Salivary MicroRNA Signature for Diagnosis of Endometriosis
Abstract
:1. Introduction
2. Material and Methods
2.1. Ethics Statement
2.2. Study Population
2.3. Saliva Sample Collection
2.4. RNA Sample Extraction, Preparation and Quality Control
3. Bioinformatics
3.1. Raw Data Preprocessing (Raw, Filtered, Aligned Reads) and Quality
3.2. Differential Expression Analysis of miRNAs
4. Statistical Analysis
4.1. Development and Validation of the Diagnostic Model
4.2. Validation of the Signature Accuracy
4.3. Other Statistical Analyses
5. Results
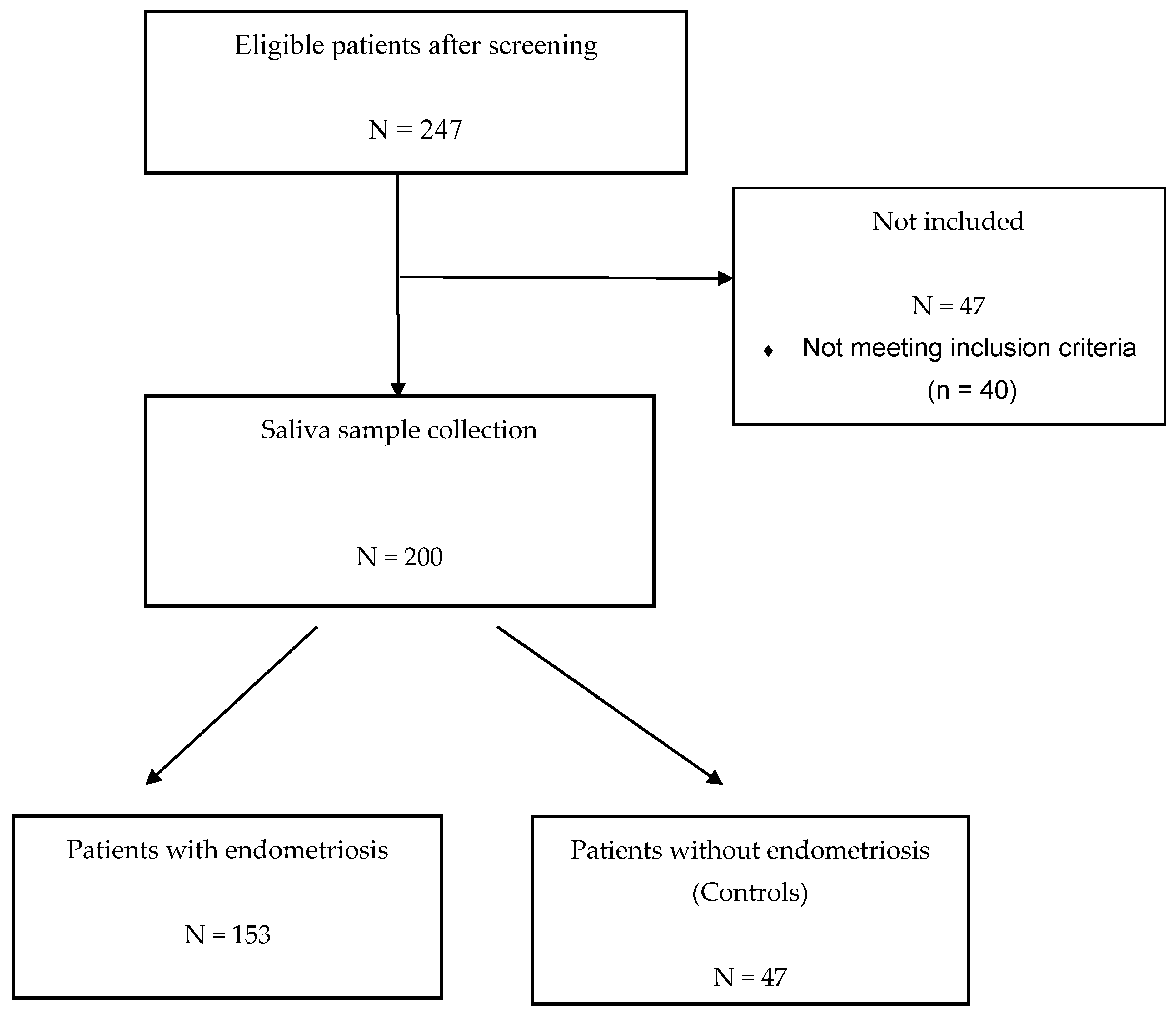
5.1. Description of the ENDO-miRNA Cohort
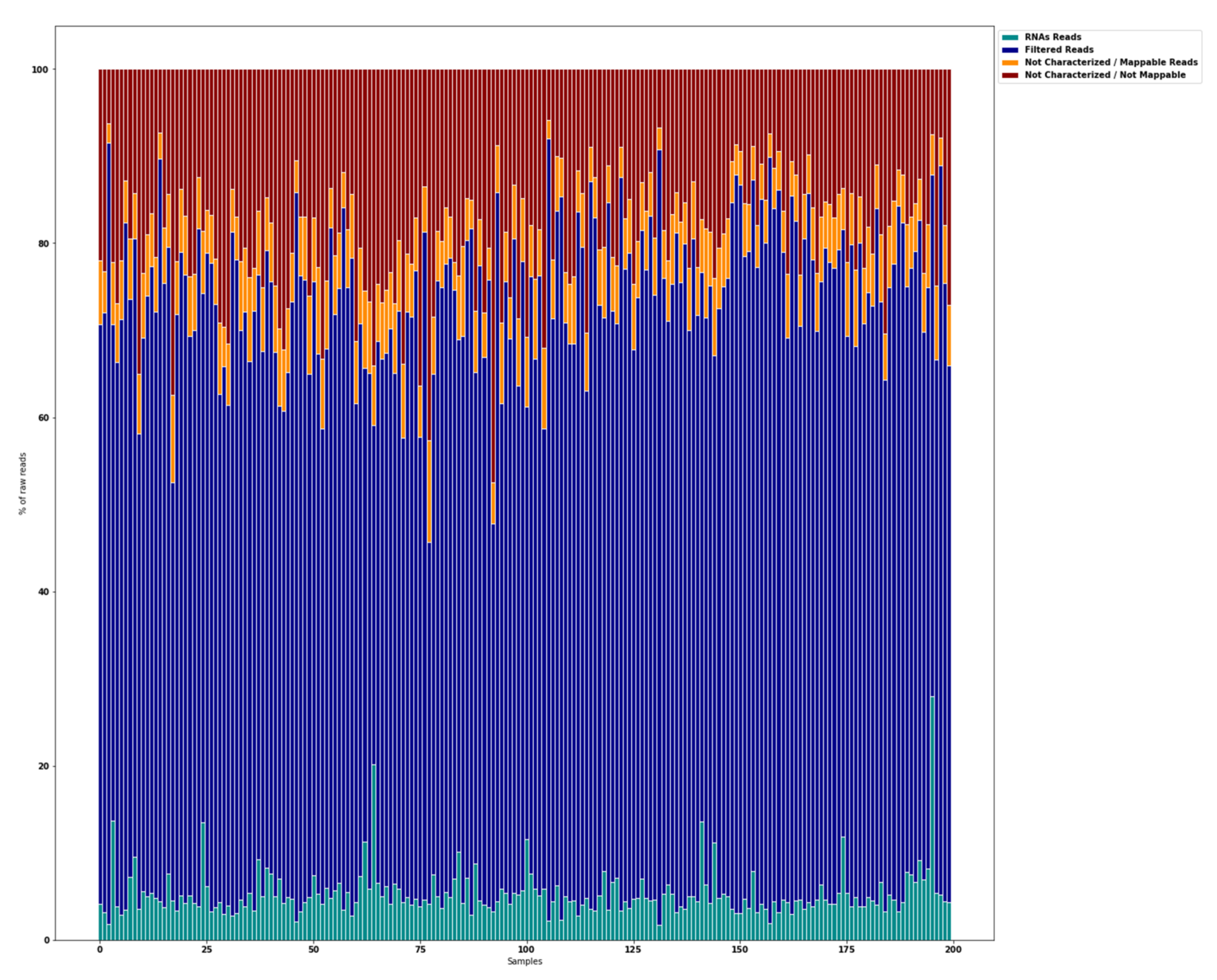
5.2. Global Overview of miRNA Transcriptome
5.3. Feature Selection of miRNAs Relevant for a Diagnosis of Endometriosis
5.4. Saliva-Based Diagnostic Signature for Endometriosis
5.5. Relation between Pathophysiology of Endometriosis and miRNA Expression
6. Discussion
7. Conclusions and Perspectives
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Giudice, L.C. Clinical Practice. Endometriosis. N. Engl. J. Med. 2010, 362, 2389–2398. [Google Scholar] [CrossRef] [PubMed]
- Zondervan, K.T.; Becker, C.M.; Missmer, S.A. Endometriosis. N. Engl. J. Med. 2020, 382, 1244–1256. [Google Scholar] [CrossRef] [PubMed]
- van der Zanden, M.; Teunissen, D.A.M.; van der Woord, I.W.; Braat, D.D.M.; Nelen, W.L.D.M.; Nap, A.W. Barriers and Facilitators to the Timely Diagnosis of Endometriosis in Primary Care in the Netherlands. Fam. Pract. 2020, 37, 131–136. [Google Scholar] [CrossRef] [PubMed]
- Duffy, J.M.N.; Adamson, G.D.; Benson, E.; Bhattacharya, S.; Bhattacharya, S.; Bofill, M.; Brian, K.; Collura, B.; Curtis, C.; Evers, J.L.H.; et al. Top 10 Priorities for Future Infertility Research: An International Consensus Development Study. Hum. Reprod. Oxf. Engl. 2020, 35, 2715–2724. [Google Scholar] [CrossRef]
- Nisenblat, V.; Bossuyt, P.M.; Farquhar, C.; Johnson, N.; Hull, M.L. Imaging Modalities for the Non-Invasive Diagnosis of Endometriosis. Cochrane Database Syst. Rev. 2016. [Google Scholar] [CrossRef]
- Bazot, M.; Lafont, C.; Rouzier, R.; Roseau, G.; Thomassin-Naggara, I.; Daraï, E. Diagnostic Accuracy of Physical Examination, Transvaginal Sonography, Rectal Endoscopic Sonography, and Magnetic Resonance Imaging to Diagnose Deep Infiltrating Endometriosis. Fertil. Steril. 2009, 92, 1825–1833. [Google Scholar] [CrossRef]
- Bazot, M.; Darai, E.; Hourani, R.; Thomassin, I.; Cortez, A.; Uzan, S.; Buy, J.-N. Deep Pelvic Endometriosis: MR Imaging for Diagnosis and Prediction of Extension of Disease. Radiology 2004, 232, 379–389. [Google Scholar] [CrossRef] [Green Version]
