The Relationship between Platelet Count and Host Gut Microbiota: A Population-Based Retrospective Cross-Sectional Study
Abstract
:1. Introduction
2. Materials and Method
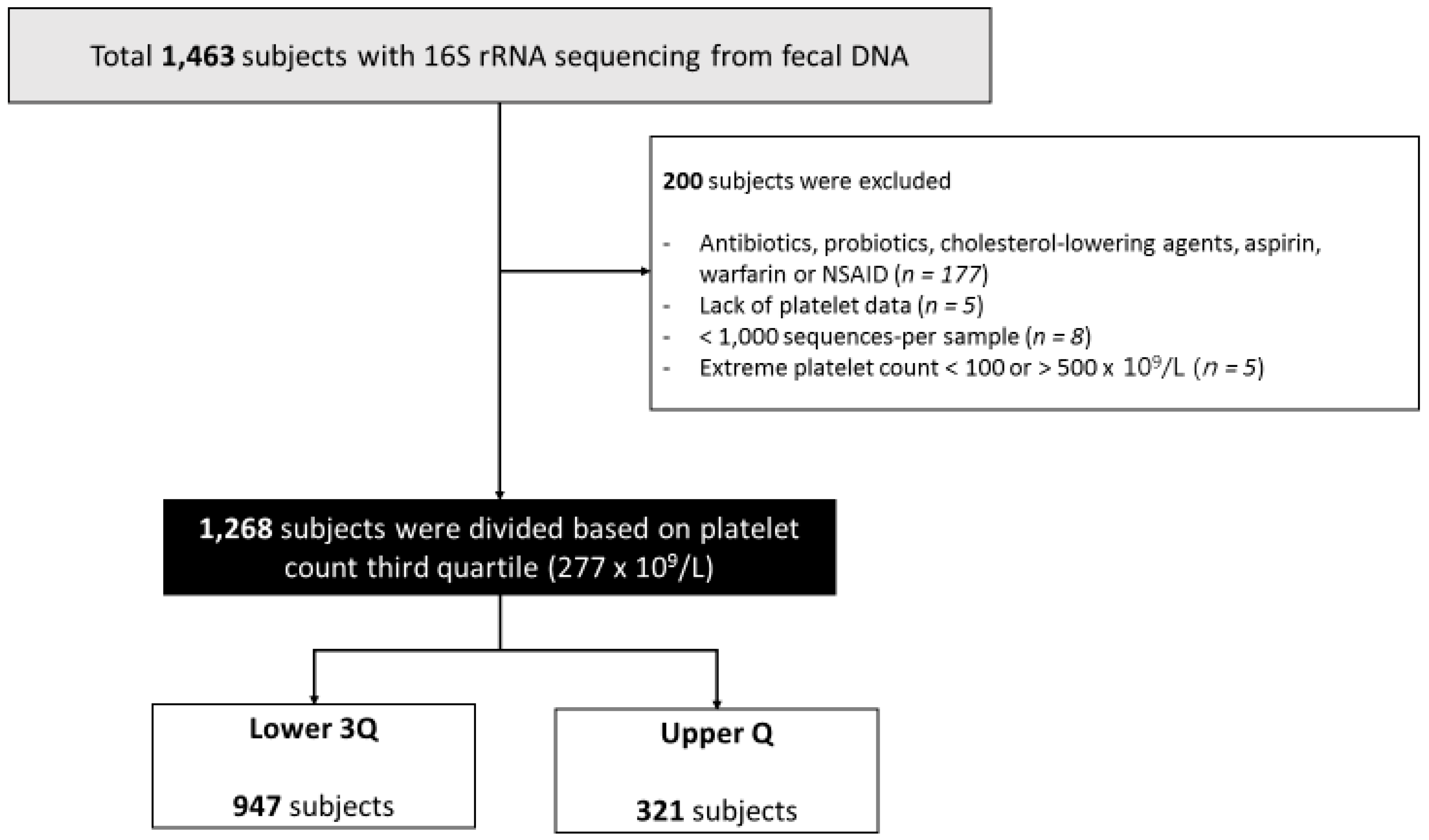
2.1. Study Population
2.2. Study Design
2.3. DNA Extraction and 16S Gene rRNA Sequencing for Bacterial Communities
2.4. Compositional Analysis of 16S rRNA Gene
2.5. Statistical Analysis
3. Results
3.1. Baseline Characteristics
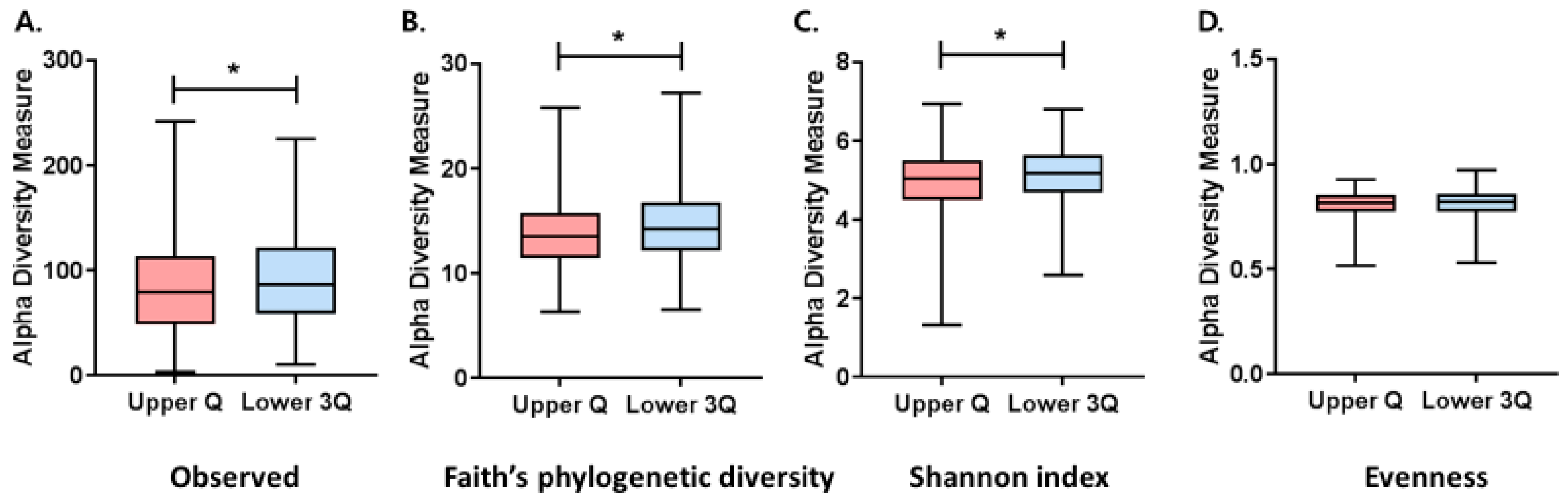
3.2. Comparison of Alpha Diversity between the Upper Quartile and Lower Three Quartiles Groups
3.3. Comparison of Beta Diversity between the Upper Quartile and Lower Three Quartiles Groups
3.4. Comparison of the Microbial Composition between the Upper Quartile and Lower Three Quartile Groups
3.5. Correlation between Platelet Count and Gut Microbiota
3.6. Comparison of the Functional Microbial Composition between the Upper Quartile and Lower Three Quartiles Groups
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing mothur: Open-Source, Platform-Independent, Community-Supported Software for Describing and Comparing Microbial Communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef] [PubMed]
- Gill, S.R.; Pop, M.; DeBoy, R.T.; Eckburg, P.B.; Turnbaugh, P.J.; Samuel, B.S.; Gordon, J.I.; Relman, D.A.; Fraser-Liggett, C.M.; Nelson, K.E.; et al. Metagenomic Analysis of the Human Distal Gut Microbiome. Science 2006, 312, 1355–1359. [Google Scholar] [CrossRef] [PubMed]
- Goodrich, J.K.; Waters, J.L.; Poole, A.C.; Sutter, J.L.; Koren, O.; Blekhman, R.; Beaumont, M.; Van Treuren, W.; Knight, R.; Bell, J.T.; et al. Human Genetics Shape the Gut Microbiome. Cell 2014, 159, 789–799. [Google Scholar] [CrossRef] [PubMed]
- Wu, G.D.; Chen, J.; Hoffmann, C.; Bittinger, K.; Chen, Y.-Y.; Keilbaugh, S.A.; Bewtra, M.; Knights, D.; Walters, W.A.; Knight, R.; et al. Linking Long-Term Dietary Patterns with Gut Microbial Enterotypes. Science 2011, 334, 105–108. [Google Scholar] [CrossRef] [PubMed]
- Preidis, G.A.; Versalovic, J. Targeting the Human Microbiome with Antibiotics, Probiotics, and Prebiotics: Gastroenterology Enters the Metagenomics Era. Gastroenterology 2009, 136, 2015–2031. [Google Scholar] [CrossRef] [PubMed]
- Rogers, M.A.M.; Aronoff, D. The influence of non-steroidal anti-inflammatory drugs on the gut microbiome. Clin. Microbiol. Infect. 2016, 22, 178.e1–178.e9. [Google Scholar] [CrossRef]
- Woodmansey, E.J. Intestinal bacteria and ageing. J. Appl. Microbiol. 2007, 102, 1178–1186. [Google Scholar] [CrossRef]
- Turnbaugh, P.J.; Ley, R.E.; Mahowald, M.A.; Magrini, V.; Mardis, E.R.; Gordon, J.I. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 2006, 444, 1027–1031. [Google Scholar] [CrossRef]
