Multiplexed Proteomic Approach for Identification of Serum Biomarkers in Hepatocellular Carcinoma Patients with Normal AFP
Abstract
1. Introduction
2. Methods
2.1. Patients and Diagnosis
2.2. Enzymatic In-Gel Digestion and Nano-LC-ESI-MS/MS Analysis
2.3. Database Searching
2.4. Target Protein Selection and LC-MRM/MS Assay
2.5. mRNA and Protein Expression of Trim22 in Tumor/Non-Tumor Tissues of HCC Patients
2.6. Western Blot Analysis of Trim22 Using Tumor/Non-Tumor Liver Tissues from HCC Patients and Other Malignancy Patients with Liver Metastasis
2.7. Western Blot Analysis of Serum Trim22 Expression in Hepatocellular Carcinoma Patients with Normal AFP
2.8. Statistical Analysis
3. Results
3.1. Baseline Characteristics
3.2. MRM/MALDI-TOF MS/MS Analysis of Serum Proteins
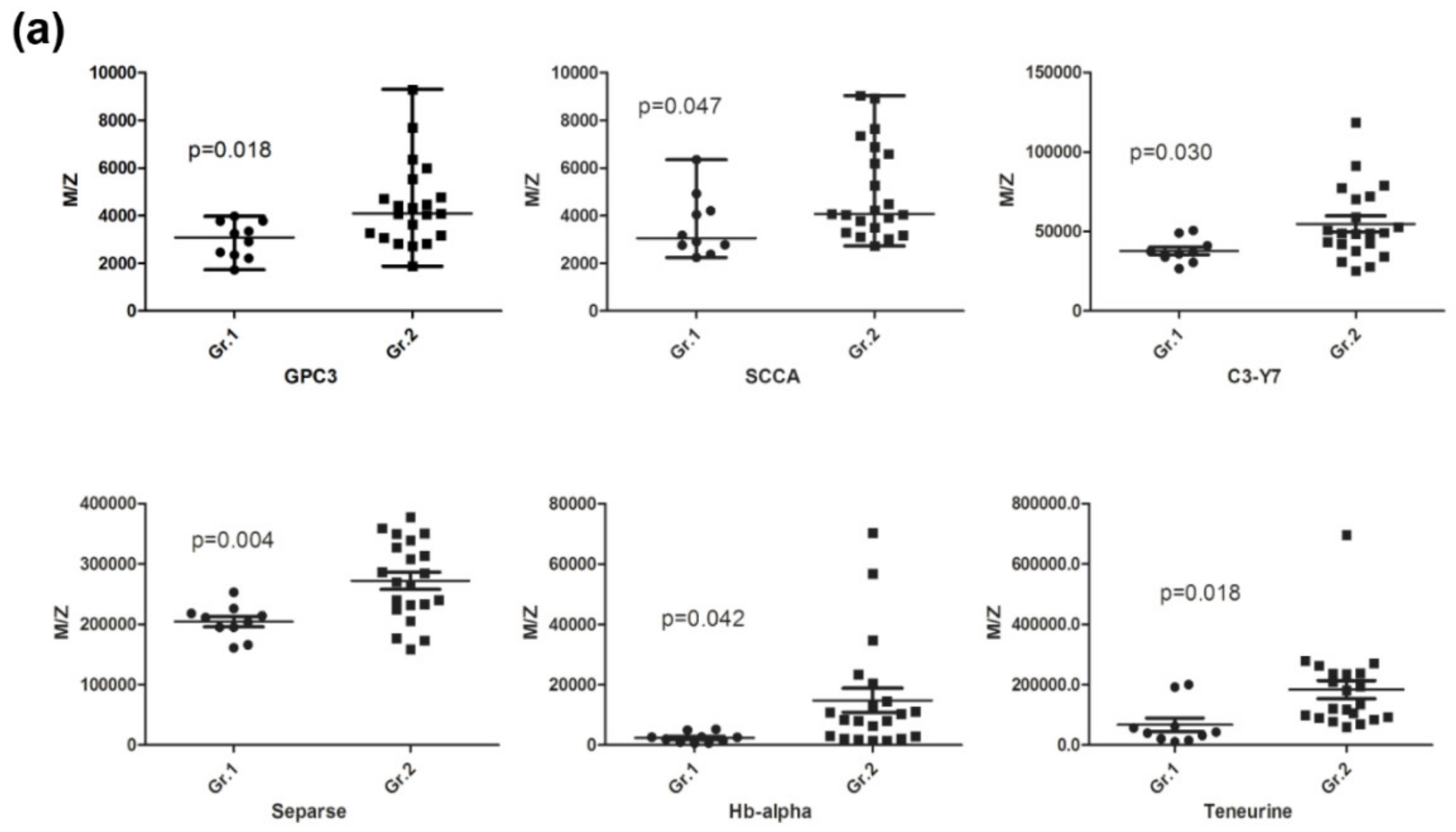
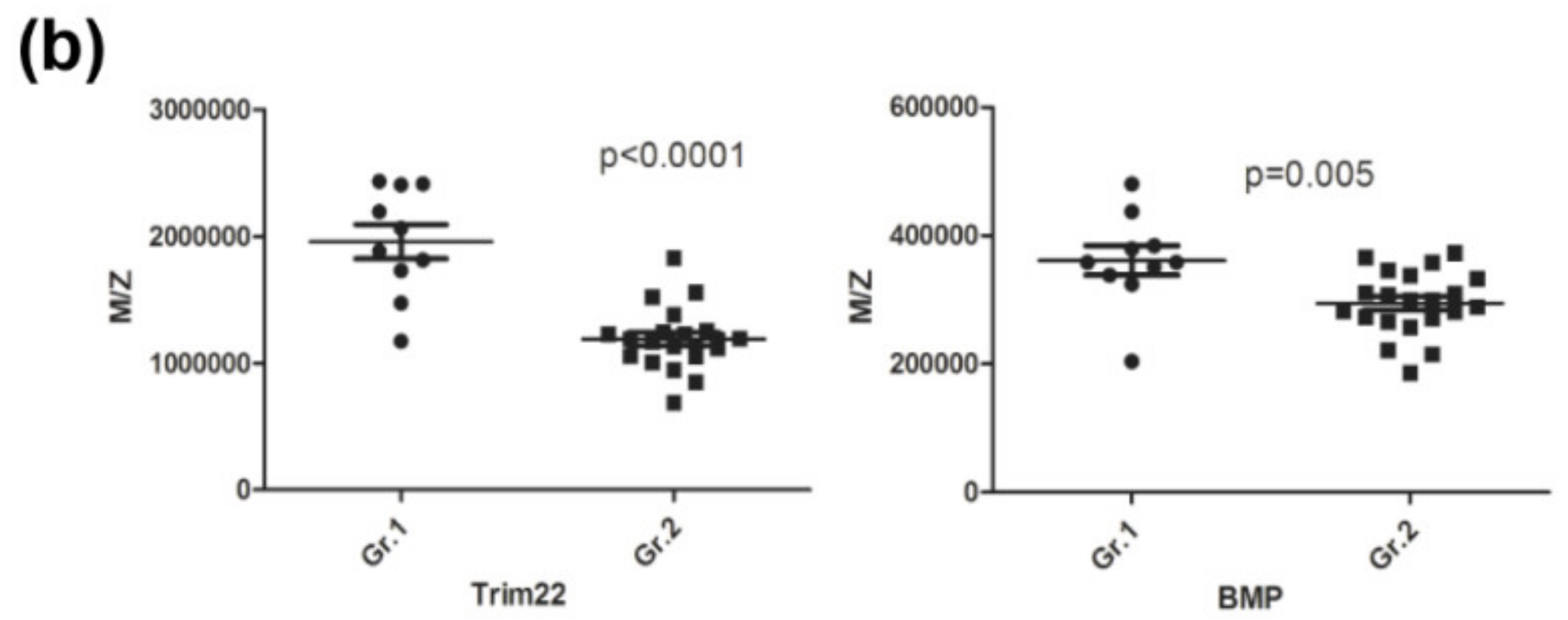
3.3. Averaged Abundance of Target Proteins in HCC Patients with Normal AFP Levels
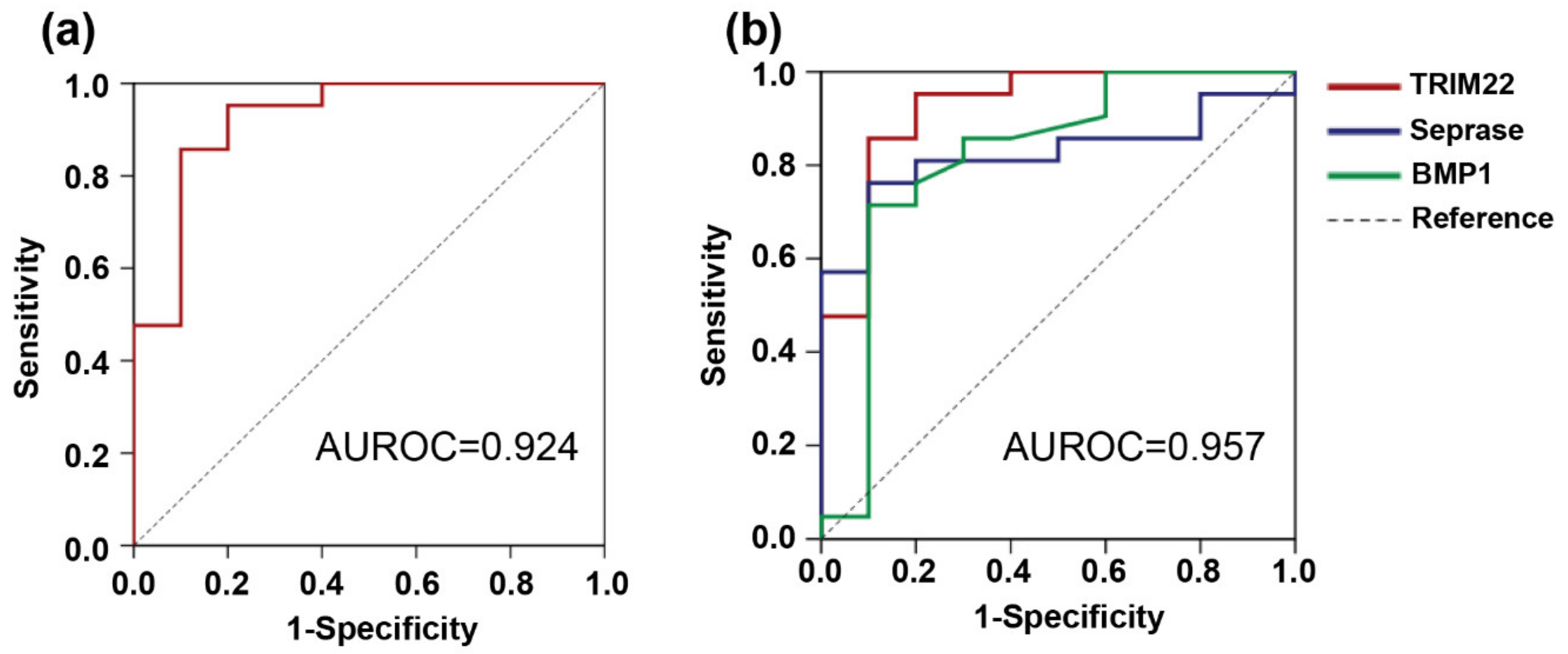
3.4. Receiver Operating Characteristic (ROC) Curves for Each of the 10 Proteins Were Significantly Altered in HCC Patients with Normal AFP Compared to Those with Liver Cirrhosis
3.5. mRNA Expression of Trim22 in Tumor/Non-Tumor Background Tissues of HCC Patients
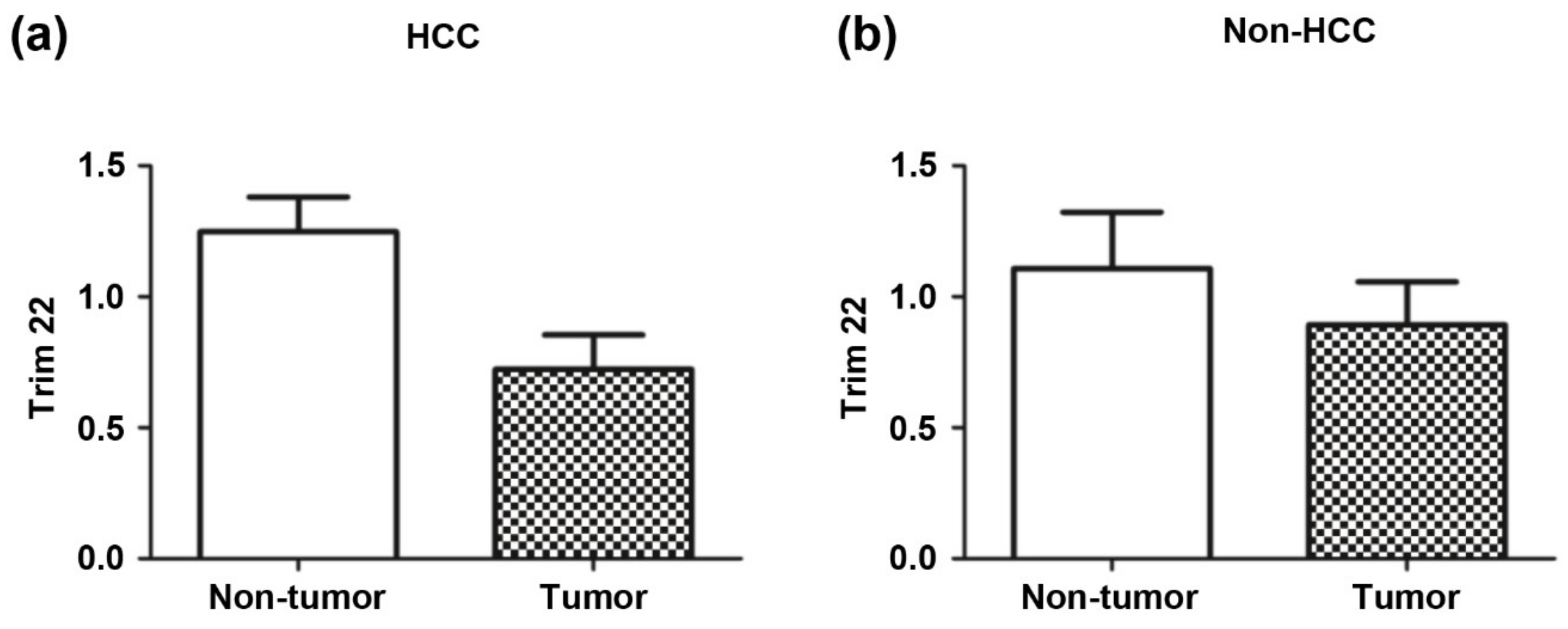
3.6. A Western Blot Analysis of Trim22 Expression in Tumor/Non-Tumor Liver Tissues in Hepatocellular Carcinoma and Non-Hepatocellular Carcinoma Patients with Liver Metastasis
3.7. Serum Trim22 Expression in Hepatocellular Carcinoma Patients with Normal AFP
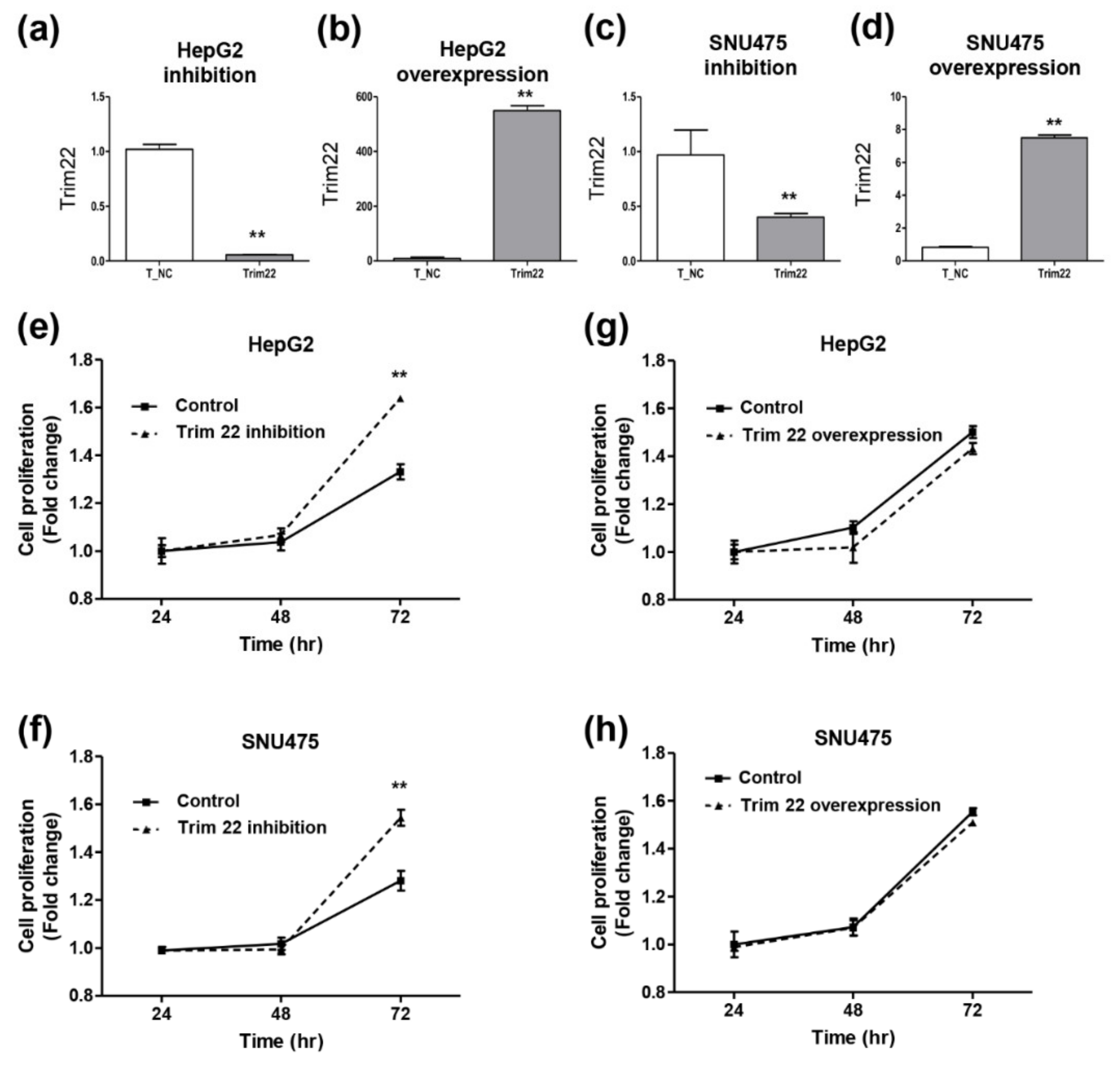
3.8. The Regulation of Hepatocytes Proliferation by Trim22
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Lok, A.S.; Sterling, R.K.; Everhart, J.E.; Wright, E.C.; Hoefs, J.C.; Di Bisceglie, A.M.; Morgan, T.R.; Kim, H.Y.; Lee, W.M.; Bonkovsky, H.L.; et al. Des-gamma-carboxy prothrombin and alpha-fetoprotein as biomarkers for the early detection of hepatocellular carcinoma. Gastroenterology 2010, 138, 493–502. [Google Scholar] [CrossRef] [PubMed]