- Lorusso, F.; Scioscia, M.; Rubini, D.; Stabile Ianora, A.A.; Scardigno, D.; Leuci, C.; De Ceglie, M.; Sardaro, A.; Lucarelli, N.; Scardapane, A. Magnetic Resonance Imaging for Deep Infiltrating Endometriosis: Current Concepts, Imaging Technique and Key Findings. Insights Imaging 2021, 12, 105. [Google Scholar] [CrossRef]
- Soliman, A.M.; Rahal, Y.; Robert, C.; Defoy, I.; Nisbet, P.; Leyland, N.; Singh, S. Impact of Endometriosis on Fatigue and Productivity Impairment in a Cross-Sectional Survey of Canadian Women. JOGC 2021, 43, 10–18. [Google Scholar] [CrossRef]
- Surrey, E.; Soliman, A.M.; Trenz, H.; Blauer-Peterson, C.; Sluis, A. Impact of Endometriosis Diagnostic Delays on Healthcare Resource Utilization and Costs. Adv. Ther. 2020, 37, 1087–1099. [Google Scholar] [CrossRef] [Green Version]
- As-Sanie, S.; Soliman, A.M.; Evans, K.; Erpelding, N.; Lanier, R.; Katz, N.P. Healthcare Utilization and Cost Burden among Women with Endometriosis by Opioid Prescription Status in the First Year after Diagnosis: A Retrospective Claims Database Analysis. J. Med. Econ. 2020, 23, 371–377. [Google Scholar] [CrossRef] [PubMed]
- Dai, Y.; Shi, B.; Huang, X.; Duan, J.; Qiu, Y.; Ha, C.; Huang, R.; Xiao, D.; Liu, J.; Xuan, J. Cost-Effectiveness Analysis of Dienogest Compared With Gonadotropin-Releasing Hormone Agonist After Conservative Surgery for Endometriosis in China. Clin. Ther. 2021, 43, 1276–1284.e1. [Google Scholar] [CrossRef] [PubMed]
- Simoens, S.; Dunselman, G.; Dirksen, C.; Hummelshoj, L.; Bokor, A.; Brandes, I.; Brodszky, V.; Canis, M.; Colombo, G.L.; DeLeire, T.; et al. The Burden of Endometriosis: Costs and Quality of Life of Women with Endometriosis and Treated in Referral Centres. Hum. Reprod. Oxf. Engl. 2012, 27, 1292–1299. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nisenblat, V.; Prentice, L.; Bossuyt, P.M.; Farquhar, C.; Hull, M.L.; Johnson, N. Combination of the Non-Invasive Tests for the Diagnosis of Endometriosis. Cochrane Database Syst. Rev. 2016, 2016, CD012281. [Google Scholar] [CrossRef]
- Nisenblat, V.; Bossuyt, P.M.; Shaikh, R.; Farquhar, C.; Jordan, V.; Scheffers, C.S.; Mol, B.W.J.; Johnson, N.; Hull, M.L. Blood Biomarkers for the Non-Invasive Diagnosis of Endometriosis. Cochrane Database Syst. Rev. 2016, 2016, CD012179. [Google Scholar] [CrossRef] [Green Version]
- Surrey, E.; Carter, C.M.; Soliman, A.M.; Khan, S.; DiBenedetti, D.B.; Snabes, M.C. Patient-Completed or Symptom-Based Screening Tools for Endometriosis: A Scoping Review. Arch. Gynecol. Obstet. 2017, 296, 153–165. [Google Scholar] [CrossRef] [Green Version]
