p53 CRISPR Deletion Affects DNA Structure and Nuclear Architecture
Abstract
1. Introduction
2. Materials and Methods
2.1. Tissue Culture and Nutlin-3 and RITA Treatments
2.2. Western Blot Analysis
2.3. Three-Dimensional Structured Illumination Microsopy (3D-SIM) Slide Preparation
2.4. Three-Dimensional Quantitative Fluorescent in situ Hybridization (3D Q-FISH)
2.5. 3D-SIM Imaging and Analysis
2.6. Widefield 3D Imaging and Analysis
2.7. Metaphase Spread Preparation
2.8. Spectral Karyotyping (SKY) Slide Preparation and Imaging
3. Results
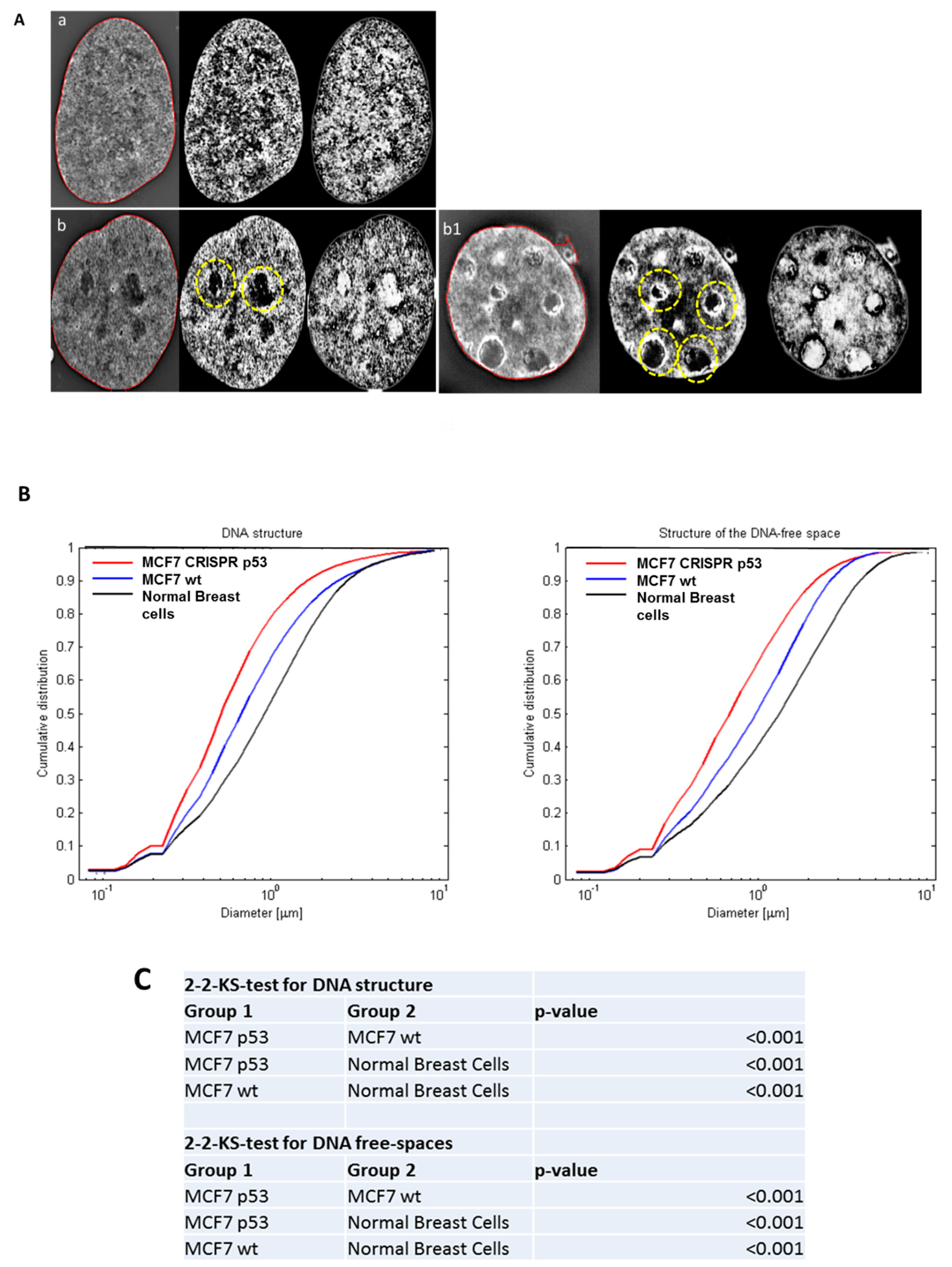
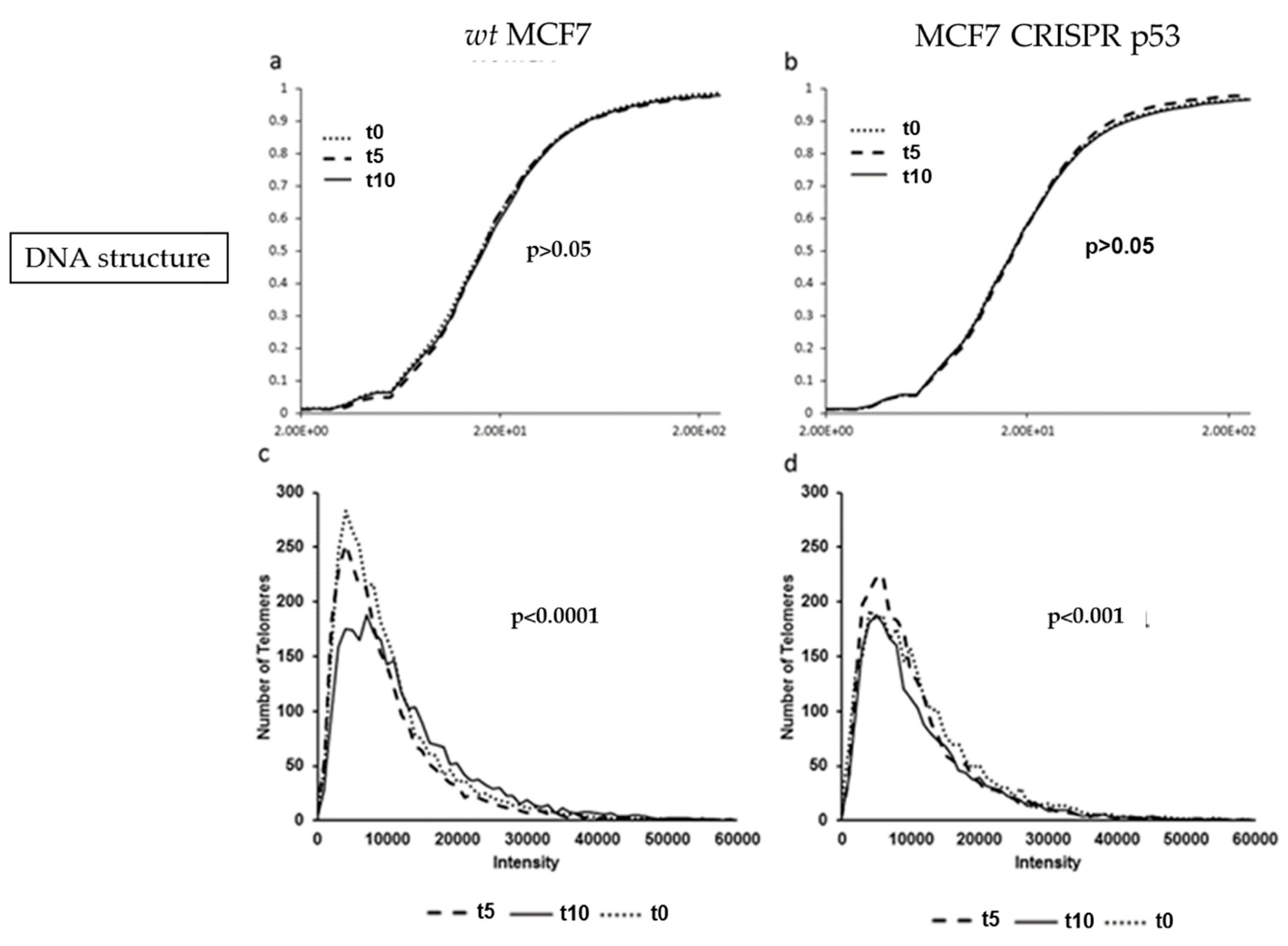
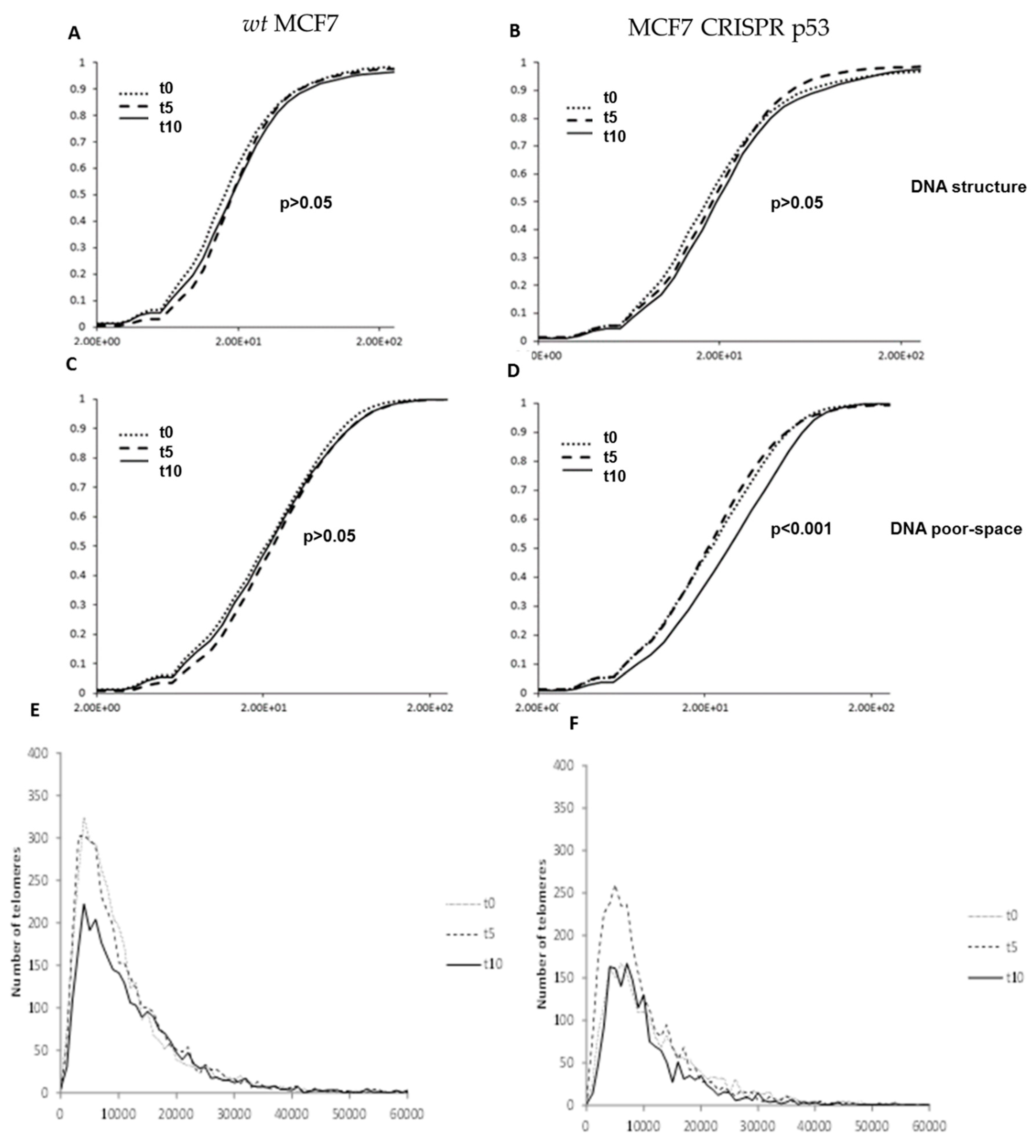
3.1. Analysis of DNA Sstructure Reveals Differences in DNA Structure and Presence of DNA Poor Spaces Between Wild-Type p53 and an Isogenic CRISPR p53 Deleted Cell Line
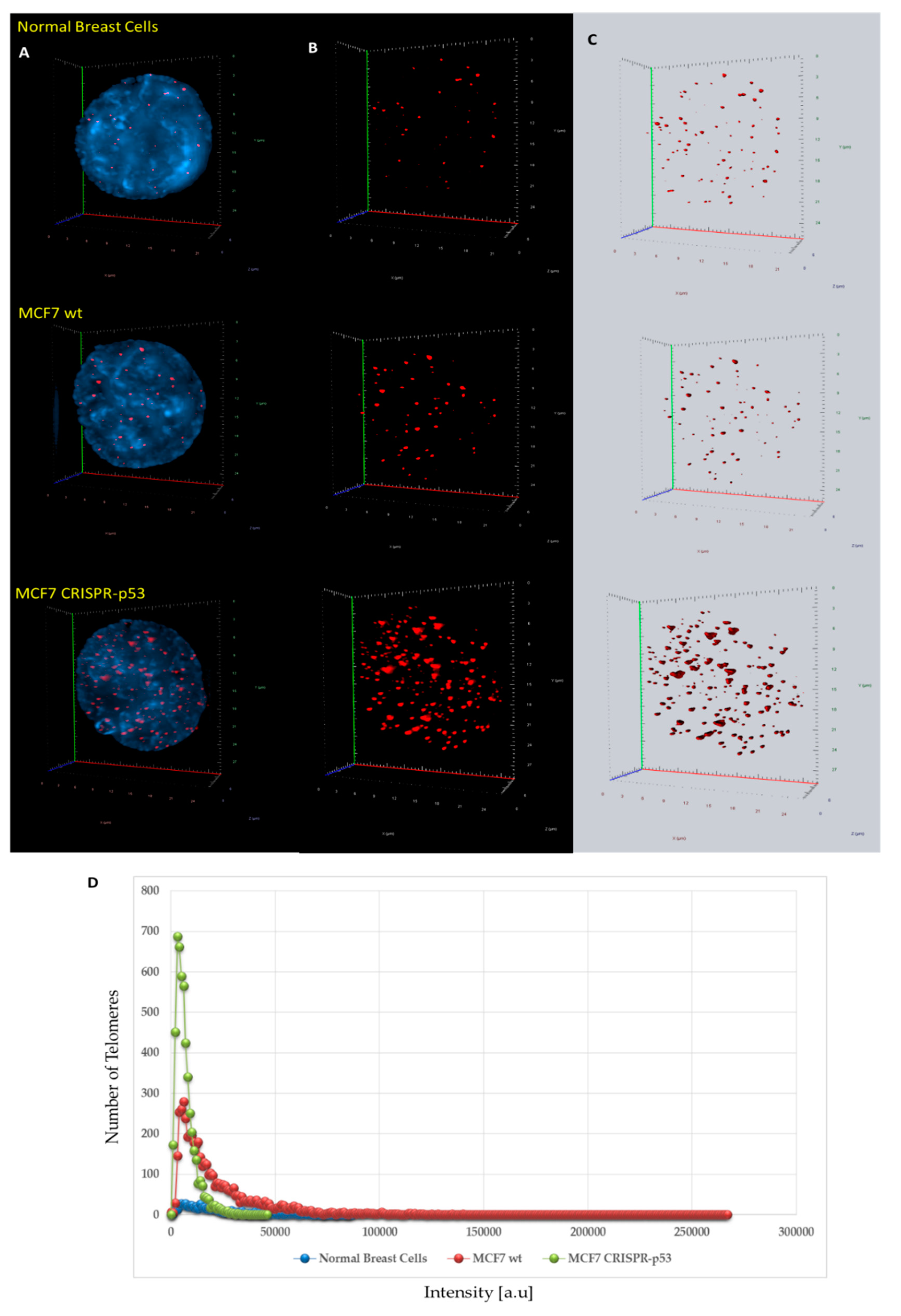
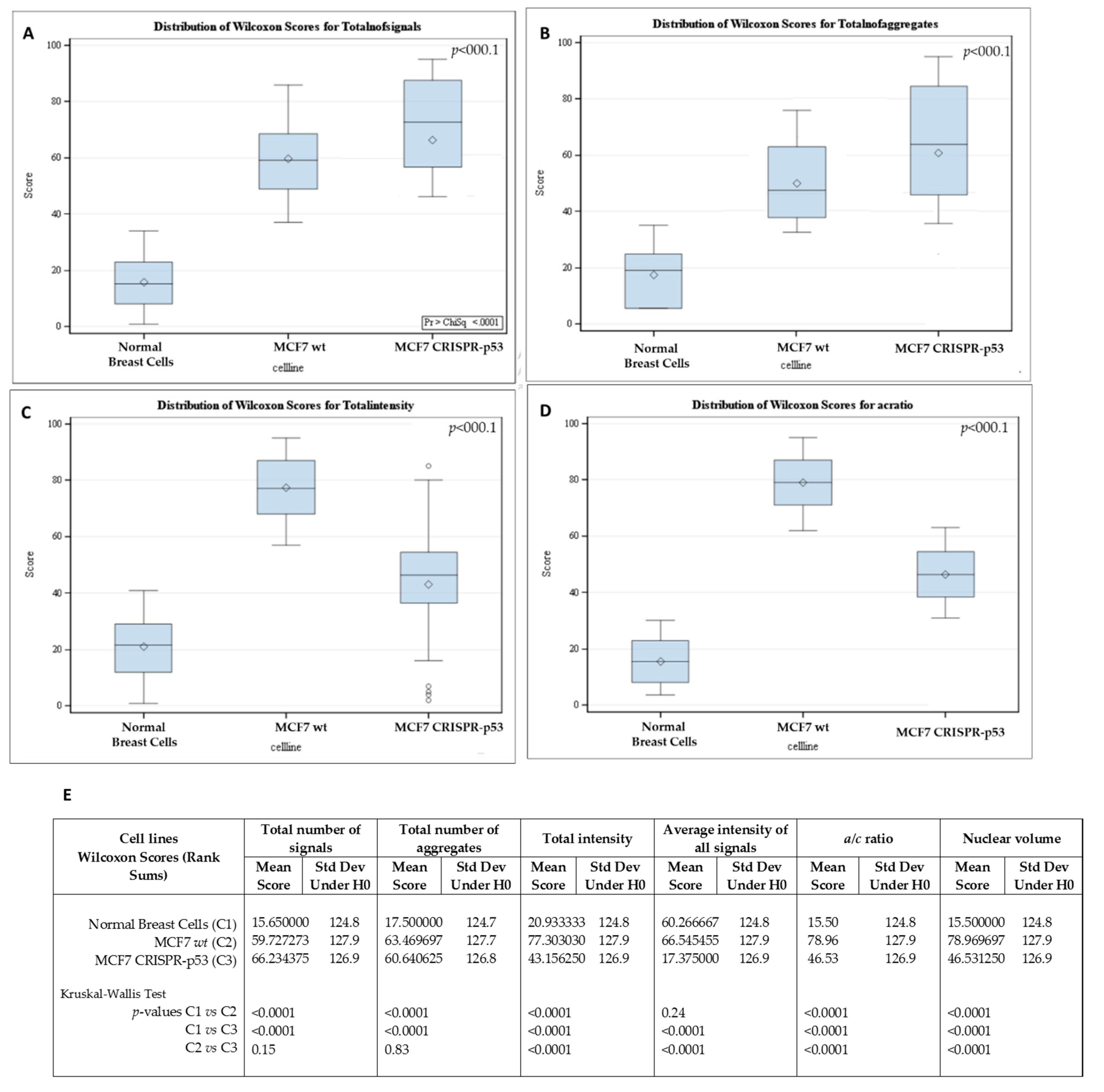
3.2. Analysis of 3D Nuclear Telomere Organization Shows Increase of Genomic Instability between Wild-Type p53 and Isogenic CRISPR p53 Deleted Cell Line
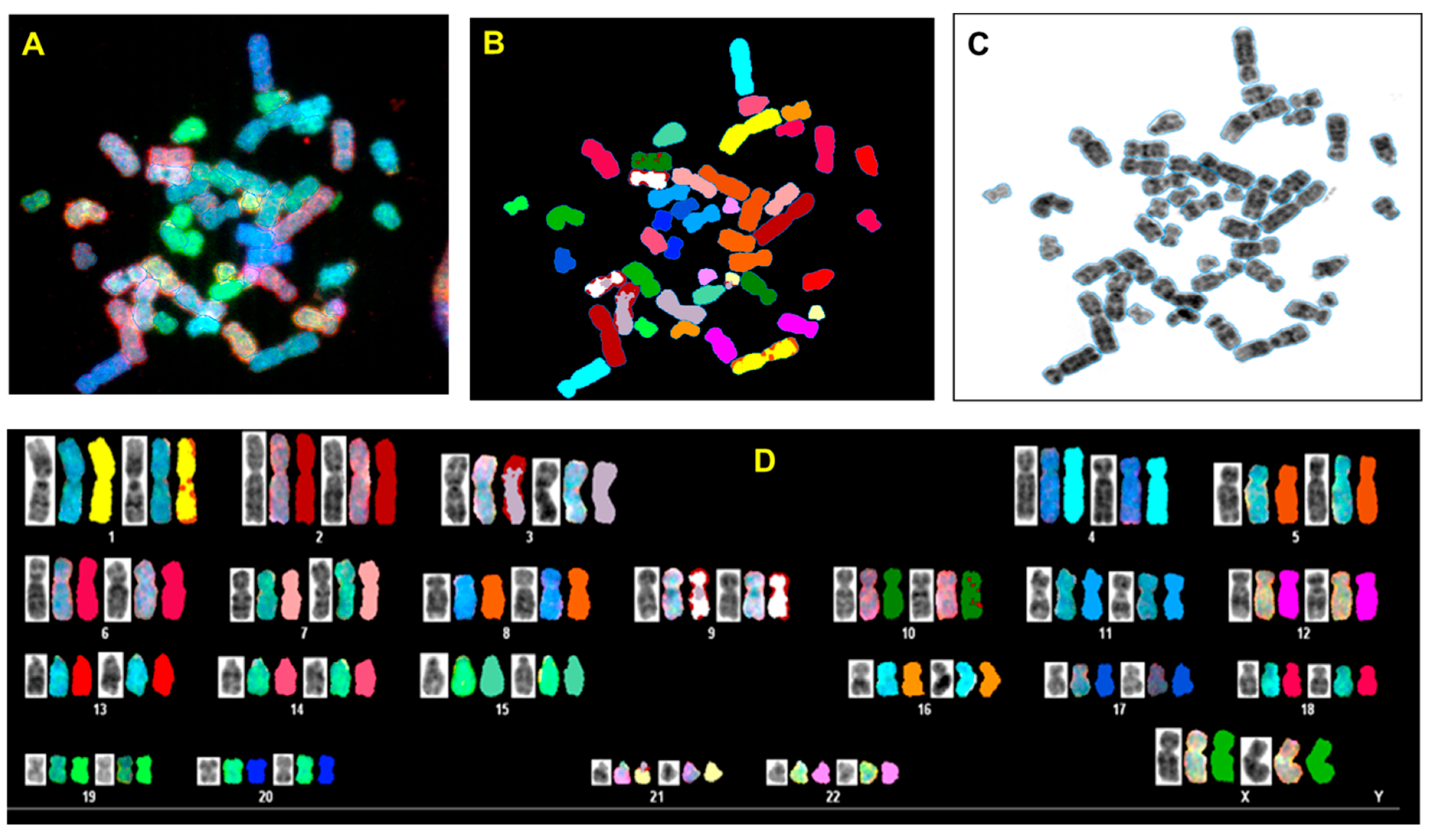
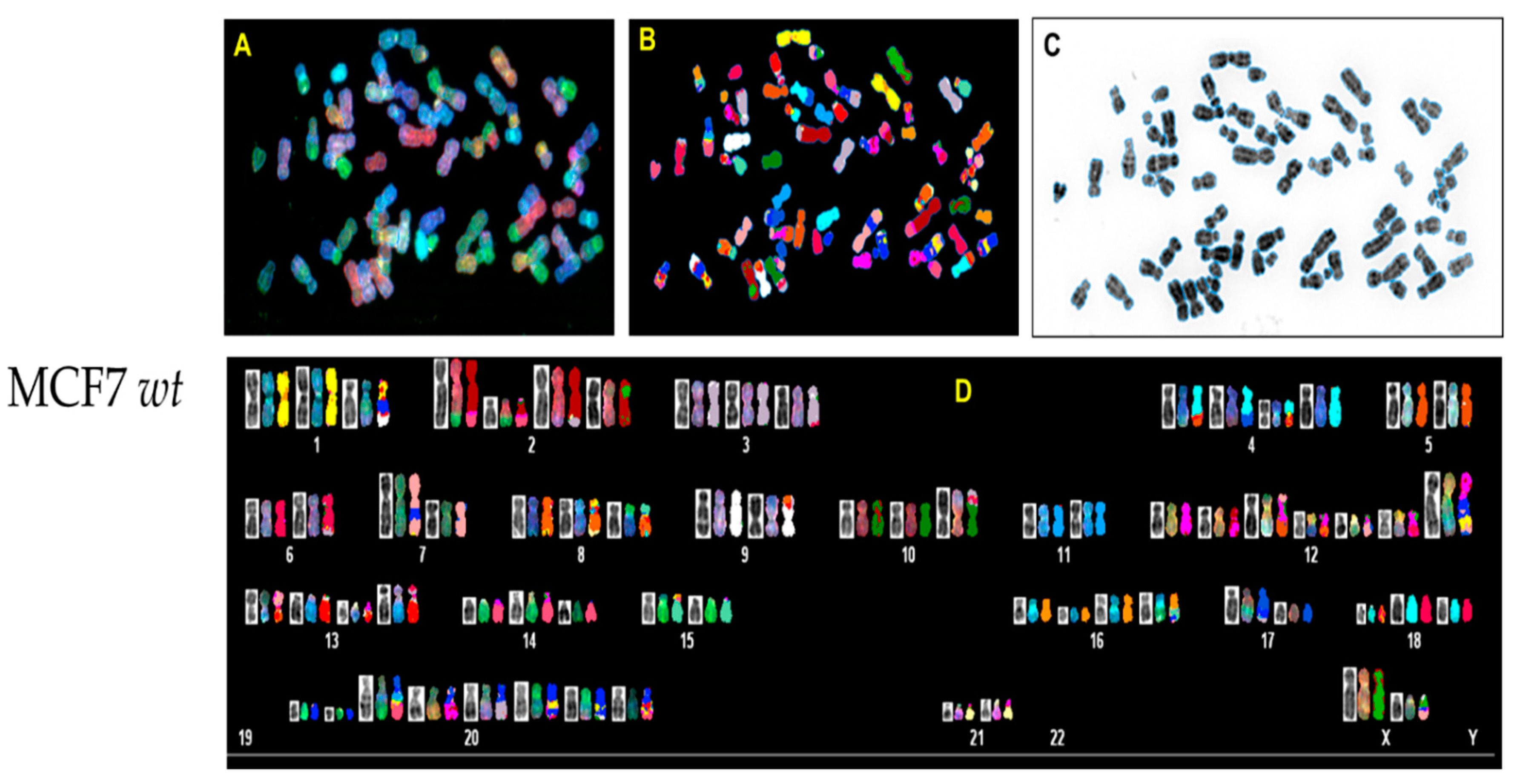
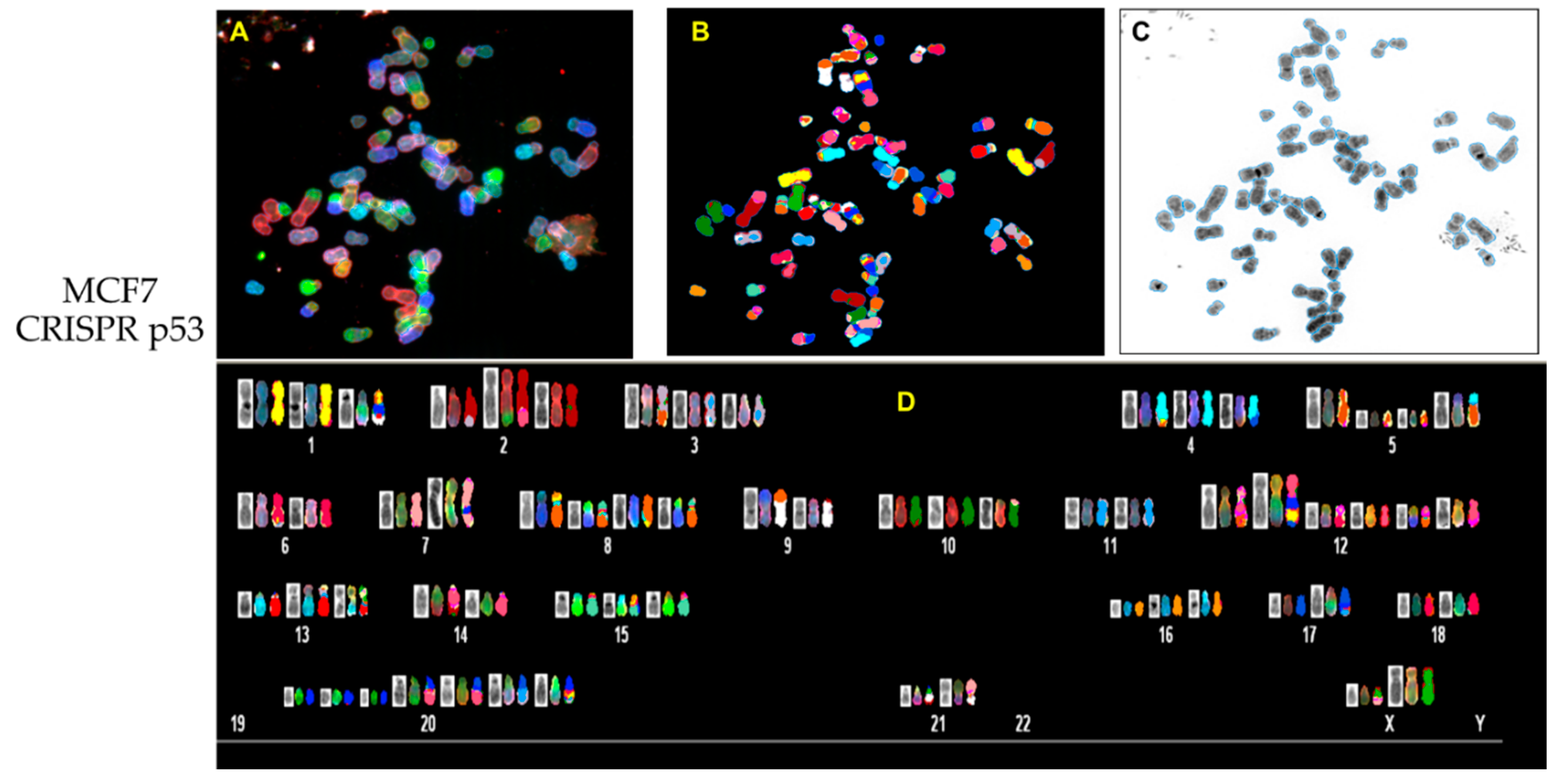
3.3. p53 CRISPR Deletion Increase Chromossomal Instability in MCF7
3.4. Nutlin-3 Affects Telomeric Nuclear Architecture But Not Super Resolution DNA Structure
3.5. RITA affects nuclear DNA structure
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Brázda, V.; Fojta, M. The Rich World of p53 DNA Binding Targets: The Role of DNA Structure. Int. J. Mol. Sci. 2019, 20, 5605. [Google Scholar] [CrossRef]
- Vousden, K.H.; Lane, D.P. p53 in health and disease. Nat. Rev. Mol. Cell Biol. 2007, 8, 275–283. [Google Scholar] [CrossRef]
- Gohler, T.; Reimann, M.; Cherny, D.; Walter, K.; Warnecke, G.; Kim, E.; Deppert, W. Specific interaction of p53 with target binding sites is determined by DNA conformation and is regulated by the C-terminal domain. J. Biol. Chem. 2002, 277, 41192–41203. [Google Scholar] [CrossRef] [PubMed]
- Meek, D.W. Regulation of the p53 response and its relationship to cancer. Biochem. J. 2015, 469, 325–346. [Google Scholar] [CrossRef] [PubMed]
- Chen, J. The Cell-Cycle Arrest and Apoptotic Functions of p53 in Tumor Initiation and Progression. Cold Spring Harb. Perspect. Med. 2016, 6, a026104. [Google Scholar] [CrossRef] [PubMed]
- Kaiser, A.M.; Attardi, L.D. Deconstructing networks of p53-mediated tumor suppression in vivo. Cell Death Differ. 2018, 25, 93–103. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Simpson, E.R.; Brown, K.A. p53: Protection against Tumor Growth beyond Effects on Cell Cycle and Apoptosis. Cancer Res. 2015, 75, 5001–5007. [Google Scholar] [CrossRef] [PubMed]