- Marrero, J.A.; Feng, Z.; Wang, Y.; Nguyen, M.H.; Befeler, A.S.; Roberts, L.R.; Reddy, K.R.; Harnois, D.; Llovet, J.M.; Normolle, D.; et al. Alpha-fetoprotein, des-gamma carboxyprothrombin, and lectin-bound alpha-fetoprotein in early hepatocellular carcinoma. Gastroenterology 2009, 137, 110–118. [Google Scholar] [CrossRef] [PubMed]
- Koike, Y.; Shiratori, Y.; Sato, S.; Obi, S.; Teratani, T.; Imamura, M.; Yoshida, H.; Shiina, S.; Omata, M. Des-gamma-carboxy prothrombin as a useful predisposing factor for the development of portal venous invasion in patients with hepatocellular carcinoma: A prospective analysis of 227 patients. Cancer 2001, 91, 561–569. [Google Scholar] [CrossRef]
- Oda, K.; Ido, A.; Tamai, T.; Matsushita, M.; Kumagai, K.; Mawatari, S.; Saishoji, A.; Kure, T.; Ohno, K.; Toyokura, E.; et al. Highly sensitive lens culinaris agglutinin-reactive alpha-fetoprotein is useful for early detection of hepatocellular carcinoma in patients with chronic liver disease. Oncol. Rep. 2011, 26, 1227–1233. [Google Scholar] [CrossRef]
- Marrero, J.A.; Kulik, L.M.; Sirlin, C.B.; Zhu, A.X.; Finn, R.S.; Abecassis, M.M.; Roberts, L.R.; Heimbach, J.K. Diagnosis, Staging, and Management of Hepatocellular Carcinoma: 2018 Practice Guidance by the American Association for the Study of Liver Diseases. Hepatology 2018, 68, 723–750. [Google Scholar] [CrossRef]
- European Association for the Study of the Liver. EASL Clinical Practice Guidelines: Management of hepatocellular carcinoma. J. Hepatol. 2018, 69, 182–236. [Google Scholar] [CrossRef]
- Korean Liver Cancer Association; National Cancer Center. 2018 Korean Liver Cancer Association-National Cancer Center Korea Practice Guidelines for the Management of Hepatocellular Carcinoma. Gut Liver 2019, 13, 227–299. [Google Scholar] [CrossRef]
- Sarvari, J.; Mojtahedi, Z.; Kuramitsu, Y.; Malek-Hosseini, S.A.; Shamsi Shahrabadi, M.; Ghaderi, A.; Nakamura, K. Differential expression of haptoglobin isoforms in chronic active hepatitis, cirrhosis and HCC related to HBV infection. Oncol. Lett. 2011, 2, 871–877. [Google Scholar] [CrossRef]
