1. Introduction
The traditional diagnostic technique, based on the expert knowledge of doctors, has a critical limitation in that the result of the diagnosis is heavily dependent on the personal knowledge and experience of the doctor. Consequently, the performance of diagnosis is limited and varies with the doctor’s experience. To surmount this limitation, a double screening scheme has been applied in some hospitals by employing an additional expert [
1]. However, this scheme is expensive and time consuming. The development of artificial intelligence [
2,
3] and imaging techniques (such as magnetic resonance imaging (MRI), X-ray, and ultrasound) in medical research has led to the development of an image-based computer-aided diagnosis system (CAD) [
4,
5], which is widely used to assist doctors in the diagnosis of various kinds of diseases. The CAD system serves as the additional expert in the double screening scheme and makes suggestions to doctors during diagnosis of diseases. This diagnosis technique is based on captured images of several parts of the human body and a computer-based program for identifying abnormal signs in these parts in lieu of the professional knowledge of doctors. This technique has been successful in many applications, such as brain [
6,
7,
8,
9,
10], breast [
11,
12,
13,
14,
15,
16,
17], lung [
18,
19,
20], and thyroid [
21,
22,
23,
24,
25,
26,
27,
28,
29,
30,
31,
32,
33,
34,
35,
36,
37,
38,
39,
40,
41,
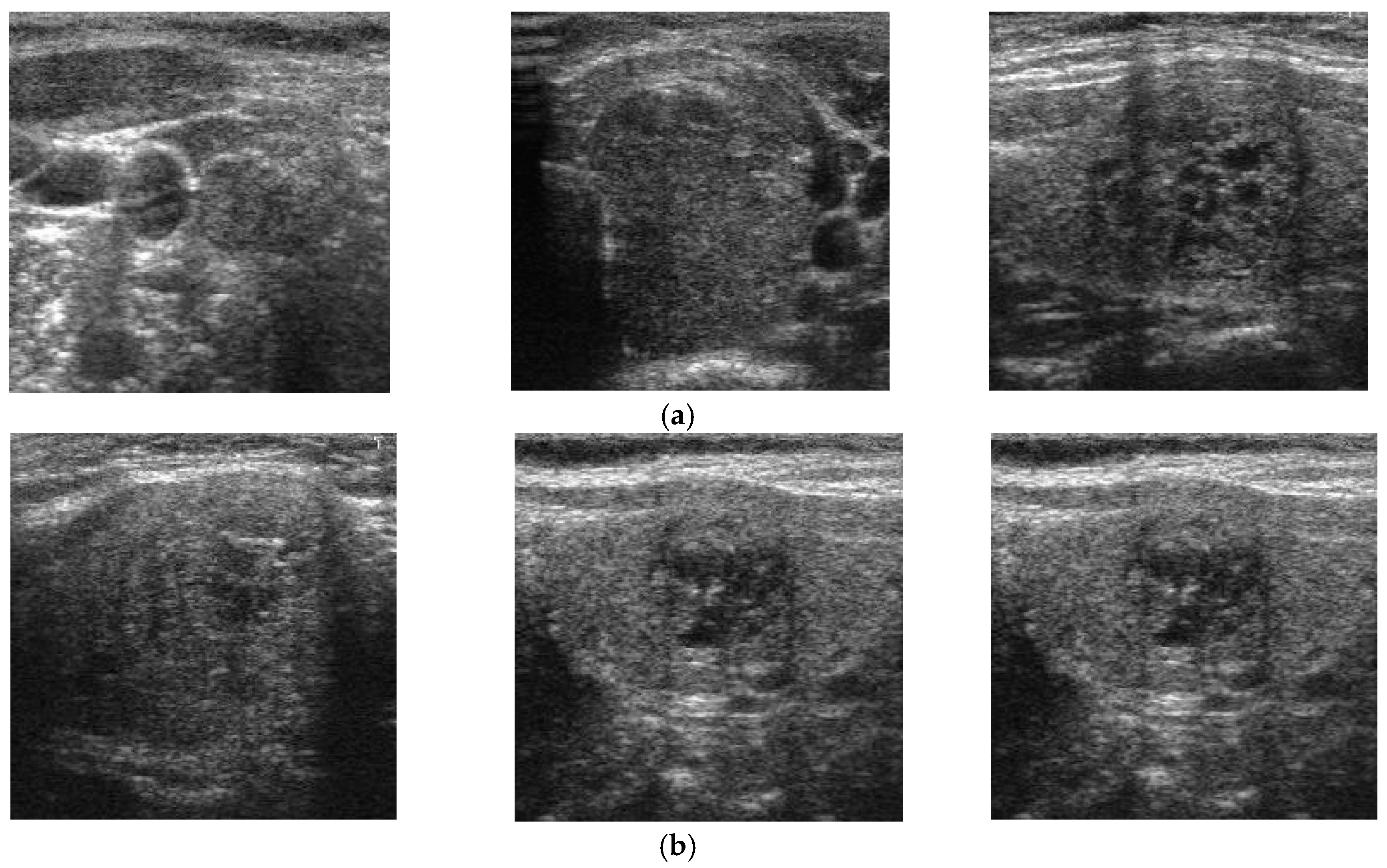
42] nodule detection/classification problems. For diagnosing thyroid nodules, previous studies focused on designing computer-based systems to perform several functions to detect and/or classify thyroid images into several classes such as nodule versus non-nodule; benign versus malign; follicles versus fibrosis, etc. In our research, we focus on the classification of benign and malign cases of thyroid nodules.