- Ranjan, A.; Iwakuma, T. Non-Canonical Cell Death Induced by p53. Int. J. Mol. Sci. 2016, 17, 2068. [Google Scholar] [CrossRef] [PubMed]
- Pfaff, M.J.; Mukhopadhyay, S.; Hoofnagle, M.; Chabasse, C.; Sarkar, R. Tumor suppressor protein p53 negatively regulates ischemia-induced angiogenesis and arteriogenesis. J. Vasc. Surg. 2018, 68, 222S–233S.e1. [Google Scholar] [CrossRef]
- Chandrangsu, S.; Sappayatosok, K. p53, p63 and p73 expression and angiogenesis in keratocystic odontogenic tumors. J. Clin. Exp. Dent. 2016, 8, e505–e511. [Google Scholar] [CrossRef]
- Williams, A.B.; Schumacher, B. p53 in the DNA-Damage-Repair Process. Cold Spring Harb. Perspect. Med. 2016, 6, a026070. [Google Scholar] [CrossRef] [PubMed]
- Nicolai, S.; Rossi, A.; Di Daniele, N.; Melino, G.; Annicchiarico-Petruzzelli, M.; Raschellà, G. DNA repair and aging: The impact of the p53 family. Aging 2015, 7, 1050–1065. [Google Scholar] [CrossRef] [PubMed]
- Rufini, A.; Tucci, P.; Celardo, I.; Melino, G. Senescence and aging: The critical roles of p53. Oncogene 2013, 2013. 32, 5129–5143. [Google Scholar] [CrossRef]
- Vogelstein, B.; Lane, D.; Levine, A.J. Surfing the p53 network. Nature 2000, 408, 307–310. [Google Scholar] [CrossRef] [PubMed]
- Mihara, M.; Erster, S.; Zaika, A.; Petrenko, O.; Chittenden, T.; Pancoska, P.; Moll, U.M. p53 has a direct apoptogenic role at the mitochondria. Mol. Cell 2003, 11, 577–590. [Google Scholar] [CrossRef]
- Vousden, K.H.; Prives, C. Blinded by the Light: The Growing Complexity of p53. Cell 2009, 137, 413–431. [Google Scholar] [CrossRef]
- Assadian, S.; El-Assaad, W.; Wang, X.Q.D.; Gannon, P.O.; Barrès, V.; Latour, M.; Mes-Masson, A.M.; Saad, F.; Sado, Y.; Dostie, J.; et al. p53 inhibits angiogenesis by inducing the production of Arresten. Cancer Res. 2012, 72, 1270–1279. [Google Scholar] [CrossRef]
- Hollstein, M.; Sidransky, D.; Vogelstein, B.; Harris, C.C. p53 mutations in human cancers. Science 1991, 253, 49–53. [Google Scholar] [CrossRef] [PubMed]
- Leroy, B.; Fournier, J.L.; Ishioka, C.; Monti, P.; Inga, A.; Fronza, G.; Soussi, T. The TP53 website: An integrative resource centre for the TP53 mutation database and TP53 mutant analysis. Nucleic Acids Res. 2013, 41, 962–969. [Google Scholar] [CrossRef]
- Momand, J.; Wu, H.H.; Dasgupta, G. MDM2--master regulator of the p53 tumor suppressor protein. Gene 2000, 242, 15–29. [Google Scholar] [CrossRef]
- Zawacka-Pankau, J.; Selivanova, G. Pharmacological reactivation of p53 as a strategy to treat cancer. J. Intern. Med. 2015, 277, 248–259. [Google Scholar] [CrossRef] [PubMed]