- Chignard, N.; Beretta, L. Proteomics for hepatocellular carcinoma marker discovery. Gastroenterology 2004, 127, S120–S125. [Google Scholar] [CrossRef]
- El-Aneed, A.; Banoub, J. Proteomics in the diagnosis of hepatocellular carcinoma: Focus on high risk hepatitis B and C patients. Anticancer Res. 2006, 26, 3293–3300. [Google Scholar]
- Luk, J.M.; Liu, A.M. Proteomics of hepatocellular carcinoma in Chinese patients. OMICS 2011, 15, 261–266. [Google Scholar] [CrossRef] [PubMed]
- Ciocan-Cartita, C.A.; Jurj, A.; Buse, M.; Gulei, D.; Braicu, C.; Raduly, L.; Cojocneanu, R.; Pruteanu, L.L.; Iuga, C.A.; Coza, O.; et al. The Relevance of Mass Spectrometry Analysis for Personalized Medicine through Its Successful Application in Cancer “Omics”. Int. J. Mol. Sci. 2019, 20. [Google Scholar] [CrossRef] [PubMed]
- Poon, T.C.; Sung, J.J.; Chow, S.M.; Ng, E.K.; Yu, A.C.; Chu, E.S.; Hui, A.M.; Leung, W.K. Diagnosis of gastric cancer by serum proteomic fingerprinting. Gastroenterology 2006, 130, 1858–1864. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.W.; Liu, H.; Shin, D.H.; Yu, G.I.; Hwang, J.S.; Kim, E.S.; Yun, J.W. Proteomic and cytokine plasma biomarkers for predicting progression from colorectal adenoma to carcinoma in human patients. Proteomics 2013. [Google Scholar] [CrossRef] [PubMed]
- Wong, J.W.; Sullivan, M.J.; Cagney, G. Computational methods for the comparative quantification of proteins in label-free LCn-MS experiments. Brief. Bioinform. 2008, 9, 156–165. [Google Scholar] [CrossRef] [PubMed]
- Ahmed Mohammed, H.F.; Roberts, L.R. Should AFP (or any biomarkers) be used for HCC surveillance? Curr. Hepatol. Rep. 2017, 16, 137–145. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.W.; Tsai, H.I.; Lee, W.C.; Huang, S.W.; Lin, C.Y.; Hsieh, Y.C.; Kuo, T.; Chen, C.W.; Yu, M.C. Normal Alpha-Fetoprotein Hepatocellular Carcinoma: Are They Really Normal? J. Clin. Med. 2019, 8. [Google Scholar] [CrossRef]
- Capurro, M.; Wanless, I.R.; Sherman, M.; Deboer, G.; Shi, W.; Miyoshi, E.; Filmus, J. Glypican-3: A novel serum and histochemical marker for hepatocellular carcinoma. Gastroenterology 2003, 125, 89–97. [Google Scholar] [CrossRef]