- Akter, S.; Xu, D.; Nagel, S.C.; Bromfield, J.J.; Pelch, K.E.; Wilshire, G.B.; Joshi, T. GenomeForest: An Ensemble Machine Learning Classifier for Endometriosis. AMIA Jt. Summits Transl. Sci. Proc. AMIA Jt. Summits Transl. Sci. 2020, 2020, 33–42. [Google Scholar]
- Kleczyk, E.J.; Peri, A.; Yadav, T.; Komera, R.; Peri, M.; Guduru, V.; Amirtharaj, S.; Huang, M. Predicting Endometriosis Onset Using Machine Learning Algorithms. Res. Sq. 2020. in review. [Google Scholar] [CrossRef]
- Vanhie, A.; O, D.; Peterse, D.; Beckers, A.; Cuéllar, A.; Fassbender, A.; Meuleman, C.; Mestdagh, P.; D’Hooghe, T. Plasma MiRNAs as Biomarkers for Endometriosis. Hum. Reprod. Oxf. Engl. 2019, 34, 1650–1660. [Google Scholar] [CrossRef]
- Agrawal, S.; Tapmeier, T.; Rahmioglu, N.; Kirtley, S.; Zondervan, K.; Becker, C. The MiRNA Mirage: How Close Are We to Finding a Non-Invasive Diagnostic Biomarker in Endometriosis? A Systematic Review. Int. J. Mol. Sci. 2018, 19, 599. [Google Scholar] [CrossRef] [Green Version]
- Santamaria-Martos, F.; Benítez, I.; Ortega, F.; Zapater, A.; Giron, C.; Pinilla, L.; Pascual, L.; Cortijo, A.; Dalmases, M.; Fernandez-Real, J.M.; et al. Circulating MicroRNA Profile as a Potential Biomarker for Obstructive Sleep Apnea Diagnosis. Sci. Rep. 2019, 9, 13456. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ghafouri-Fard, S.; Shoorei, H.; Taheri, M. Role of Non-Coding RNAs in the Pathogenesis of Endometriosis. Front. Oncol. 2020, 10, 1370. [Google Scholar] [CrossRef] [PubMed]
- Panir, K.; Schjenken, J.E.; Robertson, S.A.; Hull, M.L. Non-Coding RNAs in Endometriosis: A Narrative Review. Hum. Reprod. Update 2018, 24, 497–515. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bartel, D.P. MicroRNAs: Genomics, Biogenesis, Mechanism, and Function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef] [Green Version]
- Hammond, S.M. RNAi, MicroRNAs, and Human Disease. Cancer Chemother. Pharmacol. 2006, 58 (Suppl. 1), S63–S68. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Target Recognition and Regulatory Functions. Cell 2009, 136, 215–233. [Google Scholar] [CrossRef] [Green Version]
- Nematian, S.E.; Mamillapalli, R.; Kadakia, T.S.; Majidi Zolbin, M.; Moustafa, S.; Taylor, H.S. Systemic Inflammation Induced by MicroRNAs: Endometriosis-Derived Alterations in Circulating MicroRNA 125b-5p and Let-7b-5p Regulate Macrophage Cytokine Production. J. Clin. Endocrinol. Metab. 2018, 103, 64–74. [Google Scholar] [CrossRef] [Green Version]
- Anastasiu, C.V.; Moga, M.A.; Elena Neculau, A.; Bălan, A.; Scârneciu, I.; Dragomir, R.M.; Dull, A.-M.; Chicea, L.-M. Biomarkers for the Noninvasive Diagnosis of Endometriosis: State of the Art and Future Perspectives. Int. J. Mol. Sci. 2020, 21, 1750. [Google Scholar] [CrossRef] [Green Version]