- Vassilev, L.T.; Vu, B.T.; Graves, B.; Carvajal, D.; Podlaski, F.; Filipovic, Z.; Kong, N.; Kammlott, U.; Lukacs, C.; Klein, C.; et al. In vivo activation of the p53 pathway by small-molecule antagonists of MDM2. Science 2004, 303, 844–848. [Google Scholar] [CrossRef]
- Issaeva, N.; Bozko, P.; Enge, M.; Protopopova, M.; Verhoef, L.G.G.C.; Masucci, M.; Pramanik, A.; Selivanova, G. Small molecule RITA binds to p53, blocks p53-HDM-2 interaction and activates p53 function in tumors. Nat. Med. 2004, 10, 1321–1328. [Google Scholar] [CrossRef] [PubMed]
- Enge, M.; Bao, W.; Hedström, E.; Jackson, S.P.; Moumen, A.; Selivanova, G. MDM2-Dependent Downregulation of p21 and hnRNP K Provides a Switch between Apoptosis and Growth Arrest Induced by Pharmacologically Activated p53. Cancer Cell 2009, 15, 171–183. [Google Scholar] [CrossRef] [PubMed]
- Weilbacher, A.; Gutekunst, M.; Oren, M.; Aulitzky, W.E.; van der Kuip, H. RITA can induce cell death in p53-defective cells independently of p53 function via activation of JNK/SAPK and p38. Cell Death Dis. 2014, 5, e1318. [Google Scholar] [CrossRef] [PubMed]
- Gadji, M.; Fortin, D.; Tsanaclis, A.-M.; Garini, Y.; Katzir, N.; Wienburg, Y.; Yan, J.; Klewes, L.; Klonisch, T.; Drouin, R.; et al. Three-dimensional Nuclear Telomere Architecture Is Associated with Differential Time to Progression and Overall Survival in Glioblastoma Patients. Neoplasia 2010, 12, 183–191. [Google Scholar] [CrossRef]
- Wark, L.; Quon, H.; Ong, A.; Drachenberg, D.; Rangel-Pozzo, A.; Mai, S. Long-Term Dynamics of Three Dimensional Telomere Profiles in Circulating Tumor Cells in High-Risk Prostate Cancer Patients Undergoing Androgen-Deprivation and Radiation Therapy. Cancers 2019, 14, 11. [Google Scholar] [CrossRef]
- Drachenberg, D.; Awe, J.A.; Rangel-Pozzo, A.; Saranchuk, J.; Mai, S. Advancing Risk Assessment of Intermediate Risk Prostate Cancer Patients. Cancers 2019, 11, 855. [Google Scholar] [CrossRef]
- Rangel-Pozzo, A.; Corrêa de Souza, D.; Schmid-Braz, A.T.; de Azambuja, A.P.; Ferraz-Aguiar, T.; Borgonovo, T.; Mai, S. 3D Telomere Structure Analysis to Detect Genomic Instability and Cytogenetic Evolution in Myelodysplastic Syndromes. Cells 2019, 8, 304. [Google Scholar] [CrossRef]
- Contu, F.; Rangel-Pozzo, A.; Trokajlo, P.; Wark, L.; Klewes, L.; Johnson, N.A.; Petrogiannis-Haliotis, T.; Gartner, J.G.; Garini, Y.; Vanni, R.; et al. Distinct 3D Structural Patterns of Lamin A/C Expression in Hodgkin and Reed-Sternberg Cells. Cancers 2018, 10, 286. [Google Scholar] [CrossRef]
- Zink, D.; Fischer, A.H.; Nickerson, J.A. Nuclear structure in cancer cells. Nat. Rev. Cancer 2004, 4, 677–687. [Google Scholar] [CrossRef] [PubMed]
