The Impact of Cholecystectomy on the Gut Microbiota: A Case-Control Study
Abstract
:1. Introduction
2. Materials and Methods
2.1. Subjects
2.2. Fecal Samples and DNA Extraction
2.3. PCR Amplification and Sequencing of the Bacterial 16S rRNA Gene
2.4. 16S rRNA Gene Compositional Analysis
2.5. Statistical Analysis
3. Results
3.1. Demographic Characteristics
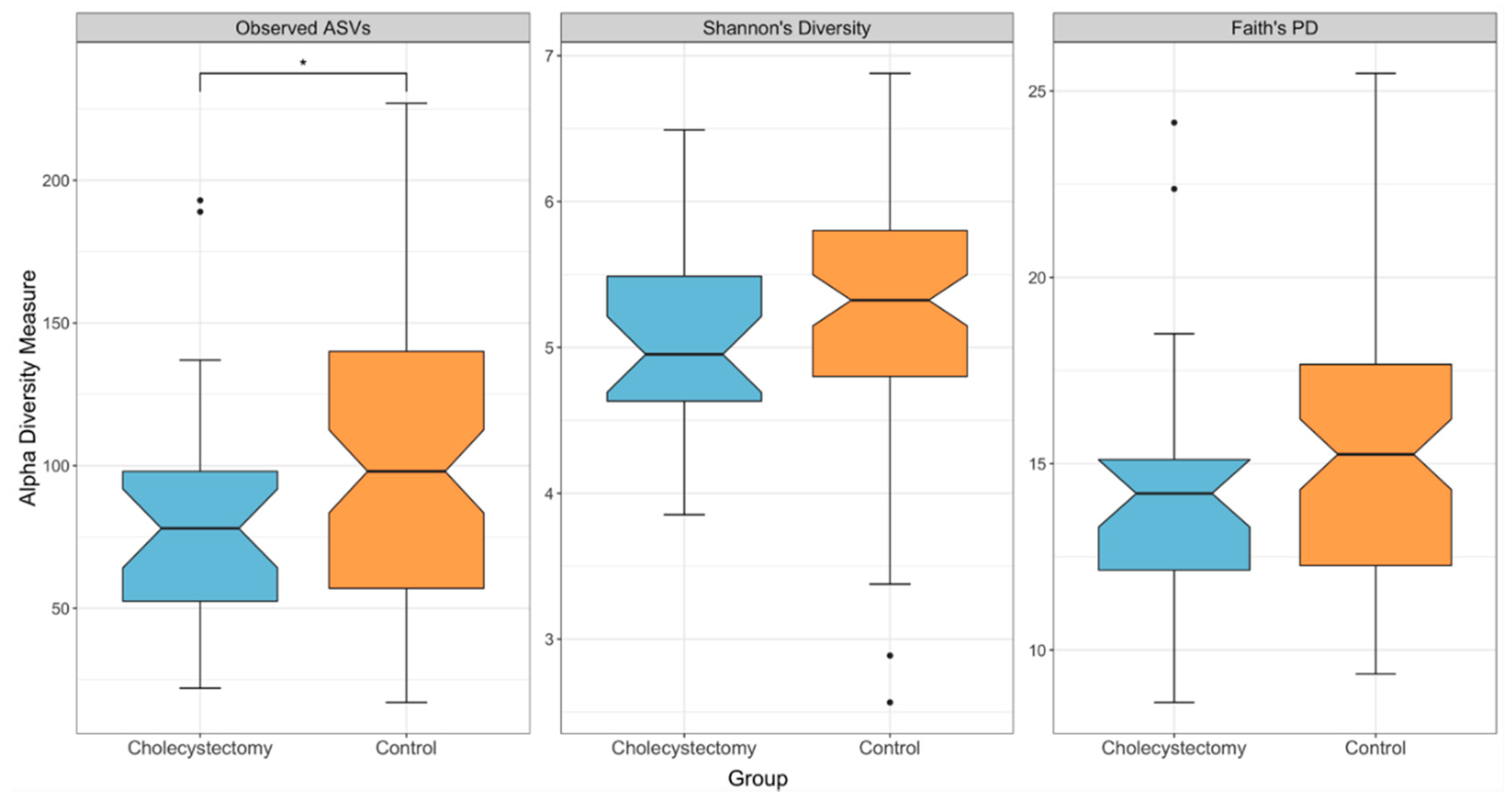
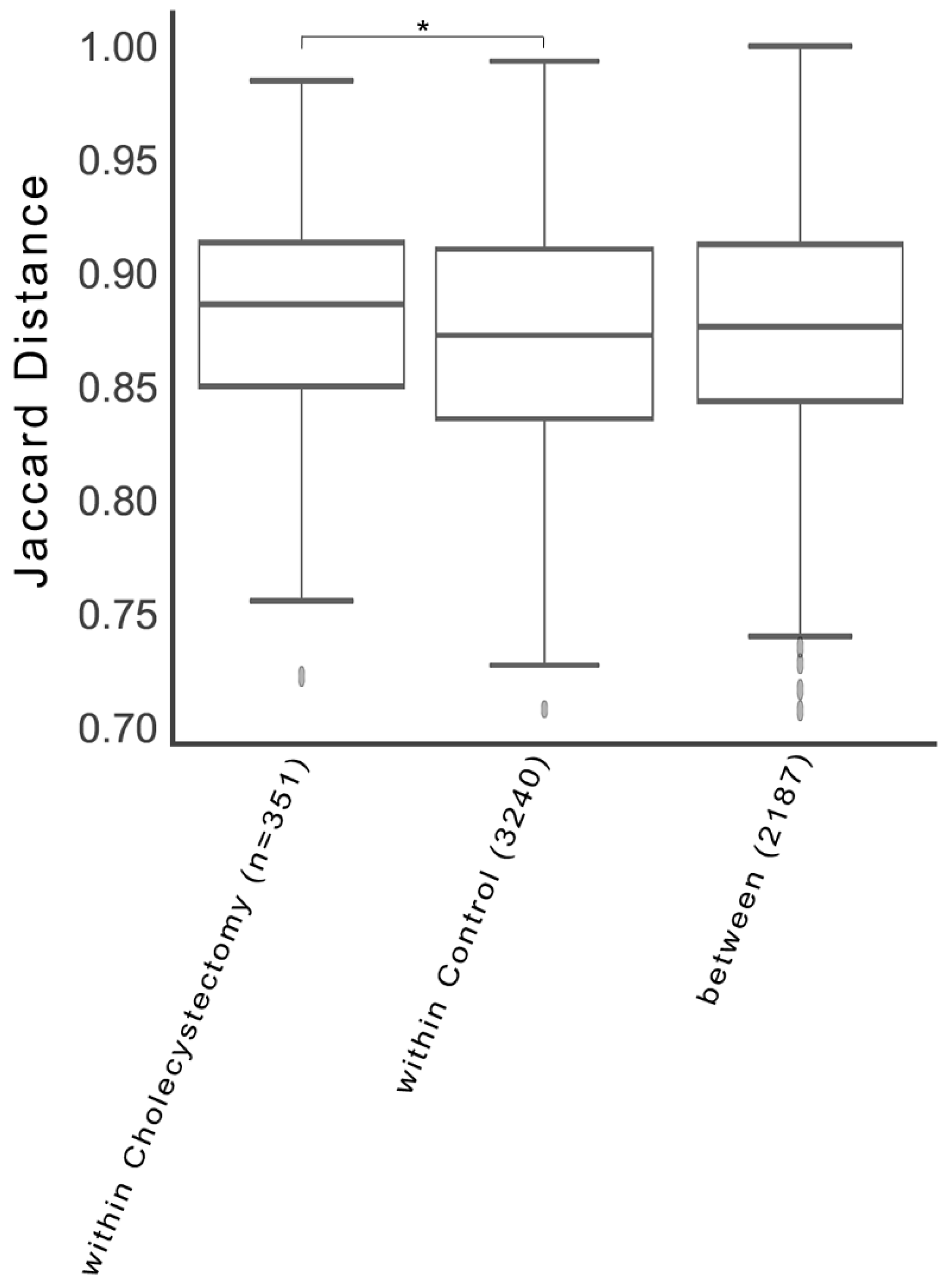
3.2. Gut Microbial Diversity
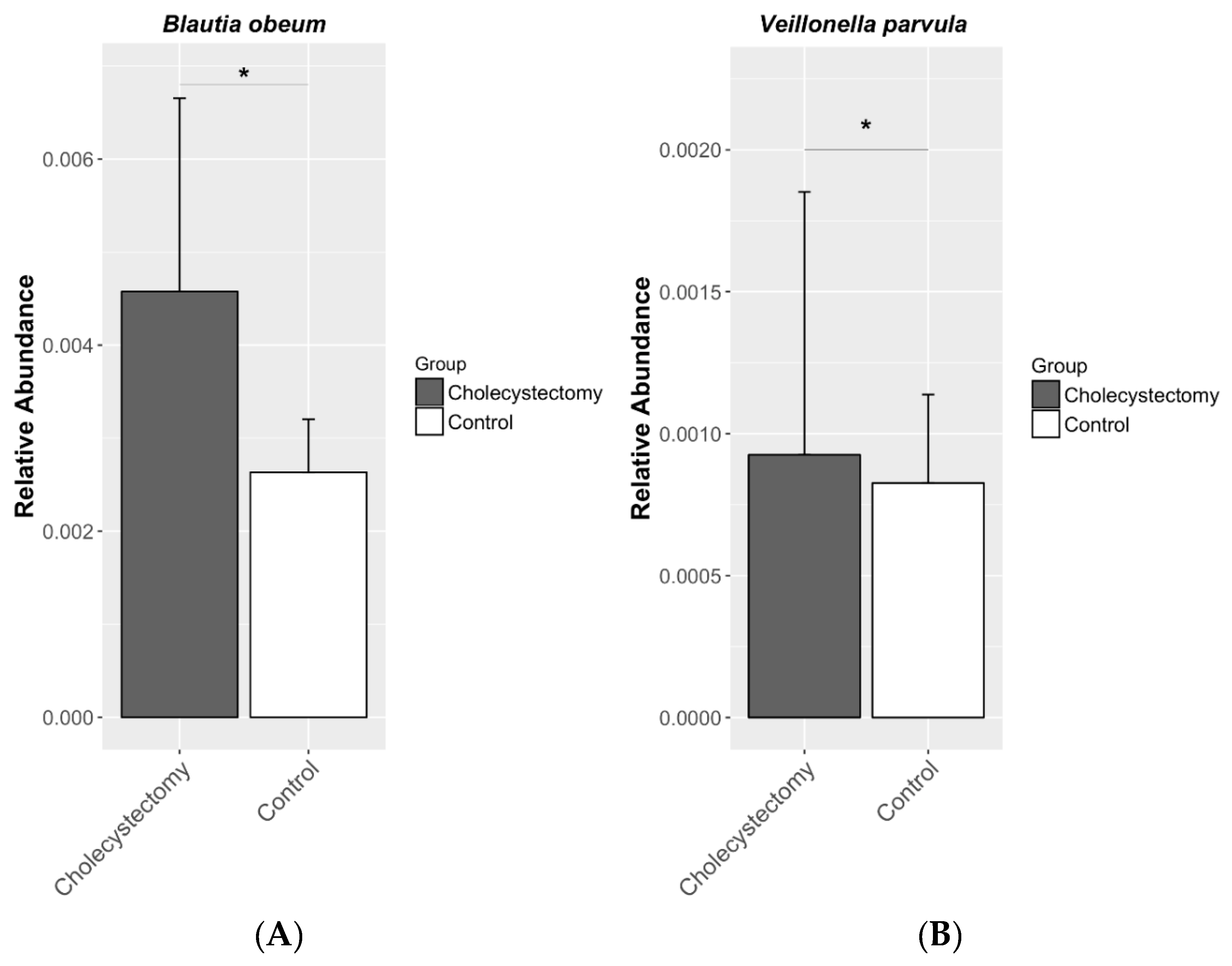
3.3. Gut Microbiota Composition
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Availability of Data and Materials
References
- Housset, C.; Chretien, Y.; Debray, D.; Chignard, N. Functions of the Gallbladder. Compr. Physiol. 2016, 6, 1549–1577. [Google Scholar] [CrossRef] [PubMed]
- Weiss, A.J.; Elixhauser, A.; Andrews, R.M. Characteristics of Operating Room Procedures in U.S. Hospitals, 2011. Available online: https://www.hcup-us.ahrq.gov/reports/statbriefs/sb170-Operating-Room-Procedures-United-States-2011.jsp (accessed on 8 February 2018).