Thyroid nodules are abnormal regions (lumps) that appear in the thyroid region of adult humans and may be an indicator of thyroid cancer [
21,
35]. Based on their characteristics, thyroid nodules are classified into two kinds: benign (negative or non-cancerous nodules that are not harmful to health) and malign (positive or cancerous nodules that can cause health problems). Fortunately, most detected thyroid nodules are benign [
22]. However, with the appearance of thyroid nodules, patients may be confronted with various health or aesthetic problems. For example, large thyroid nodules, both benign and malign, may be visible and/or make it difficult for patients to breathe and swallow. More critically, malign thyroid nodules can produce an additional hormone called thyroxine, which causes some critical problems with patient’s health, including unexplained weight loss, tremors, and rapid heartbeat, and can also cause thyroid cancer. According to a report by the American Cancer Society, there were about 1950 patients estimated to have died among 62,450 new cases of thyroid cancer in 2015 [
21]. However, the mortality rate can be reduced with early detection and treatment. Furthermore, determining whether a thyroid nodule is benign or malign is a hard task for doctors when it is based only on symptoms or experience. Consequently, detection and classification methods for thyroid nodules are critical for diagnosing problems with the thyroid.
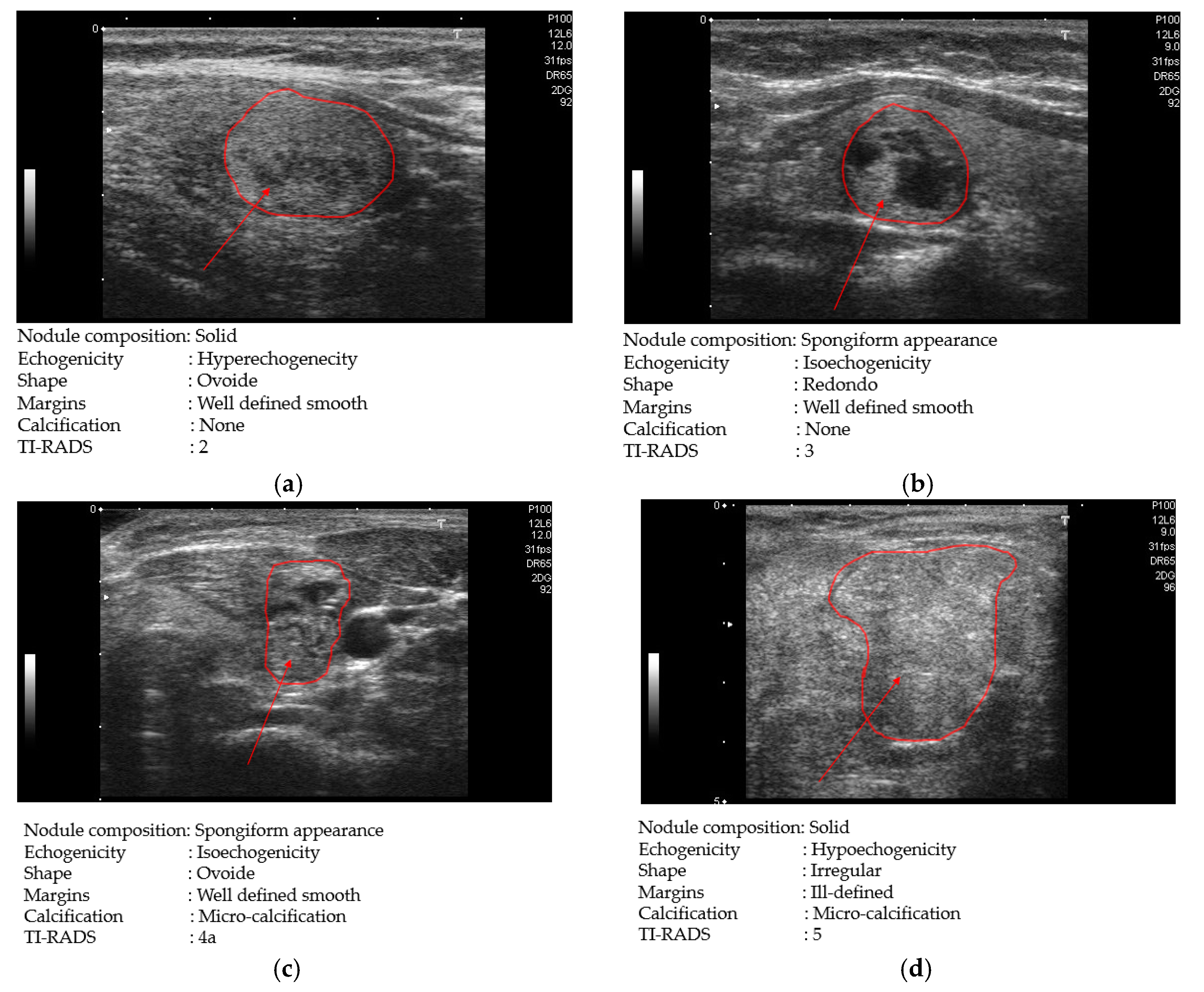