- Kang, J.-W.; Eun, Y.-G.; Lee, Y.-C. Diagnostic Value of Salivary MiRNA in Head and Neck Squamous Cell Cancer: Systematic Review and Meta-Analysis. Int. J. Mol. Sci. 2021, 22, 7026. [Google Scholar] [CrossRef]
- Rapado-González, Ó.; Majem, B.; Álvarez-Castro, A.; Díaz-Peña, R.; Abalo, A.; Suárez-Cabrera, L.; Gil-Moreno, A.; Santamaría, A.; López-López, R.; Muinelo-Romay, L.; et al. A Novel Saliva-Based MiRNA Signature for Colorectal Cancer Diagnosis. J. Clin. Med. 2019, 8, 2029. [Google Scholar] [CrossRef] [Green Version]
- Setti, G.; Pezzi, M.E.; Viani, M.V.; Pertinhez, T.A.; Cassi, D.; Magnoni, C.; Bellini, P.; Musolino, A.; Vescovi, P.; Meleti, M. Salivary MicroRNA for Diagnosis of Cancer and Systemic Diseases: A Systematic Review. Int. J. Mol. Sci. 2020, 21, 907. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Patel, R.S.; Jakymiw, A.; Yao, B.; Pauley, B.A.; Carcamo, W.C.; Katz, J.; Cheng, J.Q.; Chan, E.K.L. High Resolution of MicroRNA Signatures in Human Whole Saliva. Arch. Oral Biol. 2011, 56, 1506–1513. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bossuyt, P.M.; Reitsma, J.B.; Bruns, D.E.; Gatsonis, C.A.; Glasziou, P.P.; Irwig, L.; Lijmer, J.G.; Moher, D.; Rennie, D.; de Vet, H.C.W.; et al. STARD 2015: An Updated List of Essential Items for Reporting Diagnostic Accuracy Studies. BMJ 2015, 351, h5527. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dreiseitl, S.; Ohno-Machado, L. Logistic Regression and Artificial Neural Network Classification Models: A Methodology Review. J. Biomed. Inform. 2002, 35, 352–359. [Google Scholar] [CrossRef] [Green Version]
- Crown, W.H. Potential Application of Machine Learning in Health Outcomes Research and Some Statistical Cautions. Value Health J. Int. Soc. Pharm. Outcomes Res. 2015, 18, 137–140. [Google Scholar] [CrossRef] [Green Version]
- Ghassemi, M.; Naumann, T.; Schulam, P.; Beam, A.L.; Chen, I.Y.; Ranganath, R. A Review of Challenges and Opportunities in Machine Learning for Health. AMIA 2020, 2020, 191–200. [Google Scholar]
- Sanal, M.G.; Paul, K.; Kumar, S.; Ganguly, N.K. Artificial Intelligence and Deep Learning: The Future of Medicine and Medical Practice. J. Assoc. Physicians India 2019, 67, 71–73. [Google Scholar]
- Goyal, A.; Kuchana, M.; Ayyagari, K.P.R. Machine Learning Predicts Live-Birth Occurrence before in-Vitro Fertilization Treatment. Sci. Rep. 2020, 10, 20925. [Google Scholar] [CrossRef]
- Subramanian, M.; Wojtusciszyn, A.; Favre, L.; Boughorbel, S.; Shan, J.; Letaief, K.B.; Pitteloud, N.; Chouchane, L. Precision Medicine in the Era of Artificial Intelligence: Implications in Chronic Disease Management. J. Transl. Med. 2020, 18, 472. [Google Scholar] [CrossRef]
