1. Introduction
Glioma is the most common primary brain tumor, accounting for 70% of malignant primary brain tumors in adults. The World Health Organization (WHO) classifies brain tumors into grades I–IV [
1], with different survival times for different grades; the higher the grade of a brain tumor, the more aggressive it is, and the shorter the survival time of patients [
2], so the early diagnosis and treatment of brain tumors are crucial. Physicians usually use magnetic resonance imaging (MRI) [
3] as the main basis for the clinical diagnosis of brain tumors. The four common MRI modalities of the brain include: T1-weighted (T1), contrast-enhanced T1-weighted (T1c), T2-weighted (T2), and fluid-attenuated inversion recovery (FLAIR) [
4]. However, brain tumors are characterized by different morphologies and variable locations [
5], and manual segmentation requires doctors to rely on their professional knowledge and work experience to analyze the condition, so human factors such as fatigue, memory, and lack of work experience may make the diagnostic results wrong during the segmentation process. Medical imaging and computer-aided diagnosis (MICAD) technology has been made possible by the growing connection between computer science and medicine. Combining MICAD technology with computer vision to allow doctors to see the results of MRI image segmentation and use them as a reference can speed up diagnostics and make them much more accurate.
Traditional brain tumor segmentation algorithms include threshold-based, clustering-based, atlas-based, and supervision-based algorithms [
6,
7,
8,
9,
10]. Although the computational complexity of brain tumor image segmentation algorithms based on traditional machine learning is low, manually designed features cannot effectively utilize the rich information of MRI brain images. Moreover, such algorithms usually need to compute many features to ensure accuracy, which slows down the computation speed and increases the memory overhead.
In recent years, convolutional neural networks (CNNs) have been successfully applied in the field of medical images [
11]. Unlike traditional machine learning methods, CNNs use massive amounts of data to learn representative features automatically. This way, they do not have to go through the complex process of extracting features. CNNs have been applied successfully in medical image classification [
12], detection [
13], segmentation [
14,
15], and other image processing tasks [
16]. Many studies on brain tumor segmentation have used CNN-based methods, which have greatly improved the speed and accuracy of diagnosing brain tumors. Havaei et al. [
17] designed a cascaded dual-path CNN structure. Convolution kernels of different sizes were used to learn local and global features of images simultaneously; then local features and global features were fused to obtain more context information. Salehi et al. [
18] proposed an auto-context-based CNN algorithm to obtain the local features and global features using an approach based on three parallel two-dimensional convolutional paths in the axial, coronal, and sagittal directions. However, these algorithms are subject to necessary post-processing as they are executed based on image blocks. Ronneberger et al. [
19] proposed the encoding-decoding network U-net, which increases the number of upsampling layers compared with the traditional FCN [
20] and gradually restores the image details and image resolution, and the feature map concatenation of the encoding path and the decoding path can maintain complete contextual features. U-net has become the most popular basic network in medical image processing tasks. Researchers have improved U-net and produced variants of higher accuracy [
21,
22]. Marcinkiewicz et al. [
23] proposed an improved structure of 2D U-Net, where MR images of three modalities are input into three independent encoders. Then, the feature channels are fused and fed into the encoders. Although 2D CNN-based brain tumor segmentation methods are efficient, 3D MRI image continuity and complete 3D spatial contextual information are difficult to capture. The 3D U-net [
24] network extended the U-net architecture by replacing all 2D operations with their 3D operations. Mehta et al. [
25] proposed a 3D U-net brain tumor segmentation network using 3D convolution. Myronenko et al. [
26,
27] reconstructed the input image by adding variational auto-coding branches to regularize the shared encoder and achieved first place in the BraTS 2018 and BraTS 2019 competitions, respectively. Attention mechanisms are often introduced in brain tumor segmentation networks to make the model more focused on tumor-related regions and improve segmentation performance [
28,
29,
30,
31]. Cao et al. [
15] used a novel multi-branch 3D shuffle attention module as the attention layer in the encoder, which grouped along the channel dimension and divided the feature maps into small feature maps.