- Hold, G.L. Role of the gut microbiota in inflammatory bowel disease pathogenesis: What have we learnt in the past 10 years? World J. Gastroenterol. 2014, 20, 1192. [Google Scholar] [CrossRef]
- Tang, W.H.W.; Hazen, S.L. The Gut Microbiome and Its Role in Cardiovascular Diseases. Circulation 2017, 135, 1008–1010. [Google Scholar] [CrossRef]
- Holmes, E.; Loo, R.L.; Stamler, J.; Bictash, M.; Yap, I.K.S.; Chan, Q.; Ebbels, T.; De Iorio, M.; Brown, I.J.; Veselkov, K.A.; et al. Human metabolic phenotype diversity and its association with diet and blood pressure. Nature 2008, 453, 396–400. [Google Scholar] [CrossRef] [PubMed]
- Quigley, E.M.M. Microbiota-Brain-Gut Axis and Neurodegenerative Diseases. Curr. Neurol. Neurosci. Rep. 2017, 17, 94. [Google Scholar] [CrossRef] [PubMed]
- De Vadder, F.; Kovatcheva-Datchary, P.; Goncalves, D.; Vinera, J.; Zitoun, C.; Duchampt, A.; Bäckhed, F.; Mithieux, G. Microbiota-Generated Metabolites Promote Metabolic Benefits via Gut-Brain Neural Circuits. Cell 2014, 156, 84–96. [Google Scholar] [CrossRef] [PubMed]
- Fiebiger, U.; Bereswill, S.; Heimesaat, M.M. Dissecting the interplay between intestinal microbiota and host immunity in health and disease: Lessons learned from germfree and gnotobiotic animal models. Eur. J. Microbiol. Immunol. 2016, 6, 253–271. [Google Scholar] [CrossRef] [PubMed]
- Schafer, A.I. Thrombocytosis. N. Engl. J. Med. 2004, 350, 1211–1219. [Google Scholar] [CrossRef] [PubMed]
- Ali, R.A.; Wuescher, L.M.; Worth, R.G. Platelets: Essential components of the immune system. Curr. Trends Immunol. 2015, 16, 65–78. [Google Scholar] [PubMed]
- Słabuszewska-Jóźwiak, A.; Dmoch-Gajzlerska, E.; Kozakiewicz, B.; Jakiel, G. The prognostic significance of thrombocytosis in ovarian cancer. Ann. Agric. Environ. Med. 2015, 22, 731–735. [Google Scholar] [CrossRef]
- Mirsaeidi, M.; Peyrani, P.; Aliberti, S.; Filardo, G.; Bordon, J.; Blasi, F.; Ramirez, J.A. Thrombocytopenia and Thrombocytosis at Time of Hospitalization Predict Mortality in Patients With Community-Acquired Pneumonia. Chest 2010, 137, 416–420. [Google Scholar] [CrossRef]
- Tomita, M.; Shimizu, T.; Hara, M.; Ayabe, T.; Onitsuka, T. Prognostic impact of thrombocytosis in resectable non-small cell lung cancer. Interact. CardioVasc. Thorac. Surg. 2008, 7, 613–615. [Google Scholar] [CrossRef]
- Harrison, M.T.; Short, P.; Williamson, P.A.; Singanayagam, A.; Chalmers, J.D.; Schembri, S. Thrombocytosis is associated with increased short and long term mortality after exacerbation of chronic obstructive pulmonary disease: a role for antiplatelet therapy? Thorax 2014, 69, 609–615. [Google Scholar] [CrossRef]
- Yoon, H.-Y.; Kim, H.-N.; Lee, S.H.; Kim, S.J.; Chang, Y.; Ryu, S.; Shin, H.; Kim, H.-L.; Lee, J.H. Association between Neutrophil-to-Lymphocyte Ratio and Gut Microbiota in a Large Population: A Retrospective Cross-Sectional Study. Sci. Rep. 2018, 8, 16031. [Google Scholar] [CrossRef] [PubMed]
