Urinary MicroRNA Profiling Predicts the Development of Microalbuminuria in Patients with Type 1 Diabetes
Abstract
:1. Introduction
2. Experimental Section
2.1. Patients and Samples
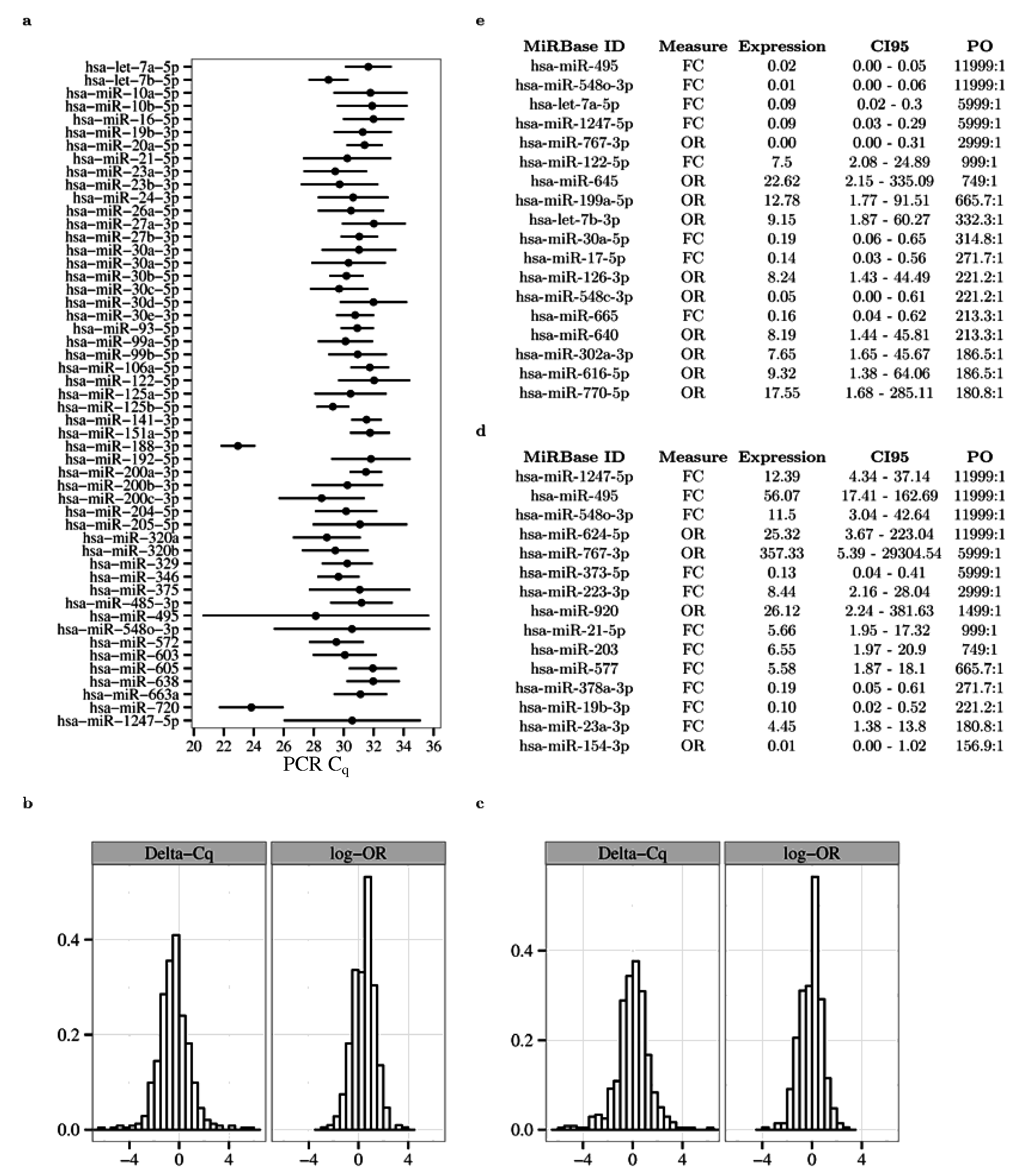
2.2. Associations between Gender, Microalbuminuria and microRNA Fold Changes
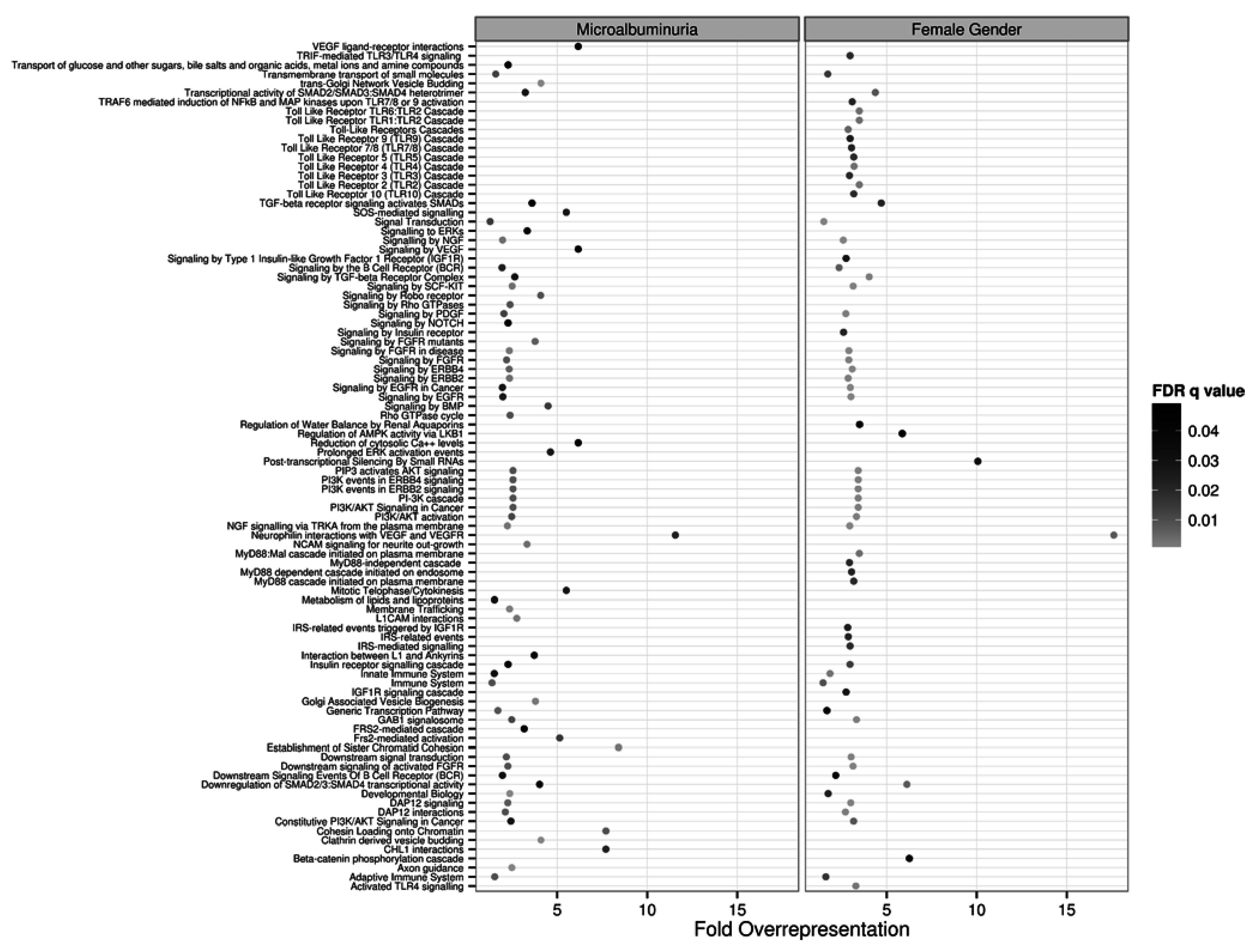
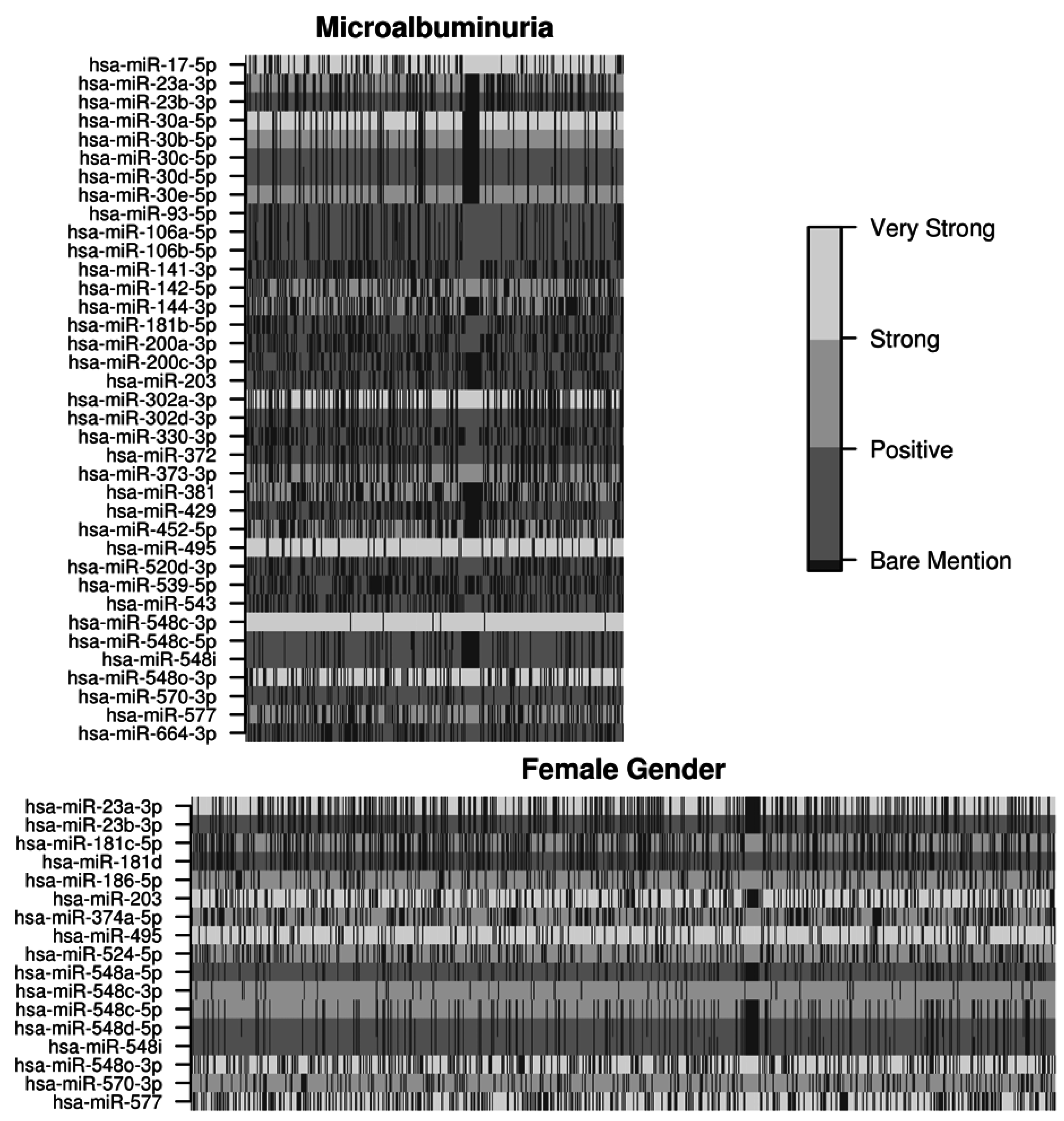
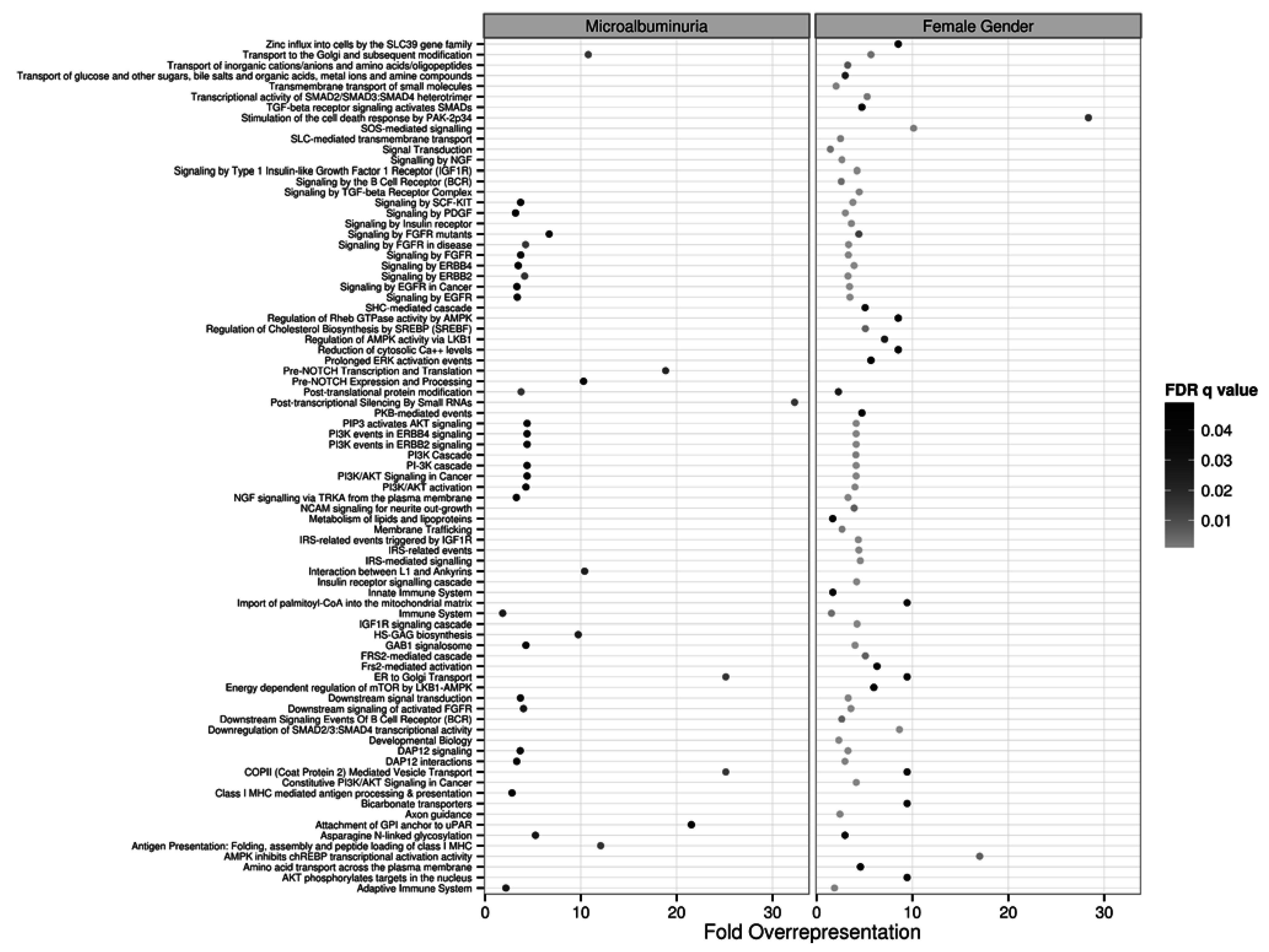
2.3. Target and Pathway Analyses
2.4. Construction of a microRNA Prognostic Index for Microalbuminuria
2.5. Validation of miRNA Features in Type 1 Diabetes
3. Results
| ID | Group | Sex | Age | HbA1c | Duration | Cycle | CAD | Stroke | PVD | Neuro | Retino | HTN |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | IMA | F | 27.4 | 10.4 | 19.2 | 3 | - | - | - | - | - | - |
| 2 | PMA | F | 22.7 | 11.8 | 20.25 | 3 | - | - | - | - | - | - |
| 3 | IMA | F | 29.9 | 11.4 | 18.6 | 5 | - | - | - | - | - | - |
| 4 | PMA | F | 26.3 | 13.1 | 18 | 5 | - | - | - | - | + | - |
| 5 | IMA | F | 24 | 10 | 21.2 | 2 | - | - | - | - | - | - |
| 6 | IMA | F | 24.3 | 14.3 | 19.7 | 2 | - | - | - | - | - | - |
| 7 | IMA | F | 26.9 | 10.4 | 18.5 | 2 | - | - | - | - | - | - |
| 8 | PMA | F | 25.2 | 8.2 | 12.9 | 2 | - | - | - | - | - | - |
| 9 | IMA | M | 30.66 | 11 | 19.97 | 3 | - | - | - | - | + | - |
| 10 | PMA | M | 23.16 | 11.5 | 22.05 | 3 | - | - | + | - | - | - |
| 11 | IMA | M | 41.7 | 6.6 | 30.54 | 6 | - | - | - | - | - | - |
| 12 | PMA | M | 38.97 | 5.2 | 31.54 | 2 | - | - | - | - | + | - |
| 13 | IMA | M | 39.08 | 12.4 | 24.52 | 6 | - | - | - | - | - | - |
| 14 | PMA | M | 28.35 | 11.6 | 27.01 | 4 | - | - | - | - | + | - |
| 15 | PMA | M | 27.16 | 13.9 | 24.3 | 2 | + | - | - | - | + | - |
| 16 | IMA | M | 23.13 | 12.1 | 9.77 | 2 | - | - | - | - | - | - |
| 17 | PMA | M | 22.8 | 13 | 12.9 | 3 | - | - | - | - | - | - |
| 18 | N | F | 40.32 | 7.1 | 29.63 | 10 | - | - | - | + | - | - |
| 19 | N | F | 48.93 | 8.3 | 36.73 | 10 | - | - | - | + | + | - |
| 20 | N | F | 51.16 | 8 | 46.96 | 10 | - | - | + | + | + | - |
| 21 | N | F | 39.45 | 7.9 | 29.77 | 10 | - | - | - | - | - | - |
| 22 | N | F | 41.19 | 9.8 | 38.13 | 10 | + | - | + | + | + | - |