- Rubio, K.; Dobersch, S.; Barreto, G. Functional interactions between scaffold proteins, noncoding RNAs, and genome loci induce liquid-liquid phase separation as organzing principle for 3-dimensional nuclear architecture: Implications in cancer. FASEB J. 2019, 33, 5814–5822. [Google Scholar] [CrossRef] [PubMed]
- Uhler, C.; Shivashankar, G.V. Regulation of genome organization and gene expression by nuclear mechanotransduction. Nat. Rev. Mol. Cell Biol. 2017, 18, 717–727. [Google Scholar] [CrossRef] [PubMed]
- Cremer, T.; Cremer, C. Chromosome territories, nuclear architecture and gene regulation in mammalian cells. Nat. Rev. Genet. 2001, 2, 292–301. [Google Scholar] [CrossRef] [PubMed]
- Albiez, H.; Cremer, M.; Tiberi, C.; Vecchio, L.; Schermelleh, L.; Dittrich, S.; Küpper, K.; Joffe, B.; Thormeyer, T.; von Hase, J.; et al. Chromatin domains and the interchromatin compartment form structurally defined and functionally interacting nuclear networks. Chromosome Res. 2006, 14, 707–733. [Google Scholar] [CrossRef] [PubMed]
- Wei, G.H.; Liu, D.P.; Liang, C.C. Chromatin domain boundaries: Insulators and beyond. Cell Res. 2005, 15, 292–300. [Google Scholar] [CrossRef]
- de Wit, E.; de Laat, W. A decade of 3C technologies : Insights into nuclear organization. Genes Dev. 2012, 26, 11–24. [Google Scholar] [CrossRef]
- Kreth, G.; Finsterle, J.; von Hase, J.; Cremer, M.; Cremer, C. Radial arrangement of chromosome territories in human cell nuclei: A computer model approach based on gene density indicates a probabilistic global positioning code. Biophys. J. 2004, 86, 2803–2812. [Google Scholar] [CrossRef]
- Misteli, T.; Soutoglou, E. The emerging role of nuclear architecture in DNA repair and genome maintenance. Nat. Rev. Mol. Cell Biol. 2009, 10, 243–254. [Google Scholar] [CrossRef]
- Lever, E.; Sheer, D. The role of nuclear organization in cancer. J. Pathol. 2010, 220, 114–125. [Google Scholar] [CrossRef]
- Louis, S.F.; Vermolen, B.J.; Garini, Y.; Young, I.T.; Guffei, A.; Lichtensztejn, Z.; Kuttler, F.; Chuang, T.C.Y.; Moshir, S.; Mougey, V.; et al. c-Myc induces chromosomal rearrangements through telomere and chromosome remodeling in the interphase nucleus. Proc. Natl. Acad. Sci. USA 2005, 102, 9613–9618. [Google Scholar] [CrossRef]
- Mai, S.; Garini, Y. The significance of telomeric aggregates in the interphase nuclei of tumor cells. J. Cell Biochem. 2006, 97, 904–915. [Google Scholar] [CrossRef]
- Mai, S.; Garini, Y. Oncogenic remodeling of the three-dimensional organization of the interphase nucleus: C-Myc induces telomeric aggregates whose formation precedes chromosomal rearrangements. Cell Cycle 2005, 4, 1327. [Google Scholar] [CrossRef]