- Eidelman, O.; Jozwik, C.; Huang, W.; Srivastava, M.; Rothwell, S.W.; Jacobowitz, D.M.; Ji, X.; Zhang, X.; Guggino, W.; Wright, J.; et al. Gender Dependence for a Subset of the Low-Abundance Signaling Proteome in Human Platelets. Hum. Genom. Proteomics 2010, 2, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Biino, G.; Santimone, I.; Minelli, C.; Sorice, R.; Frongia, B.; Traglia, M.; Ulivi, S.; Di Castelnuovo, A.; Gögele, M.; Nutile, T.; et al. Age- and Sex-Related Variations in Platelet Count in Italy: A Proposal of Reference Ranges Based on 40987 Subjects’ Data. PLoS ONE 2013, 8, e54289. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.-N.; Yun, Y.; Ryu, S.; Chang, Y.; Kwon, M.-J.; Cho, J.; Shin, H.; Kim, H.-L. Correlation between gut microbiota and personality in adults: A cross-sectional study. Brain Behav. Immun. 2018, 69, 374–385. [Google Scholar] [CrossRef] [PubMed]
- Kozich, J.J.; Westcott, S.L.; Baxter, N.T.; Highlander, S.K.; Schloss, P.D. Development of a Dual-Index Sequencing Strategy and Curation Pipeline for Analyzing Amplicon Sequence Data on the MiSeq Illumina Sequencing Platform. Appl. Environ. Microbiol. 2013, 79, 5112–5120. [Google Scholar] [CrossRef] [PubMed]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; A Johnson, A.J.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef] [PubMed]
- Callahan, B.J.; McMurdie, P.J.; Holmes, S.P. Exact sequence variants should replace operational taxonomic units in marker-gene data analysis. ISME J. 2017, 11, 2639–2643. [Google Scholar] [CrossRef] [PubMed]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Peña, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [PubMed]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. QIIME 2: Reproducible, Interactive, Scalable, and Extensible Microbiome Data Science. Available online: https://peerj.com/preprints/27295/ (accessed on 13 December 2018).
- DeSantis, T.Z.; Hugenholtz, P.; Larsen, N.; Rojas, M.; Brodie, E.L.; Keller, K.; Huber, T.; Dalevi, D.; Hu, P.; Andersen, G.L.; et al. Greengenes, a Chimera-Checked 16S rRNA Gene Database and Workbench Compatible with ARB. Appl. Environ. Microbiol. 2006, 72, 5069–5072. [Google Scholar] [CrossRef] [PubMed]
- Faith, D.P. Conservation evaluation and phylogenetic diversity. Biol. Conserv. 1992, 61, 1–10. [Google Scholar] [CrossRef]
- Neisser, U. Cognitive Psychology: Classic Edition; Psychology Press: New York, NY, USA, 2014. [Google Scholar]
- Pielou, E. The measurement of diversity in different types of biological collections. J. Theor. Biol. 1966, 13, 131–144. [Google Scholar] [CrossRef]