- Clavien, P.-A.; Baillie, J. Diseases of the Gallbladder and Bile Ducts: Diagnosis and Management, 2nd ed.; Blackwell Publishing: Malden, MA, USA, 2006. [Google Scholar]
- Hepner, G.W.; Hofmann, A.F.; Malagelada, J.R.; Szczepanik, P.A.; Klein, P.D. Increased bacterial degradation of bile acids in cholecystectomized patients. Gastroenterology 1974, 66, 556–564. [Google Scholar] [PubMed]
- Malagelada, J.R.; Go, V.L.; Summerskill, W.H.; Gamble, W.S. Bile acid secretion and biliary bile acid composition altered by cholecystectomy. Am. J. Dig. Dis. 1973, 18, 455–459. [Google Scholar] [CrossRef]
- Pomare, E.W.; Heaton, K.W. The effect of cholecystectomy on bile salt metabolism. Gut 1973, 14, 753–762. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Roda, E.; Aldini, R.; Mazzella, G.; Roda, A.; Sama, C.; Festi, D.; Barbara, L. Enterohepatic circulation of bile acids after cholecystectomy. Gut 1978, 19, 640–649. [Google Scholar] [CrossRef]
- Linos, D.; Beard, C.M.; O’Fallon, W.M.; Dockerty, M.B.; Beart, R.W., Jr.; Kurland, L.T. Cholecystectomy and carcinoma of the colon. Lancet (Lond. Engl.) 1981, 2, 379–381. [Google Scholar] [CrossRef]
- Rundgren, A.; Mellstrom, D. Cholecystectomy and colon cancer in the elderly. Age Ageing 1983, 12, 44–49. [Google Scholar] [CrossRef]
- Nielsen, G.P.; Theodors, A.; Tulinius, H.; Sigvaldason, H. Cholecystectomy and colorectal carcinoma: A total-population historical prospective study. Am. J. Gastroenterol. 1991, 86, 1486–1490. [Google Scholar]
- Ekbom, A.; Yuen, J.; Adami, H.O.; McLaughlin, J.K.; Chow, W.H.; Persson, I.; Fraumeni, J.F., Jr. Cholecystectomy and colorectal cancer. Gastroenterology 1993, 105, 142–147. [Google Scholar] [CrossRef]
- Goldbohm, R.A.; van den Brandt, P.A.; van’t Veer, P.; Dorant, E.; Sturmans, F.; Hermus, R.J. Cholecystectomy and colorectal cancer: Evidence from a cohort study on diet and cancer. Int. J. Cancer 1993, 53, 735–739. [Google Scholar] [CrossRef]
- Lagergren, J.; Ye, W.; Ekbom, A. Intestinal cancer after cholecystectomy: Is bile involved in carcinogenesis? Gastroenterology 2001, 121, 542–547. [Google Scholar] [CrossRef] [PubMed]
- Shao, T.; Yang, Y.X. Cholecystectomy and the risk of colorectal cancer. Am. J. Gastroenterol. 2005, 100, 1813–1820. [Google Scholar] [CrossRef] [PubMed]
- Goldacre, M.J.; Wotton, C.J.; Abisgold, J.; Yeates, D.G.; Collins, J. Association between cholecystectomy and intestinal cancer: A national record linkage study. Ann. Surg. 2012, 256, 1068–1072. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Jiao, L.; He, C.; Sun, H.; Cai, Q.; Han, T.; Hu, H. Alteration of gut microbiota in association with cholesterol gallstone formation in mice. BMC Gastroenterol. 2017, 17, 74. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wu, T.; Zhang, Z.; Liu, B.; Hou, D.; Liang, Y.; Zhang, J.; Shi, P. Gut microbiota dysbiosis and bacterial community assembly associated with cholesterol gallstones in large-scale study. BMC Genom. 2013, 14, 669. [Google Scholar] [CrossRef] [PubMed]
