; Palejev, D.; Wilson, M.; et al. FOXG1-Dependent Dysregulation of GABA/Glutamate Neuron Differentiation in Autism Spectrum Disorders. Cell 2015, 162, 375–390. [Google Scholar] [CrossRef]
- Saurman, V.; Margolis, K.G.; Luna, R.A. Autism Spectrum Disorder as a Brain-Gut-Microbiome Axis Disorder. Dig. Dis. Sci. 2020. [Google Scholar] [CrossRef]
- Collins, S.M.; Surette, M.; Bercik, P. The interplay between the intestinal microbiota and the brain. Nat. Rev. Microbiol. 2012, 10, 735–742. [Google Scholar] [CrossRef]
- Alessio, N.; Brigida, A.L.; Peluso, G.; Antonucci, N.; Galderisi, U.; Siniscalco, D. Stem Cell-Derived Exosomes in Autism Spectrum Disorder. Int. J. Environ. Res. Public Health 2020, 17, 944. [Google Scholar] [CrossRef]
- Li, Q.; Wang, H.; Peng, H.; Huyan, T.; Cacalano, N.A. Exosomes: Versatile Nano Mediators of Immune Regulation. Cancers 2019, 11, 1557. [Google Scholar] [CrossRef]
- Matta, S.M.; Hill-Yardin, E.L.; Crack, P.J. The influence of neuroinflammation in Autism Spectrum Disorder. Brain Behav. Immun. 2019, 79, 75–90. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Sui, H.; Zheng, Y.; Jiang, Y.; Shi, Y.; Liang, J.; Zhao, L. Curcumin-primed exosomes potently ameliorate cognitive function in AD mice by inhibiting hyperphosphorylation of the Tau protein through the AKT/GSK-3beta pathway. Nanoscale 2019, 11, 7481–7496. [Google Scholar] [CrossRef] [PubMed]
- Bhatia, S.N.; Ingber, D.E. Microfluidic organs-on-chips. Nat. Biotechnol. 2014, 32, 760–772. [Google Scholar] [CrossRef] [PubMed]
| Locus | Function | Genes | Variation | Reference |
|---|---|---|---|---|
| 3q21.1 - 3q21.2 | Abnormalities in neuronal maturation and long-term potentiation in the brain, macrocephaly, intellectual disability facial dysmorphism | KALRN | Duplication | [166,167,168] |
| 5p14.1 | Neuronal cell-adhesion molecules | Cadherin 10 (CDH10) Cadherin 9 (CDH9) | Deletion | [159,169] |
| 6q14.3 | Learning problems, intellectual disability, behavioral problems | ZNF292 (zinc finger protein 292) | Deletion | [170] |
| 12q24.23 | Neuronal cells and misregulated neural ‘microexons’ in the brains | nSR100/SRRM4 | [171] | |
| 16p11.2 | Reduced proliferation of neuronal progenitors, the increased cell death during the brain development, microcephaly | KCTD13 | Deletion Duplication | [161,162,163,164,165] |
| 16q24.3 | Cognitive impairment, brain abnormality | ANKRD11 ZNF778 | Microdeletion | [160] |
| 17q12 | Macrocephaly, neurocognitive impairment | HNF1B | Deletion | [172] |
| 20q13.12 | Releases of glutamate at the synapse | RIMS4 | [166,173] | |
| 22q11.2 | Physical, behavioral, social communication, neurocognitive impairments | Deletion | [166] | |
| 22q13 | Cognitive deficits, behavioral autistic symptoms, language and social communication problems | SHANK3 | Deletion | [96,97,174,175,176] |
| Xq27.3 | Synaptic function in the brain | FMR1 | [166,177,178] |
| Epigenetic Factors | Genes | Function | Possible Epigenetic Mechanisms | Reference |
|---|---|---|---|---|
| DNA methylation | MeCP2 | Encodes a methyl binding protein that binds to the methylated region of DNA and silence the gene. Has a role in synaptic development and long-term synaptic plasticity. | MeCP2 regulation of other genes via epigenetics: recruitment of co-repressors, chromatin looping. | [184,193] |
| UBE3A | Known for its role in Angelman syndrome. | Loss of imprinting of one copy, and production of antisense RNA that binds to UBE3A and mRNA Prevents translation. | [185,186] | |
| OXTR | G-protein coupled receptor for oxytocin. Modulates: stress, anxiety, social memory, maternal-offspring behavior, etc. | Hypermethylation and silencing.Decreased OXTR expression. | [194,195] | |
| SHANK3 | Effect on the morphology of dendritic spine and synaptic transmission | Expression of SHANK3 was strongly regulated by methylated CpG island. | [195,196] | |
| Histone modification | KDM5C | Alters the epigenetic state, which is associated with intellectual disability and frequent autistic behavior. | Involved in the regulation of transcription and chromatin remodeling. | [197] |
| HIST1H1E | Associated with the features of ASD and intellectual disorders. | To organize the higher-order chromatin structure and regulation of gene transcription. | [198] | |
| CHD8 | Inhibit the target genes of Wnt/β-catenin, and many of the genes in CHD8 targets included autism risk genes. | Encode ATP-dependent helicases that are typically involved in chromatin remodeling. | [199] | |
| ARID1B | A component of the ATP-dependent human SWI/SNF chromatin-remodeling complex. | Involved in chromatin remodeling. | [15,200] | |
| BCL11A | Encode proteins that interact directly with members of the SWI/SNF. | Involved in chromatin remodeling. | [201] | |
| ADNP | Encode proteins that interact directly with members of the SWI/SNF. | Involved in chromatin remodeling. | [201] | |
| Micro RNA | Deregulation of miRNA synthesis leads to neurodevelopmental disorders. | Epigenetic regulator that control the expression of many genes at the level of post-transcription by blocking protein synthesis or mRNA degradation. | [202,203,204,205] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yoon, S.H.; Choi, J.; Lee, W.J.; Do, J.T. Genetic and Epigenetic Etiology Underlying Autism Spectrum Disorder. J. Clin. Med. 2020, 9, 966. https://doi.org/10.3390/jcm9040966
Yoon SH, Choi J, Lee WJ, Do JT. Genetic and Epigenetic Etiology Underlying Autism Spectrum Disorder. Journal of Clinical Medicine. 2020; 9(4):966. https://doi.org/10.3390/jcm9040966
Chicago/Turabian StyleYoon, Sang Hoon, Joonhyuk Choi, Won Ji Lee, and Jeong Tae Do. 2020. "Genetic and Epigenetic Etiology Underlying Autism Spectrum Disorder" Journal of Clinical Medicine 9, no. 4: 966. https://doi.org/10.3390/jcm9040966
APA StyleYoon, S. H., Choi, J., Lee, W. J., & Do, J. T. (2020). Genetic and Epigenetic Etiology Underlying Autism Spectrum Disorder. Journal of Clinical Medicine, 9(4), 966. https://doi.org/10.3390/jcm9040966