For thyroid nodule diagnosis, thyroid imaging reporting and data system (TI-RADS) is referred to as several risk stratification systems for thyroid lesions, and TI-RADS scores using the ultrasound image have been adopted to indicate whether the patient has a problem with their thyroid [
25]. In addition, methoxyisobutylisonitrile-single photon emission computed tomography (MIBI-SPECT) has been used for the diagnosis of thyroid nodules [
43]. As the other method of thyroid nodule diagnosis, a non-surgical diagnosis method called fine needle inspiration (FNA) biopsy has been widely used [
21,
22,
23,
24]. However, because thyroid nodules are highly complex, about 10%–20% of thyroid nodule biopsies are non-diagnostic [
22]. Furthermore, the main disadvantages of the FNA method are that it is labor-intensive, invasive, and costly, resulting in the CAD system becoming popular for assisting doctors in diagnosing thyroid nodules.
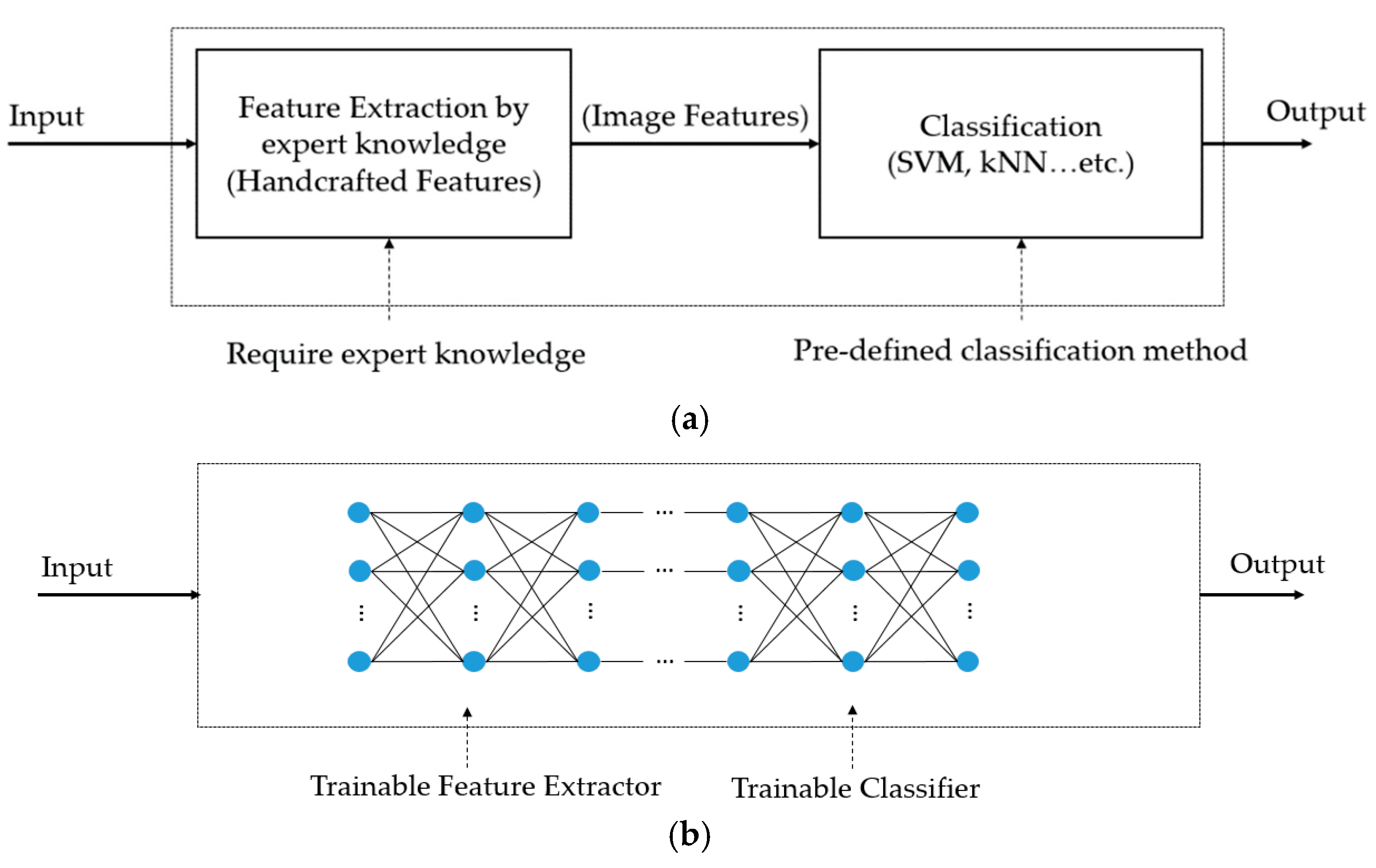
Previous studies on the thyroid nodule classification problem can be categorized into two groups: handcrafted-based and deep learning-based methods. In the first category, researchers used several traditional image feature extraction methods to extract efficient image features for the classification problem. In [
26], the authors used up to 78 textural features, such as the gray level co-occurrence matrix, statistical features, the gray level run-length matrix, and Law’s texture energy measures, to describe an input ultrasound thyroid image. These features are then inputted into a support vector machine (SVM) to classify the input image into several categories, such as nodule versus non-nodule and follicles versus fibrosis. In [