Although the segmentation networks based on encoding and decoding structures can achieve satisfactory performance, continuous upsampling does not bring back image detail information from multiple downsampling, cannot bring back spatial information well, and does not pay much attention to edges. The extracted feature approach inevitably produces a fuzzy feature mapping after multiple convolutions and loses some important details, such as the boundaries of the brain tumor. Sun et al. [
32] proposed a high-resolution network (HRNet). HRNet has been successfully applied in many tasks, such as human pose estimation [
33] and semantic segmentation [
34]. Compared with U-Net, it can maintain accurate spatial feature information due to the high-resolution branch. However, a large number of repetitive inter-fusion operations between multiple stages increase computational complexity, and intensive feature fusion computes a large amount of redundant feature information.
Although 3D convolution has good segmentation performance, it significantly increases the number of model training parameters, which results in a larger computational overhead. Nowadays, in edge computing systems, model efficiency is also an important aspect to be considered. With the rapid development of CNN technology, many lightweight networks [
35,
36,
37] have been proposed to further reduce the model complexity and the memory occupied. Ma et al. [
38] utilized three operations, channel split, depthwise separable convolutions, and channel shuffle, to enable the model to greatly reduce the computational effort of model while maintaining accuracy. Chen et al. [
39] exploited the advantage of separable 3D convolutions to reduce the network complexity but with much lower segmentation accuracy. To balance network efficiency and accuracy in 3D MRI brain tumor segmentation, Chen et al. [