- Lozupone, C.; Lladser, M.E.; Knights, D.; Stombaugh, J.; Knight, R. UniFrac: An effective distance metric for microbial community comparison. ISME J. 2010, 5, 169–172. [Google Scholar] [CrossRef] [PubMed]
- Charbonneau, M.; Subramanian, S.; Seedorf, H.; Knight, R.; Rosenbaum, M.; Faith, J.J.; Guruge, J.L.; Goodman, A.L.; Clemente, J.C.; Heath, A.C.; et al. The long-term stability of the human gut microbiota. Science 2013, 341, 1237439. [Google Scholar]
- Navas-Molina, J.A.; Peralta-Sánchez, J.M.; González, A.; McMurdie, P.J.; Vázquez-Baeza, Y.; Xu, Z.; Ursell, L.K.; Lauber, C.; Zhou, H.; Song, S.J.; et al. Advancing Our Understanding of the Human Microbiome Using QIIME. Meth. Enzymol. 2013, 531, 371–444. [Google Scholar] [PubMed]
- Mandal, S.; Van Treuren, W.; White, R.A.; Eggesbø, M.; Knight, R.; Peddada, S.D. Analysis of composition of microbiomes: A novel method for studying microbial composition. Microb. Ecol. Health Dis. 2015, 26, 27663. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.-W.; Kuan, C.-S.; Wu, L.S.-H.; Weng, J.T.-Y. Metagenome and Metatranscriptome Profiling of Moderate and Severe COPD Sputum in Taiwanese Han Males. PLoS ONE 2016, 11, e0159066. [Google Scholar] [CrossRef]
- Odamaki, T.; Kato, K.; Sugahara, H.; Hashikura, N.; Takahashi, S.; Xiao, J.-Z.; Abe, F.; Osawa, R. Age-related changes in gut microbiota composition from newborn to centenarian: A cross-sectional study. BMC Microbiol. 2016, 16, 333. [Google Scholar] [CrossRef] [PubMed]
- Markle, J.G.M.; Frank, D.N.; Mortin-Toth, S.; Robertson, C.; Feazel, L.M.; Rolle-Kampczyk, U.; Von Bergen, M.; McCoy, K.D.; MacPherson, A.J.; Danska, J.S.; et al. Sex Differences in the Gut Microbiome Drive Hormone-Dependent Regulation of Autoimmunity. Science 2013, 339, 1084–1088. [Google Scholar] [CrossRef] [PubMed]
- Tilg, H.; Moschen, A.R.; Kaser, A. Obesity and the Microbiota. Gastroenterology 2009, 136, 1476–1483. [Google Scholar] [CrossRef] [PubMed]
- Savin, Z.; Kivity, S.; Yonath, H.; Yehuda, S. Smoking and the intestinal microbiome. Arch. Microbiol. 2018, 200, 677–684. [Google Scholar] [CrossRef] [PubMed]
- Langille, M.G.I.; Zaneveld, J.; Caporaso, J.G.; McDonald, D.; Knights, D.; A Reyes, J.; Clemente, J.C.; E Burkepile, D.; Thurber, R.L.V.; Knight, R.; et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 2013, 31, 814–821. [Google Scholar] [CrossRef] [PubMed]
- Parks, D.H.; Tyson, G.W.; Hugenholtz, P.; Beiko, R.G. STAMP: Statistical analysis of taxonomic and functional profiles. Bioinformatics 2014, 30, 3123–3124. [Google Scholar] [CrossRef]
- Barcenilla, A.; Pryde, S.E.; Martin, J.C.; Duncan, S.H.; Stewart, C.S.; Henderson, C.; Flint, H.J. Phylogenetic Relationships of Butyrate-Producing Bacteria from the Human Gut. Appl. Environ. Microbiol. 2000, 66, 1654–1661. [Google Scholar] [CrossRef] [PubMed]