| 23 | N | M | 48.72 | 6.6 | 33.76 | 10 | - | - | - | - | - | - |
| 24 | N | M | 42.46 | 9.8 | 33.4 | 10 | + | - | - | - | - | + |
| 25 | N | M | 42.5 | 8.2 | 36 | 10 | - | - | - | - | - | - |
| 26 | N | M | 35.35 | 9.1 | 28.53 | 10 | + | - | - | + | + | - |
| 27 | N | M | 38.54 | 7.7 | 27.81 | 10 | - | - | - | - | - | - |
| Feature | Log-Odds ǂ | |
|---|---|---|
| Concentration—Only Model | Concentration—Binding Model | |
| Intercept | 2.725 | 3.313 |
| hsa-miR-105-3p | −0.125 | −0.196 |
| hsa-miR-122-3p | 0.022 | |
| hsa-miR-124-3p | 0.003 | |
| hsa-miR-126-3p | 0.045 | |
| hsa-miR-1972 | −0.003 | −0.054 |
| hsa-miR-28-5p | −0.316 | −0.682 |
| hsa-miR-30b-5p | −0.008 | |
| hsa-miR-363-3p | −0.141 | −0.009 |
| hsa-miR-424-5p | −0.069 | |
| hsa-miR-486-5p | 0.083 | 0.212 |
| hsa-miR-495 | −0.045 | −0.028 |
| hsa-miR-548o-3p | −0.055 | |
| hsa-miR-122-5p X Women | 0.007 | |
| hsa-miR-192-5p X Women | 0.033 | 0.03 |
| hsa-miR-200c-3p X Women | 0.07 | |
| hsa-miR-548o-3p X Women | −0.296 | −0.498 |
| hsa-miR-720 X Women | 0.059 | 0.018 |
4. Discussion
5. Conclusions
Supplementary Files
Supplementary File 1Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
References
- Gross, J.L.; de Azevedo, M.J.; Silveiro, S.P.; Canani, L.H.; Caramori, M.L.; Zelmanovitz, T. Diabetic Nephropathy: Diagnosis, Prevention, and Treatment. Diabetes Care 2005, 28, 164–176. [Google Scholar] [CrossRef] [PubMed]
- Sacks, D.B.; Arnold, M.; Bakris, G.L.; Bruns, D.E.; Horvath, A.R.; Kirkman, M.S.; Lernmark, A.; Metzger, B.E.; Nathan, D.M. Guidelines and Recommendations for Laboratory Analysis in the Diagnosis and Management of Diabetes Mellitus. Clin. Chem. 2011, 57, e1–e47. [Google Scholar] [CrossRef] [PubMed]
- Fioretto, P.; Mauer, M. Histopathology of diabetic nephropathy. Semin. Nephrol. 2007, 27, 195–207. [Google Scholar] [CrossRef] [PubMed]
- Bader, R.; Bader, H.; Grund, K.E.; Mackensen-Haen, S.; Christ, H.; Bohle, A. Structure and function of the kidney in diabetic glomerulosclerosis. Correlations between morphological and functional parameters. Pathol. Res. Pract. 1980, 167, 204–216. [Google Scholar] [CrossRef]