- Gao, R.; Gao, Z.; Huang, L.; Qin, H. Gut microbiota and colorectal cancer. Eur. J. Clin. Microbiol. Infect. Dis. 2017, 36, 757–769. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Keren, N.; Konikoff, F.M.; Paitan, Y.; Gabay, G.; Reshef, L.; Naftali, T.; Gophna, U. Interactions between the intestinal microbiota and bile acids in gallstones patients. Environ. Microbiol. Rep. 2015, 7, 874–880. [Google Scholar] [CrossRef]
- Fadrosh, D.W.; Ma, B.; Gajer, P.; Sengamalay, N.; Ott, S.; Brotman, R.M.; Ravel, J. An improved dual-indexing approach for multiplexed 16S rRNA gene sequencing on the Illumina MiSeq platform. Microbiome 2014, 2, 6. [Google Scholar] [CrossRef]
- Kozich, J.J.; Westcott, S.L.; Baxter, N.T.; Highlander, S.K.; Schloss, P.D. Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl. Environ. Microbiol. 2013, 79, 5112–5120. [Google Scholar] [CrossRef]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef] [Green Version]
- Caporaso, J.G.; Kuczynski, J.; Stombaugh, J.; Bittinger, K.; Bushman, F.D.; Costello, E.K.; Fierer, N.; Pena, A.G.; Goodrich, J.K.; Gordon, J.I.; et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 2010, 7, 335–336. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jost, L. Partitioning diversity into independent alpha and beta components. Ecology 2007, 88, 2427–2439. [Google Scholar] [CrossRef] [PubMed]
- Faith, D.P.; Baker, A.M. Phylogenetic diversity (PD) and biodiversity conservation: Some bioinformatics challenges. Evol. Bioinform. Online 2007, 2, 121–128. [Google Scholar] [CrossRef] [PubMed]
- Lozupone, C.; Lladser, M.E.; Knights, D.; Stombaugh, J.; Knight, R. UniFrac: An effective distance metric for microbial community comparison. ISME J. 2011, 5, 169–172. [Google Scholar] [CrossRef] [PubMed]
- Bray, J.R.; Curtis, J.T. An ordination of the upland forest communities of Southern Wisconsin. Ecol. Monogr. 1957, 27, 325–349. [Google Scholar] [CrossRef]
- Morgan, X.C.; Tickle, T.L.; Sokol, H.; Gevers, D.; Devaney, K.L.; Ward, D.V.; Reyes, J.A.; Shah, S.A.; LeLeiko, N.; Snapper, S.B.; et al. Dysfunction of the intestinal microbiome in inflammatory bowel disease and treatment. Genome Biol. 2012, 13, R79. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Wang, J.; Li, J.; Yan, P.; Jin, Y.; Zhang, R.; Yue, W.; Guo, Q.; Geng, J. Cholecystectomy Damages Aging-Associated Intestinal Microbiota Construction. Front. Microbiol. 2018, 9, 1402. [Google Scholar] [CrossRef] [PubMed]
- Lawson, P.A.; Finegold, S.M. Reclassification of Ruminococcus obeum as Blautia obeum comb. nov. Int. J. Syst. Evol. Microbiol. 2015, 65, 789–793. [Google Scholar] [CrossRef] [PubMed]
- Theriot, C.M.; Bowman, A.A.; Young, V.B. Antibiotic-Induced Alterations of the Gut Microbiota Alter Secondary Bile Acid Production and Allow for Clostridium difficile Spore Germination and Outgrowth in the Large Intestine. mSphere 2016, 1. [Google Scholar] [CrossRef]