27], the authors showed the efficiency of a multi-scale features approach based on wavelet analysis for thyroid nodule classification. Based on the handcrafted image features, the authors in [
31] analyzed the performance of linear and non-linear classifiers for ultrasound thyroid nodule images. They showed that the two kinds of classification methods yield similar (comparable) levels of classification accuracy. Based on the characteristics of thyroid nodules, Raghavendra et al. [
35] found that ultrasound thyroid images are subjected to different threshold values to generate corresponding binary images. Based on this observation, they used segmentation-based fractal texture analysis (SFTA) method to extract image features from binarized images at different threshold values for the thyroid nodule classification problem. Furthermore, they combined the fractal texture features with spatial handcrafted texture features to enhance the performance of the classification system. Consequently, they showed that the handcrafted-based features are efficient for characterizing and categorizing thyroid nodules. However, the accuracy of this kind of methods is limited.
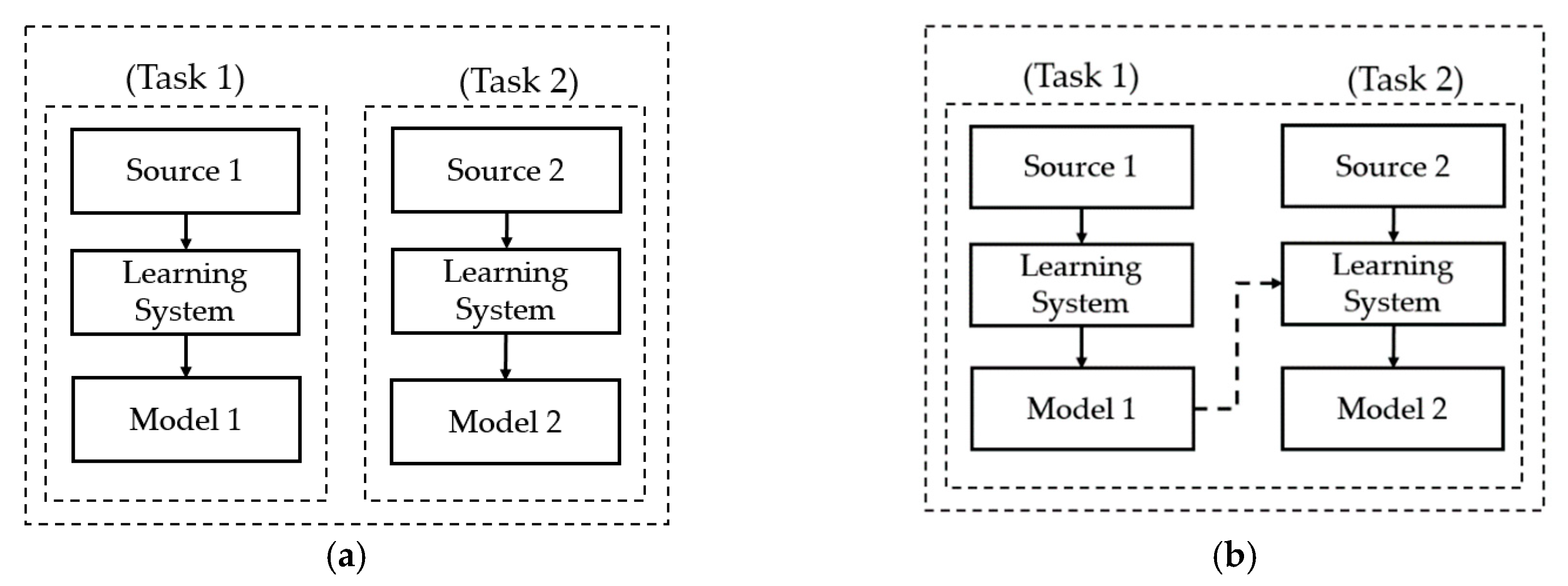
In the second category, deep learning-based methods, based on constructing a deep neural network for image classification, have been used to overcome the limitations of handcrafted-based methods. As explained in the preceding section, the handcrafted-based methods are based on pre-designed image feature extractors for extracting efficient image