- Caramori, M.L.; Kim, Y.; Huang, C.; Fish, A.J.; Rich, S.S.; Miller, M.E.; Russell, G.; Mauer, M. Cellular Basis of Diabetic Nephropathy 1. Study Design and Renal Structural-Functional Relationships in Patients with Long-Standing Type 1. Diabetes 2002, 51, 506–513. [Google Scholar] [CrossRef] [PubMed]
- Najafian, B.; Kim, Y.; Crosson, J.T.; Mauer, M. Atubular Glomeruli and Glomerulotubular Junction Abnormalities in Diabetic Nephropathy. J. Am. Soc. Nephrol. 2003, 14, 908–917. [Google Scholar] [CrossRef] [PubMed]
- Najafian, B.; Crosson, J.T.; Kim, Y.; Mauer, M. Glomerulotubular Junction Abnormalities are Associated with Proteinuria in Type 1 Diabetes. J. Am. Soc. Nephrol. 2006, 17, S53–S60. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez Suarez, M.L.; Thomas, D.B.; Barisoni, L.; Fornoni, A. Diabetic nephropathy: Is it time yet for routine kidney biopsy? World J. Diabetes 2013, 4, 245–255. [Google Scholar] [PubMed]
- Alter, M.L.; Kretschmer, A.; von Websky, K.; Tsuprykov, O.; Reichetzeder, C.; Simon, A.; Stasch, J.-P.; Hocher, B. Early urinary and plasma biomarkers for experimental diabetic nephropathy. Clin. Lab. 2012, 58, 659–671. [Google Scholar] [PubMed]
- Astor, B.C.; Matsushita, K.; Gansevoort, R.T.; van der Velde, M.; Woodward, M.; Levey, A.S.; de Jong, P.E.; Coresh, J.; el-Nahas, M.; Eckardt, K.-U.; et al. Lower estimated glomerular filtration rate and higher albuminuria are associated with mortality and end-stage renal disease. A collaborative meta-analysis of kidney disease population cohorts. Kidney Int. 2011, 79, 1331–1340. [Google Scholar] [CrossRef] [PubMed]
- Gansevoort, R.T.; Matsushita, K.; van der Velde, M.; Astor, B.C.; Woodward, M.; Levey, A.S.; de Jong, P.E.; Coresh, J.; el-Nahas, M.; Eckardt, K.-U.; et al. Lower estimated GFR and higher albuminuria are associated with adverse kidney outcomes. A collaborative meta-analysis of general and high-risk population cohorts. Kidney Int. 2011, 80, 93–104. [Google Scholar] [CrossRef] [PubMed]
- Matsushita, K.; van der Velde, M.; Astor, B.C.; Woodward, M.; Levey, A.S.; de Jong, P.E.; Coresh, J.; Gansevoort, R.T. Association of estimated glomerular filtration rate and albuminuria with all-cause and cardiovascular mortality: A collaborative meta-analysis of general population cohorts. Lancet 2010, 375, 2073–2081. [Google Scholar] [PubMed]