- Tremaroli, V.; Bäckhed, F. Functional interactions between the gut microbiota and host metabolism. Nature 2012, 489, 242–249. [Google Scholar] [CrossRef] [PubMed]
- Sokol, H.; Pigneur, B.; Watterlot, L.; Lakhdari, O.; Bermúdez-Humarán, L.G.; Gratadoux, J.-J.; Blugeon, S.; Bridonneau, C.; Furet, J.-P.; Corthier, G.; et al. Faecalibacterium prausnitzii is an anti-inflammatory commensal bacterium identified by gut microbiota analysis of Crohn disease patients. Proc. Natl. Acad. Sci. USA 2008, 105, 16731–16736. [Google Scholar] [CrossRef] [PubMed]
- Seksik, P.; Firmesse, O.; Nion-Larmurier, I.; Cosnes, J.; Sokol, H.; Furet, J.P.; Beaugerie, L.; Corthier, G.; Marteau, P.; Doré, J.; et al. Low counts of Faecalibacterium prausnitzii in colitis microbiota. Inflamm. Bowel Dis. 2009, 15, 1183–1189. [Google Scholar]
- Voudoukis, E. Multipotent role of platelets in inflammatory bowel diseases: A clinical approach. WJG 2014, 20, 3180. [Google Scholar] [CrossRef] [PubMed]
- Taniguchi, A.; Fukushima, M.; Seino, Y.; Sakai, M.; Yoshii, S.; Nagasaka, S.; Yamauchi, I.; Okumura, T.; Nin, K.; Tokuyama, K.; et al. Platelet count is independently associated with insulin resistance in non-obese Japanese type 2 diabetic patients. Metabolism 2003, 52, 1246–1249. [Google Scholar] [CrossRef]
- Sterner, G.; Carlson, J.; Ekberg, G. Raised platelet levels in diabetes mellitus complicated with nephropathy. J. Intern. Med. 1998, 244, 437–441. [Google Scholar] [CrossRef]
- Furet, J.-P.; Kong, L.-C.; Tap, J.; Poitou, C.; Basdevant, A.; Bouillot, J.-L.; Mariat, D.; Corthier, G.; Doré, J.; Henegar, C.; et al. Differential Adaptation of Human Gut Microbiota to Bariatric Surgery-Induced Weight Loss: Links with Metabolic and Low-Grade Inflammation Markers. Diabetes 2010, 59, 3049–3057. [Google Scholar] [CrossRef]
- Käser, A. Interleukin-6 stimulates thrombopoiesis through thrombopoietin: Role in inflammatory thrombocytosis. Blood 2001, 98, 2720–2725. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.-L. Faecalibacterium prausnitzii supernatant ameliorates dextran sulfate sodium induced colitis by regulating Th17 cell differentiation. WJG 2016, 22, 5201. [Google Scholar] [CrossRef]
- The Human Microbiome Project Consortium; Huttenhower, C.; Gevers, D.; Knight, R.; Abubucker, S.; Badger, J.H.; Chinwalla, A.T.; Creasy, H.H.; Earl, A.M.; Fitzgerald, M.G.; et al. Structure, function and diversity of the healthy human microbiome. Nature 2012, 486, 207–214. [Google Scholar]
- Lozupone, C.A.; Stombaugh, J.I.; Gordon, J.I.; Jansson, J.K.; Knight, R. Diversity, stability and resilience of the human gut microbiota. Nature 2012, 489, 220–230. [Google Scholar] [CrossRef] [PubMed]