- La Reau, A.J.; Meier-Kolthoff, J.P.; Suen, G. Sequence-based analysis of the genus Ruminococcus resolves its phylogeny and reveals strong host association. Microb. Genom. 2016, 2, e000099. [Google Scholar] [CrossRef] [PubMed]
- Youssef, O.; Lahti, L.; Kokkola, A.; Karla, T.; Tikkanen, M.; Ehsan, H.; Carpelan-Holmstrom, M.; Koskensalo, S.; Bohling, T.; Rautelin, H.; et al. Stool Microbiota Composition Differs in Patients with Stomach, Colon, and Rectal Neoplasms. Dig. Dis. Sci. 2018. [Google Scholar] [CrossRef] [PubMed]
- Allali, I.; Boukhatem, N.; Bouguenouch, L.; Hardi, H.; Boudouaya, H.A.; Cadenas, M.B.; Ouldim, K.; Amzazi, S.; Azcarate-Peril, M.A.; Ghazal, H. Gut microbiome of Moroccan colorectal cancer patients. Med. Microbiol. Immunol. 2018, 207, 211–225. [Google Scholar] [CrossRef] [PubMed]
- Poppleton, D.I.; Duchateau, M.; Hourdel, V.; Matondo, M.; Flechsler, J.; Klingl, A.; Beloin, C.; Gribaldo, S. Outer Membrane Proteome of Veillonella parvula: A Diderm Firmicute of the Human Microbiome. Front. Microbiol. 2017, 8, 1215. [Google Scholar] [CrossRef] [PubMed]
- Dewhirst, F.E.; Chen, T.; Izard, J.; Paster, B.J.; Tanner, A.C.; Yu, W.H.; Lakshmanan, A.; Wade, W.G. The human oral microbiome. J. Bacteriol. 2010, 192, 5002–5017. [Google Scholar] [CrossRef]
- Valm, A.M.; Mark Welch, J.L.; Rieken, C.W.; Hasegawa, Y.; Sogin, M.L.; Oldenbourg, R.; Dewhirst, F.E.; Borisy, G.G. Systems-level analysis of microbial community organization through combinatorial labeling and spectral imaging. Proc. Natl. Acad. Sci. USA 2011, 108, 4152–4157. [Google Scholar] [CrossRef] [Green Version]
- Perez-Jacoiste Asin, M.A.; Fernandez-Ruiz, M.; Serrano-Navarro, I.; Prieto-Rodriguez, S.; Aguado, J.M. Polymicrobial endocarditis involving Veillonella parvula in an intravenous drug user: Case report and literature review of Veillonella endocarditis. Infection 2013, 41, 591–594. [Google Scholar] [CrossRef] [PubMed]
- Boo, T.W.; Cryan, B.; O’Donnell, A.; Fahy, G. Prosthetic valve endocarditis caused by Veillonella parvula. J. Infect. 2005, 50, 81–83. [Google Scholar] [CrossRef] [PubMed]
- Berenger, B.M.; Chui, L.; Borkent, A.; Lee, M.C. Anaerobic urinary tract infection caused by Veillonella parvula identified using cystine-lactose-electrolyte deficient media and matrix-assisted laser desorption ionization-time of flight mass spectrometry. IDCases 2015, 2, 44–46. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, Y.C.; Ko, P.H.; Yang, C.J.; Chen, Y.C.; Lay, C.J.; Tsai, C.C.; Hsieh, M.H. Epidural abscess caused by Veillonella parvula: Case report and review of the literature. J. Microbiol. Immunol. Infect. 2016, 49, 804–808. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bhatti, M.A.; Frank, M.O. Veillonella parvula meningitis: Case report and review of Veillonella infections. Clin. Infect. Dis. 2000, 31, 839–840. [Google Scholar] [CrossRef] [PubMed]