- Van Roosbroeck, K.; Pollet, J.; Calin, G.A. miRNAs and long noncoding RNAs as biomarkers in human diseases. Expert Rev. Mol. Diagn. 2013, 13, 183–204. [Google Scholar] [CrossRef] [PubMed]
- Mraz, M.; Malinova, K.; Mayer, J.; Pospisilova, S. MicroRNA isolation and stability in stored RNA samples. Biochem. Biophys. Res. Commun. 2009, 390, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Khella, H.W.Z.; Bakhet, M.; Lichner, Z.; Romaschin, A.D.; Jewett, M.A.S.; Yousef, G.M. MicroRNAs in kidney disease: An emerging understanding. Am. J. Kidney Dis. 2013, 61, 798–808. [Google Scholar] [CrossRef] [PubMed]
- Schena, F.P.; Serino, G.; Sallustio, F. MicroRNAs in kidney diseases: New promising biomarkers for diagnosis and monitoring. Nephrol. Dial. Transplant. 2014, 29, 755–763. [Google Scholar] [CrossRef] [PubMed]
- Chandrasekaran, K.; Karolina, D.S.; Sepramaniam, S.; Armugam, A.; Wintour, E.M.; Bertram, J.F.; Jeyaseelan, K. Role of microRNAs in kidney homeostasis and disease. Kidney Int. 2012, 81, 617–627. [Google Scholar] [CrossRef] [PubMed]
- Alvarez, M.L.; Distefano, J.K. The role of non-coding RNAs in diabetic nephropathy: Potential applications as biomarkers for disease development and progression. Diabetes Res. Clin. Pract. 2013, 99, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Xiao, L.; Li, J.; Kanwar, Y.S.; Liu, F.; Sun, L. Urine miRNAs: Potential biomarkers for monitoring progression of early stages of diabetic nephropathy. Med. Hypotheses 2013, 81, 274–278. [Google Scholar] [CrossRef] [PubMed]
- Barutta, F.; Tricarico, M.; Corbelli, A.; Annaratone, L.; Pinach, S.; Grimaldi, S.; Bruno, G.; Cimino, D.; Taverna, D.; Deregibus, M.C.; et al. Urinary Exosomal MicroRNAs in Incipient Diabetic Nephropathy. PLoS ONE 2013, 8, e73798. [Google Scholar] [CrossRef] [PubMed]
- Osipova, J.; Fischer, D.-C.; Dangwal, S.; Volkmann, I.; Widera, C.; Schwarz, K.; Lorenzen, J.M.; Schreiver, C.; Jacoby, U.; Heimhalt, M.; et al. Diabetes-associated microRNAs in paediatric patients with Type 1 diabetes mellitus: A cross-sectional cohort study. J. Clin. Endocrinol. Metab. 2014, 99. [Google Scholar] [CrossRef]