features and a classification method for classification. Thus, the performance of classification methods is dependent on the selection of image feature extractors, which usually affect only limited aspects of the thyroid nodule classification problem, and a classification method. By applying the deep learning-based method to the problem, these dependencies can be removed, and the feature extractors and classification methods are driven by training data. In methods of this type, researchers construct a deep learning-based model for various purposes, such as feature extraction, or the classification or detection of thyroid nodules in captured ultrasound images. In a study by Sundar et al. [
32], they proposed a framework for ultrasound thyroid nodule image classification based on the use of a convolutional neural network (CNN). In their study, they first used a pre-trained CNN network as an image feature extractor to extract the image features of an inputted ultrasound thyroid nodule image. Based on the extracted image features, they used the SVM method to classify the input images into benign or malign classes. In their second study, they also constructed a new CNN network for image classification by either training from scratch or using a transfer learning technique (using VGG16-Net [
44] or Inception-Net [
45]). Similar to the study by Sundar et al., the study by Song et al. [
34] also applied a transfer learning technique to successfully classify thyroid nodule images. Through their research, they demonstrated that CNN networks are suitable for thyroid nodule classification problems and produce good classification results. In recent studies by Song et al. [
30] and Wang et al. [
33], a detection and classification scheme was applied to the thyroid nodule classification problem. The rough position of possible nodule regions was first detected using either a multi-scale single-shot detection network (multi-scale SSD) or a YOLOv2 network. With the region detected, a fine classification step is performed to classify the nodule into a corresponding category, i.e., benign or malign. Although this approach can achieve high classification accuracy, it is time-consuming for nodule detection and requires a complex network design. In
Table 1, we summarize our approach for ultrasound thyroid image classification.
As a hybrid approach of the methods just mentioned, a combination of deep and handcrafted image features is considered for the thyroid nodule classification problem. In this study, we propose a new method that combines the deep and handcrafted features with four novelties, as follows:
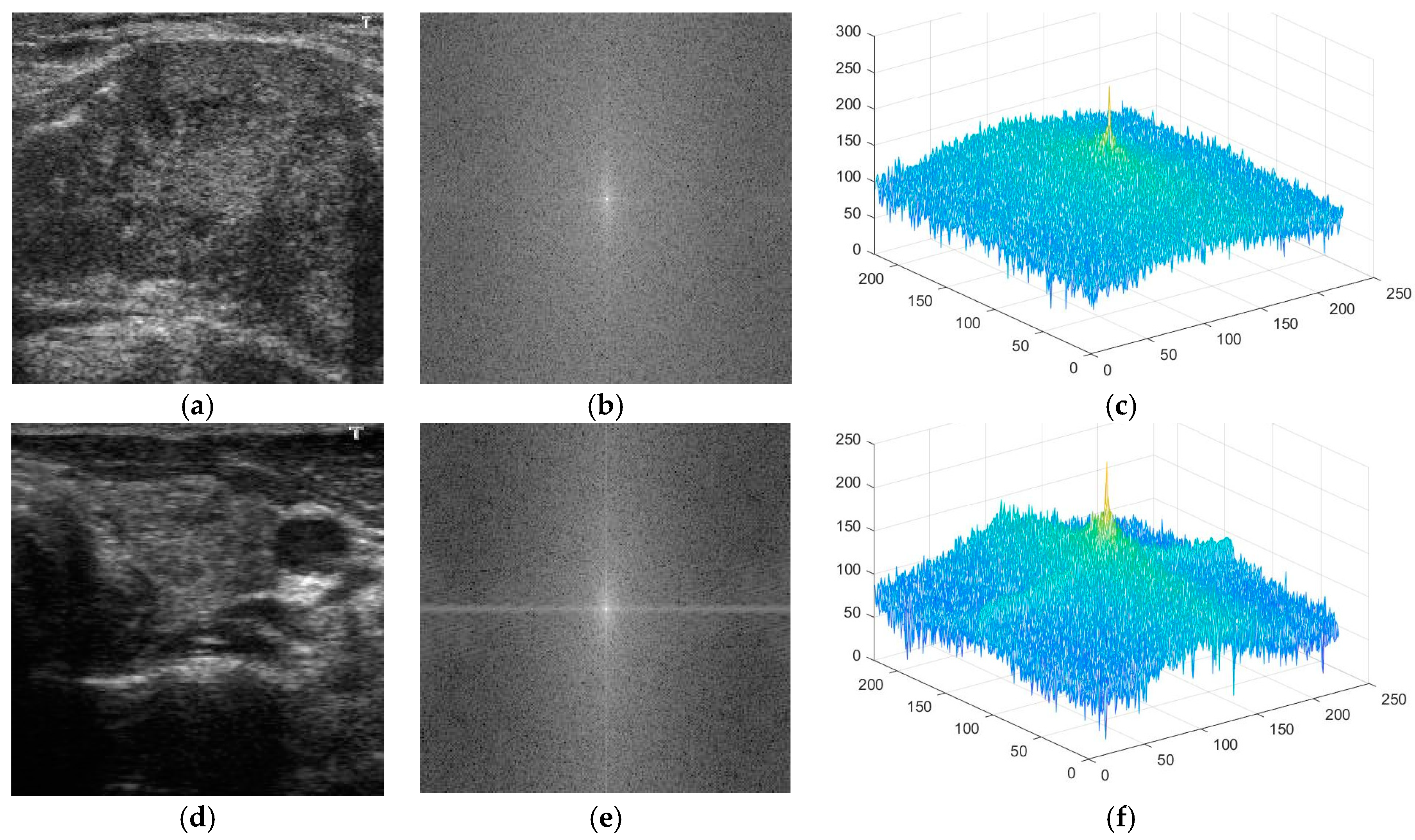
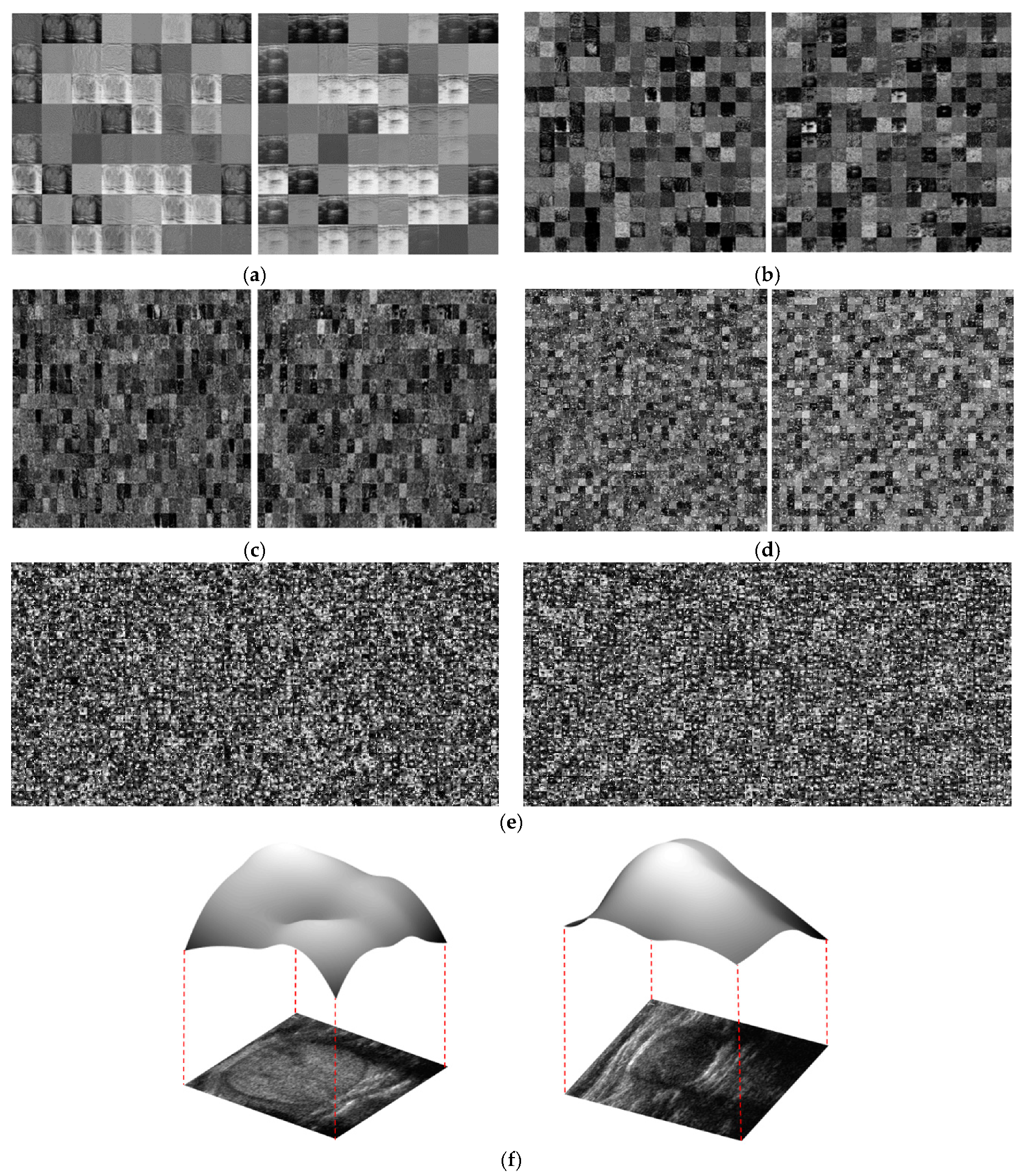
First, we propose the use of information in the frequency domain for thyroid nodule image classification. In contrast to most of the previous studies, which only consider information on thyroid nodule images in the spatial domain, our study also explores the utility of information in the frequency domain for the thyroid nodule classification problem, using the Fast Fourier Transform (FFT) method, based on our observation of the characteristics of benign and malign nodules.