- Ott, S.J. Reduction in diversity of the colonic mucosa associated bacterial microflora in patients with active inflammatory bowel disease. Gut 2004, 53, 685–693. [Google Scholar] [CrossRef] [PubMed]
- Shah, V.; Lambeth, S.M.; Carson, T.; Lowe, J.; Ramaraj, T.; Leff, J.W.; Luo, L.; Bell, C.J. Composition Diversity and Abundance of Gut Microbiome in Prediabetes and Type 2 Diabetes. J. Diabetes Obes. 2015, 2, 108–114. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Karlsson, C.; Olsson, C.; Adlerberth, I.; Wold, A.E.; Strachan, D.P.; Martricardi, P.M.; Åberg, N.; Perkin, M.R.; Tripodi, S.; et al. Reduced diversity in the early fecal microbiota of infants with atopic eczema. J. Allergy Clin. Immunol. Pract. 2008, 121, 129–134. [Google Scholar] [CrossRef] [PubMed]
- Taur, Y.; Jenq, R.; Perales, M.-A.; Littmann, E.R.; Morjaria, S.; Ling, L.; No, D.; Gobourne, A.; Viale, A.; Dahi, P.; et al. The effects of intestinal tract bacterial diversity on mortality following allogeneic hematopoietic stem cell transplantation. Blood 2014, 124, 1174–1182. [Google Scholar] [CrossRef] [PubMed]
- Malard, F.; Gasc, C.; Plantamura, E.; Dore, J. High gastrointestinal microbial diversity and clinical outcome in graft-versus-host disease patients. Bone Marrow Transplant. 2018, 53, 1493–1497. [Google Scholar] [CrossRef]
- D’Angelo, G.; Capasso, S.; Sticco, L.; Russo, D. Glycosphingolipids: Synthesis and functions. FEBS J. 2013, 280, 6338–6353. [Google Scholar] [CrossRef] [PubMed]
- Bremer, E.G.; Hakomori, S.; Bowen-Pope, D.F.; Raines, E.; Ross, R. Ganglioside-mediated modulation of cell growth, growth factor binding, and receptor phosphorylation. J. Biol. Chem. 1984, 259, 6818–6825. [Google Scholar] [PubMed]
- Yates, A.; VanBrocklyn, J.; Saqr, H.; Guan, Z.; Stokes, B.; O’Dorisio, M. Mechanisms through Which Gangliosides Inhibit PDGF-Stimulated Mitogenesis in Intact Swiss 3T3 Cells: Receptor Tyrosine Phosphorylation, Intracellular Calcium, and Receptor Binding. Exp. Cell Res. 1993, 204, 38–45. [Google Scholar] [CrossRef] [PubMed]
- Su, R.; Li, K.; Yang, M.; Zhang, X.; Tsang, K.; Fok, T.; Li, C.; Yuen, P. Platelet-derived growth factor enhances ex vivo expansion of megakaryocytic progenitors from human cord blood. Bone Marrow Transplant. 2001, 27, 1075–1080. [Google Scholar] [CrossRef] [PubMed]
- Pekny, M.; Gebre-Medhin, S.; Swolin, B.; Betsholtz, C.; Leveen, P.; Larsson, E. Mice deficient for PDGF B show renal, cardiovascular, and hematological abnormalities. Genes Dev. 1994, 8, 1875–1887. [Google Scholar]
- Lev, P.R.; Marta, R.F.; Vassallu, P.; Molinas, F.C. Variation of PDGF, TGFbeta, and bFGF levels in essential thrombocythemia patients treated with anagrelide. Am. J. Hematol. 2002, 70, 85–91. [Google Scholar] [CrossRef] [PubMed]