- Argyropoulos, C.; Wang, K.; McClarty, S.; Huang, D.; Bernardo, J.; Ellis, D.; Orchard, T.; Galas, D.; Johnson, J. Urinary microRNA profiling in the nephropathy of Type 1 diabetes. PLoS ONE 2013, 8, e54662. [Google Scholar] [CrossRef] [PubMed]
- Marques, F.Z.; Campain, A.E.; Tomaszewski, M.; Zukowska-Szczechowska, E.; Yang, Y.H.J.; Charchar, F.J.; Morris, B.J. Gene Expression Profiling Reveals Renin mRNA Overexpression in Human Hypertensive Kidneys and a Role for MicroRNAs. Hypertension 2011, 58, 1093–1098. [Google Scholar] [CrossRef] [PubMed]
- Miller, R.G.; Secrest, A.M.; Ellis, D.; Becker, D.J.; Orchard, T.J. Changing Impact of Modifiable Risk Factors on the Incidence of Major Outcomes of Type 1 Diabetes: The Pittsburgh Epidemiology of Diabetes Complications Study. Diabetes Care 2013, 36, 3999–4006. [Google Scholar] [CrossRef] [PubMed]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 2001, 29. [Google Scholar] [CrossRef]
- Argyropoulos, C.; Nikiforidis, G.C.; Theodoropoulou, M.; Adamopoulos, P.; Boubali, S.; Georgakopoulos, T.N.; Paliogianni, F.; Papavassiliou, A.G.; Mouzaki, A. Mining microarray data to identify transcription factors expressed in naïve resting but not activated T lymphocytes. Genes Immun. 2004, 5, 16–25. [Google Scholar] [CrossRef] [PubMed]
- Kass, R.E.; Raftery, A.E. Bayes Factors. J. Am. Stat. Assoc. 1995, 90, 773–795. [Google Scholar] [CrossRef]
- Fiedler, J.; Gupta, S.K.; Thum, T. Identification of cardiovascular microRNA targetomes. J. Mol. Cell. Cardiol. 2011, 51, 674–681. [Google Scholar] [CrossRef] [PubMed]
- Turner, H.; Bailey, T.; Krzanowski, W. Improved biclustering of microarray data demonstrated through systematic performance tests. Comput. Stat. Data Anal. 2005, 48, 235–254. [Google Scholar] [CrossRef]
- Matthews, L.; Gopinath, G.; Gillespie, M.; Caudy, M.; Croft, D.; de Bono, B.; Garapati, P.; Hemish, J.; Hermjakob, H.; Jassal, B.; et al. REACTOME knowledgebase of human biological pathways and processes. Nucleic Acids Res. 2009, 37, D619–D622. [Google Scholar] [CrossRef] [PubMed]