- Zittermann, S.I.; Capurro, M.I.; Shi, W.; Filmus, J. Soluble glypican 3 inhibits the growth of hepatocellular carcinoma in vitro and in vivo. Int. J. Cancer 2010, 126, 1291–1301. [Google Scholar] [CrossRef]
- Beneduce, L.; Castaldi, F.; Marino, M.; Quarta, S.; Ruvoletto, M.; Benvegnu, L.; Calabrese, F.; Gatta, A.; Pontisso, P.; Fassina, G. Squamous cell carcinoma antigen-immunoglobulin M complexes as novel biomarkers for hepatocellular carcinoma. Cancer 2005, 103, 2558–2565. [Google Scholar] [CrossRef]
- Pontisso, P.; Quarta, S.; Caberlotto, C.; Beneduce, L.; Marino, M.; Bernardinello, E.; Tono, N.; Fassina, G.; Cavalletto, L.; Gatta, A.; et al. Progressive increase of SCCA-IgM immune complexes in cirrhotic patients is associated with development of hepatocellular carcinoma. Int. J. Cancer 2006, 119, 735–740. [Google Scholar] [CrossRef] [PubMed]
- Mao, Y.; Yang, H.; Xu, H.; Lu, X.; Sang, X.; Du, S.; Zhao, H.; Chen, W.; Xu, Y.; Chi, T.; et al. Golgi protein 73 (GOLPH2) is a valuable serum marker for hepatocellular carcinoma. Gut 2010, 59, 1687–1693. [Google Scholar] [CrossRef]
- Hatakeyama, S. TRIM proteins and cancer. Nat. Rev. Cancer 2011, 11, 792–804. [Google Scholar] [CrossRef] [PubMed]
- Hattlmann, C.J.; Kelly, J.N.; Barr, S.D. TRIM22: A Diverse and Dynamic Antiviral Protein. Mol. Biol. Int. 2012, 2012, 153415. [Google Scholar] [CrossRef] [PubMed]
- Vicenzi, E.; Poli, G. The interferon-stimulated gene TRIM22: A double-edged sword in HIV-1 infection. Cytokine Growth Factor Rev. 2018, 40, 40–47. [Google Scholar] [CrossRef] [PubMed]
- Iizuka, N.; Oka, M.; Yamada-Okabe, H.; Nishida, M.; Maeda, Y.; Mori, N.; Takao, T.; Tamesa, T.; Tangoku, A.; Tabuchi, H.; et al. Oligonucleotide microarray for prediction of early intrahepatic recurrence of hepatocellular carcinoma after curative resection. Lancet 2003, 361, 923–929. [Google Scholar] [CrossRef]