- Kuzyk, A.; Gartner, J.; Mai, S. Identification of Neuroblastoma Subgroups Based on Three-Dimensional Telomere Organization. Transl. Oncol. 2016, 9, 348–356. [Google Scholar] [CrossRef] [PubMed]
- Gadji, M.; Adebayo Awe, J.; Rodrigues, P.; Kumar, R.; Houston, D.S.; Klewes, L.; Dièye, T.N.; Rego, E.M.; Passetto, R.F.; de Oliveira, F.M.; et al. Profiling three-dimensional nuclear telomeric architecture of myelodysplastic syndromes and acute myeloid leukemia defines patient subgroups. Clin. Cancer Res. 2012, 18, 3293. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; O’Connor, M.S.; Qin, J.; Songyang, Z. Telosome, a mammalian telomere-associated complex formed by multiple telomeric proteins. J. Biol. Chem. 2004, 279, 51338–51342. [Google Scholar] [CrossRef] [PubMed]
- O’Sullivan, R.J.; Karlseder, J. Telomeres: Protecting chromosomes against genome instability. Nat. Rev. Mol. Cell Biol. 2010, 11, 171–181. [Google Scholar] [CrossRef] [PubMed]
- Vermolen, B.J.; Garini, Y.; Mai, S.; Mougey, V.; Fest, T.; Chuang, T.C.Y.; Chuang, Y.C.; Wark, L.; Young, I.T. Characterizing the three-dimensional organization of telomeres. Cytom. Part. A 2005, 67, 144–150. [Google Scholar] [CrossRef] [PubMed]
- Poon, S.S.; Martens, U.M.; Ward, R.K.; Lansdorp, P.M. Telomere length measurements using digital fluorescence microscopy. Cytometry 1999, 36, 267–278. [Google Scholar] [CrossRef]
- Chuang, T.C.Y.; Moshir, S.; Garini, Y.; Chuang, A.Y.; Young, I.T.; Vermolen, B.; van den Doel, R.; Mougey, V.; Perrin, M.; Braun, M.; et al. The three-dimensional organization of telomeres in the nucleus of mammalian cells. BMC Biol. 2004, 2, 12. [Google Scholar] [CrossRef]
- Ciriello, G.; Miller, M.L.; Aksoy, B.A.; Senbabaoglu, Y.; Schultz, N.; Sander, C. Emerging landscape of oncogenic signatures across human cancers. Nat. Genet. 2013, 45, 1127–1133. [Google Scholar] [CrossRef]
- Samassekou, O.; Bastien, N.; Lichtensztejn, D.; Yan, J.; Mai, S.; Drouin, R. Different TP53 mutations are associated with specific chromosomal rearrangements, telomere length changes, and remodeling of the nuclear architecture of telomeres. Genes Chromosome Cancer 2014, 53, 934–950. [Google Scholar] [CrossRef]
- Sullivan, K.D.; Galbraith, M.D.; Andrysik, Z.; Espinosa, J.M. Mechanisms of transcriptional regulation by p53. Cell Death Differ. 2018, 25, 133–143. [Google Scholar] [CrossRef]
- Link, N.; Kurtz, P.; O’Neal, M.; Garcia-Hughes, G.; Abrams, J.M. A p53 enhancer region regulates target genes through chromatin conformations in cis and in trans. Genes Dev. 2013, 27, 2433–2438. [Google Scholar] [CrossRef]
- Melo, C.A.; Drost, J.; Wijchers, P.J.; van de Werken, H.; de Wit, E.; Oude Vrielink, J.A.; Elkon, R.; Melo, S.A.; Léveillé, N.; Kalluri, R.; et al. eRNAs are required for p53-dependent enhancer activity and gene transcription. Mol. Cell 2013, 49, 524–535. [Google Scholar] [CrossRef]