- Zou, H.; Hastie, T. Regularization and variable selection via the elastic net. J. R. Stat. Soc. Ser. B Stat. Methodol. 2005, 67, 301–320. [Google Scholar] [CrossRef]
- Alam-Faruque, Y.; Dimmer, E.C.; Huntley, R.P.; O’Donovan, C.; Scambler, P.; Apweiler, R. The renal gene ontology annotation initiative. Organogenesis 2010, 6, 71–75. [Google Scholar] [CrossRef] [PubMed]
- Alam-Faruque, Y.; Hill, D.P.; Dimmer, E.C.; Harris, M.A.; Foulger, R.E.; Tweedie, S.; Attrill, H.; Howe, D.G.; Thomas, S.R.; Davidson, D.; et al. Representing Kidney Development Using the Gene Ontology. PLoS ONE 2014, 9, e99864. [Google Scholar] [CrossRef] [PubMed]
- Baelde, H.J.; Eikmans, M.; Doran, P.P.; Lappin, D.W.P.; de Heer, E.; Bruijn, J.A. Gene expression profiling in glomeruli from human kidneys with diabetic nephropathy. Am. J. Kidney Dis. 2004, 43, 636–650. [Google Scholar] [CrossRef] [PubMed]
- Barrell, D.; Dimmer, E.; Huntley, R.P.; Binns, D.; O’Donovan, C.; Apweiler, R. The GOA database in 2009—An integrated Gene Ontology Annotation resource. Nucleic Acids Res. 2009, 37, D396–D403. [Google Scholar] [CrossRef] [PubMed]
- Wada, J.; Makino, H. Inflammation and the pathogenesis of diabetic nephropathy. Clin. Sci. 2013, 124, 139–152. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.-T.; Tsai, P.-C.; Liao, Y.-C.; Hsu, C.-Y.; Juo, S.-H.H. Circulating microRNAs have a sex-specific association with metabolic syndrome. J. Biomed. Sci. 2013, 20. [Google Scholar] [CrossRef] [PubMed]
- Klinge, C.M. Estrogen Regulation of MicroRNA Expression. Curr. Genomics 2009, 10, 169–183. [Google Scholar] [CrossRef] [PubMed]
- Ganesan, J.; Ramanujam, D.; Sassi, Y.; Ahles, A.; Jentzsch, C.; Werfel, S.; Leierseder, S.; Loyer, X.; Giacca, M.; Zentilin, L.; et al. MiR-378 controls cardiac hypertrophy by combined repression of mitogen-activated protein kinase pathway factors. Circulation 2013, 127, 2097–2106. [Google Scholar] [CrossRef] [PubMed]
- Costacou, T.; Fried, L.; Ellis, D.; Orchard, T.J. Sex Differences in the Development of Kidney Disease in Individuals with Type 1 Diabetes Mellitus: A Contemporary Analysis. Am. J. Kidney Dis. 2011, 58, 565–573. [Google Scholar] [CrossRef] [PubMed]