- Lopez-Rincon, A.; Mendoza-Maldonado, L.; Martinez-Archundia, M.; Schönhuth, A.; Kraneveld, A.D.; Garssen, J.; Tonda, A. Machine Learning-Based Ensemble Recursive Feature Selection of Circulating MiRNAs for Cancer Tumor Classification. Cancers 2020, 12, 1785. [Google Scholar] [CrossRef]
- Lopez-Rincon, A.; Martinez-Archundia, M.; Martinez-Ruiz, G.U.; Schoenhuth, A.; Tonda, A. Automatic Discovery of 100-MiRNA Signature for Cancer Classification Using Ensemble Feature Selection. BMC Bioinform. 2019, 20, 480. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Metselaar, P.I.; Mendoza-Maldonado, L.; Li Yim, A.Y.F.; Abarkan, I.; Henneman, P.; Te Velde, A.A.; Schönhuth, A.; Bosch, J.A.; Kraneveld, A.D.; Lopez-Rincon, A. Recursive Ensemble Feature Selection Provides a Robust MRNA Expression Signature for Myalgic Encephalomyelitis/Chronic Fatigue Syndrome. Sci. Rep. 2021, 11, 4541. [Google Scholar] [CrossRef] [PubMed]
- Metzemaekers, J.; Haazebroek, P.; Smeets, M.J.G.H.; English, J.; Blikkendaal, M.D.; Twijnstra, A.R.H.; Adamson, G.D.; Keckstein, J.; Jansen, F.W. EQUSUM: Endometriosis QUality and Grading Instrument for SUrgical Performance: Proof of Concept Study for Automatic Digital Registration and Classification Scoring for r-ASRM, EFI and Enzian. Hum. Reprod. Open 2020, 2020, hoaa053. [Google Scholar] [CrossRef] [PubMed]
- Potla, P.; Ali, S.A.; Kapoor, M. A Bioinformatics Approach to MicroRNA-Sequencing Analysis. Osteoarthr. Cartil. Open 2021, 3, 100131. [Google Scholar] [CrossRef]
- LaRocca, D.; Barns, S.; Hicks, S.D.; Brindle, A.; Williams, J.; Uhlig, R.; Johnson, P.; Neville, C.; Middleton, F.A. Comparison of Serum and Saliva MiRNAs for Identification and Characterization of MTBI in Adult Mixed Martial Arts Fighters. PLoS ONE 2019, 14, e0207785. [Google Scholar] [CrossRef] [Green Version]
- Gyvyte, U.; Juzenas, S.; Salteniene, V.; Kupcinskas, J.; Poskiene, L.; Kucinskas, L.; Jarmalaite, S.; Stuopelyte, K.; Steponaitiene, R.; Hemmrich-Stanisak, G.; et al. MiRNA Profiling of Gastrointestinal Stromal Tumors by Next-Generation Sequencing. Oncotarget 2017, 8, 37225–37238. [Google Scholar] [CrossRef]
- Gyvyte, U.; Kupcinskas, J.; Juzenas, S.; Inciuraite, R.; Poskiene, L.; Salteniene, V.; Link, A.; Fassan, M.; Franke, A.; Kupcinskas, L.; et al. Identification of Long Intergenic Non-Coding RNAs (LincRNAs) Deregulated in Gastrointestinal Stromal Tumors (GISTs). PLoS ONE 2018, 13, e0209342. [Google Scholar] [CrossRef]