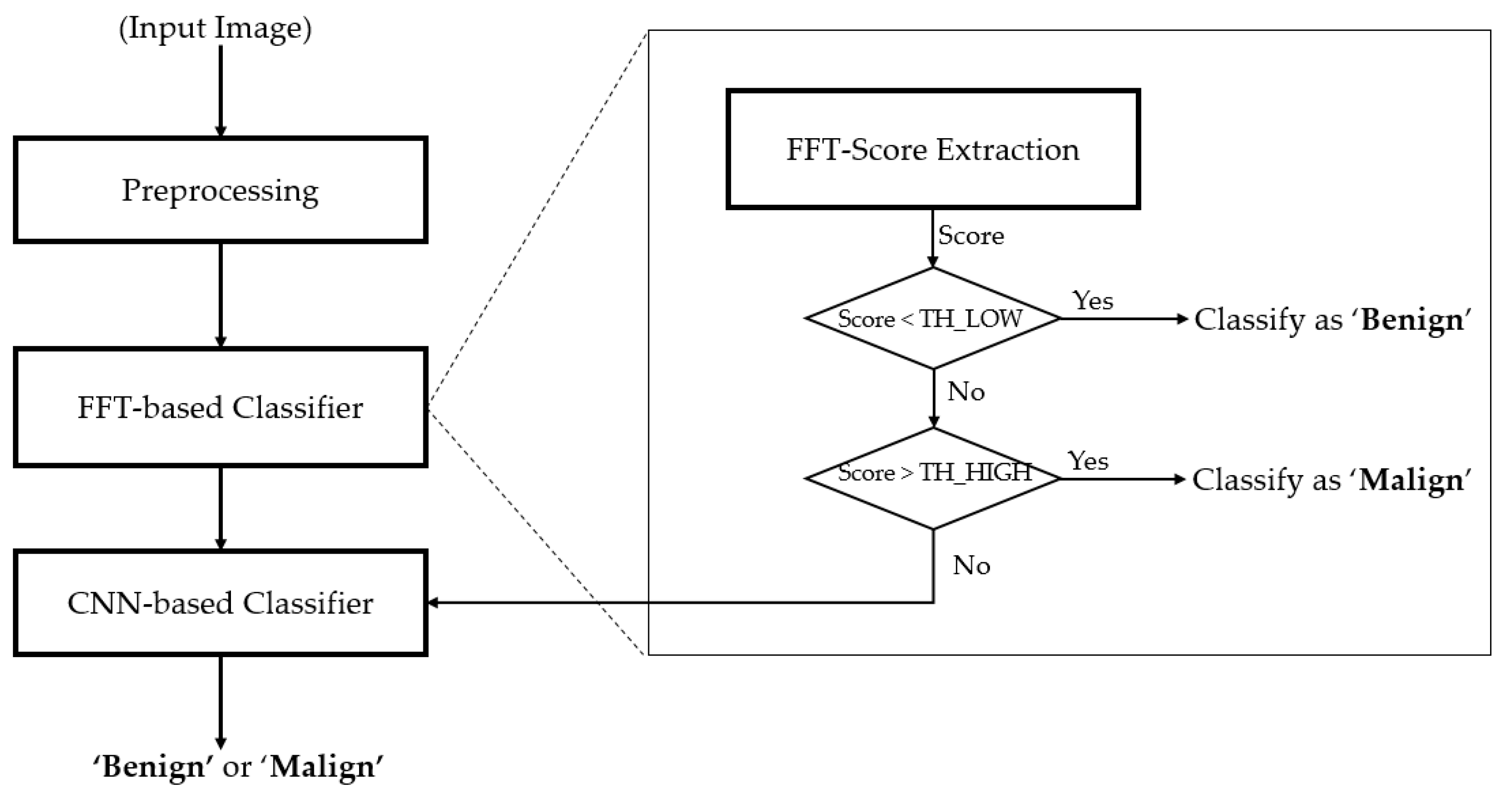
Secondly, we propose a method for extracting the information in the frequency domain to differentiate between benign and malign cases. Based on the result of this proposed method, we construct a rule for classifying input images into one of three groups: benign, benign-malign, or malign, as explained in detail in subsequent sections.