- Gao, B.; Duan, Z.; Xu, W.; Xiong, S. Tripartite motif-containing 22 inhibits the activity of hepatitis B virus core promoter, which is dependent on nuclear-located RING domain. Hepatology 2009, 50, 424–433. [Google Scholar] [CrossRef]
- Lim, K.H.; Park, E.S.; Kim, D.H.; Cho, K.C.; Kim, K.P.; Park, Y.K.; Ahn, S.H.; Park, S.H.; Kim, K.H.; Kim, C.W.; et al. Suppression of interferon-mediated anti-HBV response by single CpG methylation in the 5’-UTR of TRIM22. Gut 2018, 67, 166–178. [Google Scholar] [CrossRef]
- Matos, J.M.; Witzmann, F.A.; Cummings, O.W.; Schmidt, C.M. A pilot study of proteomic profiles of human hepatocellular carcinoma in the United States. J. Surg. Res. 2009, 155, 237–243. [Google Scholar] [CrossRef]
- Liu, Y.; He, J.; Li, C.; Benitez, R.; Fu, S.; Marrero, J.; Lubman, D.M. Identification and confirmation of biomarkers using an integrated platform for quantitative analysis of glycoproteins and their glycosylations. J. Proteome Res. 2010, 9, 798–805. [Google Scholar] [CrossRef]
- O’Brien, P.; O’Connor, B.F. Seprase: An overview of an important matrix serine protease. Biochim. Biophys. Acta 2008, 1784, 1130–1145. [Google Scholar] [CrossRef]
- Liao, Y.; Ni, Y.; He, R.; Liu, W.; Du, J. Clinical implications of fibroblast activation protein-alpha in non-small cell lung cancer after curative resection: A new predictor for prognosis. J. Cancer Res. Clin. Oncol. 2013. [Google Scholar] [CrossRef] [PubMed]
- Wikberg, M.L.; Edin, S.; Lundberg, I.V.; Van Guelpen, B.; Dahlin, A.M.; Rutegard, J.; Stenling, R.; Oberg, A.; Palmqvist, R. High intratumoral expression of fibroblast activation protein (FAP) in colon cancer is associated with poorer patient prognosis. Tumour. Biol. 2013, 34, 1013–1020. [Google Scholar] [CrossRef] [PubMed]
- Maegdefrau, U.; Amann, T.; Winklmeier, A.; Braig, S.; Schubert, T.; Weiss, T.S.; Schardt, K.; Warnecke, C.; Hellerbrand, C.; Bosserhoff, A.K. Bone morphogenetic protein 4 is induced in hepatocellular carcinoma by hypoxia and promotes tumour progression. J. Pathol. 2009, 218, 520–529. [Google Scholar] [CrossRef] [PubMed]