- Schermelleh, L.; Carlton, P.M.; Haase, S.; Shao, L.; Kner, P.; Burke, B.; Cardoso, M.C.; Agard, D.A.; Gustafsson, M.G.; Leonhardt, H.; et al. Subdiffraction Multicolor Imaging of the Nuclear Periphery with 3D Structured Illumination Microscopy. Science 2008, 320, 1332–1336. [Google Scholar] [CrossRef]
- Gustafsson, M.G. Surpassing the lateral resolution limit by a factor of two using structured illumination microscopy. J. Microsc. 2000, 198, 82–87. [Google Scholar] [CrossRef]
- Rangel-Pozzo, A.; Kuzyk, A.; Gartner, J.G.; Mai, S. MYCN overexpression is linked to significant differences in nuclear DNA organization in neuroblastoma. SPG Biomed. 2019. [Google Scholar] [CrossRef]
- Righolt, C.H.; Guffei, A.; Knecht, H.; Young, I.T.; Stallinga, S.; Van Vliet, L.J.; Mai, S. Differences in nuclear DNA organization between lymphocytes, hodgkin and reed-sternberg cells revealed by structured illumination microscopy. J. Cell Biochem. 2014, 115, 1441–1448. [Google Scholar] [CrossRef] [PubMed]
- Sathitruangsak, C.; Righolt, C.H.; Klewes, L.; Tammur, P.; Ilus, T.; Tamm, A.; Punab, M.; Olujohungbe, A.; Mai, S. Quantitative Superresolution Microscopy Reveals Differences in Nuclear DNA Organization of Multiple Myeloma and Monoclonal Gammopathy of Undetermined Significance. J. Cell Biochem. 2015, 116, 704–710. [Google Scholar] [CrossRef] [PubMed]
- Cong, L.; Ran, F.A.; Cox, D.; Lin, S.; Barretto, R.; Habib, N.; Hsu, P.D.; Wu, X.; Jiang, W.; Marraffini, L.A.; et al. Multiplex genome engineering using CRISPR/Cas systems. Science 2013, 339, 819–823. [Google Scholar] [CrossRef] [PubMed]
- Zetsche, B.; Heidenreich, M.; Mohanraju, P.; Fedorova, I.; Kneppers, J.; DeGennaro, E.M.; Winblad, N.; Choudhury, S.R.; Abudayyeh, O.O.; Gootenberg, J.S.; et al. Multiplex gene editing by CRISPR-Cpf1 using a single crRNA array. Nat. Biotechnol. 2017, 35, 31–34. [Google Scholar] [CrossRef]
- Liyanage, M.; Coleman, A.; du Manoir, S.; Veldman, T.; McCormack, S.; Dickson, R.B.; Barlow, C.; Wynshaw-Boris, A.; Janz, S.; Wienberg, J.; et al. Multicolour spectral karyotyping of mouse chromosomes. Nat. Genet. 1996, 14, 312–315. [Google Scholar] [CrossRef] [PubMed]
- Garini, Y.; Macville, M.; du Manoir, S.; Buckwald, R.A.; Lavi, M.; Katzir, N.; Wine, D.; Bar-Am, I.; Schro ̈ ck, E.; Cabib, D.; et al. Spectralkaryotyping. Bioimaging 1996, 1996 4, 65–72. [Google Scholar] [CrossRef]
- Fields, S.; Jang, S.K. Presence of a potent transcription activating sequence in the p53 protein. Science 1990, 249, 1046–1049. [Google Scholar] [CrossRef] [PubMed]