- Neugarten, J.; Acharya, A.; Silbiger, S.R. Effect of gender on the progression of nondiabetic renal disease: A meta-analysis. J. Am. Soc. Nephrol. 2000, 11, 319–329. [Google Scholar] [PubMed]
- Neugarten, J.; Golestaneh, L. Gender and the prevalence and progression of renal disease. Adv. Chronic Kidney Dis. 2013, 20, 390–395. [Google Scholar] [CrossRef] [PubMed]
- Cattran, D.C.; Reich, H.N.; Beanlands, H.J.; Miller, J.A.; Scholey, J.W.; Troyanov, S.; Genes, Gender and Glomerulonephritis Group. The impact of sex in primary glomerulonephritis. Nephrol. Dial. Transplant. 2008, 23, 2247–2253. [Google Scholar] [CrossRef] [PubMed]
- Hirst, J.A.; Taylor, K.S.; Stevens, R.J.; Blacklock, C.L.; Roberts, N.W.; Pugh, C.W.; Farmer, A.J. The impact of renin-angiotensin-aldosterone system inhibitors on Type 1 and Type 2 diabetic patients with and without early diabetic nephropathy. Kidney Int. 2012, 81, 674–683. [Google Scholar] [CrossRef] [PubMed]
- Hooten, N.N.; Fitzpatrick, M.; Wood, W.H.; De, S.; Ejiogu, N.; Zhang, Y.; Mattison, J.A.; Becker, K.G.; Zonderman, A.B.; Evans, M.K.; et al. Age-related changes in microRNA levels in serum. Aging 2013, 5, 725–740. [Google Scholar]
- Verzola, D.; Gandolfo, M.T.; Gaetani, G.; Ferraris, A.; Mangerini, R.; Ferrario, F.; Villaggio, B.; Gianiorio, F.; Tosetti, F.; Weiss, U.; et al. Accelerated senescence in the kidneys of patients with type 2 diabetic nephropathy. Am. J. Physiol. Ren. Physiol. 2008, 295, F1563–F1573. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Zhang, R.; Torreggiani, M.; Ting, A.; Xiong, H.; Striker, G.E.; Vlassara, H.; Zheng, F. Induction of diabetes in aged C57B6 mice results in severe nephropathy: An association with oxidative stress, endoplasmic reticulum stress, and inflammation. Am. J. Pathol. 2010, 176, 2163–2176. [Google Scholar] [CrossRef] [PubMed]
- Roscioni, S.S.; Lambers Heerspink, H.J.; de Zeeuw, D. Microalbuminuria: Target for renoprotective therapy PRO. Kidney Int. 2014, 86, 40–49. [Google Scholar] [CrossRef] [PubMed]
- MacIsaac, R.J.; Ekinci, E.I.; Jerums, G. Progressive diabetic nephropathy. How useful is microalbuminuria?: Contra′. Kidney Int. 2014, 86, 50–57. [Google Scholar] [CrossRef] [PubMed]