- Nukina, S.; Hibi, A.; Nishida, K. Bacterial meningitis caused by Veillonella parvula. Acta Paediatr. Jpn. 1989, 31, 609–614. [Google Scholar] [CrossRef] [PubMed]
- Fisher, R.G.; Denison, M.R. Veillonella parvula bacteremia without an underlying source. J. Clin. Microbiol. 1996, 34, 3235–3236. [Google Scholar] [PubMed]
- Singh, N.; Yu, V.L. Osteomyelitis due to Veillonella parvula: Case report and review. Clin. Infect. Dis. 1992, 14, 361–363. [Google Scholar] [CrossRef] [PubMed]
- Marriott, D.; Stark, D.; Harkness, J. Veillonella parvula discitis and secondary bacteremia: A rare infection complicating endoscopy and colonoscopy? J. Clin. Microbiol. 2007, 45, 672–674. [Google Scholar] [CrossRef]
- Malinen, E.; Rinttila, T.; Kajander, K.; Matto, J.; Kassinen, A.; Krogius, L.; Saarela, M.; Korpela, R.; Palva, A. Analysis of the fecal microbiota of irritable bowel syndrome patients and healthy controls with real-time PCR. Am. J. Gastroenterol. 2005, 100, 373–382. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Liu, H.; Li, L.; Ai, M.; Gong, Z.; He, Y.; Dong, Y.; Xu, S.; Wang, J.; Jin, B.; et al. Cholecystectomy can increase the risk of colorectal cancer: A meta-analysis of 10 cohort studies. PLoS ONE 2017, 12, e0181852. [Google Scholar] [CrossRef] [PubMed]
- Ajouz, H.; Mukherji, D.; Shamseddine, A. Secondary bile acids: An underrecognized cause of colon cancer. World J. Surg. Oncol. 2014, 12, 164. [Google Scholar] [CrossRef]
- Azcarate-Peril, M.A.; Sikes, M.; Bruno-Barcena, J.M. The intestinal microbiota, gastrointestinal environment and colorectal cancer: A putative role for probiotics in prevention of colorectal cancer? Am. J. Physiol. Gastrointest. Liver Physiol. 2011, 301, G401–G424. [Google Scholar] [CrossRef]
| Characteristics | Cholecystectomy Group | Control Group | Missing, n | p-Value |
|---|---|---|---|---|
| Number (n) | 27 | 81 | ||
| Age (years) | 48 (39–64) | 47 (38–64) | 0.823 | |
| Sex (male:female) | 15:12 | 45:36 | 1.000 | |
| BMI (kg/m2) | 23.8 (19.4–30.2) | 23.3 (18.7–29.5) | 0.321 | |
| Smoking history (never:former:current) | 15:7:2 | 50:17:10 | 7 | 0.778 * |
| Frequency of alcohol consumption (days/week) | 1 (0–6) | 1 (0–7) | 10 | 0.543 |
| Carbohydrate intake (g/day) | 221.1 (24.6–477.5) | 250.6 (0.9–474.9) | 29 | 0.673 |
| Fat intake (g/day) | 19.5 (6.8–72.6) | 23.5 (0–68.9) | 29 | 0.653 |
| Protein intake (g/day) | 40.1 (12.9–108.5) | 46.1 (0.3–107.3) | 29 | 0.748 |
| Characteristics | Cholecystectomy Group | Control Group | Missing, n | p-Value |
|---|---|---|---|---|
| Number (n) | 27 | 81 | ||
| White blood cells (×103/mm3) | 5.5 (3.9–8.8) | 5.7 (2.3–10.5) | 0.273 | |
| Hemoglobin (g/dL) | 13.8 (10.3–17.3) | 14.2 (6.9–16.7) | 0.823 | |
| Platelets (×103/mm3) | 234 (162–383) | 238 (79–465) | 0.859 | |