- Raycroft, L.; Wu, H.Y.; Lozano, G. Transcriptional activation by wild-type but not transforming mutants of the p53 anti-oncogene. Science 1990, 249, 1049–1051. [Google Scholar] [CrossRef] [PubMed]
- Kern, S.E.; Kinzler, K.W.; Bruskin, A.; Jarosz, D.; Friedman, P.; Prives, C.; Vogelstein, B. Identification of p53 as a sequence-specific DNA-binding protein. Science 1991, 252, 1708–1711. [Google Scholar] [CrossRef] [PubMed]
- Farmer, G.; Bargonetti, J.; Zhu, H.; Friedman, P.; Prywes, R.; Prives, C. Wild-type p53 activates transcription in vitro. Nature 1992, 358, 83–86. [Google Scholar] [CrossRef] [PubMed]
- Dixon, J.R.; Gorkin, D.U. Chromatin Domains: The Unit of Chromosome Organization. Mol. Cell 2016, 62, 668–680. [Google Scholar] [CrossRef]
- Rickman, D.S.; Soong, T.D.; Moss, B.; Mosquera, J.M.; Dlabal, J.; Terry, S.; MacDonald, T.Y.; Tripodi, J.; Bunting, K.; Najfeld, V.; et al. Oncogene-mediated alterations in chromatin conformation. Proc. Natl. Acad. Sci. USA 2012, 109, 9083–9088. [Google Scholar] [CrossRef]
- Wood, L.J.; Mukherjee, M.; Dolde, C.E.; Xu, Y.; Maher, J.F.; Bunton, T.E.; Williams, J.B.; Resar, L.M. HMG-I/Y, a new c-Myc target gene and potential oncogene. Mol. Cell Biol. 2000, 20, 5490–5502. [Google Scholar] [CrossRef]
- Vogel, B.; Löschberger, A.; Sauer, M.; Hock, R. Cross-linking of DNA through HMGA1 suggests a DNA scaffold. Nucleic Acids Res. 2011, 39, 7124–7133. [Google Scholar] [CrossRef] [PubMed]
- Falvo, J.V.; Thanos, D.; Maniatis, T. Reversal of intrinsic DNA bends in the IFNβ gene enhancer by transcription factors and the architectural protein HMG I(Y). Cell 1995, 83, 1101–1111. [Google Scholar] [CrossRef]
- Singh, D.; Rajeswari, M.R. Enhanced expression of HMG-Y proteins in proliferating tissues. Indian J. Clin. Biochem. 2001, 16, 72–76. [Google Scholar] [CrossRef] [PubMed]
- Dolde, C.E.; Mukherjee, M.; Cho, C.; Resar, L.M.S. HMG-I/Y in human breast cancer cell lines. Breast Cancer Res. Treat. 2002, 71, 181–191. [Google Scholar] [CrossRef] [PubMed]
- Donizy, P.; Biecek, P.; Halon, A.; Maciejczyk, A.; Matkowski, R. Nucleoli cytomorphology in cutaneous melanoma cells—A new prognostic approach to an old concept. Diagn. Pathol. 2017, 12, 88. [Google Scholar] [CrossRef] [PubMed]
- Bártová, E.; Jirsová, P.; Fojtová, M.; Soucek, K.; Kozubek, S. Chromosomal territory segmentation in apoptotic cells. Cell Mol. Life Sci. 2003, 60, 979–990. [Google Scholar] [CrossRef]
- Roh, J.L.; Ko, J.H.; Moon, S.J.; Ryu, C.H.; Choi, J.Y.; Koch, W.M. The p53-reactivating small-molecule RITA enhances cisplatin-induced cytotoxicity and apoptosis in head and neck cancer. Cancer Lett. 2012, 325, 35–41. [Google Scholar] [CrossRef]