- Glassock, R.J. Is the presence of microalbuminuria a relevant marker of kidney disease? Curr. Hypertens. Rep. 2010, 12, 364–368. [Google Scholar] [CrossRef] [PubMed]
- Galler, A.; Haberland, H.; Näke, A.; Hofer, S.; Holder, M.; Raile, K.; Holl, R.W. Natural course of untreated microalbuminuria in children and adolescents with Type 1 diabetes and the importance of diabetes duration and immigrant status: Longitudinal analysis from the prospective nationwide German and Austrian diabetes survey DPV. Eur. J. Endocrinol. 2012, 166, 493–501. [Google Scholar] [CrossRef] [PubMed]
- Amin, R.; Widmer, B.; Prevost, A.T.; Schwarze, P.; Cooper, J.; Edge, J.; Marcovecchio, L.; Neil, A.; Dalton, R.N.; Dunger, D.B.; et al. Risk of microalbuminuria and progression to macroalbuminuria in a cohort with childhood onset Type 1 diabetes: Prospective observational study. BMJ 2008, 336, 697–701. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Argyropoulos, C.; Wang, K.; Bernardo, J.; Ellis, D.; Orchard, T.; Galas, D.; Johnson, J.P. Urinary MicroRNA Profiling Predicts the Development of Microalbuminuria in Patients with Type 1 Diabetes. J. Clin. Med. 2015, 4, 1498-1517. https://doi.org/10.3390/jcm4071498
Argyropoulos C, Wang K, Bernardo J, Ellis D, Orchard T, Galas D, Johnson JP. Urinary MicroRNA Profiling Predicts the Development of Microalbuminuria in Patients with Type 1 Diabetes. Journal of Clinical Medicine. 2015; 4(7):1498-1517. https://doi.org/10.3390/jcm4071498
Chicago/Turabian StyleArgyropoulos, Christos, Kai Wang, Jose Bernardo, Demetrius Ellis, Trevor Orchard, David Galas, and John P. Johnson. 2015. "Urinary MicroRNA Profiling Predicts the Development of Microalbuminuria in Patients with Type 1 Diabetes" Journal of Clinical Medicine 4, no. 7: 1498-1517. https://doi.org/10.3390/jcm4071498
APA StyleArgyropoulos, C., Wang, K., Bernardo, J., Ellis, D., Orchard, T., Galas, D., & Johnson, J. P. (2015). Urinary MicroRNA Profiling Predicts the Development of Microalbuminuria in Patients with Type 1 Diabetes. Journal of Clinical Medicine, 4(7), 1498-1517. https://doi.org/10.3390/jcm4071498