- Langmead, B.; Trapnell, C.; Pop, M.; Salzberg, S.L. Ultrafast and Memory-Efficient Alignment of Short DNA Sequences to the Human Genome. Genome Biol. 2009, 10, R25. [Google Scholar] [CrossRef] [Green Version]
- Griffiths-Jones, S.; Saini, H.K.; van Dongen, S.; Enright, A.J. MiRBase: Tools for MicroRNA Genomics. Nucleic Acids Res. 2008, 36, D154–D158. [Google Scholar] [CrossRef] [Green Version]
- Gao, L.; Zhang, L. Construction and Comprehensive Analysis of a CeRNA Network to Reveal Potential Prognostic Biomarkers for Lung Adenocarcinoma. BMC Cancer 2021, 21, 849. [Google Scholar] [CrossRef]
- Li, Q.; Liu, G.; Bao, Y.; Wu, Y.; You, Q. Evaluation and Application of Tools for the Identification of Known MicroRNAs in Plants. Appl. Plant Sci. 2021, 9, e11414. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated Estimation of Fold Change and Dispersion for RNA-Seq Data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [Green Version]
- Bargaje, R.; Hariharan, M.; Scaria, V.; Pillai, B. Consensus MiRNA Expression Profiles Derived from Interplatform Normalization of Microarray Data. RNA 2010, 16, 16–25. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nguyen, J.-M.; Jézéquel, P.; Gillois, P.; Silva, L.; Ben Azzouz, F.; Lambert-Lacroix, S.; Juin, P.; Campone, M.; Gaultier, A.; Moreau-Gaudry, A.; et al. Random Forest of Perfect Trees: Concept, Performance, Applications, and Perspectives. Bioinform. Oxf. Engl. 2021. [Google Scholar] [CrossRef] [PubMed]
- Lecointre, L.; Bund, V.; Sangnier, E.; Ouldamer, L.; Bendifallah, S.; Koskas, M.; Bolze, P.-A.; Collinet, P.; Canlorbe, G.; Touboul, C.; et al. Status of Surgical Management of Borderline Ovarian Tumors in France: Are Recommendations Being Followed? Multicentric French Study by the FRANCOGYN Group. Ann. Surg. Oncol. 2021, 28, 7616–7623. [Google Scholar] [CrossRef] [PubMed]
- Geoffron, S.; Lier, A.; de Kermadec, E.; Sermondade, N.; Varinot, J.; Thomassin-Naggara, I.; Bendifallah, S.; Daraï, E.; Chabbert-Buffet, N.; Kolanska, K. Fertility Preservation in Women with Malignant and Borderline Ovarian Tumors: Experience of the French ESGO-Certified Center and Pregnancy-Associated Cancer Network (CALG). Gynecol. Oncol. 2021, 161, 817–824. [Google Scholar] [CrossRef] [PubMed]
- Harrell, F.E.J.; Lee, K.L.; Mark, D.B. Multivariable Prognostic Models: Issues in Developing Models, Evaluating Assumptions and Adequacy, and Measuring and Reducing Errors. Stat. Med. 1996, 15, 361–387. [Google Scholar] [CrossRef]
- Steyerberg, E.W.; Eijkemans, M.J.; Harrell, F.E.J.; Habbema, J.D. Prognostic Modelling with Logistic Regression Analysis: A Comparison of Selection and Estimation Methods in Small Data Sets. Stat. Med. 2000, 19, 1059–1079. [Google Scholar] [CrossRef]