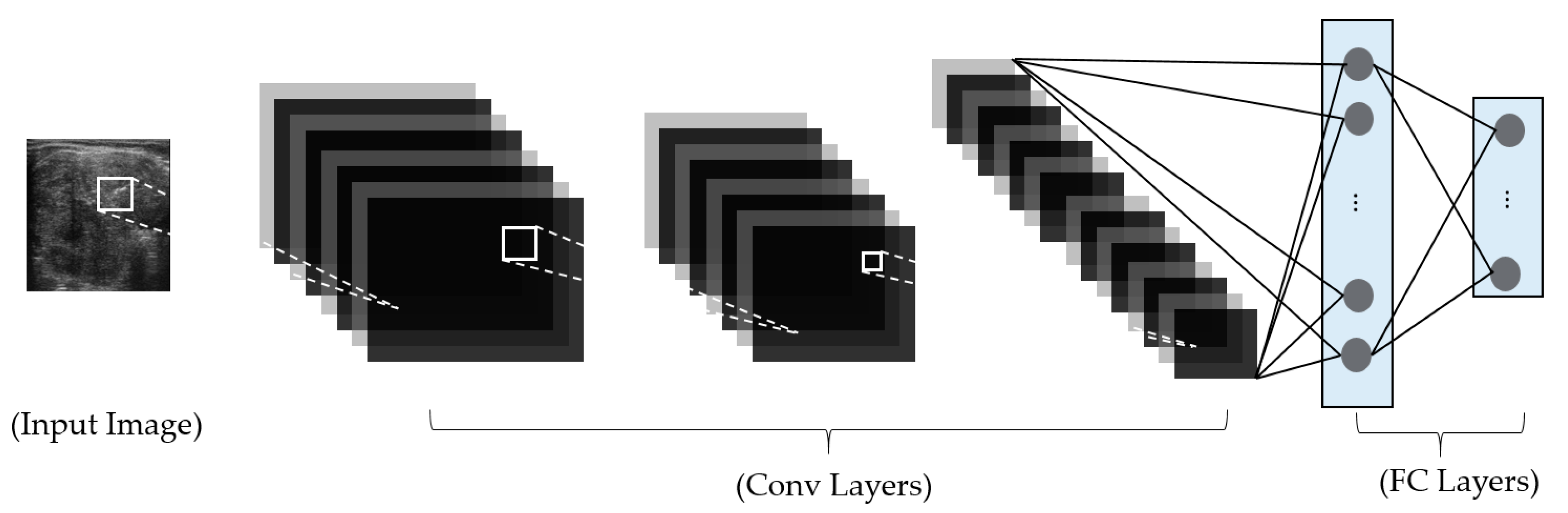
Thirdly, we propose a modified residual network (Resnet) by adopting global average pooling to summarize the feature maps of the last convolution layer, which reduces the number of network parameters, and by using batch normalization and dropout techniques to reduce the effects overfitting problem. Also, we combine the classification results using a deep learning-based method and a frequency-based method to enhance classification accuracy. For this purpose, we designed a cascade classifier classification system, based on the information extracted from the frequency and spatial domains.
Finally, we make our algorithm available to the public through [
46], so that other researchers can make fair assessments of our system.
In the rest of the paper, we first describe the proposed method for the thyroid nodule classification problem in
Section 2. Using the proposed method, we perform various experiments using the TDID dataset to evaluate the performance of our proposed approach, as well as compare it to previous studies in the literature in
Section 3. Finally, we give a conclusion of our study in
Section 4.
4. Conclusions
In this study, we proposed a thyroid nodule classification method using a cascade classifier scheme, based on the extracted information in both the spatial and frequency domains of an ultrasound thyroid image. Our proposed method processes an ultrasound image of the thyroid region to discriminate benign and malign cases in thyroid nodules. For this purpose, we first applied a classifier for the classification problem using image features in the frequency domain, which has not been done in previous studies. Secondly, the ambiguous samples with the first classifier are then further classified using a deep learning-based method. Through our expensive experiments, we show that our proposed method achieves a higher classification accuracy than various previous studies using the TDID dataset. Our algorithm is designed to provide more accurate suggestion to doctors (radiologists) in diagnosing thyroid nodules and reduce the number of operations on benign thyroid nodules. As a result, it is applicable in hospitals in daily routine. To use our algorithm requires a conventional desktop computer with a monitor and a commercial graphics processing unit (GPU) card for running deep neural networks, which costs less than $1500.
As future work, we would apply our algorithm in real applications, with the help of doctors, to enhance the performance of our algorithm, and make it applicable with the daily routine in hospitals. In addition, we would expand our work to include non-malignant thyroid pathologies, like Hashimoto and Graves’ disease patients, by collecting more data with the help of doctors in hospitals.