| Total cholesterol (mg/dL) | 185 (144–269) | 194 (126–275) | 0.333 | |
| Triglycerides (mg/dL) | 108 (32–256) | 96 (34–432) | 0.731 | |
| HDL-C (mg/dL) | 55 (33–98) | 56 (30–121) | 0.887 | |
| LDL-C (mg/dL) | 114 (85–199) | 118.5 (60–194) | 1 | 0.268 |
| Total protein (g/dL) | 7.2 (6.7–7.8) | 7.1 (6.4–8.3) | 0.751 | |
| Bilirubin (mg/dL) | 0.9 (0.3–1.8) | 0.8 (0.3–2.1) | 0.516 | |
| AST (IU/L) | 20 (13–39) | 19 (12–32) | 0.127 | |
| ALT (IU/L) | 18 (9–65) | 16 (7–40) | 0.012 | |
| GGT (IU/L) | 26 (6–180) | 17.5 (6–81) | 1 | 0.283 |
| BUN (mg/dL) | 13.8 (8.1–19.3) | 14.1 (6.7–25.1) | 0.717 | |
| Creatinine (mg/dL) | 0.8 (0.6–1.2) | 0.9 (0.5–1.2) | 0.635 | |
| Glucose (mg/dL) | 94 (76–136) | 92 (75–130) | 0.842 | |
| Hemoglobin A1c (%) | 5.5 (5.1–7.5) | 5.5 (5.1–6.8) | 0.225 |
| Taxonomic Level | Taxa | CE ¶ | Odds ¶ | n | n not 0 | p-Value | q-Value |
|---|---|---|---|---|---|---|---|
| Family | p_Bacteroidetes; c_Bacteroidia; o_Bacteroidales; f_S24_7 | −0.558 | 0.572 | 108 | 42 | 0.004 | 0.061 |
| Family | p_Firmicutes; c_Bacilli; o_Lactobacillales; f_Lactobacillaceae | 0.504 | 1.655 | 108 | 33 | 0.004 | 0.061 |
| Genus | p_Firmicutes; c_Clostridia; o_Clostridiales; f_Lachnospiraceae; g_Ruminococcus | 0.344 | 1.411 | 108 | 89 | 0.005 | 0.210 |
| Species | p_Firmicutes; c_Clostridia; o_Clostridiales; f_Lachnospiraceae; g_Blautia; s_obeum | 0.457 | 1.579 | 108 | 41 | 0.001 | 0.025 |
| Species | p_Firmicutes; c_Clostridia;o_Clostridiales; f_Veillonellaceae; g_Veillonella; s_parvula | 0.980 | 2.664 | 108 | 13 | 0.003 | 0.044 |
| Species | p_Actinobacteria; c_Actinobacteria; o_Bifidobacteriales; f_Bifidobacteriaceae; g_Bifidobacterium; s_adolescentis | 0.476 | 1.610 | 108 | 44 | 0.009 | 0.074 |
| Species | p_Firmicutes; c_Clostridia; o_Clostridiales; f_Lachnospiraceae; g_Ruminococcus; s_torques | 0.477 | 1.611 | 108 | 41 | 0.009 | 0.074 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yoon, W.J.; Kim, H.-N.; Park, E.; Ryu, S.; Chang, Y.; Shin, H.; Kim, H.-L.; Yi, S.Y. The Impact of Cholecystectomy on the Gut Microbiota: A Case-Control Study. J. Clin. Med. 2019, 8, 79. https://doi.org/10.3390/jcm8010079
Yoon WJ, Kim H-N, Park E, Ryu S, Chang Y, Shin H, Kim H-L, Yi SY. The Impact of Cholecystectomy on the Gut Microbiota: A Case-Control Study. Journal of Clinical Medicine. 2019; 8(1):79. https://doi.org/10.3390/jcm8010079
Chicago/Turabian StyleYoon, Won Jae, Han-Na Kim, Eunkyo Park, Seungho Ryu, Yoosoo Chang, Hocheol Shin, Hyung-Lae Kim, and Sun Young Yi. 2019. "The Impact of Cholecystectomy on the Gut Microbiota: A Case-Control Study" Journal of Clinical Medicine 8, no. 1: 79. https://doi.org/10.3390/jcm8010079
APA StyleYoon, W. J., Kim, H.-N., Park, E., Ryu, S., Chang, Y., Shin, H., Kim, H.-L., & Yi, S. Y. (2019). The Impact of Cholecystectomy on the Gut Microbiota: A Case-Control Study. Journal of Clinical Medicine, 8(1), 79. https://doi.org/10.3390/jcm8010079