- Kern, F.; Krammes, L.; Danz, K.; Diener, C.; Kehl, T.; Küchler, O.; Fehlmann, T.; Kahraman, M.; Rheinheimer, S.; Aparicio-Puerta, E.; et al. Validation of Human MicroRNA Target Pathways Enables Evaluation of Target Prediction Tools. Nucleic Acids Res. 2021, 49, 127–144. [Google Scholar] [CrossRef]
- Wan, N.; Weinberg, D.; Liu, T.-Y.; Niehaus, K.; Ariazi, E.A.; Delubac, D.; Kannan, A.; White, B.; Bailey, M.; Bertin, M.; et al. Machine Learning Enables Detection of Early-Stage Colorectal Cancer by Whole-Genome Sequencing of Plasma Cell-Free DNA. BMC Cancer 2019, 19, 832. [Google Scholar] [CrossRef] [Green Version]
- Monnaka, V.U.; Hernandes, C.; Heller, D.; Podgaec, S. Overview of MiRNAs for the Non-Invasive Diagnosis of Endometriosis: Evidence, Challenges and Strategies. A Systematic Review. Einstein Sao Paulo Braz. 2021, 19, eRW5704. [Google Scholar] [CrossRef] [PubMed]
- Courts, C.; Madea, B. Specific Micro-RNA Signatures for the Detection of Saliva and Blood in Forensic Body-fluid Identification. J. Forensic Sci. 2011, 56, 1464–1470. [Google Scholar] [CrossRef] [PubMed]
- El-Mogy, M.; Lam, B.; Haj-Ahmad, T.A.; McGowan, S.; Yu, D.; Nosal, L.; Rghei, N.; Roberts, P.; Haj-Ahmad, Y. Diversity and Signature of Small RNA in Different Bodily Fluids Using next Generation Sequencing. BMC Genom. 2018, 19, 408. [Google Scholar] [CrossRef] [PubMed]
- Kraus, V.B.; Burnett, B.; Coindreau, J.; Cottrell, S.; Eyre, D.; Gendreau, M.; Gardiner, J.; Garnero, P.; Hardin, J.; Henrotin, Y.; et al. Application of Biomarkers in the Development of Drugs Intended for the Treatment of Osteoarthritis. Osteoarthr. Cartil. 2011, 19, 515–542. [Google Scholar] [CrossRef] [Green Version]
- Moustafa, S.; Burn, M.; Mamillapalli, R.; Nematian, S.; Flores, V.; Taylor, H.S. Accurate Diagnosis of Endometriosis Using Serum MicroRNAs. Am. J. Obstet. Gynecol. 2020, 223, 557.e1–557.e11. [Google Scholar] [CrossRef]
- Maged, A.M.; Deeb, W.S.; El Amir, A.; Zaki, S.S.; El Sawah, H.; Al Mohamady, M.; Metwally, A.A.; Katta, M.A. Diagnostic Accuracy of Serum MiR-122 and MiR-199a in Women with Endometriosis. Int. J. Gynaecol. Obstet. Off. Organ Int. Fed. Gynaecol. Obstet. 2018, 141, 14–19. [Google Scholar] [CrossRef]
- Chen, Y.; Wu, Y.; Yao, H.; Luo, H.; Lin, B.; Zhang, X.; Liang, X.; Sun, R.; Zhao, S.; Li, Y.; et al. MiRNA Expression Profile of Saliva in Subjects of Yang Deficiency Constitution and Yin Deficiency Constitution. Cell. Physiol. Biochem. 2018, 49, 2088–2098. [Google Scholar] [CrossRef]
| Control Patients | Patients with Endometriosis | ||
|---|---|---|---|
| N (%) | N (%) | ||
| N = 47 | N = 153 | ||