40] proposed a new 3D expanded multi-fiber network (DMFNet) that uses efficient group convolution on top of multi-fiber units and obtains a multi-scale image representation for segmentation by introducing weighted 3D expanded convolution operations, which achieved precise segmentation while reducing the number of network parameters. However, the problem of the lack of exchange of channel information was not solved. Zhou et al. [
6] proposed an efficient 3D residual neural network (ERV-Net) by first utilizing the computationally efficient network 3D ShuffleNetV2 as an encoder to reduce GPU memory and improve the segmentation efficiency of the network, and then introducing a decoder with residual blocks to avoid degradation.
The challenge of this study is to design a 3D brain tumor segmentation network with high accuracy and efficiency. The networks based on the U-net network ignore the gap problem between low-level visual features and high-level semantic features, have limited ability to reconstruct spatial information of brain tumors, and pay little attention to boundaries. The purpose of this paper is to achieve fast and accurate segmentation of brain tumors using a high-resolution-based network.
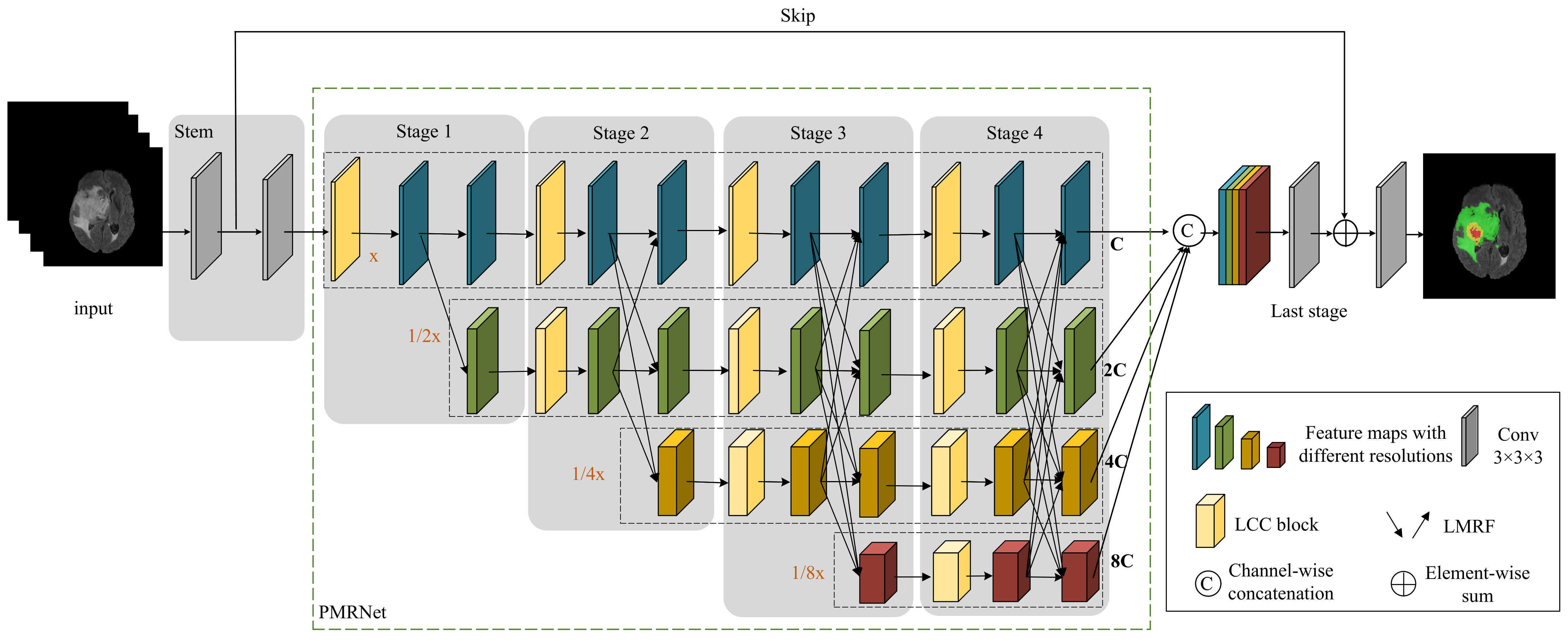
In this paper, we propose a hierarchical multi-scale brain tumor segmentation network (HMNet) to segment brain tumors with high efficiency and accuracy. Firstly, the network contains a high-resolution branch to maintain accurate spatial feature representation and multi-resolution branches to acquire multi-scale receptive fields and multi-scale features, which helps to segment brain tumors of different sizes and shapes. Secondly, to reduce the network parameters and computation to improve the efficiency of brain tumor segmentation, we exploit a 3D shuffle block [
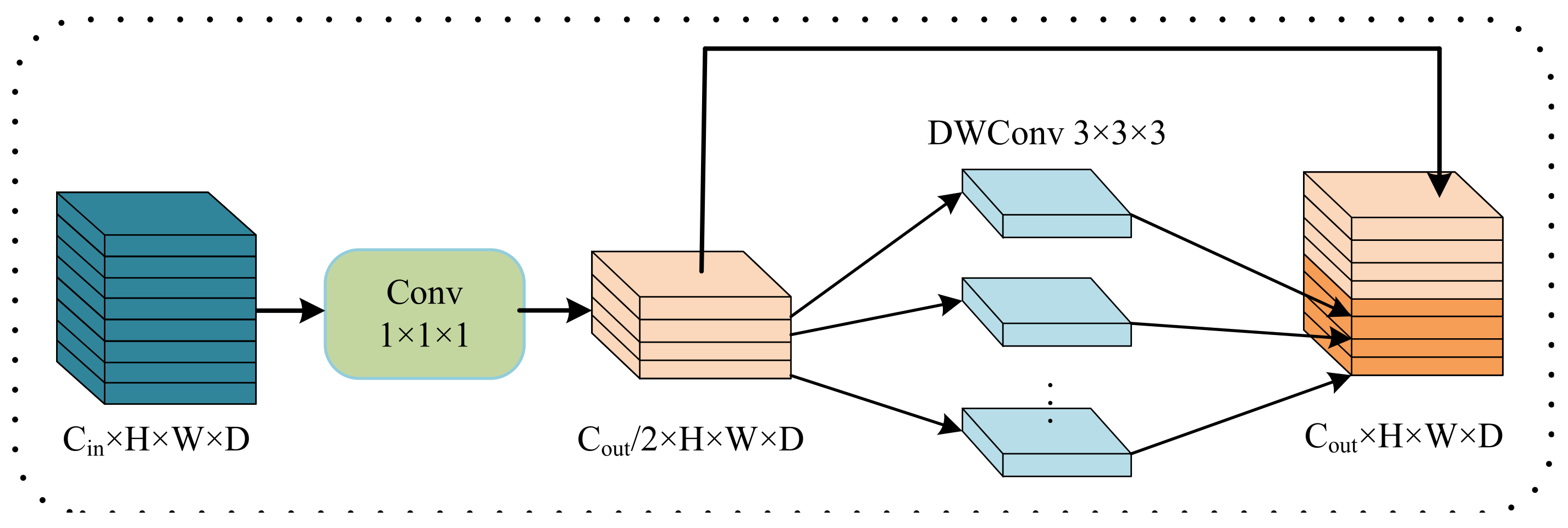
38] for feature extraction, then propose a lightweight conditional channel weighting (LCC) block to improve it. Thirdly, we propose a lightweight multi-resolution feature fusion (LMRF) module to further reduce model complexity and reduce feature redundancy. Finally, we assess our experimental results using the BraTS 2018, BraTS 2019 and BraTS 2020 datasets.