- Kavey, R.-E.W.; Allada, V.; Daniels, S.R.; Hayman, L.L.; McCrindle, B.W.; Newburger, J.W. Cardiovascular risk reduction in high-risk pediatric patients: A scientific statement from the American Heart Association expert panel on population and prevention science; the councils on cardiovascular disease in the young, epidemiology and prevention, nutrition, physical activity and metabolism, high blood pressure research, cardiovascular nursing, and the kidney in heart disease; and the interdisciplinary working group on quality of care and outcomes research: Endorsed by the American Academy of Pediatrics. Circulation 2006, 114, 2710–2738. [Google Scholar] [PubMed]
- Hua, W.; Hongwei, W.; Peixuan, C. Kawasaki disease on PDGF expression and VSMC proliferation. J. Tongji Med. Univ. 1998, 18, 243–246. [Google Scholar] [CrossRef]
- Reilly, J. Idiopathic myelofibrosis: pathogenesis, natural history and management. Blood Rev. 1997, 11, 233–242. [Google Scholar] [CrossRef]
- Greer, J.P.; Arber, D.A.; Glader, B.; List, A.F.; Means, R.T.; Paraskevas, F.; Rodgers, G.M. Wintrobe’s Clinical Hematology; Wolters Kluwer Health/Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2014. [Google Scholar]
- Kadikoylu, G.; Yavasoglu, I.; Bolaman, Z.; Senturk, T. Platelet parameters in women with iron deficiency anemia. J. Natl. Med. Assoc. 2006, 98, 398–402. [Google Scholar]
- Park, M.-J.; Park, P.-W.; Seo, Y.-H.; Kim, K.-H.; Park, S.-H.; Jeong, J.-H.; Ahn, J.-Y. The relationship between iron parameters and platelet parameters in women with iron deficiency anemia and thrombocytosis. Platelets 2012, 24, 348–351. [Google Scholar] [CrossRef]
- Frelinger, A.L.; Grace, R.F.; Gerrits, A.J.; Berny-Lang, M.A.; Brown, T.; Carmichael, S.L.; Neufeld, E.J.; Michelson, A.D. Platelet function tests, independent of platelet count, are associated with bleeding severity in ITP. Blood 2015, 126, 873–879. [Google Scholar] [CrossRef] [PubMed]
| Variables | Lower 3Q | Upper Q | Total | p-Value |
|---|---|---|---|---|
| No. | 947 | 321 | 1268 | |
| Age, years | 45.7 ± 9.0 | 44.7 ± 8.4 | 45.4 ± 8.8 | 0.093 |
| Male sex | 634 (66.9%) | 153 (47.7%) | 787 (62.1%) | <0.001 |
| Body mass index, kg/m2 | 23.6 ± 3.1 | 23.6 ± 3.1 | 23.6 ± 3.1 | 0.748 |
| Smoking status | 0.056 | |||
| Never | 505 (57.0%) | 195 (64.8%) | 700 (59.0%) | |
| Former | 216 (24.4%) | 58 (19.3%) | 274 (23.1%) | |
| Current | 165 (18.6%) | 48 (15.9%) | 213 (17.9%) | |
| Smoking amount, pack-years | 14.4 ± 10.8 | 15.4 ± 13.9 | 14.6 ± 11.6 | 0.483 |
| Laboratory finding | ||||
| Platelet, 109/L | 224.4 ± 33.5 | 314.9 ± 36.4 | 247.3 ± 52.2 | <0.001 |
| White blood cell, 103/mm3 | 5.6 ± 1.4 | 6.4 ± 1.6 | 5.8 ± 1.5 | <0.001 |
| Neutrophil, % | 54.9 ± 8.0 | 56.0 ± 7.7 | 55.2 ± 8.0 | 0.037 |
| Lymphocyte, % | 33.5 ± 7.4 | 34.9 ± 7.1 | 35.4 ± 7.3 | 0.167 |
| Eosinophil, % | 2.6 ± 2.1 | 2.4 ± 1.9 | 2.6 ± 2.4 | 0.189 |