- Saha, M.N.; Jiang, H.; Yang, Y.; Zhu, X.; Wang, X.; Schimmer, A.D.; Qiu, L.; Chang, H. Targeting p53 via JNK pathway: A novel role of RITA for apoptotic signaling in multiple myeloma. PLoS ONE 2012, 7, 1–11. [Google Scholar] [CrossRef]
- Surget, S.; Descamps, G.; Brosseau, C.; Normant, V.; Maïga, S.; Gomez-Bougie, P.; Gouy-Colin, N.; Godon, C.; Béné, M.C.; Moreau, P.; et al. RITA (Reactivating p53 and Inducing Tumor Apoptosis) is efficient against TP53abnormal myeloma cells independently of the p53 pathway. BMC Cancer 2014, 14, 437. [Google Scholar] [CrossRef]
- Wanzel, M.; Vischedyk, J.B.; Gittler, M.P.; Gremke, N.; Seiz, J.R.; Hefter, M.; Noack, M.; Savai, R.; Mernberger, M.; Charles, J.P.; et al. CRISPR-Cas9-based target validation for p53-reactivating model compounds. Nat. Chem. Biol. 2016, 12, 22–28. [Google Scholar] [CrossRef] [PubMed]
- Ma, T.; Yamada, S.; Ichwan, S.J.A.; Iseki, S.; Ohtani, K.; Otsu, M.; Ikeda, M.A. Inability of p53-reactivating compounds nutlin-3 and rita to overcome p53 resistance in tumor cells deficient in p53ser46 phosphorylation. Biochem. Biophys. Res. Commun. 2012, 417, 931–9317. [Google Scholar] [CrossRef] [PubMed]
- Girardini, J.E.; Del Sal, G. Improving pharmacological rescue of p53 function: RITA targets mutant p53. Cell Cycle 2010, 9, 2062. [Google Scholar] [CrossRef] [PubMed]
- Spinnler, C.; Hedström, E.; Li, H.; de Lange, J.; Nikulenkov, F.; Teunisse, A.F.A.S.; Verlaan-de Vries, M.; Grinkevich, V.; Jochemsen, A.G.; Selivanova, G. Abrogation of Wip1 expression by RITA-activated p53 potentiates apoptosis induction via activation of ATM and inhibition of HdmX. Cell Death Differ. 2011, 18, 1736–1745. [Google Scholar] [CrossRef] [PubMed]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rangel-Pozzo, A.; Booth, S.; Yu, P.L.I.; Singh, M.; Selivanova, G.; Mai, S. p53 CRISPR Deletion Affects DNA Structure and Nuclear Architecture. J. Clin. Med. 2020, 9, 598. https://doi.org/10.3390/jcm9020598
Rangel-Pozzo A, Booth S, Yu PLI, Singh M, Selivanova G, Mai S. p53 CRISPR Deletion Affects DNA Structure and Nuclear Architecture. Journal of Clinical Medicine. 2020; 9(2):598. https://doi.org/10.3390/jcm9020598
Chicago/Turabian StyleRangel-Pozzo, Aline, Samuel Booth, Pak Lok Ivan Yu, Madhurendra Singh, Galina Selivanova, and Sabine Mai. 2020. "p53 CRISPR Deletion Affects DNA Structure and Nuclear Architecture" Journal of Clinical Medicine 9, no. 2: 598. https://doi.org/10.3390/jcm9020598
APA StyleRangel-Pozzo, A., Booth, S., Yu, P. L. I., Singh, M., Selivanova, G., & Mai, S. (2020). p53 CRISPR Deletion Affects DNA Structure and Nuclear Architecture. Journal of Clinical Medicine, 9(2), 598. https://doi.org/10.3390/jcm9020598