The contributions of this paper can be summarized as follows.
We propose a hierarchical multi-scale brain tumor segmentation network (HMNet). HMNet uses a parallel multi-resolution feature extraction network with a high-resolution branch and parallel multi-resolution branches to figure out what brain tumors look like.
We propose a lightweight conditional channel weighting (LCC) block based on a 3D shuffle block [
38] to overcome the computational problem caused by traditional 3D convolution and enhance useful features.
We propose a lightweight multi-resolution fusion (LMRF) module to solve the problem that the traditional method of downsampling and upsampling has a lot of computational complexity.
The remaining sections of this article are structured as follows.
Section 2 describes our methodology in depth.
Section 3 contains the appropriate experimental data and analyses.
Section 4 discusses the effectiveness of our proposed network.
Section 5 summarizes the work of this paper.
4. Discussion
In this paper, we proposed a lightweight brain tumor segmentation network HMNet. Specifically, the shape, size and location of brain tumors vary greatly among patients, and noise is introduced during MRI scanning, resulting in blurred boundaries between tumor regions, making it difficult to segment brain tumors accurately. In addition, the networks based on 3D convolutions are computationally intensive, so these networks are not efficient enough for clinical practice. To balance segmentation accuracy and network complexity, we designed a hierarchical multi-scale brain tumor segmentation network.
We ran tests on the BraTS 2020 dataset to verify the effectiveness of the HMNet. The dice similarity coefficients of HMNet for ET, WT, and TC are 0.781, 0.901, and 0.823, respectively. Extensive experiments on the BraTS 2018, BraTS 2019, and BraTS 2020 datasets show that the proposed network has achieved satisfactory performance compared with the SOTA approaches.
Compared with the baseline network in
Table 8, the parameters and flops of HMNet are significantly reduced while improving segmentation accuracy. The reason for this is that we introduce the depthwise separable convolution to replace the traditional 3 × 3 × 3 convolutions in the LCC block and LMRF module and replace the costly 1 × 1 × 1 convolution in the 3D shuffle block with CWU and SWU to overcome the problem of the heavily-used 1 × 1 × 1 convolution. HMNet has higher segmentation accuracy than the baseline network because the CWU and SWU make the network more focused on the areas associated with the tumor. In addition, the LMRF module and LCC block adopt the idea of feature reuse, where shallow feature information is directly used by deeper layers to prevent network degradation or overfitting. This can reduce the computational effort while reducing feature redundancy and improving the segmentation accuracy of the network.
Compared with other non-lightweight brain tumor segmentation networks, our proposed HMNet has better segmentation performance and lower model complexity. This is mainly because the PMRNet maintains the high-resolution features during feature propagation and ensures that the features contain accurate detailed information. However, the networks based on U-net lose some image detail information from multiple downsampling, especially at the edges of brain tumors. Moreover, our proposed HMNet is competitive with other lightweight brain tumor segmentation networks. Some lightweight networks such as DMF-Net and HDC-Net are based on U-net, so they may lose some important information. Furthermore, HDC-Net is too light to segment brain tumor boundaries accurately. Overall, our network achieves a better balance of accuracy and complexity.