- Maegdefrau, U.; Bosserhoff, A.K. BMP activated Smad signaling strongly promotes migration and invasion of hepatocellular carcinoma cells. Exp. Mol. Pathol. 2012, 92, 74–81. [Google Scholar] [CrossRef]
- Shang, S.; Plymoth, A.; Ge, S.; Feng, Z.; Rosen, H.R.; Sangrajrang, S.; Hainaut, P.; Marrero, J.A.; Beretta, L. Identification of osteopontin as a novel marker for early hepatocellular carcinoma. Hepatology 2012, 55, 483–490. [Google Scholar] [CrossRef] [PubMed]
- Feng, J.T.; Liu, Y.K.; Song, H.Y.; Dai, Z.; Qin, L.X.; Almofti, M.R.; Fang, C.Y.; Lu, H.J.; Yang, P.Y.; Tang, Z.Y. Heat-shock protein 27: A potential biomarker for hepatocellular carcinoma identified by serum proteome analysis. Proteomics 2005, 5, 4581–4588. [Google Scholar] [CrossRef]
- Niu, D.; Feng, H.; Chen, W.N. Proteomic analysis of HBV-associated HCC: Insights on mechanisms of disease onset and biomarker discovery. J. Proteom. 2010, 73, 1283–1290. [Google Scholar] [CrossRef]

| LC Patients (Group 1, n = 10) | HCC Patients with Normal AFP (Group 2, n = 21) | p-Value | |
|---|---|---|---|
| Age, years | 51.2 ± 10.2 | 62.8 ± 9.9 | 0.014 |
| Sex (M/F) | 7/3 | 16/5 | 0.415 |
| Etiology HBV/HCV/Alcohol | 10/0/0 | 14/2/5 | 0.116 |
| Child-Pugh score A/B/C | 9/1/0 | 20/1/0 | 0.579 |
| Total bilirubin (mg/dL) | 0.90 ± 0.35 | 0.75 ± 0.39 | 0.303 |
| Albumin (g/dL) | 4.23 ± 0.42 | 3.90 ± 0.50 | 0.096 |
| PT (INR) | 1.07 ± 0.06 | 1.08 ± 0.06 | 0.762 |
| AFP (ng/mL) | 4.04 ± 2.16 | 4.25 ± 2.25 | 0.800 |
| Tumor number 1/2~3/4 (n, %) | NA | 15/6/0 (71.4/19.1/9.5, %) | NA |
| Tumor size <3, 3~5, >5 cm, (n, %) | NA | 4/8/9 (19/38.1/42.9, %) | NA |
| Tumor differentiation 1/2/3/4, (n, %) | NA | 3/10/6/0 (15.8/52.6/31.6/0, %) | NA |
| BCLC, 0/A/B/C/D, (n, %) | NA | 1/11/8/1 (4.8/52.4/38.1/4.8, %) | NA |
| NCBI GI | Protein | Peptide Sequence | Q1 | Q3 |
|---|---|---|---|---|
| 23271174 | Glypican 3 (GPC23) | VFGNFPK | 404.72 | 562.3 |
| 239552 | squamous cell carcinoma antigen (SCCA) | VLHFDQVTENTTGK GQWEK | 794.9 324.16 | 977.49 462.23 |
| 386783 | Haptoglobin | TEGDGVYTLNDK | 656.31 | 1081.52 |