| Basophil, % | 0.4 ± 0.3 | 0.5 ± 0.3 | 0.5 ± 0.3 | 0.003 |
| Monocyte, % | 6.5 ± 1.6 | 6.2 ± 1.5 | 6.4 ± 1.6 | 0.003 |
| Neutrophil/lymphocyte ratio | 1.7 ± 0.7 | 1.7 ± 0.6 | 1.7 ± 0.7 | 0.238 |
| Hematocrit, % | 42.3 ± 3.6 | 40.9 ± 4.0 | 42.0 ± 3.7 | <0.001 |
| Iron, µg/dL | 122.0 ± 41.4 | 115.5 ± 4.5.6 | 120.4 ± 42.6 | 0.045 |
| Ferritin, ng/mL | 161.1 ± 135.1 | 138.0 ± 128.3 | 155.3 ± 133.7 | 0.008 |
| C-reactive protein, mg/dL | 0.1 ± 0.2 | 0.1 ± 0.1 | 0.1 ± 0.2 | 0.808 |
| Beta Diversity Indices | Total | Male | Female |
|---|---|---|---|
| Unweighted UniFrac distance | 3.472 * | 2.964 * | 1.598 |
| Weighted UniFrac distance | 1.696 | 3.074 * | 1.643 |
| Jaccard dissimilarity | 1.299 * | 1.167 | 1.059 |
| Level | Taxonomic Assignment | W a | Normalized W b |
|---|---|---|---|
| Class | k__Bacteria; p__Firmicutes; c__Clostridia * | 10 | 0.32 |
| Order | k__Bacteria; p__Firmicutes; c__Clostridia; o__Clostridiales * | 21 | 0.44 |
| Family | k__Bacteria; p__Firmicutes; c__Clostridia; o__Clostridiales; f__Ruminococcaceae * | 34 | 0.40 |
| Genus | k__Bacteria; p__Firmicutes; c__Clostridia; o__Clostridiales; f__Ruminococcaceae; g__Faecalibacterium * | 140 | 0.65 |
| Order | Family | Genus | n Not to Zero (%) | CE | p-Value * | q-Value * |
|---|---|---|---|---|---|---|
| Clostridiales | Ruminococcaceae | 1257 | −0.00031 | 0.00043 | 0.0064 | |
| Clostridiales | Ruminococcaceae | Faecalibacterium | 1220 | −0.00022 | 0.00097 | 0.0016 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yoon, H.-Y.; Kim, H.-N.; Lee, S.H.; Kim, S.J.; Chang, Y.; Ryu, S.; Shin, H.; Kim, H.-L.; Lee, J.H. The Relationship between Platelet Count and Host Gut Microbiota: A Population-Based Retrospective Cross-Sectional Study. J. Clin. Med. 2019, 8, 230. https://doi.org/10.3390/jcm8020230
Yoon H-Y, Kim H-N, Lee SH, Kim SJ, Chang Y, Ryu S, Shin H, Kim H-L, Lee JH. The Relationship between Platelet Count and Host Gut Microbiota: A Population-Based Retrospective Cross-Sectional Study. Journal of Clinical Medicine. 2019; 8(2):230. https://doi.org/10.3390/jcm8020230
Chicago/Turabian StyleYoon, Hee-Young, Han-Na Kim, Su Hwan Lee, Soo Jung Kim, Yoosoo Chang, Seungho Ryu, Hocheol Shin, Hyung-Lae Kim, and Jin Hwa Lee. 2019. "The Relationship between Platelet Count and Host Gut Microbiota: A Population-Based Retrospective Cross-Sectional Study" Journal of Clinical Medicine 8, no. 2: 230. https://doi.org/10.3390/jcm8020230
APA StyleYoon, H.-Y., Kim, H.-N., Lee, S. H., Kim, S. J., Chang, Y., Ryu, S., Shin, H., Kim, H.-L., & Lee, J. H. (2019). The Relationship between Platelet Count and Host Gut Microbiota: A Population-Based Retrospective Cross-Sectional Study. Journal of Clinical Medicine, 8(2), 230. https://doi.org/10.3390/jcm8020230