| Age (years) (mean ± SD) | 30.92 (13.79) | 31.17 (10.78) | 0.1912 |
| Age range | |||
| 72% (34) | 63% (96) | |
| 28% (13) | 37% (57) | 0.294 |
| BMI (body mass index) (mean ± SD) | 24.84 (11.10) | 24.36 (8.38) | 0.525 |
| Infertility | |||
| 17% (8) | 24% (36) | |
| 83% (39) | 76% (117) | 0.556 |
| rASRM classification | - | ||
| - | 52% (80) | |
| - | 48% (73) | |
| Control diagnoses | |||
| 51% (24) | _ | - |
| 2% (1) | ||
| 11% (5) | ||
| 23% (11) | ||
| 13% (6) | ||
| Dysmenorrhea | 100% | 100% | |
| Abdominal pain outside menstruation | |||
| 66% (21) | 71% (89) | 0.6905 |
| Pain suggesting sciatica | |||
| 31% (10) | 56% (70) | 0.0214 |
| Lower back pain outside menstruation | |||
| 62% (20) | 81% (101) | 0.0498 |
| Right shoulder pain during menstruation | |||
| 9% (3) | 21% (26) | 0.2184 |
| Blood in the stools during menstruation | |||
| 12% (4) | 24% (30) | 0.2425 |
| Blood in urine during menstruation | |||
| 25% (8) | 17% (21) | 0.4172 |
| Diagnostic method | |||
| 47 (100) | 83 (54.2) | |
| - | 70 (45.8) | - |
| Random Forest | |||
|---|---|---|---|
| Dataset | AUC | Sensitivity | Specificity |
| 1 | 0.935 | 0.871 | 1 |
| 2 | 0.903 | 0.806 | 1 |
| 3 | 0.935 | 0.871 | 1 |
| 4 | 0.983 | 0.967 | 1 |
| 5 | 0.867 | 0.833 | 0.9 |
| 6 | 0.968 | 0.935 | 1 |
| 7 | 0.919 | 0.839 | 1 |
| 8 | 0.935 | 0.871 | 1 |
| 9 | 0.933 | 0.967 | 0.9 |
| 10 | 0.9 | 0.8 | 1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bendifallah, S.; Suisse, S.; Puchar, A.; Delbos, L.; Poilblanc, M.; Descamps, P.; Golfier, F.; Jornea, L.; Bouteiller, D.; Touboul, C.; et al. Salivary MicroRNA Signature for Diagnosis of Endometriosis. J. Clin. Med. 2022, 11, 612. https://doi.org/10.3390/jcm11030612
Bendifallah S, Suisse S, Puchar A, Delbos L, Poilblanc M, Descamps P, Golfier F, Jornea L, Bouteiller D, Touboul C, et al. Salivary MicroRNA Signature for Diagnosis of Endometriosis. Journal of Clinical Medicine. 2022; 11(3):612. https://doi.org/10.3390/jcm11030612
Chicago/Turabian StyleBendifallah, Sofiane, Stéphane Suisse, Anne Puchar, Léa Delbos, Mathieu Poilblanc, Philippe Descamps, Francois Golfier, Ludmila Jornea, Delphine Bouteiller, Cyril Touboul, and et al. 2022. "Salivary MicroRNA Signature for Diagnosis of Endometriosis" Journal of Clinical Medicine 11, no. 3: 612. https://doi.org/10.3390/jcm11030612
APA StyleBendifallah, S., Suisse, S., Puchar, A., Delbos, L., Poilblanc, M., Descamps, P., Golfier, F., Jornea, L., Bouteiller, D., Touboul, C., Dabi, Y., & Daraï, E. (2022). Salivary MicroRNA Signature for Diagnosis of Endometriosis. Journal of Clinical Medicine, 11(3), 612. https://doi.org/10.3390/jcm11030612