| 115298678 | complement C3 (C3) | GYTQQLAFR | 542.28 542.28 | 762.43 863.47 |
| 4504345 | Hemoglobin subunit gamma 2 (Hb-γ2) | LLVVYPWTQR | 637.87 637.87 | 687.36 850.42 |
| 6715607 | Hemoglobin subunit alpha (Hb-α) | VGAHAGEYGAEALER | 765.37 | 1094.51 |
| 122937400 | Teneurin-3 | SDETGWTTFFDYDSEGR | 671.61 | 726.31 |
| 16933540 | Seprase | TQEHIEESR | 564.77 | 770.38 |
| 148745745 | Bone morphogenetic protein 1 (BMP-1) | DGFWR.2/y3 | 340.66 | 508.27 |
| 116283348 | Tripartite motif-containing antigen 22 (Trim22) | HLANIVER | 476.27 | 701.39 |
| Protein | Averaged Abundance (fmol) | Ratio | p-Value | Sensitivity/Specificity | AUC | 95% CI | |
|---|---|---|---|---|---|---|---|
| HCC | LC | (HCC/LC) | |||||
| GPC3 | 4438 ± 1763 | 2978 ± 766 | 1.49 | 0.003 | 61.9/100 | 0.790 | 0.632–0.949 |
| SCCA | 10,004 ± 2989 | 7914 ± 165 | 1.26 | 0.019 | 57.1/90.0 | 0.733 | 0.555–0.912 |
| 5012 ± 1986 | 3578 ± 1299 | 1.40 | 0.024 | 95.2/50.0 | 0.733 | 0.537–0.929 | |
| Haptoglobin | 18,339 ± 18,773 | 8164 ± 7024 | 2.25 | 0.038 | 85.7/60.0 | 0.791 | 0.521–0.917 |
| C3 | 410,703 ± 16,297 | 31,873 ± 5311 | 1.29 | 0.027 | 76.2/70.0 | 0.679 | 0.488–0.869 |
| 54,830 ± 22,951 | 37,815 ± 7462 | 1.38 | 0.005 | 66.7/80.0 | 0.762 | 0.590–0.934 | |
| Hb-γ2 | 535,196 ± 720,000 | 144,343 ± 93,717 | 3.71 | 0.024 | 66.7/80.0 | 0.721 | 0.540–0.902 |
| 303,225 ± 390,000 | 81,560 ± 54,269 | 3.72 | 0.021 | 66.7/80.0 | 0.714 | 0.532–0.897 | |
| Hb-α | 14,830 ± 18,335 | 2372 ± 1635 | 6.25 | 0.006 | 66.7/100 | 0.848 | 0.713–0.982 |
| Teneurin-3 | 183,688 ± 138,000 | 67,378 ± 69,984 | 2.73 | 0.004 | 95.2/80.0 | 0.876 | 0.000–1 |
| Seprase | 272,230 ± 65,326 | 204,450 ± 27,254 | 1.33 | <0.001 | 76.2/90.0 | 0.824 | 0.676–0.972 |
| BMP-1 | 288,682 ± 47,235 | 361,200 ± 72,619 | 0.80 | 0.013 | 90.0/71.4 | 0.817 | 0.630–1.003 |
| Trim22 | 1,190,000 ± 240,000 | 1,950,000 ± 420,000 | 0.61 | <0.001 | 90.0/85.7 | 0.924 | 0.813–1 |
| LC Patients (n = 30) | HCC Patients with Normal AFP (n = 39) | p-Value | |
|---|---|---|---|
| Age, years | 58.4 ± 9.67 | 67.1 ± 11.4 | 0.001 |
| Sex (M/F) | 16/14 | 32/7 | 0.010 |
| Etiology HBV/HCV/Alcohol | 22/0/8/0 | 23/3/13 | 0.213 |
| Child-Pugh score A/B/C | 29/1/0 | 37/2/0 | 0.717 |
| Total bilirubin (mg/dL) | 0.81 ± 0.36 | 0.72 ± 0.31 | 0.267 |
| Albumin (g/dL) | 4.25 ± 0.37 | 3.97 ± 0.46 | 0.008 |
| PT (INR) | 1.17 ± 0.66 | 1.06 ± 0.06 | 0.313 |
| AFP (ng/mL) | 4.27 ± 3.81 | 4.31 ± 2.19 | 0.956 |
| Tumor number 1/2~3/4 (n, %) | NA | 30/7/2 (76.9/17.9/5.1, %) | NA |
| Tumor size <3, 3~5, >5 cm, (n, %) | NA | 11/15/13 (28.2/38.5/33.3, %) | NA |
| Tumor differentiation 1/2/3/4, (n, %) | NA | 8/19/12/0 (20.5/48.7/30.8/0, %) | NA |
| BCLC, 0/A/B/C/D, (n, %) | NA | 5/20/9/5/0 (12.8/51.3/23.1/12.8/0.0, %) | NA |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, Y.-S.; Ko, E.; Yoon, E.L.; Jung, Y.K.; Kim, J.H.; Seo, Y.S.; Yim, H.J.; Kim, K.-H.; Kwon, S.Y.; Yeon, J.E.; et al. Multiplexed Proteomic Approach for Identification of Serum Biomarkers in Hepatocellular Carcinoma Patients with Normal AFP. J. Clin. Med. 2020, 9, 323. https://doi.org/10.3390/jcm9020323
Lee Y-S, Ko E, Yoon EL, Jung YK, Kim JH, Seo YS, Yim HJ, Kim K-H, Kwon SY, Yeon JE, et al. Multiplexed Proteomic Approach for Identification of Serum Biomarkers in Hepatocellular Carcinoma Patients with Normal AFP. Journal of Clinical Medicine. 2020; 9(2):323. https://doi.org/10.3390/jcm9020323
Chicago/Turabian StyleLee, Young-Sun, Eunjung Ko, Eileen L. Yoon, Young Kul Jung, Ji Hoon Kim, Yeon Seok Seo, Hyung Joon Yim, Kyun-Hwan Kim, So Young Kwon, Jong Eun Yeon, and et al. 2020. "Multiplexed Proteomic Approach for Identification of Serum Biomarkers in Hepatocellular Carcinoma Patients with Normal AFP" Journal of Clinical Medicine 9, no. 2: 323. https://doi.org/10.3390/jcm9020323
APA StyleLee, Y.-S., Ko, E., Yoon, E. L., Jung, Y. K., Kim, J. H., Seo, Y. S., Yim, H. J., Kim, K.-H., Kwon, S. Y., Yeon, J. E., Um, S. H., & Byun, K. S. (2020). Multiplexed Proteomic Approach for Identification of Serum Biomarkers in Hepatocellular Carcinoma Patients with Normal AFP. Journal of Clinical Medicine, 9(2), 323. https://doi.org/10.3390/jcm9020323






